Abstract
Background
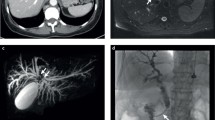
Cholangiocarcinoma can be classified in intrahepatic cholangiocarcinoma (ICC) and perihilar cholangiocarcinoma (PCC). Moreover, PCC includes two different forms: extrahepatic (EH) PCC, which arises from the perihilar EH large ducts, and intrahepatic (IH) PCC, in which a significant liver mass invades the perihilar bile ducts. In this study, we investigated the molecular profile and molecular prognostic factors in EH-PCC, IH-PCC, and ICC submitted to curative surgery.
Methods
Ninety-one patients with cholangiocarcinoma (38 EH-PCC, 18 IH-PCC, and 35 ICC), who underwent curative surgery in a single tertiary hepatobiliary surgery referral center were assessed for mutational status in 56 cancer-related genes.
Results
The most frequently mutated genes in EH-PCC were KRAS (47.4 %), TP53 (23.7 %) and ARID1A (15.8 %); in IH-PCC were KRAS (22.2 %), PBRM1 (16.7 %), and PIK3CA (16.7 %); and in ICC were IDH1 (17.1 %), NRAS (17.1 %), and BAP1 (14.3 %). The presence of mutations in ALK, IDH1, and TP53 genes was significantly associated with poor prognosis in patients with EH-PCC (p < 0.001, p = 0.043, and p = 0.019, respectively). Mutation of the TP53 gene was significantly associated with poor prognosis in patients with IH-PCC (p = 0.049). The presence of mutations in ARID1A, PIK3C2G, STK11, TGFBR2, and TP53 genes was significantly associated with poor prognosis in patients with ICC (p = 0.012, p = 0.030, p = 0.030, p = 0.011, and p = 0.011, respectively).
Conclusions
Mutational gene profiling identified different gene mutations in EH-PCC, IH-PCC, and ICC. Moreover, our study reported specific prognostic genes that can identify patients with poor prognosis after curative surgery who may benefit from traditional or target adjuvant treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Cholangiocarcinoma (CCA) is a heterogeneous group of malignancies arising from the epithelial cells of biliary tree with a poor prognosis.1,2 CCA can be classified into three different forms: intrahepatic cholangiocarcinoma (ICC), arising from intrahepatic (IH) bile ducts; PCC, arising or involving the hepatic biliary confluence; and distal cholangiocarcinoma, arising from the bile duct distal to the cystic duct origin.3,4 PCC includes two separate subtypes: EH-PCC, which arises from the perihilar extrahepatic (EH) large ducts, and IH PCC, in which a significant liver mass invades the perihilar bile ducts.5,6 In some cases the clinical discrimination between ICC and IH-PCC may be difficult.7,8
The pathogenic pathways involved in carcinogenesis of ICC and PCC are still unclear. Recently, several studies investigated the molecular mutations that characterize CCAs including PIK3CA, PTEN, AKT1, IDH1, and IDH2. 9–13 However, the prevalence of these alterations varies widely among studies. Two recent whole-exome sequencing studies of ICC revealed a key role for chromatin remodeling genes BAP1, ARID1A, and PBRM1 in the development of these neoplasms.9,14 Other study reported that IDH1 mutations occurred more frequently in ICC and ERBB2 in EH CCA.15 It has also been shown that specific molecular mutations are associated with different types of biliary tree carcinomas, supporting their pathologic and molecular heterogeneity.16 To our knowledge, no studies have investigated the differences in molecular profiling between EH-PCC and IH-PCC.
The prognostic implication of the mutational profiling of CCA after surgical resection is still under investigation.
The aims of the present study were to identify and compare the mutation profiling of EH-PCC, IH-PCC, and ICC and to identify their molecular prognostic factors in patients who underwent surgery with curative intent.
Patients and Methods
Definition of CCA Subtypes
CCAs were classified according to World Health Organization 2010 and American Joint Committee on Cancer (AJCC)/International Union Against Cancer (UICC) 7th edition criteria as ICC and PCC.3,4 PCC were defined as tumors that involve the biliary confluence, even in the presence of a liver parenchymal mass. Moreover, in this study we applied the 3rd English edition of the Japanese classification of biliary tract cancers; this classification defines as PCC every tumor that involves the perihilar bile duct (between the right side of the umbilical portion of the left portal branch and the left side of the origin of the right posterior portal branch). Tumors with a predominant liver mass component were included in the perihilar group if the center of the mass was located between the above portal landmarks.8
PCC was divided according to the macroscopic aspect of the tumors: disease with a predominant liver mass component was defined as IH-PCC, and disease without a significant liver mass was defined as EH-PCC.6 When identification of the tumor location was difficult, the presence of carcinoma-in-situ and the most extensive cancer infiltration with ductal stricture were used for reference, and a multidisciplinary team discussion including surgeons (A.G. and A.R.), pathologists (A.S., C.P., and M.F.), and a radiologist (M.D.O.) was carried out to reach a consensus.
Patients
From September 1990 to December 2012, a total of 146 patients with CCA submitted to surgical resection with radical intent in a single tertiary hepatobiliary surgery referral center. In 91 CCA specimens, the material was sufficient for the pathologic and molecular analyses and were retrieved from the formalin-fixed, paraffin-embedded (FFPE) archives of the Department of Pathology–Diagnostics and the Arc-Net biobank of the University and Hospital Trust of Verona.
The clinical and pathologic data were prospectively collected in all patients. For the 91 CCAs, tissue microarrays were also prepared using two 1 mm cores for each case. FFPE tissue was obtained under local ethics committee ARC-Net approval (prog. 1959).
DNA Extraction and PCR Amplification
DNA was prepared from tissues after enrichment for neoplastic cellularity using manual microdissection. A total of 5–15 consecutive 4 µm FFPE sections per case were used. DNA was extracted using the QIAamp DNA FFPE Tissue Kit (Qiagen, Germantown, MD). Purified DNA was quantified and its quality assessed using NanoDrop (Invitrogen; Life Technologies, Carlsbad, CA, USA) and Qubit (Invitrogen) platforms.17 DNA quality was further evaluated by PCR analysis using the BIOMED 2 PCR multiplex protocol with PCR products analyzed by DNA 1000 Assay (Invitrogen) on the Agilent 2100 Bioanalyzer on-chip electrophoresis (Agilent Technologies, Santa Clara, CA).18
Two multigene panels were used: the 50-gene Ion AmpliSeq Cancer Hotspot Panel v2 (Life Technologies) and a 7-gene AmpliSeq Custom Panel. The first explores selected regions of the following 50 cancer-associated genes, in alphabetical order: ABL1, AKT1, ALK, APC, ATM, BRAF, CDH1, CDKN2A, CSF1R, CTNNB1, EGFR, ERBB2, ERBB4, EZH2, FBXW7, FGFR1, FGFR2, FGFR3, FLT3, GNA11, GNAS, GNAQ, HNF1A, HRAS, IDH1, IDH2, JAK2, JAK3, KDR/VEGFR2, KIT, KRAS, MET, MLH1, MPL, NOTCH1, NPM1, NRAS, PDGFRA, PIK3CA, PTEN, PTPN11, RB1, RET, SMAD4, SMARCB1, SMO, SRC, STK11, TP53, and VHL. The details of the target regions may be found online (http://www.lifetechnologies.com). The 7-gene custom panel was designed to target selected regions of a gene included in the 50-gene panel (IDH2), and 6 genes were selected according to the results of a previously published ICC exome sequencing study (ARID1A, BAP1, PBRM1, PIK3C2A, PIK3C2G, TGFBR2).9
Sequencing was run on the Ion Torrent Personal Genome Machine (PGM; Life Technologies) loaded with 316 (50-gene panel) or 318 (7-gene panel) chips.
Statistical Analysis
Data were collected and analyzed by SPSS 21 statistical software (IBM, Armonk, NY). The differences between categorical variables were analyzed by the χ 2 test. Comparisons between means were carried out by t test. Survival analysis was carried out using the Kaplan–Meier method. We considered the treatment day as time 0, and patients alive at the end of follow-up were considered censored. The mean follow-up period was 28.3 ± 25.8 months. Nine patients with 90-day postoperative mortality (7 EH-PCC, 1 IH-PCC, 1 ICC) were excluded from survival analysis.
A multivariate analysis including the clinical, pathologic, and molecular factors related to survival at univariate analysis (p < 0.05) were carried out with the Cox regression model with forward and backward analysis to identify factors that were independently related to survival. p < 0.05 was regarded as statistically significant.
Results
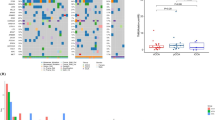
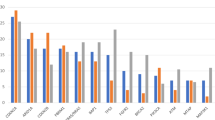
The clinical and pathologic features of the 91 patients included in the study are summarized in Table 1. The study population included 38 EH-PCC, 18 IH-PCC, and 35 ICC. A detailed description of gene mutations in EH-PCC, IH-PCC, and ICC is provided in Table 2. No mutations were identified in the following 24 genes: ABL1, AKT1, APC, ATM, CDH1, CSF1R, ERBB2, FGFR1, FGFR2, FLT3, HNF1A, JAK2, JAK3, MET, MPL, NOTCH1, NPM1, PDGFRA, RB1, RET, SMARCB1, SMO, SRC, and VHL. At least 1 gene mutation was identified in 76.9 % of the patients (n = 70).
The most frequently mutated gene in EH-PCC was KRAS, with mutation in 47.4 % of the patients. Other genes with a high rate of mutations (over 10 %) in EH-PCC were TP53 (23.7 %), ARID1A (15.8 %), and PIK3C2G (10.5 %).
In IH-PCC, the most frequently mutated gene was also KRAS, with mutation in 22.2 % of the patients. Other genes with a rate of mutations over 10 % in IH-PCC were PBRM1 (16.7 %), PIK3CA (16.7 %), ARID1A (11.1 %), PIK3C2A (11.1 %), and TP53 (11.1 %).
The most frequently mutated genes in ICC were IDH1 and NRAS, with mutation in 17.1 % % of the patients. Other genes with a high rate of mutations (over 10 %) in ICC were BAP1 (14.3 %), ARID1A (11.4 %), and PBRM1 (11.4 %).
Comparison in Molecular Profile among IH-PCC, EH-PCC, and ICC
The results of the univariate analysis of comparison in molecular profile among the 3 groups are shown in Table 2.
Comparing EH-PCC with IH-PCC, we observed a statistically significant higher frequency of mutation for KRAS, 47.4 and 22.2 %, respectively (p = 0.044). Considering the genes with a mutation rate of over 10 % in both groups, the frequency of gene mutation for ARID1A and TP53 was not statistically different between EH-PCC and IH-PCC. No mutations of ERBB4, FGFR3, and NRAS were identified in both EH-PCC and IH-PCC.
Comparing IH-PCC with ICC, no differences in gene mutation reach the statistical significance. Nevertheless, a higher frequency of mutation in ICC occurred in BAP1 and NRAS (14.3 vs. 0 %, p = 0.092, and 17.1 vs. 0 %, p = 0.062, respectively). Conversely, PIK3CA was more frequently mutated in IH-PCC compared to ICC (16.7 vs. 2.8 %, p = 0.071). Considering the genes with a mutation rate of over 10 % in both groups, the frequency of gene mutation for ARID1A and PBRM1 were not statistically different between IH-PCC and ICC. No mutations in EGFR, HRAS, KIT, MLH1, and SMAD were identified in both IH-PCC and ICC.
Higher frequencies of mutations were observed in ICC compared to EH-PCC for IDH1 (17.1 vs. 2.6 %, p = 0.035) and NRAS (17.1 vs. 0 %, p = 0.008). Conversely, KRAS and TP53 were more commonly mutated in EH-PCC compared to ICC (47.4 vs. 8.6 %, p < 0.001; and 23.7 vs. 5.7 %, p = 0.032, respectively). Considering the genes with mutation rate over 10 % in both groups, the frequency of gene mutation for ARID1A was not statistically different between EH-PCC and ICC. No mutations of CDKN2A, CTNNB1, GNAS, and PTPN11 were identified in EH-PCC and ICC.
Subgroup analysis limited to advanced stages of disease (AJCC/UICC stages III and IVa) confirmed that each tumor subtype (ICC, EH-PCC, and IH-PCC) had a different frequency of gene mutations (Supplementary Table 1).
Prognostic Factors After Surgery
Figure 1 shows overall survival after surgery for EH-PCC, IH-PCC, and ICC.
Survival curves after curative resection for extrahepatic perihilar cholangiocarcinoma (EH-PCC), intrahepatic perihilar cholangiocarcinoma (IH-PCC), and intrahepatic cholangiocarcinoma (ICC). IH-PCC had similar overall survival after surgical resection compared to EH-PCC, with median survival of 22.6 months (3-year overall survival rate 41.2 %) and 30.5 months (3 years overall survival rate 45.9 %), respectively (p = 0.435), whereas survival for ICC was significantly longer, with median survival of 52.0 months (3-year overall survival rate 57.6 %) (p = 0.023)
Univariate analysis of clinical, pathologic, and molecular prognostic factors of EH-PCC, IH-PCC, and ICC is shown in Table 3. The presence of mutations in ALK, IDH1, and TP53 genes was significantly associated with poor prognosis in patients with EH-PCC compared to wild type (median overall survival 5.0 vs. 34.9 months, p < 0.001, 9.1 vs. 29.6 months, p = 0.043; and 15.4 vs. 32.5 months, p = 0.019, respectively).
Mutation of TP53 was significantly associated with poor prognosis in patients with IH-PCC compared to wild type, with a median overall survival of 6.1 vs. 22.6 months (p = 0.049). IH-PCC patients with mutations of BRAF showed a reduced overall survival compared to wild-type patients (median overall survival of 10.2 vs. 22.6 months, p = 0.065), but the difference did not reach statistical significance.
The presence of mutations of ARID1A, PIK3C2G, STK11, TGFBR2, and TP53 genes was significantly associated with poor prognosis in patients with ICC compared to wild type (median overall survival of 14.0 vs. 52.0 months, p = 0.012; 11.8 vs. 40.1 months, p = 0.030; 11.8 vs. 40.1 months, p = 0.030; 9.3 vs. 40.1 months, p = 0.011; and 5.7 vs. 40.1 months, p = 0.011, respectively).
We performed a subgroup analysis comparing survival of patients with any mutation in specific prognostic genes identified at univariate analysis for each subtype (EH-PCC, IH-PCC, and ICC) in patients without mutations (wild type); in EH-PCC, patients with any mutation in ALK, IDH1, and TP53 showed a poorer prognosis, with a 3-year survival rate of 11.1 % (median survival 11.1 months) compared to 51.3 % of wild-type patients (median survival 40.5 months) (p = 0.001) (Fig. 2a).
a Subgroup analysis in extrahepatic perihilar cholangiocarcinoma (EH-PCC) comparing survival of patients with any mutation in ALK, IDH1, and TP53 (prognostic genes at univariate analysis, p < 0.001, p = 0.043, and p = 0.019, respectively) with patients without mutations (wild type); patients with mutations showed poorer prognosis, with 3-year survival rate of 11.1 % (median survival 11.1 months) compared to 51.3 % of wild-type patients (median survival 40.5 months, p = 0.001). b Subgroup analysis in intrahepatic perihilar cholangiocarcinoma (IH-PCC) comparing survival of patients with any mutation in TP53 and BRAF (prognostic genes at univariate analysis, p = 0.049 and p = 0.065, respectively) with patients without mutations (wild type); patients with mutations in these genes showed poorer prognosis, with no survivors after 3 years (median survival 10.2 months) compared to 30.8 % of wild-type patients surviving after 3 years (median survival 23.8 months, p = 0.007). c Subgroup analysis of overall survival in intrahepatic cholangiocarcinoma (ICC) comparing patients with any mutation in ARID1A, PIK3C2A, STK11, TGFBR2, and TP53 (prognostic genes at univariate analysis, p = 0.012, p = 0.011, p = 0.011, p = 0.030, and p = 0.030, respectively) with patients without any mutation (wild type); patients with mutations showed poor prognosis with no survival at 3 years (median survival 11.8 months) compared to 66.0 % survival at 3 years in wild-type patients (median survival 62.7 months, p < 0.001)
IH-PCC patients with mutation in TP53 and/or BRAF showed a poorer prognosis, with no survivors after 3 years (median survival 10.2 months) compared to 30.8 % of wild-type patients surviving more than 3 years (median survival 23.8 months) (p = 0.007) (Fig. 2b).
ICC patients with cancers harboring mutations in ARID1A, PIK3C2A, STK11, TGFBR2, and TP53 showed a poorer prognosis, with no survivors after 3 years (median survival 11.8 months) compared to 66.0 % of wild-type patients surviving more than 3 years (median survival 62.7 months) (p < 0.001) (Fig. 2c).
At multivariate analysis including clinical, pathologic, and genetic features, the factors independently related with survival for EH-PCC were as follows: R1 resection (odds ratio [OR] 2.699, 95 % confidence interval [CI] 0.999–7.293, p = 0.050), pN status (OR 2.883, 95 % CI 1.140–7.287, p = 0.025), and mutations in IDH1 (OR 17.844, 95 % CI 3.947–17.397, p = 0.004) and TP53 (OR 2.706, 95 % CI 1.092–8.210, p = 0.039). The independent prognostic factors for IH-PCC were as follows: pN status (OR 3.223, 95 % CI 1.090–12.803, p = 0.027) and mutations in the TP53 gene (OR 3.110, 95 % CI 1.067–9.065, p = 0.038). In the ICC group, R1 resection (OR 2.845, 95 % CI 1.019–8.805, p = 0.040), pN status (OR 2.038, 1.015–8.299, p = 0.033), and mutations in the ARID1A gene (OR 5.337, 95 % CI 1.325–21.489, p = 0.018) and the TP53 gene (OR 10.803, 95 % CI 2.022–57.727, p = 0.005) were independent factors related to survival (Supplementary Table 2).
Discussion
In this study, we analyzed the molecular features of EH-PCC, IH-PCC, and ICC in a patient series from a single tertiary hepatobiliary referral center. The main findings of our study showed specific molecular characteristics and distinctive molecular prognostic factors for EH-PCC, IH-PCC, and ICC.
From our data, the macroscopic type of CCA, EH-PCC, IH-PCC, and ICC seem to have significant differences at the molecular level, not only in prognosis and type of treatment, suggesting different carcinogenetic pathways.
Data are available on the molecular profiling of CCA, mostly regarding ICC, whereas data for PCC are from limited series.19
Previously, several studies reported IDH1 mutations in about 20 % of ICC.15,20 Recently, Zhu et al. analyzed 15 cancer-related genes in 200 resected specimens of ICC from seven different centers and reported mutation of IDH1 in 15.5 % and of KRAS in 8.6 %, but in this study there was no clear definition of ICC (more than 22 % of patients with biliary confluence invasion). In contrast, NRAS mutations were reported in only 3.1 % of ICC.21 In addition, using exome sequencing, a multicenter study on 32 ICC identified mutations of BAP1, ARID1A, PBRM1, and IDH1/2 in 25, 19, 17, and 19 %, respectively.9
Mutational data on PCC are fewer and unclear, and to our knowledge, no previous study has reported specific molecular profile for EH-PCC and IH-PCC. For example, a study on 34 CCAs reported mutation of KRAS in 38.2 % and of PIK3CA in 32.4 %, but the type of CCA, perihilar or IH, was not specified.22 A separate study on 27 PCC reported mutations of KRAS and TP53 in 40.7 % and 22.2 % of patients, respectively.19 Park et al. in a study on 104 CCA showed mutation of TP53 in 47.4 % of 41 hilar CCA.23
Moreover, we identified disease-specific molecular prognostic factors that differ between EH-PCC, IH-PCC, and ICC. Specifically, ALK and IDH1 mutation had an exclusive prognostic impact for EH-PCC, BRAF for IH-PCC, and ARID1A, PIK3C2G, STK11, and TGFBR2 for ICC. However, mutation of TP53 is a negative prognostic factor in all 3 groups.
A small number of studies described molecular prognostic factors for PCC. According to a meta-analysis, TP53 appears to be an important prognostic factor for overall survival of patients with EH CCA.24 More recently, Park et al. reported no impact on survival of TP53 mutations in PCC.23
With only a few studies investigating ICC, a clear prognostic role of molecular profiling in ICC has not yet been established.15,21,25,26 Robertson et al. reported worse survival of ICC patients with KRAS mutations compared to wild type.25 Wang et al. reported better overall and disease-free survival in ICC patients with IDH1/2 mutations.26 In contrast, a recent study showed that ICC patients with IDH1/2 mutations had a 3-year survival of 33 % compared to 81 % for wild type (p = 0.003).9 Zhu et al. showed a relationship between mutations in IDH1/2, KRAS, NRAS, and BRAF and clinicopathologic features of ICC, but there was no association between the presence of mutated genes and survival.21 Churi et al. reported a poorer prognosis of ICC patients with KRAS and TP53 mutations compared to wild type.15
A limitation of the current study is the small sample size, although data in the literature on the molecular profiling of CCA are frequently multi-institutional and limited to a small number of patients. Moreover, statistical analysis on differences between subgroups and survival analysis could be suboptimal as a result of the low frequency rate of some gene mutations. Furthermore, our series included many patients with advanced disease stage (AJCC/UICC stage III or IV), particularly for EH-PCC and IH-PCC patients.
External validation and further study are needed to confirm our results.
Conclusion
Mutational gene profiling identified different gene mutations in EH-PCC, IH-PCC, and ICC. Moreover, our study reported specific prognostic genes for EH-PCC, IH-PCC, and ICC that can identify patients with poor prognosis after curative surgery who may benefit from adjuvant treatments. The disease-specific genes we identified can be explored for new molecular therapies in clinical trials.
References
de Groen PC, Gores GJ, LaRusso NF, Gunderson LL, Nagorney DM. Biliary tract cancers. N Engl J Med. 1999;341:1368–78.
Patel T. Cholangiocarcinoma—controversies and challenges. Nat Rev Gastroenterol Hepatol. 2011;8:189–200.
International Union Against Cancer (UICC). TNM classification of malignant tumours. 7th ed. Chichester: Wiley-Blackwell; 2010.
American Joint Committee on Cancer (AJCC). Cancer staging manual. 7th ed. New York: Springer; 2010.
Sano T, Shimada K, Sakamoto Y, Ojima H, Esaki M, Kosuge T. Prognosis of perihilar cholangiocarcinoma: hilar bile duct cancer versus intrahepatic cholangiocarcinoma involving the hepatic hilus. Ann Surg Oncol. 2008;15:590–9.
Ebata T, Kamiya J, Nishio H, Nagasaka T, Nimura Y, Nagino M. The concept of perihilar cholangiocarcinoma is valid. Br J Surg. 2009;96:926–34.
Nakeeb A, Pitt HA, Sohn TA, et al. Cholangiocarcinoma. A spectrum of intrahepatic, perihilar, and distal tumors. Ann Surg. 1996;224:463–73.
Miyazaki M, Ohtsuka M, Miyakawa S, et al. Classification of biliary tract cancers established by the Japanese Society of Hepato-Biliary-Pancreatic Surgery: 3rd English edition. J Hepatobiliary Pancreat Sci. 2015;22:181–96.
Jiao Y, Pawlik TM, Anders RA, et al. Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nat Genet. 2013;45:1470–3.
Voss JS, Holtegaard LM, Kerr SE, et al. Molecular profiling of cholangiocarcinoma shows potential for targeted therapy treatment decisions. Hum Pathol. 2013;44:1216–22.
Borger DR, Zhu AX. IDH mutations: new genetic signatures in cholangiocarcinoma and therapeutic implications. Exp Rev Anticancer Ther. 2012;12:543–6.
Chuang SC, Lee KT, Tsai KB, et al. Immunohistochemical study of DPC4 and p53 proteins in gallbladder and bile duct cancers. World J Surg. 2004;28:995–1000.
Yanagisawa N, Mikami T, Saegusa M, Okayasu I. More frequent beta-catenin exon 3 mutations in gallbladder adenomas than in carcinomas indicate different lineages. Cancer Res. 2001;61:19–22.
Chan-On W, Nairismagi ML, Ong CK, et al. Exome sequencing identifies distinct mutational patterns in liver fluke–related and non-infection-related bile duct cancers. Nat Genet. 2013;45:1474–8.
Churi CR, Shroff R, Wang Y, et al. Mutation profiling in cholangiocarcinoma: prognostic and therapeutic implications. PLoS One. 2014;9:e115383.
Simbolo M, Fassan M, Ruzzenente A, et al. Multigene mutational profiling of cholangiocarcinomas identifies actionable molecular subgroups. Oncotarget. 2014;5:2839–52.
Simbolo M, Gottardi M, Corbo V, et al. DNA qualification workflow for next generation sequencing of histopathological samples. PLoS One. 2013;8:e62692.
Zamo A, Bertolaso A, van Raaij AW, et al. Application of microfluidic technology to the BIOMED-2 protocol for detection of B-cell clonality. J Mol Diagn. 2012;14:30–37.
Sturm PD, Baas IO, Clement MJ, et al. Alterations of the p53 tumor-suppressor gene and K-ras oncogene in perihilar cholangiocarcinomas from a high-incidence area. Int J Cancer. 1998;78:695–8.
Borger DR, Tanabe KK, Fan KC, et al. Frequent mutation of isocitrate dehydrogenase (IDH) 1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. Oncologist. 2012;17:72–9.
Zhu AX, Borger DR, Kim Y, et al. Genomic profiling of intrahepatic cholangiocarcinoma: refining prognosis and identifying therapeutic targets. Ann Surg Oncol. 2014;21:3827–34.
Xu RF, Sun JP, Zhang SR, et al. KRAS and PIK3CA but not BRAF genes are frequently mutated in Chinese cholangiocarcinoma patients. Biomed Pharmacother. 2011;65:22–6.
Park KW, Jung ES, Kim DG, et al. ERCC1 can be a prognostic factor in hilar cholangiocarcinoma and extrahepatic bile duct cancer, but not in intrahepatic cholangiocarcinoma. Cancer Res Treat. 2013;45:63–9.
Wang J, Wang X, Xie S, et al. p53 status and its prognostic role in extrahepatic bile duct cancer: a meta-analysis of published studies. Dig Dis Sci. 2011;56:655–62.
Robertson S, Hyder O, Dodson R, et al. The frequency of KRAS and BRAF mutations in intrahepatic cholangiocarcinomas and their correlation with clinical outcome. Hum Pathol. 2013;44:2768–73.
Wang P, Dong Q, Zhang C, et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene. 2013;32:3091–100.
Acknowledgment
Supported in part by Associazione Italiana per la Ricerca sul Cancro (AIRC) Grants 12182 and 6421, FP7 EU project CAM-PaC 602783, the Italian Cancer Genome Project Grant from the Italian Ministry of Research (FIRB-RBAP10AHJB), and FIMP–Ministry of Health (CUP_J33G13000210001).
Disclosure
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Andrea Ruzzenente and Matteo Fassan shared first authorship. Calogero Iacono and Aldo Scarpa shared last authorship.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ruzzenente, A., Fassan, M., Conci, S. et al. Cholangiocarcinoma Heterogeneity Revealed by Multigene Mutational Profiling: Clinical and Prognostic Relevance in Surgically Resected Patients. Ann Surg Oncol 23, 1699–1707 (2016). https://doi.org/10.1245/s10434-015-5046-6
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-015-5046-6