Abstract
The evolution of the modern human brain was accompanied by distinct molecular and cellular specializations, which underpin our diverse cognitive abilities but also increase our susceptibility to neurological diseases. These features, some specific to humans and others shared with related species, manifest during different stages of brain development. In this multi-stage process, neural stem cells proliferate to produce a large and diverse progenitor pool, giving rise to excitatory or inhibitory neurons that integrate into circuits during further maturation. This process unfolds over varying time scales across species and has progressively become slower in the human lineage, with differences in tempo correlating with differences in brain size, cell number and diversity, and connectivity. Here we introduce the terms ‘bradychrony’ and ‘tachycrony’ to describe slowed and accelerated developmental tempos, respectively. We review how recent technical advances across disciplines, including advanced engineering of in vitro models, functional comparative genetics and high-throughput single-cell profiling, are leading to a deeper understanding of how specializations of the human brain arise during bradychronic neurodevelopment. Emerging insights point to a central role for genetics, gene-regulatory networks, cellular innovations and developmental tempo, which together contribute to the establishment of human specializations during various stages of neurodevelopment and at different points in evolution.
Similar content being viewed by others
Main
The age-old question of what sets the human brain apart from those of other species touches on diverse fields including evolutionary biology, anthropology, psychology and neuropathology. There is profound interest in understanding the distinct molecular and cellular structures of the human brain, as well as how these come about during neurodevelopment, in order to ultimately link these to our broad range of cognitive abilities. Many of these abilities have often been touted as being unique to our species; however, recent reinvestigations have revealed that other animals exhibit cognitive abilities that were previously considered to be human-specific, such as communication, theory of mind and self-awareness1,2. Similarly, animals have evolved characteristic cognitive abilities that humans lack—for example, long-distance navigational memory3. Thus, rather than a singular skill, it appears that the diversity and degree of cognitive abilities—for example, advanced language skills, complex problem solving and rich social and cultural societies2,4,5— is characteristic in humans, which also seems to make our species more prone to neuropsychiatric diseases6,7.
Similar to our cognitive abilities, most of the genetic, cellular and structural features of the human brain are shared with our relatives8,9,10, indicating that many aspects of human neurodevelopment and function can be extrapolated from other species. Indeed, important insights from a broad spectrum of model organisms have laid the foundation for our current understanding of human brain development and function. However, the human brain also exhibits distinct features at the functional, cellular and molecular levels that manifest during brain development. Thus, there is a keen interest in understanding exactly what these specializations are and how human neurodevelopment diverges to give rise to them. Investigating the neurobiological basis of human-specific specializations has long been challenging and has until recently relied on snapshot views of the human brain compared with model organisms, as well as limited in-depth molecular profiling. Technical advances in in vitro models, comparative genetics and single-cell profiling are now providing an unprecedented opportunity to explore human-specific features across a wide range of neurodevelopmental stages and at high molecular resolution11,12,13,14,15,16,17. Recent studies have begun to provide a deeper mechanistic understanding of the molecular and cellular innovations of the human brain, with a central role for genetics and gene-regulatory networks, as well as developmental tempo, one of the most notable developmental differences in human brain evolution. Although the field is still nascent, insights into specializations that arise at various points along the human lineage, from distant relatives such as rodents to our closest hominin ancestors, as well as a mechanistic understanding of how they come about during development, are emerging.
Distinct features of the human brain
The adult human brain exhibits certain features that are likely to be evolutionary adaptations (Box 1) that arose at different times during evolution. These can be unique to humans (human-specific), or shared with other great apes (hominid-specific), primates (primate-specific) or mammals (mammal-specific) (Fig. 1a). One of the most distinct anatomical specializations of the human brain is its notable relative size (Fig. 1b,c). Although primates exhibit a high degree of encephalization (a metric for brain size relative to body mass), this feature is more pronounced in humans18. The human brain is about three times larger than those of our closest living relatives—chimpanzees, gorillas and bonobos19. A similar increase is observed when comparing modern humans to Australopithecus, an archaic ancestor of humans with a brain size of around 450 cm3, based on endocranial volumes of endocast fossils20. However, brain size began to increase with the birth of the genus Homo two million years ago, where a slight increase in endocranial volume (to around 600 cm3) is first visible, followed by a more marked increase to approximately 1,000 cm3 in Homo erectus and finally the jump to 1,400–1,500 cm3 in Homo sapiens and Homo neanderthalensis21. Thus, the evolutionary expansion of the human brain was a gradual one involving stepwise enlargement, suggesting multiple evolutionary events.
a, Phylogenetic tree of the human lineage, highlighting brain specializations that have evolved at each point (see b–f for details and references). b, Comparative analysis across species of specializations emerging at different times during human evolution, including the expansion of cortical, cerebellar and white matter structures38,44,235 (top), the relative increase of supragranular neurons44 (middle) and the increase in neuronal spine number and dendritic arborization66,67,71,236 (bottom). c, Scaled representation of a coronal brain section of mouse237, chimpanzee and human (modified with permission from Welker and Johnson Comparative Brain Collection and http://brainmuseum.org). White and grey matter regions are highlighted, indicating a relative increased evolutionary expansion of white matter. d, Schematic representation of the mouse, chimpanzee and human brain displayed at similar sizes, highlighting regions that show increased relative expansion during evolution. MC, motor cortex; PFC, prefrontal cortex. e, Cross-species analysis of the proportion of different cortical layers relative to the total cortical thickness44. f, Schematic representation of cortices of mouse, chimpanzee (prefrontal cortex) and human (prefrontal cortex and motor cortex), with a magnified representation of a dendritic shaft with spines. Illustrations display species-specific and regional differences in cortical thickness44, relative proportion of supragranular and infragranular neurons44, and dendritic arborization and spine number66,67,71,236.
Expansion of brain regions
The evolutionary expansion of the brain was not entirely uniform, and specific regions, such as the neocortex, cerebellum and white matter structures, have undergone increased expansion (Fig. 1b,c). Although most of this increase aligns with expected allometric scaling patterns, a fraction remains that is unaccounted for22. This has sparked discussions of whether some regions expanded merely owing to an overall increase in brain size23 or if they could also have evolved independently24. Within these debates regarding the target of selection, there is consensus that these brain regions, particularly the neocortex, are integral for our distinct cognitive abilities.
Indeed, the expansion and specialization of the neocortex has been a focal point of recent research owing to its link with complex cognitive functions and related pathologies. Compared with other primates, the human neocortex has undergone 50% more expansion relative to the rest of the brain25, with an accompanying increase in gyri and sulci. Accordingly, following allometric scaling, humans have the largest number of cortical neurons compared to close and more distant relatives26,27,28. Compared to chimpanzees, humans display a reduced genetic heritability in cortical anatomy and size, particularly in the association cortices, suggesting greater environmental influences on our brain development29. Parts of the association cortices, including the prefrontal, temporal and parietal lobes, show additional selective enlargement in humans30. Of these, the prefrontal cortex has attracted substantial attention, as this region in the frontal lobe is key for flexible integration of various functional modalities and is more susceptible to neuropsychiatric disorders, including schizophrenia, bipolar disorder and post-traumatic stress disorder31,32,33. Differences in the frontal lobe in modern humans have been suggested to have evolved more recently than initially thought, as a fossil study reported that this reorganization occurred around the time of the initial migrations of Homo from Africa34. Thus, the expansion and macrostructural remodelling of the neocortex, particularly the association cortices, have evolved along the human lineage, with a notable human-specific increase.
Another brain region that has shown relatively increased expansion in humans, yet remains relatively less explored, is the cerebellum35,36,37,38 (Fig. 2b,d). Humans exhibit further selective enlargements of cerebellar regions connected to the frontal and parietal association cortices compared with chimpanzees35,36,37,38. This regional expansion is disproportionate when compared with the expansion of cerebellar cortical motor areas, which has led to the emerging hypothesis that the cerebellum, in addition to its well-known role in motor functions, may contribute to specialized cognitive functions in humans37.
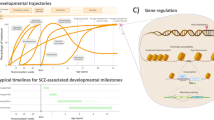
a, Schematic illustration of bradychrony and tachychrony, which describe slowed and accelerated developmental timing, respectively. b, Relation between neonatal brain weight and fetal brain growth tempo across species. Data are presented in Supplementary Table 1. Nonlinear regression, R2 = 0.86, with 10 degrees of freedom. c, Relation between adult brain weight and adult brain growth tempo across species. Data are presented in Supplementary Table 2. Nonlinear regression, R2 = 0.96, with 6 degrees of freedom. SB tamarins, saddle-back tamarins. d, Comparative analysis across species of the time window of total brain development, from the start of gastrulation until the end of myelination. Data are presented in Supplementary Table 3. e, Comparative analysis across species of the relative time of total brain development (as shown in c) compared to the average life expectancy in captivity. The red line indicates complete brain development (as shown in d). Data are presented in Supplementary Table 3.
In parallel with the evolutionary expansion of cortical and cerebellar grey matter, which harbour somatodendritic structures and local axon projections, there is an even greater increase in white matter volume along the human lineage (Fig. 2b,c). This is mostly attributable to the long-range axon projections in the white matter, which must cover greater distances in larger brains. Indeed, the disproportionate increase in white matter volume in different mammalian species is largely explained by a universal power law, suggesting that animals from different lineages with large brains share global mechanisms that drive these changes39. Additionally, there is further regional expansion of white matter structures that is human-specific and could underlie human-specific differences in connectivity. For instance, cross-species analyses in humans, great apes and other primates show that the expansion of the prefrontal cortex is mostly explained by an increase in white matter structures rather than grey matter structures40,41, indicating a human-specific increase in connectivity of the prefrontal cortex with other brain structures.
Cellular conservation and divergence
Despite large species-specific variations in the size of brains and certain brain regions, the overall tissue architecture is shared across mammals, indicating that the basic ontogenetic plan is developmentally and genetically conserved. Overall homology of neuronal and non-neuronal subtypes is also largely similar across mammals. The mammalian neocortex exhibits a six-layered cytoarchitecture, with pyramidal neurons, the predominant glutamatergic excitatory subtypes, categorized by their layer-specific localizations. These include infragranular neurons (layer V and VI), which largely project to subcortical regions, supragranular neurons (layer II and III), which mainly project to other cortical regions, and layer IV neurons that innervate local structures while receiving input from the thalamus and other regions (Figs. 2e and 3a). The cortex displays regional variations in overall thickness, as well as differences in layering and thickness of individual layers42. Granular cortical regions contain all six layers, whereas agranular areas lack layer IV42. The granular prefrontal cortex has received particular attention in the context of human brain evolution, as it emerged initially in early primates and further diversified in simians, and its evolutionary expansion occurred in concert with other association areas43.
Although overall cortical cytoarchitecture is shared across mammals, the structure is expanded and more elaborate in primates, exhibiting thickened supragranular layers throughout the cortex compared with other mammals44 (Fig. 1e). The expansion of supragranular layers extends to apes, with a further thickened cortical grey matter exhibiting subdivision of layers II and III into as many as four sublayers in certain regions45 that has remained unchanged in humans compared with other apes44,46 (Fig. 1e). This expansion in cytoarchitecture is also reflected in the wider diversification of glutamatergic neuron types in humans compared to mice, which is observed at the morphological, biophysical and transcriptomic levels47.
In mice, the cortical neuronal population is made up of approximately 15% GABAergic inhibitory neurons, a proportion that is largely invariant across neocortical areas48,49. In primates, however, these proportions vary by neocortical area and generally favour higher proportions of interneurons, reaching as much as 30–40%, particularly in higher-order association areas48,49,50,51. Ferrets and other non-rodent mammals also have higher proportions of interneurons than mice, suggesting that laboratory mice may be an outlier51. However, in the motor cortex humans exhibit an even larger number of interneurons than marmosets, suggesting that an additional change was acquired at some point in our more recent lineage49. Thus, these various neuronal populations have undergone substantial changes in their diversification at various points in human evolution.
The mammalian cerebellar cortex also follows a shared ontogenetic plan, comprising molecular, Purkinje cell and granule cell layers, with each layer containing specific neuronal subtypes including interneurons52. Although the cerebellar cortex has been described as a uniform sheet of invariant circuits, recent cellular profiling in mouse cerebellar cortex has revealed subtle regional and functional heterogeneity in each of these neuronal classes53. The developing human cerebellum shows features that are not seen in other species, including macaques, which in particular involve its protracted development and the appearance of a progenitor pool in the posterior lobe54. It is presently unknown whether cortical cerebellar cell types have further diversified during human evolution, highlighting an area in need of further research.
Evolutionary diversification and specialization have also been described for non-neuronal brain cells, which include oligodendrocytes, astrocytes, microglia, vascular and leptomeningeal types, ependymal cells, and choroid plexus epithelial cells, which together have wide-ranging functions in neuronal transmission, immune responses, providing barriers and homeostasis. A cross-species transcriptomic and morphological analysis demonstrated a greater diversification of subtype diversity and abundance of astrocytes, microglia and oligodendrocytes in humans compared with other primates and mammals55,56,57,58. In an analysis of single-nucleus RNA sequencing from the middle temporal cortex of five primates, including humans and other great apes, microglia showed the greatest transcriptional divergence with evolutionary distance50. Moreover, differentially expressed genes in human non-neuronal types were more likely to be associated with regions of the genome exhibiting signatures of evolutionary selection such as human-accelerated regions (HARs) or human deletions in conserved regions50 (hCONDELs). Cortical astrocytes also show distinct features in humans compared with rodents, marked by their larger cell shapes with more complex processes, and their notably faster propagation of Ca2+ waves59. Additionally, oligodendrocytes exhibit greater gene expression divergence and acceleration compared with neurons58, with a human-specific increase in oligodendrocyte progenitor cells compared with mature oligodendrocytes60, and they are enriched for psychiatric disorder variants58.
Most of the identified evolutionary changes within the human brain seem to come from more subtle shifts in homologous cell-type counterparts, such as differences in cell number, diversification, transcriptomic expression profiles and chromatin accessibility49,61,62, which together may have marked consequences in terms of overall cell numbers and brain size. Although changes in cell number and diversification of cellular homologues are prevalent, recent reports have also identified a number of important qualitative differences, including the emergence of unique cell types, specific loss of certain cells or distinct reallocations of homologous cell types. For instance, various unique interneuron types have recently been identified in the human striatum that are specific for humans or primates51,63. One specific inhibitory neuron subtype, the ‘ivy cell’, is abundantly present in cortical and hippocampal regions in primates, but appears in only the hippocampus in mice and ferrets51. Another example of reallocation of conserved types is found in a population of lateral ganglionic eminence-derived interneurons that migrate primarily to the olfactory bulb in mice. In humans and other primates, this type instead becomes abundant in the deep cortical white matter along the cingulate cortex, suggesting divergence in migration between primates and rodents64.
Specializations in neuronal connectivity
The human neocortex displays structural, functional and subcellular differences in neuronal connectivity when compared to close relatives, as well as between distinct structures within the human brain. Most of these connectivity-related features are typically not shared with other great apes, emphasizing their specific emergence within the human lineage. For instance, humans display higher connectivity between association areas and lower interhemispheric connectivity compared with chimpanzees65. Subcellular differences associated with connectivity, including changes in synaptic connections and dendritic arborization, have also been observed in humans when compared with closely and more distantly related species. Despite constraints imposed by inevitable small sample sizes, these studies appear to show consistent trends across species and cortical regions. For instance, compared to chimpanzees, layer III pyramidal neurons in various human cortical areas contain notably longer and more branched dendrites with relatively more spines, and less profound differences are also observed in spine density66 (Fig. 1b,f). By contrast, differences in spine number, spine densities and dendritic complexity are less profound between macaques and rodents than between humans and macaques, despite their considerably larger evolutionary distance66,67,68,69,70,71 (Fig. 1b,f). In humans and other great apes, connectivity-related changes are also detected across cortical regions; for instance, the prefrontal and temporal cortex exhibit relative increases in spine number and dendritic arborization compared with primary sensory cortices66,67,68,69,70,71 (Fig. 1b,f).
Whilst comprehensive profiling of connectivity-related differences in humans compared with other great apes is technically and ethically challenging, comparisons with other primates and mammals have yielded valuable insights. For instance, proteomic analysis of synaptosomes from the human, macaque and mouse motor cortex revealed a relative enrichment of presynaptic components and microtubules72. This points to variations in molecular composition at individual synapses, which are likely to contribute to human-specific differences in connectivity72. Indeed, functional assays have revealed distinct biophysical properties of both glutamatergic neurons73,74,75,76 and GABAergic (γ-aminobutyric-acid-producing) neurons77,78,79, resulting in differences in transmission capabilities in neural circuits in humans. Moreover, a human-specific modulator of neuronal excitability, encoded by the hominid-specific gene LRRC37B, was recently identified and functions by inhibiting voltage-gated sodium channels80.
The cerebellum in humans also exhibits notable changes in neural connections compared with other species. Cerebellar Purkinje cells in humans exhibit increased dendritic complexity and spine numbers compared with rodents, leading to enhanced cellular computational capacity81. However, it remains to be determined whether these changes are also evident when comparing humans with more closely related species.
Distinct features of neurodevelopment in humans
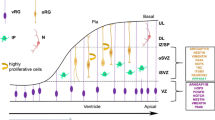
Specializations of the human brain arise during distinct stages of neurodevelopment, a spatiotemporally controlled process whose ontogeny is generally shared between mammals. Neurodevelopment starts with the proliferation of neuroepithelial cells that convert into different populations of neural progenitors (Fig. 3a). These progenitors give rise to various neuronal subtypes that migrate to specific brain regions. In the cortex, the first progenitors that arise are apical radial glia, which continue to expand and produce additional amplifying progenitor types, such as intermediate progenitors and basal or outer radial glia. This pool of cortical progenitors eventually gives rise to cortical neurons, which then migrate towards the outer surface and form the distinct layers of the cortex in an inside-out order. The subsequent development of various neuron types is characterized by a series of drastic morphological changes and the assembly of specialized subcellular structures, a process collectively referred to as neuronal maturation. First, during neuronal migration, neurons grow out a single axon, which bundles with other axons to form distinct tracts projecting to specific brain regions (Fig. 3b). Axon outgrowth is followed by the sequential processes of dendrite formation, synaptogenesis, synaptic pruning and axon myelination, which together enable the establishment and refinement of neuronal circuits.
a, Left, schematic illustration of prolonged neural stem cell expansion, increased progenitor proliferation and delayed supragranular neurogenesis in humans compared with non-human primates95,97,100. Bottom right, scaled cross-species analysis of the timing of neural stem cell expansion91,93,102,103,104,105 and neurogenesis91,92,93. aRG, apical radial glia; bRG, basal radial glia; IP, intermediate progenitor; NEC, neuroepithelium cell; NSC, neural stem cell. b, Left, schematic illustration of neuronal maturation stages in humans and chimpanzees. Humans develop relatively longer axons, more arborized dendrites and more spines per neuron. Bottom right, scaled cross-species analysis of the timing of synaptogenesis132,133,134,145, synaptic pruning132,133,139,145 and myelination135,136,137,138, showing that later maturation stages are more protracted than early maturation stages in humans compared with chimpanzees.
Variations in neurodevelopmental tempo
Although the sequence of neurodevelopmental events is shared amongst mammals, the tempo at which these events unfold greatly varies between species. The protraction of these developmental stages becomes increasingly pronounced from primates to great apes, culminating in a further delayed progression in humans compared with close relatives. At birth, the human brain is 28% of the adult size, compared with 40% in chimpanzees82. A similar trend is observed within hominins, where this proportion has gradually decreased from 38% in Australopithecus to 28% in middle pleistocene pre-modern Homo83. During subsequent postnatal development, humans reach adult brain size later than chimpanzees and the white matter of the prefrontal cortex develops more slowly82,84.
Such developmental delays have often been described as neoteny. This term, derived from Greek meaning ‘extended youth’, was originally used to describe how certain amphibians, such as the axolotl, retain larval traits into adulthood85. Although the term has been extended to also indicate other developmental delays, its varied usage may lead to confusion. Indeed, in humans neoteny is often referred to as a retention of juvenile characteristics, due to our rather immature physical features, such as fine body hair and delicate craniofacial structures, features that also distinguish us from other apes and primates. However, protracted development is already present in primates compared with the next closest mammalian clade made up of rodents and lagomorphs, and apes exhibit a further protraction, yet these species do not exhibit such juvenile features. Moreover, developmental delay is not limited to pre-adult traits persisting into adulthood, but occurs throughout embryonic and postnatal development. Thus, we propose the term ‘bradychrony’, meaning slowed time, to describe developmental delay (Fig. 2a and Box 2). Similarly, certain mammals, such as mice, exhibit accelerated development, which we term ‘tachychrony’ (Fig. 2a and Box 2), probably owing to their unique environmental and seasonal pressures86. Because the degree of bradychrony has progressively increased in human evolutionary history, whereas neotenic features have appeared rather suddenly in our recent history, the mechanisms underlying the two may be distinct.
This divergence in developmental timing may provide the extended window that is necessary for distinct specializations to take shape in the developing human brain. Consistent with this idea, brain growth tempo—defined as the relative rate of brain development to reach neonatal or adult brain weight—is slower in humans compared with our close relatives (Fig. 2b,c and Supplementary Table 1 and 2). The degree of bradychrony of brain growth tempo aligns with what is expected for our brain size, indicating that allometric scaling in primate evolution applies not only to brain size and neuron number26,27,28, but also to the phase in which this is achieved. Nevertheless, it is worth noting that the human brain continues to mature and refine neural circuits until about 30 years of age, with no increase in brain weight after 22 years of age. These later stages of neurodevelopment are particularly extended in humans, potentially leading to a more pronounced bradychrony in overall brain development. Indeed, in humans it takes 30 years to complete brain development, from gastrulation to completion of myelination, which is nearly half our average lifetime, compared with around 30% of the lifespan in chimpanzees and macaque, and around 5% in mouse (Fig. 2d,e and Supplementary Table 3). This delayed maturity until well after birth also enables increased environmental influence on brain development, thereby providing opportunities for enhanced behavioural plasticity and cultural transmission87. Studies on more species are needed to determine whether this profound bradychrony of these late maturation stages also follows allometric scaling or is slower than expected for our species.
Divergent aspects of neurogenesis
How humans obtain their increased cortical neuron numbers, diversification of homologous cell types and unique cell types is also a key question. Over the past decade, many studies have pinpointed various neurodevelopmental mechanisms that distinguish humans from other mammals, such as rodents, and progress has been made in understanding differences between humans and mice during glutamatergic neurogenesis13,88,89,90. However, human and primate-specific differences that deviate from other mammals are only now beginning to emerge. Additionally, whereas glutamatergic neurogenesis has been the subject of much research, attention is also turning to the less well-explored GABAergic neurogenesis.
Investigating human-specific aspects of brain development, such as exploring the timing of embryonic neurodevelopmental stages, is challenging owing to the limited accessibility of fetal tissues of great apes. Compared with other primates, the in vivo neurogenic time window in humans is bradychronic, lasting about four months, which is about twice as long as in macaques and ten times longer than in mice91,92,93 (Fig. 3a). Although the exact duration of non-human ape neurogenesis remains uncertain, ultrasound imaging has revealed a similar rate of brain growth in chimpanzees to humans. However, humans exhibit both a longer phase of growth and a larger brain at the earliest investigated time point of 15 weeks of gestation94, suggesting that the human brain is larger even before the majority of neurons have been produced.
Whereas it is challenging to compare the developing human brain with those of other apes, in vitro models provide an alternative window and have begun to uncover ape and human-specific delays in neurogenic transitions. Two- and three-dimensional differentiation of human and chimpanzee cortical neurons revealed a prolonged proliferative state of progenitors followed by a delayed switch to supragranular identities compared with macaque95 (Fig. 3a). The same study reported no difference in the timing of onset of supragranular neurogenesis between human and chimpanzee95, consistent with the ape-acquired elaboration of laminar architecture seen in vivo that is unchanged in humans. This is also in line with in vivo reports showing that the proliferative phase of basal progenitors is bradychronic in humans compared with rodents, similarly resulting in increased numbers of supragranular neurons91,96,97.
Organoids—in vitro self-organizing neural tissues—also revealed a switch to gliogenesis earlier in chimpanzee than human98, as well as a difference in cell cycle that relates to neurogenic versus proliferative divisions99. Such heterochrony also appears to be present at the earliest stages of neuroepithelial expansion, with human compared to non-human apes showing enhanced proliferative capacity and bradychrony in the switch from symmetrically expanding neuroepithelial cells to radial glia100 (Fig. 3a). Although there are no in vivo data on non-human apes during these early stages, comparisons between humans, macaque and mice show that the time window of tangential cortical expansion, which encompasses the proliferation of neuroepithelial cells and symmetrically dividing apical radial glia—collectively termed neural stem cells101—is bradychronic in humans. Specifically, this period, spanning from neural tube closure until neurogenic onset, takes about 4 weeks in humans91,102, compared to 2.5 weeks in macaque103,104 and 2 days in mouse93,105 (Fig. 3a). Together, the expansion of these neural stem cells results in an increased founder neurogenic progenitor pool and—consistent with in vivo observations and the radial unit hypothesis103—helps explain how human-specific tangential cortical expansion can arise without a change in cortical thickness or relative proportions of cortical layers compared with other apes.
Although GABAergic progenitors and neuron differences in humans compared with other great apes are less well explored, comparisons with other primates and rodents reveal notable specializations that have evolved in the human lineage. Single-cell epigenomic analysis reveals an impressive diversification amongst neural progenitors across brain regions in humans106. Although less is known about the proliferative dynamics of human and other primate telencephalic GABAergic progenitors, the relative proportions of neuron types in the adult neocortex suggests that population dynamics of inhibitory and excitatory neurons are coordinated. For example, primate V1 has vastly higher neuronal density than elsewhere in the neocortex107. The expansion of the progenitor-rich outer subventricular zone proximal to presumptive V1 during mid-gestation could account for increased excitatory neuron numbers108. Interneuron populations also markedly increase in absolute numbers in primate V1109, resulting in similar overall proportions of inhibitory to excitatory neuron ratios to other cortical areas51. Whether this population matching is controlled by differential rates of interneuron neurogenesis timing, migration or cell death is not yet known.
Similar to other neural progenitors and cell types, progenitor classes within the ganglionic eminences, which give rise to telencephalic GABAergic neurons, appear generally well conserved between primates and rodents64. The emergence of human-specific cell types or brain structures is likely to be exceptionally rare, yet some primate- or human-specific features may arise from novel or specialized progenitor domains. For example, a recently-discovered striatal GABAergic interneuron type51 may arise from a specialized progenitor domain in the ganglionic eminences in primates64. Whereas mice have a single detectable initial class for medial ganglionic eminence-derived interneurons destined for striatum, in primates this class diverges early (after conception day 65) into two distinct postmitotic classes, one of which goes on to mature into a specialized TAC3+ population in primates64.
One intriguing evolutionary modification to primate neocortical progenitors entails the possibility that dorsal cortical progenitors give rise to both glutamatergic and GABAergic neurons, rather than exclusively glutamatergic neurons (as is found in rodents). Whereas past evidence for multipotency of primate dorsal progenitors has been mixed110,111,112,113,114, recent in vitro lineage tracing in human organotypic cortical slice cultures has found that a substantial proportion of cortical interneurons and glutamatergic neurons share a common lineage115. This multipotency is relatively rare early in human cortical development and becomes more prevalent at mid-gestational timepoints110,114,116. No clonal relationships between cortical GABAergic and glutamatergic neurons were found when using the same lineage-tracing approach in mice in vivo115,117. Furthermore, recent examination of intrinsic genetic mosaicism revealed a subtype of DLX1+ inhibitory neurons in the human brain that share an origin with excitatory neurons118, suggesting that local proliferation of cortical GABAergic types could be one potential route by which cortical interneuron abundances increase in primates within particular cortical regions.
While novel primate- or human-specific cell types are fascinating, changes in cell proportions, gene expression programmes and spatial reallocations of conserved types are much more prevalent. For example, whereas rodent thalamic nuclei are dominated by glutamatergic neurons, primate thalamus contains high proportions of GABAergic neurons119—in general, GABAergic neuron composition of dorsal thalamic nuclei appears to scale with species complexity120. Although thalamic structures largely derive from the diencephalon, there appears to be some recruitment of GABAergic neurons from other developmental compartments in rodents and non-human primates121,122. Human organotypic slice culture experiments show that streams of GABAergic neurons of telencephalic origin migrate into thalamic nuclei and populate higher-order associative thalamic nuclei123. The developmental origin and cellular composition of the medial pulvinar—another associative thalamic nucleus—remain to be determined, but there is some evidence that it may be initially formed from diencephalic progenitors along the third ventricle, after which there is contribution from a second wave of telencephalic-origin progenitors124. These examples reveal that the cell-type-specific developmental processes that control proliferation and response to guidance cues are likely to be critical for understanding human specializations of cellular diversity.
Divergent aspects of neuronal maturation
As previously highlighted, many connectivity-related specializations in humans are absent in other great apes, emphasizing their relevance in understanding distinct aspects of human brain function. Thus there is growing interest in deciphering how these differences in neuronal connectivity, all of which arise during neuronal maturation stages, come about during development125,126 (Fig. 3b). Technical advancements, many of which have emerged in the past few years, have begun to enable in vitro modelling of these later stages, thereby paving the way for more mechanistic insights127,128,129. Together with in vivo data, a picture is emerging of evolutionary divergence in neuronal maturation—a process that spans from mid-fetal development to early adolescence and appears to be particularly distinctive in humans. A comprehensive comparative transcriptomic analysis in human and macaque samples from early prenatal stage to adulthood reported the most notable differences that occur during maturation stages130. Specifically, the transcriptome was most divergent during mid-fetal development, aligning with axon outgrowth, dendritogenesis and the onset of synaptogenesis, and during early adolescence, when synaptic pruning and circuitry refinement occur130. Another recent study on proteomic analysis of postsynaptic densities in human, macaque and mouse reported that morphologic and functional synapse maturation is bradychronic in humans, mediated by Rho guanine nucleotide exchange factor signalling131.
Moreover, the degree of bradychrony during neuronal maturation appears to be most distinct in humans, with later maturation stages becoming increasingly protracted in humans compared with earlier stages. For instance, initial synaptogenesis spans approximately five years in humans, four years in chimpanzees and three months in macaques132,133,134 (Fig. 3b), which is notably more bradychronic than the neurogenic stages (Fig. 3a). Late neuronal maturation events, such as synaptic pruning and axon tract myelination, take 30 years in humans, and are increasingly more delayed compared with the 10 years in chimpanzees and 4.5 years in macaques132,133,135,136,137,138,139 (Fig. 3b). Further, this pattern shows that the increased degree of slowing during late neuronal maturation is especially marked in humans when set against other great apes, whereas the differences between non-human apes and other primates and mammals is less profound (Fig. 3b). This suggests that the degree of bradychrony during neuronal maturation in humans may be the result of a relatively recent evolutionary event. Moreover, a recent report found that the timing of synaptogenesis in mice and macaques was remarkably similar, contradicting the long-held belief that the timing of these processes is primarily related to a species’ lifespan132. It should be noted, however, that different timeframes for synaptogenesis in mice have been documented in another study140.
Although the appearance of dendritic and synaptic specializations are relatively more explored, humans also develop extended axonal projections as reflected by increased white matter volume40,41. In human neuron cultures, axon formation was found to be bradychronic141,142 and to proceed through an additional developmental stage involving an increase in microtubule remodelling and relocation of axon initial segment proteins from the distal axon (classified as stage 3a) to the proximal axon (stage 3b, corresponding to the classic stage 3143), which has not been reported in other species142. Another study in human neuron cultures revealed that the onset of axon development is mediated by centrosomes, with the axon initial segment protein TRIM46 localizing to centrosomes in human cells, but not in rodent and monkey cells, before it relocates to the axon as it develops144. Further research is needed to shed light on divergent aspects of axon development that underlie related specializations in the human brain.
In addition to species-specific variations, spatial differences in neuronal maturation tempo within the human brain are also evident, and are likely to contribute to regional differences in neuronal connectivity. Indeed, neuronal maturation is more bradychronic in neocortical regions that have more specialized functions and higher synaptic densities133. For instance, development in the prefrontal cortex is particularly slow, with synaptogenesis peaking 15 months after birth and synaptic pruning continuing up to 30 years, probably contributing to the cognitive and behavioural changes that are observed during childhood and adolescence133. By comparison, the auditory cortex has a faster tempo, with synaptogenesis peaking 3 months after birth and synaptic pruning extending until 12 years of age133. These regional differences in neuronal maturation tempo contrast with chimpanzees and macaques, in which synaptogenesis is more synchronized across cortical regions133,145. However, similar to humans, chimpanzees do exhibit relatively bradychronic dendritic development in the prefrontal cortex compared with other brain regions145.
Thus, the degree of bradychrony, either across brain regions or between different species, is associated with increased connectivity and cognitive functions, sparking an emerging interest in exploring the mechanisms underlying developmental tempo differences. An important insight is that differences in timing appear to be driven by cell-intrinsic mechanisms, as the species-specific tempo of developing cells is maintained when human cortical neurons are xenografted into the mouse brain or co-cultured with non-human primate neurons95,129,141. While the nature of differences in timing between primates is still unclear, recent comparison of human and mouse motor neuron differentiation revealed a correlation with protein stability146, and comparative studies of the segmentation clock have similarly demonstrated a correlation with protein turnover147,148. Besides global timing mechanisms, the varying degree of bradychrony of neuronal maturation compared with other developmental stages suggests the existence of stage-specific factors influencing tempo. Although the exact mechanism of human-specific bradychrony of neuronal maturation is unknown, a recent study comparing human and mouse revealed a slower pattern of mitochondria development and metabolism in human neurons, with neuronal maturation being accelerated upon stimulation of mitochondrial metabolic activity149. Whether this mechanism is specific for humans or shared with other primates is still unclear. As discussed below, epigenetic changes may also drive the pace of neuronal maturation150. Further investigations are needed to understand global and stage-specific mechanisms that drive the tempo of neurodevelopment, as well as to what extent these contribute to human-specific specializations versus those shared with other primates.
Molecular mechanisms of human features
The human genome differs from those of other primates owing to nucleotide substitutions, short insertions and deletions (indels), and larger structural variants (losses, gains, inversions, translocations and rearrangements of DNA) that arise through processes such as transposon activity, recombination, replication and repair151,152,153,154,155,156. With the generation of telomere-to-telomere genome assemblies157, we are approaching a complete list of all fixed differences between modern humans and our closest relatives. Research is also shedding light on how these genetic changes alter protein function and gene-regulatory networks, which is key in establishing their roles in human-specific aspects of neurodevelopment and neurological disease.
Divergent proteins
Both gene duplications and modifications to coding sequences have altered the human proteome. Amino acid substitutions and indels have altered about one third of proteins in humans and a similar number in chimpanzees since our lineages split156. Further proteome differences have arisen through gene birth158,159, duplications, fusions and truncations155,160,161. In several cases, these protein-coding changes have been linked to distinct characteristics of the human brain. For instance, a single lysine-to-arginine substitution in transketolase-like 1 (TKTL1)—which arose after the divergence of modern humans from Neanderthals—increases neural production by basal radial glia in human cerebral organoids and in mice, potentially through greater production of fatty acids required for the outgrowth of processes on radial glia162. Radial glial biology has also been modified by three human-specific paralogues of the NOTCH2 receptor (NOTCH2NL), which promote clonal expansion of neural progenitors and delay their differentiation via inhibition of Delta–Notch interactions and activation of the Notch pathway163,164. NOTCH2NL genes serve as breakpoints for human polymorphic structural variants that are associated with variation in brain size and autism, supporting the hypothesis that NOTCH2NL contributed to cortical expansion during human evolution.
Another set of gene duplication events with effects on brain development is SRGAP2. SRGAP2C evolved from gene duplication before the major evolutionary increase of brain size in humans165. Mice expressing the human-specific SRGAP2C showed enhanced excitatory synaptogenesis in upper layer neurons, increased local and long-range cortico-cortical connections and improved learning abilities of cortex-dependent tasks165. Furthermore, the human-specific paralogues SRGAP2B and SRGAP2C slow synaptic maturation by antagonizing and downregulating the ancestral protein encoded by SRGAP2A. This results in prolonged upregulation of postsynaptic SYNGAP1166, modulating neuronal spine formation and promoting bradychronic maturation161,167,168,169. Another example is Rho GTPase-activating protein 11B (ARHGAP11B), a human-specific gene that enlarges the neocortex and induces gyrification when expressed in mice and marmosets170,171. Thus, although most of the proteome is identical between humans and chimpanzees, the search for genes underlying human neurodevelopmental specializations has yielded a number of variants and gene duplications that modify intracellular signalling in the human cortex with effects on downstream metabolic and transcriptional processes, anatomy and developmental timing.
Divergent gene-regulatory networks
Human genetic variants also affect non-coding DNA that contains regulatory elements such as enhancers, repressors and insulators172,173,174,175. For example, during early and mid-fetal human development, there is a spatially restricted patterning of retinoic acid in the prefrontal cortex that is important for promoting enhanced intra-prefrontal cortex dendritic synaptogenesis and prefrontal cortex–mediodorsal thalamus connectivity176. The onset of synaptogenesis in the prefrontal cortex during mid-fetal development aligns with the local, transient and laminar-specific upregulation of CBLN2, NRXN and GRID in humans, an expression pattern that is not observed in macaque177. Increased levels of CBLN2 in the prefrontal cortex promote spine formation, a regulatory effect that is shared between human and chimpanzees, driven by a hominin-specific deletion in SOX5-binding sites within a retinoic acid-responsive CBLN2 enhancer177. Thousands of other human-derived regulatory elements have been proposed on the basis of cross-species differences in gene expression, chromatin accessibility and histone modifications in post mortem brain tissues178,179,180,181,182,183,184,185,186, neuronal cell lines187 and cerebral organoids17,98. Many of these elements have human variants that may explain their differential function in the human genome—for example, with changes in binding of transcription factors or chromatin modifiers172.
The fastest-evolving regulatory elements are HARs, conserved non-coding sequences with an excess of nucleotide changes in the human genome188,189,190. Many HARs function as gene-regulatory enhancers in neurodevelopment191. Using massively parallel reporter assays, hundreds of HARs have been validated as enhancers where the modern human sequences show differential activity188,192,193,194,195,196, and dozens show differential enhancer activity in mice191,197. In the case of HARE5, a human-accelerated enhancer of the WNT receptor FRIZZLED8, differential activity has been linked to human brain features. Compared to the chimpanzee sequence, human HARE5 accelerates the cell cycle in neural progenitors and increases brain size in transgenic mice198. Thus, innovations in human brain development are now being linked not only to variants affecting proteins, but also to variants in gene-regulatory elements.
Beyond point mutations, structural variants have translocated, duplicated, and deleted thousands of regulatory elements during human evolution199,200. Not surprisingly, these large-scale changes in regulatory information can lead to developmental phenotypes. For instance, loss of a hCONDEL that regulates GADD45G is correlated with expansion of specific brain regions in humans199. Structural variants also alter the relationship between nearby genes and regulatory elements by reorganizing and modifying the linear genome161,201. Furthermore, indels and substitutions affecting chromatin boundaries can reorganize three-dimensional chromatin without major changes in the genomic distance between regulatory elements and promoters. These disruptions to three-dimensional genome folding can separate regulatory elements from their target genes, thereby reducing their ability to control gene expression, and conversely they may bring together gene promoters with regulatory elements from which they were previously insulated (enhancer hijacking). Indeed, HARs are enriched in loci with structural variants that alter genome folding202, suggesting that their accelerated evolution could be attributed, in part, to their being hijacked enhancers.
Other mechanisms that could explain regulatory elements that lack cis sequence variation yet show epigenetic differences in the human brain include genetic variants in upstream transcription factors203, altered cell-type specificity of these factors204, and differences in the chemical environment inside human cells. A role for epigenetics in controlling neurodevelopmental timing in humans150 as well as more distant species205 is emerging. A study using in vitro human neuron cultures identified an epigenetic barrier that controls timing of neuronal maturation, which is mediated by the gradual release of the epigenetic factors EHZ2, EHMT1, EHMT2 and DOT1L, which are otherwise retained in neuronal precursor cells150. Post-transcriptional regulation has also contributed to evolution of the human brain206,207,208. In sum, changes to regulatory element sequences, trans factors and genome organization all have roles in the evolution of human gene-regulatory networks.
Elucidating the functional effects of this re-wiring on neurodevelopment is an active area of current research. An important future direction is studying regulatory elements in their endogenous genomic context, as opposed to probing them via reporter assays. This will become easier as genome-editing tools with nucleotide resolution and high-throughput capabilities continue to be developed. The demonstration that deletion of SOX5-binding sites within a CBLN2 enhancer causally contribute to its human-specific upregulation in the prefrontal cortex177 exemplifies the discovery potential of endogenous editing.
Outlook
Various specializations of the human brain have been identified in recent years. Some of these features are likely to be adaptive, such as the vast increase in neuron numbers, providing a fitness advantage and enabling more complex cognitive and social abilities. Many of these adaptations also appear to be shared with other primates, such as the increase in neuronal density and the relative increase of supragranular layers, which may lead to advanced cognitive functions that are already evident in primates. Other specializations involve those that appear early in evolution but show a profound divergence in the human lineage, including neocortical and cerebellar expansion emerging during neurogenesis, and the increased dendritic arborization, synapse number and axon projections established during neuronal maturation. The human-specific features set during neuronal maturation stages remain relatively unknown, largely owing to technical hurdles in modelling these stages. Emerging evidence is highlighting the role of specific gene-regulatory networks and developmental tempo in driving variations in synapse numbers. Further research is needed to further explore the downstream intracellular programmes that they act on, as well as how evolutionary differences in axon projections and dendritic arborization come about during development. An exciting avenue for future investigations would be to examine the possible influence of neuronal activity during these developmental processes, because in non-primates, neuronal activity in established circuits is known to feedback by modulating gene transcription and subsequent neurodevelopment209,210,211,212,213.
In addition to specializations with a profound divergence in the human lineage, there is an accompanying marked difference in the degree of bradychrony in humans compared with our close relatives. Although the increased bradychrony in humans may imply that an extended timeframe is important for developing a more expanded brain with divergent features, more experimental evidence is needed to support this. Methodologies to perturb species-specific timing are being developed214, and will be instrumental in gaining functional insights. Thus, further research is needed to identify mechanisms that modulate developmental timing, as well as their relevance for establishing the different human-specific specializations.
The presence of specializations in non-cortical structures in the human brain is becoming increasingly apparent, and offers another intriguing direction for future research. Notably, modifications to subcortical structures are pronounced in regions that are coupled to distributed networks of higher-order association cortex, suggesting coordinated evolution. For example, non-motor and non-sensory structures in the human cerebellum have expanded in conjunction with the expansion of human higher-order neocortical networks36,215. Similarly, the associative mediodorsal and medial pulvinar nuclei of the thalamus have expanded in primate evolution216, a feat that may in part have been enabled by adaptations to the expansion and migration properties of telencephalic progenitors124. Applying cell-type resolved lineage-tracing technologies115,217 to developing primates in vivo could help to establish how and when primate-specific adaptations to progenitors arise, which could then be further modelled in vitro. Furthermore, technologies that enable cell-type-resolved readouts of synaptic connectivity218 or projection patterns219 in primates will be key for understanding the relationship between expanded neuronal populations and changes in connectivity properties. Moreover, investigating features that greatly diverge in humans and rodents, such as the CA1 region of the hippocampus—which is more packed with pyramidal neurons of distinct morphologies in humans—in other primates could enhance our understanding of the evolutionary context of these features220.
The emerging mechanistic understanding of human brain evolution largely stems from various technical advances. Much progress has been made in developing human in vitro models, including neural cultures and brain organoids, with varying degrees of complexity. A growing body of literature using these methods has consistently demonstrated that these models recapitulate key stages of neurodevelopment, from early proliferation to neurogenesis and gliogenesis, and that they successfully model the anticipated timing differences across species. The latest developments in this field include in vitro systems that model later stages of human neurodevelopment, including optimized methods for long-term brain cultures, in vivo transplantations and assembloids, as well as the modelling of more brain structures, such as choroid plexus and cerebellum127,128,129,221,222,223,224. Future advances in in vitro models may include functional vascularization of organoids, recapitulating long-range axon tracts and modelling more brain structures. As these tools become more commonplace, further comparative studies would also benefit from more individuals per species to increase confidence that any identified differences represent species-specific features rather than technical artefacts.
Additionally, important advances in high-throughput single-cell profiling techniques, such as single-cell RNA sequencing, ATAC (assay for transposase-accessible chromatin) sequencing and proteomics, have enabled scaling up cell numbers and reads per cell, improved quality of readouts, the integration of multiple modalities and spatiotemporal reconstructions225,226,227. This has led to a large number of detailed maps of genetic, gene accessibility and transcriptomic patterns at different neurodevelopmental stages across species98,228,229, as well as the increased identification of human-specific genomic and gene-regulatory changes during neurodevelopment. Although some have been functionally explored, further investigations are needed to uncover how these changes are mechanistically linked to human-specific specializations arising during neurodevelopment. Moreover, recent technical developments in imaging and microscopy, such as three-dimensional live imaging and clearing methods, have begun to unravel the subcellular structures and temporal dynamics of developing human neurons230,231,232,233,234. This multidisciplinary application of new techniques will continue to be at the foundation of understanding the distinct aspects of human brain development and function.
References
Lage, C. A., Wolmarans, D. W. & Mograbi, D. C. An evolutionary view of self-awareness. Behav. Process. 194, 104543 (2022).
Roth, G. & Dicke, U. Origin and evolution of human cognition. Prog. Brain Res. 250, 285–316 (2019).
Mouritsen, H., Heyers, D. & Güntürkün, O. The neural basis of long-distance navigation in birds. Annu. Rev. Physiol. 78, 133–154 (2015).
Jarvis, E. D. Evolution of vocal learning and spoken language. Science 366, 50–54 (2019).
Whiten, A., Hinde, R. A., Laland, K. N. & Stringer, C. B. Culture evolves. Philos. Trans. R. Soc. B 366, 938–948 (2011).
Varki, A. & Gagneux, P. in On Human Nature: Biology, Psychology, Ethics, Politics, and Religion (eds Tibayrenc, M. et al.) 151–160 (Academic, 2017).
Pattabiraman, K., Muchnik, S. K. & Sestan, N. The evolution of the human brain and disease susceptibility. Curr. Opin. Genet. Dev. 65, 91–97 (2020).
Martinez, P. & Sprecher, S. G. Of circuits and brains: the origin and diversification of neural architectures. Front. Ecol. Evol. 8, 82 (2020).
Martín-Durán, J. M. & Hejnol, A. A developmental perspective on the evolution of the nervous system. Dev. Biol. 475, 181–192 (2019).
Vallender, E. J. Genetics of human brain evolution. Prog. Brain Res. 250, 3–39 (2019).
Uzquiano, A. & Arlotta, P. Brain organoids: the quest to decipher human-specific features of brain development. Curr. Opin. Genet. Dev. 75, 101955 (2022).
Vinsland, E. & Linnarsson, S. Single-cell RNA-sequencing of mammalian brain development: insights and future directions. Development 149, dev200180 (2022).
Kelley, K. W. & Pașca, S. P. Human brain organogenesis: toward a cellular understanding of development and disease. Cell 185, 42–61 (2022).
Perkel, J. M. Single-cell analysis enters the multiomics age. Nature 595, 614–616 (2021).
Chiaradia, I. & Lancaster, M. A. Brain organoids for the study of human neurobiology at the interface of in vitro and in vivo. Nat. Neurosci. 23, 1496–1508 (2020).
Benito-Kwiecinski, S. & Lancaster, M. A. Brain organoids: human neurodevelopment in a dish. Cold Spring Harb. Perspect. Biol. 12, a035709 (2019).
Pollen, A. A. et al. Establishing cerebral organoids as models of human-specific brain evolution. Cell 176, 743–756.e17 (2019).
Jerison, H. J. The theory of encephalization. Ann. NY Acad. Sci. 299, 146–160 (1977).
Beaudet, A., Du, A. & Wood, B. Evolution of the modern human brain. Prog. Brain Res. 250, 219–250 (2019).
Du, A. et al. Pattern and process in hominin brain size evolution are scale-dependent. Proc. R. Soc. B 285, 20172738 (2018).
Tilot, A. K. et al. The evolutionary history of common genetic variants influencing human cortical surface area. Cereb. Cortex 31, 1873–1887 (2020).
Finlay, B. L. & Darlington, R. B. Linked regularities in the development and evolution of mammalian brains. Science 268, 1578–1584 (1995).
Finlay, B. L., Darlington, R. B. & Nicastro, N. Developmental structure in brain evolution. Behav. Brain Sci. 24, 263–278 (2001).
Montgomery, S. H., Mundy, N. I. & Barton, R. A. Brain evolution and development: adaptation, allometry and constraint. Proc. R. Soc. B 283, 20160433 (2016).
Dunbar, R. I. M. Neocortex size as a constraint on group size in primates. J. Hum. Evol. 22, 469–493 (1992).
Dicke, U. & Roth, G. Neuronal factors determining high intelligence. Philos. Trans. R. Soc B 371, 20150180 (2016).
Herculano-Houzel, S. The human brain in numbers: a linearly scaled-up primate brain. Front. Hum. Neurosci. 3, 31 (2009).
Azevedo, F. A. C. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled‐up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Gómez-Robles, A., Hopkins, W. D., Schapiro, S. J. & Sherwood, C. C. Relaxed genetic control of cortical organization in human brains compared with chimpanzees. Proc. Natl Acad. Sci. USA 112, 14799–14804 (2015).
Hill, J. et al. Similar patterns of cortical expansion during human development and evolution. Proc. Natl Acad. Sci. USA 107, 13135–13140 (2010).
Arnsten, A. F. T. Stress signalling pathways that impair prefrontal cortex structure and function. Nat. Rev. Neurosci. 10, 410–422 (2009).
Holmes, A. J. et al. Prefrontal functioning during context processing in schizophrenia and major depression: An event-related fMRI study. Schizophr. Res. 76, 199–206 (2005).
Rougier, N. P., Noelle, D. C., Braver, T. S., Cohen, J. D. & O’Reilly, R. C. Prefrontal cortex and flexible cognitive control: rules without symbols. Proc. Natl Acad. Sci. USA 102, 7338–7343 (2005).
León, M. S. Pde et al. The primitive brain of early Homo. Science 372, 165–171 (2021).
Sereno, M. I. et al. The human cerebellum has almost 80% of the surface area of the neocortex. Proc. Natl Acad. Sci. USA 117, 19538–19543 (2020).
Buckner, R. L., Krienen, F. M., Castellanos, A., Diaz, J. C. & Yeo, B. T. T. The organization of the human cerebellum estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 2322–2345 (2011).
Balsters, J. H. et al. Evolution of the cerebellar cortex: the selective expansion of prefrontal-projecting cerebellar lobules. Neuroimage 49, 2045–2052 (2010).
Sultan, F. & Braitenberg, V. Shapes and sizes of different mammalian cerebella. A study in quantitative comparative neuroanatomy. J. Hirnforsch. 34, 79–92 (1993).
Zhang, K. & Sejnowski, T. J. A universal scaling law between gray matter and white matter of cerebral cortex. Proc. Natl Acad. Sci. USA 97, 5621–5626 (2000).
Schoenemann, P. T., Sheehan, M. J. & Glotzer, L. D. Prefrontal white matter volume is disproportionately larger in humans than in other primates. Nat. Neurosci. 8, 242–252 (2005). A cross-species analysis of 11 primate species, including human and other great apes, shows a profound disproportionate increase of relative prefrontal white matter volume in humans.
Donahue, C. J., Glasser, M. F., Preuss, T. M., Rilling, J. K. & Essen, D. C. V. Quantitative assessment of prefrontal cortex in humans relative to nonhuman primates. Proc. Natl Acad. Sci. USA 115, E5183–E5192 (2018).
von Economo, C. Cellular Structure of the Human Cerebral Cortex (Karger, 2009).
Preuss, T. M. & Wise, S. P. Evolution of prefrontal cortex. Neuropsychopharmacology 47, 3–19 (2022).
Hutsler, J. J., Lee, D.-G. & Porter, K. K. Comparative analysis of cortical layering and supragranular layer enlargement in rodent carnivore and primate species. Brain Res. 1052, 71–81 (2005).
Balaram, P. & Kaas, J. H. Towards a unified scheme of cortical lamination for primary visual cortex across primates: insights from NeuN and vGLUT2 immunoreactivity. Front. Neuroanat. 8, 81 (2014).
Sousa, A. Ade. et al. Comparative cytoarchitectural analyses of striate and extrastriate areas in hominoids. Cereb. Cortex 20, 966–981 (2010). Refs. 44 and 46 show that a relative increase of supragranular layers is observed in great apes compared with other primates, but is not further increased in humans compared with other apes.
Berg, J. et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature 598, 151–158 (2021).
Loomba, S. et al. Connectomic comparison of mouse and human cortex. Science 377, eabo0924 (2022).
Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 111–119 (2021).
Jorstad, N. L. et al. Comparative transcriptomics reveals human-specific cortical features. Science 382, eade9516 (2023).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020). This study shows that the ivy cell, an interneuron subtype, is present in the cortex and hippocampus in primates, but localizes to the hippocampus only in mice and ferrets.
Cerminara, N. L., Lang, E. J., Sillitoe, R. V. & Apps, R. Redefining the cerebellar cortex as an assembly of non-uniform Purkinje cell microcircuits. Nat. Rev. Neurosci. 16, 79–93 (2015).
Kozareva, V. et al. A transcriptomic atlas of mouse cerebellar cortex comprehensively defines cell types. Nature 598, 214–219 (2021).
Haldipur, P. et al. Spatiotemporal expansion of primary progenitor zones in the developing human cerebellum. Science 366, 454–460 (2019).
Falcone, C. et al. Redefining varicose projection astrocytes in primates. Glia 70, 145–154 (2022).
Geirsdottir, L. et al. Cross-species single-cell analysis reveals divergence of the primate microglia program. Cell 179, 1609–1622.e16 (2019).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Berto, S. et al. Accelerated evolution of oligodendrocytes in the human brain. Proc. Natl Acad. Sci. USA 116, 24334–24342 (2019).
Oberheim, N. A. et al. Uniquely hominid features of adult human astrocytes. J. Neurosci. 29, 3276–3287 (2009).
Caglayan, E. et al. Molecular features driving cellular complexity of human brain evolution. Nature 620, 145–153 (2023).
Fang, R. et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56–62 (2022).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Sousa, A. M. M. et al. Molecular and cellular reorganization of neural circuits in the human lineage. Science 358, 1027–1032 (2017).
Schmitz, M. T. et al. The development and evolution of inhibitory neurons in primate cerebrum. Nature 603, 871–877 (2022).
Ardesch, D. J. et al. Evolutionary expansion of connectivity between multimodal association areas in the human brain compared with chimpanzees. Proc. Natl Acad. Sci. USA 116, 201818512 (2019).
Bianchi, S. et al. Dendritic morphology of pyramidal neurons in the chimpanzee neocortex: regional specializations and comparison to humans. Cereb. Cortex 23, 2429–2436 (2013). This study demonstrates that pyramidal neurons in different neocortical structures exhibit increased dendritic arborization and synapse number per neuron in humans compared with chimpanzees.
Mohan, H. et al. Dendritic and axonal architecture of individual pyramidal neurons across layers of adult human neocortex. Cereb. Cortex 25, 4839–4853 (2015).
Elston, G. N. et al. Specializations of the granular prefrontal cortex of primates: implications for cognitive processing. Anat. Rec. A 288A, 26–35 (2006).
Elston, G. N., Benavides-Piccione, R. & DeFelipe, J. A study of pyramidal cell structure in the cingulate cortex of the macaque monkey with comparative notes on inferotemporal and primary visual cortex. Cereb. Cortex 15, 64–73 (2005).
Elston, G. N., Benavides-Piccione, R. & DeFelipe, J. The pyramidal cell in cognition: a comparative study in human and monkey. J. Neurosci. 21, RC163 (2001).
Jacobs, B. et al. Regional dendritic and spine variation in human cerebral cortex: a quantitative Golgi study. Cereb. Cortex 11, 558–571 (2001).
Koopmans, F. et al. Comparative hippocampal synaptic proteomes of rodents and primates: differences in neuroplasticity-related proteins. Front. Mol. Neurosci. 11, 364 (2018).
Beaulieu-Laroche, L. et al. Allometric rules for mammalian cortical layer 5 neuron biophysics. Nature 600, 274–278 (2021).
Gidon, A. et al. Dendritic action potentials and computation in human layer 2/3 cortical neurons. Science 367, 83–87 (2020).
Beaulieu-Laroche, L. et al. Enhanced dendritic compartmentalization in human cortical neurons. Cell 175, 643–651.e14 (2018).
Eyal, G. et al. Unique membrane properties and enhanced signal processing in human neocortical neurons. eLife 5, e16553 (2016).
Lee, B. R. et al. Signature morphoelectric properties of diverse GABAergic interneurons in the human neocortex. Science 382, eadf6484 (2023).
Chartrand, T. et al. Morphoelectric and transcriptomic divergence of the layer 1 interneuron repertoire in human versus mouse neocortex. Science 382, eadf0805 (2023).
Molnár, G. et al. Human pyramidal to interneuron synapses are mediated by multi-vesicular release and multiple docked vesicles. eLife 5, e18167 (2016).
Libé-Philippot, B. et al. LRRC37B is a species-specific regulator of voltage-gated channels and excitability in human cortical neurons. Cell 186, 5766–5783.e25 (2023).
Masoli, S. et al. Human outperform mouse Purkinje cells in dendritic complexity and computational capacity. Commun. Biol. 7, 5 (2024). This comparative study shows that Purkinje cells exhibit increased dendritic arborization and spine numbers in humans compared with rodents.
DeSilva, J. & Lesnik, J. Chimpanzee neonatal brain size: Implications for brain growth in Homo erectus. J. Hum. Evol. 51, 207–212 (2006).
DeSilva, J. M. & Lesnik, J. J. Brain size at birth throughout human evolution: a new method for estimating neonatal brain size in hominins. J. Hum. Evol. 55, 1064–1074 (2008).
Sakai, T. et al. Differential prefrontal white matter development in chimpanzees and humans. Curr. Biol. 22, 171 (2012).
Kollmann, J. Das Ueberwintern von europäischen Frosch- und Tritonlarven und die Umwandlung des mexikanischen Axolotl. Verh. Naturforsch. Ges. Basel 7, 387–398 (1885).
Berry, R. J. & Bronson, F. H. Life history and bioeconomy of the house mouse. Biol. Rev. 67, 519–550 (1992).
Guyer, A. E., Pérez‐Edgar, K. & Crone, E. A. Opportunities for neurodevelopmental plasticity from infancy through early adulthood. Child Dev. 89, 687–697 (2018).
Libé-Philippot, B. & Vanderhaeghen, P. Cellular and molecular mechanisms linking human cortical development and evolution. Annu Rev Genet 55, 555–581 (2021).
Villalba, A., Götz, M. & Borrell, V. The regulation of cortical neurogenesis. Curr. Top. Dev. Biol. 142, 1–66 (2020).
Miller, D. J., Bhaduri, A., Sestan, N. & Kriegstein, A. Shared and derived features of cellular diversity in the human cerebral cortex. Curr. Opin. Neurobiol. 56, 117–124 (2019).
Stepien, B. K., Vaid, S. & Huttner, W. B. Length of the neurogenic period—a key determinant for the generation of upper-layer neurons during neocortex development and evolution. Front. Cell Dev. Biol. 9, 676911 (2021).
Silbereis, J. C., Pochareddy, S., Zhu, Y., Li, M. & Sestan, N. The cellular and molecular landscapes of the developing human central nervous system. Neuron 89, 248–268 (2016).
Haubensak, W., Attardo, A., Denk, W. & Huttner, W. B. Neurons arise in the basal neuroepithelium of the early mammalian telencephalon: a major site of neurogenesis. Proc. Natl Acad. Sci. USA 101, 3196–3201 (2004).
Sakai, T. et al. Fetal brain development in chimpanzees versus humans. Curr. Biol. 22, R791–R792 (2012).
Otani, T., Marchetto, M. C., Gage, F. H., Simons, B. D. & Livesey, F. J. 2D and 3D stem cell models of primate cortical development identify species-specific differences in progenitor behavior contributing to brain size. Cell Stem Cell 18, 467–480 (2016).
Wang, X., Tsai, J.-W., LaMonica, B. & Kriegstein, A. R. A new subtype of progenitor cell in the mouse embryonic neocortex. Nat. Neurosci. 14, 555–561 (2011).
Hansen, D. V., Lui, J. H., Parker, P. R. L. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010).
Kanton, S. et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422 (2019).
Mora-Bermúdez, F. et al. Differences and similarities between human and chimpanzee neural progenitors during cerebral cortex development. eLife 5, e18683 (2016).
Benito-Kwiecinski, S. et al. An early cell shape transition drives evolutionary expansion of the human forebrain. Cell 184, 2084–2102.e19 (2021).
Taverna, E., Götz, M. & Huttner, W. B. The cell biology of neurogenesis: toward an understanding of the development and evolution of the neocortex. Annu. Rev. Cell Dev. Biol. 30, 465–502 (2014).
O’Rahilly, R. & Müller, F. Developmental stages in human embryos: revised and new measurements. Cells Tissues Organs 192, 73–84 (2010).
Rakic, P. Specification of cerebral cortical areas. Science 241, 170–176 (1988).
Davignon, R. W., Parker, R. M. & Hendrickx, A. G. Staging of the early embryonic brain in the baboon (Papio cynocephalus) and rhesus monkey (Macaca mulatta). Anat. Embryol. 159, 317–334 (1980).
Pryor, S. E., Massa, V., Savery, D., Greene, N. D. E. & Copp, A. J. Convergent extension analysis in mouse whole embryo culture. Methods Mol. Biol. 839, 133–146 (2012).
Ziffra, R. S. et al. Single-cell epigenomics reveals mechanisms of human cortical development. Nature 598, 205–213 (2021).
Collins, C. E., Airey, D. C., Young, N. A., Leitch, D. B. & Kaas, J. H. Neuron densities vary across and within cortical areas in primates. Proc. Natl Acad. Sci. USA 107, 15927–15932 (2010).
Smart, I. H. M., Dehay, C., Giroud, P., Berland, M. & Kennedy, H. Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb. Cortex 12, 37–53 (2002).
Krienen, F. M. et al. A marmoset brain cell census reveals regional specialization of cellular identities. Sci. Adv. 9, eadk3986 (2023).
Ma, T. et al. Subcortical origins of human and monkey neocortical interneurons. Nat. Neurosci. 16, 1588–1597 (2013).
Jakovcevski, I., Mayer, N. & Zecevic, N. Multiple origins of human neocortical interneurons are supported by distinct expression of transcription factors. Cereb. Cortex 21, 1771–1782 (2011).
Yu, X. & Zecevic, N. Dorsal radial glial cells have the potential to generate cortical interneurons in human but not in mouse brain. J. Neurosci. 31, 2413–2420 (2011).
Petanjek, Z., Berger, B. & Esclapez, M. Origins of cortical GABAergic neurons in the cynomolgus monkey. Cereb. Cortex 19, 249–262 (2008).
Letinic, K., Zoncu, R. & Rakic, P. Origin of GABAergic neurons in the human neocortex. Nature 417, 645–649 (2002).
Delgado, R. N. et al. Individual human cortical progenitors can produce excitatory and inhibitory neurons. Nature 601, 397–403 (2022).
Hansen, D. V. et al. Non-epithelial stem cells and cortical interneuron production in the human ganglionic eminences. Nat. Neurosci. 16, 1576–1587 (2013).
Bandler, R. C. et al. Single-cell delineation of lineage and genetic identity in the mouse brain. Nature 601, 404–409 (2022).
Chung, C. et al. Cell-type-resolved mosaicism reveals clonal dynamics of the human forebrain. Nature 629, 384–392 (2024).
Bakken, T. E. et al. Single-cell and single-nucleus RNA-seq uncovers shared and distinct axes of variation in dorsal LGN neurons in mice, non-human primates, and humans. eLife 10, e64875 (2021).
Arcelli, P., Frassoni, C., Regondi, M. C., Biasi, S. D. & Spreafico, R. GABAergic neurons in mammalian thalamus: a marker of thalamic complexity? Brain Res. Bull. 42, 27–37 (1997).
Jager, P. et al. Dual midbrain and forebrain origins of thalamic inhibitory interneurons. eLife 10, e59272 (2021).
Jager, P. et al. Tectal-derived interneurons contribute to phasic and tonic inhibition in the visual thalamus. Nat. Commun. 7, 13579 (2016).
Letinic, K. & Rakic, P. Telencephalic origin of human thalamic GABAergic neurons. Nat. Neurosci. 4, 931–936 (2001).
Rakić, P. & Sidman, R. L. Telencephalic origin of pulvinar neurons in the fetal human brain. Z. Anat. Entwickl. Gesch. 129, 53–82 (1969).
Wallace, J. L. & Pollen, A. A. Human neuronal maturation comes of age: cellular mechanisms and species differences. Nat. Rev. Neurosci. 25, 7–29 (2024).
Vanderhaeghen, P. & Polleux, F. Developmental mechanisms underlying the evolution of human cortical circuits. Nat. Rev. Neurosci. 24, 213–232 (2023).
Revah, O. et al. Maturation and circuit integration of transplanted human cortical organoids. Nature 610, 319–326 (2022).
Giandomenico, S. L. et al. Cerebral organoids at the air-liquid interface generate diverse nerve tracts with functional output. Nat. Neurosci. 22, 669–679 (2019).
Linaro, D. et al. Xenotransplanted human cortical neurons reveal species-specific development and functional integration into mouse visual circuits. Neuron 104, 972–986.e6 (2019).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018).
Wang, L. et al. A cross-species proteomic map reveals neoteny of human synapse development. Nature 622, 112–119 (2023).
Wildenberg, G., Li, H. & Kasthuri, N. The development of synapses in mouse and macaque primary sensory cortices. Preprint at bioRxiv https://doi.org/10.1101/2023.02.15.528564 (2023).
Huttenlocher, P. R. & Dabholkar, A. S. Regional differences in synaptogenesis in human cerebral cortex. J. Comp. Neurol. 387, 167–178 (1997).
Rakic, P., Bourgeois, J.-P., Eckenhoff, M. F., Zecevic, N. & Goldman-Rakic, P. S. Concurrent overproduction of synapses in diverse regions of the primate cerebral cortex. Science 232, 232–235 (1986).
Zeiss, C. J. Comparative milestones in rodent and human postnatal central nervous system development. Toxicol. Pathol. 49, 1368–1373 (2021).
Meng, Y., Jiang, J., Bachevalier, J., Zhang, X. & Chan, A. W. S. Developmental whole brain white matter alterations in transgenic Huntington’s disease monkey. Sci. Rep. 7, 379 (2017).
Sakai, T. et al. Developmental trajectory of the corpus callosum from infancy to the juvenile stage: comparative MRI between chimpanzees and humans. PLoS ONE 12, e0179624 (2017).
Semple, B. D., Blomgren, K., Gimlin, K., Ferriero, D. M. & Noble-Haeusslein, L. J. Brain development in rodents and humans: identifying benchmarks of maturation and vulnerability to injury across species. Prog. Neurobiol. 106, 1–16 (2013).
Bourgeois, J. & Rakic, P. Changes of synaptic density in the primary visual cortex of the macaque monkey from fetal to adult stage. J. Neurosci. 13, 2801–2820 (1993).
De Felipe, J., Marco, P., Fairén, A. & Jones, E. G. Inhibitory synaptogenesis in mouse somatosensory cortex. Cereb. Cortex 7, 619–634 (1997).
Schörnig, M. et al. Comparison of induced neurons reveals slower structural and functional maturation in humans than in apes. eLife 10, e59323 (2021).
Lindhout, F. W. et al. Quantitative mapping of transcriptome and proteome dynamics during polarization of human iPSC-derived neurons. eLife 9, e58124 (2020). This study reports the identification of an intermediate axon developmental stage in human neurons.
Dotti, C., Sullivan, C. & Banker, G. The establishment of polarity by hippocampal neurons in culture. J. Neurosci. 8, 1454–1468 (1988).
Lindhout, F. W. et al. Centrosome‐mediated microtubule remodeling during axon formation in human iPSC‐derived neurons. EMBO J. 40, e106798 (2021). This study shows that centrosomes modulate axon development in human neurons, with a human-specific localization of axon initial segment protein TRIM46 to centrosomes prior to axon formation.
Bianchi, S. et al. Synaptogenesis and development of pyramidal neuron dendritic morphology in the chimpanzee neocortex resembles humans. Proc. Natl Acad. Sci. USA 110, 10395–10401 (2013).
Rayon, T. et al. Species-specific pace of development is associated with differences in protein stability. Science 369, eaba7667 (2020).
Lázaro, J. et al. A stem cell zoo uncovers intracellular scaling of developmental tempo across mammals. Cell Stem Cell 30, 938–949.e7 (2023).
Matsuda, M. et al. Species-specific segmentation clock periods are due to differential biochemical reaction speeds. Science 369, 1450–1455 (2020).
Iwata, R. et al. Mitochondria metabolism sets the species-specific tempo of neuronal development. Science 379, eabn4705 (2023). In vitro experiments show that mitochondrial metabolism regulates species-specific neurodevelopmental tempo.
Ciceri, G. et al. An epigenetic barrier sets the timing of human neuronal maturation. Nature 626 626, 881–890 (2024). This study presents dentification of a set of chromatin modifiers orchestrating neuronal maturation tempo.
Wang, J., Weatheritt, R. & Voineagu, I. Alu-minating the mechanisms underlying primate cortex evolution. Biol. Psychiatry 92, 760–771 (2022).
Nesta, A. V., Tafur, D. & Beck, C. R. Hotspots of human mutation. Trends Genet. 37, 717–729 (2021).
Suntsova, M. V. & Buzdin, A. A. Differences between human and chimpanzee genomes and their implications in gene expression, protein functions and biochemical properties of the two species. BMC Genomics 21, 535 (2020).
Saitou, M. & Gokcumen, O. An evolutionary perspective on the impact of genomic copy number variation on human health. J. Mol. Evol. 88, 104–119 (2020).
Dennis, M. Y. & Eichler, E. E. Human adaptation and evolution by segmental duplication. Curr. Opin. Genet. Dev. 41, 44–52 (2016).
Waterson, R. H., Lander, E. S. & Wilson, R. K. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87 (2005).
Nurk, S. et al. The complete sequence of a human genome. Science 376, 44–53 (2022).
An, N. A. et al. De novo genes with an lncRNA origin encode unique human brain developmental functionality. Nat. Ecol. Evol. 7, 264–278 (2023).
Vakirlis, N., Vance, Z., Duggan, K. M. & McLysaght, A. De novo birth of functional microproteins in the human lineage. Cell Rep. 41, 111808 (2022).
Levchenko, A., Kanapin, A., Samsonova, A. & Gainetdinov, R. R. Human accelerated regions and other human-specific sequence variations in the context of evolution and their relevance for brain development. Genome Biol. Evol. 10, 166–188 (2017).
Franchini, L. F. & Pollard, K. S. Genomic approaches to studying human-specific developmental traits. Development 142, 3100–3112 (2015).
Pinson, A. et al. Human TKTL1 implies greater neurogenesis in frontal neocortex of modern humans than Neanderthals. Science 377, eabl6422 (2022).
Suzuki, I. K. et al. Human-specific NOTCH2NL genes expand cortical neurogenesis through Delta/Notch regulation. Cell 173, 1370–1384.e16 (2018).
Fiddes, I. T. et al. Human-specific NOTCH2NL genes affect Notch signaling and cortical neurogenesis. Cell 173, 1356–1369.e22 (2018).
Schmidt, E. R. E. et al. A human-specific modifier of cortical connectivity and circuit function. Nature 599, 640–644 (2021).
Libé-Philippot, B. et al. Human synaptic neoteny requires species-specific balancing of SRGAP2–SYNGAP1 cross-inhibition. Preprint at bioRxiv 10.1101/2023.03.01.530630 (2023).
Lucas, B. & Hardin, J. Mind the (sr)GAP—roles of Slit–Robo GAPs in neurons, brains and beyond. J. Cell Sci. 130, 3965–3974 (2017).
Charrier, C. et al. Inhibition of SRGAP2 function by its human-specific paralogs induces neoteny during spine maturation. Cell 149, 923–935 (2012). This study demonstrates that the human-specific gene SRGAP2C inhibits SRGCAP2A activity, delaying spine maturation.
Dennis, M. Y. et al. Evolution of human-specific neural SRGAP2 genes by incomplete segmental duplication. Cell 149, 912–922 (2012).
Heide, M. et al. Human-specific ARHGAP11B increases size and folding of primate neocortex in the fetal marmoset. Science 369, 546–550 (2020).
Florio, M. et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science 347, 1465–1470 (2015).
Franchini, L. F. & Pollard, K. S. Human evolution: the non-coding revolution. BMC Biol. 15, 89 (2017).
Romero, I. G., Ruvinsky, I. & Gilad, Y. Comparative studies of gene expression and the evolution of gene regulation. Nat. Rev. Genet. 13, 505–516 (2012).
Sholtis, S. J. & Noonan, J. P. Gene regulation and the origins of human biological uniqueness. Trends Genet. 26, 110–118 (2010).
Noonan, J. P. Regulatory DNAs and the evolution of human development. Curr. Opin. Genet. Dev. 19, 557–564 (2009).
Shibata, M. et al. Regulation of prefrontal patterning and connectivity by retinoic acid. Nature 598, 483–488 (2021).
Shibata, M. et al. Hominini-specific regulation of CBLN2 increases prefrontal spinogenesis. Nature 598, 489–494 (2021). Refs. 176 and 177 report molecular mechanisms of increased connectivity in the prefrontal cortex in humans.
Jeong, H. et al. Evolution of DNA methylation in the human brain. Nat. Commun. 12, 2021 (2021).
Kozlenkov, A. et al. Evolution of regulatory signatures in primate cortical neurons at cell-type resolution. Proc. Natl Acad. Sci. USA 117, 28422–28432 (2020).
Xu, C. et al. Human-specific features of spatial gene expression and regulation in eight brain regions. Genome Res. 28, 1097–1110 (2018).
Babbitt, C. C., Haygood, R., Nielsen, W. J. & Wray, G. A. Gene expression and adaptive noncoding changes during human evolution. BMC Genomics 18, 435 (2017).
Vermunt, M. W. et al. Epigenomic annotation of gene regulatory alterations during evolution of the primate brain. Nat. Neurosci. 19, 494–503 (2016).
Reilly, S. K. et al. Evolutionary changes in promoter and enhancer activity during human corticogenesis. Science 347, 1155–1159 (2015).
Shibata, Y. et al. Extensive evolutionary changes in regulatory element activity during human origins are associated with altered gene expression and positive selection. PLoS Genet. 8, e1002789 (2012).
Zeng, J. et al. Divergent whole-genome methylation maps of human and chimpanzee brains reveal epigenetic basis of human regulatory evolution. Am. J. Hum. Genet. 91, 455–465 (2012).
Oldham, M. C., Horvath, S. & Geschwind, D. H. Conservation and evolution of gene coexpression networks in human and chimpanzee brains. Proc. Natl Acad. Sci. USA 103, 17973–17978 (2006).
Prescott, S. L. et al. Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 163, 68–83 (2015).
Mangan, R. J. et al. Adaptive sequence divergence forged new neurodevelopmental enhancers in humans. Cell 185, 4587–4603.e23 (2022).
Hubisz, M. J. & Pollard, K. S. Exploring the genesis and functions of Human Accelerated Regions sheds light on their role in human evolution. Curr. Opin. Genet. Dev. 29, 15–21 (2014).
Pollard, K. S. et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 443, 167–172 (2006). This study reports on the identification of HARs, the fastest evolving regulatory elements in the genome.
Whalen, S. & Pollard, K. S. Enhancer function and evolutionary roles of human accelerated regions. Annu. Rev. Genet. 56, 423–439 (2022).
Whalen, S. et al. Machine learning dissection of human accelerated regions in primate neurodevelopment. Neuron 111, 857–873.e8 (2023).
Pizzollo, J., Zintel, T. M. & Babbitt, C. C. Differentially active and conserved neural enhancers define two forms of adaptive noncoding evolution in humans. Genome Biol. Evol. 14, evac108 (2022).
Uebbing, S. et al. Massively parallel discovery of human-specific substitutions that alter enhancer activity. Proc. Natl Acad. Sci. USA 118, e2007049118 (2021).
Girskis, K. M. et al. Rewiring of human neurodevelopmental gene regulatory programs by human accelerated regions. Neuron 109, 3239–3251.e7 (2021).
Weiss, C. V. et al. The cis-regulatory effects of modern human-specific variants. eLife 10, e63713 (2021).
Reilly, S. K. & Noonan, J. P. Evolution of gene regulation in humans. Annu. Rev. Genomics Hum. Genet. 17, 45–67 (2014).
Boyd, J. L. et al. Human–chimpanzee differences in a FZD8 enhancer alter cell-cycle dynamics in the developing neocortex. Curr. Biol. 25, 772–779 (2015).
McLean, C. Y. et al. Human-specific loss of regulatory DNA and the evolution of human-specific traits. Nature 471, 216–219 (2011).
Kostka, D., Hahn, M. W. & Pollard, K. S. Noncoding sequences near duplicated genes evolve rapidly. Genome Biol. Evol. 2, 518–533 (2010).
Johansson, P. A. et al. A cis-acting structural variation at the ZNF558 locus controls a gene regulatory network in human brain development. Cell Stem Cell 29, 52–69.e8 (2022).
Keough, K. C. et al. Three-dimensional genome rewiring in loci with human accelerated regions. Science 380, eabm1696 (2023).
Berto, S. & Nowick, K. Species-specific changes in a primate transcription factor network provide insights into the molecular evolution of the primate prefrontal cortex. Genome Biol. Evol. 10, evy149 (2018).
Florio, M. et al. Evolution and cell-type specificity of human-specific genes preferentially expressed in progenitors of fetal neocortex. eLife 7, e32332 (2018).
Aughey, G. N., Forsberg, E., Grimes, K., Zhang, S. & Southall, T. D. NuRD‐independent Mi‐2 activity represses ectopic gene expression during neuronal maturation. EMBO Rep. 24, e55362 (2023).
Salamon, I. & Rasin, M.-R. Evolution of the neocortex through RNA-binding proteins and post-transcriptional regulation. Front. Neurosci. 15, 803107 (2022).
Duffy, E. E. et al. Developmental dynamics of RNA translation in the human brain. Nat. Neurosci. 25, 1353–1365 (2022).
Li, M. & Larsen, P. A. Primate-specific retrotransposons and the evolution of circadian networks in the human brain. Neurosci. Biobehav. Rev. 131, 988–1004 (2021).
Ibarra, I. L. et al. Comparative chromatin accessibility upon BDNF stimulation delineates neuronal regulatory elements. Mol. Syst. Biol. 18, e10473 (2022).
Beagan, J. A. et al. Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression. Nat. Neurosci. 23, 707–717 (2020).
Tyssowski, K. M. et al. Different neuronal activity patterns induce different gene expression programs. Neuron 98, 530–546.e11 (2018).
Su, Y. et al. Neuronal activity modifies the chromatin accessibility landscape in the adult brain. Nat. Neurosci. 20, 476–483 (2017).
Kim, T.-K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Hergenreder, E. et al. Combined small-molecule treatment accelerates maturation of human pluripotent stem cell-derived neurons. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-02031-z (2024).
Kebschull, J. M. et al. Cerebellar nuclei evolved by repeatedly duplicating a conserved cell-type set. Science 370, eabd5059 (2020).
Homman‐Ludiye, J. & Bourne, J. A. The medial pulvinar: function, origin and association with neurodevelopmental disorders. J. Anat. 235, 507–520 (2019).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Saunders, A. et al. Ascertaining cells’ synaptic connections and RNA expression simultaneously with barcoded rabies virus libraries. Nat. Commun. 13, 6993 (2022).
Yuan, L., Chen, X., Zhan, H., Gilbert, H. L. & Zador, A. M. Massive multiplexing of spatially resolved single neuron projections with axonal BARseq. Preprint at Biorxiv https://doi.org/10.1101/2023.02.18.528865 (2023).
Benavides-Piccione, R. et al. Differential structure of hippocampal CA1 pyramidal neurons in the human and mouse. Cereb. Cortex 30, 730–752 (2020).
Atamian, A. et al. Human cerebellar organoids with functional Purkinje cells. Cell Stem Cell 31, 39–51.e6 (2024).
Chen, Y. et al. Generation of advanced cerebellar organoids for neurogenesis and neuronal network development. Hum. Mol. Genet. 32, 2832–2841 (2023).
Pellegrini, L. et al. Human CNS barrier-forming organoids with cerebrospinal fluid production. Science 369, eaaz5626 (2020).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Rao, A., Barkley, D., França, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Longo, S. K., Guo, M. G., Ji, A. L. & Khavari, P. A. Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nat. Rev. Genet. 22, 627–644 (2021).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Zhu, K. et al. Multi-omic profiling of the developing human cerebral cortex at the single-cell level. Sci. Adv. 9, eadg3754 (2023).
Domcke, S. et al. A human cell atlas of fetal chromatin accessibility. Science 370, eaba7612 (2020).
Pesce, L. et al. 3D molecular phenotyping of cleared human brain tissues with light-sheet fluorescence microscopy. Commun. Biol. 5, 447 (2022).
Rodriguez-Gatica, J. E. et al. Imaging three-dimensional brain organoid architecture from meso- to nanoscale across development. Development 149, dev200439 (2022).
Michalska, J. M. et al. Uncovering brain tissue architecture across scales with super-resolution light microscopy. Preprint at bioRxiv https://doi.org/10.1101/2022.08.17.504272 (2022).
Sarkar, D. et al. Revealing nanostructures in brain tissue via protein decrowding by iterative expansion microscopy. Nat. Biomed. Eng. 6, 1057–1073 (2022).
Rakotoson, I. et al. Fast 3-D imaging of brain organoids with a new single-objective planar-illumination two-photon microscope. Front. Neuroanat. 13, 77 (2019).
Krafft, P. R. et al. Etiology of stroke and choice of models. Int. J. Stroke 7, 398–406 (2012).
Benavides-Piccione, R., Ballesteros-Yáñez, I., DeFelipe, J. & Yuste, R. Cortical area and species differences in dendritic spine morphology. J. Neurocytol. 31, 337–346 (2002).
Mikula, S., Trotts, I., Stone, J. M. & Jones, E. G. Internet-enabled high-resolution brain mapping and virtual microscopy. NeuroImage 35, 9–15 (2007).
Barrett, R. D. H. & Hoekstra, H. E. Molecular spandrels: tests of adaptation at the genetic level. Nat. Rev. Genet. 12, 767–780 (2011).
Acknowledgements
The authors thank their colleagues for inspiring scientific discussions. This work was supported by the Netherlands Organisation for Scientific Research (NWO-Rubicon, 019.211EN.032ß) and an European Molecular Biology Organisation non-stipendiary postdoctoral fellowship (EMBO, ALTF 845–2021) to F.W.L.; an Klingenstein-Simons fellowship, a Simons Foundation BTI award, and an NIH BRAIN Initiative grant (UM1MH130981) to F.M.K.; the Gladstone Institutes, Chan Zuckerberg Biohub San Fransisco and the NIMH (U01-MH116438) to K.S.P.; and the Medical Research Council (MC_UP_1201/9) and the European Research Council (ERC-STG, 757710) to M.A.L.
Author information
Authors and Affiliations
Contributions
All authors contributed to writing and reviewing the manuscript. Specifically, F.W.L. and M.A.L. planned and outlined the manuscript, F.W.L. drafted the manuscript and figures, F.M.K. contributed to sections on cellular diversity, K.S.P. contributed to sections on evolutionary genetics and mechanisms, M.A.L. contributed to sections on human-specific brain evolution and nomenclature, and all authors contributed to editing and finalizing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Javier DeFelipe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1–3 and references.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lindhout, F.W., Krienen, F.M., Pollard, K.S. et al. A molecular and cellular perspective on human brain evolution and tempo. Nature 630, 596–608 (2024). https://doi.org/10.1038/s41586-024-07521-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07521-x
- Springer Nature Limited