Abstract
Azathioprine (AZA) and its metabolite, mercaptopurine (6-MP), are widely used immunosuppressant drugs. Polymorphisms in genes implicated in AZA/6-MP metabolism, reportedly, could account in part for their potential toxicity. In the present study we performed a systematic review and a meta-analysis, comprising 30 studies and 3582 individuals, to investigate the putative genetic association of two inosine triphosphatase (ITPA) polymorphisms with adverse effects in patients treated with AZA/6-MP. We found that rs1127354 is associated with neutropenia in general populations and in children (OR: 2.39, 95%CI: 1.97–2.90, and OR: 2.43, 95%CI: 2.12–2.79, respectively), and with all adverse effects tested herein in adult populations (OR: 2.12, 95%CI: 1.22–3.69). We also found that rs7270101 is associated with neutropenia and leucopenia in all-ages populations (OR: 2.93, 95%CI: 2.36–3.63, and OR: 2.82, 95%CI: 1.76–4.50, respectively) and with all adverse effects tested herein in children (OR: 1.74, 95%CI: 1.06–2.87). Stratification according to background disease, in combination with multiple comparisons corrections, verified neutropenia to be associated with both polymorphisms, in acute lymphoblastic leukemia (ALL) patients. These findings suggest that ITPA polymorphisms could be used as predictive biomarkers for adverse effects of thiopurine drugs to eliminate intolerance in ALL patients and clarify dosing in patients with different ITPA variants.
Similar content being viewed by others
Introduction
The thiopurine drugs, mercaptopurine (6-MP) and its prodrug, azathioprine (AZA), are purine analogs which are widely used immunosuppressant agents to treat patients with inflammatory bowel disease (IBD), acute lymphoblastic leukemia (ALL), rheumatoid arthritis (RA) and other diseases, as well as for patients having received a transplant [1,2,3,4]. Although these drugs are potent and effective medications, 20% of the patients present serious toxicity that may lead to therapy discontinuation, while 9% of the patients are resistant to thiopurine treatment [5,6,7,8,9,10,11]. The most common reported adverse effects include leucopenia, neutropenia, myelosuppression, pancreatitis, hepatotoxicity, while flu-like symptoms, skin reaction, rash, alopecia, nausea and vomiting seem to be rarer [10]. Because adverse effects are idiosyncratic, thiopurine dosing is difficult to be calculated [7]. In some people, AZA/6-MP can accumulate in their bodies and become toxic, while others have no negative reaction.
AZA is the pro-drug and is metabolized to 6-MP in the liver by the enzyme glutathione-S-transferase (GST) [12]. 6-MP can be metabolized to 6-methylmercaptopurine (6-MMP) by the enzyme thiopurine methyltransferase (TPMT) and 6-thiouric acid (6-TU) by the enzyme xanthine oxidase (XO) [13]. Both 6-MMP and 6-TU are inactive metabolites of MP. In the main pathway 6-MP is metabolized to 6-thioinosine 5ʹ-monophosphate (6-TIMP) and finally to 6-thioguanine nucleotide (6-TGN), the active metabolite of the drug. However, 6-TIMP can also be converted to thioinosine triphosphate (6-TITP) a metabolite with potential toxicity to cells [14, 15]. The enzyme inosine triphosphatase (ITPA) catalyzes the pyrophosphohydrolysis of 6-TITP back to 6-TIMP to eventually be metabolized to 6-TGN, the active metabolite for immune modulation. Though the exact mechanism of 6-TGN function is not fully elucidated yet, it is speculated (based on the structural similarity to the purine guanine), that immunosupression and cytotoxity is the result of inhibition of nucleotide synthesis (RNA, DNA) and hence protein synthesis, leading to inhibition of lymphocytes proliferation [16,17,18].
The cytotoxic and immunosuppressive properties of AZA/6-MP are mediated by 6-TGN [13, 19,20,21]. TPMT is a major determinant for the inactivation of AZA/6-MP. Among individuals, polymorphisms in TPMT have been identified, and there is high variability in the activity of the enzyme, being high, medium, and low metabolizers [7]. More recently, it has also been shown that NUDT15 polymorphisms are implicated in the tolerance of AZA/6-MP and the development of various types of adverse effects [22, 23]. A common practice has emerged to scan the individuals that are going to receive AZA/6-MP treatment for TPMT activity [7, 24, 25]. However, only TPMT or NUDT15 polymorphisms do not suffice for the prediction of all adverse effects, since about 70% of patients that show bone marrow suppression as adverse effect, have normal TPMT activity [24, 26, 27]. The above observations suggest that other factors are also in play to control adverse effects [28]. On the other hand, it is also known that genetic ITPA deficiency results in the cellular accumulation of the toxic 6-TITP that follows exposure to thiopurines [7]. ITPA is considered as a “house cleaning” gene since it hydrolyzes 6-TITP back to 6-TIMP, thus preventing the accumulation of 6-TITP (the rogue nucleotides) and promoting the production of 6-TGN [7]. It seems that a fine-tuned regulation between 6-TITP toxicity and 6-TGN therapeutic effect is accomplished by the proper expression of ITPA.
Among polymorphisms of ITPA, two are reported to show decreased enzymatic activity, and thus, they have been investigated for thiopurine-associated toxicity [29,30,31,32]. According to data obtained from the resolution of the crystal structure [33] and biochemical experiments [34], ITPA acts as a homodimer. Genotype and biochemical data from humans [35] further support the function of ITPA as homodimer. It is reported that decreased ITPA activity, is caused by a missense nucleotide change in exon 2 (rs1127354, or 94 C > A), resulting in a proline-to-threonine substitution (P32T), and heterozygotes present around 22% activity compared to wild type (wt) homozygotes, whereas risk homozygotes present zero activity [34,35,36]. For proteins participating in homodimers in an heterozygote individual the possible combinations for wt/wt, wt/mt and mt/mt dimers are 1:2:1, meaning that only one homodimer in four is fully functional [34]. This explanation is absolutely supported by the above mentioned 22% activity in rs1127354 heterozygotes. In terms of biological implication, this is translated to a dominant mode of inheritance [37, 38]. Another polymorphism, rs7270101 (IVS2 + 21 A > C) is located in the second intron, it modifies splicing and heterozygotes display around 60% activity of the wt homozygotes [35]. Thus, these two variations lead to lower ITPA enzymatic activity and subsequently to accumulation of 6-TITP in erythrocytes and increased toxicity of purine analog drugs [7, 29,30,31,32, 36, 39].
Several research groups have performed pharmacogenetic cohort studies in order to investigate the association between ITPA polymorphisms and AZA/6-MP toxicity. In 2007 a meta-analysis [40] was published but since then more data have become available. Herein, we conducted a systematic review and a meta-analysis to investigate putative associations of the ITPA polymorphisms rs1127354 and rs7270101 with adverse reactions after AZA/6-MP treatment in patients with any disease that had been treated for. We integrated data from thirty studies and 3582 individuals presenting all kinds of adverse effects described in the literature. The results of the present meta-analysis may help health professionals in decision making procedures for patients that are going to receive AZA/6-MP.
Material and methods
Literature search strategy
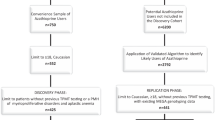
A systematic, computerized literature search in PubMed, SCOPUS, GOOGLE SCHOLAR and PharmGKB until December 2020 was performed to identify all articles in the literature reporting allele or genotype data. Effort was made to include conference papers and dissertations. Studies were identified and subsequently selected according to PICO framework [41] using the following search terms: ITPA or “Inosine Triphosphatase” combined with AZA or Azathioprine or Mercaptopurine or “6-MP” and “gene” or “polymorphism” or “mutation” or “SNP” combined with “side effects” or “adverse effects”. Thus, an eligible study had to include all kinds of patients that had received AZA or 6-MP, provide data on alleles or genotypes of ITPA polymorphisms and follow them during treatment to test if adverse effects occurred. The overall procedure of data extraction has been performed according to PRISMA guidelines [42] (Fig. 1). After an initial screening of titles and abstracts, only relevant articles were analyzed. Studies were examined carefully and only the ones that provided sufficient data to estimate Odds Ratio and its 95% confidence interval (95% CI) were included in the qualitative and quantitative analysis. The reference lists were scrutinized to consider unpublished studies (conference papers, dissertations) to avoid gray literature publication bias. No restrictions in the study selection procedure (study design, language or other quality measures) were imposed. Non-English manuscripts were also considered for review [43, 44].
Risk of bias assessment and data extraction
Since all included studies were non-randomized, risk of bias was assessed by two reviewers (EB and GB) using the seven domains of ROBINS-I criteria [45].
The following data were extracted from each study: PubMed ID, first author’s name, year of publication, racial descent of participants, country of populations studied, and the total number of the subjects (with or with no adverse effects). Furthermore, type of disease, type of adverse effect of AZA/6-MP and the distribution of genotypes and alleles for all participants were recorded.
Statistical analysis
In order to evaluate the distribution of alleles and genotypes between cases and controls, the specific odds ratios along with their 95% confidence intervals were calculated for each polymorphism and for each comparison applying the random-effects method of DerSimonian and Laird [46]. In case of a zero cell, a continuity correction was applied by adding 0.5 to all cells of the contingency table. The percentage of between-studies variability, due to heterogeneity, was evaluated by the inconsistency index I2 [47].
Initially, a meta-analysis was performed on a unique comprehensive group of patients that had at least one of the adverse effects, regardless of the background disease. It has to be mentioned that many studies did not specify the adverse effect and thus the term “all adverse effects” was used in the present study to include cumulatively all the mentioned and the non-mentioned adverse effects. When articles reported more than one adverse effect on the same population, the study with the highest number of cases was included in the “all adverse effects” meta-analysis so that each individual patient was counted only once in each meta-analysis. Two studies [10, 15] were considered to have duplicate data and thus only one was included [10]. Stratification analyses were also performed for adults and children, as well as for subpopulations according to race, where possible.
Three different contrasts were investigated corresponding to co-dominant, dominant and recessive modes of inheritance (A vs. C, AC + AA vs. CC and ΑΑ vs. ΑC + CC for rs1127354 and C vs. A, CC + ΑC vs. AA, and CC vs. AC + AA genotypes for rs7270101). However, because the numbers of risk homozygotes were in all cases either very low or zero, only results for the dominant mode of inheritance are presented herein. For all analyses, the statistical package STATA 13 was used. Results with p value < 0.05 were considered statistically significant.
The Begg and Mazumdar rank correlation test [48] and the fixed-effects regression method of Egger [49] were used to evaluate publication bias. Influential meta-analysis was performed, by checking the effect of removing an individual study and each time repeating the analysis on the overall significance of the estimate and on the heterogeneity. The standard cumulative meta-analysis approach and a more recently proposed regression-based method were used for the detection of the time-trend [50]. Multiple comparisons testing was performed according to methods proposed by Bonferroni [51], Holm [52], Sidak [53], and Holland [54].
Results
Characteristics of the studies included in the meta-analysis
A comprehensive, computerized search in the literature yielded a total of 155 studies that examined the association of rs1127354 and rs7270101 polymorphisms with the appearance of adverse effects of AZA/6-MP. For rs1127354, 30 studies (with 3582 patients) met the inclusion criteria and their data were included in the meta-analysis (Fig. 1). The characteristics of each study along with information on alleles and genotypes are shown in Table 1. In particular, three studies were on neutropenia as adverse effect [10, 14, 55], ten studies [14, 31, 32, 56,57,58,59,60,61,62] addressed leucopenia, myelosuppression was investigated in eight studies [1, 31, 60, 63,64,65,66,67], two studies were on hematotoxicity [68, 69], two studies on agranulocytosis [14, 31], twelve studies [10, 29, 31, 59, 60, 63,64,65,66, 68,69,70] provided data on hepatotoxicity, and nine studies [10, 56, 59, 60, 63,64,65,66, 70] reported pancreatitis. In addition, flu-like symptoms were examined in five studies [9, 10, 59, 64, 68], nausea & vomiting in three studies [10, 59, 64], rash in two studies [10, 64], skin reaction in two studies [59, 65], alopecia in two studies [31, 65] and other (non-mentioned) adverse effects were reported in four studies [1, 10, 64, 69] (Table 2). For rs7270101, twelve studies (with 1562 patients) were included (Fig. 1). Three of these studies [10, 14, 55] reported on neutropenia, five on leucopenia [14, 32, 56, 57, 60], four studies [10, 29, 60, 65] on hepatotoxicity, four studies [10, 56, 60, 65] on pancreatitis, four on myelosuppression [1, 60, 65, 67] and two studies examined other (non-mentioned) adverse effects [1, 10] (Table 3). AZA/6-MP was received by patients suffering from different background diseases. The above studies included information about three major background disease states for rs1127354. In particular, 2169 patients suffered from IBD, 974 patients from ALL and 370 patients had undergone transplantation. For rs7270101, IBD was found in five studies with 778 patients, ALL in five studies with 569 patients and two studies described adverse effects in 215 transplant recipients (Table 4).
Risk of bias for the included studies was assessed with the use of ROBINS-I tool. As shown in Supplementary Fig. 1, most studies had low to moderate risk of bias in the domains: selection, classification of intervention, missing data and selection of reported results. However, confounding bias was diverse because in many studies the intervention was discontinued or adjusted and the time of intervention was not completely specified. Bias in the measurement outcomes was not low because of the method of study recruitment; in all studies all patients had received intervention.
rs1127354 polymorphism of ITPA gene
In total, thirty studies (with 3582 patients) [1, 9, 10, 14, 29,30,31,32, 55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76] were included in our meta-analysis to test the association of rs1127354 polymorphism of ITPA with AZA/6-MP toxicity. Among them, seventeen studies were conducted in Caucasian populations, eleven in Asian, one in Egyptian and one study was carried out in a mixed population with Caucasian, Asian and African origin (Table 1).
Meta-analysis according to dominant mode of inheritance (AC + AA vs. CC) and using the random effects model showed association of rs1127354 with AZA/6-MP-caused all adverse effects in adults (OR: 2.12, 95%CI: 1.22–3.69) (Table 2). Heterogeneity was rather high (I2 = 71.10%) and neither evidence for publication bias (p value = 0.245) nor for time trend was observed (Table 2). Of note is the fact that in all tested populations, the numbers of risk homozygotes were zero or extremely low or genotypes’ data were given as sum of heterozygotes and risk homozygotes (Supplementary Table 1). Thus, meta-analysis using the recessive or co-dominant mode of inheritance would be carried out only by adding pseudo-counts to all cells of the 2 × 2 table and consequently would lead to meaningless biological interpretations, thus they were not performed.
Because of the racial differences in the frequency of rs1127354, we stratified our analysis according to the races reported in the included studies. Our results did not show any different outcome between Asians and Caucasians (Fig. 2a).
We, next, created groups of patients according to their adverse effects and meta-analyses were conducted separately for each adverse effect. This stratification analysis revealed rs1127354 to be associated with neutropenia (irrespective of the age of the patients) with OR: 2.39 (95%CI: 1.97–2.90) and particularly in children with OR: 2.43 (95%CI: 2.12–2.79) (Table 2 and Fig. 2b).
Meta-analyses investigating putative association of the rest of the separate adverse effects (leucopenia, myelosuppression, hepatotoxicity, pancreatitis, skin reaction, alopecia, flu-like, rash and other non-mentioned) and rs1127354 polymorphism showed no significant association according to the random effects model. There was no evidence for publication bias except for analysis of all adverse effects for pediatric patients, nor was time trend observed in any of the contrasts (Table 2).
After multiple comparisons testing with Bonferroni, Holm, Sidak, and Holland methods, rs1127354 was confirmed to be associated only with neutropenia (Supplementary Table 3).
rs7270101 polymorphism of ITPA gene
Meta-analysis was performed for the association of rs7270101 polymorphism of ITPA with AZA/6-MP toxicity. In total, twelve studies [1, 10, 14, 29, 32, 55,56,57, 60, 65, 67, 74] were found eligible and provided data for 1562 subjects (Table 1). Ten studies enrolled Caucasian populations, one study Egyptian population and one mixed population with most patients of Caucasian origin while only few were Asians or Africans (60:9:3). Initially, a meta-analysis was performed with patients presenting all adverse effects (regardless of the disease they were taking AZA/6-MP for); 598 patients presented adverse effects and 964 patients were without adverse effects.
Meta-analysis was performed according to dominant mode of inheritance (AC + CC vs AA) and no association was found with the random effects model (OR: 1.14, 95%CI: 0.67-1.93) (Table 3 and Fig. 3a). However, the association of all adverse effects was significant in pediatric patients with OR: 1.74 and 95%CI: 1.06-2.87 (Table 3). Once more, the nearly zero frequency of risk homozygotes in all tested populations (Supplementary Table 2) would have resulted to erroneous results in case of using the co-dominant or recessive models of inheritance.
To identify any putative effect of frequency differences between reported ethnic populations, we stratified our analysis according to races. We observed no different outcome in different races as shown in Fig. 3a.
Next, stratification of the meta-analysis by separate adverse effects revealed significant association of rs7270101 with neutropenia and leucopenia in the general population and in pediatric patients. ORs for neutropenia in the general population was 2.93 (95%CI: 2.36–3.63) and in children 3.00 (95%CI: 2.63–3.41). For leucopenia ORs were 2.82 (95%CI: 1.76–4.50) in general population and 2.92 (95%CI: 2.31–3.70) in children (Table 3 and Fig. 3b). For the rest of the adverse effects no associations were found (Table 3). No publication bias was observed for most analyses; exceptions are analysis of all adverse effects, and myelosuppression. No time trend was identified in any of the analyses, except for analysis of leucopenia (Table 3).
To adjust the level of significance of the above associations we performed multiple comparison tests with Bonferroni, Holm, Sidak and Holland multiple testing methods. Remarkably, rs7270101 polymorphisms was confirmed to be associated with neutropenia and leucopenia (Supplementary Table 3).
Association between rs1127354 and rs7270101 and adverse effects in a stratification analysis according to background disease
Since the disease, an individual may suffer from, can be partially attributed to a specific genetic background, we set out to investigate whether certain background diseases can affect the existence and the nature of AZA/6-MP toxic effects. Towards this direction, additional sub-group analyses were performed for each polymorphism stratifying for the background disease.
For rs1127354, we encountered three different background disease situations i.e., IBD, ALL and renal or liver transplant recipients. In patients with IBD, rs1127354 polymorphism was significantly associated with all adverse effects in adults (OR: 2.12, 95%CI: 1.06–4.24) (Table 4). Further stratification according to background disease and type of adverse effect revealed neutropenia, as an adverse effect, to be significantly associated with rs1127354 polymorphism in pediatric patients with ALL (OR: 2.43, 95%CI: 2.12–2.79). Heterogeneity was not detected (Table 4).
In our attempt to stratify data for rs7270101 polymorphism according to background disease entities, three subgroup analyses were formulated. No significant association was depicted in the groups of IBD patients and transplant recipients. Remarkably, all adverse effects and neutropenia, separately, were statistically significantly associated with rs7270101 in pediatric patients suffering from ALL (OR: 1.87, 95%CI: 1.13–3.08 and OR: 3.00, 95%CI: 2.64–3.41, respectively) (Table 4).
p-values of all tested associations were tested according to Bonferroni, Holm, Sidak, and Holland methods in order to prevent the inflation of false positive rates that could have occurred in the above multiple statistical tests. The association of both polymorphisms with neutropenia in ALL patients was confirmed (Supplementary Table 4).
Discussion
The present meta-analysis is an attempt to investigate the putative effect of ITPA polymorphisms in the development of adverse effects in AZA/6-MP treated patients. We found that the risk allele A of rs1127354 (94 C > A) is statistically significantly associated with increased risk for cumulatively all adverse effects in adults and separately with neutropenia. We also found that rs7270101 (IVS2 + 21 A > C) is associated with all adverse effects in children and with neutropenia and leucopenia development in all-ages and pediatric AZA/6-MP treated patients. The adjusted p-values, after multiple comparisons corrections, revealed significance for the associations with neutropenia and leucopenia. The evidence suggesting associations with all adverse effects in distinct populations was not confirmed denoting that more studies are needed to lead to robust results or that adverse effects should better be studied separately.
Stratification according to background disease revealed some interesting findings. IBD adult patients with rs1127354 polymorphism are more likely to develop all kinds of adverse effects. Importantly, pediatric ALL patients with either rs1127354 or rs7270101 polymorphism have almost two and half and three times, respectively, higher risk to develop neutropenia. The latter associations were confirmed after multiple comparisons tests. Though speculative, association with neutropenia in ALL patients may suggest either that ITPA deficiency can regulate the function of genes that predispose to neutropenia and leukemia [77, 78] or that the 6-thio-ITP, the metabolite that is the substrate of ITPA, exert inhibitory or mutagenic effects (directly or indirectly) on DNA replication or transcription in leukocytes [79]. Based on differences in the frequencies of gene variants of the other two thiopurine related pharmacogenes (TPMT and NUDT15) among individuals of different origin [28], we also stratified our analysis according to geographical ancestry; no association was depicted for any of the two ITPA polymorphisms in any specific population.
As we already discussed, functional and biochemical data in the literature show that ITPA forms a homodimer in order to exert its action. This finding is consistent with the notion that, in heterozygotes of rs1127354, only one out of four dimers can be functional. Heterozygotes of rs1127354 show, also, only 22% of the wt homozygotes mean activity, whereas homozygotes show zero ITPase activity [10, 34,35,36]. Taken together these facts imply clearly a dominant mode of inheritance, similar to what was seen in other dimers such the melanocortin-4 receptor [37], or the calcium-sensing receptor (CASR) [38, 80]. Importantly, in most of the included studies the risk homozygotes were zero or very few or no information was given. Thus, analysis for the co-dominant or recessive modes of inheritance, would need adding pseudo-counts and, would lead to biologically erroneous interpretations. Beyond that, testing additional null hypotheses would entail an increased risk of Type I error rate.
In the present meta-analysis, we investigated the association of two ITPA polymorphisms with the presence of adverse effects after AZA/6-MP administration. Compared to a previous meta-analysis [40] published in 2007, our study has certain advantages. First it provides an update of the global literature and robustly associates rs1127354 with adverse effects of AZA/6-MP, namely neutropenia. It contains data from 30 studies with 3712 patients as compared to 6 studies and 751 patients of the previous one that received AZA/6-MP treatment. Second, it presents, for the first time, a meta-analysis on rs7270101 showing association only with neutropenia and leucopenia. Third, we stratified our analysis according to the age of patients (children and adults) while the previous meta-analysis investigated only the rs1127354 polymorphism in all-ages populations. Fourth, in this study we investigated all types of adverse effects separately adding eight more adverse effect categories compared to the previous one [40] (twelve types of adverse effects as compared to four types of adverse effects). Fifth, the previous meta-analysis [40] was limited to IBD patients, while the present study includes all kinds of background disease and stratifies the analysis into IBD, ALL and transplant recipient patients. Sixth, because rs1127354 has different frequencies in various ethnic populations, showing the highest frequency in Asians (11-19%), the lowest in South Americans (1–2%) and intermediate in Caucasians (9%) and Africans (5%) [35, 81, 82], we explored the putative association of both polymorphisms with adverse effects in separate populations. Finally, we investigated methodological issues such as heterogeneity, publication bias, time-trend, sensitivity analysis, multiple comparison testing and assessment of the risk of bias.
We also acknowledge some important issues and limitations that could have affected the credibility of our results. The existence of gray literature cannot be precisely identified; thus publication bias could not be ruled out. The studies investigated in the present meta-analysis encompassed a diverse range of drug intervention and time settings. In addition, confounding factors can potentially influence the quality parameters assessed. In addition, tests for the severity of toxicity in a drug dose dependent manner could not be performed since neither data for toxicity classification was given nor comparable and stable (within studies and within individuals) AZA/6-MP dosage schemes were used. Moreover, in four cases methotrexate was co-administered. Thus, heterogeneity in study designs and diversity in the intervention times could have affected our results. However, these factors were considered in risk of bias assessments. Nevertheless, we performed the meta-analysis carefully taking all these considerations into account. More detailed analyses, for instance to account for co-administered drugs, various dosage schemes, different follow-up times, or the co-existence of the adverse effects, would need access to individual data, and subsequently an individual patients’ data (IPD) meta-analysis.
In the last few years, developments in molecular techniques have allowed genes of pharmacological importance to emerge as biomarkers for predicting adverse effects of drug administration. It is increasingly acknowledged that clinical effectiveness emerges from the knowledge of association of gene variants and variable drug responses in individual patients [83, 84]. TPMT was the first gene characterized as pharmacogene for thiopurine treated individuals since some of its polymorphisms (pharmacovariants) were investigated and proved to be associated with more adverse effects in the variant carriers [84, 85]. In 2018, the Clinical Pharmacogenetics Implementation Consortium updated their guidelines for thiopurine dosing by advising the pre-emptive genotyping of TPMT and NUDT15 genes [25]. Optimization of treatment approaches can substantially reduce the risk of adverse drug reactions without compromising therapeutic outcomes [28]. Pre-emptive genotype testing of pharmacogenes is increasingly adopted in many national healthcare systems since cost-utility analyses showed that it can reduce the systems’ expenses for unnecessary or harmful treatments [86]. However, it is similarly acknowledged that other, rarer or not well studied pharmacovariants and/or pharmacogenes are in play to dictate thiopurine adverse responses in individuals that are genotyped for TPMT and NUDT15 common variants and still present severe adverse effects [25, 28]. ITPA has been investigated (although less, compared to TPMT and NUDT15) for its utilization as a pharmacogene for thiopurine-based treatments, with conflicting results [85]. Additional, pre-emptive genotyping, before AZA/6-MP administration, of other than TPMT and NUDT15 genes is already a well-educated perspective that awaits documentation [25, 28], and rs1127354 and rs7270101 of ITPA have the potential to be used as predictive biomarkers. In the present meta-analysis, we quantitatively summarized the up-to-date global data of the literature on ITPA related adverse effects with the hope that our results can contribute to new treatment algorithms, protocols and dosage individualization schedules, to eliminate adverse effects and improve individual patients’ drug response. Future, large-scale studies, testing association of TPMT-NUDT15-ITPA combined genotypes and drug adverse effects and effectiveness will help assess the significance of adding ITPA testing to the list of preemptive tests before thiopurine treatment.
References
Breen DP, Marinaki AM, Arenas M, Hayes PC. Pharmacogenetic association with adverse drug reactions to azathioprine immunosuppressive therapy following liver transplantation. Liver Transpl. 2005;11:826–33.
Pui CH, Evans WE. Treatment of acute lymphoblastic leukemia. N Engl J Med. 2006;354:166–78.
Pui CH, Robison LL, Look AT. Acute lymphoblastic leukaemia. Lancet. 2008;371:1030–43.
Teml A, Schaeffeler E, Herrlinger KR, Klotz U, Schwab M. Thiopurine treatment in inflammatory bowel disease: clinical pharmacology and implication of pharmacogenetically guided dosing. Clin Pharmacokinetics. 2007;46:187–208.
Connell WR, Kamm MA, Ritchie JK, Lennard-Jones JE. Bone marrow toxicity caused by azathioprine in inflammatory bowel disease: 27 years of experience. Gut. 1993;34:1081–5.
Floyd A, Pedersen L, Nielsen GL, Thorlacius-Ussing O, Sorensen HT. Risk of acute pancreatitis in users of azathioprine: a population-based case-control study. Am J Gastroenterol. 2003;98:1305–8.
Moon W, Loftus EV Jr. Review article: recent advances in pharmacogenetics and pharmacokinetics for safe and effective thiopurine therapy in inflammatory bowel disease. Alimentary Pharmacol Therapeut. 2016;43:863–83.
Schwab M, Schaffeler E, Marx C, Fischer C, Lang T, Behrens C, et al. Azathioprine therapy and adverse drug reactions in patients with inflammatory bowel disease: impact of thiopurine S-methyltransferase polymorphism. Pharmacogenetics. 2002;12:429–36.
Ansari A, Arenas M, Greenfield SM, Morris D, Lindsay J, Gilshenan K, et al. Prospective evaluation of the pharmacogenetics of azathioprine in the treatment of inflammatory bowel disease. Alimentary Pharmacol Therapeut. 2008;28:973–83.
Marinaki AM, Duley JA, Arenas M, Ansari A, Sumi S, Lewis CM, et al. Mutation in the ITPA gene predicts intolerance to azathioprine. Nucleosides Nucleotides Nucleic Acids. 2004;23:1393–7.
Chaparro M, Ordas I, Cabre E, Garcia-Sanchez V, Bastida G, Penalva M, et al. Safety of thiopurine therapy in inflammatory bowel disease: long-term follow-up study of 3931 patients. Inflamm Bowel Dis. 2013;19:1404–10.
Watanabe A, Hobara N, Nagashima H. Demonstration of enzymatic activity converting azathioprine to 6-mercaptopurine. Acta Med Okayama. 1978;32:173–9.
Siegel CA, Sands BE. Review article: practical management of inflammatory bowel disease patients taking immunomodulators. Alimentary Pharmacol Therapeut. 2005;22:1–16.
Hareedy MS, El Desoky ES, Woillard JB, Thabet RH, Ali AM, Marquet P, et al. Genetic variants in 6-mercaptopurine pathway as potential factors of hematological toxicity in acute lymphoblastic leukemia patients. Pharmacogenomics. 2015;16:1119–34.
Marinaki AM, Ansari A, Duley JA, Arenas M, Sumi S, Lewis CM, et al. Adverse drug reactions to azathioprine therapy are associated with polymorphism in the gene encoding inosine triphosphate pyrophosphatase (ITPase). Pharmacogenetics. 2004;14:181–7.
Tiede I, Fritz G, Strand S, Poppe D, Dvorsky R, Strand D, et al. CD28-dependent Rac1 activation is the molecular target of azathioprine in primary human CD4+ T lymphocytes. J Clin Investig. 2003;111:1133–45.
Fairchild CR, Maybaum J, Kennedy KA. Concurrent unilateral chromatid damage and DNA strand breakage in response to 6-thioguanine treatment. Biochemical Pharmacol. 1986;35:3533–41.
Lennard L. TPMT in the treatment of Crohn’s disease with azathioprine. Gut. 2002;51:143–6.
Aberra FN, Lichtenstein GR. Review article: monitoring of immunomodulators in inflammatory bowel disease. Alimentary Pharmacol Therapeut. 2005;21:307–19.
Derijks LJ, Gilissen LP, Engels LG, Bos LP, Bus PJ, Lohman JJ, et al. Pharmacokinetics of 6-thioguanine in patients with inflammatory bowel disease. Therapeutic Drug Monit. 2006;28:45–50.
Stocco G, Cheok MH, Crews KR, Dervieux T, French D, Pei D, et al. Genetic polymorphism of inosine triphosphate pyrophosphatase is a determinant of mercaptopurine metabolism and toxicity during treatment for acute lymphoblastic leukemia. Clin Pharmacol Therapeut. 2009;85:164–72.
Yin D, Xia X, Zhang J, Zhang S, Liao F, Zhang G, et al. Impact of NUDT15 polymorphisms on thiopurines-induced myelotoxicity and thiopurines tolerance dose. Oncotarget. 2017;8:13575–85.
Zhang AL, Yang J, Wang H, Lu JL, Tang S, Zhang XJ. Association of NUDT15 c.415C>T allele and thiopurine-induced leukocytopenia in Asians: a systematic review and meta-analysis. Ir J Med Sci. 2018;187:145–53.
Nielsen OH, Bjerrum JT, Herfarth H, Rogler G. Recent advances using immunomodulators for inflammatory bowel disease. J Clin Pharmacol. 2013;53:575–88.
Relling MV, Schwab M, Whirl-Carrillo M, Suarez-Kurtz G, Pui CH, Stein CM, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) guideline for thiopurine dosing based on TPMT and NUDT15 genotypes: 2018 update. Clin Pharmacol Therapeutic. 2019;105:5.
Gisbert JP, Luna M, Mate J, Gonzalez-Guijarro L, Cara C, Pajares JM. Choice of azathioprine or 6-mercaptopurine dose based on thiopurine methyltransferase (TPMT) activity to avoid myelosuppression. A prospective study. Hepato-Gastroenterol. 2006;53:399–404.
Colombel JF, Ferrari N, Debuysere H, Marteau P, Gendre JP, Bonaz B, et al. Genotypic analysis of thiopurine S-methyltransferase in patients with Crohn’s disease and severe myelosuppression during azathioprine therapy. Gastroenterology. 2000;118:1025–30.
Koutsilieri S, Caudle KE, Alzghari SK, Monte AA, Relling MV, Patrinos GP. Optimizing thiopurine dosing based on TPMT and NUDT15 genotypes: It takes two to tango. Am J Hematol. 2019;94:737–40. Wiley-Liss Inc
Azimi F, Mortazavi Y, Alavi S, Khalili M, Ramazani A. Frequency of ITPA gene polymorphisms in Iranian patients with acute lymphoblastic leukemia and prediction of its myelosuppressive effects. Leuk Res. 2015;39:1048–54.
Ma X, Zheng J, Jin M, Li W, Gao C, Zhang D, et al. Inosine triphosphate pyrophosphohydrolase (ITPA) polymorphic sequence variants in Chinese ALL children and possible association with mercaptopurine related toxicity. Int J Clin Exp Pathol. 2014;7:4552–6.
Uchiyama K, Nakamura M, Kubota T, Yamane T, Fujise K, Tajiri H. Thiopurine S-methyltransferase and inosine triphosphate pyrophosphohydrolase genes in Japanese patients with inflammatory bowel disease in whom adverse drug reactions were induced by azathioprine/6-mercaptopurine treatment. J Gastroenterol. 2009;44:197–203.
Zelinkova Z, Derijks LJ, Stokkers PC, Vogels EW, van Kampen AH, Curvers WL, et al. Inosine triphosphate pyrophosphatase and thiopurine s-methyltransferase genotypes relationship to azathioprine-induced myelosuppression. Clin Gastroenterol Hepatol. 2006;4:44–9.
Stenmark P, Kursula P, Flodin S, Gräslund S, Landry R, Nordlund P, et al. Crystal structure of human inosine triphosphatase. Substrate binding and implication of the inosine triphosphatase deficiency mutation P32T. J Biol Chem. 2007;282:3182–7.
Lin S, McLennan AG, Ying K, Wang Z, Gu S, Jin H, et al. Cloning, expression, and characterization of a human inosine triphosphate pyrophosphatase encoded by the itpa gene. J Biol Chem. 2001;276:18695–701.
Shipkova M, Lorenz K, Oellerich M, Wieland E, von Ahsen N. Measurement of Erythrocyte Inosine Triphosphate Pyrophosphohydrolase (ITPA) Activity by HPLC and Correlation of ITPA Genotype-Phenotype in a Caucasian Population. Clin Chem. 2006;52:240–47.
Sumi S, Marinaki AM, Arenas M, Fairbanks L, Shobowale-Bakre M, Rees DC, et al. Genetic basis of inosine triphosphate pyrophosphohydrolase deficiency. Hum Genet. 2002;111:360–7.
Biebermann H, Krude H, Elsner A, Chubanov V, Gudermann T, Grüters A. Autosomal-dominant mode of inheritance of a melanocortin-4 receptor mutation in a patient with severe early-onset obesity is due to a dominant-negative effect caused by receptor dimerization. Diabetes. 2003;52:2984–88.
Hendy GN, Geoffrey N, Vito Guarnieri, Lucie Canaff. Chapter 3 calcium-sensing receptor and associated diseases. Prog Mol Biol Transl Sci. 2009;89:31–95.
Kouwenberg TW, van den Bosch BJC, Bierau J, Te Loo DMWM, Coenen MJH, Hagleitner MM. Dosage of 6-mercaptopurine in relation to genetic TPMT and ITPA variants: toward individualized pediatric acute lymphoblastic leukemia maintenance treatment. J Pediatr Hematol/Oncol. 2020;42:E94–97.
Van Dieren JM, Hansen BE, Kuipers EJ, Nieuwenhuis EE, Van der Woude CJ. Meta-analysis: Inosine triphosphate pyrophosphatase polymorphisms and thiopurine toxicity in the treatment of inflammatory bowel disease. Alimentary Pharmacol Therapeut. 2007;26:643–52.
Counsell C. Formulating questions and locating primary studies for inclusion in systematic reviews. Ann Intern Med. 1997;127:380–7. American College of Physicians
Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA Statement. PLoS Med 2009;6:e1000097.
Hopewell S, McDonald S, Clarke M, Egger M. Grey literature in meta-analyses of randomized trials of health care interventions. Cochrane Database Syst Rev. 2007;18:MR000010.
Stroup DF, Berlin JA, Morton SC, Olkin I, Williamson GD, Rennie D, et al. Meta-analysis of observational studies in epidemiology: a proposal for reporting. Meta-analysis Of Observational Studies in Epidemiology (MOOSE) group. JAMA. 2000;283:2008–12.
Sterne JA, Hernán MA, Reeves BC, Savović J, Berkman ND, Viswanathan M, et al. ROBINS-I: A tool for assessing risk of bias in non-randomised studies of interventions. BMJ. 2016;355:i4919. https://doi.org/10.1136/bmj.i4919.
DerSimonian R, Laird N. Meta-analysis in clinical trials. Controlled Clin Trials. 1986;7:177–88.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–60.
Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994;50:1088–101.
Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315:629–34.
Bagos PG, Nikolopoulos GK. Generalized least squares for assessing trends in cumulative meta-analysis with applications in genetic epidemiology. J Clin Epidemiol. 2009;62:1037–44.
Aickin M, Gensler H. Adjusting for multiple testing when reporting research results: the Bonferroni vs Holm methods. Am J Public Health. 1996;86:726–8.
Holm S. A simple sequentially rejective multiple test procedure. Scand J Stat. 1979;6:65–70.
Šidák Z. Rectangular confidence regions for the means of multivariate normal distributions. J Am Stat Assoc. 1967;62:626–33.
Keselman HJ, Cribbie R, Holland B. Controlling the rate of Type I error over a large set of statistical tests. Br J Math Stat Psychol. 2002;55:27–39.
Wahlund M, Nilsson A, Kahlin AZ, Broliden K, Myrberg IH, Appell ML, et al. The role of TPMT, ITPA, and NUDT15 variants during mercaptopurine treatment of swedish pediatric patients with acute lymphoblastic leukemia. J Pediatr. 2020;216:150–.e1.
De Ridder L, Van Dieren JM, Van Deventer HJ, Stokkers PC, Van der Woude JC, Van, et al. Pharmacogenetics of thiopurine therapy in paediatric IBD patients. Alimentary Pharmacol Therapeut. 2006;23:1137–41.
Kurzawski M, Dziewanowski K, Lener A, Drozdzik M. TPMT but not ITPA gene polymorphism influences the risk of azathioprine intolerance in renal transplant recipients. Eur J Clin Pharmacol. 2009;65:533–40.
Odahara S, Uchiyama K, Kubota T, Ito Z, Takami S, Kobayashi H, et al. A prospective study evaluating metabolic capacity of thiopurine and associated adverse reactions in Japanese patients with inflammatory bowel disease (IBD). PloS One. 2015;10:e0137798.
Palmieri O, Latiano A, Bossa F, Vecchi M, D’Inca R, Guagnozzi D, et al. Sequential evaluation of thiopurine methyltransferase, inosine triphosphate pyrophosphatase, and HPRT1 genes polymorphisms to explain thiopurines’ toxicity and efficacy. Alimentary Pharmacol Therapeut. 2007;26:737–45.
Steponaitiene R, Kupcinskas J, Survilaite S, Varkalaite G, Jonaitis L, Kiudelis G, et al. TPMT and ITPA genetic variants in Lithuanian inflammatory bowel disease patients: prevalence and azathioprine-related side effects. Adv Med Sci. 2016;61:135–40.
Kim JH, Cheon JH, Hong SS, Eun CS, Byeon JS, Hong SY, et al. Influences of thiopurine methyltransferase genotype and activity on thiopurine-induced leukopenia in Korean patients with inflammatory bowel disease: a retrospective cohort study. J Clin Gastroenterol. 2010;44:e242–8.
Tanaka Y, Nakadate H, Kondoh K, Nakamura K, Koh K, Manabe A. Interaction between NUDT15 and ABCC4 variants enhances intolerability of 6-mercaptopurine in Japanese patients with childhood acute lymphoblastic leukemia. Pharmacogenomics J. 2018;18:275–80.
van Dieren JM, van Vuuren AJ, Kusters JG, Nieuwenhuis EE, Kuipers EJ, van der Woude CJ. ITPA genotyping is not predictive for the development of side effects in AZA treated inflammatory bowel disease patients. Gut. 2005;54:1664.
Gearry RB, Roberts RL, Barclay ML, Kennedy MA. Lack of association between the ITPA 94C>A polymorphism and adverse effects from azathioprine. Pharmacogenetics. 2004;14:779–81.
Zabala-Fernandez W, Barreiro-de Acosta M, Echarri A, Carpio D, Lorenzo A, Castro J, et al. A pharmacogenetics study of TPMT and ITPA genes detects a relationship with side effects and clinical response in patients with inflammatory bowel disease receiving Azathioprine. J Gastrointest Liver Dis. 2011;20:247–53.
Hindorf U, Lindqvist M, Peterson C, Soderkvist P, Strom M, Hjortswang H, et al. Pharmacogenetics during standardised initiation of thiopurine treatment in inflammatory bowel disease. Gut. 2006;55:1423–31.
Hawwa AF, Millership JS, Collier PS, Vandenbroeck K, McCarthy A, Dempsey S, et al. Pharmacogenomic studies of the anticancer and immunosuppressive thiopurines mercaptopurine and azathioprine. Br J Clin Pharmacol. 2008;66:517–28.
Xiong H, Xin HW, Wu XC, Li Q, Xiong L, Yu AR. Association between inosine triphosphate pyrophosphohydrolase deficiency and azathioprine-related adverse drug reactions in the Chinese kidney transplant recipients. Fundamental Clin Pharmacol. 2010;24:393–400.
Tanaka Y, Manabe A, Fukushima H, Suzuki R, Nakadate H, Kondoh K, et al. Multidrug resistance protein 4 (MRP4) polymorphisms impact the 6-mercaptopurine dose tolerance during maintenance therapy in Japanese childhood acute lymphoblastic leukemia. Pharmacogenomics J. 2015;15:380–4.
von Ahsen N, Armstrong VW, Behrens C, von Tirpitz C, Stallmach A, Herfarth H, et al. Association of inosine triphosphatase 94C>A and thiopurine S-methyltransferase deficiency with adverse events and study drop-outs under azathioprine therapy in a prospective Crohn disease study. Clin Chem. 2005;51:2282–8.
Al-Judaibi B, Schwarz UI, Huda N, Dresser GK, Gregor JC, Ponich T, et al. Genetic predictors of azathioprine toxicity and clinical response in patients with inflammatory bowel disease. J Popul Therapeutics Clin Pharmacol. 2016;23:e26–36.
Suzuki R, Fukushima H, Noguchi E, Tsuchida M, Kiyokawa N, Koike K, et al. Influence of SLCO1B1 polymorphism on maintenance therapy for childhood leukemia. Pediatr Int. 2015;57:572–7.
Kim H, Kang HJ, Kim HJ, Jang MK, Kim NH, Oh Y, et al. Pharmacogenetic analysis of pediatric patients with acute lymphoblastic leukemia: a possible association between survival rate and ITPA polymorphism. PLoS One. 2012;7:e45558.
Smid A, Karas-Kuzelicki N, Milek M, Jazbec J, Mlinaric-Rascan I. Association of ITPA genotype with event-free survival and relapse rates in children with acute lymphoblastic leukemia undergoing maintenance therapy. PLoS One. 2014;9:e109551.
Tsuchiya A, Aomori T, Sakamoto M, Takeuchi A, Suzuki S, Jibiki A, et al. Effect of genetic polymorphisms of azathioprine-metabolizing enzymes on response to rheumatoid arthritis treatment. Pharmazie. 2017;72:22–28.
Kishibe M, Nozaki H, Fujii M, Iinuma S, Ohtsubo S, Igawa S, et al. Severe thiopurine-induced leukocytopenia and hair loss in Japanese patients with defective NUDT15 variant: Retrospective case-control study. J Dermatol. 2018;45:1160–5.
Bierau J, Lindhout M, Bakker JA. Pharmacogenetic significance of inosine triphosphatase. Pharmacogenomics. 2007;8:1221–1228.
Walkovich K, Connelly JA. Congenital neutropenia and rare functional phagocyte disorders in children. Hematol/Oncol Clin North Am. 2019;33:533–51. W.B. Saunders
Marinaki AM, Arenas-Hernandez M. Reducing risk in thiopurine therapy. Xenobiotica. 2020;50:101–9. Taylor and Francis Ltd
Pollak MR, Brown EM, Estep HL, McLaine PN, Kifor O, Park J, et al. Autosomal dominant hypocalcaemia caused by a Ca(2+)-sensing receptor gene mutation. Nat Genet. 1994;8:303–7.
Cao H, Hegele RA. DNA polymorphisms in ITPA including basis of inosine triphosphatase deficiency. J Hum Genet. 2002;47:620–22.
Marsh S, King CR, Ahluwalia R, McLeod HL. Distribution of ITPA P32T Alleles in Multiple World Populations. J Hum Genet. 2004;49:579–81.
Lakiotaki K, Kanterakis A, Kartsaki E, Katsila T, Patrinos GP, Potamias G. Exploring public genomics data for population pharmacogenomics. PLoS One. 2017;8:12.
Katara P, Yadav A Pharmacogenes (PGx-genes): Current understanding and future directions. Gene. Nov, 2019;718. Elsevier B.V.
Al-Mahayri ZN, Patrinos GP, Ali BR. Pharmacogenomics in pediatric acute lymphoblastic leukemia: Promises and limitations. Pharmacogenomics. 2017;18:687–99. Future Medicine Ltd
Simeonidis S, Koutsilieri S, Vozikis A, Cooper DN, Mitropoulou C, Patrinos GP. Application of economic evaluation to assess feasibility for reimbursement of genomic testing as part of personalized medicine interventions. Front Pharmacol. 2019;10:830.
Funding
PIK and PGB acknowledge support of this work by the project “ELIXIR-GR: The Greek Research Infrastructure for Data Management and Analysis in Life Sciences”, Grant Number (MIS) 5002780.
Author information
Authors and Affiliations
Contributions
GGB participated in the conception of the study, in data collection, in the analysis and in the interpretation of the results. PGB participated in the conception of the study, in data collection, in the analysis and in the interpretation of the results. EB did most of data collection, performed most of the analysis, participated in the interpretation of the results, and drafted the initial version of the manuscript. PIK participated in the analysis and in interpretation of the results. IM participated in the interpretation of the results. All authors participated in drafting the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Barba, E., Kontou, P.I., Michalopoulos, I. et al. Association of ITPA gene polymorphisms with adverse effects of AZA/6-MP administration: a systematic review and meta-analysis. Pharmacogenomics J 22, 39–54 (2022). https://doi.org/10.1038/s41397-021-00255-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-021-00255-3
- Springer Nature Limited