Abstract
The archaeal composition of permafrost samples taken during the drilling of frozen marine sediments in the area of the Barentsburg coal mine on the east coast of Grønfjord Bay of Western Spitsbergen has been studied. This study is based on an analysis of the V4 region of the 16S rRNA gene, carried out using next-generation sequencing. The general phyla of the Archaea domain are Euryarchaeota, Bathyarchaeota, Thaumarchaeota, and Asgardarchaea. As a result of a phylogenetic analysis of the dominant operational taxonomic units, representatives of methanogenic and methane- and ammonium-oxidizing archaea, as well as heterotrophic archaea, are found. The methanogenic archaea of Euryarchaeota phylum, Methanobacteria class, are found in permafrost with controversial genesis, while the methane-oxidizing archaea of Methanomicrobia class Methanosarcinales order are found in the marine permafrost at cape Finneset: the ANME-2a, -2b group in layers of 8.6 and 11.7 m and the ANME-2d group (Candidatus Methanoperedens) in a layer of 6.5 m. Ammonium-oxidizing archaea of phylum Thaumarchaeota is present in all types of permafrost, while the order of Nitrososphaerales is found in permafrost with controversial genesis and the order Nitrosopumilales is in permafrost with marine and controversial genesis. Representatives of phylum Bathyarchaeota are found stratigraphically in the most ancient samples under study. Asgardarchaeota superfylum is excluded in the layers of permafrost with marine genesis and is represented by the phyla Lokiarchaeota, Thorarchaeota, and an unclassified group belonging to this superphylum. The presence of methane, ethylene, and ethane in the permafrost of the first sea terrace of Cape Finnemet at a depth of 11.7 m, as well as the composition of the archaeal community, give us reason to assume that, before freezing, microbiological processes of anaerobic methane oxidation took place in it, probably received from Tertiary rocks. The results of both this and previous works present the Spitsberen permafrost as a rich archive of genetic information of little-studied prokaryotic groups.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
INTRODUCTION
The Spitsbergen is a unique region of the Arctic where fast processes associated with climate warming are recorded. According to meteorological data from Longyearbyen, the average annual air temperature increased during the 20th century from about –9 to ‒4°С (Humlum et al., 2003). The area of glaciers in the vicinity of Longyearbyen and Barentsburg decreased by about 50% from 1936 to 2017 (Chernov et al., 2018). Spitsbergen permafrost is the warmest within the High Arctic due to the warming influence of currents and air masses carried from the Atlantic with the West Spitsbergen current. In 2016, as part of the creation of a permafrost test site of the Russian Science Center on the eastern bank of Grønfjord Bay, boreholes were drilled to study the structural features of permafrost and monitor ground temperatures (Demidov et al., 2016, 2019). Permafrost coastal–marine sediments frozen in the late Pleistocene and Holocene after they emerged from under the sea level, were drilled. This study is a continuation of the ongoing work on a comprehensive study of permafrost in the vicinity of the Barentsburg mine. Comparative microbiological studies of the Arctic systems of seawater–marine sediment, as well as terrestrial–subsea permafrost, show the differences between the dominant prokaryotic groups. For the Archaea domain, in the same seawater–marine sediments transition, an increase in the proportions of the orders of Methanomicrobia and Methanococci and a decrease in the order of Methanobacteria of the phylum Euarchaeota, as well as a significant increase in the proportion of Thermoprotei (Crenarchaeota) and the appearance of the order Thermoplasmata (Euarychaeota), were observed (Hamdan et al., 2013). The dominant groups of the Archaea domain in permafrost in the Cape Mamontov Klyk area were anaerobic methanogenic, ammonium-oxidizing, and methane-oxidizing Archaea. The permafrost in the Buor-Khaya Bay (southern Laptev Sea) was dominated by groups of methanogenic and methane-oxidizing Achaea (Winkel et al., 2018). In West Antarctica in the South Atlantic Ocean, the composition of the Archaea domain of King George Island—rich in microbial methane and about 7500 years old—the dominance of methanogenic and ammonium-oxidizing Archaea was revealed using the V3-V5 and V1-V5 regions of 16S rRNA (Karaevskaya et al., 2014). In the study of 2-m strata of marine sediments from the offshore of Northwestern Spitsbergen with a nonvertical methane emission, representatives of phylum Crenarchaeota, Miscellaneous Crenarchaeotal group (at the top of the strata), as well as phylum Euarchaeota, ANME-1 group of anaerobally methane-oxidizing archaea (in the lower part of the strata), were discovered (Treude et al., 2020). In the biofilms from boreholes drilled in marine gas hydrate pingo approximately 50 km from the south-southwest of South Cape Island (Spitsbergen archipelago), representatives of Methanomicrobia class, ANME-1a and -b groups of phylum Eurarchaeota; representatives of the recently discovered C. methanofastidiosales order, carrying out metallotrophic methanogenesis (Vanwonterghem et al., 2016); representatives of phylum Bathyarchaeota, which is supposedly capable of methanogenesis (Evans et al., 2015); and representatives of phyla Thermoplasmata and Woesearchaeota (Gründer et al., 2019) were discovered. However, the studies on the ancient, deep permafrost on Spitsbergen using the new generation sequencing method have not yet been carried out. The aim of our work was to characterize permafrost coastal marine sediments covering the entire altitude and age range of the ladder of marine terraces in the Barentsburg region using the methods of analyzing the V4 region of the 16S rRNA gene in order to gain an idea of the structure of archaeal communities and corresponding microbiological processes that took place in marine sediments before their freezing for a long time. This work represents the second part of the study on the Archaeal domain; the first part was dedicated to the Bacterial domain.
SAMPLING SITE
This study used permafrost cores from borehole (Bh) 1 (78.02289° N, 14.29845° E, 2.0 m above sea level (MSL)) drilled at the mouth of the Grøn River; Bh 2 (78.09504° N, 14.24096° E, 75.5 m MSL) and 5 (78.09856° N, 14.23299° E, 43.0 m MSL), drilled on the eastern bank of the Isfjord Bay; and Bh 7 (78.04703° N, 14.21962° E, 8.0 m MSL), drilled at Cape Finneset (Fig. 1). These sediments were represented by sands, sandy loams, loams, and clays of coastal–marine genesis (Fig. 2) which accumulated during the middle and late Holocene and then, during a sharp drop in relative sea level, came to the surface, were frozen, and then were covered with a thin cover of continental sediments of various genesis (Forman et al., 2004; Svendsen, Mangerud, 1997). The average annual temperature of Bh 2 (September 25, 2018–August 25, 2019) at a depth of 5.5 m was –2.17°С and a one-time measurement (September 12, 2016) of the Bh 7 temperature at a depth of 12.5 m was –0.87°С (Demidov et al., 2020). Despite the fact that in the modern era these sediments are frozen, samples from depths less than 3 m could have been thawed during the Holocene warming, when the seasonal thawing depth exceeded the modern one. In the case of Bh 1, a short-term rise in sea level might have caused the temporarily thawing of sediments which are now frozen (Solovieva et al., 2018; Salvigsen, Høgvard, 2005). Samples taken for next-generation sequencing were numbered according to Bh numbers and sampling depths (S1-1, S2-4, S5-3, S7-7, S7-9, and S7-12).
Region of the study area in the Spitsbergen archipelago (a); satellite image of a permafrost test site in Barentsburg showing the locations of boreholes 1, 2, 5, and 7 (b); and the precise locations of boreholes 2 and 5 (c) (using maps from https://google.ru/maps and https://toposvalbard.npolar.no).
METHODS
Sampling. Drilling was carried out in August and September 2016 using the UKB 12/25 drilling machine (Vorovskiy Head, Yekaterinburg, Russia). The coring was undertaken without washing and without adding chemical reagents. Thin-walled core pipes with an external diameter of 76 to 112 mm were used. After cleaning the surface of the frozen core segments with a sterile scalpel, the samples were placed in sterile bags (Whirl-Pak®, Nasko, United States) and stored at temperatures –4 to –10°С. On 28 October 2016, they were transported to the laboratory, where they were stored in a freezer at –18°С until analysis in the period from October 2016 to February 2020.
Analysis of carbon monoxide, carbon dioxide, methane and ethylene concentrations. For an analysis of gases, we selected Bh7, since it is the deepest (12 m) and abundant in samples for gas extraction from permafrost. Gas was collected using headspace degassing in 150 mL syringes (Alperin et al., 1985).
Analyses of carbon monoxide, carbon dioxide, methane, ethylene, and ethane concentrations were carried out on a Chromatek-Kristall 5000 chromatograph with packed and capillary columns and a temperature in the column oven by isotherm at 80°C (NRC Kurchatov Institute – IREA, Moscow, Russia). Three detectors were used: PID-1 (for carbon oxides and methane) and PID-2 (for C2-C4 hydrocarbons); the temperature of both was 200°C; RTA was used to determine nitrogen, oxygen, and hydrogen; the temperature was 160°C. Reference gas consumption (helium) was 15 mL/min. The carrier gas was helium (99.9999%); pressure was 25 kPa. Hydrogen consumption was 25 mL/min and air consumption was 500 mL/min. The gas mixtures used were O2/CO/CO2/CH4/N2/He with mole fraction of components (ppm) 2.2/2.2/2.3/2.3/2.6/remainder and 7.8/7.7/7.6/7.7/7.6/remainder respectively, C2H4, C2H2, C2H6, C3H6, C3H8,i-C4H10/n-C4H10,N2 with mole fraction of components (%) 0.00105/0.00097/ 0.00108/0.00104/0.00105/0.00111/0.00107/remainder.
DNA isolation and preparation and sequencing of amplicon libraries. DNA from the samples was isolated using Fast DNA Spin Kit for Soil according to the manufacturer’s method (MP Biomedicals, United States). The concentration was measured using a Qubit 2.0 fluorimeter with a dsDNA HS reagent KIT (InvitrogenTM, United States). Amplicon libraries were created using PCR with universal primers for region V4 in accordance with the previously described methodology (Fadrosh et al., 2014) at the Vernadsky Institute of Microbiology. Primers were selected for the most objective ratio of Bacteria and Archaea domains: 515F (5′-GTGBCAGCMGCCGCGGTAA-3′) (Hugerth et al., 2014) and Pro-mod-805R (5'-GACTACNVGGGTMTCTAATCC-3′) (Merkel et al., 2019). Sequencing was performed on a MiSeq system (Illumina, United States) at Biospark (Moscow, Troitsk, Russia) using a reaction MiSeq Reagent Micro KIT v2 that reads 150 nucleotides from each end. Each sample was read in two technical duplicates, including the reagents and air control sample used to subtract the contaminant sequences from the study sample. A total of 164 843 sequences were obtained.
Bioinformatic and statistical analysis. Demultiplexing, as well as subsequent processing and sequence analysis, were performed using the appropriate scripts in QIIME 2 ver2019.1 software (Bolyen et al., 2019). Operational Taxonomic Units (OTUs) were identified using the SILVAngs 1.4 pipeline (https://ngs.arb-silva. de/silvangs/) and BLAST (http://blast.ncbi.nlm. nih.gov/Blast.cgi) programs. The coverage index of amplicon libraries was calculated by the formula
where n is the number of OTUs represented by one amplicon and N is the total number of amplicons (Good, 1953). The Chao1 index (Chao, 1984) was calculated by the formula
where Sobs is the identified number of phylotypes (OTU), a is the number of phylotypes (OTU) represented by one amplicon, and b is the number of phylotypes represented by two amplicons. The Shannon–Weaver Index (Magurran, 1988) was calculated according to the formula
where pi is the relative abundance of the ith philotype (OTU). The relative abundance of technical replications was combined to indicate the relative content for analysis of the bacterial community using bubble graph diagrams. The analysis of main components and cluster analysis were performed using the Past3 statistical package (Hammer, 2017) using the paired group algorithm (UPGMA) and the Euclidean affinity index.
Deposit at GenBank. The sequence of regions V4 16S rRNA were loaded into the NCBI database as bioproject PRJNA625477. The gene sequences of 16S rRNA isolates were transferred to the NCBI database under the numbers MN599988-MN599993.
RESULTS
Methane and Other Carbon-Containing Gases in Samples of Bh 7 Near Cape Finneset
The methane content in Bh 7 samples ranged from 0.09029 to 0.48315 mL/kg (Table 1). The concentration of carbon dioxide was about 58–98 times higher than the methane concentration and 2.2–5.8 times higher than the carbon monoxide concentration. In the sample S7-12, ethylene and ethane were detected at comparable levels with methane content concentrations (Table 1).
Profiling of the V4 region of 16S rRNA. Genomic DNA was isolated from all samples: in concentrations from 0.1 to 0.8 ng/μL for Bh 1 and 7 samples and about 0.02 ng/μL for Bh 2 and 5 samples. The content of archaeal DNA in the total prokaryotic DNA did not exceed 1% (Table 2).
A total of 44 OTUs of the Archaea domain were revealed (all of them are dominant, ≥1%): 7, 5, 7, 8, 1, and 16 for samples S1-1, S2-4, S5-3, S7-7, S7-9, and S7-12, respectively.
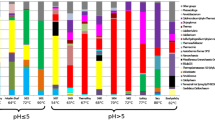
It was represented by the phyla Euryarchaeota (5.6–100%), Bathyarchaeota (22.1–81.0%), Thaumarchaeota (5.6–54.0%), Asgardarchaea (6.3–79.2%), and Woesearchaeota (5.6–52.2%) (Figs. 3, 4). Methanogenic Archaea were found in samples S1-1, S2-4, S5-3, and S7-7 and were represented by four genera: Methanocalculus, Methanobacterium, Methanobrevibacter, and Methanothermobacter. In phylum Euryarchaeota, the Methanobacteria class was the most diverse in the studied samples. Its representatives were found in all boreholes. Therefore, in the S1-1 sample, 32.7% was the genus Methanobacterium and 15.4% was the genus Methanothermobacter. This is the only sample out of six in which the proportion of methanogenic Archaea exceeded 20.0%, amounting to 46%. This genus was also discovered as the only representative of methanogenic Archaea in sample S7-7. The genus Methanothermobacter was also found in sample S2-4. The genus Methanobrevibacter was the only identified representative of methanogens in sample S5-3. The Methanomicrobia class is represented in our samples by the genus Methanocalculus and was found only in sample S2-4, where it amounted to 18% of the archaeal community. The Thermoplasmata class was represented by the SG8-5 group and was found only in sample S7-12 (2.8%). Other members of the Methanosarcinales order belonging to Archaea involved in anaerobic methane oxidation were found in Bh 7: groups ANME-2a, -2b (S7-9 100% and S7-12 30.3%), and -2d (Candidatus Methanoperedens) (S7-7 36.1%). The phylum Bathyarchaeota was found in samples S2-4 (80.6%) and S7-12 (17.8%). Representatives of ammonium-oxidizing Archaea of the order of Nitrososphaerales were present in samples S1-1 (50%), S7-7 (3.6%), and S7-12 (5.6%) and those of the order Nitrosopumilales in samples S1-1 (3.9%), S5-3 (10.0%), and S7-12 (2.8%). Inside the Asgardarchaeota superfila, phyla Lokiarchaeota (S5-3 22.2%, S7-7 6.8%, and S7-12 12.2%), Candidatus Thorarchaeota (S5-3 27.7% and S7-12 29.3%), and unclassified Asgardarchaeota by the 16S rRNA gene (S5-3 33.3%) were discovered. Representatives of Lokiarchaeota in sample S5-3, as well as unclassified Asgardarchaeota in samples S5-3 and S7-12, were identified thanks to the phylogenetic tree on which they appeared on common branches by representatives of Lokiarchaeota from samples S7-7 and S7-12. Woesearchaeota phylum was found inside the DPANN superfila (S5-3 3.9%, S7-7 52.2%, and S7-12 9.4%).
DISCUSSION
The content of carbon-containing gases in permafrost of Cape Finneset. The presence of ethylene and ethane in the sample S7-12 can be explained both by the natural composition of the gas phase rising through the permafrost to the surface from the bedrock, and by the anaerobic processes of ethylene splitting by archaea to ethane and methane described earlier (Koene-Cottaar and Schraa, 1997; Xie et al., 2017). Ethylene and acetylene are known to inhibit melanogenesis in marine sediments (Oremland, Taylor, 1975).
Characteristics of amplicon libraries (the characteristics are also given in the first part of the study about Bacteria domain (Karaevskaya et al., 2021)). Sample S1-1 was characterized by the greatest variety; its Shannon–Weaver index was 5.0 ± 0.1 and the Chao1 index was 8540 ± 1156, while the library coverage was 96.3 ± 4.3% (Table 2). The samples of Bh 7 were less diverse, for which the Shannon–Weaver index was 4.1 ± 0.6 to 4.7 ± 0.1 and Chao1 was 452 ± 169 to 1242 ± 83. Sample S7–9, with the lowest diversity indices and low library coverage, 53.6 ± 9.6%, was knocked out of the general picture when compared with the other two samples, S7–7 and S7–12 (96.8 ± 0.4 – 98.2 ± 0.1%). This phenomenon may exist due to unfavorable conditions for the conservation or isolation of DNA in this sample, which differs from others in its high content of plant residues. Samples S2-4 (2.5 ± 0.1) and S5-3 (2.8 ± 0.1) were characterized by the lowest diversity by the Shannon–Weaver index, while the library coverage in them was very good and amounted to 98.3 ± 1.1 and 98.7 ± 0.1, respectively. This may be due to the very low output concentrations of DNA, and this, in turn, to their older age when compared to the samples of the lower marine terraces exposed by Bh 1 and 7.
Taxonomy of OTUs of the Archaea domain. Here we considered the taxonomy of OTUs comprising the Archaea domain (Figs. 3, 4; Table 3), excluding those that make up less than 5% of all archaeal amplicons, which is below the error level of the method, marked with an asterisk in Table 3.
The OTUs of the phylum Euryarchaeota of the genus Methanobacterium revealed in samples S1-1 and S7-7 were found to be related to the type strain of hydrogenotrophic freshwater methanogenic archaea M. lacus 17A1T (Borrel et al., 2012). The genus Methanocalculus unites halotolerant species and was represented in the S2-4 OTU sample, related to the type strain of M. pumilus MHT-1T hydrogenotrophic archaea (Mori et al., 2000), which is resistant to high concentrations of several heavy metals, such as CdCl2 and CuSO4 (Fargeau et al., 2019).
OTUs of marine anaerobic methane-oxidizing archaea ANME-2a, -2b (Beuling et al., 2019) were found in samples S7-9 and S7-12, and freshwater ones—ANME-2d (Kurth et al., 2019)—were found in sample S7-7.
The phylum Bathyarchaeota, found in samples S2-4 and S7-12, is an archaea that occurs inter alia in Na2+-saline habitats in symbiosis with the class Methanomicrobia. In the Bathyarchaeota metagenome, genes encoding the methyl-coenzyme M-reductase (MKP) complex were found (Evans et al., 2015; Kallistova et al., 2017), participating in methanogenesis and anaerobic oxidation of methane (Thauer, 1998; Shima and Thauer, 2005).
The family Nitrososphaeraceae (found in samples S1-1, S7-7, and S7-12) and the family Nitrosopumilaceae (genus Candidatus Nitrosopumilus) (found in samples S1-1, S5-3, and S7-12) of the phylum Thaumarchaeota are anaerobic ammonium-oxidizing archaea; the former is found in soil and in freshwater and marine ecosystems and the latter is found in deep-sea marine ecosystems (Park et al., 2012).
Archaea of the superphylum Asgardarchaeota were found only in wells 5 and 7. It is assumed that the archaea of this superphylum are obligate anaerobes; can have autotrophic, heterotrophic, and phototrophic types of nutrition; and can also participate in the reduction of iron and manganese in the presence of methanogenic archaea and sulfate-reducing bacteria (Jørgensen et al., 2013). The phylum Lokiarchaeota, found in samples S5-3, S7-7, and S7-12, was found in hydrothermal vents in Japan (Takai, Horikoshi, 1999), bottom marine sediments of the Atlantic Ocean (Vetriani et al., 1999), and in terrestrial anaerobic/microaerophilic aquatic ecosystems (Sorensen, Teske, 2006) and deep-sea hydrotherms of the North Atlantic on the Knipovich Ridge (Jørgensen et al., 2012; 2013). The phylum Thorarchaeota found in samples S5-3 and S7-12 was found in lacustrine, mangrove, and hydrothermal marine bottom sediments; presumably they are capable of transforming proteins and hydrocarbons, as well as acetogenesis and sulfate reduction (Seitz et al., 2016). All representatives of Asgardarchaeota closest to the OTUs obtained by us were found in marine ecosystems. It should be noted that the identified OTUs related to the phylum Asgardarchaeota practically did not intersect with each other in different samples, with the exception of AS5-3-2 and AS7-12-3, AS5-3-3 and AS7-12-4, which indicates their varied composition depending on the location and time of sedimentation (Fig. 4).
Genetic relationship of samples. Based on the analysis of the principal components, according to the taxonomic diversity of the Archaea, the samples most similar each other were S7-7 and S7-12, as well as S1-1. The isolation of sample S7-9 in Bh 7 can be explained by the increased content of fragments of plant material in it, which is most likely due to sedimentation conditions that are different from the other two samples. By the presence of superphylum, Asgardarchaeota made it possible to combine samples from Bh 5 (S5-3) and 7 (S7-7 and S7-12) and phylum Thorarchaeota (S5-3 and S7-12) (Fig. 5b). Another cluster is formed by samples S5-3 and S2-4 (Fig. 5b). The archaeal diversity of sample S2-4 is the most different of all samples. This may be due to the different conditions of sediment formation and the different age of the sediments of Bh 1, 3–7, and Bh 2.
PCA of phyla content for two technical repetitions of domain Archaea (a). Vectors show selected taxonomical factors that are mainly responsible for the variance between samples. Cluster analysis of average abundance values using the paired group algorithm (UPGMA) and the Euclidean affinity index (b).
Most likely, the archaeal communities of the layers corresponding to samples S5-3 and S7-9 were formed in strictly anaerobic conditions favorable for the processes of the anaerobic oxidation of methane and ammonium. Whereas the communities of the layers corresponding to samples S1-1, S2-4, S7-7, and S7-12 formed in both aerobic and anaerobic conditions, methanogenesis was added to other processes. The reason for the appearance of methanogenesis may be connected not so much with aerobic conditions arising in the process of the evolution of the sediments, but with the influence of saturated carbon dioxide in groundwaters on land ecosystems (Demidov et al., 2020).
CONCLUSIONS
A comparative study of Spitsbergen permafrost representing marine terraces located in different sites of the east bank of Grønfjord was carried out for the first time, using mutually complementary methods: profiling the V4 region of the 16S rRNA gene for the Archaeal domain.
Characterized by the predominance of archaeal phyla Euryarchaeota, Bathyarchaeota, Thaumarchaeota, and Asgardarchaea, the communities were similar to the communities of modern coastal and marine coastal sediments. Presumably, they were formed mainly under anaerobic, but also under mixed aerobic–anaerobic conditions.
It had been previously established that the isotopic composition of methane and carbon dioxide in Bh 7, as well as the predominance of methane oxidizing of the ANME-2a, -2b and -2d groups over methanogenic of ther Methanobacteria order, suggests that these deposits could have been formed under conditions of gas intake from Tertiary rocks.
The presence of methane, ethylene, and ethane in the sample from Bh 7 at a depth of 11.7 m, as well as the structure of archaeal communities, suggests the presence of microbiological processes of anaerobic oxidation of methane in this layer before freezing, probably coming from Tertiary rocks.
Thus, combining the results from domains Bacteria and Archaea, the functional role of the studied prokaryotic communities seems to be reduced to heterotrophic psychrophilic activity in all samples; methanogenesis in samples S1-1 and S2-4; the anaerobic oxidation of methane by bacteria of the genus Methylobacter in sample S1-1; the anaerobic oxidation of methane by archaeal groups ANME-2a, -2b, and -2d in samples S7-7, S7-9, and S7-12; the sulfate-reducing activity of bacteria of the phyla Firmicutes and Nitrospirae in samples S2-4, S7-7, and S7-9; and anaerobic ammonium oxidation by Archaea of the phylum Thaumarchaeota in samples S1-1, S5-3, S7-7, and S7-12. Also, all samples assume the presence of microbiological processes of conversion of hydrocarbons.
The results obtained here and in our previous article (Karaevskaya et al., 2021) give motivation for further research in the field of the phylogeny of prokaryotic communities of Spitsbengen’s permafrost, in particular that of marine archaea of the recently discovered superphylum Asgardarchaeota. Their metabolism, as well as the reasons for their quite wide variety in this permafrost, is of great interest. Due to the very low fraction of the Archea domain in the prokaryotic communities under study, approaches with primers for the gene of interest of certain archaeal groups must be used for molecular genetic studies in this area.
REFERENCES
Alain, K., Holler, T., Musat, F., Elvert, M., Treude, T., and Krüger, M., Microbiological investigation of methane- and hydrocarbon-discharging mud volcanoes in the Carpathian Mountains, Romania, Environ. Microbiol., 2006, vol. 8, no. 4, pp. 574–590. https://doi.org/10.1111/j.1462-2920.2005.00922.x
Alperin, M.J. and Reeburgh, W.S., Inhibition experiments on anaerobic methane oxidation, Appl. Environ. Microbiol., 1985, vol. 50, no. 4, pp. 940–945. https://doi.org/10.1128/aem.50.4.940-945.1985
Amaral-Zettler, L.A., Rocca, J.D., Lamontagne, M.G., Dennett, M.R., and Gast, R.J., Changes in microbial community structure in the wake of Hurricanes Katrina and Rita, Environ. Sci. Technol., 2008, vol. 42, no. 24, pp. 9072–9078. https://doi.org/10.1021/es801904z
Beulig, F., Røy, H., McGlynn, S.E., and Jørgensen, B.B., Cryptic CH4 cycling in the sulfate–methane transition of marine sediments apparently mediated by ANME-1 archaea, ISME J., 2019, vol. 13, no. 2, pp. 250–262. https://doi.org/10.1038/s41396-018-0273-z
Bolyen, E., Rideout, J.R., Dillon, M.R., Bokulich, N.A., Abnet, C.C., Al-Ghalith, G.A., Alexander, H., Alm, E.J., Arumugam, M., Asnicar, F., Bai, Y., Bisanz, J.E., Bittinger, K., Brejnrod, A., Brislawn, C.J., et al., Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2, Nat. Biotechnol., 2019, vol. 37, no. 8, pp. 852–857; Erratum: Nat. Biotechnol., 2019, vol. 37, no. 9, p. 1091. https://doi.org/10.1038/s41587-019-0209-9
Borrel, G., Joblin, K., Guedon, A., Colombet, J., Tardy, V., Lehours, A.C., and Fonty, G., Methanobacterium lacus sp. nov., isolated from the profundal sediment of a freshwater meromictic lake, Int. J. Syst. Evol. Microbiol., 2012, vol. 62, no. 7, pp. 1625–1629. https://doi.org/10.1099/ijs.0.034538-0
Bueno de Mesquita, C.P.B., Schmidt, S.K., and Suding, K.N., Litter-driven feedbacks influence plant colonization of a high elevation early successional ecosystem, Plant Soil, 2019, vol. 444, no. 1, pp. 71–85. https://doi.org/10.1007/s11104-019-04242-3
Chao, A., Nonparametric estimation of the number of classes in a population, Scand. J. Stat.,1984, vol. 11, pp. 265–270.
Chernov, R.A. and Murav’ev, A.Y., Contemporary changes in the area of glaciers in the western part of the Nordenskjold Land (Svalbard), Led Sneg, 2018, vol. 58, no. 4, pp. 462–472.
Cho, H., Hyun, J.H., You, O.R., Kim, M., Kim, S.H., Choi, D.L., Green, S.J., and Kostka, J.E., Microbial community structure associated with biogeochemical processes in the sulfate–methane transition zone (SMTZ) of gas-hydrate-bearing sediment of the Ulleung Basin, East Sea, Geomicrobiol. J., 2017, vol. 34, no. 3, pp. 207–219. https://doi.org/10.1080/01490451.2016.1159767
Conrad, R., Chan, O.C., Claus, P., and Casper, P., Characterization of methanogenic Archaea and stable isotope fractionation during methane production in the profundal sediment of an oligotrophic lake (Lake Stechlin, Germany), Limnol. Oceanogr., 2007, vol. 52, no. 4, pp. 1393–1406. https://doi.org/10.4319/lo.2007.52.4.1393
Demidov, N.E., Verkulich, S.R., Karaevskaya, E.S., Nikulina, A.L., and Savatyugin, L.M., First results of permafrost monitoring on the cryospheric site of the Russian Scientific Center on Spitsbergen (RSCS), Probl. Arkt. Antarkt., 2016, vol. 1, no. 4, pp. 67–79.
Demidov, N., Wetterich, S., Verkulich, S., Ekaykin, A., Meyer, H., Anisimov, M., Schirrmeister, L., Demidov, V., and Hodson, A.J., Geochemical signatures of pingo ice and its origin in Grøndalen, West Spitsbergen, Cryosphere, 2019, vol. 13, pp. 3155–3169. https://doi.org/10.5194/tc-13-3155-2019
Demidov, N.E., Borisik, A.L., Verkulich, S.R., Wetterich, S., Gunar, A.Yu., Demidov, V.E., Zheltenkova, N.V., Koshurnikov, A.V., Mikhailova, V.M., Nikulina, A.L., Novikov, A.L., Savatyugin, L.M., Sirotkin, A.N., Terekhov, A.V., Ugrumov, Yu.V., and Schirrmeister, L., Geocryological and Hydrogeological Conditions of the Western Part of Nordenskiold Land (Spitsbergen Archipelago), Izv., Atmos. Ocean. Phys., 2020, vol. 56, no. 11, pp. 1376–1400. https://doi.org/10.1134/S000143382011002X
De Wever, A., Muylaert, K., Van der Gucht, K., Pirlot, S., Cocquyt, C., Descy, J.P., Plisnier, P.-D., and Vyverman, W., Bacterial community composition in Lake Tanganyika: Vertical and horizontal heterogeneity, App-l. Environ. Microbiol., 2005, vol. 71, no. 9, pp. 5029–5037. https://doi.org/10.1128/AEM.71.9.5029-5037.2005
Doerfert, S.N., Reichlen, M., Iyer, P., Wang, M., and Ferry, J.G., Methanolobus zinderi sp. nov., a methylotrophic methanogen isolated from a deep subsurface coal seam, Int. J. Syst. Evol. Microbiol., vol. 59, no. 5, pp. 1064–1069. https://doi.org/10.1099/ijs.0.003772-0
Evans, P.N., Parks, D.H., Chadwick, G.L., Robbins, S.J., Orphan, V.J., Golding, S.D., and Tyson, G.W., Methane metabolism in the archaeal phylum Bathyarchaeota revealed by genome-centric metagenomics, Science, 2015, vol. 350, no. 6259, pp. 434–438. https://doi.org/10.1126/science.aac7745
Fadrosh, D.W., Ma, B., Gajer, P., Sengamalay, N., Ott, S., Brotman, R.M., and Ravel, J., An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform, Microbiome, 2014, vol. 2, no. 1, p. 6. https://doi.org/10.1186/2049-2618-2-6
Fardeau, M.-L., Cayol, J.-L., and Ollivier, B., Methanocalculus, in Bergey’s Manual of Systematics of Archaea and Bacteria (BMSAB), Hoboken, N.J.: Wiley, 2019. https://hal.archives-ouvertes.fr/hal-02883889.
Forman, S.L., Lubinski, D.J., Ingolfsson, O., Zeeberg, J.J., Snyder, J.A., Siegert, M.J., and Matishov, G.G., A review of postglacial emergence on Svalbard, Franz Josef Land and Novaya Zemlya, northern Eurasia, Quat. Sci. Rev., 2004, vol. 23, pp. 1391–1434. https://doi.org/10.1016/j.quascirev.2003.12.007
Good, I.J., The population frequencies of species and the estimation of population parameters, Biometrika, 1953, vol. 40, pp. 237–264. https://doi.org/10.1093/biomet/40.3-4.237
Hamdan, L.J., Coffin, R.B., Sikaroodi, M., Greinert, J., Treude, T., and Gillevet, P.M., Ocean currents shape the microbiome of Arctic marine sediments, ISME J., 2013, vol. 7, no. 4, pp. 685–696.
Hammer, Ø., Harper, D.A.T., and Ryan, P.D., PAST: Paleontological statistics software package for education, 2017.
Hernández, E.A., Piquet, A.M.T., Lopez, J.L., Buma, A.G., and Mac Cormack, W.P., Marine archaeal community structure from Potter Cove, Antarctica: High temporal and spatial dominance of the phylum Thaumarchaeota, Polar Biol., 2015, vol. 38, no. 2, pp. 117–130. https://doi.org/10.1007/s00300-014-1569-8
Hoshino, T. and Inagaki, F., A comparative study of microbial diversity and community structure in marine sediments using poly (A) tailing and reverse transcription-PCR, Front. Microbiol., 2013, vol. 4, p. 160. https://doi.org/10.3389/fmicb.2013.00160
Hugerth, L.W., Wefer, H.A., Lundin, S., Jakobsson, H.E., Lindberg, M., Rodin, S., Engstrand, L., and Andersson, A.F., DegePrime, a program for degenerate primer design for broad-taxonomic-range PCR in microbial ecology studies, Appl. Environ. Microbiol., 2014, vol. 80, no. 16, pp. 5116–5123. https://doi.org/10.1128/AEM.01403-14
Humlum, O., Instanes, A. and Sollid, J.L., Permafrost in Svalbard: A review of research history, climatic background and engineering challenges, Polar Res., 2003, vol. 22, pp. 191–215. https://doi.org/10.1111/j.1751-8369.2003.tb00107.x
Imachi, H., Nobu, M.K., Nakahara, N., Morono, Y., Ogawara, M., Takaki, Y., Takano, Y., Uematsu, K., Ikuta, T., Ito, M., Matsui, Y., Miyazaki, M., Murata, K.M., Saito, Y., Sakai, S., et al., Isolation of an archaeon at the prokaryote–eukaryote interface, Nature, 2000, vol. 577, no. 7791, pp. 519–525. https://doi.org/10.1038/s41586-019-1916-6
Joblin, K.N., Naylor, G.E., and Williams, A.G., Effect of Methanobrevibacter smithii on xylanolytic activity of anaerobic ruminal fungi, Appl. Environ. Microbiol., 1990, vol. 56, no. 8, pp. 2287–2295. https://doi.org/10.1128/aem.56.8.2287-2295.1990
Jarvis, G.N., Strömpl, C., Burgess, D.M., Skillman, L.C., Moore, E.R., and Joblin, K.N., Isolation and identification of ruminal methanogens from grazing cattle, Curr. Microbiol., 2000, vol. 40, no. 5, pp. 327–332. https://doi.org/10.1007/s002849910065
Jørgensen, S.L., Hannisdal, B., Lanzén, A., Baumberger, T., Flesland, K., Fonseca, R., Øvreås, L., Steen, I.H., Thorseth, I.H., Pedersen, R.B., and Schleper, C., Correlating microbial community profiles with geochemical data in highly stratified sediments from the Arctic Mid-Ocean Ridge, Proc. Natl. Acad. Sci., 2012, vol. 109, no. 42, pp. E2846–E2855. https://doi.org/10.1073/pnas.1207574109
Jørgensen, S.L., Thorseth, I.H., Pedersen, R.B., Baumberger, T., and Schleper, C., Quantitative and phylogenetic study of the Deep Sea Archaeal Group in sediments of the Arctic mid-ocean spreading ridge, Front. Microbiol., 2013, vol. 4, p. 299. https://doi.org/10.3389/fmicb.2013.00299
Jørgensen, B.B., Beulig, F., Egger, M., Petro, C., Scholze, C., and Røy, H., Organoclastic sulfate reduction in the sulfate-methane transition of marine sediments, Geochim. Cosmochim. Acta, 2019, vol. 254, pp. 231–245. https://doi.org/10.1016/j.gca.2019.03.016
Kallistova, A.Y., Merkel, A.Y., Tarnovetskii, I.Y., and Pimenov, N.V., Methane formation and oxidation by prokaryotes, Microbiology, 2017, vol. 86, no. 6, pp. 671–691. https://doi.org/10.1134/S0026261717060091
Karaevskaya, E.S., Demchenko, L.S., Demidov, N.E., Rivkina, E.M., Bulat, S.A., and Gilichinsky, D.A., Archaeal diversity in permafrost sediments of Bunger Hills Oasis and King George Island (Antarctica) according to the 16S rRNA gene sequencing, Microbiology, 2014, vol. 83, no. 4, pp. 398–406. https://doi.org/10.1134/S0026261714040092
Karaevskaya, E.S., Demidov, N.E., Kazantsev, V.S., Elizarov, I.M., Kaloshin, A.G., Petrov, A.P., Karlov, D.S., Schirrmeister, L., Belov, A.A., and Wetterich, S., Bacterial communities of frozen quaternary sediments of marine origin on the coast of Western Spitsbergen, Geofiz. Protsessy Biosfera, 2021, vol. 20, no. 2, pp. 75–98. https://doi.org/10.21455/GPB2021.2-5
Kato, S., Ikehata, K., Shibuya, T., Urabe, T., Ohkuma, M., and Yamagishi, A., Potential for biogeochemical cycling of sulfur, iron and carbon within massive sulfide deposits below the seafloor, Environ. Microbiol., 2015, vol. 17, no. 5, pp. 1817–1835. https://doi.org/10.1111/1462-2920.12648
Kemnitz, D., Kolb, S., and Conrad, R., Phenotypic characterization of Rice Cluster III archaea without prior isolation by applying quantitative polymerase chain reaction to an enrichment culture, Environ. Microbiol., 2005, vol. 7, no. 4, pp. 553–565. https://doi.org/10.1111/j.1462-2920.2005.00723.x
Koene-Cottaar F.H. and Schraa G. Anaerobic reduction of ethene to ethane in an enrichment culture, FEMS Microbiol. Ecol. 1998, vol. 25, no. 3, pp. 251–256.
Kumar, S., Stecher, G., and Tamura, K., MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for bigger datasets, Mol. Biol. Evol., 2016, vol. 33, no. 7, pp. 1870–1874. https://doi.org/10.1093/molbev/msw054
Kurth, J.M., Smit, N.T., Berger, S., Schouten, S., Jetten, M.S.M., and Welte, C.U. Anaerobic methanotrophic archaea of the ANME-2d clade feature lipid composition that differs from other ANME archaea, FEMS Microbiol. Ecol., 2019, vol. 95, no. 7, p. fiz082. https://doi.org/10.1093/femsec/fiz082
Lai, M., Laonitisatr, W., Chen, Y., Ding, J., Wu, C., and Lai, S., Microbial communities of spring pits in Jing-Mei River at the southeastern Taipei basin, in Proceedings from the 2010 AGU Western Pacific Geophysics Meeting, 2010.
Lane, D.J., 16S/23S rRNA sequencing, in Nucleic Acid Techniques in Bacterial Systematics, Stackebrandt, E. and Goodfellow, M., Eds., Chichester, UK: Wiley, 1991, pp. 115– 175.
Lynch, R.C., King, A.J., Farías, M.E., Sowell, P., Vitry, C., and Schmidt, S.K., The potential for microbial life in the highest-elevation (> 6000 masl) mineral soils of the Atacama region, J. Geophys. Res.: Biogeosci., 2012, vol. 117, no. G2. https://doi.org/10.1029/2012JG001961
Magurran, A.E., Ecological Diversity and Its Measurement, Princeton Univ. Press, 1988; Moscow: Mir, 1992.
Mangerud, J., Radiocarbon dating of marine shells, including a discussion of apparent age of recent shells from Norway, Boreas, 1972, vol. 1, no. 2, pp. 143–172.
Merkel, A.Yu., Tarnovetskii, I.Yu., Podosokorskaya, O.A., and Toshchakov, S.V., Analysis of 16S rRNA primer systems for profiling of thermophilic microbial communities, Microbiology, 2019, vol. 13, no. 6, pp. 671–680. https://doi.org/10.1134/S0026261719060110
Miller, T.L., Wolin, M.J., de Macario, E.C., and Macario, A.J., Isolation of Methanobrevibacter smithii from human feces, Appl. Environ. Microbiol., 1982, vol. 43, no. 1, pp. 227–232. https://doi.org/10.1128/aem.43.1.227-232.1982
Mitzscherling, J., Horn, F., Winterfeld, M., Mahler, L., Kallmeyer, J., Overduin, P, pp., Schirrmeister, L., Winkel, M., Grigoriev, M.N., Wagner, D., and Liebner, S., Microbial community composition and abundance after millennia of submarine permafrost warming, Biogeosciences, 2019, vol.16, pp. 3941–3958. https://doi.org/10.5194/bg-16-3941-2019
Mori, K., Yamamoto, H., Kamagata, Y., Hatsu, M., and Takamizawa, K., Methanocalculus pumilus sp. nov., a heavy-metal-tolerant methanogen isolated from a waste-disposal site, Int. J. Syst. Evol. Microbiol., 2000, vol. 50, no. 5, pp. 1723–1729. https://doi.org/10.1099/00207713-50-5-1723
Okita, N., Hoaki, T., Suzuki, S., Hatamoto, M., Characteristics of microbial community structure at the seafloor surface of the Nankai Trough, J. Pure Appl. Microbiol., 2019, vol. 13, pp. 1917–1928. https://doi.org/10.22207/JPAM.13.4.04
Oremland R.S. and Taylor B.F., Inhibition of methanogenesis in marine sediments by acetylene and ethylene: Validity of the acetylene reduction assay for anaerobic microcosms, Applied Microbiology, 1975, vol. 30, no. 4. pp. 707–709.
Oshiki, M., Segawa, T., and Ishii, S., Nitrogen cycle evaluation (nice) chip for simultaneous analysis of multiple n cycle-associated genes, Appl. Environ. Microbiol., 2018, vol. 84, no. 8. https://doi.org/10.1128/AEM.02615-17
Pan, J., Chen,Y., Wang, Y., Zhou, Z., and Li, M., Vertical distribution of Bathyarchaeotal communities in mangrove wetlands suggests distinct niche preference of Bathyarchaeota subgroup 6, Microb. Ecol., 2019, vol. 77, no. 2, pp. 417–428. https://doi.org/10.1007/s00248-018-1309-7
Park, S.J., Kim, J.G., Jung, M.Y., Kim, S.J., Cha, I.T., Ghai, R., Martín-Cuadrado, A.-B., Rodrígues-Valera, F., and Rhee, S.K., Draft genome sequence of an ammonia-oxidizing archaeon,“Candidatus Nitrosopumilus sediminis” AR2, from Svalbard in the Arctic Circle, J. Bacteriol., 2012, vol. 194, no. 24, pp. 6948–6949.
Pires, A.C., Cleary, D.F., Almeida, A., Cunha, Â., Dealtry, S., Mendonça-Hagler, L.C., Smalla, K., and Gomes, N.N.C., Denaturing gradient gel electrophoresis and barcoded pyrosequencing reveal unprecedented archaeal diversity in mangrove sediment and rhizosphere samples, Appl. Environ. Microbiol., 2012, vol. 78, no. 16, pp. 5520–5528. https://doi.org/10.1128/AEM.00386-12
Probst, A.J. and Moissl-Eichinger, C. “Altiarchaeales”: Uncultivated archaea from the subsurface, Life, 2015, vol. 5, no. 2, pp. 1381–1395. https://doi.org/10.3390/life5021381
Qin, W., Heal, K.R., Ramdasi, R., Kobelt, J.N., Martens-Habbena, W., Bertagnolli, A.D., Amin, Sh.A., Walker, Ch.B., Urakawa, H., Könneke, M., Moffett, J.W., Armbrust, E.V., Ingalls, A.E., Jensen, G.J., Stahl, D.A., and Devol, A.H., Nitrosopumilus maritimus gen. nov., sp. nov., Nitrosopumilus cobalaminigenes sp. nov., Nitrosopumilus oxyclinae sp. nov., and Nitrosopumilus ureiphilus sp. nov., four marine ammonia-oxidizing archaea of the phylum Thaumarchaeota, Int. J. Syst. Evol. Microbiol., 2017, vol. 67, no. 12, pp. 5067–5079. https://doi.org/10.1099/ijsem.0.002416
Roussel, E.G., Sauvadet, A.L., Chaduteau, C., Fouquet, Y., Charlou, J.L., Prieur, D., and Cambon Bonavita, M.A., Archaeal communities associated with shallow to deep subseafloor sediments of the New Caledonia Basin, Environ. Microbiol., 2009, vol. 11, no. 9, pp. 2446–2462. https://doi.org/10.1111/j.1462-2920.2009.01976.x
Salvigsen, O. and Høgvard, K., Glacial history, Holocene shoreline displacement and palaeoclimate based on radiocarbon ages in the area of Bockfjorden, north-western Spitsbergen, Svalbard, Polar Res., 2005, vol. 25, no. 1, pp. 15–24. https://doi.org/10.1111/j.1751-8369.2006.tb00147.x
Seidel, M., Graue, J., Engelen, B., Köster, J., Sass, H., and Rullkötter, J., Advection and diffusion determine vertical distribution of microbial communities in intertidal sediments as revealed by combined biogeochemical and molecular biological analysis, Org. Geochem., 2012, vol. 52, pp. 114–129. https://doi.org/10.1016/j.orggeochem.2012.08.015
Seitz, K.W., Lazar, C.S., Hinrichs, K.U., Teske, A.P., and Baker, B.J., Genomic reconstruction of a novel, deeply branched sediment archaeal phylum with pathways for acetogenesis and sulfur reduction, ISME J., 2016, vol. 10, no. 7, pp. 1696–1705. https://doi.org/10.1038/ismej.2015.233
Shima, S. and Thauer, R.K., Methyl-coenzyme M reductase and the anaerobic oxidation of methane in methanotrophic Archaea, Curr. Opin. Microbiol., 2005, vol. 8, no. 6, pp. 643–648. https://doi.org/10.1016/j.mib.2005.10.002
Smith, D.R., Doucette-Stamm, L.A., Deloughery, C., Lee, H., Dubois, J., Aldredge, T., Bashirzadeh, R., Blakely, D., Cook, R., Gilbert, K., Harrison, D., Hoang, L., Keagle, P., Lumm, W., Pothier B., et al., Complete genome sequence of Methanobacterium thermoautotrophicum Delta H: Functional analysis and comparative genomics, J. Bacteriol., 1997, vol. 179, pp. 7135–7155. https://doi.org/10.1128/jb.179.22.7135-7155.1997
Solov’eva, D.A., Savel’eva, L.A., Verkulich, S.R., and Zazovskaya, E.P., Postglacial changes in the natural environment in the area of the Barentsburg mine (West Spitsbergen Island), in Theory and Methods of Polar Science: Proceedings of International Youth Scientific Conference on the Polar Geodesy, Glaciology, Hydrology and Geophysics, St. Petersburg, 2018, pp. 213–222.
Sørensen, K.B. and Teske, A., Stratified communities of active archaea in deep marine subsurface sediments, Appl. Environ. Microbiol., 2006, vol. 72, no. 7, pp. 4596–4603. https://doi.org/10.1128/AEM.00562-06
Srinivas, T.N.R., Rao, S.N., Reddy, P.V.V., Pratibha, M.S., Sailaja, B., Kavya, B., and Shivaji, S., Bacterial diversity and bioprospecting for cold-active lipases, amylases and proteases, from bacteria of Kongsfjorden and Ny-Ålesund, Svalbard, Arctic, Curr. Microbiol., 2009, vol. 59, no. 5, pp. 537–547. https://doi.org/10.1007/s00284-009-9473-0
Stauffert, M., Duran, R., Gassie, C., and Cravo-Laureau, C., Response of archaeal communities to oil spill in bioturbated mudflat sediments, Microb. Ecol., 2014, vol. 67, no. 1, pp. 108–119. https://doi.org/10.1007/s00248-013-0288-y
Stieglmeier, M., Klingl, A., Alves, R.J., Rittmann, S.K.M., Melcher, M., Leisch, N., and Schleper, C., Nitrososphaera viennensis gen. nov., sp. nov., an aerobic and mesophilic, ammonia-oxidizing archaeon from soil and a member of the archaeal phylum Thaumarchaeota, Int. J. Syst. Evol. Microbiol., 2014, vol. 64, no. 8, pp. 2738–2752. https://doi.org/10.1099/ijs.0.063172-0
Svendsen, J.I. and Mangerud, J., Holocene glacial and climatic variations on Spitsbergen, Svalbard, Holocene, 1997, vol. 7, no. 1, pp. 45–57. https://doi.org/10.1177/095968369700700105
Takai, K. and Horikoshi, K., Genetic diversity of Archaea in deep-sea hydrothermal vent environments, Genetics, 1999, vol. 152, no. 4, pp. 1285–1297.
Tamura, K. and Nei, M., Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees, Mol. Biol. Evol., 1993, vol. 10, pp. 512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Teske, A., Durbin, A., Ziervogel, K., Cox, C., and Arnosti, C., Microbial community composition and function in permanently cold seawater and sediments from an Arctic fjord of Svalbard, Appl. Environ. Microbiol., 2011, vol. 77, no. 6, pp. 2008–2018. https://doi.org/10.1128/AEM.01507-10
Thauer, R.K., Biochemistry of methanogenesis: A tribute to Marjory Stephenson: 1998 Marjory Stephenson Prize lecture, Microbiology, 1998, vol. 144, no. 9, pp. 2377–2406. https://doi.org/10.1099/00221287-144-9-2377
Timmers, P.H., Suarez-Zuluaga, D.A., van Rossem, M., Diender, M., Stams, A.J., Plugge, C.M., Anaerobic oxidation of methane associated with sulfate reduction in a natural freshwater gas source, ISME J., 2016, vol. 10, no. 6, pp. 1400–1412.
Treude, T., Krause, S., Steinle, L., Burwicz, E., Hamdan, L.J., Niemann, H., Feseker, T., Liebertau, V., Krastel, S., and Berndt, C., Biogeochemical consequences of nonvertical methane transport in sediment offshore northwestern Svalbard, J. Geophys. Res.: Biogeosci., 2020, vol. 125, no. 3, p. e2019JG005371. https://doi.org/10.1029/2019JG005371
Vetriani, C., Jannasch, H.W., MacGregor, B.J., Stahl, D.A., and Reysenbach, A.L., Population structure and phylogenetic characterization of marine benthic Archaea in deep-sea sediments, Appl. Environ. Microbiol., 1999, vol. 65, no. 10, pp. 4375–4384. https://doi.org/10.1128/AEM.65.10.4375-4384.1999
Webster, G., Sass, H., Cragg, B.A., Gorra, R., Knab, N.J., Green, C.J., Mathes, F., Fry, J.C., Weightman, A.J., and Parkes, R.J., Enrichment and cultivation of prokaryotes associated with the sulphate–methane transition zone of diffusion-controlled sediments of Aarhus Bay, Denmark, under heterotrophic conditions, FEMS Microbiol. Ecol., 2011, vol. 77, no. 2, pp. 248–263. https://doi.org/10.1111/j.1574-6941.2011.01109.x
Winkel, M., Mitzscherling, J., Overduin, P.P., Horn, F., Winterfeld, M., Rijkers, R., Grigoriev, M.N., Knoblauch, Ch., Mangelsdorf, K., Wagner, D., and Liebner, S., Anaerobic methanotrophic communities thrive in deep submarine permafrost, Sci. Rep., 2018, vol. 8, no. 1, p. 1291. https://doi.org/10.1038/s41598-018-19505-9
Wright, J.J., Microbial community structure and ecology of Marine Group A bacteria in the oxygen minimum zone of the Northeast subarctic Pacific Ocean, Ph.D. Dissertation, University of British Columbia, 2013. https://doi.org/10.14288/1.0074090.
Xie, L.S., Xu, L., He, Y., Zhang, Y., Wang, J.H., and Xu, J., Archaeal diversity in the euphotic seawater at a slope in the northern South China Sea, Chin. J. Appl. Environ. Biol., 2017, vol. 23, pp. 21–27. https://doi.org/10.3724/SP.J.1145.2016.03005
Yanagawa, K., Nunoura, T., McAllister, S., Hirai, M., Breuker, A., Brandt, L., House, Ch. H., Moyer, C.L., Birrien, J.-L., Aoike, K., Sunamura, M., Urabe, T., Mottl, M.J., and Takai, K., The first microbiological contamination assessment by deep-sea drilling and coring by the D/V Chikyu at the Iheya North hydrothermal field in the Mid-Okinawa Trough (IODP Expedition 331), Front. Microbiol., 2013, vol. 4, p. 327. https://doi.org/10.3389/fmicb.2013.00327
Zeglin, L.H., Wang, B., Waythomas, C., Rainey, F., and Talbot, S.L., Organic matter quantity and source affects microbial community structure and function following volcanic eruption on Kasatochi Island, Alaska, Environ. Microbiol., 2016, vol. 18, no. 1, pp. 146–158. https://doi.org/10.1111/1462-2920.12924
ACKNOWLEDGMENTS
We thank A.Yu. Merkel (Vinogradsky Institute of Microbiology, Russian Academy of Sciences) for performing NGS analysis and consulting on the method; A.S. Nartov (NRC Kurchatov Institute – IREA) for measuring carbon-containing gases in permafrost; and M.Y. Cherbunina and D.G. Shmelev for consulting on the properties of methane in permafrost.
Funding
This research was supported by a grant from the Russian Science Foundation, project no. 19-77-10066 (to Nikita Demidov). Field work at the cryosphere test site near Barentsburg was carried out as part of the Russian Arctic Expedition to Spitsbergen.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Karaevskaya, E.S., Demidov, N.E., Kazantsev, V.S. et al. Archaeal Communities of Frozen Quaternary Sediments of Marine Origin on the Coast of Western Spitsbergen. Izv. Atmos. Ocean. Phys. 57, 1254–1270 (2021). https://doi.org/10.1134/S0001433821100066
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0001433821100066