Abstract
Non-alcoholic fatty liver disease (NAFLD) comprises a range of chronic liver diseases that result from the accumulation of excess triglycerides in the liver, and which, in its early phases, is categorized NAFLD, or hepato-steatosis with pure fatty liver. The mortality rate of non-alcoholic steatohepatitis (NASH) is more than NAFLD; therefore, diagnosing the disease in its early stages may decrease liver damage and increase the survival rate. In the current study, we screened the gene expression data of NAFLD patients and control samples from the public dataset GEO to detect DEGs. Then, the correlation betweenbetween the top selected DEGs and clinical data was evaluated. In the present study, two GEO datasets (GSE48452, GSE126848) were downloaded. The dysregulated expressed genes (DEGs) were identified by machine learning methods (Penalize regression models). Then, the shared DEGs between the two training datasets were validated using validation datasets. ROC-curve analysis was used to identify diagnostic markers. R software analyzed the interactions between DEGs, clinical data, and fatty liver. Ten novel genes, including ABCF1, SART3, APC5, NONO, KAT7, ZPR1, RABGAP1, SLC7A8, SPAG9, and KAT6A were found to have a differential expression between NAFLD and healthy individuals. Based on validation results and ROC analysis, NR4A2 and IGFBP1b were identified as diagnostic markers. These key genes may be predictive markers for the development of fatty liver. It is recommended that these key genes are assessed further as possible predictive markers during the development of fatty liver.
Similar content being viewed by others
Introduction
Non-alcoholic fatty liver disease (NAFLD) is a range of chronic liver diseases resulting from the accumulation of excess triglycerides in the liver1. NAFLD affects about 25–30% of the population globally and is associated with an increased risk of other diseases, including type 2 diabetes mellitus, atherosclerotic cardiovascular disease, and chronic kidney disease (CKD)2,3,4,5. The frequency of NAFLD is anticipated to increase from 83 million in 2015 to 100 million by 20306. The main reasons are an unhealthy diet and a sedentary lifestyle7. The advanced type of fatty liver disease is non-alcoholic steatohepatitis8, with features that include: fibrosis, necrotic inflammation leading to cirrhosis, and hepatocellular carcinoma1,7,9,10. The morbidity and mortality rates of cirrhosis and hepatocellular carcinoma are much greater than NAFLD11. Therefore, diagnosing and treating disease in the early stages might reduce the risk of liver damage and increase the survival rate12,13.
The gold standard for detecting NASH is liver biopsy, an invasive approach14. The other diagnostic tools ,resonance imaging15, and computed tomography (CT) whichare high-cost and time-consuming methods that burden the healthcare system financially16,17. Additionally, recognizing the higher level of lipid content and inflammatory factors such as C-reactive protein (CRP), IL-6, IL-18, IL-1b, IL-8, and TNF-a signify chronic inflammation in NASH development18,19. Therefore low-cost, reliable, and non-invasive methods are required to identify specific diagnostic biomarkers in the early stage of NAFLD.
In the progression of NAFLD, the molecular pathways are altered, leading to differential expression genes (DEGs). In the new era of technology, Machine learning (ML) is a novel artificial intelligence that has been widely performed to screen DEGs in different diseases and discover new diagnostic and prognostic biomarkers. Artificial intelligence enables processing data sets using programmed algorithms in logical models for performance tasks20,21,22. ML has various advantages, including automation, Handling multi-dimensional data, nonlinearity, low fault, and wide applications23,24.
In the current investigation, we screened the gene expression data of NAFLD patients and control samples from the public dataset GEO (Gene Expression Omnibus) to detect DEGs. Then, the correlation between the top selected DEGs and clinical data was evaluated.
Methods and materials
Workflow
The RNASeq data of fatty liver patients and clinical features were downloaded from the GEO dataset (GSE126848 and GSE48452). Filtering and normalization were performed as preprocessing, and the data quality was controlled using Principal Component Analysis (PCA). Before classification, feature selection was implemented using Relief-based algorithms to calculate the higher score for each feature. Then Penalize machine learning technique was used to detect the most important biomarkers. Eventually, the candidate genes were validated by other datasets.
Data source
In the present study, two datasets from GEO were downloaded. The first dataset included 33,297 array-based expression profiling of 73 samples which were grouped into C (control = 14), O (obese = 27), S (steatosis = 14), and N (NASH = 18), and the second dataset consisted of 19786 gene expression from normal individuals (n = 14), obese (n = 12), NAFLD (n = 15) and NASH (n = 16) patients. The two datasets were extracted from https://www.ncbi.nlm.nih.gov/geo/geo2r/?acc=GSE48452 and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE126848, respectively. The two datasets also have clinical and demographic variables considered in the analysis.
Differential expression analysis (preprocessing)
Gene expression data were screened by filtering, and the zero expressions were eliminated; then, data were normalized with limma in R 4.1 software. The adjusted p < 0.05 and − 1.5 <|Log2FC (fold change) |< 1.5 were identified for subsequent analysis as significant genes. After that, Principal Component Analysis (PCA) which is a statistical procedure for visualizing whether the sample groups (control and patients) were separable and correlated was applied..
Identifying Important genes and correlation between clinical/demographic factors with fatty liver
The effect coefficient of all factors on the fatty liver was calculated using Regularization regressions (LASSO, () Ridge, and Elastic Net) models. These models will be described as follows. Before the modeling, Relief-based feature selection was implemented. Weight by Relief is applied to calculate the weights of the attributes in the polynomial dataset. Chi-square and One way-ANOVA also were used to evaluate the relationship between clinical variables and disease, and Kolmogorov–Smirnov was used for normality test distribution. The binary correlation of some variables was examined using a correlation matrix. R4.1.and EVIEWS12 software was utilized for analysis.
Regularization regression
In statistics and machine learning, Regularization regression is a type of regression analysis for variable selection and is used when train and test data are varying. To better manage many parameters or Multicollinearity between variables and reduce complexity, a “penalty” is added to cost function (Regularization) for the best fitting of training data. This reduce the variance of the test data, prevent over-fitting and enhance the prediction accuracy. Here are briefly introduced three Regularization regressions methods.
Least Absolute Shrinkage and Selection Operator (LASSO) regression
The term Lasso stands for “least absolute shrinkage and selection operator”. Lasso uses shrinkage by shrinking data values to a central point such as mean. In this model, the regularization method is based on the absolute value of loss function. As a result, the target function in "Lasso Regression" is written as follows
Ridge regression
In the Ridge regression, the Quadratic Loss Function is used. In such way, the amount of penalty is determined as the sum of squares of coefficients.
Thus, if we consider the regression model as follows:
The Ridge regression model is performed by minimizing the following function.
Note that argmin refers to values of ββ that minimize the desired function.
To estimate the regression parameters in the Ridge method, there is a constraint on the parameters which is written as follows.
This constraint specifies that the sum of the squares of the parameters must be less than a constant or threshold value. In this way, the method of estimating the parameters will be as follows. It is clear that a balance is established between the existence of ββ parameters and their zeroing in the constraint section, and the num ber of related parameters and variables is optimized.
The λλ parameter here is called the Penalty Regulation (Regularization Parameter).
Note that regularization is done only for parameters β1β1 to βnβn, and intercept of β0β0 is an exception in this regard. Estimation of the parameters of the Ridge regression model according to the mentioned constraint will be as follows.
Elastic Net regression
Elastic Net Regression, by combining lasso regression and Ridge regression, overcomes their disadvantages and is a reliable alternative to them. Thus, if you are faced with a model whose descriptive variables are correlated with each other, it is better to use Elastic Net regression. In this method, Loss Function and Quadratic Loss Function are applied to the model simultaneously. As a result, the target function in the elastic network regression will be written as follows.
Considering the multiple linear regression model, it can also be written as follows.
Note that, like the lasso regression and the Ridge regression, in the Elastic Net regression there is no assumption that the residual is normal. Also, the intercept is not involved in the regularization25,26.
Protein–protein interaction network
The online string tool (https://www.string-db.org/) was performed to analyze DEGs’ protein–protein interaction with a score of 0.4. Moreover, all the networks were depicted using R software.
GO pathway analysis
The enrichment GO analyses were performed to detect the molecular function of DEGs in NAFLD using Go package, nrichGO, and gseGO package.
Validation of biomarkers gene expression
The expression levels of candidate genes in patients were verified by using Gene Expression Omnibus (GEO) dataset (GSE89632 and GSE63067). The validation datasets consisted of data from patients with fatty liver, which were downloaded from this web tool, and the pre-processing was performed.
Combine ROC curve
The receiver operating characteristic (ROC) curve was performed to evaluate the efficacy of the diagnostic model. Specificity, sensitivity, area under the ROC curve, positive predictive value, negative predictive value, and cut-off value were assessed for each gene and their combination. All the procedures were analyzed by package combioROC in R.
Results
Data description
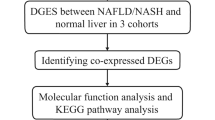
Figure 1A shows the overall workflow. Tables 1 and 2 show the mean and standard deviation of the quantitative variables. The frequency and percentage of attributes in the study are also mentioned. The result of PCA indicated the discrimination between patients and healthy samples (Fig. 1B and C).
Weight by Relief
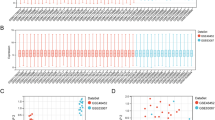
The weight of the variables of the two datasets can be seen in Fig. 2. The data show a significant correlation between DEGs and fatty liver.
Comparison of three methods for identifying important coefficients (GSE126848)
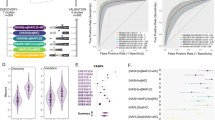
Three methods of Regularization regression, including LASSO, Ridge, and Elastic Net, were candidate to identify the effect coefficient of variables on fatty liver. Each of the color lines belongs to the coefficient of one variable, which with increasing Lambda parameter, the number of non-zero coefficients decreases, and the size of the coefficients becomes smaller and approaches zero. After fitting the model, with five k-fold cross-validation, the optimal value of the Lambda parameter was determined, and the results of the final model were reported. The model’s cross-validation results were plotted in a graph containing different values of Lambda versus Train/Test error, which shows the Train/Test Error related fitted models in different Lambda sizes (Fig. S1). Among the three implemented methods with five k-fold cross-validation for evaluation, the Elastic Net method had the highest performance (Lambda at minimum error: 11.87, R2 = 0.999 and alpha = 0.5, l1 Norm = 1.31). The area under the curve was approximately 0.99 with a confidence interval (0.95,1). The Elastic Net is an extension of the lasso robust to extreme correlations among the predictors. The results of Elastic Net method for identifying important factors can be seen in Table 3.
Comparison of three methods for identifying important coefficients (GSE48452)
The three methods of Regularization regression were used to identify candidate genes that may be used to identify the effect coefficient of variables on fatty liver. Each of the colored lines represents the coefficient of one variable, which with increasing Lambda parameter, the number of non-zero coefficients decreases, and the size of the coefficients becomes smaller and approaches zero. After fitting the model, with five k-fold cross-validation, the optimal value of Lambda parameter was gained, and the results of the final model were reported. The results of cross-validation of the model were plotted in a graph containing different values of Lambda versus Train/Test error, which shows the Train/Test Error related fitted models in different Lambda sizes (Fig. S2). Among the three implemented methods with five k-fold cross-validation for evaluation, the Elastic Net method had the highest performance (Lambda at minimum error: 0.00, R2 = 0.999 and alpha = 0.5, l1 Norm = 213.66). The area under the curve was approximately 0.99 with a confidence interval (0.95, 1). The Elastic Net is an extension of the lasso robust to extreme correlations among the predictors. The results of Elastic Net method for identifying important factors can be seen in Table 3.
Comparison of three methods for identifying common genes between two datasets
After normalization with significant p-value and log fold change, the common genes between GSE126848 and GSE48452 were 155, which were used to identify the most important candidate genes using Lasso Machine Learning technique. For GSE126848 dataset with 57 samples, among the three implemented methods with five k-fold cross validation for evaluation, the Lasso method had the highest performance (Lambda at minimum error: 1.451, R2 = 0.999 and alpha = 1, L1 Norm = 15.96)(Fig. S3). For GSE48452 dataset with 73 samples, among the three implemented methods with five k-fold cross-validation for evaluation, the Lasso method had the highest performance (Lambda at minimum error: 0.01388, R2 = 0.999 and alpha = 1, L1 Norm = 15.96) (Fig. S4).
Identification of dysregulate expression genes (DEGs)
The GSE48452 chip contained 14 NAFLD, 18 NASH, and 27 obese samples, among which 15,000 genes and 1400 DEGs were identified. Moreover, the GSE126848 chip had 15 NAFLD, 16 NASH, and 12 obese 9540 genes, and 843 DEGs were found in this dataset based on specific criteria (Table 2). Furthermore, the commonality of novel genes between two datasets was assessed after normalization. Then Penalize machine learning technique was used to detect the most important common genes between two data sets. The results indicated that eighty-eight genes were common between two datasets (Table 3).
PPI network construction
As seen in Fig. 3, the PPI interaction network of DEGs was analyzed and depicted by String, and the interaction score was set at 0.4. As we can see in the network analysis, the KATA6A and KAT7 genes were strongly correlated, as well as, a significant correlation was detected between the SART3 and RNPS1 genes.
Gene ontology (GO) functional annotation of top DEGs enrichment terms in fatty liver disease; molecular function (MF) of DEGs was mainly enriched in histone acetyltransferase activity, peptide − lysine − N − acetyltransferase activity, histone binding, and peptide N − acetyltransferase activity. The biological process (BP) consisted of RNA splicing, hematopoietic stem cell proliferation, and histone H3 acetylation. The cell component (CC) was detected in nuclear speck and H3 histone acetyltransferase complexes.
Gene ontology analyses of DEGs
R software results showed that the molecular function of DEGs was enriched in histone acetyltransferase activity, peptide–lysine–N–acetyltransferase activity, histone binding, and peptide N-acetyltransferase activity. The biological process includes RNA splicing, hematopoietic stem cell proliferation, and histone H3 acetylation. Furthermore, the cell component was detected in nuclear speck and H3 histone acetyltransferase complexes (Fig. 3).
Validation using validation datasets
The five common genes between two datasets, GEO126848 and GEO48452, were validated by two other datasets, consisting of GSE89632 and GSE63067. The results indicated the five most important novel genes in fatty liver, including NR4A2, ZEB2, IGFBP1b, AKR1B10, DHRS2, and UGT2B17 (Table 4).
GO pathway analyses
Enrichment analysis results showed that the molecular function of shared DEGs was mainly enriched in structural molecule activity. The biological processes were peptide biosynthetic process and translation. Moreover, the main involved cell components were ribonucleoprotein complex and ribosome. Reactom pathway analysis revealed that metabolism of RNA and cellular responses to stress and stimuli were the most significant dysregulated pathways in fatty liver (Fig. 3).
ROC curve for identification of diagnostic markers
Our finding showed that NR4A2 alone (AUC of 0.92, 95% CI with a sensitivity of 1.00and specificity of 0.71), and also, its combination with ZEB2 (AUC of 0.92, 95% CI with a sensitivity of 0.90 and specificity of 0.85) had the highest rank of ROC analysis and can be considered as diagnostic markers (Fig. S5 and Table S1). Moreover, our data revealed that IGFBP1b alone (AUC of 0.90, 95% CI with a sensitivity of 0.89 and specificity of 0.87), and its combination with AKR1B10, DHRS2, IGFBP1, and UGT2B17 with AUC of 0.96, 95% CI with a sensitivity of 0.94 and specificity of 0.95, also had the highest rank (Fig. S6 and Table S2).
Association between Clinical/Demographic factors and fatty liver
A significant relationship was obtained between fat, fibrosis, BMI, inflammation, and fatty liver.
Investigation of the binary correlations of Clinical/Demographic influence variables on fatty liver
Using the correlation matrix, we examined the correlation between pairs of variables. The results are shown in Fig. 4. Note that a correlation coefficient of less than 0.3 is considered weak, the coefficient between 0.3 and 0.6 is moderate, and a coefficient greater than 0.6 is considered strong. Coefficients with a P-value less than 0.05 are also significant. As we concluded from Fig. 4, BMI, Lar, Leptin, Fat, and Nas have correlated significantly with the disease in positive direct and Adiponectin correlated with fatty liver negatively.
Discussion
For the first time in the present study, we have used machine learning approaches to compare the gene expression profile of individuals with NAFLD, NASH, and obesity with healthy individuals.
Firstly, we analyzed GSE126848 and GSE48452 datasets separately, and the results detected 9540 and 1400 DEGs genes in the two datasets, respectively. We reported genes with higher coefficients in each dataset. Six genes, including ABCF1, SART3, APC5, NONO, KAT7, and ZPR1 were identified in GSE48452 datasets, as well as four genes, including RABGAP1, SLC7A8, SPAG9, and KAT6A were detected in GSE126848 dataset with a different expression between NAFLD and healthy samples. Subsequently, we identified six common genes between the two datasets and validated them in other datasets. Further analysis demonstrated that two genes, including NR4A2 and IGFBP1b with higher AUC, sensitivity, and specificity, were diagnostic biomarkers in fatty liver.
ABCF1, also named ABC50, is a member of the ABC transporter superfamily protein localized on the cytosol and endoplasmic reticulum (ER), which transport different molecules, including carbohydrates, amino acids, and ions. Furthermore, ABCF1 is critical in regulating innate immune and inflammatory responses27,28. This protein is considered an oncofetal protein significantly expressed in the fetal liver, not healthy adult cells. Fung et al. showed that the expression of ABCF1 was increased in hepatocellular carcinoma (HCC), and was associated with chemoresistance29. Cheung et al. demonstrated that upregulated ABCF1 gene is associated with poor recurrence-free survival (RFS) in liver cancer30. A significant association between other members of the ABC family and NAFLD has been proven in previous studies. ABCB1 plays a crucial role in transporting phospholipids and cholesterol into the liver cells. An animal study exhibited that the level of transporter proteins such as ABCB1, ABCC1-6, and ABCG2 increased during the progression of NASH31. The ABCB1 is overexpressed in liver diseases such as cholestatic, biliary cirrhosis, and obstructive jaundice32,33,34. The SART3 and RNPS1 are the genes with the highest score in the advanced stage of NAFLD; moreover, the result of PPI revealed that there is a strong correlation between SART3 and RNPS1, both of them are members of the post-splicing complex. SART3 is known as tumor-associated antigens detected in HCC and makes hepatocytes sensitive to immunotherapy35. A previous study used two datasets of GEO (GSE33814 and GSE89632) and showed that RNPS1 is one of the top genes overexpressed in NAFLD cells compared to the control group. RNPS1 is a member of the post-splicing complex role in RNA processing and apoptosis36. One of the other key genes detected in our investigation was APC5, a subunit of the anaphase-promoting complex (APC). Zhang et al. showed APC5 plays a critical role in activating the cell cycle during adipose tissue proliferation37. A study showed that after feeding, the expression of NONO gene significantly increased to uptake glucose. Furthermore, the results revealed that the deficient-NONO gene in mice reduces triglyceride storage and increases hepatocyte lipid catabolism38. In a current study, Wu et al. indicated that the expression of NONO gene was highly elevated in NAFLD mice39. our result indicates that CNTNAP1 is upregulated in NAFLD, which agrees with the previous study. CNTNAP1 has a positive role in triglyceride metabolism40. KAT7 gene, also known as HBO1, belongs to the lysine acetyltransferase family, which is a key factor in forming a replication complex, regulating the immune system and developing embryonic development. Information confirmed that the expression of KAT7 in mRNA and protein levels elevated in HCC cells leads to the proliferation and invasion of tumor cells. Zhong et al. reported that silencing the KAT7 gene using short hairpin RNA (shRNA) and CRISPR/Cas9 in the xenograft HCC model inhibited tumorigenesis41. ZPR1 is a zinc finger family member, and Wo et al. showed patients with severe NAFLD had ZPR1 rs964184 polymorphism. we hypothesized that this polymorphism could be associated with high expression of ZPR1 in patients42.
The analysis of the GSE126848 dataset revealed that the expression of RABGAP1 gene is associated with NAFLD. The previous studies showed Rabgap1 expression raised in perirenal fat and brown fat in Gpr21 knockout mice when fed with a high-fat diet43. Rabgap1 GTPase Activating protein which transited the cells from metaphase to anaphase. SLC7A8 and SPAG9 are two novel DEGs identified in our study. SLC7A8, the light-chain subunit solute carrier family 7, member 8, is a vital gene in inducing hypertrophy in adipose tissue and inflammation. Pitere et al. reported that the SLC7A8 deficiency in mice with diet-induced obesity decreases lipid accumulation in the liver44. SPAG9 is expressed explicitly in the testis and has a vital role in fertility. A study on chicken illustrated that the samples that overexpressed the SPAG9 gene have more fat content on the abdominal and liver tissues45. Furthermore, SPAG9 increases the proliferation of HCC cells through the interaction with MAPK/Jun pathway45. KAT6A is another member of the lysine acetyltransferase family, which epigenetically regulates the transcription of different genes involved in DNA repairing systems, cell cycle, metabolism, and autophagy. Many studies confirmed the overexpression of KAT6A related to HCC progression and chemoresistance46,47.
Our result revealed a significant relationship between clinical and demographic data, including fat, fibrosis, body mass index (BMI), inflammation, and fatty liver. In many studies, BMI is announced as a critical index for increasing the risk of fatty liver. The BMI score of patients is a 4 to 14-fold change higher than healthy individuals. Fan et al. reported that 73% of patients with NAFLD were obese and overweight48. BMI measurement is a helpful and non-invasive marker for predicting fatty liver. They suggested triple approaches comprising examining the lipid panel, BMI measurement, and radiological techniques49,50. Inflammation and fibrosis are the major pathological consequences of NAFLD. Fibrogenesis is stimulated by the activation of hepatic stellate cells and Kupffer cells, resulting from high plasma levels of glucose and lipids51. The activated hepatic stellate cells express different myogenic and pro-inflammatory markers such as myocyte enhancer factor-2 (Mef2), c-myb, and TGF-β. Moreover, inflammation results from increasing the level of reactive oxygen species (ROS) and cytokines in liver tissue52,53. The result of a meta-analysis revealed that the fibrosis stage significantly correlates with the risk of mortality in NAFLD54.
We reported NR4A2 and IGFBP1b as novel diagnostic biomarkers in fatty liver. Insulin-like growth factor binding protein (IGFBP) binds to insulin-like growth factors (IGFs) and regulates cellular metabolism. Hepatocytes largely produce IGFBP and secrete it into the serum. Previous studies are in line with our results, Pan et al. reported a high expression of IGFBP in NAFLD patients, L02 cells, and also in mice models of NAFLD55. NR4A2 is a transcription factor that plays a pivotal role in regulating fatty acid beta-oxidation. Therefore, the dysregulation of NR4A2 causes fat accumulation in the liver56. Chen et al. showed that NR4A2 overexpression prevents Hepatic stellate cell (HSCs) proliferation which plays a key role in liver fibrogenesis57.
Previous evidence confirmed that novel approaches, including machine learning, are promising strategies for diagnosing, preventing, and managing diseases. Wu et al. compared four machine learning algorithms in predicting fatty liver disease, and they showed that the random forest model has a higher performance in the early diagnosis of fatty liver58. The result of a cross-sectional investigation showed that machine learning is a predictive model of NAFLD. They revealed that this method enhances clinical decisions and reduces end-stage disease59. Furthermore, previous studies used machine learning methods for identifying novel biomarkers in various conditions, such as cancer60,61,62, cardiovascular diseases63,64, pulmonary diseases65,66, and neurological disorders67,68.
In conclusion, using a bioinformatic approach; twelve key genes were detected that are significantly related to the fatty liver. It is recommended that these key genes are assessed further as possible predictive markers during the development of the fatty liver.
Data availability
The datasets generated and/or analysed during the current study are available in the GEO repository, https://www.ncbi.nlm.nih.gov/geo/geo2r/?acc=GSE48452 and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE126848.
References
Yu, J., Marsh, S., Hu, J., Feng, W. & Wu, C. The pathogenesis of nonalcoholic fatty liver disease: Interplay between diet, gut microbiota, and genetic background. Gastroenterol. Res. Pract. 2016, 13 (2016).
Younossi, Z. M. et al. Global epidemiology of nonalcoholic fatty liver disease—meta-analytic assessment of prevalence, incidence, and outcomes. Hepatology 64(1), 73–84 (2016).
Kaya, E. & Yilmaz, Y. Non-alcoholic Fatty Liver Disease: A Global Public Health Issue 321–333 (Springer, 2020).
Tanaka, N. et al. Current status, problems, and perspectives of non-alcoholic fatty liver disease research. World J. Gastroenterol. 25(2), 163 (2019).
Sanyal, A. J. et al. Nonalcoholic steatohepatitis: Association of insulin resistance and mitochondrial abnormalities. Gastroenterology 120(5), 1183–1192 (2001).
Estes, C., Razavi, H., Loomba, R., Younossi, Z. & Sanyal, A. J. Modeling the epidemic of nonalcoholic fatty liver disease demonstrates an exponential increase in burden of disease. Hepatology 67(1), 123–133 (2018).
Mirmiran, P., Amirhamidi, Z., Ejtahed, H.-S., Bahadoran, Z. & Azizi, F. Relationship between diet and non-alcoholic fatty liver disease: A review article. Iran. J. Public Health 46(8), 1007 (2017).
Koppad, S., Basava, A., Nash, K., Gkoutos, G. V. & Acharjee, A. Machine learning-based identification of colon cancer candidate diagnostics genes. Biology 11(3), 365 (2022).
Ibrahim, S. H., Hirsova, P. & Gores, G. J. Non-alcoholic steatohepatitis pathogenesis: Sublethal hepatocyte injury as a driver of liver inflammation. Gut 67(5), 963–972 (2018).
Ratziu, V., Bellentani, S., Cortez-Pinto, H., Day, C. & Marchesini, G. A position statement on NAFLD/NASH based on the EASL 2009 special conference. J. Hepatol. 53(2), 372–384 (2010).
Kim, D. et al. Changing trends in etiology-based and ethnicity-based annual mortality rates of cirrhosis and hepatocellular carcinoma in the United States. Hepatology 69(3), 1064–1074 (2019).
Cotter, T. G. et al. Nonalcoholic fatty liver disease: Impact on healthcare resource utilization, liver transplantation and mortality in a large, integrated healthcare system. J. Gastroenterol. 55(7), 722–730 (2020).
Schuppan, D. & Afdhal, N. H. Liver cirrhosis. The Lancet 371(9615), 838–851 (2008).
Sumida, Y., Nakajima, A. & Itoh, Y. Limitations of liver biopsy and non-invasive diagnostic tests for the diagnosis of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. World J. Gastroenterol.: WJG 20(2), 475 (2014).
Stangeland, B. et al. Combined expressional analysis, bioinformatics and targeted proteomics identify new potential therapeutic targets in glioblastoma stem cells. Oncotarget 6(28), 26192 (2015).
Bedogni, G. et al. The Fatty Liver Index: A simple and accurate predictor of hepatic steatosis in the general population. BMC Gastroenterol. 6(1), 1–7 (2006).
Schwenzer, N. F. et al. Non-invasive assessment and quantification of liver steatosis by ultrasound, computed tomography and magnetic resonance. J. Hepatol. 51(3), 433–445 (2009).
Baranova, A., Birerdinc, A. & Younossi, Z. M. Adipokines in Nonalcoholic Fatty Liver Disease. Adipose Tissue and Adipokines in Health and Disease 249–283 (Springer, 2014).
Lee, Y.-H. & Pratley, R. E. The evolving role of inflammation in obesity and the metabolic syndrome. Curr. Diabetes Rep. 5(1), 70–75 (2005).
Mohammed, M., Khan, M. B. & Bashier, E. B. M. Machine Learning: Algorithms and Applications (CRC Press, 2016).
Zhang, X., Jonassen, I. & Goksøyr, A. Machine Learning Approaches for Biomarker Discovery Using Gene Expression Data 53–64 (Exon Publications, 2021).
Chang, C.-H., Lin, C.-H. & Lane, H.-Y. Machine learning and novel biomarkers for the diagnosis of Alzheimer’s disease. Int. J. Mol. Sci. 22(5), 2761 (2021).
Wuest, T., Weimer, D., Irgens, C. & Thoben, K.-D. Machine learning in manufacturing: Advantages, challenges, and applications. Prod. Manuf. Res. 4(1), 23–45 (2016).
Khanzode, K. C. A. & Sarode, R. D. Advantages and disadvantages of artificial intelligence and machine learning: A literature review. Int. J. Libr. Inf. Sci. (IJLIS) 9(1), 3 (2020).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33(1), 1 (2010).
Dorugade, A. & Kashid, D. Alternative method for choosing ridge parameter for regression. Appl. Math. Sci. 4(9), 447–456 (2010).
Lee, M. N. et al. Identification of regulators of the innate immune response to cytosolic DNA and retroviral infection by an integrative approach. Nat. Immunol. 14(2), 179–185 (2013).
Carney, E. F. ABCF1 switches off inflammation in sepsis. Nat. Rev. Nephrol. 15(5), 255–355 (2019).
Fung, S. W. et al. The ATP-binding cassette transporter ABCF1 is a hepatic oncofetal protein that promotes chemoresistance, EMT and cancer stemness in hepatocellular carcinoma. Cancer Lett. 457, 98–109 (2019).
Cheung, S. T., Cheung, P. F., Cheng, C. K. & Fan, S. T. Genomics approach to identify drug transporter ABCF1 associated with liver cancer recurrence and chemo-resistance: The 103rd Annual Meeting of the American Association for Cancer Research (AACR 2012), Chicago, IL., 31 March-4 April 2012
Hardwick, R. N., Fisher, C. D., Canet, M. J., Scheffer, G. L. & Cherrington, N. J. Variations in ATP-binding cassette transporter regulation during the progression of human nonalcoholic fatty liver disease. Drug Metabol. Dispos. 39(12), 2395–2402 (2011).
Lickteig, A. J. et al. Efflux transporter expression and acetaminophen metabolite excretion are altered in rodent models of nonalcoholic fatty liver disease. Drug Metabol. Dispos. 35(10), 1970–1978 (2007).
Kojima, H. et al. Disturbed colocalization of multidrug resistance protein 2 and radixin in human cholestatic liver diseases. J. Gastroenterol. Hepatol. 23(7), e120–e128 (2008).
Kojima, H. et al. Changes in the expression and localization of hepatocellular transporters and radixin in primary biliary cirrhosis. J. Hepatol. 39(5), 693–702 (2003).
Mizukoshi, E. et al. Comparative analysis of various tumor-associated antigen-specific t-cell responses in patients with hepatocellular carcinoma. Hepatology 53(4), 1206–1216 (2011).
Li, L. et al. Identification of key genes in non-alcoholic fatty liver disease progression based on bioinformatics analysis. Mol. Med. Rep. 17(6), 7708–7720 (2018).
Zhang, J. et al. Differential expression of cell cycle regulators during hyperplastic and hypertrophic growth of broiler subcutaneous adipose tissue. Lipids 50(10), 965–976 (2015).
Benegiamo, G. et al. The RNA-binding protein NONO coordinates hepatic adaptation to feeding. Cell Metabol. 27(2), 404–418 (2018).
Wu, P., Zhang, M. & Webster, N. J. Alternative RNA splicing in fatty liver disease. Front. Endocrinol. 12, 58 (2021).
Naderi, M. et al. Two triacylglycerol pathway genes, CTDNEP1 and LPIN1, are down-regulated by hsa-miR-122–5p in hepatocytes. Arch. Iran. Med. 20(3), 1 (2017).
Zhong, W., Liu, H., Deng, L., Chen, G. & Liu, Y. HBO1 overexpression is important for hepatocellular carcinoma cell growth. Cell Death Dis. 12(6), 1–13 (2021).
Esteve-Luque, V. et al. Implication between genetic variants from APOA5 and ZPR1 and NAFLD severity in patients with hypertriglyceridemia. Nutrients 13(2), 552 (2021).
Wang, J. et al. GPR21 KO mice demonstrate no resistance to high fat diet induced obesity or improved glucose tolerance. F1000Research 2016, 5 (2016).
Pitere, R. R., van Heerden, M. B., Pepper, M. S. & Ambele, M. A. Slc7a8 deletion is protective against diet-induced obesity and attenuates lipid accumulation in multiple organs. Biology 11(2), 311 (2022).
Zhang, H. et al. Detection of genome-wide copy number variations in two chicken lines divergently selected for abdominal fat content. BMC Genom. 15(1), 1–12 (2014).
Jin, Y. et al. KAT6A is associated with sorafenib resistance and contributes to progression of hepatocellular carcinoma by targeting YAP. Biochem. Biophys. Res. Commun. 585, 185–190 (2021).
Wiesel-Motiuk, N. & Assaraf, Y. G. The key roles of the lysine acetyltransferases KAT6A and KAT6B in physiology and pathology. Drug Resist. Updates 53, 100729 (2020).
Fan, R., Wang, J. & Du, J. Association between body mass index and fatty liver risk: A dose-response analysis. Sci. Rep. 8(1), 15273. https://doi.org/10.1038/s41598-018-33419-6 (2018).
Fan, R., Wang, J. & Du, J. Association between body mass index and fatty liver risk: A dose-response analysis. Sci. Rep. 8(1), 1–7 (2018).
Abangah, G. et al. Correlation of body mass index and serum parameters with ultrasonographic grade of fatty change in non-alcoholic fatty liver disease. Iran. Red Crescent Med. J. 16(1), 859 (2014).
Pourali, G. et al. Therapeutic potential of herbal medicine against non-alcoholic fatty liver disease. Curr. Drug Targets 2023, 859 (2023).
Basaranoglu, M., Basaranoglu, G. & Sentürk, H. From fatty liver to fibrosis: A tale of “second hit”. World J. Gastroenterol.: WJG 19(8), 1158 (2013).
Zafari, N. et al. Role of gut bacterial and non-bacterial microbiota in alcohol-associated liver disease: Molecular mechanisms, biomarkers, and therapeutic prospective. Life Sci. 305, 120760. https://doi.org/10.1016/j.lfs.2022.120760 (2022).
Taylor, R. S. et al. Association between fibrosis stage and outcomes of patients with nonalcoholic fatty liver disease: A systematic review and meta-analysis. Gastroenterology 158(6), 1611–1625 (2020).
Pan, J. et al. Insulin-like growth factor binding protein 1 ameliorates lipid accumulation and inflammation in nonalcoholic fatty liver disease. J. Gastroenterol. Hepatol. 36(12), 3438–3447 (2021).
Holla, V. R., Wu, H., Shi, Q., Menter, D. G. & DuBois, R. N. Nuclear orphan receptor NR4A2 modulates fatty acid oxidation pathways in colorectal cancer. J. Biol. Chem. 286(34), 30003–30009 (2011).
Chen, P. et al. Orphan nuclear receptor NR4A2 inhibits hepatic stellate cell proliferation through MAPK pathway in liver fibrosis. PeerJ 3, e1518 (2015).
Wu, C.-C. et al. Prediction of fatty liver disease using machine learning algorithms. Comput. Methods Progr. Biomed. 170, 23–29 (2019).
Ma, H., Xu, C.-F., Shen, Z., Yu, C.-H. & Li, Y.-M. Application of machine learning techniques for clinical predictive modeling: A cross-sectional study on nonalcoholic fatty liver disease in China. BioMed Res. Int. 2018, 9 (2018).
Khalili-Tanha, G. et al. Identification of ZMYND19 as a novel biomarker of colorectal cancer: RNA-sequencing and machine learning analysis. J. Cell Commun. Signal. 2023, 1–17 (2023).
Xie, Y. et al. Early lung cancer diagnostic biomarker discovery by machine learning methods. Transl. Oncol. 14(1), 100907 (2021).
Nazari, E. et al. Identification of potential biomarkers in stomach adenocarcinoma using machine learning approaches. Curr. Bioinform. 18(4), 320–333 (2023).
Dogan, M. V. et al. Blood-based biomarkers for predicting the risk for five-year incident coronary heart disease in the Framingham Heart Study via machine learning. Genes 9(12), 641 (2018).
Poss, A. M. et al. Machine learning reveals serum sphingolipids as cholesterol-independent biomarkers of coronary artery disease. J. Clin. Investig. 130(3), 1363–1376 (2020).
Zhang, Y. et al. Machine-learning algorithm-based prediction of diagnostic gene biomarkers related to immune infiltration in patients with chronic obstructive pulmonary disease. Front. Immunol. 13, 740513 (2022).
Dhong, K.-R., Lee, J.-H., Yoon, Y.-R. & Park, H.-J. Identification of TRPC6 as a novel diagnostic biomarker of pm-induced chronic obstructive pulmonary disease using machine learning models. Genes 14(2), 284 (2023).
Su, C., Tong, J. & Wang, F. Mining genetic and transcriptomic data using machine learning approaches in Parkinson’s disease. NPJ Parkinson’s Dis. 6(1), 24 (2020).
Ren, J., Zhang, B., Wei, D. & Zhang, Z. Identification of methylated gene biomarkers in patients with Alzheimer’s disease based on machine learning. BioMed Res. Int. 2020, 11 (2020).
Author information
Authors and Affiliations
Contributions
S.M.H., A.A., M.G.-M., and G.A.F. conceived of the presented idea. A.A., M.N., M.A.K., M.M., and M.K. developed the theory and performed the data collection and analysis. G.P., E.N., and G.K.T. provided the initial draft of the manuscript. All authors commented on previous versions of the manuscript, discussed the results, and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nazari, E., Khalili-Tanha, G., Asadnia, A. et al. Bioinformatics analysis and machine learning approach applied to the identification of novel key genes involved in non-alcoholic fatty liver disease. Sci Rep 13, 20489 (2023). https://doi.org/10.1038/s41598-023-46711-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46711-x
- Springer Nature Limited