Abstract
Alternative splicing represents a critical posttranscriptional regulation of gene expression, which contributes to the protein complexity and mRNA processing. Defects of alternative splicing including genetic alteration and/or altered expression of both pre-mRNA and trans-acting factors give rise to many cancers. By integrally analyzing clinical data and splicing data from TCGA and SpliceSeq databases, a number of splicing events were found clinically relevant in tumor samples. Alternative splicing of KLK2 (KLK2_51239) was found as a potential inducement of nonsense-mediated mRNA decay and associated with poor survival in prostate cancer. Consensus K-means clustering analysis indicated that alternative splicing events could be potentially used for molecular subtype classification of cancers. By random forest survival algorithm, prognostic prediction signatures with well performances were constructed for 31 cancers by using survival-associated alternative splicing events. Furthermore, an online tool for visualization of Kaplan–Meier plots of splicing events in 31 cancers was explored. Briefly, alternative splicing was found of significant clinical relevance with cancers.
Similar content being viewed by others
Introduction
Alternative splicing, compared with constitutive pre-mRNA splicing, refers to the process of producing different mRNA isoforms, which has been found widespread in metazoan, including human, plants, animals, and fungi, and plays important roles in controlling cellular differentiation and organismal development [1, 2]. Alternative splicing is also a critical method of posttranscriptional regulation, which may transform products of a coding gene to protein isoforms deactivated or delocalized by varying exons composition and mRNA isoforms with no translations or degraded by nonsense-mediated decay (NMD) [3,4,5,6]. It has been reported that in humans most multi-exon genes are alternatively spliced, and only 60% of the spliced products will encode protein isoforms [7, 8]. Alternative splicing involves seven basic splicing patterns [8], including alternate acceptor sites (AA), alternate donor sites (AD), alternate promoter (AP), alternate terminator (AT), exon skipping (ES), mutually exclusive exons (ME), and retained intron (RI) (Supplementary Fig. S1B). Although the mechanisms of AP have been reported to be different from those of typical alternative splicing, they both involve the alternative use of variable exons and production of different protein isoforms with distinct functional activities [8, 9]. Since the processed alternative splicing data include the AP splicing method, we have retained these seven splicing modes throughout the article.
The different protein isoforms produced by alternative splicing may have varied or even opposite biological functions, which greatly generating abundance and complexity of the organism’s protein–protein interactions [3]. However, splicing abnormalities producing irregular protein isoforms may also lead to many diseases, including cancers [10]. Many researches indicated that outlier alternative splicing of cancer-related genes involve in regulation of various biological processes, prediction of cancer prognoses, and treatment responses. For instance, alternative splicing of exon 2 (AD) of BCL2 causes the upregulation of antiapoptotic BCL-XL in tumors, leading to apoptosis tolerance of tumor cells [11]. Compared with prostate cancer patients that characterized as a negative androgen receptor variant-7 (AR-V7), in our previous study, the AR-V7-positive patients suffered significantly lower PSA response rates to androgen-deprivation therapy and worse survival for castration-resistant prostate cancer (CRPC) [12].
In recent years, many anticancer target drugs have been developed based on genetic alterations including gene mutations, and altered genes expression, drugs targeting PD-L1 is a powerful example. Determining the molecular mechanisms and regulatory networks of alternative splicing is necessary for designing individual treatments targeting cancer-related splicing events. With the flourishing of algorithm explored for detecting alternative splicing, currently, several comprehensive analyses were performed in cancers focused on the landscape of alternative splicing in genetic alterations, tissue specificity and splicing regulatory networks. For example, a pan-cancer analysis in 32 human cancers by Kahles et al. showed that alternative splicing events increased in TCGA tumors compared with GTEx normal tissues and associated with identified variations of trans-acting factors [13]. Another team analyzed the splicing quantitative trait loci (sQTLs) and explored a database that provides informative resource for potential relationships of SNPs and irregular spliced transcript isoforms in human cancers [14]. Jayasinghe et al. discovered 1964 mutations that induce evidently splice site creation and produce alternative splice junctions among 32 TCGA cancers by the developed tool MiSplice [15]. Prognostic values of alternative splicing events were also analyzed in some individual cancers, including non-small cell lung cancer [16]. Integrated pan-cancer analysis about clinical relevance of alternative splicing events, however, has not been reported yet. To comprehensively and comparably explore clinical relevant splicing events that may function in the tumorigenesis, progression, molecular classification, and prognostic prediction of different cancer types, RNA-seq (level 3), clinical information, and splicing data of 31 cancers were integrated and analyzed in this study. Besides, a website (www.oncosplicing.com) has been explored for Kaplan–Meier plotting of splicing events in 31 human cancers.
Results
Profiles of alternative splicing events in 31 human cancers
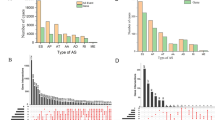
Integrated analysis of mRNA splicing profiles in 31 human cancers were performed in patients of TCGA cohort (Table 1). For different cancer types, the numbers of splicing events ranged from 28,281 of UCEC to 50,342 of ESCA. The most common splice type ES and the least common type ME account for 34.5–41.4% and 0.3–0.6% of total events, separately. The percentages of splice types AP, AT, AA, AD, and RI account for the total events in different cancers ranged by 15.8–20.8%, 16.8–27.6%, 7.6–8.4%, 6.3–7.3%, and 6.0–7.2%, respectively (Fig. 1a). We have observed that proportions of different splice types varied much more peaceably than that of the total splicing events between these different cancers. The total sizes of splicing events and genes across these 31 cancers were 64,174 and 12,692, separately, and the intersection size was 21 908 splicing events within 6693 genes. For most splicing events crossed in two or more cancer types, only a few events were found cancer specific. LGG have 1166, but both of COAD and BLCA have zero exclusive events compared with other cancers (Fig. 1c). Furthermore, we found that the ratio of average events and average genes of 31 cancers fluctuated significantly among splice patterns ranging from 1.02 of ME to 2.54 of ES (Fig. 1b; Supplementary Fig. S2A).
Profiles of alternative splicing events in 31 cancers. a The numbers and proportions of splicing events in different splice types. b Comparison of average splicing events and allied genes of 31 cancers and their ratios between different splice types. c The numbers of cancer-specific events in different cancers. There are 21,908 (central circle) events shared commonly between 31 cancers, but only a few events were cancer specific (inner circles) for each individual cancer. The numbers in the medial circles represent that the total events of each cancer, except commonly shared events
There were 12,692 genes (a half of the known ~20,530 coding genes) detected with splicing events spreading in these 31 cancers. The numbers of gene allied events were 5.05 in average and 685 in maximum, and most genes were detected with more than two splicing events (Supplementary Fig. S2B). Upset graph and heatmap were used to display the distribution of spliced genes in different splice types (Supplementary Fig. S2C, D). Though many genes were spliced in multi different types, there were still 4636 genes that spliced singly by one of the seven pattern, the most 2175 of which were spliced in ES. Events number of genes spliced in ES were significantly larger than that in other splice types.
Different alternative splicing events in 12 cancers
Different alternative splicing events (DASEs) were defined as median PSI value varied more than 0.1 between tumor tissues and adjacent normal tissues and corrected p-value < 0.05, which was assessed by Wilcoxon test and corrected for multiple testing by Benjamini–Hochberg method. These cancers with paired samples more than 20 were contained in this analysis, including BLCA (19 paired samples), BRCA, COAD, HNSC, KICH, KIRC, KIRP, LIHC, LUAD, LUSC, STAD, and UCEC (Table 1). There were an average of 1924 DASEs in these 12 cancers ranged from 1036 of LIHC to 3471 of LUSC (Fig. 2a).
Different alternative splicing events (DASEs) in the 12 cancers. dasTN represents median PSI values differed in tumor tissues and adjacent normal tissues. a The numbers of DASEs and allied genes in different 12 cancers are shown by bar plots. b,c Molecular complex detection between genes related to DASEs of LIHC. d Twenty-eight DASEs of cancer-related genes in more than five cancer types are shown in the bubble plot
To describe the different potential mechanisms of tumorigenesis of these cancers, genes related to cancer-specific DASEs were submitted to DAVID website (https://david.ncifcrf.gov/) for enrichment analyses, separately [17]. The comparison of top five KEGG pathways enriched in each cancer showed that there were several communal pathway terms in many cancer types (Supplementary Fig. S3B). For example, adherens junction and focal adhesion pathways were found enriched in more than six cancers. In STAD, genes were characteristically enriched to the cGMP-PKG signaling pathway; in UCEC genes were characteristically enriched to the MAPK signaling pathway; and in KIRC genes were characteristically enriched to the PPAR signaling pathway. Similarly, these genes of each cancer were variably enriched to different GO pathway terms (Supplementary Fig. S3A). Furthermore, a clustering algorithm named molecular complex detection (MCODE) was used to detect densely connected regions correspond to known molecular complexes in the large protein–protein interaction networks [18]. Genes related to DASEs were submitted to Metascape (a website for gene annotation & analysis, http://metascape.org) for MCODE analysis. The results showed that genes of the largest sets MCODE1 and MCODE2 of LIHC were mostly enriched in cell cycle and mRNA splicing pathways (Fig. 2b, c and Table 2).
DASEs of 574 cancer-related genes (CRGs) were also analyzed in the 12 cancers. A total of 398 DASEs within 184 CRGs were involved in one or more cancer types (Supplementary Table S1). These DASEs that spliced in more than five cancers were shown using bubble plots (Fig. 2d). When compared with adjacent normal tissues in most cancers, the trends were consistent for most upregulated or downregulated DASEs in tumor tissues, except for the splicing of TPM4, PBRM1, NCOR2, and CASP8 in KIRC.
Survival-associated alternative splicing events in 31 cancers
Survival-associated alternative splicing events (SASEs) were derived from univariate cox regression analysis with p-value < 0.05 (wald χ2 test). There are average 1580 SASEs of these 31 cancers ranged from 440 of DLBC to 7391 of KIRC, including the second most 5363 of LGG (Supplementary Fig. S4A, B). Fewer SASEs were found overlapping with DASEs for each type of the 12 cancers (Supplementary Fig. S4C).
There were more than one splicing event for most splicing-related genes (Supplementary Fig. S2B). To study the varied functions of allied splicing events, genes related to SASEs with good or poor survival were analyzed, respectively, in each cancer type. There were 4417 genes found with survival-bipolar events in different cancers, and totally 11,384 times in 31 cancers (Fig. 3a). Importantly, these survival-bipolar genes are mostly associated with splicing events involved in pattern AP or AT (Fig. 3d). These results validating that irregular alternative splicing of a coding gene would deactivate the product function and even reverse it in some conditions. Furthermore, there were 506 paired survival-bipolar events of 189 validated CRGs found in 31 cancers and 57 of them with p-value < 0.001 in 13 cancers were integrally displayed (Fig. 3b; Supplementary Table S2). Kallikrein-related peptidase 2 (KLK2), for example, was considered as an important biomarker of prostate cancer and associated with bone metastasis of CRPC [19, 20]. KLK2 allied SASEs KLK2_51234 (AT) and KLK2_51239 (ES) were found survival opposite in PRAD (Fig. 3c). Speckle-type BTB/POZ protein (SPOP) that promotes tumorigenesis of renal cancer is a potential target for tumor therapy [21]. SPOP allied events SPOP_42302 (AP) and SPOP_42303 (AP) were found as two survival-bipolar events in KIRC (Fig. 3c). It is also notable that survival-bipolar events of few genes may also appear oppositely in different cancers, such as TFEB_76123 (AT) and TFEB_76124 (AT) in LAML and PRAD.
Survival-associated alternative splicing events (SASEs) in 31 cancers. a The numbers of genes with at least two survival-bipolar SASEs and genes with only survival good or poor SASEs of each cancer type are shown by bar plots. b Sixty-seven SASEs of 58 survival-bipolar cancer-related genes in 13 cancers with P (p-value) < 0.001 were integrally displayed. P-values and HRs were derived from univariate cox regression analysis. c SPOP in KIRC and KLK2 in PRAD were listed as two examples to show gene allied splicing events with opposite survival results. d The number of genes with survival-bipolar events in different splice pattern. AT-AT and AP-AP were the most two patterns producing survival opposite events
Clinical relevant splicing events identified in the 12 cancers
TNM-stage system, a standard reference for treatment decision and prognosis prediction, is widely used for classifying cancer patients. Patients in stage IV, often found with metastasis in most cancers, may surfer much shorter survival and worse life quality, whom need much more specific individual treatment. Splicing events with median PSI varied more than 0.1 and corrected p-value < 0.05 between stage IV and non-stage IV patients were considered as advanced stage-related events (AASEs) in our study. There are average 70 AASEs of 8 cancers ranged from 17 of COAD to 148 of KIRC.
Splicing events that involved in any one of the DASEs, SASEs, and AASEs were considered as clinical relevant events, which appeared in more cancer types may associated greater significantly with tumorigenesis or clinical prognosis [22]. Totally, there are 95 splicing events of 65 genes that relevant to clinical characters in ten and more cancer types (Fig. 4). Given SASEs and AASEs are much more characteristic for each cancer, those clinical relevant events were mostly involved in DASEs. Furthermore, there are 83 splicing events of 51 CRGs that relevant to clinical characters in 5 and more cancers (Supplementary Fig. S5A). GO enrichment analysis showed that these cancer-related genes were mainly enriched in regulation of the transcription pathway (Supplementary Fig. S5B).
Splicing events potentially used for molecular classification
Pan-cancer clustering analysis for molecular subtype classification of protein expression data and genome variations, including mRNA/ncRNA expression, mutation, DNA methylation, and copy number alterations, have been performed in several studies [22,23,24,25]. Alternative splicing is an important regulator of gene expression of coding or noncoding genes and proportions of protein transforms, which may work in molecular classification. In our study, we performed an unsupervised hierarchical clustering analysis first between 31 cancers by mean PSI values based on the most varied 1590 splicing events (Supplementary Fig. S6A). These events appeared significant tissues specificity especially in cancers of blood and central nerve system. By using consensus k-means clustering method furtherly, a total of 9172 tumor samples with clinical data in 31 cancers were clustered in different 12 clusters based on the most varied 1590 splicing events (Fig. 5a; Supplementary Fig. S6B, C). For most clusters, they clustered with different types of cancer samples symmetrically such as clusters C1-3. For cluster C12, C11, and C4, however, they predominately clustered with LGG, GBM, and LAML, respectively (Supplementary Fig. S6D). For most cancer types, they were clustered to different clusters similarly, except specific cancers like LGG, GBM, and LAML (Fig. 5a, b). Interestingly, COAD, READ, ESCA, STAD, UCEC, and OV were similarly clustered together to C10, C6, C1, and C3. Thus, we furtherly analyzed microsatellite instability (MSI) signatures of those samples in COAD, READ, ESCA, STAD, and UCEC, which were attached with MSI information in clinical data sets, and observed that the MSI-high (MSI-H) account for a larger proportion in cluster C3 than other clusters (Supplementary Fig. S6E).
Clustering analysis in 9172 samples of 31 cancers based on 1590 most varied events revealed potentials of alternative splicing for molecular subtypes classification. a Distribution of cancer types by clusters are shown in heatmap and annotations. Top 200 cluster-specific events of cluster C3 and C12 are marked by rectangles. b The proportion of each type of cancer that clustered in different clusters. c Different clinical outcomes of 12 clusters are shown by Kaplan–Meier plot. Samples in cluster C12 were considered with the best survival, while samples in C3 suffer the worst survival. d GO enrichment analyses of genes related to cluster-specific events of cluster C3 and C12 were performed, respectively, and top ten terms of each cluster are displayed in bubble plots
Kaplan–Meier plot showed that samples in cluster C12 have significantly better survival, while samples in cluster C3 oppositely have worse survival (Fig. 5c). To identify cluster-specific events that may contribute to those two opposite results, Mann–Whitney test was performed between cluster C3 vs non-C3 and C12 vs non-C12 samples, and top 200 splicing events with the most significant p-value were identified and ranked (Fig. 5a). Enrichment analyses showed that genes of cluster C12 specific events were mainly enriched to the cell–cell adhesion pathway. Genes of cluster C3 specific events were mainly enriched to terms of protein transport and regulation of proteinase activity. Though quite different of the enriched genes (C3: SEC23A, SEC31A, LGMN, DCTN5, DCTN6; and C12: SEC31A, AP1B1, DCTN1, DCTN2), both of these two clusters significantly enriched to term of antigen processing and presentation exogenous peptide antigen via MHC class II, indicating that regulations of immune responses by alternative splicing play important roles in tumor patients’ survival (Fig. 5d).
Splicing networks constructed by RNA-binding proteins and splicing events
RNA-binding proteins (RBPs) including splicing factors (SFs) are trans-acting factors that influence splicing sites selection by recognizing trans-regulatory elements of the pre-mRNA to regulate alternative splicing. There are thus far about 1355 genes encoding known and predicted RNA-binding proteins, including 228 splicing factors [26, 27]. Though alternative splicing regulated by lncRNA or histone modification were reported recently, it is common sense that the mainly regulators of alternative splicing are still RBPs [28]. Correlation analysis of RBPs and splicing events have been implemented to explore potential disease-related splicing regulatory networks in several researches [16, 27].
To efficiently identify the potential relationships between splicing events (PSI value) and specific RBPs (mRNA level), we supposed that the more cancers involved, the greater significantly to identify RBPs-splicing regulatory networks. Therefore, correlations of PSI values of the splicing events and mRNA levels of the specific RBPs were calculated separately in 31 human cancers, and each network with correlation coefficient >0.7 or <−0.7 were recorded. Finally, significant RBP-splicing networks in more than 11 cancers were shown integrally (Fig. 6a, b). Specially, for known splicing factors, the networks in more than four cancers were also reported (Supplementary Fig. S7A, B).
RBPs-splicing regulatory networks identifying in 31 cancers. a Splicing regulatory networks with correlation coefficients of RBPs and splicing events > 0.7 in more than seven cancers. b Splicing regulatory networks with correlation coefficients of RBPs and splicing events < −0.7 in more than seven cancers
Prognostic signatures constructed by SASEs predicting clinical outcomes
Prognostic signatures constructed by abnormal gene expression, mutation, and methylation were reported in many researches. Recently, prognostic signatures of alternative splicing were studied to predict clinical outcome in several tumors [16]. Generally, these signature-composed splicing events were selected by ranking p-value (pRank) that generated from univariate cox regression model, which was restricted to widely use especially for a larger number of variables. For a huge dimension data, there are 2n ways of combination for n variables, which is intractable to choose an applicable one as the optimal predicting signature. In our study, we have implemented a machine-learning method named random forest survival model (rfsModel) to identify critical events for efficiently signature constructing. For comparing, AUCs at 1, 3, 5 years of signatures constructed by rfsModel and pRank were calculated in 31 cancers, respectively. Significance of differences of AUCs derived from these two methods was accessed by paired sample Student’s test in each type of cancer (Fig. 7a). Splicing events consisted in prognostic signatures with best performance were integrally presented (Supplementary Table S3). The results showed that rfsModel was better than pRank when comparing AUCs of their signatures in many cancers. BLCA, for example, there were 17 SASEs selected by rfsModel to construct predicting signature. The AUC values of this signature for survival prediction were 0.86, 0.87 and 0.90, separately, at 1, 3, and 5 year. Kaplan–Meier plot showed that patients with high risk and low risk have quite different survival (Fig. 7b, c).
Critical events selecting and prediction signatures constructing in 31 cancers. a AUCs at 1, 3, 5 years of each cancer-specific prediction signatures constructed by pRank and rfsModel were compared by median with range in bar plots. Paired samples Student’s test were implemented for accessing the significances. “**”p < 0.01; “*”p < 0.05; “.”p < 0.1. b Predicting signature for BLCA constructed by rfsModel was valued by AUCs at 1, 3, 5 years and showed by time-dependent ROC curve. c Predicting signature for BLCA constructed by rfsModel was evaluated by Kaplan–Meier plot. Patients in high risk suffer a significant worse survival than the low risk
Different splicing of KRAS, TCF7L2, and NUMA1 validated in renal cell carcinoma
In colorectal cancer, prognostic value of KRAS splicing was reported that low relative KRAS-4A expression was associated with higher level of KRAS signaling and lower overall survival [29]. In the analysis section of different alternative splicing events in this study, 398 DASEs within 184 CRGs were identified across 12 cancer types (Supplementary Table S1). For splice type ES, KRAS_20820_ES, TCF7L2_13156_ES, and NUMA1_17515_ES were three of the most significant DASEs in KIRC. RT-PCR analysis revealed that the shorter isoforms of KRAS and TCF7L2 were highly expressed in tumor tissues compared with adjacent normal tissues of KIRC patients, while the shorter isoform of NUMA1 was lowly expressed in tumor tissues (Fig. 8a–c). It is still unclear whether the alternative splicing of these genes is the cause or result of tumorigenesis. Therefore, more work needs to be done to explore how alternative splicing of these genes can play a role in the development of renal cell carcinoma.
Validation of different splicing of KRAS, TCF7L2, and NUMA1 in renal cell carcinoma. a The shorter isoforms of KRAS and TCF7L2 were found highly expressed while the shorter isoform of NUMA1 lowly expressed in tumor tissues by RT-PCR analysis. b Wilcoxon signed rank test validated that different alternative splicing of KRAS, TCF7L2, and NUMA1 was significant in human samples. c Sanger sequencing conformed the sequence information of alternative splicing of KRAS
Discussion
Alternative splicing represents a critical posttranscriptional regulation of gene expression. Defects of alternative splicing contribute to numerous diseases including cancers. Many years recently, difficulties in precision detection and quantitation of alternative spliced mRNA isoforms restrict the application of alternative splicing in tumor diagnosis, prognosis prediction, and targeting treatment. Based on RNA-seq data, several algorithms to detect alternative splicing have been explored, including SpliceSeq [30], Cufflinks [31], SplAdder [32]. Instead of detecting mRNA isoforms, however, these algorithms were applied for splicing events, using PSI to describe the inclusion or exclusion of exons, not to mention the quantitative analysis of isoforms. Therefore, the precise prediction and deterministic causality of cancers based on splicing events are still challenged. More recently, a comprehensive study of splicing events accompanying with genetic alterations were performed using SplAdder, which had detected five alternative splicing patterns (AA, AD, ES, ME, and RI), excluding AP and AT [13]. The mechanisms involved in AP and AT may be different from those of the other splicing modes, but they all result in variable exons using and producing of different protein isoforms. AP and AT also known as AFE and ALE, separately, both of which often confer splicing isoforms specific localization and control survival and migration of cancer cells [33,34,35,36]. In our study, we have found that a large number of splicing events in AP and AT were clinical relevant in cancers, which may function in tumor genesis and development. Alternative splicing of DNAJB6 (AP) modulated by CDK12, for example, promoted migration capacity and invasiveness of breast tumor cells [36]. All these data indicate that alternative splicing including splicing events in AP and AT play important roles in tumors.
CRGs represent a catalogue of genes with mutations that causally implicated in one or more cancer types. Genetic alterations including irregular alternative splicing of many genes especially for CRGs, that change genes expression or activity, were involved in cancer development and progression and used for tumor causality analysis. For instance, alternative splicing of exon 8 (AA) of VEGFA transform its anti-angiogenic function in normal tissues to angiogenesis function in tumors [37]. Alternative splicing of exon 9 of spleen tyrosine kinase (SYK) producing two different isoforms: the longer isoform SYK-L promoting cancer cells survival and tumor malignancy and the shorter isoform SYK-S inducing apoptosis of cancer cells [38]. Given tissue specificity of alternative splicing and locality of splicing events, the same splicing events were also reported with varied functions in different cancers in previous studies and in our analysis.
A number of splicing events were found survival associated and/or differentially spliced in tumor tissues. For example, PBRM1 that involved in transcriptional activation and negative regulation of cell proliferation was found second most mutated in KIRC [39]. Inactivation of PBRM1 were reported associated with tumor aggressiveness and advanced stage of KIRC [40,41,42]. When blast exon sequence of PBRM1 to Ensembl database, exclusion of exon 28 and 29 of PBRM1 (lower PSI of PBRM1_65236) resulting defect of high mobility group (HMG) box domain, which is essential for DNA binding of PBRM1 [43], were found overregulated in KIRC (lowed PSI of PBRM1_65236 in tumors).
Nonsense-mediated mRNA decay (NMD) is one of conserved RNA surveillance system, which recognizes and eliminates deleterious mRNA isoforms that contain premature translation termination codons (PTCs) [44, 45]. Defects in NMD pathways, including genetic alteration of UPFs (UPF1, UPF2, UPF3A, and UPF3B) that compile the most important complex function in NMD, producing accumulation of toxic truncated proteins, were confirmed associated with cancers [4, 46,47,48]. PTCs were often introduced by cassette exon or retained intron as well as nonsense mutations or frame‐shift mutations [49, 50]. Derived from alternative splicing, ~30% of mRNA isoforms have been estimated to contain a nonsense codon (PTC) gave rise to NMD [51]. For example, increase expression of polypyrimdine tract binding protein (PTBP1) targeting itself leads to the skipping of exon 11 of PTBP1 pre-mRNA, which in turn induces a frameshift and a PTC within exon 12 and results NMD consuming redundant PTBP1 products [6]. Thus, alternative splicing producing varied transcript isoforms not only contribute to diversity of proteins but also participate in regulation of gene expression by cooperating with the NMD pathway.
KLK2 encodes protein hk2 which consist 261 amino acids (isoform KLK2-201). Genetic alterations of KLK2 were verified to be associated with prostate cancer as well as KLK3 encoding protein prostate-specific antigen (PSA) [52]. In our study, we found that inclusion of exon 2.2 (higher PSI of KLK2_51239) and AT splicing of exon 4.4 (higher PSI of KLK2_51235) were both associated with worse survival of prostate cancer, while AT splicing of exon 5 (higher PSI of KLK2_51234) was related to better survival. When blast exon sequence of KLK2 to Ensembl, we found that AT splicing of exon 5 was associated with integrality (isoforms KLK2-201 and KLK2-203), while inclusion of exon 2.2 (isoform KLK2-205) and AT splicing of exon 4.4 (isoforms KLK2-208,214) were associated with nonsense-mediated mRNA decay of KLK2. Furthermore, correlation analysis showed that expression of UPFs in PRAD were significantly positively correlated with KLK2_51239, while negatively correlated with KLK2_51234, especially for UPF3B (Supplementary Fig. S8). Thus, we infer that inclusion of exon 2.2 (86 base) of KLK2 (higher PSI of KLK2_51239) induces a frameshift and a PTC (codon: TAA), resulting NMD and regulating gene expression of KLK2 in prostate cancer.
In normal, alternative splicing varied transitionally when tissues development contribute to the acquisition of adult tissue functions and identity [1, 53]. Continuous evolving produces tissue-, cell-, and development-specific alternative splicing and regulatory networks by RBPs, which involved in tissues normal development and disease genesis. For example, thousands of cell-specific alternative splicing events were identified between neuronal and nonneuronal cells, and SLM2 individually activating a highly cell-specific alternative splicing program was reported to control the properties of glutamatergic synapses [54, 55]. Tissue specificity of splicing networks regulated by RBPs were also illustrated, that 90% of totally 82 tissue-specific RBPs were identified in germline, brain, muscle, bone marrow, or liver cells, indicating tissue specificity of numerous alternative splicing events targeted by these RBPs [26]. Similarly in our study, LGG, GBM, and LAML that derived from blood and brain systems significantly characterized from other solid tumors and clustered predominately in specific clusters, such as C12, C11, and C4. Furthermore, cancers of the digestive system such as COAD, READ, STAD, and ESCA that commonly ascribed with similar characteristics were clustered closely together. We also observed LUSC, CESC, and HNSC that derived from different tissues were clustered to the same clusters tagged by pathologic characteristic of squamous cell carcinoma. These results indicating that a number of splicing events are tissue- and/or cell-specific and benefit to molecular classification of cancers.
The RBPs wear many hats for regulations of posttranscriptional modification and RNA processing. Defection of RBPs including mutations and altered expression commonly associated with irregular alternative splicing as well as mutations in pre-mRNA splice sites among cancers [14, 15, 27]. RBPs were also reported to control each other’s function and expression at several levels, including by synergistic or antagonistic modulation of their binding activities and by mutual or automatic splicing [6, 27]. For the 20,530 coding genes of 31 cancers, there are 64,174 events within 12,692 (61.8%) genes in total. While for the 1355 RBPs, there are 6467 splicing events within 1032 (76.2%) genes, which indicate that RBPs are preferred genes for splicing. Similarly, genes allied to significant different splicing events in different cancers enriched to the mRNA splicing pathway (Table 2), regulation of RNA splicing or mRNA splice site selection (Supplementary Fig. S3) pathways, indicating that irregular splicing of these RNA regulatory genes may play important roles in tumor genesis and development.
Compared with the enormous number of detected alternative splicing events, however, only a few irregular spliced isoforms have been validated as direct contributors to tumors. Here, we have identified numerous of clinical relevant splicing events especially for events of CRGs, which may function as tumor suppressor or cancer driver events in specific cancers. We have also explored an online resource for visualization of clinical relevance of splicing events, which could be used for identifying cancer relevant events by combining with genetic information. The increasing efforts to identify cancer-related defects in pre-mRNA splicing and develop new chemotherapies targeting the varied alternative splicing would offer more individual treatments for cancer patients. Indeed, these fields have recently offered many new therapeutic opportunities, including RNA-binding regulators, anti-sense oligonucleotides, molecules against core spliceosome machinery, and chemical inhibitors of splicing factors kinase [56, 57]. By the development of detecting alternative splicing, especial for the increasing application of single-cell sequence and long-read sequence, there will be many more cancer-related splicing isoforms identified and targeted in the future [58, 59].
Materials and methods
Data obtaining and processing
Splicing data indicated as percent splice in (PSI) for all cancers were obtained from the SpliceSeq database (http://bioinformatics.mdanderson.org/TCGASpliceSeq/index.jsp). The PSI value of a splicing event, representing the relative expression level of a longer transcript (Supplementary Fig. S1C), is an intuitive ratio for calculating the splicing efficiency of a gene sequence into a transcript isoform [30, 60]. The primary PSI data with no more than 25% missing values were analyzed. In total, 9708 tumor samples of 31 cancers and 557 paired tumor and adjacent normal samples of 12 cancers were available for analysis. In most cancer types, about 80% splicing events were found with values in more than 90% samples, which were maintained to performing further analyses (Supplementary Fig. S1A). Events with standard deviation more than 0.05 were used in the survival study, and the missing values were filled up with average PSI value of each events. Samples were then divided into two groups (indicated by 0 = lower and 1 = higher) by the median of PSI values for univariate cox regression analysis and Kaplan–Meier plot. Besides, clinical data and level 3 RNA-seq data of 31 cancers were obtained from TCGA cohort by UCSC Xena website. Known cancer-related genes (tier = 1) were obtained from COSMIC database (https://cancer.sanger.ac.uk/cosmic/download).
Differential and survival-associated splicing events
Twelve cancers (including BLCA that with 19 paired samples) with more than 20 paired tumor and adjacent normal tissues were analyzed to identify different alternative splicing events (DASEs). Wilcoxon test was performed to evaluate the significances of DASEs in paired tumor and adjacent normal tissues, and Benjamini–Hochberg method was used to correct for multiple testing. Splicing events with varied median PSI > 0.1 in absolute and corrected p-value < 0.05 were considered as significant different alternative splicing events (DASEs) [27]. Univariate cox regression analyses were performed for splicing events with standard deviation >0.05, and percent samples with values >90% by the survival package in R software. Survival-associated splicing events (SASEs) were identified with p-value < 0.05. For patients with clinical events of overall survival less than ten in PRAD, TGCT, and THCA, cox regression analyses and Kaplan–Meier plots of these cancers were performed based on recurrence free survival data.
Identification of clinical relevant splicing events
Advanced stage associated splicing events (AASEs) were identified in eight cancers (BLCA, BRCA, COAD, HNSC, KIRC, LUAD, STAD, and UCEC), of which there are >20% or absolute 20 patients in stage IV. Median PSI values were calculated in stage IV and non-stage IV patients, separately, for each splicing event, and Mann–Whitney test was used to evaluate the difference in two groups, and Benjamini–Hochberg method was used to correct for multiple testing. As result, splicing events with PSI values varied >0.1 and corrected p-value < 0.05 were considered as significant AASEs. Splicing events that involved in any one of the DASEs, SASEs, and AASEs were considered as clinical relevant events. Clinical relevant events that appeared in more cancers were depicted as more critical cancer-related events that may associated with tumorigenesis or prognosis [22].
Clustering analysis and identification of cluster-specific splicing events
Average PSI value of each events in all cancer types were calculated separately, then standard deviation of average PSI value between these cancers were calculated, and 1590 splicing events with standard deviation (s.d.) >0.1 were considered as most variable events and selected for further study. Unsupervised hierarchical clustering were performed first between mean PSI of these 31 cancers based on the 1590 splicing events. By the consensus k-means clustering method (using the “ConsensusClusterPlus” R package), furthermore, the 1590 splicing events with the most variation were taken for clustering analysis in 9172 tumor samples with clinical data of 31 cancers. Consensus matrix and delta area plots affiliated to ConsensusClusterPlus program were used for selecting k-value of clustering. For cluster C3 and C12, Mann–Whitney test was performed to identify top 200 significant cluster-specific events.
Critical splicing events identified by random forest survival analysis
To professionally recognize the panel of splicing events with optimal clinical prognostic value from the characteristic SASEs, we performed a random forest survival analysis using machine-learning algorithm, which is a highly integrated method. A developed R package named RandomForestSRC was used in this study. First, whole SASEs were selected to construct the random forest survival model (rfsModel) (parameters setting: ntree = 1000, mtry = p/3, and nodesize = 3, p represent number of SASEs). Second, formula var.select() depending on minimal tree depth algorithm was used to identify important variable (parameters setting: method = “vh” and nrep = 100). The top important variables selected were furthermore used to fit a multivariate coxPH model finally, and variables with p-value < 0.1 were considered as individual prognostic factors.
Prognostic signatures generation
The traditional method that the most significant SASEs ranked by p-value (pRank) were further fit a multivariate cox regression model in each type of cancer separately, generating a panel of signature to predict clinical outcome. We also constructed another signature in each cancer with critical SASEs selected by the rfsModel. The area under receiver-operating characteristics curve (AUCs) of signatures were calculated at 1, 3, 5 years, and compared by paired sample Student’s test in each cancer. AUCs of each signature constructed by these two methods were then used to access their performance on predicting patients’ survival. Risk scores of these signatures were calculated by coefficients of their component features and corresponding PSI levels, by which patients were re-split into two groups: the higher and the lower. Kaplan–Meier plots were generated to evaluate their impacts on clinical outcomes.
Validation in human samples
Ten pairs of tumor and adjacent normal tissues were obtained from KIRC patients treated at the Department of Urology of Tongji Hospital (Wuhan, China) after their written informed consent. All the tissues were kept in liquid nitrogen before RNA extraction. The total RNA was extracted using Trizol (Invitrogen, USA), and reverse-transcribed to cDNA using revertase (TaKaRa, Dalian, China) as recommended by the manufacturer. RT-PCR was performed in paired samples by splicing-specific primers (Supplementary Table S4) using 2 × Green PCR Mix (Vazyme, Jinan, China). Splicing-specific products were isolated by agarose gel electrophoresis and grayscale measurement by software Image J (Rawak Software Inc., Stuttgart, Germany). Splicing isoforms isolated by electrophoresis were recovered and performed Sanger sequencing (Tsingke, Wuhan, China) to conform the gene sequence of KRAS splicing.
Statistical analyses and visualization
Statistical analyses were performed using the R software (version 3.4.2) and Python (version 3.6). The data visualization tools including the ggplot2, pheatmap, survival, survminer, survivalROC, ConsensusClusterPlus, and randomforestSRC packages of Rwere used in this study. Bar and some other graphs of splicing landscape were visualized by Graphpad Prism version 7.01 (GraphPad Software Inc., San Diego, CA, USA).
Construction of online resource
The OncoSplicing website was developed in JSP using a Struts2 framework and was deployed on a Tomcat 6.0.44 web server that ran under a Redhat 6.4 system. All data in OncoSplicing were stored and managed by using MySQL (version 5.7.18). Rserve (version 1.86) and two pieces of R code were used to manage the results and visualization.
Code availability
Research code that was used to implement methods described in this study is publicly available on GitHub: https://github.com/yjzhang2013/OncoSplicing.
References
Baralle FE, Giudice J. Alternative splicing as a regulator of development and tissue identity. Nat Rev Mol Cell Biol. 2017;18:437–51.
Kelemen O, Convertini P, Zhang Z, Wen Y, Shen M, Falaleeva M, et al. Function of alternative splicing. Gene. 2013;514:1–30.
Blencowe BJ. The relationship between alternative splicing and proteomic complexity. Trends Biochem Sci. 2017;42:407–8.
da Costa PJ, Menezes J, Romao L. The role of alternative splicing coupled to nonsense-mediated mRNA decay in human disease. Int J Biochem Cell Biol. 2017;91:168–75.
Tabrez SS, Sharma RD, Jain V, Siddiqui AA, Mukhopadhyay A. Differential alternative splicing coupled to nonsense-mediated decay of mRNA ensures dietary restriction-induced longevity. Nat Commun. 2017;8:306.
Wollerton MC, Gooding C, Wagner EJ, Garcia-Blanco MA, Smith CW. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol Cell. 2004;13:91–100.
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008;40:1413–5.
Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, et al. Alternative isoform regulation in human tissue transcriptomes. Nature. 2008;456:470–6.
Davuluri RV, Suzuki Y, Sugano S, Plass C, Huang TH. The functional consequences of alternative promoter use in mammalian genomes. Trends Genet. 2008;24:167–77.
Climente-Gonzalez H, Porta-Pardo E, Godzik A, Eyras E. The functional impact of alternative splicing in cancer. Cell Rep. 2017;20:2215–26.
Bauman JA, Li SD, Yang A, Huang L, Kole R. Anti-tumor activity of splice-switching oligonucleotides. Nucleic Acids Res. 2010;38:8348–56.
Li H, Wang Z, Xiao W, Yan L, Guan W, Hu Z, et al. Androgen-receptor splice variant-7-positive prostate cancer: a novel molecular subtype with markedly worse androgen-deprivation therapy outcomes in newly diagnosed patients. Mod Pathol. 2018;31:198–208.
Kahles A, Lehmann KV, Toussaint NC, Huser M, Stark SG, Sachsenberg T, et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell. 2018;34:211–24 e6.
Tian J, Wang Z, Mei S, Yang N, Yang Y, Ke J, et al. CancerSplicingQTL: a database for genome-wide identification of splicing QTLs in human cancer. Nucleic Acids Res. 2018;47:D909–16.
Jayasinghe RG, Cao S, Gao Q, Wendl MC, Vo NS, Reynolds SM, et al. Systematic analysis of splice-site-creating mutations in cancer. Cell Rep. 2018;23:270–81 e3.
Li Y, Sun N, Lu Z, Sun S, Huang J, Chen Z, et al. Prognostic alternative mRNA splicing signature in non-small cell lung cancer. Cancer Lett. 2017;393:40–51.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Bader GD, Hogue CW. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinf. 2003;4:2.
Hong SK. Kallikreins as biomarkers for prostate cancer. Biomed Res Int. 2014;2014:526341.
Ylitalo EB, Thysell E, Jernberg E, Lundholm M, Crnalic S, Egevad L, et al. Subgroups of castration-resistant prostate cancer bone metastases defined through an inverse relationship between androgen receptor activity and immune response. Eur Urol. 2017;72:E147–7.
Guo ZQ, Zheng T, Chen B, Luo C, Ouyang S, Gong S, et al. Small-molecule targeting of E3 ligase adaptor SPOP in kidney cancer. Cancer Cell. 2016;30:474–84.
Han L, Diao L, Yu S, Xu X, Li J, Zhang R, et al. The genomic landscape and clinical relevance of A-to-I RNA editing in human cancers. Cancer Cell. 2015;28:515–28.
Chen F, Zhang Y, Gibbons DL, Deneen B, Kwiatkowski DJ, Ittmann M, et al. Pan-cancer molecular classes transcending tumor lineage across 32 cancer types, multiple data platforms, and over 10,000 cases. Clin Cancer Res. 2018;24:2182–93.
Karpinski P, Pesz K, Sasiadek MM. Pan-cancer analysis reveals presence of pronounced DNA methylation drift in CpG island methylator phenotype clusters. Epigenomics. 2017;9:1341–52.
Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell. 2014;158:929–44.
Gerstberger S, Hafner M, Tuschl T. A census of human RNA-binding proteins. Nat Rev Genet. 2014;15:829–45.
Sebestyen E, Singh B, Minana B, Pages A, Mateo F, Pujana MA, et al. Large-scale analysis of genome and transcriptome alterations in multiple tumors unveils novel cancer-relevant splicing networks. Genome Res. 2016;26:732–44.
Fu XD, Ares M Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat Rev Genet. 2014;15:689–701.
Eilertsen IA, Sveen A, Stromme JM, Skotheim RI, Nesbakken A, Lothe RA. Alternative splicing expands the prognostic impact of KRAS in microsatellite stable primary colorectal cancer. Int J Cancer. 2018;144:841–7.
Ryan MC, Cleland J, Kim R, Wong WC, Weinstein JN. SpliceSeq: a resource for analysis and visualization of RNA-Seq data on alternative splicing and its functional impactsfunctional impacts. Bioinforma. 2012;28:2385–7.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012;7:562–78.
Kahles A, Ong CS, Zhong Y, Ratsch G. SplAdder: identification, quantification and testing of alternative splicing events from RNA-Seq data. Bioinforma. 2016;32:1840–7.
Taliaferro JM, Vidaki M, Oliveira R, Olson S, Zhan L, Saxena T, et al. Distal alternative last exons localize mRNAs to neural projections. Mol cell. 2016;61:821–33.
Miller MB, Yan Y, Wu Y, Hao B, Mains RE, Eipper BA. Alternate promoter usage generates two subpopulations of the neuronal RhoGEF Kalirin-7. J Neurochem. 2017;140:889–902.
Ushijima T, Hanada K, Gotoh E, Yamori W, Kodama Y, Tanaka H, et al. Light controls protein localization through phytochrome-mediated alternative promoter selection. Cell. 2017;171:1316.
Tien JF, Mazloomian A, Cheng SG, Hughes CS, Chow CCT, Canapi LT, et al. CDK12 regulates alternative last exon mRNA splicing and promotes breast cancer cell invasion. Nucleic Acids Res. 2017;45:6698–716.
Ladomery MR, Harper SJ, Bates DO. Alternative splicing in angiogenesis: the vascular endothelial growth factor paradigm. Cancer Lett. 2007;249:133–42.
Prinos P, Garneau D, Lucier JF, Gendron D, Couture S, Boivin M, et al. Alternative splicing of SYK regulates mitosis and cell survival. Nat Struct Mol Biol. 2011;18:673–9.
Varela I, Tarpey P, Raine K, Huang D, Ong CK, Stephens P, et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature. 2011;469:539–42.
Gao W, Li W, Xiao T, Liu XS, Kaelin WG Jr. Inactivation of the PBRM1 tumor suppressor gene amplifies the HIF-response in VHL-/- clear cell renal carcinoma. PNAS. 2017;114:1027–32.
Hogner A, Krause H, Jandrig B, Kasim M, Fuller TF, Schostak M, et al. PBRM1 and VHL expression correlate in human clear cell renal cell carcinoma with differential association with patient’s overall survival. Urol Oncol. 2018;36:94e1–e14.
Kim JY, Lee SH, Moon KC, Kwak C, Kim HH, Keam B, et al. The impact of PBRM1 expression as a prognostic and predictive marker in metastatic renal cell carcinoma. J Urol. 2015;194:1112–9.
Stros M, Launholt D, Grasser KD. The HMG-box: a versatile protein domain occurring in a wide variety of DNA-binding proteins. Cell Mol Life Sci. 2007;64:2590–606.
Popp MW, Maquat LE. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu Rev Genet. 2013;47:139–65.
Chang YF, Imam JS, Wilkinson MF. The nonsense-mediated decay RNA surveillance pathway. Annu Rev Biochem. 2007;76:51–74.
Lu JW, Plank TD, Su F, Shi XJ, Liu C, Ji Y, et al. The nonsense-mediated RNA decay pathway is disrupted in inflammatory myofibroblastic tumors. J Clin Invest. 2016;126:3058–62.
Lindeboom RG, Supek F, Lehner B. The rules and impact of nonsense-mediated mRNA decay in human cancers. Nat Genet. 2016;48:1112–8.
Shum EY, Jones SH, Shao A, Dumdie J, Krause MD, Chan WK, et al. The antagonistic gene paralogs Upf3a and Upf3b govern nonsense-mediated RNA decay. Cell. 2016;165:382–95.
Ge Y, Porse BT. The functional consequences of intron retention: Alternative splicing coupled to NMD as a regulator of gene expression. Bioessays. 2014;36:236–43.
Yan Q, Weyn-Vanhentenryck SM, Wu J, Sloan SA, Zhang Y, Chen K, et al. Systematic discovery of regulated and conserved alternative exons in the mammalian brain reveals NMD modulating chromatin regulators. PNAS. 2015;112:3445–50.
Green RE, Lewis BP, Hillman RT, Blanchette M, Lareau LF, Garnett AT, et al. Widespread predicted nonsense-mediated mRNA decay of alternatively-spliced transcripts of human normal and disease genes. Bioinforma. 2003;19:i118–21.
Nam RK, Zhang WW, Klotz LH, Trachtenberg J, Jewett MA, Sweet J, et al. Variants of the hK2 protein gene (KLK2) are associated with serum hK2 levels and predict the presence of prostate cancer at biopsy. Clin Cancer Res. 2006;12:6452–8.
Vuong CK, Black DL, Zheng S. The neurogenetics of alternative splicing. Nat Rev Neurosci. 2016;17:265–81.
Traunmuller L, Gomez AM, Nguyen TM, Scheiffele P. Control of neuronal synapse specification by a highly dedicated alternative splicing program. Science. 2016;352:982–6.
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S, et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci. 2014;34:11929–47.
Bates DO, Morris JC, Oltean S, Donaldson LF. Pharmacology of Modulators of Alternative Splicing. Pharmacol Rev. 2017;69:63–79.
Han T, Goralski M, Gaskill N, Capota E, Kim J, Ting TC, et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science. 2017;356:eaal3755.
Arzalluz-Luque A, Conesa A. Single-cell RNAseq for the study of isoforms-how is that possible? Genome Biol. 2018;19:110.
Tilgner H, Jahanbani F, Gupta I, Collier P, Wei E, Rasmussen M, et al. Microfluidic isoform sequencing shows widespread splicing coordination in the human transcriptome. Genome Res. 2018;28:231–42.
Schafer S, Miao K, Benson CC, Heinig M, Cook SA, Hubner N. Alternative splicing signatures in RNA-seq data: percent spliced in (PSI). Curr Protoc Hum Genet. 2015;87:11.16.1–14.
Acknowledgements
The authors would like to thank Xudong Zhang of Genek Company for helpful advices of data processing. This work was funded by the National Natural Science Foundation of China (81702522, 81602236) and National Major Scientific and Technological Special Project for Significant New Drugs Development (2017ZX09304022).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Y., Yan, L., Zeng, J. et al. Pan-cancer analysis of clinical relevance of alternative splicing events in 31 human cancers. Oncogene 38, 6678–6695 (2019). https://doi.org/10.1038/s41388-019-0910-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0910-7
- Springer Nature Limited
This article is cited by
-
Alternative splicing in prostate cancer progression and therapeutic resistance
Oncogene (2024)
-
Alternative splicing: a new breakthrough for understanding tumorigenesis and potential clinical applications
Genes & Genomics (2023)
-
Dissection of molecular and histological subtypes of papillary thyroid cancer using alternative splicing profiles
Experimental & Molecular Medicine (2022)
-
Broad misappropriation of developmental splicing profile by cancer in multiple organs
Nature Communications (2022)
-
AS-CMC: a pan-cancer database of alternative splicing for molecular classification of cancer
Scientific Reports (2022)