Abstract
The fungus Candida albicans frequently colonizes the human gastrointestinal tract, from which it can disseminate to cause systemic disease. This polymorphic species can transition between growing as single-celled yeast and as multicellular hyphae to adapt to its environment. The current dogma of C. albicans commensalism is that the yeast form is optimal for gut colonization, whereas hyphal cells are detrimental to colonization but critical for virulence1,2,3. Here, we reveal that this paradigm does not apply to multi-kingdom communities in which a complex interplay between fungal morphology and bacteria dictates C. albicans fitness. Thus, whereas yeast-locked cells outcompete wild-type cells when gut bacteria are absent or depleted by antibiotics, hyphae-competent wild-type cells outcompete yeast-locked cells in hosts with replete bacterial populations. This increased fitness of wild-type cells involves the production of hyphal-specific factors including the toxin candidalysin4,5, which promotes the establishment of colonization. At later time points, adaptive immunity is engaged, and intestinal immunoglobulin A preferentially selects against hyphal cells1,6. Hyphal morphotypes are thus under both positive and negative selective pressures in the gut. Our study further shows that candidalysin has a direct inhibitory effect on bacterial species, including limiting their metabolic output. We therefore propose that C. albicans has evolved hyphal-specific factors, including candidalysin, to better compete with bacterial species in the intestinal niche.
Similar content being viewed by others
Main
Fungi are life-long components of the human microbiota and are acquired soon after birth7. Despite being minor constituents of the intestinal microbiota, commensal fungi have key roles in stimulating both local and systemic immune responses. These eukaryotic species can serve protective or pathologic functions that include roles in inflammatory bowel disease and allergic reactions5,8,9,10,11,12,13,14,15. Candida species can also escape the commensal niche and cause life-threatening systemic infections, particularly in the immunocompromised host16,17,18.
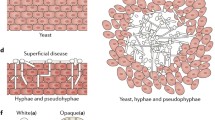
One of the most clinically relevant fungal species is C. albicans, which can grow as budding yeast or as filamentous hyphae or pseudohyphae19,20. This species is a common component of the human gut microbiota and yet the fungal factors promoting commensalism are poorly defined, particularly under homeostatic conditions in which fungi are vastly outnumbered by bacteria. Previous studies have established that the yeast-to-hyphal transition is critical for systemic virulence20,21,22,23, and yet yeast-locked forms are optimal for commensalism in germ-free (GF) or antibiotic-treated mice1,2,3,24,25. Indeed, serial passaging of C. albicans in the gut of antibiotic-treated mice repeatedly gave rise to yeast-locked forms, establishing that loss of the hyphal state is advantageous in this environment2.
These observations have led to the paradigm that yeast cells are optimal for gut colonization whereas the hyphal form is detrimental to colonization but critical for disease. However, if filamentation is detrimental to commensal fitness, this raises the question of why this programme has been retained. Indeed, the vast majority of clinical isolates are capable of undergoing the yeast-to-hyphal transition, whereas a minority are naturally yeast-locked25. These observations led us to speculate whether hyphal formation is maintained by certain selective pressures during C. albicans colonization of the mammalian host.
Here, we examined C. albicans gut colonization in diverse mouse models, both in the presence and absence of bacterial competition. We demonstrate that intestinal colonization involves a complex interplay between the morphological state of the fungus and the bacterial microbiome. Thus, yeast-locked cells demonstrate increased fitness when intestinal bacteria are absent or depleted, whereas hyphae formation is critical for gut colonization when commensal bacteria are prevalent. We show that filamentation is advantageous for colonization owing to the production of hyphal-specific factors including the toxin candidalysin—the first true virulence factor identified in a human fungal pathogen4,26,27. Hyphal growth therefore enables C. albicans to propagate in bacterially-colonized hosts, establishing that fungal virulence factors act as ‘commensalism factors’ during inter-kingdom competition in the intestinal niche.
Colonization by yeast-locked C. albicans cells
We performed a series of competition assays to compare the fitness of wild-type (WT) C. albicans cells with those lacking the Efg1 transcription factor, which are yeast-locked under most conditions22,28,29,30. Previous studies have often utilized antibiotics to promote stable gut colonization with C. albicans1,2,3,24,25,31. Here, colonization was compared in hosts fed a standard diet (SD) (FormuLab 5001) versus those fed a low-fibre, high-sucrose purified diet (PD) (AIN-93G), both with and without antibiotics. PD-fed hosts were previously shown to support long-term C. albicans colonization even without antibiotic dysbiosis32.
We first compared the relative colonization fitness of the standard C. albicans WT strain SC5314 with efg1Δ/Δ cells constructed in this background. Consistent with previous studies1,2,3,24,25, efg1Δ/Δ cells outcompeted WT SC5314 cells in mice treated with antibiotics (penicillin/streptomycin), reaching more than 90% of the fecal population by two days after infection, independent of whether mice were fed the SD or the PD (Fig. 1a–c). efg1Δ/Δ cells also dominated in gastrointestinal organs taken from these hosts (Extended Data Fig. 1a,b). In contrast to colonization of antibiotic-treated hosts, however, efg1Δ/Δ cells exhibited a marked fitness defect relative to WT SC5314 cells in both SD-fed and PD-fed mice in the absence of antibiotics (Fig. 1d,e). For PD-fed hosts, WT SC5314 cells made up 96% of the population recovered from fecal pellets by day 3 and 100% of the population by day 7 (Fig. 1e) and dominated in gastrointestinal organs recovered at the experimental endpoint (Extended Data Fig. 1c). WT cells also dominated in SD-fed hosts, showing significantly higher fitness than efg1Δ/Δ cells (Fig. 1d). C. albicans WT SC5314 cells show limited colonization of SD-fed hosts without antibiotics; we therefore also compared WT versus efg1Δ/Δ fitness using C. albicans isolates 529L and CHN1, which stably colonize SD-fed mice even without antibiotics33. WT versions of both 529L and CHN1 outcompeted efg1Δ/Δ derivatives in antibiotic-naive SD-fed mice and colonization levels were stably maintained, in contrast to the outcome observed with SC5314 cells (Extended Data Fig. 1d–f).
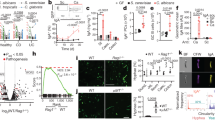
a, Schematic of C. albicans colonization models. BALB/c mice were gavaged with a 1:1 mixture of WT and efg1Δ/Δ cells (or EFG1/efg1Δ and efg1Δ/Δ cells) and fed a SD or a PD. Drinking water was supplemented with penicillin/streptomycin (P/S) as indicated. C. albicans colonies were examined in fecal pellets at the indicated time points. b–e, WT (CAY2698) versus efg1Δ/Δ (CAY11750) competition in the SD + P/S (b), PD + P/S (c) or SD (d) models; EFG1/efg1Δ (CAY7064) and efg1Δ/Δ (CAY7769) were competed in the PD model (e). f, PD-fed mice were colonized with C. albicans and the number of colony forming units (CFU) per gram of fecal pellets was determined. n = 4. LOD, limit of detection. g, Analysis of cell morphology of WT SC5314, efg1Δ/Δ and nrg1Δ/Δ cells. C. albicans cells were stained with anti-Candida antibody (green) and epithelial cells were stained with DAPI (blue) in the SD + P/S model. Scale bar, 100 μm. h, Quantitative analysis of yeast and filamentous morphotypes from colonic sections (C57BL/6 mice) using the SD + P/S model. For WT (CAY2698), n = 10; efg1Δ/Δ (CAY10965), n = 5; nrg1Δ/Δ (CAY1398), n = 5. i, Analysis of cell morphology of WT SC5314, efg1Δ/Δ and nrg1Δ/Δ cells cultured at 37 °C in caecal contents from PD-fed BALB/c mice. Scale bar, 20 μm. j, Quantitative analysis of yeast and filamentous morphotypes in caecal contents in i. n = 3. GI, gastrointestinal. Data are mean ± s.e.m.; each data point represents an individual mouse. In j, each data point represents an individual experiment. b–e, Paired t-test (two-tailed). f,h,j, Mann–Whitney (two-tailed) test.
The colonization properties of SC5314 WT and efg1Δ/Δ strains were further compared by individually inoculating these strains into mice alongside an SC5314 strain lacking NRG1 that is constitutively filamentous34,35,36. Both efg1Δ/Δ and nrg1Δ/Δ cells were defective for colonization of PD-fed mice (Fig. 1f), in line with a study showing that hyphal-locked cells are also defective in colonizing antibiotic-treated mice37. WT SC5314 cells adopted both yeast and hyphal morphologies in antibiotic-treated hosts, whereas more than 98% of efg1Δ/Δ cells were yeast cells and more than 90% of nrg1Δ/Δ cells were filamentous cells (Fig. 1g,h). Fungal morphotypes were also evaluated in caecal contents from antibiotic-naive, PD-fed mice, which again showed that WT SC5314 cells adopted both yeast and hyphal forms, whereas efg1Δ/Δ and nrg1Δ/Δ cells were essentially yeast-locked and hyphal-locked, respectively (Fig. 1i,j).
We also evaluated the fitness of yeast-locked flo8Δ/Δ cells, as FLO8 is a second transcription factor that is important for hyphal development2 (Extended Data Fig. 2a,b). flo8Δ/Δ SC5314 cells outcompeted their WT counterparts in GF and antibiotic-treated hosts (Extended Data Fig. 2c–e) but, similar to efg1Δ/Δ cells, showed decreased relative fitness in antibiotic-naive PD- and SD- fed mice (Extended Data Fig. 2f,g). Deletion of FLO8 in strains 529L and CHN1 also resulted in reduced fitness relative to their WT controls in antibiotic-naive hosts (Extended Data Fig. 2h,i).
Together, these results reveal that yeast-locked cells exhibit a colonization fitness advantage over filamentation-competent WT cells in antibiotic-treated mice that are depleted for bacteria. By contrast, the relative fitness of these strains is reversed in antibiotic-naive mice, in which WT cells are fitter than yeast-locked (and hyphal-locked) mutants. Both fungal morphotypes (or transitions between morphotypes) are therefore required for gastrointestinal colonization of antibiotic-naive hosts. Moreover, these results were independent of the mouse diet, fungal strain or mouse background, establishing the generality of these findings.
Diet and antibiotics alter the bacterial microbiome
Given that a change in diet or antibiotics can alter fungal gut commensalism, we examined the effects of these factors on the bacterial microbiome, both with and without C. albicans colonization. In the absence of antibiotics, SD-fed mice displayed decreasing levels of C. albicans in the gut and most lost fungal colonization by day 21, whereas PD-fed mice and those on antibiotics exhibited stable C. albicans colonization throughout the experiment (Extended Data Fig. 3a,b). SD-fed and PD-fed mice showed average bacterial loads of 1010 to 1011 CFU g−1 in fecal pellets that decreased 103- to 105-fold upon antibiotic supplementation (Extended Data Fig. 3c,d). Under these conditions, antibiotic-naive mice therefore harbour an ‘intact’ bacterial microbiota with more than 109 bacterial CFU g−1. Notably, C. albicans colonization did not significantly affect total gut bacterial loads in any of the groups tested (Extended Data Fig. 3c,d).
To assess intestinal bacterial communities, we performed 16S rRNA sequencing. A shift in diet had the largest effect on 16S composition (Extended Data Fig. 3e), with a significantly lower Shannon Diversity in PD-fed mice than in SD-fed mice at days 7 and 10 (Extended Data Fig. 3f). PD-fed mice also harboured higher Deferribacterota (day 1, 3, 14, 21 and SI), Proteobacteria (day 10) and Verrucomicrobia (SI), but lower Bacteroidota (day 0 and 3) than SD-fed mice (Extended Data Fig. 3g). SD-fed mice were initially associated with more Firmicutes, including Bacilli and Clostridia, as well as increased Mycoplasmota (Anaeroplasma), whereas PD-fed mice were associated with more Alphaproteobacteria (Rhodospirales) (Extended Data Fig. 4a–d). In addition to depleting intestinal bacterial levels, antibiotic treatment shifted the composition of the bacterial microbiome, with a larger effect in PD-fed mice than in SD-fed mice (Extended Data Fig. 3h,i). The most significant effects of antibiotics included an increase in Proteobacteria and Actinobacteria (multiple time points with both diets), as well as a decrease in Bacteroidota at early time points (Extended Data Fig. 3h,i).
C. albicans colonization did not affect overall bacterial levels or alter Shannon diversity, but still had a significant effect on microbiome composition (Extended Data Figs. 3c,d,j–m and 5a–c). C. albicans decreased Actinobacteria in PD-fed mice (day 7 and 21) and in SD-fed mice given antibiotics (day 3 and 21) (Extended Data Fig. 3k,m). Eggerthellaceae was the major family represented in the Actinobacteria phylum and the relative abundance of this phylum decreased in several groups that harboured C. albicans at day 21 (Extended Data Fig. 3n). C. albicans also impacted the Verrucomicrobia phylum; it decreased this phylum in SD-fed mice (day 1, 14 and 21) and PD-fed mice with antibiotics (day 10) yet increased this phylum in PD-fed mice without antibiotics (day 7) (Extended Data Fig. 3j–l). At the genus level, C. albicans colonization in PD-fed mice led to decreased abundance of Enterorhabdus and Gordonibacter from the Actinobacteria phylum (Extended Data Fig. 4e–h).
Together, these data demonstrate that antibiotic treatment or changing of the diet (from SD to PD) caused substantial changes to gut bacterial populations, including a higher abundance of aerobic Proteobacteria, a marker of microbiome dysbiosis38. Disruption of the bacterial microbiome was therefore associated with increased fungal colonization. By contrast, introduction of C. albicans was associated with limited changes to bacterial populations including the restriction and expansion of certain specific phyla.
Bacteria govern the fitness of C. albicans cells
To further address how changes in the bacterial microbiome affect fungal fitness, we performed competition experiments in GF mice and in mice that harboured specific bacterial communities. Consistent with previous studies24, GF mice were stably colonized by WT SC5314 cells. Moreover, yeast-locked efg1Δ/Δ cells showed equal or higher fitness than WT cells in these hosts; efg1Δ/Δ cells outcompeted WT cells in GF C57BL/6 and BALB/c mice (Fig. 2a,b and Extended Data Fig. 6a,b), whereas these cell types showed equivalent fitness in GF NMRI mice (Extended Data Fig. 6c).
a, Schematic of gut colonization assays in gnotobiotic hosts that are GF or harbour different bacterial consortia. Mice were inoculated with a 1:1 mixture of WT and yeast-locked efg1Δ/Δ (CAY11750) cells (SC5314 background). b–d, Competition outcomes in GF C57BL/6 mice (b), NMRI mice harbouring the OMM12 consortium (c), C57BL/6 mice harbouring the OMM15 consortium (d) or the ASF consortium (e). f, Schematic of competition experiments using SD-fed mice supplemented with ampicillin (Amp). Mice were colonized with AmpR bacteria prior to gavage with C. albicans or were given C. albicans without bacteria. g, Total bacterial levels in fecal pellets were determined by quantitative PCR. No treatment, samples collected prior to ampicillin supplementation. Ca, C. albicans; Ec, E. coli; Ef, E. faecium; Kp, K. pneumoniae. h,i, Bacterial (h) and C. albicans (i) CFUs quantified in fecal pellets at 1, 3 and 7 days after infection. n = 8 in ampicillin-treated mice without bacteria, n = 6 in mice with E. coli, n = 3 in mice with K. pneumoniae, n = 4 in mice with E. faecium, and n = 6 in mice with no added bacteria treatment. j, WT versus efg1Δ/Δ competition in ampicillin-treated mice. n = 8. k–m, Competitions performed in ampicillin-treated mice pre-colonized with AmpR E. coli (k), K. pneumoniae (l) or E. faecium (m). n = 6 (k), n = 3 (l), n = 4 (m). Each data point represents an individual mouse. b–e,g–m, Data are mean ± s.e.m. b–e,j–m, Paired t-test (two-tailed). g–i, Mann–Whitney (two-tailed) test.
To evaluate the effect of different bacterial species on C. albicans colonization, WT and efg1Δ/Δ cells were inoculated into gnotobiotic mice harbouring the oligo-mouse-microbiota of 12 (OMM12) or 15 (OMM15) defined bacterial species, or the altered Schaedler flora (ASF) of 8 species39,40. These consortia, to varying degrees, can limit colonization by enteric pathogens such as Salmonella enterica serovar Typhimurium39. C. albicans competition assays revealed that in contrast to GF mice, WT cells rapidly outcompeted efg1Δ/Δ cells in the presence of each of the three bacterial consortia (Fig. 2c–e). For example, WT cells represented more than 99% of the colonizing population by day 7 in mice carrying either the OMM15 or ASF consortium (Fig. 2d,e). The presence of each of these defined bacterial populations therefore strongly selects for filamentation-competent WT cells over yeast-locked efg1Δ/Δ cells. Total Candida colonization levels were also two to three orders of magnitude lower in hosts harbouring OMM12 or OMM15 than in comparable GF hosts (Extended Data Fig. 6e,f).
Similar competitions were performed in ampicillin-treated SD-fed mice into which an ampicillin-resistant (AmpR) Gram-negative or Gram-positive strain had been introduced prior to C. albicans inoculation (Fig. 2f). Ampicillin treatment led to a decrease of three orders of magnitude in gut bacterial loads consistent with previous studies41, whereas AmpR Escherichia coli, Klebsiella pneumoniae or Enterococcus faecium colonized to high levels in ampicillin-treated mice (Fig. 2g,h). The presence of AmpR bacteria restricted, but did not prevent, fungal colonization; fungal CFUs were more than 107 cells per gram in fecal material from ampicillin-treated mice but decreased to 1.5–4.0 × 106 cells per gram when mice were pre-colonized with AmpR bacteria (at 7 days after infection; Fig. 2i). Yeast-locked efg1Δ/Δ cells outcompeted WT C. albicans cells in ampicillin-treated mice as expected (Fig. 2j), but inclusion of any of the three AmpR bacterial species resulted in WT cells rapidly outcompeting efg1Δ/Δ cells (population >90% WT by day 5; Fig. 2k–m). Indeed, introduction of E. coli cells into a GF host also strongly selected for WT cells over efg1Δ/Δ cells (Extended Data Fig. 6d), establishing that bacterial monocolonization is sufficient, at high CFU levels, to drive the selection for filamentation-competent WT cells over yeast-locked cells. By contrast, gavage of heat-killed E. coli cells did not select for WT cells over efg1Δ/Δ cells (Extended Data Fig. 6g) indicating that dead bacteria do not affect fungal fitness in the same way as live bacteria.
Adaptive immunity in C. albicans colonization
Recent studies have shown that C. albicans morphotypes are differentially targeted by adaptive immunity in the gut. Hyphal cells preferentially induce intestinal immunoglobulin A (IgA) which binds to prominent hyphal-specific factors including Als1, Als3 and Hwp11,6. Loss of IgA increases the proportion of hyphae present in the gut, indicating that the filamentous state is suppressed by antibody responses. This in turn can promote the long-term fitness of C. albicans populations in GF mice by favouring colonization by cells in the yeast form1,6,42.
Induction of IgA takes at least a week1, and therefore C. albicans-specific IgA did not contribute to the fitness advantages of WT over yeast-locked cells described here, as these arise early (within 1–3 days) following inoculation (Fig. 1e). To further address this point, the fitness of WT and efg1Δ/Δ cells was compared between WT and Rag1−/− mice, as the latter lack B cells and T cells. We observed that C. albicans WT cells rapidly outcompeted efg1Δ/Δ cells in both control and Rag1−/− SD-fed mice without antibiotics (Extended Data Fig. 6h,i), establishing that the increased fitness of WT over yeast-locked C. albicans cells is independent of IgA responses at early stages of colonization. Of note, overall fungal colonization levels were similar between control and Rag1−/− mice up to 7–10 days, yet by 14 days fungal loads were significantly higher in Rag1−/− mice (Extended Data Fig. 6j), consistent with IgA targeting WT C. albicans cells at the later time points1,6.
These results establish that hyphae-competent C. albicans cells exhibit an early fitness advantage over yeast-locked cells in mice with an intact (antibiotic-naive) bacterial microbiome and that this advantage is present in both control and Rag1−/− mice. At later time points, however, adaptive immunity can potentially restrict the fitness of WT C. albicans cells, in line with IgA targeting of hyphal-specific cell surface adhesins1,6.
Hyphal-specific effectors promote gut colonization
We tested whether the increased colonization fitness of C. albicans WT cells over yeast-locked forms was owing to the expression of certain hyphal-specific genes20 (HSGs). HSGs are highly expressed in the gut, where their expression is lost upon deletion of EFG13. Prominent HSGs include established virulence factor genes such as ECE1, which encodes the peptide toxin candidalysin4,26,27,43,44. This factor damages epithelial cells and has been linked to inflammatory bowel disease5 and to colonization fitness in antibiotic-treated mice45. Other prominent HSGs include ALS3, which encodes an adhesin46 and invasin47 that also mediates iron acquisition48, and SOD5, which encodes a superoxide dismutase that protects against oxidative stress49,50.
Each of these HSGs was deleted from SC5314 and the colonization fitness of each mutant was compared with WT controls. Loss of either ECE1 or ALS3, but not SOD5, resulted in a fitness defect in antibiotic-naive hosts (Fig. 3a,b and Extended Data Fig. 7a,b,d). For example, competition of WT and ece1Δ/Δ cells showed that the colonizing population consisted entirely of WT cells in fecal pellets as early as 7 days after infection in PD-fed hosts (Fig. 3b) as well as in gastrointestinal organs from these mice (Extended Data Fig. 8a,b). A similar colonization defect was observed with ece1Δ/Δ cells constructed in the 529L and CHN1 strain backgrounds in antibiotic-naive hosts (Extended Data Fig. 8c,d). By contrast, loss of ECE1 (as with loss of EFG1) did not substantially decrease fitness in antibiotic-treated or GF mice (Fig. 3c,d and Extended Data Fig. 8e,f), whereas loss of ALS3 caused fitness defects in both antibiotic-treated and antibiotic-naive hosts (Extended Data Fig. 7d,e). The fitness advantage of WT cells over ece1Δ/Δ cells was restored in GF mice by pre-colonization with E. coli (Fig. 3e and Extended Data Fig. 8g). These results indicate that bacterial colonization levels determine the relative fitness of WT populations versus ece1Δ/Δ populations in the gut, as they do for WT cells versus efg1Δ/Δ cells.
a, Experimental design of C. albicans WT versus ece1Δ/Δ or ece1Δ/Δ + ECE1ΔpIII in conventional or gnotobiotic hosts. b–e, Competition between WT and ece1Δ/Δ cells in PD-fed BALB/c mice (b), SD-fed BALB/c mice on penicillin/streptomycin antibiotics (c), GF C57BL/6 mice (d) and GF C57BL/6 mice pre-colonized with E. coli (e). WT (CAY12202) versus ece1Δ/Δ (CAY8578) in b,d,e. WT (CAY11533) versus ece1Δ/Δ (CAY11507) in c. n = 4 in b, n = 5 in c–e. f,g, Competitions between WT (CAY12202) and ece1Δ/Δ+ECE1ΔpIII (CAY8580) in GF C57BL/6 mice (f) and PD-fed BALB/c mice (g). n = 4. h, Colonization levels of WT (CAY2698) and ece1Δ/Δ (CAY8785) strains individually colonizing SD + P/S mice. n = 3. i, Colonization levels of WT (CAY2698), ece1Δ/Δ (CAY8785) and ece1Δ/Δ + ECE1ΔpIII (CAY8580) strains individually colonizing PD-fed mice. n = 4. j,k, Analysis of cells in the colon of BALB/c SD + P/S mice infected with WT (CAY2698) or ece1Δ/Δ (CAY8785) cells at 7 days post-infection. j, Percentage of filamentous cells. n = 3. k, Staining with anti-Candida antibody (green) and DAPI (blue). Hyphal cells indicated by arrows. Scale bars, 20 μm. l,m, Analysis of cell morphology of WT (CAY2698) and ece1Δ/Δ (CAY8785) cells cultured at 37 °C in caecal contents from PD-fed BALB/c mice, n = 4. l, Quantitative analysis of yeast and filamentous morphotypes. m, Staining with anti-Candida antibody (green) and DAPI (blue). Scale bar, 50 μm. Each data point represents an individual mouse. b–j,l, Data are mean ± s.e.m. l, Each data point represents an individual experiment. b–g, Paired t-test (two-tailed). h–j, Mann–Whitney (two-tailed) test. l, Unpaired t-test (two-tailed).
The ECE1 gene product is proteolytically processed into eight peptides, of which the third peptide (pIII) represents the candidalysin toxin. WT cells were competed against ece1Δ/Δ + ECE1ΔpIII cells, which lack only the toxin-encoding peptide and, similar to full ECE1 gene deletions, ECE1ΔpIII cells showed equivalent fitness levels to WT cells in GF mice (Fig. 3f and Extended Data Fig. 8h), but were rapidly outcompeted by WT cells in conventionally-housed antibiotic-naive mice (Fig. 3g and Extended Data Fig. 8i). Mice were also individually colonized with WT, ece1Δ/Δ or ece1Δ/Δ + ECE1ΔpIII strains, both with and without antibiotic supplementation. Each of these strains colonized to similar levels in mice treated with antibiotics (Fig. 3h), in contrast to a previous analysis of ece1Δ/Δ cells45. WT and ece1Δ/Δ cells also showed similar filamentation profiles in these hosts (Fig. 3j,k), consistent with in vitro studies4. However, ece1Δ/Δ and ece1Δ/Δ + ECE1ΔpIII cells both showed substantial colonization defects relative to the WT strain in antibiotic-naive mice (CFU levels 26- to 32-fold lower than WT; Fig. 3i). Filamentation of ece1Δ/Δ cells was also similar to WT cells when evaluated in caecal contents from these mice (Fig. 3l,m). These results establish that the hyphal programme promotes C. albicans commensalism in mice harbouring an intact bacterial microbiome and does so, at least in part, owing to the expression of HSGs such as ECE1.
Candidalysin exhibits anti-bacterial properties
To determine how ECE1 affects gut commensalism, we evaluated the global transcriptomes of WT and candidalysin-deficient ece1Δ/Δ and ece1Δ/Δ + ECE1ΔpIII strains. Loss of either the ECE1 gene or the pIII peptide had little effect on the transcriptome, even under filamentation-promoting conditions, including the expression of other HSGs (Extended Data Fig. 9a). We engineered WT and ece1Δ/Δ cells to express GFP or RFP to enable in vitro head-to-head competitions. Competitions were performed under a variety of biologically relevant conditions including growth in RPMI medium at 37 °C with CO2, acidic pH (pH 4.7), high salt (1 M NaCl), hypoxia (1% O2), oxidative stress (2 mM H2O2), or in the presence of short-chain fatty acids (2% acetate) (Extended Data Fig. 9b–g). We did not observe any differences in fitness between WT and ece1Δ/Δ cells in any of these conditions. The ability of ece1Δ/Δ and WT cells to grow on epithelial cells also revealed no difference between the two cell types (Extended Data Fig. 9h). WT and candidalysin-deficient cells therefore exhibit similar abilities to propagate under in vitro conditions including those associated with physiological stress.
Given that the contribution of Ece1 to commensal fitness depends on bacterial competition, we tested the direct effect of synthetic candidalysin peptide on the bacterial species used in our gastrointestinal colonization assays. Notably, candidalysin peptide significantly reduced the number of CFUs generated by E. coli, K. pneumoniae and E. faecium, resulting in a twofold to threefold decrease in CFU formation by each of these species (Fig. 4a,b). Candidalysin also decreased the metabolic activity of all three bacterial species in a dose-dependent manner, with 70 µM candidalysin peptide decreasing E. coli metabolism by around 80% (Fig. 4c). Candidalysin peptide decreased glucose consumption in two of the three species (E. coli and K. pneumoniae) by 1.5- to 2.5-fold (Fig. 4d). Together, these results reveal that candidalysin toxin can directly inhibit the growth and metabolism of commensal bacterial species that compete for resources with C. albicans in the gut, in addition to the established role of candidalysin in mediating host cell damage and inflammation.
a, C. albicans can grow as yeast or filamentous forms, including pseudohyphae or true hyphae. Hyphal cells express a set of HSGs that include ECE1, which encodes the candidalysin toxin. The Ece1 proprotein is proteolytically processed by Kex2 and Kex1 to produce mature candidalysin peptide (Cal). b, Quantification of E. coli, E. faecium and K. pneumoniae colonies following challenge with 70 μM candidalysin peptide in PBS at 37 °C for the indicated times. n = 3. c, Effect of candidalysin on bacterial metabolism as determined by XTT assays. Activity is shown as a percentage of the no-candidalysin control (dotted line). n = 4. d, Effect of candidalysin concentration on bacterial glucose uptake. n = 3. RLU, relative light units. Data are mean ± s.e.m.; each data point represents independent experiments. b–d, Unpaired t-test (two-tailed).
Discussion
C. albicans is a commensal of the human gastrointestinal tract, where it has key roles in local and systemic immune responses, in addition to its ability to escape this niche and cause systemic disease. Currently, the yeast form of the species is thought to be optimal for colonization, whereas the filamentous form is deemed detrimental to gut fitness but critical for pathogenesis1,2,3,24,25. From an evolutionary perspective, this raises the question of why filamentation has been retained by the species if it reduces fitness in the natural environmental niche. Similarly, although candidalysin toxin is an established virulence factor that is essential for epithelial cell damage4 and is associated with escape from macrophages43, it is unclear what intrinsic advantage expressing this toxin provides to commensal fungal cells.
Here we reveal that both yeast-locked and hyphal-locked C. albicans cells are defective in intestinal colonization when an intact (antibiotic-naive) bacterial microbiome is present. The ability of cells to exist in both morphotypes, or to transition between morphotypes, is therefore as critical to the commensal fitness of the species as it is for virulence20,21,22,23. We show that hyphal-specific factors promote initial colonization of the gut owing to the expression of HSGs including ECE1 (encoding candidalysin) and ALS3 (encoding a multifactorial adhesin and invasion), establishing that these factors provide an intrinsic benefit to the cells producing them.
Fitness differences between WT and yeast-locked cells arise within 1–3 days of inoculation, ruling out a role for adaptive immunity in driving this selection, and analysis of colonization in Rag1−/− mice further established that such fitness differences are independent of the involvement of adaptive immunity. Notably, WT C. albicans colonization levels at later time points (14 days) were higher in Rag1−/− mice than in control mice, indicating that adaptive responses restrict fungal colonization, and consistent with the observation that mucosal IgA preferentially targets hyphal C. albicans cells1,6. Hyphal cells are therefore the target of both positive and negative selective pressures in the gut environment.
Our work also establishes that the fitness of different C. albicans strains in the gut is determined by bacterial populations in this niche. Most studies of fungal commensalism have utilized antibiotic-treated hosts to promote colonization, yet this dysbiosis alters the relative fitness attributes of WT and mutant C. albicans strains. As shown in the current study, yeast-locked cells are hyperfit in mice given antibiotics, but exhibit a lower fitness than WT cells in antibiotic-naive mice, in which bacterial loads are orders of magnitude higher. Moreover, different bacterial populations can select for WT cells over yeast-locked cells, implicating bacterial colonization levels, rather than specific bacterial species, as responsible for driving this selection.
The effect of ECE1 on fitness was, similar to that of yeast-locked cells, most evident in antibiotic-naive mice harbouring high bacterial loads in the gut. Under these conditions, WT cells were fitter than candidalysin-deficient cells during early colonization, both when tested individually and in head-to-head competitions. The latter result establishes that toxin production provides an intrinsic benefit to toxin-producing cells rather than a change to the environment that favours all commensal C. albicans cells. Loss of ECE1 did not markedly alter the C. albicans transcriptome or fitness in vitro, including during infection of epithelial cells. Candidalysin had a direct inhibitory effect, however, on the growth and metabolism of bacterial species shown to compete with C. albicans in vivo. These results implicate candidalysin in promoting gut commensalism through the direct targeting of bacteria that are competing for resources with C. albicans. This toxin therefore supports the establishment of fungal colonization by mechanisms distinct from, or in addition to, its established roles in cell damage and mucosal inflammation4,5. Future studies will further address the specificity of candidalysin on different bacterial species and whether this toxin has additional functions that enable fungal gut colonization.
In summary, we reveal that virulence factors produced by C. albicans hyphal cells are also bona fide commensalism factors that support fungal propagation in the gut when bacterial competition is high. These factors provide intrinsic benefits to the fitness of fungal cells, consistent with commensal niches acting as ‘training grounds’ for the selection of traits that are critical to both commensalism and pathogenicity51.
Methods
Mice
Age- and sex-matched mice were used in this study. Mice were monitored under care of full-time staff, given access to food and water ad libitum and maintained under a 12-h light:dark cycle, with temperature maintained at 22–25 °C and a relative humidity of 30–70%. Unless stated otherwise, conventional WT 7- to 8-week-old, female BALB/c mice (strain code 028) and 7- to 8-week-old, female C57BL/6 J mice (stock no. 000664) were obtained from Charles River Laboratories and The Jackson Laboratory, respectively, to perform experiments at Brown University. Male and female GF BALB/c mice (4–5 weeks old) obtained from Taconic Biosciences (Extended Data Figs. 2b and 6b) and female GF WT C57BL/6 (9–10 weeks old) from the National Gnotobiotic Rodent Resource Center (NGRRC) (University of North Carolina, Chapel Hill, NC) were housed and experiments conducted in a flexible isolator bubble at Brown University (Fig. 3d,f and Extended Data Fig. 8d,e). NMRI mice were bred and raised under GF conditions in the Central Animal Facility of the Hannover Medical School (Hanover, Germany) (Extended Data Fig. 6c,d). Mice of both sexes were used at 7–10 weeks of age. Littermates of the same sex were randomly assigned to experimental groups. After inoculation with C. albicans, mice were maintained in ventilated cages. Female GF C57BL/6 mice (7–8 weeks old) at the New York University Grossman School of Medicine Gnotobiotics Animal Facility were used to conduct the experiments in Bioexclusion cages (Tecniplast) (Fig. 2b,c). Male ASF52 C57BL/6 mice (6 weeks old) from Taconic Biosciences were used to perform the mouse study in Taconic’s Isolator Breeding Solutions (IBS) facility housed in a gnotobiotic study isolator (Fig. 2e). Female GF C57BL/6 J mice (11–12 weeks old) from NGRRC were used for experiments performed at University of North Carolina (UNC) (Fig. 3e and Extended Data Fig. 8h). Mice were assigned randomly to experimental groups and colonization assays were performed unblinded to know the identity of each group.
Media and reagents
Yeast extract peptone dextrose (YPD) and synthetic complete dextrose (SCD) medium were made as previously described53,54. YPD containing 200 µg ml−1 nourseothricin (Jena Biosciences) was used to select nourseothricin-resistant (NATR) strains. SC + maltose plates were prepared as for SCD but with 2% maltose instead of 2% dextrose. E. coli and K. pneumoniae were grown and maintained in Luria broth (LB) and E. faecium was grown in Todd Hewitt broth.
Construction of plasmids and C. albicans strains
Oligonucleotides and C. albicans strains used in the present study are listed in Supplementary Tables 1 and 2, respectively. To create the efg1Δ/Δ strain, pRB72125 was linearized with ApaI and SacI and transformed into SC5314, 529L or CHN1 to generate efg1Δ/EFG1 derivatives. Integration was checked by PCR with oligonucleotides 2284/4438 and 2286/4439 for 5′ and 3′ junctions, respectively. The SAT1 marker was recycled after growing on SC + maltose medium and the second allele of EFG1 was deleted in SC5314 and CHN1 backgrounds by transformation with linearized pRB721 to generate efg1Δ/Δ strains. PCR with oligonucleotides 819/828, 2284/4438 and 2286/4439 was used to check the open reading frame (ORF), 5′ junction and 3′ junction, respectively. Mutant strains were grown on SC + maltose to recycle the SAT1 marker, resulting in nourseothricin-sensitive (NATs) efg1Δ/Δ strains (CAY10195 and CAY10965 in SC5314, and CAY11184 in CHN1). To delete the second EFG1 allele in 529L, PCR with oligonucleotides 5948/5949 was used to amplify a hygromycin B (HYG) resistance cassette from pRB19555 and transformed into efg1Δ/EFG1 to generate efg1Δ/Δ (CAY11482). PCR using oligonucleotides 819/828, 4476/1458 and 2910/1459 was used to check ORF, 5′ junction and 3′ junction, respectively.
To delete FLO8, a pSFS-FLO8 knockout plasmid (pRB989) was generated by PCR amplification of the FLO8 5′ and 3′ flanking regions using oligonucleotides 4988/4989 and 4990/4991, respectively, and inserting into ApaI/XhoI and SacI/SacII sites in pSFS2A56. pRB989 was linearized by ApaI and SacI and transformed into WT 529L and CHN1 cells to generate flo8Δ/FLO8 cells. PCR with oligonucleotides 4982/2274 and 5076/739 was used to check cassette integration. After recycling the SAT1 marker by growing on SC + maltose medium, linearized pRB989 was again used to delete the second FLO8 allele to generate flo8Δ/Δ. PCRs were performed to check the ORF, 5′ junction and 3′ junction using oligonucleotides 4986/5200, 4982/2274 and 5076/739, respectively. Cells were grown on SC + maltose medium to produce NATs flo8Δ/Δ mutants in 529L (CAY11180) and CHN1 (CAY11186).
To generate SC5314 ece1Δ/Δ (CAY8785), oligonucleotides 4248/4249 were used to PCR amplify the ARG4 and HIS1 cassettes from CAY8578 (ece1Δ/Δ4) and transformed into strain SN9557. Cells were grown on SC medium without arginine to select transformants. Junction PCR checks were performed using oligonucleotides 4252/4287 and 4286/4253. The transformation was repeated to delete the second ECE1 allele and transformants grown on SC medium lacking histidine and arginine. Transformants were PCR checked using oligonucleotides 4250/4251 for ORF, 4252/4289 for the 5′ junction and 4288/4253 for the 3′ junction.
To delete ECE1 in 529L and CHN1, a pSFS-ECE1 knockout plasmid (pRB1481) was generated by inserting the 5′ and 3′ flanking regions of ECE1, amplified by PCR using oligonucleotides 6207/6208 and 6205/6206, into SacI/SacII and KpnI/XhoI sites in pSFS2A56. pRB1481 was linearized by KpnI/SacI and transformed into 529L and CHN1. Cassette integration was examined by PCR using oligonucleotides 4438/4248 and 4439/6277. The SAT1 marker was recycled, and the transformation was repeated to delete the second ECE1 copy. The 5′ junction, 3′ junction and ORF were analysed by PCR using oligonucleotides 4438/4248, 4439/6277 and 201/2810, respectively. The resulting strain was grown on medium containing maltose to recycle the SAT1 marker resulting in a NATS ece1Δ/Δ mutant in 529L and CHN1 strain backgrounds (CAY12441 and CAY12446, respectively).
To create an isogenic and prototrophic control SC5314 strain, HIS1 and ARG4 genes were integrated into strain SN9557. PCR with oligonucleotides 4114/4115 was used to amplify HIS1 from SC5314 and the amplicon was transformed into SN95, with selection on SC medium without histidine. Cassette integration was checked by PCR using oligonucleotides 2623/4289 and 2624/4288. The ARG4 gene was integrated using the plasmid pEM00358 digested with MfeI/SacI, and transformants selected on SC medium lacking histidine and arginine. Integration was PCR checked using oligonucleotides 6044/6045 and 6046/6047.
To generate NATR strains, the pDis3 plasmid was used to integrate into the NEUT5L locus as described previously59. The plasmid was linearized by NgoMIV and transformed into SC5314 efg1Δ/Δ (CAY10195), flo8Δ/Δ (yLM79460) and ece1Δ/Δ (CAY8785) to create NATR versions CAY11750, CAY9796 and CAY11507, respectively. PCR checks were performed using oligonucleotides 3055/3056.
To delete SOD5, plasmids were constructed where the HIS1 and LEU2 markers were flanked by 500 bp of homology from SOD5. The sod5::HIS1 plasmid (pRB2151) was constructed via Golden Gate Assembly using BsmBI. A 500-bp fragment from upstream of SOD5 was amplified from SC5314 using oligonucleotides 8551/8552. HIS1 was amplified by PCR from pSN5257 in two parts to mutagenize an endogenous BsmBI site. The 5′ and 3′ regions of HIS1 were amplified by PCR with oligonucleotides 7006/7007 and 7008/7009, respectively. A 500-bp fragment downstream of the SOD5 ORF was amplified by PCR from gDNA with oligonucleotides 8553/8554. These four fragments were assembled into pGGASelect (NEB). The sod5::LEU2 plasmid (pRB2153) was constructed using the same upstream and downstream 500-bp fragments, combined with the LEU2 marker amplified from pSN4057. The knockout cassette from pRB2151 was released by digestion with PacI and transformed into SN8757 to create sod5::HIS1/SOD5 strains CAY14705/14706. Integration was PCR checked using oligonucleotides 7510/7834 and 8596/8558. The second SOD5 allele was disrupted with LEU2 using PacI-digested pRB2153 and junction checked by PCR using oligonucleotides 7510/5056.
Mouse gastrointestinal infection
Mouse gastrointestinal infections were performed by oral gavage as previously described25. C. albicans cells were cultured in YPD medium overnight at 30 °C. The overnight cultures were diluted 50-fold into fresh YPD medium and grown for 4 h at 30 °C. Cells were washed with sterile water three times. To perform C. albicans competitions, the inoculum was composed of one NATS and one NATR isolate at a ratio of 1:1. Mice were orally gavaged with a total of ~108 fungal CFU (counted by haemocytometer) in 0.5 ml of water using plastic feeding tubes (Instech Laboratories) for experiments done at Brown University, New York University and University of Wuerzburg. An inoculum of ~108 CFU in 0.2 ml of water was used to inoculate mice in Taconic Biosciences. For the study performed in UNC, an inoculum of ~5 × 107 CFU in 0.1 ml of water was used. Due to the difficulty of performing oral gavage with GF mice in gnotobiotic conditions, C. albicans was added to the drinking water at a final concentration of ~2 × 106 CFU ml−1. Inoculum water was replaced with sterile water after 24 h. Two mice were cohoused in each cage. Fecal pellets were collected at the indicated time points and gastrointestinal organs (stomach, small intestine, caecum and colon) were collected at the end of the experiment. Gastrointestinal organs were homogenized in PBS with antibiotics (500 µg ml−1 penicillin, 500 µg ml−1 ampicillin, 250 µg ml−1 streptomycin, 225 µg ml−1 kanamycin, 125 µg ml−1 chloramphenicol and 25 µg ml−1 doxycycline) and cultured on YPD and YPD + NAT media to determine CFU of NATR and NATS strains.
Conventional mouse models utilized SD (FormuLab 5001, PMI Nutrition International) or PD (AIN-93G; Dyets). When antibiotics were included, they were present in the drinking water (1,500 U ml−1 of penicillin, 2 mg ml−1 of streptomycin and 2.5% glucose for taste) for four days prior to C. albicans inoculation. Mice remained on the indicated diet with or without antibiotics throughout the experiment.
Colonization experiments with individual C. albicans strains were carried out for WT SC5314, efg1Δ/Δ and nrg1Δ/Δ strains using female BALB/c mice from Charles River Laboratory that were 7–8 weeks of age and that were allowed to acclimatize in the animal facility for 4 days. Mice were housed together initially and were fed the PD for one week. Colonization of mice with C. albicans strains was performed by gavaging 108 fungal CFU in water.
To perform C. albicans competitions with AmpR bacteria (K. pneumoniae B425, E. coli B427 or Enterococcus faecium B460), ampicillin was dosed into the drinking water at 0.5 g l−1 for four days prior to bacterial inoculation for conventional mice. Bacterial strains were cultured in LB medium at 37 °C overnight. The overnight cultures were washed twice with PBS and resuspended in PBS at half the original volume. To generate the heat-killed E. coli (B427), bacterial cells were heated at 70 °C for 30 min. Bacterial cell death was verified by plating the inoculum on LB + carbenicillin (100 µg ml−1). Heat-killed E. coli were inoculated every other day after the first inoculation throughout the experiment. Inoculation of C. albicans cells was performed three days following bacterial inoculation. Mice were housed individually after supplementing ampicillin into the drinking water. Mice were fed a SD and remained on ampicillin throughout the experiment. CFU of AmpR bacteria was determined by plating on LB + carbenicillin. Bacterial strains used in this study are listed in Supplementary Table 3.
GF NMRI mice were maintained carrying the OMM12 consortium39. OMM12 contains 12 bacterial strains including Acutalibacter muris KB18, Flavonifractor plautii YL31, Clostridium clostridioforme YL32, Blautia coccoides YL58, Clostridium innocuum I46, Lactobacillus reuteri I49, Enterococcus faecalis KB1, Bacteroides caecimuris I48, Muribaculum intestinale YL27, Bifidobacterium longum subsp. animalis YL2, Turicimonas muris YL45 and Akkermansia muciniphila YL44. The OMM15 consortium was previously described39,61 and contains OMM12 plus E. coli Mt1B1, Staphylococcus xylosus 33ERD13C, and Streptococcus danieliae ERD01G.
Male C57BL/6 mice carrying the ASF52 (consisting of Parabacteroides goldsteinii ASF519, Eubacterium plexicaudatum ASF492, Schaedlerella arabinosiphila ASF502, Pseudoflavonifractor Sp. ASF500, Clostridium sp. ASF356, Mucispirillum schaedleri ASF457, Ligilactobacillus murinus ASF 361 and Lactobacillus intestinalis ASF360) were inoculated with C. albicans cells immediately after transfer from the gnotobiotic breading isolator into a gnotobiotic isolator in the Taconic IBS facility. Fecal samples were collected at indicated time points and stored in 25% glycerol at −80 °C and shipped to Brown University on dry ice, where they were processed to determine fungal populations.
Female GF C57BL/6 mice were orally inoculated with E. coli promptly after transfer out of the gnotobiotic isolator. Mice were inoculated with C. albicans three days post E. coli inoculation. Fecal samples were collected at indicated time points and stored in PBS buffer with 25% glycerol at −80 °C and shipped to Brown University on dry ice where they were processed to determine the fungal populations.
Analysis of C. albicans morphology in vitro and in vivo
C. albicans morphology was analysed in vitro by overnight culture in YPD at 30 °C. From overnight cultures, 107 cells were subcultured into concentrated and filtered caecal contents (10 mg ml-1) collected from PD-fed BALB/c mice for 3 h. Cells were washed twice with PBS and stained with 100 μg ml−1 Calcofluor White for 15 min. Cells were further washed twice with PBS, resuspended in PBS and imaged using a Zeiss Axio Observer microscope.
C. albicans cells in the different gastrointestinal sections were imaged by immunofluorescence analysis. Pieces of 1–2 cm of gastrointestinal tract segments (ileum and colon) were fixed in methacarn (American Master Tech Scientific) immediately after collection and stored overnight at 22 °C. The following day, fixed tissues were washed twice with 70% ethanol and embedded into paraffin blocks (Leica EG1150C). To evaluate Candida morphology in the mouse gastrointestinal tract, 10-µm tissue sections, made by an automated microtome (Leica RM2265), were deparaffinized, blocked with PBS plus 5% FBS for 30 min at 22 °C, and incubated with an anti-Candida antibody coupled to fluorescein isothiocyanate (FITC) (1:500 dilution; PA173154, Thermo Fisher Scientific) overnight at 4 °C. This was followed by three washes with PBS at 22 °C and staining of the epithelial nuclei with 4,6-diamidino-2-phenylindole (DAPI, Invitrogen). Cell counting was carried out using a Zeiss Axio Observer Microscope. Tissue sections from a group of 3–5 mice were stained and 100 to 1,200 Candida cells per section were assessed for morphology.
16S rRNA sequencing
Nucleic acids (DNA) were isolated from samples utilizing the ZymoBIOMICS Quick-DNA Fecal/Soil Microbe 96 Kit (Zymo Research, D6011) following the manufacturer’s guidelines. DNA was eluted in nuclease-free water, and quantification performed using the dsDNA-HS assay on a QubitTM 3.0 fluorometer (Thermo Fisher Scientific). The 16S rRNA V4 hypervariable region was amplified from the entire DNA pool by utilizing the barcoded 515 F forward primer and the 806 R reverse primers, as established by the Earth Microbiome Project62. This involved generating amplicons using 5X Phusion High-Fidelity DNA Polymerase: initial denaturation at 98 °C for 30 s, succeeded by 25 cycles of 98 °C for 10 s, 57 °C for 30 s, and 72 °C for 30 s, finalized by extension at 72 °C for 5 min. Following amplification, samples underwent visualization via gel electrophoresis and were combined in equal proportions. The consolidated amplicon library was analysed by the Rhode Island Genomics and Sequencing Center at University of Rhose Island using an Illumina MiSeq platform. Sequencing encompassed paired-end sequencing (2 × 250 bp) employing the 600-cycle kit along with established protocols. The raw reads were submitted to the NCBI Sequence Read Archive (SRA) and can be accessed under BioProject PRJNA1008281.
16S rRNA sequencing analysis
Raw paired-end FASTQ reads were demultiplexed using idemp (https://github.com/yhwu/idemp/blob/master/idemp.cp). Reads were subjected to quality filtering, trimming, de-noising with DADA263 (via q2-dada2), and merging using the Qiime2 (Quantitative Insights Into Microbial Ecology 2 program) pipeline (version 2021.8)64. Ribosomal sequence variants were aligned with mafft65 (via q2-alignment), and phylogenetic tree construction was done with fasttree266 (via q2-phylogeny). Taxonomic assignment was done with the pre-trained naive Bayes classifier and the q2-feature-classifier67 trained on the SILVA 132 99% database68. Alpha diversity (Shannon, Faith’s phylogenetic diversity) and beta diversity (Bray–Curtis dissimilarity)69,70 were calculated using the phyloseq package (version 1.42.0) in R (version 4.2.3)71. Significance was determined by the Benjamini, Krieger and Yekutieli test to correct for false discoveries with adjusted P value < 0.0572. Linear discriminant analysis (LDA) analysis was conducted using LEfSe (linear discriminant analysis effect size) (Galaxy Version 1.0, http://huttenhower.sph.harvard.edu/galaxy)73,74.
Quantitative PCR of fecal bacteria
Quantitative PCR (iTaq Universal SYBR Green Supermix, Bio-Rad) was performed to determine the bacterial loads in fecal pellets using universal 16S rRNA primers (oligonucleotide 7212 and 7213) as previously described31. Bacterial genomic DNA (gDNA) was purified using a ZymoBIOMICS DNA Miniprep Kit (Zymo Research). Bacterial abundance was determined using standard curves with gDNA from in vitro E. coli culture.
Transcriptional profiling
Isolation of RNA was performed as described75. In brief, C. albicans cells were collected from 30 °C YPD overnight cultures, washed with PBS and 107 yeast cells per ml incubated in petri dishes for 3 h at 37 °C 5% CO2 in RPMI. Hyphal cells were collected by scraping and subsequent centrifugation at 3,000g for 2 min. RNA was isolated using the RNeasy Minikit (Qiagen) combined with an upstream glass bead disruption protocol. RNA concentration and quality were verified using the NanoDrop 1000 and the Agilent 2100 Bioanalyzer Nanochip system, respectively, according to the manufacturers’ instructions. Microarray analyses were performed as described previously76,77. In brief, the one-colour QuickAmp labelling kit (Agilent) was used to generate cRNA with fluorescently labelled CTP (Cy5 or Cy3 CTP; GE Healthcare) from high-quality total fungal RNA. Labelled cRNA was purified using the RNeasy Minikit (Qiagen), and sufficient dye incorporation was verified by spectrophotometry with the NanoDrop instrument. C. albicans arrays were purchased from Agilent technologies (GEO accession number GPL19932). The Gene Expression Hybridization kit (Agilent) was used according to the manufacturer’s instructions, using a two-colour hybridization system. The arrays were hybridized in triplicate for each strain. The arrays were scanned with a GenePix 4200AL scanner (GenePix Pro 6.1; 635 and 594 nm; automatically determined photomultiplier tube gains; pixel size 5 nm). Data were extracted using the Agilent Feature Extractor (Version 12.0) and imported into GeneSpring 14.9 (Agilent) for analysis. All microarray data are available at ArrayExpress under accession number MTAB-13349.
In vitro competition experiments
Assays were conducted between WT and ece1Δ/Δ reporter strains under hypoxia, low pH, oxidative stress, osmotic stress, or in the presence of a short-chain fatty acid (sodium acetate (Sigma-Aldrich)). Strains were grown overnight in 1% YPD at 30 °C and 180 rpm. 500 µl of each strain was washed twice with sterile PBS and resuspended in the same volume. Mixtures of 1:1 WT and ece1Δ/Δ reporter strains were incubated for 24 h in liquid RPMI to a final concentration of 2 × 105 cells per ml in the 12-well plate. After 24 h, the 12-well plate was centrifuged for 10 min at 4,000g and the supernatant was carefully removed. The hyphal aggregates were resuspended in 1 ml PBS containing 1 mg Zymolyase 20 T (Amsbio) and incubated for 3 h at 37 °C. Following Zymolyase treatment, hyphal fragments were transferred to a 1.5 ml microcentrifuge, collected at 10,000g for 5 min, washed once with PBS, then resuspended in the same volume. Quantification of the two labelled populations in a mixture was carried out via BD FACSVerse flow cytometer (BD Bioscience) equipped with an argon laser emitting at 488 nm. The fluorescence signal of the GFP reporter strains was detected using the FITC-A channel equipped with a 527 nm band-pass filter (bandwidth 15 nm) while the RFP signal was detected using the PerCP-A channel equipped with a 700 nm band-pass filter (bandwidth 54 nm). Gating of single hyphal cells was achieved as follows. First, the fragmented hyphal populations were separated from cell debris and selected via SSC-A (side scatter) and FSC-A (forward scatter) signals. Second, duplet cells were excluded via FSC-W and FSC-H signals. A total of 100,000 events was analysed per sample at a flow rate of approximately 1,000 events per second. Both qualitative and quantitative analyses of the C. albicans labelled strains were performed with FlowJo (Version 10.2) software.
In vitro competition experiments on epithelial cell lines were conducted using TR146 human buccal epithelial cells from ECACC (European Collection of Authenticated Cell Cultures; Sigma-Aldrich, 10032305-1VL). The cells were cultured in DMEM-F12 with 10% FBS at 37 °C and 5% CO2. For experiments, TR146 cells were seeded and incubated till confluency. Reporter strains of C. albicans from overnight cultures were mixed in equal proportions in DMEM-F12 to a concentration of 1 × 107 cells per ml. Infection mixtures were added to TR146 cells to a final concentration of 2 × 105 Candida cells per ml. After 24 h, cells were centrifuged (10 min, 4,000g), washed with PBS, and treated with DNase I (2 µl; 1 U µl−1) in DNase I buffer (10 mM Tris-HCl, 2.5 mM MgCl2, 0.5 mM CaCl2; pH 7.6) for 1 h at 37 °C. Cells were centrifuged and the supernatant removed. Hyphal aggregates were fragmented using 1 mg ml−1 Zymolyase 20 T (Amsbio) in PBS for 3 h at 37 °C. Samples were centrifuged and resuspended in 1 ml PBS and analysed by flow cytometry as described above (500,000 events analysed at a flow rate of 2,000 events per s). Single reporter strains seeded on TR146 cells and mixed in a 1:1 ratio after 24 h growth were used as controls.
The competitive index (CI) was used to estimate the relative fitness of a single strain in competition assays. To calculate the CI between two competing strains, the following formula was used:
In vitro candidalysin assays
Candidalysin peptide used in this study was synthetically synthesized corresponding to the processed third peptide of the precursor protein Ece1 of C. albicans (SIIGIIMGILGNIPQVIQIIMSIVKAFKGNK)4. Peptides were obtained from Peptide Protein Research, dissolved in ultra-pure water (MQ water, Merck Millipore) to a concentration of 1.4 mM and stored at −20 °C.
Bacterial CFU assays with candidalysin
E. coli, K. pneumoniae and E. faecium were grown in LB at 37 °C at 180 rpm overnight. Stationary phase cultures were washed twice with PBS and set to OD 0.04 in PBS in 1.5 ml tubes. Cell suspensions were incubated for 2 h at 37 °C with shaking. Suspensions were re-diluted to OD600 = 0.02 with PBS with or without 70 µM of the peptide and incubated at 37 °C with shaking. Every 30 min, cells were removed and plated on LB agar plates and incubated overnight. Analysis was performed in biological triplicates for each bacterial species.
XTT assay
XTT assays were performed to analyse the effect of candidalysin on the metabolic activity of bacteria by adaption of assays from Miramón et al.78. Stationary phase overnight cultures of bacteria were washed twice with PBS and adjusted to an OD600, which is equivalent to 109 CFU ml−1. Candidalysin dilutions were prepared in MQ water. The test conditions contained 50 µl of the respective peptide dilution (final concentration: 20–70 µM), 50 µl of the bacterial suspension and 100 µl DMEM. For controls, peptide dilutions were replaced with MQ water, and the bacteria suspension was replaced by PBS. Plates were incubated at 37 °C, 18% O2 and 5% CO2. After 200 min, 100 µl of XTT mixture (final 1 mg ml−1 XTT (Fluka) and 0.1 mg ml−1 coenzyme Q0 (2,3-dimethoxy-5-methyl-p-benzoquinone) (Sigma)) was added to each well and the plate was incubated for another 60 min. The plate was centrifuged at 250g for 10 min. The supernatant was transferred to a clean 96-well plate and measured in a microplate reader (absorbance 451 nm, absorbance 600 nm for reference). All conditions were done in technical duplicates and the assay was performed at least three times for each of the selected bacteria. The value of the reference measurement and the value of the media control were subtracted from the data. The data of the test conditions was evaluated as a percentage of the normal control.
Glucose consumption assay
Stationary phase overnight cultures of bacteria were washed twice with PBS and adjusted to an OD600, which is equivalent to 109 CFU ml−1. The test conditions contained 50 µl of the respective peptide dilution (final concentration: 70 µM), 50 µl of bacterial suspension and 100 µl minimal media (0.67% YNB, 0.2% glucose, 1% casamino acids, 25 mM MES pH 6). For controls, peptide dilutions were replaced with MQ water. The plate was incubated at 37 °C, 18% O2 and 5% CO2 for 200 min before the plate was centrifuged at 250g for 10 min. The supernatant was diluted in PBS in a clean 96-well plate and measured with the GlucoseGlo-Kit (Promega) according to the manufacturer’s instructions. The background measurement (PBS only) was subtracted from each sample. All conditions were done in technical duplicates and the assay was performed at least three times for each of the selected bacteria.
Statistical analysis
For significant differences between experimental groups MaAsLin2 (microbiome multivariable association with linear models)79 was used with R version 4.2.3. Effect size calculations were done with PERMANOVA (permutational multivariate ANOVA) with the adonis function in phyloseq version 1.42.0 with R version 4.2.3. For LDA analysis by LEfSe, differentially abundant species with LDA score >2 is shown. Kruskal–Wallis test was used to test to compare features between diets (P < 0.05). Pairwise Wilcoxon test was used to compare between taxa (P < 0.05). Paired parametric t-tests were used to determine significant differences in the proportion of two C. albicans populations in the in vivo competition assays. Unpaired nonparametric Mann–Whitney U-tests were used to determine the significance between colonization levels of different strains in the monocolonization assays. Unpaired parametric t-tests were used to determine significant differences in the treated and control samples for in vitro bacterial CFU assays, XTT assays and glucose consumption assays. P values for all t-tests were calculated using GraphPad Prism (GraphPad Software v.10) and are indicated in the figure legends. P < 0.05 is considered significant.
Ethics
Animal studies were performed according to approved protocols by the Institutional Animal Care and Use Committee (IACUC) of each institution in the US and the animal studies and protocols (permission numbers 16/2307 and 16/2372) were approved by the local government of Lower Saxony, Germany.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The 16S sequencing data are publicly available at the NCBI Sequence Read Archive (SRA) and can be accessed under BioProject PRJNA1008281. Microarray data for analysis of C. albicans expression are available at ArrayExpress under accession MTAB-13349. Source data are provided with this paper.
References
Ost, K. S. et al. Adaptive immunity induces mutualism between commensal eukaryotes. Nature 596, 114–118 (2021).
Tso, G. H. W. et al. Experimental evolution of a fungal pathogen into a gut symbiont. Science 362, 589–595 (2018).
Witchley, J. N. et al. Candida albicans morphogenesis programs control the balance between gut commensalism and invasive infection. Cell Host Microbe 25, 432–443.e436 (2019).
Moyes, D. L. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016).
Li, X. V. et al. Immune regulation by fungal strain diversity in inflammatory bowel disease. Nature 603, 672–678 (2022).
Doron, I. et al. Mycobiota-induced IgA antibodies regulate fungal commensalism in the gut and are dysregulated in Crohn’s disease. Nat Microbiol 6, 1493–1504 (2021).
Rao, C. et al. Multi-kingdom ecological drivers of microbiota assembly in preterm infants. Nature 591, 633–638 (2021).
Iliev, I. D. & Cadwell, K. Effects of intestinal fungi and viruses on immune responses and inflammatory bowel diseases. Gastroenterology 160, 1050–1066 (2021).
Swidergall, M. & LeibundGut-Landmann, S. Immunosurveillance of Candida albicans commensalism by the adaptive immune system. Mucosal Immunol. 15, 829–836 (2022).
Shao, T. Y., Haslam, D. B., Bennett, R. J. & Way, S. S. Friendly fungi: symbiosis with commensal Candida albicans. Trends Immunol. 43, 706–717 (2022).
Li, Q. et al. Dysbiosis of gut fungal microbiota is associated with mucosal inflammation in Crohn’s disease. J. Clin. Gastroenterol. 48, 513–523 (2014).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Bacher, P. et al. Human anti-fungal Th17 immunity and pathology rely on cross-reactivity against Candida albicans. Cell 176, 1340–1355.e1315 (2019).
Shao, T. Y. et al. Commensal Candida albicans positively calibrates systemic Th17 immunological responses. Cell Host Microbe 25, 404–417.e406 (2019).
Yeung, F. et al. Altered immunity of laboratory mice in the natural environment is associated with fungal colonization. Cell Host Microbe 27, 809–822.e806 (2020).
Zhai, B. et al. High-resolution mycobiota analysis reveals dynamic intestinal translocation preceding invasive candidiasis. Nat. Med. 26, 59–64 (2020).
Pappas, P. G., Lionakis, M. S., Arendrup, M. C., Ostrosky-Zeichner, L. & Kullberg, B. J. Invasive candidiasis. Nat. Rev. Dis. Primers 4, 18026 (2018).
Koh, A. Y., Kohler, J. R., Coggshall, K. T., Van Rooijen, N. & Pier, G. B. Mucosal damage and neutropenia are required for Candida albicans dissemination. PLoS Pathog. 4, e35 (2008).
Noble, S. M., Gianetti, B. A. & Witchley, J. N. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat. Rev. Microbiol. 15, 96–108 (2017).
Kadosh, D. Morphogenesis in Candida albicans: Cellular and Molecular Biology (ed. Prasad, R.) 41–62 (Springer, 2017).
Saville, S. P., Lazzell, A. L., Monteagudo, C. & Lopez-Ribot, J. L. Engineered control of cell morphology in vivo reveals distinct roles for yeast and filamentous forms of Candida albicans during infection. Eukaryot. Cell 2, 1053–1060 (2003).
Lo, H. J. et al. Nonfilamentous C. albicans mutants are avirulent. Cell 90, 939–949 (1997).
Carlisle, P. L. et al. Expression levels of a filament-specific transcriptional regulator are sufficient to determine Candida albicans morphology and virulence. Proc. Natl Acad. Sci. USA 106, 599–604 (2009).
Bohm, L. et al. The yeast form of the fungus Candida albicans promotes persistence in the gut of gnotobiotic mice. PLoS Pathog. 13, e1006699 (2017).
Liang, S. H. et al. Hemizygosity enables a mutational transition governing fungal virulence and commensalism. Cell Host Microbe 25, 418–431.e416 (2019).
Mogavero, S. et al. Candidalysin delivery to the invasion pocket is critical for host epithelial damage induced by Candida albicans. Cell Microbiol. 23, e13378 (2021).
Naglik, J. R., Gaffen, S. L. & Hube, B. Candidalysin: discovery and function in Candida albicans infections. Curr. Opin. Microbiol. 52, 100–109 (2019).
Stoldt, V. R., Sonneborn, A., Leuker, C. E. & Ernst, J. F. Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J. 16, 1982–1991 (1997).
Braun, B. R. & Johnson, A. D. TUP1, CPH1 and EFG1 make independent contributions to filamentation in Candida albicans. Genetics 155, 57–67 (2000).
Wakade, R. S., Huang, M., Mitchell, A. P., Wellington, M. & Krysan, D. J. Intravital imaging of Candida albicans identifies differential in vitro and in vivo filamentation phenotypes for transcription factor deletion mutants. mSphere 6, e0043621 (2021).
Fan, D. et al. Activation of HIF-1α and LL-37 by commensal bacteria inhibits Candida albicans colonization. Nat. Med. 21, 808–814 (2015).
Yamaguchi, N. et al. Gastric colonization of Candida albicans differs in mice fed commercial and purified diets. J. Nutr. 135, 109–115 (2005).
McDonough, L. D. et al. Candida albicans isolates 529L and CHN1 exhibit stable colonization of the murine gastrointestinal tract. mBio 12, e0287821 (2021).
Braun, B. R., Kadosh, D. & Johnson, A. D. NRG1, a repressor of filamentous growth in C. albicans, is down-regulated during filament induction. EMBO J. 20, 4753–4761 (2001).
Murad, A. M. et al. NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans. EMBO J. 20, 4742–4752 (2001).
Wakade, R. S., Kramara, J., Wellington, M. & Krysan, D. J. Candida albicans filamentation does not require the cAMP–PKA pathway in vivo. mBio 13, e0085122 (2022).
Vautier, S. et al. Candida albicans colonization and dissemination from the murine gastrointestinal tract: the influence of morphology and Th17 immunity. Cell Microbiol 17, 445–450 (2015).
Miller, B. M., Liou, M. J., Lee, J. Y. & Baumler, A. J. The longitudinal and cross-sectional heterogeneity of the intestinal microbiota. Curr. Opin. Microbiol. 63, 221–230 (2021).
Brugiroux, S. et al. Genome-guided design of a defined mouse microbiota that confers colonization resistance against Salmonella enterica serovar Typhimurium. Nat Microbiol 2, 16215 (2016).
Trexler, P. C., & Orcutt, R.P. Chapter Sixteen: Development of Gnotobiotics and Contamination Control in Laboratory Animal Science. In: 50 Years of Laboratory Animal Science. Memphis, TN: Am Assoc Lab Anim Sci, 121–128 (2000).
Caballero, S. et al. Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell Host Microbe 21, 592–602.e594 (2017).
Dambuza, I. M. & Brown, G. D. Managing the mycobiota with IgA. Nat. Microbiol. 6, 1471–1472 (2021).
Kasper, L. et al. The fungal peptide toxin candidalysin activates the NLRP3 inflammasome and causes cytolysis in mononuclear phagocytes. Nat. Commun. 9, 4260 (2018).
Swidergall, M. et al. Candidalysin is required for neutrophil recruitment and virulence during systemic Candida albicans infection. J. Infect. Dis. 220, 1477–1488 (2019).
White, S. J. et al. Self-regulation of Candida albicans population size during GI colonization. PLoS Pathog. 3, e184 (2007).
Hoyer, L. L., Payne, T. L., Bell, M., Myers, A. M. & Scherer, S. Candida albicans ALS3 and insights into the nature of the ALS gene family. Curr. Genet. 33, 451–459 (1998).
Phan, Q. T. et al. Als3 is a Candida albicans invasin that binds to cadherins and induces endocytosis by host cells. PLoS Biol. 5, e64 (2007).
Almeida, R. S. et al. The hyphal-associated adhesin and invasin Als3 of Candida albicans mediates iron acquisition from host ferritin. PLoS Pathog. 4, e1000217 (2008).
Martchenko, M., Alarco, A. M., Harcus, D. & Whiteway, M. Superoxide dismutases in Candida albicans: transcriptional regulation and functional characterization of the hyphal-induced SOD5 gene. Mol. Biol. Cell 15, 456–467 (2004).
Fradin, C. et al. Granulocytes govern the transcriptional response, morphology and proliferation of Candida albicans in human blood. Mol. Microbiol. 56, 397–415 (2005).
Hube, B. Fungal adaptation to the host environment. Curr. Opin. Microbiol. 12, 347–349 (2009).
Dewhirst, F. E. et al. Phylogeny of the defined murine microbiota: altered Schaedler flora. Appl. Environ. Microbiol. 65, 3287–3292 (1999).
Guthrie, C. & Fink, G. R. Guide to Yeast Genetics and Molecular Biology (Academic Press, 1991).
Liu, H., Kohler, J. & Fink, G. R. Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 266, 1723–1726 (1994).
Park, S. O., Frazer, C. & Bennett, R. J. An adjuvant-based approach enables the use of dominant HYG and KAN selectable markers in Candida albicans. mSphere 7, e0034722 (2022).
Reuss, O., Vik, A., Kolter, R. & Morschhauser, J. The SAT1 flipper, an optimized tool for gene disruption in Candida albicans. Gene 341, 119–127 (2004).
Noble, S. M. & Johnson, A. D. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot. Cell 4, 298–309 (2005).
Mancera, E. et al. Genetic modification of closely related Candida species. Front. Microbiol. 10, 357 (2019).
Gerami-Nejad, M., Zacchi, L. F., McClellan, M., Matter, K. & Berman, J. Shuttle vectors for facile gap repair cloning and integration into a neutral locus in Candida albicans. Microbiology 159, 565–579 (2013).
Hollomon, J. M. et al. The Candida albicans Cdk8-dependent phosphoproteome reveals repression of hyphal growth through a Flo8-dependent pathway. PLoS Genet. 18, e1009622 (2022).
Dallari, S. et al. Enteric viruses evoke broad host immune responses resembling those elicited by the bacterial microbiome. Cell Host Microbe 29, 1014–1029.e1018 (2021).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Bokulich, N. A. et al. q2-longitudinal: longitudinal and paired-sample analyses of microbiome data. mSystems 3, e00219–18 (2018).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Bray, J. R. & Curtis, J. T. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monogr. 27, 325–349 (1957).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6, 90 (2018).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Benjaminii, Y. & Hichberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Mogavero, S. & Hube, B. Candida albicans interaction with oral epithelial cells: adhesion, invasion, and damage assays. Methods Mol. Biol. 2260, 133–143 (2021).
Gerwien, F. et al. A novel hybrid iron regulation network combines features from pathogenic and nonpathogenic yeasts. mBio 7, e01782-16 (2016).
Ramirez-Zavala, B. et al. The Snf1-activating kinase Sak1 is a key regulator of metabolic adaptation and in vivo fitness of Candida albicans. Mol. Microbiol. 104, 989–1007 (2017).
Miramon, P. et al. A family of glutathione peroxidases contributes to oxidative stress resistance in Candida albicans. Med. Mycol. 52, 223–239 (2014).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Acknowledgements
This work was supported by NIH grants AI166869, AI141893, AI081704 and AI168222 to R.J.B. Y.-H.C. was supported by the Charles H. Revson Senior Fellowship in Biomedical Science. B.H. and T.B.S. were supported by the German Research Foundation (Deutsche Forschungsgemeinschaft (DFG)) within the Cluster of Excellence ‘Balance of the Microverse’, under Germany’s Excellence Strategy, EXC 2051, project ID 390713860. B.H. and S.A. were further supported by the DFG, project HU 528/20-1. J.C.P was supported by AI175081, P.B. was supported by DK125382 and S.P. was supported by a Graduate Research Fellowship from the NSF under award number 1644760. I.V.E. was supported by the Institut Pasteur and is a CIFAR Azrieli Global Scholar in the CIFAR Program Fungal Kingdom: Threats & Opportunities. We thank E. Pamer, I. Jacobsen and T. Hohl for sharing strains and the National Gnotobiotic Rodent Resource Center (NIH grants P40 OD010995 and P30 DK034987) for support.
Author information
Authors and Affiliations
Contributions
R.J.B. conceived the majority of the experiments and wrote the initial manuscript draft, with input from S.-H.L. and S.S. S.-H.L. and S.S. performed the majority of the experiments. P.K. performed the microscopic analysis of yeast and hyphal cells. L.D.M. constructed strains and analysed in vivo phenotypes. J.D. performed analyses of C. albicans strains with individual bacterial strains in mice. C.F. assisted with design and construction of C. albicans mutant strains. S.V. provided gnotobiotic mice and advised on the project. J.C.P., K.C. and Y.-H.C. performed consortia experiments. S.P. and P.B. analysed 16S data and generated related figures. T.B.S., S.A., M.H., S.M. and O.E. performed analyses of C. albicans with candidalysin. O.E. performed in vitro C. albicans competition experiments. S.A. and S.M. performed transcriptional profiling of the ece1 mutant. B.H. conceived the in vitro experiments, provided the C. albicans ece1 mutant strains and advised on the project. I.V.E. performed in vivo experiments and advised on the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of the fitness of WT and yeast-locked efg1Δ/Δ cells from different C. albicans strain backgrounds.
a, Experiments were performed as described in Fig. 1. Mouse GI organs (St; stomach, Si; small intestine, Ce; cecum, Co; colon) were homogenized to identify WT and efg1Δ/Δ cells 14 days post-C. albicans inoculation. WT (CAY2698) versus efg1Δ/Δ (CAY11750) competitions in SD + P/S (P/S; penicillin/streptomycin) model (a), PD + P/S model (b) and PD model (c). dpi; days post-inoculation. d, Schematic of competition between WT and yeast-locked efg1Δ/Δ cells. Mice fed the SD without antibiotics were gavaged with a 1:1 mixture of WT and efg1Δ/Δ cells and C. albicans colonies were examined in fecal pellets at the indicated time points. e,f, Competition outcomes (e) and fecal CFUs (f) of WT versus efg1Δ/Δ in 529L (CAY5016 versus CAY11482) and CHN1 (CAY11170 versus CAY11184). Experiments were performed in BALB/c and C57BL/6 J mice as indicated. Each data point represents an individual mouse, data are mean ± s.e.m in a, b, and f. n = 3 for competitions in BALB/c mice and n = 4 for competitions in C57BL/6 J mice. A paired t-test (two-tailed) was used in e and a Mann-Whitney (two-tailed) test in f.
Extended Data Fig. 2 Comparison of the fitness of WT and flo8Δ/Δ cells in different colonization models.
a, Microscopic images of cells in the colon of BALB/c SD + P/S mice infected with WT (CAY2698) or flo8Δ/Δ (CAY9796) cells at 7 days post-infection. b, Quantitative analysis of yeast and filamentous morphotypes. n = 2. c, Schematic of competition between WT and flo8Δ/Δ cells. n = 2. Each data point represents an individual mouse. A 1:1 mix of WT and flo8Δ/Δ cells were inoculated by oral gavage and fecal samples were collected at the indicated time points. d-f, Results for SC5314 WT (CAY2698) versus flo8Δ/Δ (CAY9796) cells in GF BALB/c mice (d), conventional PD-fed BALB/c mice treated with antibiotics (P/S; penicillin/streptomycin) (e), and conventional PD-fed BALB/c mice (without antibiotics) (f). dpi; days post-inoculation. n = 3 in e and f. g-i, Competition of WT versus flo8Δ/Δ cells in BALB/c mice fed a SD (without antibiotics). Experiments were performed using SC5314 (CAY2698 vs. CAY9796) (g), 529 L (CAY11168 vs. CAY11180) (h), or CHN1 (CAY11170 vs. CAY11186) (i) strain backgrounds. n = 3. Each data point represents an individual mouse. Data are mean ± s.e.m in b. Significance was determined by a Mann-Whitney (two-tailed) test in b and a paired two-tailed t-test in d-i.
Extended Data Fig. 3 Analysis of microbiome composition upon changes in diet, antibiotic treatment, and C. albicans colonization.
a, Schematic of colonization experiments. BALB/c mice were fed the SD or PD, with or without antibiotics (P/S; penicillin/streptomycin). These groups were compared with and without colonization with C. albicans SC5314 cells. b, C. albicans CFUs in fecal pellets collected at the indicated time points. LOD, limit of detection. n = 8. c,d, Total bacterial levels in fecal pellets collected at the indicated time points by quantitative PCR in SD- or PD-fed mice with antibiotics (c) and without antibiotics (d). n = 8. e, Bubble plot depicting the amount of variation in gut microbial composition determined by Permutational Multivariate Analysis of Variance (PERMANOVA) analysis using the adonis function and Bray-Curtis distances of beta diversity. Effect size refers to the magnitude of the differences or dissimilarities between groups. f, Shannon diversity of control mice on SD or PD (n = 8). g-m, Distribution of bacterial phyla in different models as determined by MaAsLin2. PD vs. SD (g), SD vs. SD + P/S (h), PD vs. PD + P/S (i), SD +/- C. albicans (j), PD +/- C. albicans (k), PD + P/S +/- C. albicans (l) and SD + P/S +/- C. albicans (m). Coefficient with standard error shown on x-axis. adj p-val cutoff 0.05. n, Relative abundance of Actinobacteria across different diet fed mice (+/− Candida) on Day 21. n = 8. Each data point represents an individual mouse. Data are mean ± s.e.m in b-m. A Mann-Whitney test was used in b,f,and n.
Extended Data Fig. 4 Linear discriminant analysis Effect Size (LEfSe) analysis to compare the alterations in gut bacterial populations in mice on different diets.
Linear discriminant analysis (LDA) effect size for significant taxa in the microbiome of mice fed the SD vs. PD are plotted onto a cladogram for Day 0 (a), Day 7 (b), Day 14 (c), Day 21 (d). and for mice fed the PD (+ /− C. albicans) for Day 0 (e), Day 7 (f), Day 14 (g) and Day 21 (h). Analysis was performed on mice as shown in Extended Data Fig. 3 (n = 8 per group). Differentially abundant species with LDA score >2 are shown in nodes represented with red (PD) and green (SD) and non-significant species are represented with yellow. A Kruskal-Wallis test was used to compare features between diets (p < 0.05) and the Pairwise Wilcoxon test was used to compare between taxa (p < 0.05).
Extended Data Fig. 5 Analysis of gut bacterial populations in different murine models.
16S rRNA sequencing data was used to determine the relative abundance of bacterial phyla in BALB/c colonization experiments using mice fed the SD or PD, and supplemented or not supplemented with antibiotics (P/S; penicillin/streptomycin). Each group of mice were also compared +/− inoculation with C. albicans SC5314 cells, as shown in Extended Data Fig. 3. a,b, Shannon diversity for bacterial populations from fecal pellets for the shown experiments. n = 8. c, Relative abundance of bacteria shown for fecal pellets for days 0, 10, 14, and 21, and for the small intestine at day 21. n = 8. Data are mean ± s.e.m in a-c.
Extended Data Fig. 6 C. albicans WT cells outcompete efg1Δ/Δ cells in gnotobiotic mice harboring different bacterial populations.
a, Schematic of competition between WT and yeast-locked efg1Δ/Δ cells in gnotobiotic colonization models. Mice were gavaged with a 1:1 mixture of WT SC5314 (CAY2698) and efg1Δ/Δ (CAY11750) cells. b-d, C. albicans cells were tested in GF BALB/c mice (b), GF NMRI mice (c), or GF C57BL/6 mice colonized with E. coli prior to inoculation with C. albicans (d). n = 4 in b, 8 in c, and 5 in d. e, C. albicans CFUs in fecal pellets collected at the indicated time points in GF NMRI mice (e) and GF C57BL/6 mice (f). n = 8 in e, n = 3–5 in f. LOD, limit of detection. g, WT v. efg1Δ/Δ competition in Amp-treated mice gavaged with heat-killed AmpR E. coli. n = 4. h,i, Competition between CHN1 WT (CAY11170) and efg1∆/∆ (CAY11184) cells in WT mice (h) or Rag1-/- mice (i) on the SD (no antibiotics). n = 4. j, C. albicans CFU levels in the fecal pellets from WT and Rag1-/- mice. n = 4. Each data point represents an individual mouse. Data are mean ± s.e.m in j. A paired t-test (two-tailed) was used in b-d and g-i, and a Mann-Whitney (two-tailed) test in e,f and j.
Extended Data Fig. 7 Examining the role of ALS3 and SOD5 in GI colonization fitness.
a, Schematic of competition between WT C. albicans and sod5Δ/Δ or als3Δ/Δ cells (SC5314 background). b-e, BALB/c mice fed a PD without antibiotics or a SD with antibiotics (P/S; penicillin/streptomycin). Experiments used WT (CAY8785) and sod5Δ/Δ (CAY14738) in b and c. n = 4. WT (CAY8785) and als3Δ/Δ (CAY14696) in d and e. n = 8. Each data point represents an individual mouse. A paired t-test (two-tailed) was used for statistical significance.
Extended Data Fig. 8 Colonization fitness of WT and ece1Δ/Δ mutants.
Experiments were performed as described in Fig. 3 Mouse GI organs (St; stomach, Si; small intestine, Duo; duodenum, Je; jejunum, Ile; ileum, Ce; cecum, Co; colon) were homogenized to identify the ratio of WT and ece1Δ/Δ cells 14 days post C. albicans infection. b, Competition between CAY11533 (WT) versus CAY11507 (ece1Δ/Δ) cells in BALB/c fed a PD. n = 5. c, Competition between WT (CAY11168) and ece1Δ/Δ cells (CAY12441) in the 529L background BALB/c mice fed a PD. n = 4. d, Competition between WT (CAY11170) and ece1Δ/Δ cells (CAY12446) in the CHN1 background in SD-fed BALB/c mice. n = 4. e-g, Competition between CAY12202 (WT) versus CAY8578 (ece1Δ/Δ) BALB/c mice fed a SD plus antibiotics (P/S; penicillin/streptomycin) (e), GF C57BL/6 mice (f), and GF C57BL/6 mice colonized with E. coli prior to C. albicans inoculation (g). n = 4 in e, and 5 in f and g. Competition outcomes of WT (CAY12202) versus ece1Δ/Δ+ECE1ΔpIII (CAY8580) in GF C57BL/6 mice (h) and in BALB/c mice fed a PD (i). n = 4 in h and i. Each data point represents an individual mouse. Data are mean ± s.e.m in c–e, h, i. A paired t-test (two-tailed) was used to determine the significance between two populations (b-h).
Extended Data Fig. 9 Comparison of C. albicans WT and ece1Δ/Δ cells in vitro.
a, Heat map of transcriptome analysis representing the fold change (FC) in expression of fungal genes when strains were grown under hyphal-inducing conditions (3 h, 37 °C, 5% CO2). b-h, Competitive fitness of SC5314 WT and ece1Δ/Δ cells in RPMI (37 °C, 5% CO2) (b), acidic pH (pH 4.7) (c), high salt (1 M NaCl) (d), hypoxia (1% O2) (e), oxidative stress (2 mM H2O2) (f), in the presence of 2% acetate (g), and during incubation with TR146 epithelial cells (h). Competitive index was calculated using the formula: log2 [(MUTcompetition/WTcompetition)/ (MUTsingle/WTsingle)]. Fitness of the ece1Δ/Δ relative to the WT is represented as log2 competitive index. Data are representative of three biological replicates (n = 3). Graphs show the mean ± sem in b - h. Statistical analysis was performed using 1-way ANOVA with Bonferroni post hoc test to detect significance.
Supplementary information
Supplementary Tables
Supplementary Tables 1–3
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, SH., Sircaik, S., Dainis, J. et al. The hyphal-specific toxin candidalysin promotes fungal gut commensalism. Nature 627, 620–627 (2024). https://doi.org/10.1038/s41586-024-07142-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07142-4
- Springer Nature Limited
This article is cited by
-
Hyphae promote Candida albicans fitness and commensalism in the gut
Nature Reviews Microbiology (2024)
-
Candida albicans — der Pilz in uns
BIOspektrum (2024)
-
The pathobiology of human fungal infections
Nature Reviews Microbiology (2024)