Abstract
APOE ɛ4, the most significant genetic risk factor for Alzheimer disease (AD), may mask effects of other loci. We re-analyzed genome-wide association study (GWAS) data from the International Genomics of Alzheimer’s Project (IGAP) Consortium in APOE ɛ4+ (10 352 cases and 9207 controls) and APOE ɛ4− (7184 cases and 26 968 controls) subgroups as well as in the total sample testing for interaction between a single-nucleotide polymorphism (SNP) and APOE ɛ4 status. Suggestive associations (P<1 × 10-4) in stage 1 were evaluated in an independent sample (stage 2) containing 4203 subjects (APOE ɛ4+: 1250 cases and 536 controls; APOE ɛ4−: 718 cases and 1699 controls). Among APOE ɛ4− subjects, novel genome-wide significant (GWS) association was observed with 17 SNPs (all between KANSL1 and LRRC37A on chromosome 17 near MAPT) in a meta-analysis of the stage 1 and stage 2 data sets (best SNP, rs2732703, P=5·8 × 10−9). Conditional analysis revealed that rs2732703 accounted for association signals in the entire 100-kilobase region that includes MAPT. Except for previously identified AD loci showing stronger association in APOE ɛ4+ subjects (CR1 and CLU) or APOE ɛ4− subjects (MS4A6A/MS4A4A/MS4A6E), no other SNPs were significantly associated with AD in a specific APOE genotype subgroup. In addition, the finding in the stage 1 sample that AD risk is significantly influenced by the interaction of APOE with rs1595014 in TMEM106B (P=1·6 × 10−7) is noteworthy, because TMEM106B variants have previously been associated with risk of frontotemporal dementia. Expression quantitative trait locus analysis revealed that rs113986870, one of the GWS SNPs near rs2732703, is significantly associated with four KANSL1 probes that target transcription of the first translated exon and an untranslated exon in hippocampus (P⩽1.3 × 10-8), frontal cortex (P⩽1.3 × 10-9) and temporal cortex (P⩽1.2 × 10−11). Rs113986870 is also strongly associated with a MAPT probe that targets transcription of alternatively spliced exon 3 in frontal cortex (P=9.2 × 10−6) and temporal cortex (P=2.6 × 10−6). Our APOE-stratified GWAS is the first to show GWS association for AD with SNPs in the chromosome 17q21.31 region. Replication of this finding in independent samples is needed to verify that SNPs in this region have significantly stronger effects on AD risk in persons lacking APOE ɛ4 compared with persons carrying this allele, and if this is found to hold, further examination of this region and studies aimed at deciphering the mechanism(s) are warranted.
Similar content being viewed by others
Introduction
The common late-onset form of Alzheimer disease (AD) has a strong genetic component,1 a portion of which is explained by APOE and several other genes identified by positional mapping, targeted gene analysis and genome-wide association studies (GWAS).2, 3, 4 Together, these loci account for less than one-half of the heritable component in AD susceptibility, of which 20–25% is due to APOE.4, 5 Because many of the known AD loci cluster in biological pathways, including those involved in inflammation, lipid metabolism and processing and intracellular trafficking of Aβ, there are likely more AD risk loci that are difficult to detect because of very weak effect size, allelic heterogeneity or rare variants. To examine yet another hypothesis, namely, that associations for some loci may be obscured by confounding or interaction with other loci, we conducted a two-stage GWAS in APOE genotype subgroups using the large resources of the International Genomics of Alzheimer’s Project (IGAP).
Methods
Study population
Details of the stage 1 sample from the IGAP Consortium, including subject recruitment, genotyping, imputation, quality control, population substructure and statistical methods for association analyses, were previously described.4 In brief, phenotype and genotype data, including APOE genotypes, for a total of 53 711 subjects were assembled by IGAP from the Alzheimer’s Disease Genetic Consortium (ADGC), the Cohorts for Heart and Ageing Research in Genomic Epidemiology (CHARGE) consortium, the European Alzheimer's Disease Initiative (EADI) and the Genetic and Environmental Risk in Alzheimer’s Disease (GERAD) consortium. Characteristics of this sample are in Supplementary Table S1. The stage 2 data set included GWAS and APOE genotype data for 4203 subjects of European ancestry from the ADC4, ADC5, ADC6, MTV, Pfizer and TARCC data sets in the ADGC. These individuals were recruited under protocols approved by the appropriate Institutional Review Boards. Details of the individual data sets are provided in the Supplementary Materials and summarized in Supplementary Table S1.
Procedures
QC, imputation and population substructure in stage 2 data sets. Quality control of the clinical and genotype data in these cohorts was performed using the procedures described elsewhere.4 Single-nucleotide polymorphism (SNP) genotypes in each stage 2 data set were imputed with IMPUTE2 using reference haplotypes from the March 2012 release of 1000 Genomes. We compared imputation results for selected variants in the stage 1 data sets using the March 2012 release of 1000 Genomes and prior imputation on the December 2010 release and found no significant difference in the distribution of genotype probabilities between old and new imputations for the same samples among the original ADGC data sets. We used actual APOE genotypes when available, because previously we observed that imputation in this region using the 1000 Genomes reference panel is unreliable.5 Population substructure was evaluated within each data set by principal components analysis using EIGENSTRAT (http://www.hsph.harvard.edu/alkes-price/software/) and a subset of 21 109 SNPs common to all genotyping platforms.
Statistical analysis
Genome-wide association study. Within each stage 1 data set, genome-wide association analyses were conducted separately in subgroups of subjects with and without the APOE ɛ4 allele using a logistic generalized linear model in case–control data sets and a logistic generalized estimating equation in family-based data sets. The potential independent effect of the APOE ɛ2 allele was not examined because of the paucity of carriers of this allele, thus rendering very small cell sizes particularly among AD cases and in smaller data sets. Cox-proportional hazards models were used to evaluate association with incident AD in three CHARGE cohorts. A quantitative estimate between 0 and 2 for the dose of the reference allele for a SNP was used to incorporate the uncertainty of the imputation estimates. Interaction between a SNP and APOE genotype was evaluated in the APOE genotype subgroups combined within each data set using regression models, including age, sex, the first three principal components and terms for the SNP, APOE ɛ4 status and interaction between the SNP and APOE ɛ4 status. Results for each model across data sets were combined by meta-analysis using the inverse variance method implemented in the software package METAL (http://www.sph.umich.edu/csg/abecasis/Metal/). Effect sizes were weighted by their inverse variance and a combined estimate was calculated by summing the weighted estimates and dividing by the summed weights. SNPs with a minor allele frequency >5% that were available in at least 50% of the data sets were included in the meta-analysis. The meta-analysis P-value for association was estimated by the summarized test statistic, after applying genomic control within each individual study.
Follow-up analysis in stage 2 data sets. SNPs attaining a P-value <10−4 in the stage 1 GWAS were evaluated in each of the stage 2 GWAS data sets, containing a total of 1786 APOE ɛ4+ and 2417 APOE ɛ4− subjects (Supplementary Table S1), using the same approach described above.
Gene expression analysis
The effect of top-ranked SNPs on gene expression was evaluated using an open access database of control brain microarray data (BRAINEAC) made publically available by the UK Human Brain Expression Consortium (http://caprica.genetics.kcl.ac.uk/BRAINEAC). This data set contains information generated by analysis of tissue samples obtained from 12 different central nervous system regions in 134 individuals. Details of the expression quantitative trait locus (eQTL) analysis are reported elsewhere.6 In this study, the experiment-wise significance threshold for association of a genetic marker with expression was determined to be 1.6 × 10−7 at the gene level and 1.8 × 10−6 for individual exons. Potential for functionality of the top-ranked SNPs was assessed using the Regulome database (http://www.regulomedb.org).
Results
We conducted a genome-wide association study for AD using data sets stratified by APOE genotype assembled by IGAP, which were from the ADGC, CHARGE consortium, EADI and GERAD consortium. Meta-analyses were performed separately in APOE ɛ4+ (10 246 cases and 11 924 controls) and APOE ɛ4− (7231 cases and 19 603 controls) subgroups, as well as the total sample, using a model including a term for the interaction of the SNP with the APOE ɛ4 status. There was limited genomic inflation in the GWAS results in the APOE ɛ4+ (λ=1.05) and APOE ɛ4− (λ=1.06) groups, but not in the total sample (λ=0.98) testing the ɛ4 × SNP interaction (Supplementary Figure S1). Genome-wide significant (GWS) association (P<5 × 10−8) for AD was found in five distinct regions (CR1, BIN1, CLU, PICALM and APOE) in the APOE ɛ4+ subgroup (Supplementary Figure S2A, Supplementary Table S2) and four distinct regions (BIN1, HBEGF, MS4A6A/MS4A4A, SLC24A4 and APOE) in the APOE ɛ4− subgroup (Supplementary Figure S2B, Supplementary Table S2). No significant SNP × APOE interactions were found in the total group (Supplementary Figure S2C). Suggestive association (P<10−6) was observed with SNPs in five novel loci in the APOE ɛ4− subgroup (SOX14/CLDN18, ACSL6, FAM20C, MAPT region and CDR2L; Supplementary Figure S2B, Supplementary Table S3) and with 21 TMEM106B SNPs (top result: rs1595014, P=1.6 × 10-7) (Supplementary Figure S2C, Supplementary Table S3).
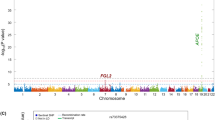
Approximately 1130 SNPs from 38 regions (including 7 previously established AD loci) were tested in stage 2 (Supplementary Table S3). Follow-up analyses of the novel loci confirmed association with SNPs in CDC42SE2-ACSL6, KANSL1/LRRC37A and CDR2L in the stage 2 sample (Table 1, Supplementary Table S2), but only SNPs near MAPT and between KANSL1 and LRRC37A (Figure 1a) were GWS after combining results from the stages 1 and 2 samples (best SNP: rs2732703, meta-analysis: P=5.8 × 10−9). The association was consistent in nearly all data sets which contained rs2732703 information (Figure 1b). To verify the reliability of the association with rs2732703, an imputed SNP, we compared rs2732703 allele dosages obtained directly by genotyping using a Taqman assay with those derived from imputation among 1010 subjects from the ACT, ADC4, ADC5 and ADC6 data sets. The correlation of these values, 0.813 in the entire sample and 0.834 among APOE ɛ4− subjects, as well as a genotype misclassification rate of only 3.5% among subjects with imputed probability scores>0.8 for a particular genotype, suggest that our association findings were not influenced substantially by imputation quality.
Association of Alzheimer’s disease with single-nucleotide polymorphisms (SNPs) in chromosome 17q21.31 in the combined stages 1 and 2 samples. (a) Regional Manhattan plot in the APOE ɛ4+ (upper panel) and the APOE ɛ4− (lower panel) subgroups. SNPs with the lowest P-value are indicated with a purple diamond. Computed estimates of linkage disequilibrium (r2) of SNPs in this region with the most significant SNP are shown as red circles for r2⩾0.8, orange circles for 0.6⩽r2<0.8, green circles for 0.4⩽r2<0.6, light blue circles for 0.2⩽r2<0.4, and blue circles for r2<0.2. Unannotated SNPs are shown as grey circles. (b) Forest plot of association results for rs2732703 in the stages 1 and 2 and total samples among APOE ɛ4− subjects. CI, confidence interval; MAF, minor allele frequency; OR, odds ratio.
Further examination of this region in the total sample revealed an association peak spanning >1.25 Mb that contains 15 genes (Figure 1a). Within this region, 17 SNPs were GWS, have minor allele frequencies ranging from 0.13 to 0.17 and are located in a 10.2-kb segment upstream of both KANSL1 and LRRC37A (Supplementary Table S4). Nominally significant association was observed with only one of these SNPs among ɛ4+ subjects (rs2732703, P=0.02) (Supplementary Table S3). Although the odds ratios (OR) for effect of the effect of minor allele on AD risk were substantially lower for all of the GWS SNPs in the ɛ4− group (0.54<OR<0.86) than in the ɛ4+ group (0.76<β<1.04), there was no evidence of interaction with APOE genotype (Supplementary Table S3). The minor alleles of these SNPs reduced AD risk by 20–37% in the ɛ4− group. The 350-kb gap in the broad association signal is punctuated at one end by a ‘cliff’ adjacent to the MAPT–KANSL1–LRRC37A association peak (Figure 1). This gap is populated by relatively few SNPs and contains several copy-number variation polymorphisms.7, 8 To explore the possibility that the association observed in the present analysis is explained by previously identified haplotypes H1/H2 in the MAPT region,8 we evaluated six models in the entire data set conditioning on rs8070723 (an H1/H2 tagging SNP), rs2732703 or rs199533. Rs2732703 remained significant in models conditioning on rs8070723 (P=0.013) or rs199533 (P=0.0020), and rs8070723 was marginally significant in the model conditioning on rs199533 (P=0.043) (Supplementary Table S5, Supplementary Figure S3). These results suggest that KANSL1/LRRC37A is the only AD risk locus in this region.
We also examined the effect of APOE ɛ4 status on previously established AD loci (Supplementary Table S2). Four of these loci attained genome-wide significance in at least one of the APOE subgroups (Table 2), and the association signal in the MS4A cluster region was evident primarily in the APOE ɛ4− subgroup (Supplementary Figure S4). The association of AD with CR1, BIN1 and CLU was supported in both the APOE subgroups.
Next we interrogated the BRAINEAC database to determine whether any of the 17 GWS SNPs located between KANSL1 and LRRC37A are cis-eQTLs. Data were available for only one of these SNPs (rs113986870) that is in high linkage disequilibrium (LD) with and 2461 base pairs away from rs2732703 (r2 and D’>0.9). Ten exon probes from four genes (KANSL1, LRRC37A4P, MAPT and C17orf69) were significantly associated with rs113986870 when averaged across all brain regions (Table 3). Rs113986870 was significantly associated with gene-level expression (Figure 2a), as well as with exon-level expression, (Figure 2b) in hippocampus, temporal cortex and cerebellum. In these brain regions, rs113986870 was significantly associated with KANSL1 probes 3762011, 3762012 and 3762013 that measure expression of the first translated exon. Additionally, we observed that expression of probe 3760518 (Supplementary Figure S5A) present in all three transcripts (NM_001193466, NM_015443 and NM_001193465) and 3760219 in transcript variant 2 (NM_015443) was significantly associated with rs113986870 (Supplementary Figure S5B), while expression of probe 3760217 in transcript variant 1 (NM_001193466) was not significant (Supplementary Figure S5C), indicating that alternative splicing may be a crucial mechanism for regulating KANSL1 expression. Rs113986870 was also strongly associated with MAPT transcription (Supplementary Figure S6A) and in particular with probe 3723712 that targets transcription of alternatively spliced exon 3 in frontal cortex (P⩽9.2 × 10−6) and temporal cortex (P⩽2.6 × 10−6) (Supplementary Figure S6B). The rs113986870 minor allele (A), which is associated with reduced risk of AD (Supplementary Table S4), increased the expression of the target exons in KANSL1 and MAPT (Figure 2, Supplementary Figure S6, Supplementary Figure S7). The association with LRRC37A4P exon probe 3759898 was significant in all three AD-related brain regions (P⩽3.6 × 10−9). The association of rs113986870 with exon probe 3723594 for C17orf69 was significant in hippocampus only (P=1.6 × 10−7). Five of the GWS SNPs, including rs2732703 and rs113986870, are located within a transcription factor-binding site or a DNase sensitivity peak, and two of these five SNPs, including rs2668626 which is only 47 bp from rs2732703, have also been identified within an eQTL (Supplementary Table S4).
Genotype specific effect of the expression quantitative trait locus (eQTL) rs113986870 on expression of KANSL1. (a) Gene-level expression of KANSL1 transcript t3760137. Transcript-level expression represents the average across all KANSL1 exon probe sets. (b) Expression of exon probe 3760212. Probes 3760211, 3760212 and 3760213 measure expression of the first translated exon, are present in all three transcript variants and were significantly associated with the eQTL. Expression profiles for probes 3760211 and 3760213 were similar to those for probe 3760212 (Table 3). The distance from 3760212 to rs113986870 is 85 431 base pairs. Log2 scale of expression (y axis) is shown for 10 regions of cognitively normal human brains (x axis) ordered by mean expression level. Rs113986870 genotype counts: AA=0, AG=56, and GG=76. Rs113986870 allele frequencies are 0.21 (A) and 0.79 (G). CRBL, cerebellum; FCTX, frontal cortex; HIPP, hippocampus; MEDU, medulla (specifically inferior olivary nucleus); OCTX, occipital cortex (specifically primary visual cortex); PUTM, putamen, SNIG, substantia nigra; TCTX, temporal cortex; THAL, thalamus; WHMT, intralobular white matter.
Discussion
This study was undertaken to identify loci whose effect on AD risk may be obscured by confounding or interaction with APOE genotype. Our APOE-stratified GWAS is the first to show GWS association for AD with SNPs in the chromosome 17q21.31 region, including MAPT, KANSL1 and LRRC37A. Among the genes expected to emerge from GWAS but never seen before is MAPT, which encodes the microtubule-associated protein tau (MAPT) found in AD neurofibrillary tangles. The association peak is located between KANSL1 and LRRC37A, approximately 200 kb downstream of MAPT, in a subset of subjects who do not possess the APOE ɛ4 allele. Although the association signal includes MAPT, conditional analysis suggests that the causal variant(s) are more likely located in a DNA segment between the 5′ end of KANSL1 and 5′ end of LRRC37A and not within MAPT or another gene distal to LRRC37A.
The nature of the AD-related functional variant could not be discerned from our genetic association findings. None of the GWS SNPs are within 42.1 kb of the KANSL1 start site or 16.8 kb of the LRRC37A start site, suggesting that the functional variant is not within the promoter region of either gene. KANSL1 is a widely expressed gene encoding a member of the nonspecific lethal complex. The KANSL1 protein is an evolutionarily conserved regulator of the chromatin modifier KAT8, which influences gene expression through histone H4 lysine 16 acetylation.9 Notably, mutations in KANSL1 cause the 17q21.31 microdeletion syndrome, which is associated with a wide range of abnormalities, including intellectual disability and developmental delay, and is therefore thought to be involved in neuronal development.10, 11 LRRC37A encodes a member of the leucine-rich repeat containing 37 family. Leucine-rich repeats (LRRs) are protein–ligand interaction motifs found in a large number of proteins with different structure, localization and function.12 LRR motifs are important for intermolecular or intercellular interactions with exogenous factors in the immune system and/or with different cell types in the developing nervous system.12
However, expression analysis of exon array data in control brain tissue revealed that rs113986870, which is in high LD with the top-ranked SNP (rs2732703) in the GWAS, is an eQTL for expression of the first translated exon in KANSL1 and the alternatively spliced exon 3 in MAPT. Previous studies suggest that splicing of MAPT may be a crucial regulatory mechanism in the brain and tauopathies in particular13 and that increased expression of exon 3 protects against neurodegeneration.14 Although rs113986870 is apparently not an eQTL for its adjacent gene LRRC37A, it was significantly associated with a closely related gene, LRRC37A4P, in all three AD-related brain regions. These results suggest that rs113986870 may have a potential function as a cis-acting regulatory element for multiple genes in this region. Another confounding feature of this region are copy-number variations that in part overlap with the 5′ end of KANSL1 and possibly influence expression.7, 8 Thus it is possible that the exon probes targeting the first translated in KANSL1 may be tagging this duplication. In addition, interrogation of a database curating information about DNA features and regulatory regions revealed that five of the GWS SNPs, including rs2732703 and rs113986870, may have strong regulatory potential.
The association peak for AD on chromosome 17q21.31 is located in a well-recognized and perplexing genomic region containing a 900-kb inversion.8 Previous GWAS identified associations of variants within and at the edges of this inversion with Parkinson disease15 and progressive supranuclear palsy,16 but the most significant associations were not with SNPs between KANSL1 and LRRC37A (Supplementary Table S6). Multiple studies have identified >40 MAPT deletions, missense mutations and splice site mutations that cause frontotemporal dementia (FTD).17 Although AD is only nominally associated with common variants in MAPT, previously we observed association of a rare MAPT variant (A152T) with increased risk for FTD and AD in a large sample,18 a finding which was supported by a subsequent smaller study.19 Ikram et al.20 identified a GWS association peak with a KANSL1 SNP approximately 166 kb away from our most significant AD SNP (rs2732703) for a continuous measure of intracranial volume in a sample of nearly 10 000 community-dwelling elders (Supplementary Table S6). These two SNPs are moderately correlated (r2=0.71), which indicates that they may tag the same functional variant.
Other studies have focused on two divergent extended MAPT haplotypes, H1 and H2, which are in near complete LD with status of the inversion and contain independently derived partial duplications of KANSL1.8, 16 The common H1 haplotype is associated with increased risk of FTD,21 Parkinson disease,22 progressive supranuclear palsy23 and corticobasal degeneration23 while H2 is linked to recurrent deletion events associated with the 17q21.31 microdeletion syndrome.10 Among these non-AD forms of dementia, it is possible for FTD to masquerade clinically as AD and thereby cases of FTD could be present in our study group; however, any inadvertent inclusion of FTD cases is expected to be very small as the minimum age of dementia onset in our study group was 60 years and onset of dementia from FTD after age 69 years is relatively rare compared with AD that in most cases occurs after age 69 years.24 Furthermore, a recent review of almost 5000 autopsy brains from a subset of cases in the ADGC cohort failed to identify any case of FTD.25 Myers et al.26 reported association of AD with H1 and with common MAPT SNPs, but this association is controversial27 and did not reach genome-wide significance in our study or previous GWAS. Another recent study showed that carriers of at least one H2 allele had a 5.4-fold increased risk of worsening hallucinations, but this result was marginally significant.28 Previously, we observed in a subset of the sample studied here that the H2-haplotype tagging rs8070723-G allele was associated with reduced risk of AD.29 However, this variant is no longer associated after conditioning on rs2732703 (Supplementary Table S5). In carriers of H2, the ancestral haplotype in both humans and chimpanzees,30 increased expression of exon 3 in MAPT has been associated with an eQTL located approximately 1500 bp from rs113986870, which decreases aggregation of microtubules.6, 31 These observations are consistent with our results showing that the rs113986870 minor allele is protective for AD and associated with elevated exon3 expression.
There is a large body of experimental evidence linking tau protein to AD pathogenesis,32 and some studies show evidence of association of AD with common MAPT SNPs.29, 33 However, analysis of the MAPT coding sequence did not reveal disease-causing variants for early-onset AD,34 and other studies examining association of MAPT SNPs with late-onset AD were negative.27, 35 Recently, Allen et al.29 reported that the rs8070723-G allele was associated with reduced MAPT expression in the cerebellum and temporal cortex of AD subjects. Robust genetic associations have also been identified for AD with several genes in cytoskeletal and axonal transport pathways, including tau, or leading to neurofibrillary tangles, most notably BIN1, EPHA1, RIN3, CASS4 and FERMT2.4
Based on the observation that overexpression of human ApoE4 in transgenic mouse neurons results in hyperphosphorylation of tau,36 it is possible that associations with AD-related loci in the chromosome 17q21.31 region are obscured by the much stronger effect of APOE ɛ4 on MAPT expression or function.37 This idea is consistent with lack of GWS association with 17q21.31 SNPs in the same data set without stratification by APOE genotype,4 and no evidence for interaction between APOE and any SNPs in the MAPT–KANSL1–LRRC37A region in the current study. Another possible explanation for the significant association of 17q21.31 SNPs with AD only among subjects lacking APOE ɛ4 is genetic heterogeneity, suggesting that variation at the chromosome 17q21.31 locus is associated with a distinct etiological subtype of AD where tau is the primary disease activator.38 Finally, the diagnosis of AD for most subjects in this data set was established, clinically suggesting the possibility of misdiagnosis or AD accompanied by other processes associated with other dementing illnesses. Further studies are needed to determine whether this subtype can be distinguished clinically or neuropathologically.
Our study also showed that the previously established association with the MS4A gene cluster is derived almost completely from subjects lacking APOE ɛ4, suggesting the contribution of the MS4A locus to AD may be mechanistically different than AD-related processes that are associated with APOE ɛ4. Members of the MS4A gene family encode membrane proteins, some of which have known roles in immune cell function;39 however, little is known about the function of MS4A6A, MS4A4A or MS4A6E in humans. Karch et al.40 showed that expression of MS4A6A was upregulated in the brains of AD patients compared with the brains of controls and significantly correlated with AD status, AIF1 expression (a marker for microglia, which is the immune cell of the brain), cognitive dementia rating score and extent of AD neuropathological change.
The observed statistical interaction of genotypes for TMEM106B with APOE on AD risk in the stage 1 GWAS is noteworthy (rs1595014, P=1.6 × 10−7) even though it is not supported by results in the comparatively small stage 2 sample. TMEM106B is a glycoprotein predominantly localized at the lysosomal membrane where it might interact with intracellular progranulin (GRN).41, 42 TMEM106B variants, particularly the p. T185S (rs3173615) mutation, are risk factors for FTD, especially among persons carrying a GRN mutation.43 TMEM106B variants are also associated with development of cognitive impairment in amyotrophic lateral sclerosis44 and implicated in the pathological presentation of AD.45 Cruchaga et al.46 observed association of the TMEM106B SNP rs1990622 risk allele with younger onset of the frontotemporal lobar degeneration subtype with TAR DNA-binding protein inclusions (FTLD-TDP), a pattern reminiscent of the association of APOE ɛ4 with increased risk and younger onset of AD. The biological underpinning of the interaction of TMEM106B with APOE affecting AD risk is unclear.
Our top findings, including those that are GWS, should be confirmed in independent samples. Functional studies will be needed to understand the relationship between APOE and the causative variant(s) in 17q21.31 once they are identified, as well as with other loci showing much stronger association with AD in particular APOE genotype strata (for example, MS4A6A/MS4A4A/MS4A6E) or through interaction with APOE (for example, TMEM106B). Our study provides a firm genetic connection of AD to several other pathologically distinct disorders in which dementia is a cardinal or common characteristic.
References
Gatz M, Reynolds CA, Fratiglioni L, Johansson B, Mortimer JA, Berg S et al. Role of genes and environments for explaining Alzheimer disease. Arch Gen Psychiatry 2006; 63: 168–174.
Farrer LA, Cupples LA, Haines JL, Hyman B, Kukull WA, Mayeux R et al. Effects of age, gender and ethnicity on the association of apolipoprotein E genotype and Alzheimer disease. JAMA 1997; 278: 1349–1356.
Sherva R, Farrer LA . Power and pitfalls of the genome wide association study approach to identify genes for Alzheimer disease. Cur Psychiatry Rep 2011; 13: 138–146.
Lambert J-C, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C et al. Meta-analysis of 74,538 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 2013; 45: 1452–1458.
Jun G, Vardarajan BN, Buros J, Yu CE, Hawk MV, Dombroski BA et al. A comprehensive search for Alzheimer disease susceptibility loci in the APOE region. Arch Neurol 2012; 69: 1270–1279.
Trabzuni D, Ryten M, Walker R, Smith C, Imran S, Ramasamy A et al. Quality control parameters on a large dataset of regionally dissected human control brains for whole genome expression studies. J Neurochem 2011; 119: 275–282.
Steinberg KM, Antonacci F, Sudmant PH, Kidd JM, Campbell CD, Vives L et al. Structural diversity and African origin of the 17q21.31 inversion polymorphism. Nat Genet 2012; 44: 872–880.
Boettger LM, Handsaker RE, Zody MC, McCarroll SA . Structural haplotypes and recent evolution of the human 17q21.31 region. Nat Genet 2012; 44: 881–885.
Li X, Wu L, Corsa CA, Kunkel S, Dou Y . Two mammalian MOF complexes regulate transcription activation by distinct mechanisms. Mol Cell 2009; 36: 290–300.
Koolen DA, Kramer JM, Neveling K, Nillesen WM, Moore-Barton HL, Elmslie FV et al. Mutations in the chromatin modifier gene KANSL1 cause the 17q21.31 microdeletion syndrome. Nat Genet 2012; 44: 639–641.
Zollino M, Orteschi D, Murdolo M, Lattante S, Battaglia D, Stefanini C, Mercuri E et al. Mutations in KANSL1 cause the 17q21.31 microdeletion syndrome phenotype. Nat Genet 2012; 44: 636–638.
Giannuzzi G, Siswara P, Malig M, Marques-Bonet TNISC Comparative Sequencing Program Mullikin JC et al. Evolutionary dynamism of the primate LRRC37 gene family. Genome Res 2013; 23: 46–59.
Trabzuni D, Wray S, Vandrovcova J, Ramasamy A, Walker R, Smith C et al. MAPT expression and splicing is differentially regulated by brain region: relation to genotype and implication for tauopathies. Hum Mol Genet 2012; 21: 4094–4103.
Caffrey TM, Joachim C, Wade-Martins R . Haplotype-specific expression of the N-terminal exons 2 and 3 at the human MAPT locus. Neurobiol Aging 2008; 29: 1923–1929.
Simón-Sánchez J, Schulte C, Bras JM, Sharma M, Gibbs JR, Berg D et al. Genome-wide association study reveals genetic risk underlying Parkinson's disease. Nat Genet 2009; 41: 1308–1312.
Höglinger GU, Melhem NM, Dickson DW, Sleiman PM, Wang LS, Klei L et al. Identification of common variants influencing risk of the tauopathy progressive supranuclear palsy. Nat Genet 2011; 43: 699–705.
Ferrari R, Hardy J, Momeni P . Frontotemporal dementia: from Mendelian genetics towards genome wide association. J Mol Neurosci 2011; 45: 500–515.
Coppola G, Chinnathambi S, Lee JJ, Dombroski BA, Baker MC, Soto-Ortolaza AI et al. Evidence for a role of the rare p.A152T variant in MAPT in increasing the risk for FTD-spectrum and Alzheimer's diseases. Hum Mol Genet 2012; 21: 3500–3512.
Lee SE, Tartaglia MC, Yener G, Genç S, Seeley WW, Sanchez-Juan P et al. Neurodegenerative disease phenotypes in carriers of MAPT p.A152T, a risk factor for frontotemporal dementia spectrum disorders and Alzheimer disease. Alzheimer Dis Assoc Disord 2013; 27: 302–309.
Ikram MA, Fornage M, Smith AV, Seshadri S, Schmidt R, Debette S et al. Common variants at 6q22 and 17q21 are associated with intracranial volume. Nat Genet 2012; 44: 539–544.
Verpillat P, Camuzat A, Hannequin D, Thomas-Anterion C, Puel M, Belliard S et al. Association between the extended tau haplotype and frontotemporal dementia. Arch Neurol 2002; 59: 935–939.
Zabetian CP, Hutter CM, Factor SA, Nutt JG, Higgins DS, Griffith A et al. Association analysis of MAPT H1 haplotype and subhaplotypes in Parkinson's disease. Ann Neurol 2007; 62: 137–144.
Pittman AM, Myers AJ, Abou-Sleiman P, Fung HC, Kaleem M, Marlowe L et al. Linkage disequilibrium fine mapping and haplotype association analysis of the tau gene in progressive supranuclear palsy and corticobasal degeneration. J Med Genet 2005; 42: 837–846.
Knopman DS, Petersen RC, Edland SD, Cha RH, Rocca WA . The incidence of frontotemporal lobar degeneration in Rochester, Minnesota, 1990 through 1994. Neurology 2004; 62: 506–508.
Beecham GW, Hamilton K, Naj A, Martin ER, Huentelman M, Myers AJ et al. Genome-wide association meta-analysis of neuropathologic features of Alzheimer’s disease and related dementias. PLoS Genet 2014; 10: e1004606.
Myers AJ, Kaleem M, Marlowe L, Pittman AM, Lees AJ, Fung HC et al. The H1c haplotype at the MAPT locus is associated with Alzheimer’s disease. Hum Mol Genet 2005; 14: 2399–2404.
Abraham R, Sims R, Carroll L, Hollingworth P, O'Donovan MC, Williams J et al. An association study of common variation at the MAPT locus with late-onset Alzheimer's disease. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 1152–1155.
Creese B, Corbett A, Jones E, Fox C, Ballard C . Role of the extended MAPT haplotype in the worsening of psychotic symptoms and treatment response in Alzheimer disease. J Am Med Dir Assoc 2014; 15: 934–937, In press.
Allen M, Kachadoorian M, Quicksall Z, Zou F, Chai HS, Younkin C et al. Association of MAPT haplotypes with Alzheimer's disease risk and MAPT brain gene expression levels. Alz Res Ther 2015; 6: 39, In press.
Zody MC, Jiang Z, Fung HC, Antonacci F, Hillier LW, Cardone MF et al. Evolutionary toggling of the MAPT 17q21.31 inversion region. Nat Genet 2008; 40: 1076–1083.
Zhong Q, Congdon EE, Nagaraja HN, Kuret J . Tau isoform composition influences the rate and extent of filament formation. J Biol Chem 2012; 287: 20711–20719.
Krstic D, Knuesel I . Deciphering the mechanism underlying late-onset Alzheimer disease. Nat Rev Neurol 2013; 9: 25–34.
Laws SM, Friedrich P, Diehl-Schmid J, Müller J, Eisele T, Bäuml J et al. Fine mapping of the MAPT locus using quantitative trait analysis identifies possible causal variants in Alzheimer's disease. Mol Psychiatry 2007; 12: 510–517.
Roks G, Dermaut B, Heutink P, Julliams A, Backhovens H, Van de Broeck M et al. Mutation screening of the tau gene in patients with early-onset Alzheimer's disease. Neurosci Lett 1999; 277: 137–139.
Cousin E, Macé S, Rocher C, Dib C, Muzard G, Hannequin D et al. No replication of genetic association between candidate polymorphisms and Alzheimer's disease. Neurobiol Aging 2011; 32: 1443–1451.
Tesseur I, Van Dorpe J, Spittaels K, Van den Haute C, Moechars D, Van Leuven F . Expression of human apolipoprotein E4 in neurons causes hyperphosphorylation of protein tau in the brains of transgenic mice. Am J Pathol 2000; 156: 951–964.
Cruchaga C, Kauwe JS, Harari O, Jin SC, Cai Y, Karch CM et al. GWAS of cerebrospinal fluid tau levels identifies risk variants for Alzheimer's disease. Neuron 2013; 72: 256–268.
Morris GP, Clark IA, Vissel B . Inconsistencies and controversies surrounding the amyloid hypothesis of Alzheimer’s disease. Acta Neuropathol Commun 2014; 2: 135.
Zuccolo J, Bau J, Childs SJ, Goss GG, Sensen CW, Deans JP . Phylogenetic analysis of the MS4A and TMEM176 gene families. PLoS One 2010; 5: e9369.
Karch CM, Jeng AT, Nowotny P, Cady J, Cruchaga C, Goate AM . Expression of novel Alzheimer's disease risk genes in control and Alzheimer's disease brains. PLoS One 2012; 7 (11): e50976.
Chen-Plotkin AS, Unger TL, Gallagher MD, Bill E, Kwong LK, Volpicelli-Daley L et al. TMEM106B, the risk gene for frontotemporal dementia, is regulated by the microRNA-132/212 cluster and affects progranulin pathways. J Neurosci 2012; 32: 11213–11227.
Lang CM, Fellerer K, Schwenk BM, Kuhn PH, Kremmer E, Edbauer D et al. Membrane orientation and subcellular localization of transmembrane protein 106B (TMEM106B), a major risk factor for frontotemporal lobar degeneration. J Biol Chem 2012; 287: 19355–19365.
Van Deerlin VM, Sleiman PM, Martinez-Lage M, Chen-Plotkin A, Wang LS, Graff-Radford NR et al. Common variants at 7p21 are associated with frontotemporal lobar degeneration with TDP-43 inclusions. Nat Genet 2010; 42: 234–239.
Vass R, Ashbridge E, Geser F, Hu WT, Grossman M, Clay-Falcone D et al. Risk genotypes at TMEM106B are associated with cognitive impairment in amyotrophic lateral sclerosis. Acta Neuropathol 2011; 121: 373–380.
Rutherford NJ, Carrasquillo MM, Li M, Bisceglio G, Menke J, Josephs KA et al. TMEM106B risk variant is implicated in the pathologic presentation of Alzheimer disease. Neurology 2012; 79: 717–718.
Cruchaga C, Graff C, Chiang HH, Wang J, Hinrichs AL, Spiegel N et al. Association of TMEM106B gene polymorphism with age at onset in granulin mutation carriers and plasma granulin protein levels. Arch Neurol 2011; 68: 581–586.
Acknowledgements
ADGC
The National Institutes of Health, National Institute on Aging (NIH-NIA) supported this work through the following grants: ADGC, U01 AG032984, RC2 AG036528; NACC, U01 AG016976; NCRAD, U24 AG021886; NIAGADS, U24-AG041689; NIA LOAD, U24 AG026395, U24 AG026390; MIRAGE: R01 AG025259; Banner Sun Health Research Institute P30 AG019610; Boston University, P30 AG013846, U01 AG10483, R01 CA129769, R01 MH080295, R01 AG017173, R01AG33193; Columbia University, P50 AG008702, R37 AG015473; Duke University, P30 AG028377, AG05128; Emory University, AG025688; Group Health Research Institute, UO1 AG06781, UO1 HG004610; U01 HG006375; Indiana University, P30 AG10133; Johns Hopkins University, P50 AG005146, R01 AG020688; Massachusetts General Hospital, P50 AG005134; Mayo Clinic, P50 AG016574; Mount Sinai School of Medicine, P50 AG005138, P01 AG002219; New York University, P30 AG08051, MO1RR00096, and UL1 RR029893; Northwestern University, P30 AG013854; Oregon Health & Science University, P30 AG008017, R01 AG026916; Rush University, P30 AG010161, R01 AG019085, R01 AG15819, R01 AG17917, R01 AG30146; TGen, R01 NS059873; University of Alabama at Birmingham, P50 AG016582, UL1RR02777; University of Arizona, R01 AG031581; University of California, Davis, P30 AG010129; University of California, Irvine, P50 AG016573, P50, P50 AG016575, P50 AG016576, P50 AG016577; University of California, Los Angeles, P50 AG016570; University of California, San Diego, P50 AG005131; University of California, San Francisco, P50 AG023501, P01 AG019724; University of Kentucky, P30 AG028383; University of Michigan, P50 AG008671; University of Pennsylvania, P30 AG010124; University of Pittsburgh, P50 AG005133, AG030653, AG041718; University of Southern California, P50 AG005142; University of Texas Southwestern, P30 AG012300; University of Miami, R01 AG027944, AG010491, AG027944, AG021547, AG019757; University of Washington, P50 AG005136; Vanderbilt University, R01 AG019085; and Washington University, P50 AG005681, P01 AG03991. The Kathleen Price Bryan Brain Bank at Duke University Medical Center is funded by NINDS grant no. NS39764, NIMH MH60451 and by Glaxo Smith Kline. Genotyping of the TGEN2 cohort was supported by Kronos Science. The TGen series was also funded by NIA grant AG034504 to AJM, The Banner Alzheimer’s Foundation, The Johnnie B. Byrd Sr. Alzheimer’s Institute, the Medical Research Council, and the state of Arizona and also includes samples from the following sites: Newcastle Brain Tissue Resource (funding via the Medical Research Council, local NHS trusts and Newcastle University), MRC London Brain Bank for Neurodegenerative Diseases (funding via the Medical Research Council),SouthWest Dementia Brain Bank (funding via numerous sources, including the Higher Education Funding Council for England (HEFCE), Alzheimer’s Research Trust (ART), BRACE as well as North Bristol NHS Trust Research and Innovation Department and DeNDRoN), The Netherlands Brain Bank (funding via numerous sources, including Stichting MS Research, Brain Net Europe, Hersenstichting Nederland Breinbrekend Werk, International Parkinson Fonds, Internationale Stiching Alzheimer Onderzoek), Institut de Neuropatologia, Servei Anatomia Patologica, Universitat de Barcelona. Marcelle Morrison-Bogorad, PhD, Tony Phelps, PhD and Walter Kukull PhD are thanked for helping to co-ordinate this collection. ADNI Funding for ADNI is through the Northern California Institute for Research and Education by grants from Abbott, AstraZeneca AB, Bayer Schering Pharma AG, Bristol-Myers Squibb, Eisai Global Clinical Development, Elan Corporation, Genentech, GE Healthcare, GlaxoSmithKline, Innogenetics, Johnson and Johnson, Eli Lilly, Medpace, Merck, Novartis AG, Pfizer, F Homan-La Roche, Schering-Plough, Synarc, Alzheimer’s Association, Alzheimer’s Drug Discovery Foundation, the Dana Foundation, and by the National Institute of Biomedical Imaging and Bioengineering and NIA grants U01 AG024904, RC2 AG036535, K01 AG030514. We thank Dr D Stephen Snyder and Dr Marilyn Miller from NIA who are ex-officio ADGC members. Support was also from the Alzheimer’s Association (LAF, IIRG-08-89720; MP-V, IIRG-05-14147) and the US Department of Veterans Affairs Administration, Office of Research and Development, Biomedical Laboratory Research Program. P.S.G.-H. is supported by the Wellcome Trust, Howard Hughes Medical Institute, and Canadian Institute of Health.
CHARGE
AGES: The AGES-Reykjavik Study is funded by NIH contract N01-AG-12100 (NIA with contributions from the NEI, NIDCD and NHLBI), the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association) and the Althingi (the Icelandic Parliament).
ASPS/PRODEM: The Austrian Stroke Prevention Study and The Prospective Dementia Register of the Austrian Alzheimer Society was supported by The Austrian Science Fond (FWF) grant number P20545-P05 (H Schmidt) and P13180; The Austrian Alzheimer Society; The Medical University of Graz.
Cardiovascular Health Study (CHS): This research was supported by NHLBI contracts HHSN268201200036C, HHSN268200800007C, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083, N01HC85086; and NHLBI grants U01HL080295, R01HL087652, R01HL105756, R01HL103612, and R01HL120393 with additional contribution from the National Institute of Neurological Disorders and Stroke (NINDS). Additional support was provided through R01AG023629, R01AG15928, R01AG20098, R01AG027058 and R01AG033193 (Seshadri) from the National Institute on Aging (NIA). A full list of principal CHS investigators and institutions can be found at CHS-NHLBI.org. The provision of genotyping data was supported in part by the National Center for Advancing Translational Sciences, CTSI grant UL1TR000124, and the National Institute of Diabetes and Digestive and Kidney Disease Diabetes Research Center (DRC) grant DK063491 to the Southern California Diabetes Endocrinology Research Center.
Framingham Heart Study (FHS): This work was supported by the National Heart, Lung and Blood Institute’s Framingham Heart Study (Contract No. N01-HC-25195) and its contract with Affymetrix, Inc for genotyping services (Contract No. N02-HL-6-4278). A portion of this research utilized the Linux Cluster for Genetic Analysis (LinGAII) funded by the Robert Dawson Evans Endowment of the Department of Medicine at Boston University School of Medicine and Boston Medical Center. This study as also supported by grants from the National Institute on Aging: AG08122 and AG033193 (Seshadri). Dr Seshadri and Dr DeStefano were also supported by additional grants from the National Institute on Aging: (R01 AG16495; AG031287, AG033040), the National Institute of Neurological Disorders and Stroke (R01 NS17950), and the National Heart, Lung and Blood Institute (U01 HL096917, HL093029 and K24HL038444, RC2-HL102419 and UC2 HL103010.
Fundació ACE: would like to thank patients and controls who participated in this project. This work has been funded by the Fundación Alzheimur (Murcia), the Ministerio de Educación y Ciencia (PCT-010000-2007-18), (DEX-580000-2008-4), (Gobierno de España), Corporación Tecnológica de Andalucía (08/211) and Agencia IDEA (841318) (Consejería de Innovación, Junta de Andalucía). We thank Ms Trinitat Port-Carbó and her family for their generous support of Fundació ACE research programs.
Erasmus Rucphen Family Study: We thank the participants from the Genetic Research in Isolated Populations in the Erasmus Rucphen Family Study who made this work possible. This study is financially supported by the Netherlands Organization for Scientific Research (NWO), the Internationale Stichting Alzheimer Onderzoek (ISAO), the Hersenstichting Nederland (HSN) and the Centre for Medical Systems Biology (CMSB1 and CMSB2) in the framework of the Netherlands Genomics Initiative (NGI).
The Rotterdam Study: The Rotterdam Study was funded by Erasmus Medical Center and Erasmus University, Rotterdam; the Netherlands Organization for Health Research and Development; the Research Institute for Diseases in the Elderly; the Ministry of Education, Culture and Science; the Ministry for Health, Welfare and Sports; the European Commission; and the Municipality of Rotterdam; by grants from the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), Internationale Stichting Alzheimer Onderzoek, Hersenstichting Nederland, the Netherlands Genomics Initiative–Netherlands Organization for Scientific Research (Center for Medical Systems Biology and the Netherlands Consortium for Healthy Aging), the Seventh Framework Program (FP7/2007-2013), the ENGAGE project (grant agreement HEALTH-F4-2007-201413), MRACE-grant from the Erasmus Medical Center and the Netherlands Organization for Health Research and Development (ZonMW Veni-grant no. 916.13.054).
ARIC: The Atherosclerosis Risk in Communities Study (ARIC) is carried out as a collaborative study supported by National Heart, Lung, and Blood Institute contracts N01-HC-55015, N01-HC-55016, N01-HC-55018, N01-HC-55019, N01-HC-55020, N01-HC-55021, N01-HC-55022 and grants R01-HL087641, RC2-HL102419 (Boerwinkle, CHARGE-S), UC2 HL103010, U01 HL096917 (Mosley) and R01-HL093029; NHGRI contract U01-HG004402; and NIH contract HHSN268200625226C and NIA: R01 AG033193 (Seshadri). Infrastructure was partly supported by Grant Number UL1RR025005, a component of the National Institutes of Health and NIH Roadmap for Medical Research.
EADI
This work was supported by the National Foundation for Alzheimer’s disease and related disorders, the Institut Pasteur de Lille and the Centre National de Génotypage. The Three-City Study was performed as part of a collaboration between the Institut National de la Santé et de la Recherche Médicale (Inserm), the Victor Segalen Bordeaux II University and Sanofi-Synthélabo. The Fondation pour la Recherche Médicale funded the preparation and initiation of the study. The 3C Study was also funded by the Caisse Nationale Maladie des Travailleurs Salariés, Direction Générale de la Santé, MGEN, Institut de laLongévité, Agence Française de Sécurité Sanitaire des Produits de Santé, the Aquitaine and Bourgogne RegionalCouncils, Fondation de France and the joint French Ministry of Research/INSERM ‘Cohortes et collections de données biologiques’ programme. Lille Génopôle received an unconditional grant from Eisai.
Belgium sample collection: Research at the Antwerp site is funded in part by the Interuniversity Attraction Poles program of the Belgian Science Policy Office, the Foundation for Alzheimer Research (SAO-FRA), a Methusalem Excellence Grant of the Flemish Government, the Research Foundation Flanders (FWO), the Special Research Fund of the University of Antwerp, Belgium. KB is a postdoctoral fellow of the FWO. The Antwerp site authors thank the personnel of the VIB Genetic Service Facility, the Biobank of the Institute Born-Bunge and the Departments of Neurology and Memory Clinics at the Hospital Network Antwerp and the University Hospitals Leuven.
Finish sample collection: Financial support for this project was provided by the Health Research Council of the Academy of Finland, EVO grant 5772708 of Kuopio University Hospital and the Nordic Centre of Excellence in Neurodegeneration.
Italian sample collections: The Bologna site (FL) obtained funds from the Italian Ministry of research and University as well as Carimonte Foundation. The Florence site was supported by a grant from the Italian ministry of Health (RFPS-2006-7-334858) and grant RF-2010-2319722. The Milan site was supported by a grant from the ‘fondazione Monzino’. We thank the expert contribution of Mr. Carmelo Romano. The Roma site received financial support from Italian Ministry of Health, Grant RF07-08 and RC08-09-10-11-12. The Pisa site is grateful to Dr Annalisa LoGerfo for her technical assistance in the DNA purification studies.
Spanish sample collection: The Madrid site (MB) was supported by grants of the Ministerio de Educación y Ciencia and the Ministerio de Sanidad y Consumo (Instituto de Salud Carlos III), and an institutional grant of the FundaciónRamón Areces to the CBMSO. We thank I Sastre and Dr A Martínez-García for the preparation and control of the DNA collection, and Dr P Gil and Dr P Coria for their cooperation in the cases/controls recruitment. We are grateful to the Asociación de Familiares de Alzheimer de Madrid (AFAL) for continuous encouragement and help.
Swedish sample collection: Financially supported in part by the Swedish Brain Power network, the Marianne and Marcus Wallenberg Foundation, the Swedish Research Council (521-2010-3134), the King Gustaf V and Queen Victoria’s Foundation of Freemasons, the Regional Agreement on Medical Training and Clinical Research (ALF) between Stockholm County Council and the Karolinska Institutet, the Swedish Brain Foundation and the Swedish Alzheimer Foundation.
GERAD
Cardiff University was supported by the Wellcome Trust, Medical Research Council (MRC), Alzheimer’s Research UK (ARUK) and the Welsh Assembly Government. Cambridge University and Kings College London acknowledge support from the MRC. ARUK supported sample collections at the South West Dementia Bank and the Universities of Nottingham, Manchester and Belfast. The Belfast group acknowledges support from the Alzheimer's Society, Ulster Garden Villages, N. Ireland R&D Office and the Royal College of Physicians/Dunhill Medical Trust. The MRC and Mercer’s Institute for Research on Ageing supported the Trinity College group. The South West Dementia Brain Bank acknowledges support from Bristol Research into Alzheimer's and Care of the Elderly. The Charles Wolfson Charitable Trust supported the OPTIMA group. Washington University was funded by NIH grants, Barnes Jewish Foundation and the Charles and Joanne Knight Alzheimer's Research Initiative. Patient recruitment for the MRC Prion Unit/UCL NIHR Department of Neurodegenerative Disease collection was supported by the UCLH/UCL NIHR Biomedical Centre and Queen Square Dementia Biomedical Research Unit. LASER-AD was funded by Lundbeck SA. The Bonn group was supported by the German Federal Ministry of Education and Research (BMBF), Competence Network Dementia and Competence Network Degenerative Dementia and by the Alfried Krupp von Bohlen und Halbach-Stiftung. The GERAD1 Consortium also used samples ascertained by the NIMH AD Genetics Initiative.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
Celine Bellenguez received personal fees from Genoscrenn unrelated to current study. HS reports grants from Austrian Science Fond (FWF) and Österreichische Nationalbank Anniversary Fund during the conduct of the study. FP reports grants from DISTALZ Labex of Excellence (ANR). NF reports payment from Janessen Alzheimer’s Immunotherapty and Pfizer to University College London for contracted image analyses and payment from Bristol Myers Squibb, Eli Lilly, GE Healthcare, Novartis and Sanofi to University College London for consultant services. JH received personal fees from Eisai, Eli Lilly and Roche, outside the current study. AMG received grants from NIA, Genentech, Pfizer and Astra Zeneca, as well as personal fees from Finnegan HC, Cognition Therapeutics, Dickstein Shapiro, Genentech and Amgen, all unrelated to the current study. In addition, AMG has a patent (US20070258898) issued and a patent with royalties paid by Taconic. WM received grants from Eli Lilly and Lundbeck. J-FD received grants and personal fees from IPSEN and Novartis and personal fees from Newron. HH reports personal fees and non-financial support from Boehringer-Ingelheim, Bristol-Myers-Squibb, Elan Corporation, Novartis, Eisai Inc., Pfizer, Sanofi-Aventis, Roche Pharmaceutical Diagnostics, GlaxoSmithKline-Biologicals, Jung-Diagnostics, Avid, Eli Lilly and Cytox and personal fees from GE Healthcare. In addition, HH has pending patents entitled ‘Method for predicting whether subjects with mild cognitive impairment (MCI) will develop Alzheimer's disease’, ‘3-Hydroxykynurenin im Serum als diagnostischer Marker fUr die Demenz vom Alzheimer-Typ’, ‘Neurodegenerative markers for psychiatric conditions’, ‘Ratio A~2/40 im Plasma in der Fruh- und Differentialdiagnose der Alzheimer Krankheit’, Liquordiagnostisches in vitro Verfahren zur Diagnose von Demenz-ErkrankUngen und neuroinfJammatorischen Erkrankungen’, ‘In vitro Verfahren zur Diagnose von neurodegenerativen Erkrankungen’ and ‘In vitro Verfahren zur Diagnose und Fruhdiagnose von neurodegenerativen Erkrankungen’. EBL received personal fees from Up to Date unrelated to the current study. MO received research grants from the Medical Research Council during the course of, but unrelated to, the current study. EMR received personal fees from AstraZeneca, CereSpir, Eisai, Eli Lilly, GlaxoSmithKline and Sanofi and research contracts from Avid/Eli Lilly and Genetech, all unrelated to the current study. JW is a full-time employee and shareholder of Pfizer (2012–present) and a former full-time employee of F. Hoffmann-La Roche (2010–2012). AW is a full-time employee and shareholder of Pfizer (2013–present). MAP-V received personal fees from Athena Neurosciences unrelated to the current study. The other authors declare no conflict of interest.
Additional information
International Genomics of Alzheimer’s Project Members Alzheimer Disease Genetics Consortium (ADGC) Perrie M Adams, PhD; Marilyn S Albert, PhD; Roger L Albin, MD; Liana G Apostolova, MD; Steven E Arnold, MD; Sanjay Asthana, MD; Craig S Atwood, PhD; Clinton T Baldwin, PhD; Michjael M Barmada, PhD; Lisa L Barnes, PhD; Thomas G Beach, MD PhD; James T Becker, PhD; Eileen H Bigio, MD; Thomas D Bird, MD; Deborah Blacker, MD; Bradley F Boeve, MD; James D Bowen, MD; Adam Boxer, MD PhD, James R Burke, MD PhD; Nigel J Cairns, PhD FRCPath; Chuanhai Cao, PhD; Chris S Carlson; PhD; Cynthia M Carlsson, MD; Regina M Carney, MD; Minerva M Carrasquillo, PhD; Steven L Carroll, MD PhD; Helena C Chui, MD; David G Clark, MD; Jason Corneveaux, BS; David H Cribbs, PhD; Elizabeth A Crocco, MD; Carlos Cruchaga, PhD; Philip L De Jager, MD PhD; Charles DeCarli, MD; Steven T DeKosky, MD; F Yesim Demirci, MD; Malcolm Dick, PhD; Dennis W Dickson, MD; Rachelle S Doody, MD PhD; Ranjan Duara, MD; Nilufer Ertekin-Taner, MD PhD; Kelley M Faber, MS; Thomas J Fairchild, PhD; Kenneth B Fallon, MD; Martin R Farlow, MD; Steven Ferris, PhD; Matthew P Frosch, MD PhD; Douglas R Galasko, MD; Marla Gearing, PhD; Daniel H Geschwind, MD PhD; Bernardino Ghetti, MD; John R Gilbert PhD; Jonathan D Glass, MD; Neill R Graff-Radford, MD; Robert C Green, MD MPH; John H Growdon, MD; Hakon Hakonarson, MD PhD; Ronald L Hamilton, MD; John Hardy, PhD; Lindy E Harrell, MD PhD; Elizabeth Head, PhD; Lawrence S Honig, MD PhD; Ryan M Huebinger, PhD, Matthew J Huentelman, PhD; Christine M Hulette, MD; Bradley T Hyman, MD PhD; Gail P Jarvik, MD PhD; Gregory A Jicha, MD PhD; Lee-Way Jin, MD PhD; Anna Karydas, BA; John SK Kauwe, PhD; Jeffrey A Kaye, MD; Ronald Kim, MD; Edward H Koo, MD; Neil W Kowall, MD; Joel H Kramer, PsyD; Frank M LaFerla, PhD; James J Lah, MD PhD; James B Leverenz, MD; Allan I Levey, MD PhD; Ge Li, MD PhD; Andrew P Lieberman, MD PhD; Chiao-Feng Lin, PhD; Oscar L Lopez, MD; Constantine G Lyketsos, MD MHS; Wendy J Mack, PhD; Daniel C Marson, JD PhD; Frank Martiniuk, PhD; Deborah C Mash, PhD; Eliezer Masliah, MD; Wayne C McCormick, MD MPH; Susan M McCurry, PhD; Andrew N McDavid, BA; Ann C McKee, MD; Marsel Mesulam, MD; Bruce L Miller, MD; Carol A Miller, MD; Joshua W Miller, PhD; John C Morris, MD; Shubhabrata Mukherjee, PhD; Jill R Murrell, PhD, Amanda J Myers, PhD; Sid O’Bryant, PhD; John M Olichney, MD; Vernon S Pankratz, PhD; Joseph E Parisi, MD; Amanda Partch, MS; Henry L Paulson, MD PhD; William Perry, MPH; Elaine Peskind, MD; Ronald C Petersen, MD PhD; Aimee Pierce, MD; Wayne W Poon, PhD; Huntington Potter, PhD; Joseph F Quinn, MD; Ashok Raj, MD; Murray Raskind, MD; Barry Reisberg, MD; Joan S Reisch, PhD; Christiane Reitz, MD PhD; John M Ringman; MD; Erik D Roberson, MD PhD; Ekaterina Rogaeva, PhD; Howard J Rosen, MD; Roger N Rosenberg, MD; Donald R Royall, MD; Mark A Sager, MD; Mary Sano, PhD; Andrew J Saykin, PsyD; Julie A Schneider, MD; Lon S Schneider, MD; William W Seeley, MD; Amanda G Smith, MD; Joshua A Sonnen, MD; Salvatore Spina, MD; Robert A Stern, PhD; Rudolph E Tanzi, PhD; Tricia A Thornton-Wells, PhD; John Q Trojanowski, MD PhD; Juan C Troncoso, MD; Debby W Tsuang, MD; Vivianna M Van Deerlin, MD PhD; Linda J Van Eldik, PhD; Badri N Vardarajan, PhD; Harry V Vinters, MD; Jean Paul Vonsattel, MD; Sandra Weintraub, PhD; Kathleen A Welsh-Bohmer, PhD; Jennifer Williamson, MS; Sarah Wishnek, MPH; Randall L Woltjer, MD PhD; Clinton B Wright, MD MS; Chuang-Kuo Wu, MD PhD; Chang-En Yu, PhD; Lei Yu, PhD. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium Rhoda Au, PhD; Philip A Wolf, MD; Alexa Beiser, PhD; Claudia Satizabal, PhD; Andre G Uitterlinden, PhD; Fernando Rivadeneira, MD, PhD; Peter J Koudstaal, MD, PhD; William T Longstreth, Jr, MD; James T Becker, PhD; Lewis H Kuller, MD; Thomas Lumley, PhD; Kenneth Rice, PhD; Tamara B Harris, MD; Michael Nalls, PhD; Josef JM Marksteiner, MD; Peter Dal-Bianco, MD; Anna Maria Töglhofer, BSc; Paul Freudenberger, MSc; Gerhard Ransmayr, MD; Thomas Benke, MD; Anna M Toeglhofer, MSc; Eric Boerwinkle, PhD; Jan Bressler, PhD; Myriam Fornage, PhD; Francisco J Morón, PhD; Isabel Hernández, MD; Maitee Rosende Roca, MD; Ana Mauleón, MD; Montserrat Alegret, PhD; Reposo Ramírez-Lorca, PhD; Antonio González-Perez, MS. European Alzheimer’s Disease Initiative (EADI) Annick, Alpérovitch, MD, PhD; Victoria Alvarez, PhD; Pascale Barberger-Gateau, MD; Karolien Bettens, PhD; Paola Bossù, MD; Alexis Brice, MD, PhD; Maria Bullido, MD; Paolo Caffara, MD; Jordi Clarimon, MD; Onofre Combarros, MD; Eliecer Coto, PhD; Maria del Zampo, PhD; Marc Delepine, PhD; Maria Candida Deniz Naranjo, PhD; Jacques Epelbaum, PhD; Laura Fratiglioni, MD, PhD; Daniela Galimberti, PhD; Caroline Graff, MD, PhD; Mikko Hiltunen, MD, PhD; Martin Ingelsson, MD, PhD; Lina Keller, MD; Lars Lannfelt, MD; Alberto Llèo, MD; Michelangelo Mancuso, MD; Ignacio Mateo, MD; Patrizia Mecocci, MD; Benedetta Nacmias, PhD; Francesco Panza, MD; Alberto Pilotto, MD; Florentino Sanchez Garcia, MD; Elio Scarpini, MD; Davide Seripa, PhD; Kristel Sleegers, PhD; Hlikka Soininen, MD; Sandro Sorbi, MD; Gianfranco Spalletta, MD; David Wallon, MD, PhD. Genetic and Environmental Risk in Alzheimer's Disease (GERAD) Consortium Charlene Thomas, BSc; Amy Gerrish, PhD; Jade Chapman, PhD; Alexandra Stretton, PhD; Angharad Morgan; Harriet Oldham, PhD; Michael J Owen, MD, PhD; Patrick G Kehoe, PhD; Christopher Medway, PhD; Kristelle Brown, PhD; Jenny Lord; James Turton; Nigel M Hooper; Emma Vardy; Jason D Warren; Jonathan M Schott; James Uphill; Paul Hollingworth, PhD, DClinPsy; Natalie Ryan; Martin Rossor, MD PhD; John Collinge, MD, PhD; Yoav Ben-Shlomo; Daniilidou Makrina; Olymbia Gkatzima; Michelle Lupton, PhD; Maria Koutroumani; Despoina Avramidou; Antonia Germanou; Frank Jessen, PhD; Steffi Riedel-Heller; Martin Dichgans, MD, PhD; Reiner Heun; Heike Kölsch; Britta Schürmann, PhD; Christine Herold; André Lacour; Dmitriy Drichel; Per Hoffmann; Johannes Kornhuber, MD, PhD; Wei Gu; Thomas Feulner; Manuel Mayhaus, PhD; Sabrina Pichler, PhD; Matthias Riemenschneider, MD; Hendrik van den Bussche, PhD; Brian Lawlor, MD, PhD; Aoibhinn Lynch, MD; David Mann, PhD; A David Smith, DPhil; Donald Warden; Gordon Wilcock; Isabella Heuser, PhD; Jens Wiltfang, MD, PhD; Lutz Frölich, PhD; Michael Hüll, MD, PhD; Kevin Mayo, PhD; Gill Livingston, MD, PhD; Nicholas J Bass, MD, PhD; Hugh Gurling, MD, PhD; Andrew McQuillin, PhD; Rhian Gwilliam; Panagiotis Deloukas; Ammar Al-Chalabi, MB ChB; Christopher E Shaw, MD, PhD; Andrew B Singleton, PhD; Rita Guerreiro, PhD; Giancarlo Russo; Karl-Heinz Jöckel, PhD; Susanne Moebus, PhD; Norman Klopp, PhD; H-Erich Wichmann, MD, PhD; Dennis W Dickson; Neill R Graff-Radford; Li Ma; Gina Bisceglio; Elizabeth Fisher, PhD; Nick Warner MB ChB; Stuart Pickering-Brown, PhD; David Craig, MD; Janet A Johnston, PhD; Bernadette McGuinness, MD, PhD; Stephen Todd, MD, PhD; David C Rubinsztein, MB ChB, PhD; Simon Lovestone, PhD; Anthony Bayer, MD; John Gallacher, MD; Petroula Proitsi, PhD; Sara Ortega-Cubero, MD.
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Jun, G., Ibrahim-Verbaas, C., Vronskaya, M. et al. A novel Alzheimer disease locus located near the gene encoding tau protein. Mol Psychiatry 21, 108–117 (2016). https://doi.org/10.1038/mp.2015.23
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.23
- Springer Nature Limited
This article is cited by
-
Inflammation and neuronal gene expression changes differ in early versus late chronic traumatic encephalopathy brain
BMC Medical Genomics (2023)
-
Mapping the genetic landscape of early-onset Alzheimer’s disease in a cohort of 36 families
Alzheimer's Research & Therapy (2022)
-
Genomics and Functional Genomics of Alzheimer's Disease
Neurotherapeutics (2022)
-
FMNL2 regulates gliovascular interactions and is associated with vascular risk factors and cerebrovascular pathology in Alzheimer’s disease
Acta Neuropathologica (2022)
-
Meta-analysis of genome-wide association studies identifies ancestry-specific associations underlying circulating total tau levels
Communications Biology (2022)