Abstract
The Streptomyces sp. VKM Ac-2618D strain has been identified, and its morphological and physiological features have been studied in relation to the production of the immunosuppressant tacrolimus. The phenotypic variability of the strain was analyzed, and a dissociant with a high level of tacrolimus production was selected. Based on a comprehensive study of morphological, physiological, and chemotaxonomic properties and on phylogenetic analysis, the strain was named Streptomyces tsukubensis VKM Ac-2618D. The strain genome contains the full version of the tacrolimus biosynthetic gene cluster. The advantages of fed-batch cultivation mode for tacrolimus biosynthesis are shown. The results broaden the understanding of the characteristics of polyketide biosynthesis and can be used in the development of technology for tacrolimus production.
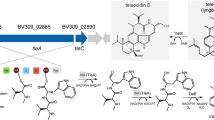
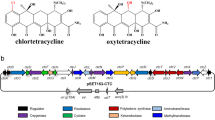
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Soil actinobacteria of the Streptomyces genus are the subject of active research due to their unique ability to synthesize biologically active substances, including antibiotics, antineoplastic agents, immunosuppressants, and antifungal and antihelminthic compounds, as well as herbicides and other substances [1, 2].
Tsukuba macrolide immunosuppressant FK-506 (Tacrolimus) was detected for the first time in 1984 in Streptomyces tsukubaensis culture broth as a result of screening by the Fujisawa Pharmaceutical Co. (now Astellas Pharma Inc.). The producer strain was isolated from a soil sample in the Tsukuba region (Japan), and tacrolimus became the first known immunosuppressant with a macrolide structure [3, 4]. Tacrolimus is currently recognized as the most effective immunosuppressive agent and is widespread in various fields of medicine: transplantology [5, 6], dermatology [7], the treatment of autoimmune [8] and viral [9] diseases, etc. The range of its clinical application is constantly expanding. Recent studies indicate its effectiveness in combination with anti-inflammatory corticosteroid drugs in the treatment of patients with serious complications caused by the new coronavirus infection (COVID-19) [11].
All of the known tacrolimus producers belong to the Streptomyces genus. A strain capable of synthesizing tacrolimus was discovered in 1984. It was patented first as S. tsukubaensis 9993 [3, 4] and deposited later as S. tsukubaensis NRRL 18488. Despite the number of publications on the features of tacrolimus biosynthesis by the NRRL 18488 strain, the species did not receive a proper taxonomic description until 2013. Due to a deep study of the morphological and physiological properties of the 9993T = NRRL 18488T (S. tsukubaensis) strain, which had become typical by that time, its phylogenetic position was confirmed and its species name S. tsukubensis was clarified [12].
Many aspects of tacrolimus biosynthesis by various S. tsukubensis strains have been elucidated to date: the features of the genome structure have been clarified; the gene cluster encoding enzyme systems for tacrolimus biosynthesis and examples of obtaining mutant producer strains have been described [13, 14]; the relationship of tacrolimus biosynthesis and central cellular metabolism has been studied [15, 16]; and approaches are being developed to increase the productivity of strains via the creation of nutrient media and optimization of the culturing conditions [17, 18]. However, the productivity levels of some strains are not yet sufficient, and the formation of side products such as ascomycin (FK-520), which complicates the isolation and purification of the tacrolimus substance to pharmacopoeial quality, as well as the unstable biosynthesis of the target compound in known producers, are key problems in tacrolimus production that are still to be resolved. There is practically no information in the literature on the morphostructural characteristics of the known producers and the effect of their phenotypic variability on the biosynthesis of the target compound.
The Streptomyces sp. VKM Ac-2618D strain is a highly active tacrolimus producer. The technique to obtain tacrolimus based on the use of this strain and various types of polymeric sorbents [19] made it possible for the first time to discover the effect of stimulation of tacrolimus biosynthesis by yeast [20], to propose some new solutions based on the addition of high- and low-molecular starches of various structures [21] that increase the tacrolimus yield, and to obtain a substance of pharmacopoeial quality [22]. However, there is a significant gap due to the lack of an evidence base for the reliable identification of the strain species. In addition, the practical use of the strain was also complicated by the low stability of the activity associated with the target compound biosynthesis due to the phenotypic variability of the strain.
The goal of the present work was to identify the tacrolimus-producing strain and to study its morphological and physiological properties associated with tacrolimus production.
EXPERIMENTAL
Reagents
Soluble starch (Kupavnareaktiv, Russia), glucose and α-asparagine (Dia-M, Russia), baker’s yeast (Saff-Moment, France), corn extract (Sigma-Aldrich, United States), malt extract (Fluka, United States), L‑lysine monohydrocloride (PanReac, United States), a sorbent of XAD type (Sigma-Aldrich), Aspergillus oryzae α-amylase (Serva, Germany), and the tacrolimus reference preparation for HPLC (Zhejiang Hisun Pharmaceutical Co. Ltd., China) were used in this work. The rest of the reagents and solvents were of chemically pure or pure for analysis grades from Russian manufacturers.
Microorganism and Conditions for Its Maintenance
The Streptomyces sp. strain VKM Аc-2618D was received from the All-Russia Collection of Microorganisms of the Institute of Biochemistry and Physiology of Microorganisms of the Russian Academy of Sciences (VKM). The Streptomyces sp. VKM Ас-2618D culture was maintained on MDA medium of the following composition, g/L: soluble starch—10; yeast extract—4; malt extract—10; and agar—20; pH 7.0.
Media and Culturing Conditions
The taxonomic analysis was carried out during culture growth on International Streptomyces Project (ISP) media [23], which contains malt extract/yeast extract agar (ISP-2), oatmeal agar (ISP-3), starch agar with inorganic salts (ISP-4), and asparagine-glycerol agar (ISP-5).
To study chemotaxonomic and molecular genetic properties, the strain was cultured in the following medium, g/L: soluble starch—20; glucose—20; peptone—18, pH 7.0. It was cultured under aerobic conditions on a shaker (220 rpm) at 25°С in two stages of 48 h and 24 h, respectively.
The growth conditions for the inoculum and tacrolimus biosynthesis were as described earlier [20, 21]. The process was carried out in the presence of a XAD polymer sorbent for 10 days in a batch or fed-batch mode (the latter with the addition of sterile 6% starch from the second day to the seventh day).
Biomass Evaluation
The amount of biomass was estimated by the dry weight, as described previously [20, 21].
Study of Phenotype Characteristics
The standard ISP protocol was used to describe the morphology of the strain colonies [23]. The colony color was assessed with the RAL K5 Classic gGmbH scale (Germany). The type of spore chain and ornamentation of the spore surface were studied with a JSM-6510 electron microscope (JEOL, Japan) in 14‑day cultures grown on ISP-2 medium at 30°С according to [24]. The phenotypic dissociation of the strain was analyzed via plating on MDA medium and incubation at 30°С for 14 days, after which the number of colonies of various phenotypes was counted. The phenotypic variability was assessed in culture samples taken on days 1, 3, 6, and 10 of cultivation; samples for scanning electron microscopy were prepared as described in [24].
The spore formation during submerged cultivation was additionally assessed via heating of the culture broth at 65°С for 15 min, followed by plating on agar medium MDA.
Study of Biochemical and Chemotaxonomic Characteristics
The use of substrates as a sole carbon source was determined with an API 50 СН test kit (bioMerieux, United States) in accordance with the manufacturer’s recommendations.
The enzymatic activity of Streptomyces sp. VKM Ас-2618D was assessed with API ZYM express tests (bioMerieux) according to the manufacturer’s recommendations.
The purified preparations of cell walls were obtained with a known method [25]. The amino-acid composition of peptidoglycans was determined with a Hitachi amino acid analyzer (Japan). Isoprenoid quinones were isolated with the protocol [26]; the menaquinone composition was determined via mass-spectrometry (МАТ 8430, Finnigan MAT, Germany). The fatty-acid composition was analyzed as described in [27]. Methyl esters of fatty acids were studied with a Perkin-Elmer F-17 gas chromatograph (Perkin Elmer, Germany).
The content of GC pairs was evaluated based on the DNA melting temperature [28, 29].
Phylogenetic Analysis
Chromosomal DNA was isolated and purified with a modified method [30]. The DNA concentration was measured on a Shimadzu UV-160 spectrophotometer (Japan).
The 16S RNA was amplified on a Tertsyk amplifier (DNK-Teknologii, Russia). The universal bacterial primers 27f (AGAGTTTGATC(A/C)TGGCTCAG) and 1492r (ACGG(С/Т)TACCTTGTTACGACTT) were used for this purpose. The nucleotide sequences of the 16S RNA gene were analyzed with an ABI PRISM 3730 automatic sequencer (Applied Biosystems, United States) with an ABI PRISM BigDyeTM reagent set (Applied Biosystems, Terminator v.3.1) according to the manufacturer’s recommendations. Phylogenetic analysis and dendrogram building were carried out with the MEGA 4 software package [31] and the neighbor-joining method [32].
The phylogenetic position of the strain was confirmed via full-genome sequencing and subsequent annotation. The library of DNA fragments 300–400 bp long was obtained with a NEBNext Ultra II DNA Library Prep Kit for Illumina (Illumina, United States). The library was sequenced on HiSeq 2500 with the use of a HiSeq Rapid PE Cluster Kit v2 (Illumina) and a HiSeq Rapid SBS Kit v2 (Illumina) (500 cycles). The quality of the obtained reads was assessed with FastQC 0.11.8 [33]. Adapter fragments were eliminated from the initial reads and the reads containing artifact sequences and PhiX were filtered with BBDuk 38.35. After that, the reads were purified of possible contamination with human DNA. To this end, the reads were mapped with BBMap 38.35 in the human genome and excluded if their identity with this genome exceeded 95%. Then, low-quality reads were eliminated with BBDuk 38.35. The purified reads were assembled with the help of SPAdes 3.13.0 [34]; the resulting contigs were removed if their size was less than 200 bp. The genome was annotated with the NCBI Prokaryotic Genomes Automatic Annotation Pipeline service. The average nucleotide identity based on BLAST values (ANIb) and digital DNA-DNA hybridization (dDDH) values were calculated with the JSpecies 1.2.1 software [35] and the GGDC 2.1 service [36], respectively.
Analysis of the Bound Glucose Content
For this purpose, culture-broth samples were hydrolyzed with 3 M trifluoroacetic acid at 100°С for 3 h. The bound glucose was quantified with the method in [37]. The sugar concentration was measured on a carbohydrate analyzer (Biotronik, Germany) at a wavelength of 570 nm.
Analysis of the Tacrolimus Content
The compound was determined via HPLC as described previously [20, 21].
Statistical Analysis
The experimental data were obtained in triplicate. Results are presented as means with sample standard deviations.
RESULTS AND DISCUSSION
Strain Taxonomic Position
The Streptomyces sp. strain VKM Ас-2618D forms on solid media in dense, opaque colonies with pronounced differentiation of the structural surface. The colonies range in color from pinkish orange to brick orange. The strain releases a pinkish orange pigment into the medium (Fig. 1a); it is characterized by the development of a branched substrate mycelium, which differentiates into straight chains of cylindrical spores with a smooth surface (Fig. 1b).
(а) Morphology (photographs) of the Streptomyces sp. VKM Ас-2618D strain colonies after 14 days of incubation at 30°C on ISP media: (а) ISP-2, (b) ISP-3, (c) ISP-4, and (d) ISP-5; (b) morphology (scanning electron micrographs) of spore chains after Streptomyces sp. VKM Ас-2618D growth on ISP 2 for 14 days at 30°C (scale bar is 5 μm).
Analysis of the Streptomyces sp. VKM Ас-2618D cell-wall composition revealed the presence of LL-diaminopimelic acid and a lack of diagnostic sugars, which is characteristic of type-I cells. The bulk of fatty acids in cell walls is represented by iso-С14:0 (18.81%), anteiso-C15:0 (17.05%), iso-C16:0 (15.16%), and iso-C15:0 (15.11%). The main menaquinones are MK-9 (H8) and MK-9 (H6).
The strain actively utilized D-glucose, dextrin, glycerol, starch, D-ribose, and N-acetylglucosamine, while poor growth was observed on maltose and salicin. High esterase, leucine-arylamidase, naphtol-AS-BI-phosphohydrolase, and α-glycosidase activities were also typical for the strain.
Preliminary analysis of the 16S rRNA sequences showed high similarity (99.9%) of Streptomyces sp. VKM Ас-2618D with the typical Streptomyces tsukubensis strain NRRL 18488Т (Fig. 2).
Phylogenetic tree based on the sequencing of a 16S rRNA gene fragment of Streptomyces species with grouping via the neighbor-joining method. Bootstrap values based on 1000 replicates are shown at branch nodes. A bar corresponds to 0.005 substitutions per nucleotide position. The 16S rRNA gene nucleotide sequence of Micromonospora rifamycinica AM 105T (AY561829) was used as an outgroup.
We recently performed full-genome sequencing and genome annotation to confirm the taxonomic identification of the VKM Ас-2618D strain [38]. The genomic sequence of the strain was deposited in the DDBJ/ENA/GenBank as SGFG00000000. The size of the Streptomyces sp. VKM Ас-2618D genome was shown to be 7.93 Mb with an average G+C content of 71.9%. The number of candidate protein-encoding genes was 6265 (1227 of which code for hypothetical proteins); 66 tRNA, 23 whole and partial rRNA, and 3 ncRNA were also found.
In this work, we carried out a comparative analysis of the genomes of the VKM Ac-2618D strain and the typical S. tsukubensis strain NRRL 18488T (CP029157.1-CP029159.1). It was shown that the values of ANI and dDDH between the sequences of these genomes were 99.99 and 99.9%, respectively, which is much higher than the threshold values for species distinction (95–96% for ANI and 70% for dDDH). This indicated that the VKM Ac-2618D strain belongs to the species Streptomyces tsukubensis.
Detailed analysis of the genome VKM Ас-2618D sequence also made it possible to detect the gene cluster of the tacrolimus biosynthesis.
It is known that the cluster of the tacrolimus biosynthesis genes encodes the polyketide synthase I–non-ribosomal peptide synthetase (PKSI-NRPS) system. Two types of this cluster have been described: those with a reduced gene set, which including 19 genes (S. tacrolimicus and S. kanamyceticus strains), and those with the complete (extended) gene set, which contains 25–26 genes (S. tsukubaensis and Streptomyces sp. KCTC 11604BP strains) [39]. The VKM Ac-2618D genome was shown to contain seven genes of the so-called full version of the tacrolimus biosynthesis cluster (Fig. 3) that are absent in the short version typical of the S. tacrolimicus ATCC 55098 and S. kanamyceticus KCTC 9225 producer strains.
Similar clusters with the full gene set for tacrolimus biosynthesis were found in the S. tsukubensis NRRL 18488T, S. tsukubensis L19, and Streptomyces sp. KCTC 11604BP strains [40]. We previously noted differences between the gene clusters in the S. tsukubensis VKM Ac-2618D and S. tsukubensis NRRL 18488T strains; however, the later publication of the full genome sequence for the S. tsukubensis NRRL 18488T (CP029157.1-CP029159.1) strain changed the situation. Comparative analysis of the genomic data available by the time of the present work publication showed that the gene clusters of tacrolimus synthesis in the S. tsukubensis VKM Ac-2618D (SGFG01000008.1:83700-166586), S. tsukubensis NRRL 18488T (CP029159.1:c7554111-7471225), and Streptomyces sp. KCTC 11604BP (HM116537.1:8712-91598) strains are completely identical.
Phenotypic Variability and Tacrolimus Biosynthesis
A high degree of morphological and physiological heterogeneity of the population (or phenotypic dissociation) is a typical feature of streptomycetes. This characteristic is expressed in the appearance of morphologically different colonies from a single predecessor colony, which are designated below as dissociants. Four dissociants with different morphologies were observed in the studied strain when grown on MDA (Fig. 4a).
Cultures grown from single colonies of different dissociants were analyzed for their capacity for tacrolimus production. The differences in tacrolimus productivity accounted for 30–86% (Fig. 4b).
The highest biosynthetic potential was characteristic of dissociant D2, which was represented by round-shaped colonies with a diameter of 7–13 mm of a bright-brick color with a dry, opaque surface. It grew weakly in agar but rose noticeably above its surface and had a central depression of a rounded or irregular shape, bounded by a roller. The lateral slopes of the colonies had a comb-wrinkled structure; the colonies were surrounded by a thin, flattened, hyaline layer of dull yellow or pastel orange; the spore formation was weak or completely absent (Fig. 4a). The dissociant provided a high and stable tacrolimus yield. We determined and optimized the conditions to maintain a high biosynthetic activity in dissociant D2 and to minimize the influence of dissociation on biosynthesis. This was achieved with the storage of dissociant D2 at –70°С with monitoring of its biosynthetic potential via plating on MDA medium and a series of alternating passages on liquid and solid media. As a result, the number of colonies of dissociant D2 with low activity during cultivation on solid MDA after growth in a liquid medium did not exceed 1–2%.
Earlier, we showed the positive effect of fed-batch cultivation on tacrolimus biosynthesis, which was expressed by a more than twofold increase in the immunosuppressant yield as compared to the batch-growth regime [21]. In this work, we assessed micro- and macromorphological changes in the producer’s mycelium. During the first two days, the mycelium biomass grew by 22–25 times (Fig. 5a), which was accompanied by an almost twofold decrease in the carbon source content (Fig. 5b). The amount of biomass then decreased (Fig. 5a), which indicated the beginning of the stationary growth phase and tacrolimus biosynthesis (Fig. 5b). The starch-fed batch mode provided a smoother decrease in biomass during tacrolimus biosynthesis as compared to batch mode; the average difference in biomass content was 36% (Fig. 5a).
(a) Dynamics of biomass growth and tacrolimus biosynthesis by S. tsukubensis VKM Ac-2618D in the batch (1) and fed-batch (2) cultivation modes. (b) Dynamics of the content of bound glucose (1a and 1b) and the target product (2a and 2b) in the medium in batch (1a and 2a) and fed-batch (1b and 2b) cultivation modes during tacrolimus biosynthesis by S. tsukubensis VKM Ac-2618D.
According to SEM data, the 1-day-old mycelium developing during tacrolimus biosynthesis in batch mode was represented by hyphae with a smooth outline without signs of destruction (Fig. 6a).
Morphological changes in S. tsukubensis VKM Ac-2618D during tacrolimus biosynthesis: (a–c) batch cultivation mode, (d–f) fed-batch cultivation mode; (a) and (d) 1 day; (b) and (e) 6 days; (c) and (f) 10 days (scanning electron microscopy, scale bar is 5 μm). Arrows indicate destructive changes, such as hyphal profile curvature and loss of hyphal intracellular content.
On the sixth day (the stage of active biosynthesis), the structure of the productive mycelium hyphae changed from cylindrical to sinuous; the clarity of their boundaries and the cytoplasm density were lost, and empty hyphal envelopes appeared (Fig. 6b). On the tenth day (the maximal production of tacrolimus and a decrease in the biosynthesis rate), enhancement of destructive processes were observed. This was expressed by a severe curvature of the hyphal profile and loss of the integrity of part of the mycelium (empty envelopes) (Fig. 6c).
Similar changes in the mycelium structure were also observed as a result of fed-batch cultivation; however, the degradation was less pronounced in this case (Figs. 6d–6f). The formation of sporiferous cultures was not observed during submerged growth and biosynthesis of the target product.
Fed-batch cultivation did not give the expected effect of pronounced mycelium intactness; however, it was accompanied by a higher level of the tacrolimus production (Figs. 5b, 6d–6f). The tacrolimus yield during fed-batch cultivation was higher than the 100% titer in control and amounted to 500 ± 47 and 245 ± 22 mg/L in fed-batch and batch modes, respectively. In terms of biomass content, the difference was not so large, only 36%.
The different levels of the tacrolimus production may be a consequence of the different amounts of NADPH in each of the used cultivation modes. It is known that the biosynthesis of most antibiotics (including tacrolimus) is an energy-consuming process, in which NADPH acts as the main reducing cofactor. In most microorganisms, the largest portion of NADPH is synthesized through the pentosophosphate pathway (PPP), which involves the glucose-6-phosphate dehydrogenase and phosphogluconate dehydrogenase activities. For instance, it was shown that the carbon flow through the PPP stimulated an increase in methylenomycin production by S. coelicolor [41]. It is most likely that the products of the enzymatic hydrolysis of starch, which was also introduced during tacrolimus biosynthesis by S. tsukubensis VKM Ac-2618D, were metabolized through the PPP with the direct uptake of synthesized NADPH for the needs of biosynthetic processes.
On the one hand, the synthesis of antibiotics depends on the occurrence of both NADPH and ATP, and, on the other hand, a high energetic charge in the cell inhibits its secondary metabolism [42]. The fact that the derepression of the genes of antibiotic biosynthesis is due to the decrease in the intracellular ATP supports the latter statement. For instance, it was reported that a high intracellular content of ATP is required for the synthesis of gramicidin C by a B. brevis strain. Conversely, the presence of reduced NADPH has an extremely strong effect on tetracycline synthesis that the tetracycline yield increases with decreases in the intensity of the TCA reactions [43]. A positive relationship was established between the production of FK-506 and the accumulation of some metabolites of both PPP and TCA [15, 16]. At the same time, the presence of inorganic phosphate at high concentrations in the medium is a negative factor for tacrolimus biosynthesis by S. tsukubaensis NRRL 18488Т, which is usually the result of intracellular ATP formation [18].
We previously showed that baker’s yeasts inactivated via medium sterilization have a stimulating effect on tacrolimus biosynthesis [20]. On the first day of cultivation, inactivated yeasts were represented by whole cells with intact cell walls and internal contents, while noticeable destructive changes were observed by the tenth day of biosynthesis. Empty yeast-cell envelopes appeared at the phase of active tacrolimus synthesis; these structures significantly differed from cells of saccharomycetes on the first days of incubation, when intracellular contents were seen in them. The yeast cells remained unchanged in the control medium without inoculation with Streptomyces culture.
The final stage of tacrolimus biosynthesis under the studied cultivation modes (Fig. 5b) was characterized by the predominance of mycelial destruction over the growth processes (Fig. 6).
CONCLUSIONS
Thus, the Streptomyces sp. strain VKM Ас-2618D was identified as S. tsukubensis VKM Ac-2618D based on a complex of chemotaxonomic and molecular biological studies, as well as in silico DNA-DNA hybridization assay. An in-depth analysis of the genome showed the presence of a cluster for tacrolimus biosynthesis with a full set of genes (26 pieces), which correlated with a high biosynthetic activity of the strain. The phenotypic variability of the strain was studied, and the D2 dissociant with the highest tacrolimus productivity was selected. The advantages of fed-batch cultivation over batch mode were shown; it was assumed that these advantages are mainly associated with the formation of higher amounts of cofactors required for the FK-506 biosynthesis, in particular, NADPH. The results can be used for the development of a tacrolimus production process.
REFERENCES
Chaudhary, A.K., Dhakal, D., and Sohng, J.K., An insight into the “-omics” based engineering of streptomycetes for secondary metabolite overproduction, Biomed Res. Int., 2013, vol. 2013, article ID 968518. https://doi.org/10.1155/2013/968518
Procópio, R.E., Silva, I.R., Martins, M.K., et al., Antibiotics produced by Streptomyces, Braz. J. Infect. Dis., 2012, vol. 16, no. 5, pp. 466–471. https://doi.org/10.1016/j.bjid.2012.08.014
Kino, T., Hatanaka, H., Hashimoto, M., et al., Fk506, a novel immunosuppressant isolated from a Streptomyces. I. Fermentation, isolation, and physico-chemical and biological characteristics, J. Antibiot. (Tokyo), 1987, vol. 40, no. 9, pp. 1249–1255. https://doi.org/10.7164/antibiotics.40.1249
Kino, T., Hatanaka, H., Miyata, S., et al., Fk506, a novel immunosuppressant isolated from a Streptomyces. II. Immunosuppressive effect of FK-506 in vitro, J. Antibiot. (Tokyo), 1987, vol. 40, no. 9, pp. 1256–1265. https://doi.org/10.7164/antibiotics.40.1256
Trede, N.S., Warwick, A.B., Rosoff, P.M., et al., Tacrolimus (FK506) in allogeneic bone marrow transplantation for severe aplastic anemia following orthotopic liver transplantation, Bone Marrow Transplant., 1997, vol. 20, no. 3, pp. 257–260. https://doi.org/10.1038/sj.bmt.1700872
McCormack, P.L. and Keating, G.M., Tacrolimus: in heart transplant recipients, Drugs, 2006, vol. 66, no. 17, pp. 2269–2279. https://doi.org/10.2165/00003495-200666170-00010
Remitz, A. and Reitamo, S., Long-term safety of tacrolimus ointment in atopic dermatitis, Expert Opin. Drug Saf., 2009, vol. 8, no. 4, pp. 501–506. https://doi.org/10.1517/14740330902969441
Akimoto, K., Kusunoki, Y., Nishio, S., et al., Safety profile of tacrolimus in patients with rheumatoid arthritis, Clin. Rheumatol., 2008, vol. 27, no. 11, pp. 1393–1397. https://doi.org/10.1007/s10067-008-0931-z
Karpas, A., Lowdell, M., Jacobson, S.K., and Hill, F., Inhibition of human immunodeficiency virus and growth of infected T cells by the immunosuppressive drugs cyclosporine A and FK 506, Proc. Natl. Acad. Sci. U. S. A., 1992, vol. 89, no. 17, pp. 8351–8355. https://doi.org/10.1073/pnas.89.17.8351
Reis, S.A., Moussatché, N., and Damaso, C.R.A., FK506, a secondary metabolite produced by Streptomyces, presents a novel antiviral activity against Orthopoxvirus infection in cell culture, J. Appl. Microbiol., 2006, vol. 100, no. 6, pp. 1373–1380. https://doi.org/10.1111/j.1365-2672.2006.02855.x
Russell, B., Moss, C., George, G., et al., Associations between immune-suppressive and stimulating drugs and novel COVID-19—a systematic review of current evidence, Ecancermedicalscience, 2020, vol. 14, p. 1022. https://doi.org/10.3332/ecancer.2020.1022
Muramatsu, H. and Nagai, K., Streptomyces tsukubensis sp. nov., a producer of the immunosuppressant tacrolimus, J. Antibiot. (Tokyo), 2013, vol. 66, no. 4, pp. 251–254. https://doi.org/10.1038/ja.2012.116
Mo, S., Lee, S.K., Jin, Y.Y., et al., Application of a combined approach involving classical random mutagenesis and metabolic engineering to enhance FK506 production in Streptomyces sp. RM7011, Appl. Microbiol. Biotechnol., 2013, vol. 97, no. 7, pp. 3053–3062. https://doi.org/10.1007/s00253-012-4413-5
Ban, Y.H., Park, S.R., and Yoon, Y.J., The biosynthetic pathway of FK506 and its engineering: from past achievements to future prospects, J. Ind. Microbiol. Biotechnol., 2016, vol. 43, nos. 2–3, pp. 389–400. https://doi.org/10.1007/s10295-015-1677-7
Huang, D., Li, S., Xia, M., et al., Genome-scale metabolic network guided engineering of Streptomyces tsukubaensis for FK506 production improvement, Microb. Cell Fact., 2013, vol. 12, no. 1, p. 52. https://doi.org/10.1186/1475-2859-12-52
Huang, D., Xia, M., Li, S., et al., Enhancement of FK506 production by engineering secondary pathways of Streptomyces tsukubaensis and exogenous feeding strategies, J. Ind. Microbiol. Biotechnol., 2013, vol. 40, no. 9, pp. 1023–1037. https://doi.org/10.1007/s10295-013-1301-7
Singh, B.P. and Behera, B.K., Regulation of tacrolimus production by altering primary source of carbons and amino acids, Lett. Appl. Microbiol., 2009, vol. 49, no. 2, pp. 254–259. https://doi.org/10.1111/j.1472-765X.2009.02652.x
Martínez-Castro, M., Salehi-Najafabadi, Z., Romero, F., et al., Taxonomy and chemically semi-defined media for the analysis of the tacrolimus producer Streptomyces tsukubaensis, Appl. Microbiol. Biotechnol., 2013, vol. 97, no. 5, pp. 2139–2152. https://doi.org/10.1007/s00253-012-4364-x
Sukhodol'skaya, G.V. and Lobastova, T.G., A method for obtaining tacrolimus by microbiological synthesis, RF Patent no. 2495937, Byull. Izobret., 2013, no. 29.
Poshekhontseva, V.Yu., Fokina, V.V., Sukhodol’skaya, G.V., et al., Influence of lower fungi on the biosynthesis of tacrolimus (FK-506) by the Streptomyces tsukubensis strain VKM Ac-2618D, Biotekhnologiya, 2019, vol. 35, no. 5, pp. 42–50. https://doi.org/10.21519/0234-2758-2019-35-5-42-50
Poshekhontseva, V.Yu., Fokina, V.V., Sukhodol’skaya, G.V., et al., Effect of starch composition on the biosynthesis of immunosuppressant tacrolimus (FK-506) by Streptomyces tsukubaensis VKM Ac-2618D strain, 2618, Appl. Biochem. Microbiol., 2019, vol. 55, no. 5, pp. 534–543. https://doi.org/10.1134/S0003683819040148
Salionov, D.S., Poshekhontseva, V.Yu., Fokina, V.V., et al., Biosynthesis of tacrolimus by the Streptomyces tsukubensis VKM Ac-2618D strain in the presence of polymeric sorbents and development of a method for its isolation and purification, Appl. Biochem. Microbiol., 2020, vol. 56, no. 6, pp. 699–707.
Shirling, E.B. and Gottlieb, D., Method for characterization of Streptomyces species, Int. J. Syst. Evol. Microbiol., 1966, vol. 16, no. 3, pp. 313–340. https://doi.org/10.1099/00207713-16-3-313
Chemodurova, A.A., Kaparullina, E.N., Machulin, A.V., et al., Ancylobacter lacus sp. nov. and Ancylobacter plantiphilus sp. nov., novel aerobic facultative methylotrophic bacteria, Microbiology (Moscow), 2020, vol. 89, no. 1, pp. 35–43. https://doi.org/10.1134/S0026261720010051
Schleifer, K.H. and Kandler, O., Peptidoglycan type of bacterial cell walls and their taxonomic implications, Bacteriol. Rev., 1972, vol. 36, no. 4, pp. 407–477. https://doi.org/10.1128/MMBR.36.4.407-477.1972
Collins, M.D. and Jones, D., The distribution of isoprenoid quinone structural types in bacteria and their taxonomic implication, Microbiol. Rev., 1981, vol. 45, no. 2, pp. 316–354.
Evtushenko, L.I., Taptykova, S.D., Akimov, V.N., and Dobritsa, S.V., A new species of actinomycete, Amycolata alni, Int. J. Syst. Bacteriol., 1989, vol. 39, no. 1, pp. 72–77. https://doi.org/10.1099/00207713-39-1-72
Mesbah, N.M., Whitman, W.B., and Mesbah, M., Determination of the G+C content of prokaryotes, in Methods in Microbiology, Rainey, F. and Oren, A., Eds., Waltham, MA: Academic, 2011, vol. 38, pp. 299–324.
Owen, R.J. and Pitcher, D., Current methods for estimating DNA base composition and levels of DNA–DNA hybridization, in Chemical Methods in Bacterial Systematics, Goodfellow, M. and Minnikin, D.E., Eds., London: Academic, 1985, pp. 67–93.
Marmur, J., A procedure for the isolation of deoxyribonucleic acid from microorganisms, Mol. Biol., 1961, vol. 3, no. 2, pp. 208–216.
Tamura, K., Didley, J., Nei, M., and Kumar, S., M-EGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0, Mol. Biol. Evol., 2007, vol. 24, no. 8, pp. 1596–1599. https://doi.org/10.1093/molbev/msm092
Saitou, N. and Nei, M., The neigbour-joining method: new method for reconstructing phylogenetic trees, Mol. Biol. Evol., 1987, vol. 4, no. 4, pp. 406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Andrews, S., FastQC: a quality control tool for high throughput sequence data, 2010. www.bioinformatics. babraham.ac.uk/projects/fastqc.
Bankevich, A., Nurk, S., Antipov, D., et al., SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing, J. Comput. Biol., 2012, vol. 19, no. 5, pp. 455–477. https://doi.org/10.1089/cmb.2012.0021
Richter, M. and Rossello-Mora, R., Shifting the genomic gold standard for the prokaryotic species definition, Proc. Natl. Acad. Sci. U. S. A., 2009, vol. 106, no. 45, pp. 19126–19131. https://doi.org/10.1073/pnas.0906412106
Meier-Kolthoff, J.P., Auch, A.F., Klenk, H.P., and Goker, M., Genome sequence-based species delimitation with confidence intervals and improved distance functions, BMC Bioinformatics, 2013, vol. 14, no. 1, p. 60. https://doi.org/10.1186/1471-2105-14-60
Sinner, M. and Puls, J., Non-Corrosive dye reagent for detection of reducing sugars in borate complex ion-exchange chromatography, J. Chromatogr. A, 1978, vol. 156, no. 1, pp. 197–204. https://doi.org/10.1016/S0021-9673(00)83140-4
Poshekhontseva, V.Y., Bragin, E.Y., Fokina, V.V., et al., Draft genome sequence of FK506-producing Streptomyces tsukubensis strain VKM Ac-2618D, Microbiol. Resour. Announc., 2019, vol. 8, no. 24, pii: e00510-19. https://doi.org/10.1128/MRA.00510-19
Goranovič, D., Blažič, M., Magdevska, V., et al., FK506 biosynthesis is regulated by two positive regulatory elements in Streptomyces tsukubaensis, BMC Microbiol., 2012, vol. 12, pp. 238. https://doi.org/10.1186/1471-2180-12-238
Ordóñez-Robles, M., Santos-Beneit, F., and Martín, J.F., Unraveling nutritional regulation of tacrolimus biosynthesis in Streptomyces tsukubaensis through omic approaches, Antibiotics (Basel), 2018, vol. 7, no. 2, p. 39.https://doi.org/10.3390/antibiotics7020039
Obanye, A.I.C., Hobbs, G., and Gardner, D.C.J., Oliver. S.G. Correlation between carbon flux through the pentose phosphate pathway and production of the antibiotic methylenomycin in Streptomyces coelicolor A3 (2), Microbiology, 1996, vol. 142, no. 1, pp. 133–137. https://doi.org/10.1099/13500872-142-1-133
Martin, J.F., Phosphate control of the biosynthesis of antibiotics and other secondary metabolites is mediated by the PhoR–PhoP system: an unfinished story, J. Bacteriol., 2004, vol. 186, no. 16, pp. 5197–5201. https://doi.org/10.1128/JB.186.16.5197-5201.2004
Zheldakova, R.A., Mekhanizmy biosinteza antibiotikov i ikh deistvie na kletki mikroorganizmov. Uchebno-metodicheskii kompleks dlya studentov spetsial’nosti 1-31 01 01 “Biologiya” (Mechanisms of Biosynthesis of Antibiotics and Their Effect on Cells of Microorganisms. Educational-Methodical Complex for Students of Specialty 1-31 01 01 “Biology”), Minsk: Belorus. Gos. Univ., 2004.
Funding
The research was carried out within the framework of the state assignment АААА-А16-116062110077-6.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest.
This article does not contain any studies involving animals performed by any of the authors.
This article does not contain any studies involving human participants performed by any of the authors outside the scope of people’s normal professional activities.
Additional information
Translated by I. Gordon
Abbreviations: ANI—average nucleotide identity; dDDH—digital DNA-DNA hybridization; FK-506—tacrolimus; FK-520—ascomycin; HPLC—high-performance liquid chromatography; NADPH–nicotinamide adenine dinucleotide phosphate; PPP—pentosophosphate pathway; SEM—scanning electron microscopy; TCA—tricarboxylic acid cycle.
Rights and permissions
About this article
Cite this article
Poshekhontseva, V.Y., Fokina, V.V., Tarlachkov, S.V. et al. Streptomyces tsukubensis VKM Aс-2618D—an Effective Producer of Tacrolimus. Appl Biochem Microbiol 57, 939–948 (2021). https://doi.org/10.1134/S0003683821090064
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683821090064