Abstract
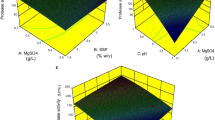
Among various strains of Bacillus subtilis, the strain RNZ-79 was selected for this study due to the highest uricase productivity in solid state cultures containing shrimp shell wastes, highest Vmax (0.42 μM mg−1min−1) and lowest value of Km (56 μM). Maximum productivity was observed at pH 7.6 and 45°C with an inducer concentration of 0.4% (w/v) uric acid and moisture content of 80% (v/w). An inoculum of 7% (v/v) and particle size of 354–500 μm for substrate were optimal for productivity after 54 h of fermentation. None of the tested carbon and inorganic nitrogen sources had stimulatory effect on uricase productivity. KH2PO4 at 0.12% (w/v) was the best source of phosphorus. Final uricase productivity was 5.05-fold more than the initial productivity. Enzyme purification increased the specific activity to 25-fold with a recovery of 36%. Furthermore, the purified enzyme showed a molecular mass of 33.7 kDa. Purified uricase was optimally active at pH 8.0 and 40°C and maximally stable at pH 7.0–10.0 and till 70°C for 30 min. The half-life (t1/2) at 60, 70, 80, 90 and 100°C were 87.0, 56.6, 30.0, 24.2 and 15.6 min, respectively, and the calculated midpoint temperature (Tm) was 66.85°C. Interestingly, purified enzyme exhibited a good storage stability for 3 months and none of uric acid analogues were competitive inhibitor, indicating a high specificity of uricase.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Abbreviations
- CB:
-
corn bran
- LSCs:
-
liquid state cultures
- LSF:
-
liquid state fermentation
- RB:
-
rice bran
- SBP:
-
soybean powder
- SSCs:
-
solid state cultures
- SSF:
-
solid state fermentation
- SSP:
-
shrimp shell powder
- WB:
-
wheat bran
References
Abd El Fattah Y., Saeed H., Gohar Y. & El-Baz M. 2005. Improved production of Pseudomonas aeruginosa uricase by optimization of process parameters through statistical experimental designs. Process Biochem. 40: 1707–1714.
Abdel-Fattah G. & Abo-Hamed N. 2002. Bioconversion of poultry wastes I-factors influencing the assay and productivity of crude uricase by three uricolytic filamentous fungi. New Microbiol. 25: 57–64.
Adámek V., Králová B., Suchová M., Valentová O. & Demnerová K. 1989. Purification of microbial uricase. J. Chromatogr. 497: 268–275.
Adámek V., Suchová M., Demnerová K., Králová B., Fořt I. & Morava P. 1990. Fermentation of Candida utilis for uricase production. J. Ind. Microbiol. 6: 85–90.
Alamillo J., Cardenas J. & Pineda M. 1991. Purification and molecular properties of urate oxidase from Chlamydomonas reinhardtii. Biochim. Biophys. Acta 1076: 20–208.
Anderson A. & Vijayakumar S. 2011. Purification and optimization of uricase enzyme produced by Pseudomonas aeruginosa. J. Exp. Sci. 2: 5–8.
Bailey W., Scott E., Finegold S. & Baron E. 1986. Bailey and Scott’s Diagnostic Microbiology. Mosby Co., USA.
Baysal Z., Uyar F. & Aytekin C. 2003. Solid state fermentation for production of α-amylase by a thermotolerant Bacillus subtilis from hot-spring water. Process Biochem. 38: 1665–1668.
Bongaerts G., Uitzetter J., Brouns R. & Vogels G. 1978. Uricase of Bacillus fastidiosus. Properties and regulation of synthesis. Biochim. Biophys. Acta 527: 348–358.
Brandenburg J., Wray L., Beier L., Jarmer H., Saxild H. & Fisher S. 2002. Roles of PucR, GInR, and TnrA in regulating expression of the Bacillus subtilis ure P3 promoter. J. Bacteriol. 84: 6060–6064.
Hassanein W., Kotb E., Awny N. & El-Zawahry Y. 2011. Fibrinolysis and anticoagulant potential of a metallo protease produced by Bacillus subtilis K42. J. Biosci. 36: 77–779.
Hibi T., Hayashi Y., Fukada H., Itoh T., Nago T. & Nishiya Y. 2014. Intersubunit salt bridges with a sulfate anion control subunit dissociation and thermal stabilization of Bacillus sp. TB-90 urate oxidase. Biochemistry 53: 3879–3888.
Huang S. & Wu T. 2004. Modified colorimetric assay for uricase activity and a screen for mutant Bacillus subtilis uricase genes following StEP mutagenesis. Eur. J. Biochem. 271: 517–523.
Ishikawa J., Yamashita A., Mikami Y., Hoshino Y., Kurita H., Hotta K., Shiba T. & Hattori M. 2004. The complete genomic sequence of Nocardia farcinica IFM 10152. Proc. Natl. Acad. Sci. USA 101: 14925–14930.
Kai L., Ma X., Zhou X., Jia X., Li X. & Guo K. 2008. Purification and characterization of a thermostable uricase from Microbacterium sp. strain ZZJ4-1. World J. Microbiol. Biotechnol. 24: 401–406.
Kotb E. 2015a. Characterization of a thermostable uricase isolated from Bacillus firmus DWD-33 and its application for uric acid quantification in human serum. Protein Pept. Lett. 22: 402–409.
Kotb E. 2015b. The biotechnological potential of subtilisin-like fibrinolytic enzyme from a newly isolated Lactobacillus plantarum KSK-II in blood destaining and antimicrobials. Biotechnol. Prog. 31: 316–324.
Koyama Y., Ichikawa T. & Nakano E. 1996. Cloning, sequence analysis, and expression in Escherichia coli of the gene encoding the Candida utilis urate oxidase (uricase). J. Biochem. 120: 969–973.
Krishna C. & Chandrasekaran M. 1996. Banana waste as substrate for a-amylase production by Bacillus subtilis (CBTK 106) under solid-state fermentation. Appl. Microbiol. Biotechnol. 46: 106–111.
Laemmli U. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Liu J., Li G., Liu H. & Zhou X. 1994. Purification and properties of uricase from Candida sp. and its application in uric acid analysis in serum. Appl. Biochem. Biotechnol. 47: 57–63.
Lonsane B., Saucedo-Castaneda G., Raimbault M., Roussos S., Viniegra-Gonzalez G., Ghildyal N., Ramakrishna M. & Krishnaiah M. 1992. Scale-up strategies for solid state fermentation systems. Process Biochem. 27: 259–273.
Lotfy W. 2008. Production of a thermostable uricase by a novel Bacillus thermocatenulatus strain. Bioresour. Technol. 99: 699–702.
Lowry O., Rosebrough N., Farr A. & Randall R. 1951. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193: 265–275.
Montalbini P., Aguilar M. & Ineda M. 1999. Isolation and characterization of uricase from bean leaves and its comparison with uredospore enzyme. Plant Sci. 147: 139–147.
Nakagawa T., Mazzali M., Kang D., Sanchez-Lozada L., Herrera-Acosta J. & Johnson R. 2006. Uric acid - a uremic toxin? Blood Purif. 24: 67–70.
Nanda P. & Babu P. 2014. Isolation, screening and production studies of uricase producing bacteria from poultry sources. Prep. Biochem. Biotechnol. 44: 811–821.
Pandey A., Selvakumar P., Soccol C. & Nigam P. 1999. Solid state fermentation for the production of industrial enzymes. Curr. Sci. 77: 149–162.
Pandey A., Soccol C. & Larroche C. 2007. Current Developments in Solid-State Fermentation. Springer Science/Asiatech Publishers, New York, USA/New Delhi, India.
Pfrimer P., de Moraes L., Galdino A., Salles L., Reis V., De Marco J., Prates M., Bloch C. & Torres F. 2010. Cloning, purification, and partial characterization of Bacillus subtilis urate oxidase expressed in Escherichia coli. Biomed. Biotechnol. 2010: 1–6.
Saeed H., Abdel-Fattah Y., Gohar Y. & Elbaz M. 2004. Purification and characterization of extracellular Pseudomonas aeruginosa urate oxidase enzyme. Pol. J. Microb. 53: 45–52.
Schiavon O., Caliceti P., Ferruti P. & Veronese F. 2000. Therapeutic proteins: a comparison of chemical and biological properties of uricase conjugated to linear or branched poly (ethylene glycol) and poly(N-acryloylmorpholine). Farmaco 55: 264–269.
Schultz C., Nygaard P. & Saxild H. 2001. Functional analysis of 14 genes that constitute the purine catabolic pathway in Bacillus subtilis and evidence for a novel regulon controlled by the PucR transcription activator. J. Bacteriol. 183: 329–3302.
Suzuki K., Sakasegawa, S., Misaki H. & Sugiyamac M. 2004. Molecular cloning and expression of uricase gene from Arthrobacter globiformis in Escherichia coli and characterization of the gene product. J. Biosci. Bioeng. 98: 15–158.
Swan D., Hale R., Dhillon N. & Leadlay P. 1987. A bacterial calcium-binding protein homologous to calmodulin. Nature 329: 84–85.
Tanaka A., Yamamura M., Kawamoto S. & Fukui S. 1977. Production of uricase by Candida tropicalis using n-alkane as substrate. Appl. Environ. Microbiol. 34: 342–346.
Wallrath L. & Friedman T. 1991. Species differences in the temporal pattern of Drosophila urate oxidase gene expression are attributed to transacting regulatory changes. Proc. Natl. Acad. Sci. USA 88: 5489–5493.
Werner A. & Witte C. 2011. The biochemistry of nitrogen mobilization: purine ring catabolism. Trends Plant Sci. 16: 381–387.
Yamamoto K., Kojima Y., Kikuchi T., Shigyo T., Sugihara K., Takashio M. & Emi S. 1996. Nucleotide sequence of the uri-case gene from Bacillus sp. TB-90. J. Biochem. 119: 80–84.
Yazdi M., Zarrini G., Mohit E., Faramarzi M., Setayesh N., Sedighi N. & Mohseni F. 2006. Mucor hiemalis: a new source for uricase production. World J. Microbiol. Biotechnol. 22: 325–330.
Yokoyama S., Ogawa A. & Obayashi A. 1988. Rapid extraction of uricase from Candida utilis cells by use of reducing agent plus surfactant. Enzyme Microb. Technol. 10: 52–55.
Zhao Y., Zhao L., Yang G., Tao J., Bu Y. & Liao F. 2006. Characterization of a uricase from Bacillus fastidious A.T.C.C. 26904 and its application to serum uric acid assay by a patented kinetic uricase method. Biotechnol. Appl. Biochem. 45: 75–78.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kotb, E. Improvement of uricase production from Bacillus subtilis RNZ-79 by solid state fermentation of shrimp shell wastes. Biologia 71, 229–238 (2016). https://doi.org/10.1515/biolog-2016-0040
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1515/biolog-2016-0040