Abstract
Chitinases are promising enzymes for a multitude of applications, including chitooligosaccharide (COS) synthesis for food and pharmaceutical uses and marine waste management. Owing to fungal diversity, fungal chitinases may offer alternatives for chitin degradation and industrial applications. The rapid reproduction cycle, inexpensive growth media, and ease of handling of fungi may also contribute to reducing enzyme production costs. Thus, this study aimed to identify fungal species with chitinolytic potential and optimize chitinase production by submerged culture and enzyme characterization using shrimp chitin. Three fungal species, Coriolopsis byrsina, Trichoderma reesei, and Trichoderma harzianum, were selected for chitinase production. The highest endochitinase production was achieved in C. byrsina after 168 h cultivation (0.3 U mL− 1). The optimal temperature for enzyme activity was similar for the three fungal species (up to 45 and 55 ºC for endochitinases and exochitinases, respectively). The effect of pH on activity indicated maximum hydrolysis in acidic pH (4–7). In addition, the crude T. reesei extract showed promising properties for removing Candida albicans biofilms. This study showed the possibility of using shrimp chitin to induce chitinase production and enzymes that can be applied in different industrial sectors.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The constant increase in the industrial shellfish processing generates approximately 1012–1014 tons chitinous waste per year [1]. Chitin is the most common biopolymer in the marine environment and its disposal, whether through ocean dumps, incineration, or landfills, contributes to natural resource wastage, economic losses, and environmental pollution [2].
Chitin is found in crustacean shells, insects, fungal cell walls, and mollusks. Crustaceans (shrimp and crabs) are the most critical sources of chitin and usually contain proteins (30–40%), CaCO3 (30–50%), chitin (20–30%), and several other compounds. Insects contain > 30% chitin in their exoskeletons and may be an alternative source of chitin [1, 3].
Chitin is a polysaccharide composed of β(1,4)-N-acetylglucosamine units. Chitinases catalyze the cleavage of glycosidic bonds in chitin [4]. They comprise a diverse group of enzymes with varied structures and mechanisms that determine their activity and suitability for applications such as chitooligosaccharide (COS) synthesis for use in the food and pharmaceutical industries, marine waste management, biocontrol of insect pests and pathogenic fungi, and biofuel production [5].
Currently, examples of commercial chitinases include Chitinase from Aspergillus niger (food grade)/Creative Enzymes®, Native Trichoderma viride Chitinase/Creative Enzymes®, Native Streptomyces griseus Chitinase/Creative Enzymes®, Chitinase from Streptomyces griseus/Merck (Sigma-Aldrich) and Chitinase (Clostridium thermocellum)/Megazyme [6]. Despite their significant biotechnological potential, chitinases have not been extensively exploited commercially to the same extent as other glycosidases. This limited commercial utilization is due to factors such as the low number of organisms that show high chitinase production rates, low enzyme activity and stability, and high cost of production [7].
Understanding the effect of culture medium composition on microbial growth is essential to optimize enzyme production. Considering the importance of bioconverting chitin-rich biomasses and the role of chitinases in developing new value-added products, in this work, we evaluated the composition of the submerged medium for chitinase production by fungi, in addition to biochemical studies of these enzymes and their ability to remove Candida biofilms.
Materials and methods
Microorganisms
The fungi used for chitinase production, Trichoderma reesei, Trichoderma harzianum, and Coriolopsis byrsina, were obtained from a collection of microorganisms at the Laboratory of Biochemistry and Applied Microbiology, UNESP, São José do Rio Preto/SP, Brazil.
Colloidal chitin preparation
To prepare colloidal chitin, 10 g shrimp chitin (Sigma Aldrich) was added to 200 mL 36–37% HCl and constantly agitated for 4 h at 4 ºC. Then, 4 L cold distilled water was added and the mixture was centrifuged at 4 ºC and 10,000 rpm for 10 min. Several washing and centrifugation cycles were performed until the pH 7.0 was reached. The chitin was dried completely at 80 °C, crushed into a powder, sieved through a 425 mm sieve to remove larger granules, and subsequently stored at − 20 ºC.
Submerged culture
First, to obtain biomass for seeding the submerged media, a pre-inoculum of each fungus was used. The fungi were inoculated in 125 mL Erlenmeyer flasks containing 30 mL of potato dextrose agar (PDA) medium and incubated at 30 ºC for 5 days. Next, 12 mL sterile distilled water was added to the PDA flasks. Using a Neubauer chamber for spore counting, 1 × 107 spores mL− 1 submerged medium were added to the fermentative flasks.
The optimal composition of the culture medium was evaluated and four different formulations were tested. The minimal medium contained 0.7% KH2PO4, 0.2% K2HPO4, 0.01% MgSO4, 0.01% CaCl2, and 0.5% colloidal chitin: 1- Minimal medium (C + minimal medium); 2- Minimal medium with 0.1% yeast extract (C + Y); 3- Minimal medium with 0.1% yeast extract and 0.1% peptone (C + Y + P); 4- Minimal medium with 0.1% yeast extract and 0.1% ammonium sulfate (C + Y+(NH4)2SO4). Where C (chitin), Y (yeast extract), and P (peptone).
The volume of the culture medium was 50 mL (250 mL Erlenmeyer flask), pH 6.0, and the flasks were incubated at 30 ºC for 10 days (240 h). One flask was harvested every 24 h, filtered using Whatman® qualitative filter paper (grade 1), and the fermentative extract was used to quantify the chitinolytic activity (endo and exochitinase). After determining the best conditions for enzyme production, another larger-scale culture (1 L) was performed to produce enough enzymatic extract for biochemical characterization and application.
The extract was concentrated using a tangential membrane filtration system with a 10 kDa cut-off filter membrane, surface area 420 cm2 (Hollow Fiber cartridge, Model UFP-10-E-4MA, GE Healthcare).
Enzyme activity assay
The reaction mixture for measuring endochitinase activity (0.01 g colloidal chitin, 100 µL enzyme, and 1 mL 0.2 mol L− 1 sodium acetate buffer, pH 5.0) was incubated at 45 °C for 20 h with stirring at 180 rpm. Next, 100 µL reaction mixture was added to 100 µL dinitrosalicylic acid (DNS) solution and incubated at 95 °C for 10 min to quantify the reducing sugars. The absorbance at 540 nm was measured using a spectrophotometer. A standard curve with N-acetyl-D-glucosamine as the reference was used for determining the activity. One unit of enzyme activity was defined as the amount of enzyme required to produce 1.0 µmoL of N-acetyl-D-glucosamine mL− 1 h− 1 under the assay conditions.
We used p-nitrophenyl-N-acetyl-β-D-glucosaminide as the substrate for measuring the exochitinase activity; the reaction mixture comprised 5 µL enzyme and 90 µL 4 mmol L− 1 substrate dissolved in 0.2 mol L− 1 sodium acetate buffer (pH 5.0). The reaction was carried out in a microplate for 15 min at 45 ºC. Then, 100 µL 2 mol L− 1 sodium carbonate was added and the enzymatic activity was measured at 410 nm. A standard curve with p-nitrophenol as the reference was used to determine the activity. One unit of enzyme activity was defined as the amount of enzyme required to produce 1.0 µmoL of p-nitrophenol mL− 1 min− 1 under the assay conditions.
Functional biochemical characterization of the crude extract
Effect of pH and temperature on enzyme activity and stability
The optimal pH for enzyme activity was determined at 45 °C using 0.2 mol L− 1 acetate (pH 4.0 and 5.0), MES (pH 5.5, 6.0, and 6.5), HEPES (pH 7.0 and 7.5), bicine (pH 8.0, 8.5, and 9.0), and CAPS (pH 9.5 and 10.0) buffers. The effect of temperature on enzyme activity was investigated at 30–60 ºC.
The thermal stability of the extracts was studied after incubating them at 30–70 ºC for 1 h. While the effect of pH on stability was studied after incubating the extracts at pH 4.0–10.0 for 1–24 h at 4 °C. In both cases, after incubation, the enzyme reaction was carried out at the optimum pH and temperature for determining their activity.
Effect of metal ions on enzyme activity
The enzyme activity in the presence of 5 mmol L− 1 (final concentration) iron III (FeCI3), cadmium (CdCl2), barium (BaCl2), calcium (CaCl2), cobalt (CoCl2), lithium (LiCl), magnesium (MgCl2), manganese (MnCl2), nickel (NiSO4), and zinc (ZnSO4) was determined. In all the tests, the enzymes were pre-incubated with their respective salts for 5 min at room temperature. The reactions were performed under optimal pH and temperature conditions for each extract.
Hydrolysis of different substrates
In addition to chitinase activities, β–xylosidase, β–glucosidase, and α–L-arabinofuranosidase activities were determined using 10 µL enzyme extract with 90 µL 0.2 mol L− 1 sodium acetate buffer solution (pH 5.0) with 4 mmol L− 1 dissolved substrate (p-nitrophenyl-β-D-xylopyranoside [pNPX; Sigma Aldrich], p-nitrophenyl-β-D-glucopyranoside [pNPG], and p-nitrophenyl-α-L-arabinofuranoside [pNPA, Sigma Aldrich] for β–xylosidase, β–glucosidase, and α–L-arabinofuranosidase, respectively). The reaction mixtures were incubated at 40 °C for 30 min and then interrupted with 100 µL 2 mol L− 1 Na2CO3. The released p-nitrophenol was quantified spectrophotometrically at 410 nm. A standard curve with p-nitrophenol as the reference was used for determining enzyme activity. One unit of enzyme activity was defined as the amount of enzyme required to produce 1.0 µmoL of p-nitrophenol mL− 1 min− 1 under the assay conditions.
For carboxymethylcellulose (CMC) and β-1,3-glucan, the DNS method was used to quantify reducing sugars. The reaction mixture, comprising 10 µL enzyme extract and 90 µL substrate (1% in 0.2 mol L− 1 acetate buffer pH 5.0), was incubated for 30 min at 40 ºC. Then, the reaction mixture was incubated with 100 µL DNS at 95 °C for 10 min. The amount of reducing sugar released was quantified at 540 nm using a spectrophotometer (SPECTRAmax Plus 384). A standard curve with D-glucose as the reference was used to determine the reducing sugar released. One unit of enzyme activity was defined as the amount of enzyme required to produce 1.0 µmoL of glucose mL− 1 h− 1 under the assay conditions.
To evaluate the caseinolytic activity, 50 µL enzyme extract was incubated with 500 µL 1% casein diluted in 0.2 mol L− 1 sodium phosphate buffer (pH 6.5) and incubated at 40 °C for 30 min. Next, the reaction was interrupted by adding 300 µL 10% of trichloroacetic acid (TCA). Blank tubes were prepared by adding 10% TCA to the reaction mixture before adding the substrate. The reaction tubes and blank tubes were then centrifuged at 10,000× g for 10 min at 25 °C, the supernatant was collected, and the absorbance was measured spectrophotometrically at 280 nm. Caseinolytic activity was expressed as U mL− 1, where one unit of enzyme activity (U) was defined as the amount of enzyme required to increase the absorbance (A280nm) by 0.01 per minute under the assay conditions [8].
Biofilm removal
The experiment was conducted as described by Menezes et al. [9] with some modifications. The yeast Candida albicans ATCC 90028 (initial inoculum: 0.1 at A600nm) was cultivated in a sterile 6-well polystyrene plate containing 1.5 mL yeast extract peptone dextrose (YPD) at 30 ºC for 5 days under static conditions for biofilm formation. Then, the culture medium was removed, the plates were washed twice with sterile water, and 2 mL enzyme extract from the three fungal species and 1 mL 0.2 mol L− 1 sodium acetate buffer (pH 5.0) was added. The enzyme extracts were previously filtered using 0.22 μm syringe filter.
The enzyme solution used for biofilm removal was a pool of the fermentative extract collected at the best enzyme production time for each species; in this experiment the sample presented 490 µg mL− 1 total protein and 0.85 U mL− 1 (chitinolytic activity); an experiment containing the heat-inactivated enzyme was used as a control. Then, we removed the enzyme solution, washed the plates with water, added 3 mL 1% (w/v) crystal violet, and incubated for 20 min at 25 ºC. Next, we removed the stain, washed the plates with water, and dried them by incubation for 1 h at 40 ºC. We imaged biofilm removal and quantified the biofilm by adding 3 mL 30% (v/v) acetic acid solution, followed by measuring the absorbance in a spectrophotometer at 595 nm. The control experiment (denatured enzyme) was considered as 100%.
Total protein quantification
This test was performed using the Bradford method [10] to quantify the microbial extract proteins in. A standard curve with bovine serum albumin was used to determine the protein concentration at 595 nm.
Data analysis
All measurements were performed with at least three independent replicates, and the control experiment was performed under the same conditions. Experimental results were expressed as the mean of replicate determinations and standard deviations (mean ± SD).
For the ion interference tests, statistical significance was assessed using one-way analysis of variance (ANOVA), followed by Dunnett’s test. Results were considered statistically significant at P ≤ 0.05. Statistical analyses were performed using IBM SPSS Statistics 20 and GraphPad Prism 9 softwares.
Results
Endo and exochitinase production
We tested four different culture media formulations to evaluate the profile of fungal chitinase production. After 168 h of cultivation (Fig. 1a), C. byrsina showed a remarkable increase in endochitinase production in C + Y+(NH4)2SO4, C + Y + P, and C + Y (0.3 U mL− 1). The best production time for C + minimal medium was 240 h (0.179 U mL− 1), which was considerably different from that for the other media used.
Submerged culture with colloidal chitin as a substrate in different culture media. a, b, and c show graphs for endochitinase activity, whereas d, e, and f show graphs for exochitinase activity. Coriolopsis byrsina: (a) endochitinase activity, (d) exochitinase activity; Trichoderma harzianum: (b) endochitinase activity, (e) exochitinase activity; Trichoderma reesei: (c) endochitinase activity, and (f) exochitinase activity
For the four culture media tested, the production of endochitinase by C. byrsina was low between 24 h and 72 h, increasing considerably from 120 h for C + Y + P and C+(NH4)2SO4, while for C + Y, enzyme production only increased after 144 h. After 168 h, chitinase production remained stable until 240 h in C + Y + P and C + Y+(NH4)2SO4(Fig. 1a). We observed that exochitinase production gradually increased with fungal growth, with the highest activity at 240 h (Fig. 1d). The enzyme activity profile was similar for C + Y + P and C + Y+(NH4)2SO4, these had the best enzyme productivity (2 U mL− 1).
T. harzianum produced the maximum endochitinase (0.280 U mL− 1) in C + Y + P after 240 h (Fig. 1b). The minimum medium showed a low endochitinase yield (0.05 U mL− 1). This value was also lower than that of C. byrsina and similar to that observed of T. reesei.
Compared to the other fungi, T. reesei showed the lowest endochitinase activity. A similar profile was observed in all media, with the maximum activity observed at 192 h (0.183 U mL− 1) in C + Y + P (Fig. 1c). It showed strong exochitinase activity in all tested media, reaching approximately 7 and 5 U mL− 1 in C + Y + P and C + Y, respectively, at 240 h (Fig. 1f).
Functional biochemical characterization of endo and exochitinases
We used the C. byrsina extract cultivated in C + Y + P for 168 h. The maximum endochitinase activity was observed between pH 5 and 6.5. The activity decreased as the pH increased, and the activity reached 50% at pH 7, gradually decreasing at alkaline pH levels (Fig. 2a). The endochitinase remained stable at all pH levels when incubated for 1–24 h. The optimum temperature for activity was noted between 40 ºC and 45 °C. The enzyme activity sharply dropped at higher temperatures and was not detected at 55 ºC. Endochitinase was stable up to 45 ºC, considering that its activity at 50 ºC was reduced to less than 30% (Fig. 2d).
We used the T. harzianum extract cultivated in C + Y + P for 144 h. T. harzianum endochitinase was more active between pH 5 and 6.5 and 45 ºC. The enzyme remained stable from pH 5 to 10, with more than 70% and 60% residual activity after 1 and 24 h, respectively (Fig. 2b). The endochitinase was also thermostable up to 45 ºC, with approximately 70% residual activity. After incubation at 50 ºC for 1 h, the enzyme lost approximately 90% activity (Fig. 2e).
We used the T. reesei extract cultivated in C + Y + P for 192 h. The optimum pH of the T. reesei endochitinase was the most acidic. Maximum activity was reached at pH 4.5-5. Enzyme stability decreased as pH increased under the two tested conditions (1 and 24 h). The highest stability was observed at pH 4–5 and 61% activity was retained at pH 10 (Fig. 2c). The endochitinase was stable up to 45 ºC, with > 80% activity, but the enzyme performance drops abruptly at 50 ºC, as with the other endochitinases studied (Fig. 2f).
The maximum exochitinase activity for C. byrsina extract was detected at pH 5.5–7. Exochitinases remained stable over a wide pH range (5–10), with > 60% residual activity at both 1 and 24 h (Fig. 3a). Under the effect of temperature (Fig. 3d), a sharp peak for activity at 55 ºC and stability up to 50 ºC (100% residual activity) were observed. However, the enzyme did not tolerate prolonged exposure to temperatures above 50 ºC, which considerably decreased the enzyme activity at 55 ºC; no activity was detected above 60 ºC (Fig. 3d).
The exochitinase activity of T. harzianum extract was more pronounced at 55 ºC and between pH 5.5–7. The enzyme remained stable over pH 4–10, maintaining > 80% and > 60% activity for 1 and 24 h, respectively (Fig. 3b). The enzyme was stable up to 45 ºC with relative activity above 80%; after incubating for 1 h at 55 ºC, it had < 10% activity (Fig. 3e).
The optimal enzyme activity of T. reesei extract was detected at 55 ºC and between pH 4.5–5.5. Moreover, the enzyme remained stable between pH 4–9, with > 70% activity after incubation for both 1 and 24 h (Fig. 3c). The enzyme maintained 40% activity up to 40 ºC, gradually decreasing at other temperatures, with no activity at 70 ºC (Fig. 3f).
Interference of metal ions on the enzyme activity
The highest positive modulation of endochitinase activity was observed with Mn2+. Ions Mg2+, Fe3+, Li+, Zn2+, Ca2+ and Ni2+ did not significantly affect the endochitinase activity of C. byrsina and T. harzianum extracts (Table 1). However, Co2+, Ba2+ and Mn2+ improved the endochitinase activity of C. byrsina extract (by 19%, 14%, and 78%, respectively) and T. harzianum extract (by 15%, 11%, and 32%, respectively). Almost all metal ions tested, except Mg2+ and Fe3+, positively modulated the endochitinase activity of T. reesei extract (Zn2+: +9%; Ni2+: +12%; Ca2+: +15%; Cd2+: +21%; Ba2+: +23%; Li+: +26%; Co2+: +30%; and Mn2+: +56%).
The main positive and negative modulation of the exochitinase activity of C. byrsina extract was caused by Fe3+ (+ 53%) and Li+ (− 16%), respectively (Table 2). Cd2+ (+ 7%), Mn2+ (+ 9%), and Ba2+ (+ 16%) also improved exochitinase activity. Fe3+ and Ni2+ decreased (approximately 18%), while Mg2+ (+ 26%), Co2+ (+ 34%), and Ba2+ (+ 18%) increased the catalytic activity of T. harzianum fermentative extract. The exochitinase activity of T. reesei extract was considerably reduced in the presence of Zn2+ (− 20.5%) and increased in the presence of Cd2+ (+ 12%), Co2+ (+ 21%), Ni2+ (+ 33%), Mg2+ (+ 55%), Li+ (+ 66%), and Fe3+ (+ 190%).
Hydrolysis of different substrates
Enzyme activities of C. byrsina, T. harzianum, and T. reesei fermentative extracts on different substrates are summarized in Table 3. For comparison, we also tested the activity of the commercial T. harzianum enzyme (Sigma Aldrich). In addition to chitinase activity, we mainly detected pNP-arabinofuranoside, pNP-Glucopyranoside, β-1,3-Glucan, Carboxymethylcellulose and casein hydrolysis.
The T. reesei extract did not show significant activity against pNPA, pNPX or pNPG, but it showed the best exochitinase activity (11.53 U mg− 1). The T. harzianum enzyme extract exhibited the highest activity against casein and colloidal chitin (121.04 and 27.12 U mg− 1, respectively).
C. albicans biofilm removal
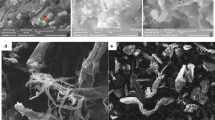
Given that the fermentation extracts of the three fungal species exhibited significant activities on various substrates, including pNP-Glucopyranoside, pNP-Arabinofuranoside, pNP-Glucosaminide, β-1,3-Glucan, Carboxymethylcellulose, colloidal chitin and casein, we subsequently assessed the effectiveness of these fermentative extracts in dispersing Candida albicans biofilm. Only the extract from the T. reesei showed the ability to remove biofilms. In plates treated with the inactive enzyme (control), we observed intense crystal violet coloration, while in the presence of active enzymes we observed a decrease in staining intensity. After 5 h of incubation, the biofilm removal was 46%, and at 20 h, it was approximately 79%. In the Fig. 4 it is possible to visualize the removal of the biofilm after enzymatic treatment. The mean triplicate values and SD are shown.
Candida albicans biofilm dispersion after treatment with Trichoderma reesei enzyme extract (490 µg mL− 1 final protein concentration and 0.85 U mL− 1 chitinolytic activity) for 5 h and 20 h. Relative absorbance (595 nm) of biofilms stained with 1% crystal violet solubilized with 30% glacial acetic acid. (a) Mean triplicate values and standard deviation (SD) are shown, and (b) visualization of biofilm removal after enzyme treatment
Discussion
Chitinases have recently gained increasing attention due to their wide application potential in various fields, particularly chitin bioconversion and biocontrol in agriculture [11]. Among chitinase-producing microorganisms, Trichoderma spp. are of great industrial interest and have long been used in agriculture as biological controls [12].
Since phytopathogens and insects are considered the greatest threats to important crops (wheat, rice, corn, and potatoes), the abusive use of chemical pesticides against these agricultural pests has been practiced for years [13]. This has caused worldwide concern regarding soil and groundwater contamination as well as human health [14, 15].
However, suitable ecofriendly, biodegradable, and economical bioproducts can be found presently. The presence of chitin in insects, fungi, and nematodes makes them logical targets for chitinases, which can function as biopesticides and are harmless to plants and vertebrates that lack chitin in their tissues [13].
Further studies are needed to enhance chitinase production at the laboratory scale. Optimizing the culture medium for producing specific enzymes is essential to not only maximize yield, but also minimize production costs, as the culture medium components can enhance growth and accelerate microbial metabolism [16].
Colloidal chitin has been widely used as an inducer for chitinase production in several studies, including the expression of chitinases from Acremonium sp. YS2-2, Trichoderma asperellum PQ34, Myxococcus fulvus UM01, and Achromobacter xylosoxidans in response to 1% colloidal chitin [17]. In addition to chitin, other nitrogen sources increased the production of chitinases by Glutamicibacter uratoxydans up to eight times [18].
Although yeast extract is an alternative vitamin, mineral, and amino acid sources, culturing Aspergillus niveus in minimal medium with crab shell chitin (96 h, 30 °C, 100 rpm), without yeast extract, had the best production of chitinase (6.5 U mg− 1 protein). In minimal medium (in the absence of yeast extract), fungal metabolism is directed toward chitin degradation, which is the only available carbon/nitrogen source [19]. In another study, the production of chitinase by Aspergillus terreus was also influenced by the source of nitrogen incorporated into the medium, and the highest chitinase activity (6.28 U mg− 1 of protein) was recorded from growth in a medium containing ammonium sulfate at 1% [20].
Typically, chitinase activity is optimal under acidic conditions. The optimal temperature and thermal stability of mesophilic fungi are similar to those found in this report (40–65 ºC). The optimal endochitinase and exochitinase activities described for the three fungal species are between pH 4–7 and up to 45 and 55 °C for endochitinases and exochitinases, respectively.
Recent reports on chitinases from different fungal species have focused on producing and determining the functional characteristics of chitinases toward colloidal chitin, including the enzyme from Thermomyces lanuginosus, which exhibited maximum activity at pH 3–4 and 50 °C, with gradual decreased activity after pH 6.0 [21]. The maximum activity for recombinant chitinase expressed in Escherichia coli (gene from Aspergillus fumigatus df347) was at pH 5 and 45 °C [22]. The chitinase purified from Penicillium oxalicum k10 showed maximum activity (100%) at pH 5 and 40 °C. The enzyme was stable up to 40 °C, with > 90% activity at 40 °C for 60 min [17]. Study with thermostable and acidic chitinase from Paecilomyces thermophila J18 (expressed in Pichia pastoris) reported optimal enzyme activity at pH 5.5 and 60 °C, and stability within pH 3.5–9.0 at 45 °C for 30 min (more than 70% residual activity) and approximately 90% residual activity up to 55 °C for 30 min [23].
Metal ions play a significant role in biological catalysis by forming complexes with proteins and interfering with enzyme structure and activity. Endochitinase activity mainly increased in the presence of MnCl2 in all three fungal species. Chitinase activity of Aspergillus niveus also increased in the presence of MnCl2 (by approximately 122%) than that in the control [19]. In the presence of 10 mM Zn2+ and Mn2+, the activity of A. terreus chitinase decreased to 46.81% and increased by 25.62%, respectively [24]. Deng et al. [25] reported that Zn2+ ions decreased the enzymatic activity of Trichoderma harzianum GIM 3442 by 49.4%. In contrast, the enzyme activity of chitinase from Penicillium oxalicum k10 increased by Zn2+ and K+, while Ag+ and Fe2+ decreased the chitinolytic activity to 65.9% and 63.9%, respectively. The addition of Cu2+ decreased chitinolytic activity by 60% [17].
The enzyme activity profile of the three fungal culture extracts highlights the synergistic role of chitinases, in addition to the presence of other enzymes, particularly proteases and β-1,3-glucanases, in trace amounts. A broad spectrum of enzymatic actions may also play an important role in biocontrol mechanisms [26]. In the context of fungal biofilms, chitinases are capable of degrading the chitin present in fungal cell walls [17]. When used in conjunction with proteolytic enzymes, which degrade proteins that are components of extracellular polymeric substances (EPS), this cocktail of enzymes is potentially effective in reducing the adhesion of microbial cells to surfaces and improving the accessibility of antimicrobial agents to the biofilm [9, 27].
In the study with Metschnikowia species tested as biocontrol agents against postharvest fungal decays on lemons, different enzymes such as chitinase, protease, pectinase and β-1,3 glucanase were detected. Among the tested yeasts, Metschnikowia aff. fructicola was the most antagonistic against the phytopathogens Penicillium digitatum and P. expansum [28].
In addition to their biocontrol applications in agriculture, enzymes are also promising agents for biofilm dispersion. We must highlight that Candida spp. infections are emerging as major health problems, with high mortality rates and increasing medical costs for governments and hospitalized patients. Mortality can be attributed to the increasing incidence of invasive systemic candidiasis and septicemia, particularly in immunocompromised patients [29].
We found that T. reesei fermentative extract was useful for removing C. albicans biofilms. Commercial enzymes, including chitinases, lipases, proteases, DNAse and lyticase (final enzyme concentration: 50 µg mL− 1) were also tested for removing C. albicans biofilms [27]. The same study evaluated C. tropicalis biofilm removal for 2 h at 37 or 25 °C. Under these conditions, biofilm detachment by chitinase treatment was approximately 23% for C. albicans and 29% for C. tropicalis. Lyticase exhibited the best biofilm removal with 85% and 54% biofilm dispersion for C. albicans and C. tropicalis, respectively. In future studies, tests combining T. reesei extract with other enzymes, such as DNAse and lyticase, as well as antibiotics, should be carried out to improve the effect on biofilm removal.
Conclusions
The chitinases secreted by the fungal species T. reesei, T. harzianum and C. byrsina are promising for chitin degradation, mainly due to their stability at different pH levels, temperatures up to 45 °C, and in the presence of various metallic ions. Chitinases from Trichoderma are well described, but we did not find any reports on chitinases from C. byrsina.
Fungal chitinases are proficient candidates for producing value-added chitin degradation products such as COS and GlcNAc. We also demonstrated the potential of the fermentative extract of T. reesei for removing Candida biofilms, which is an important finding given the emerging need to study environment-friendly compounds capable of removing biofilms.
References
Lv J et al (2023) Chitin and chitin-based biomaterials: a review of advances in processing and food applications. Carbohydr Polym 299:120142. https://doi.org/10.1016/j.carbpol.2022.120142
Chakravarty J, Edwards TA (2022) Innovation from waste with biomass-derived chitin and chitosan as green and sustainable polymer: a review. Energy Nexus 8:100149. https://doi.org/10.1016/j.nexus.2022.100149
Khajavian M et al (2022) Chitin and derivative chitosan-based structures — Preparation strategies aided by deep eutectic solvents: a review. Carbohydr Polym 275:118702. https://doi.org/10.1016/j.carbpol.2021.118702
Younes I, Rinaudo M (2015) Chitin and Chitosan Preparation from Marine sources. Structure, Properties and Applications. Mar Drugs 13:1133–1174. https://doi.org/10.3390/md13031133
Oyeleye A, Normi YM (2018) Chitinase: diversity, limitations, and trends in engineering for suitable applications. Biosci Rep 38:4. https://doi.org/10.1042/BSR20180323
Poria V et al (2021) Current perspectives on Chitinolytic Enzymes and their agro-industrial applications. Biology 10:1319. https://doi.org/10.3390/biology10121319
Baldoni DB et al (2020) Chitinase production by Trichoderma Koningiopsis UFSMQ40 using solid state fermentation. Braz J Microbiol 51:1897–1908. https://doi.org/10.1007/s42770-020-00334-w
Duffeck CE et al (2020) Citrobacter diversus-derived keratinases and their potential application as detergent-compatible cloth-cleaning agents. Braz J Microbiol 51:969–977. https://doi.org/10.1007/s42770-020-00268-3
Menezes CLA et al (2023) The degradation of chicken feathers by Ochrobactrum intermedium results in antioxidant and metal chelating hydrolysates and proteolytic enzymes for staphylococcal biofilm dispersion. 3 Biotech 13(6):202. https://doi.org/10.1007/s13205-023-03619-7
Bradford MM (1976) A rapid and sensitive for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Zhang W et al (2021) Biochemical characterization of a novel acidic chitinase with antifungal activity from Paenibacillus xylanexedens Z2–4. Int J Biol Macromol 182:1528–1536. https://doi.org/10.1016/j.ijbiomac.2021.05.111
del Urbina-Salazar A R et al (2018) Chitinase production by Trichoderma Harzianum grown on a chitin-rich mushroom byproduct formulated medium. Waste Biomass Valoriz 10:2915–2923. https://doi.org/10.1007/s12649-018-0328-4
Silva RR, Santos RC (2019) Pest Control: can Chitinases help to reduce Pesticide use? J Agric Food Chem 67(29):8071–8073. https://doi.org/10.1021/acs.jafc.9b03219
Singh G, Arya SK (2019) Antifungal and insecticidal potential of chitinases: a credible choice for the eco-friendly farming. Biocatal Agric Biotechnol 20:101289. https://doi.org/10.1016/j.bcab.2019.101289
Rajput M et al (2022) Myco-chitinases as versatile biocatalysts for translation of coastal residual resources to eco-competent chito-bioactives. Fungal Biol Rev 41:52–69. https://doi.org/10.1016/j.fbr.2022.04.001
Akeed Y et al (2020) Partial purification and characterization of chitinase produced by Bacillus licheniformis B307. Heliyon 6:03858. https://doi.org/10.1016/j.heliyon.2020.e03858
Xie X-H et al (2021) A broad-specificity chitinase from Penicillium Oxalicum k10 exhibits antifungal activity and Biodegradation properties of Chitin. Mar Drugs 19:356. https://doi.org/10.3390/md19070356
Asif T et al (2019) Bioconversion of Colloidal Chitin using Novel Chitinase from Glutamicibacter uratoxydans exhibiting anti-fungal potential by Hydrolyzing Chitin within Fungal Cell Wall. Waste Biomass Valoriz 11:4129–4143. https://doi.org/10.1007/s12649-019-00746-2
Alves TB et al (2018) Production and characterization of a thermostable antifungal chitinase secreted by the filamentous fungus aspergillus niveus under submerged fermentation. 3 Biotech 8:369. https://doi.org/10.1007/s13205-018-1397-6
Farag AM et al (2014) Production, optimization, characterization and antifungal activity of chitinase produced by aspergillus terrus. Afr J Biotechnol 13(14):1567–1578. https://doi.org/10.5897/AJB2014.13628
Suryawanshi N, Eswari JS (2022) Purification and characterization of chitinase produced by thermophilic fungi Thermomyces Lanuginosus. Prep Biochem Biotechnol 52:1087–1095. https://doi.org/10.1080/10826068.2022.2028639
Wu Y-L et al (2022) The Discovery, Enzymatic characterization and functional analysis of a newly isolated chitinase from Marine-Derived Fungus Aspergillus Fumigatus df347. Mar Drugs 20:520. https://doi.org/10.3390/md20080520
Han S et al (2024) High level production of a novel acidic and thermostable chitinase from Paecilomyces Thermophila for the extraction of bioactive components from Ganoderma lucidum spores. Process Biochem 136:182–190. https://doi.org/10.1016/j.procbio.2023.11.023
Farag AM et al (2016) Purification, characterization and antimicrobial activity of chitinase from marine-derived aspergillus terreus. Egypt J Aquat Res 42(2):185–192. https://doi.org/10.1016/j.ejar.2016.04.004
Deng J-J et al (2019) Heterologous expression and characterization of an antifungal chitinase (Chit46) from Trichoderma Harzianum GIM 3.442 and its application in colloidal chitin conversion. Int J Biol Macromol 134:113–121. https://doi.org/10.1016/j.ijbiomac.2019.04.177
Silva RR (2019) Enzyme technology in food preservation: a promising and sustainable strategy for biocontrol of post-harvest fungal pathogens. Food Chem 277:531–532. https://doi.org/10.1016/j.foodchem.2018.11.022
Al-Fattani MA, Douglas LJ (2006) Biofilm matrix of Candida albicans and Candida tropicalis: chemical composition and role in drug resistance. Med Microbiol 55(8):999–1008. https://doi.org/10.1099/jmm.0.46569-0
Oztekin S, Karbancioglu-Guler F (2021) Bioprospection of Metschnikowia sp. isolates as biocontrol agents against postharvest fungal decays on lemons with their potential modes of action. Postharvest Biol Technol 181:111634. https://doi.org/10.1016/j.postharvbio.2021.111634
Pereira R et al (2020) Biofilm of Candida albicans: formation, regulation and resistance. J Appl Microbiol 131(1):11–22. https://doi.org/10.1111/jam.14949
Funding
Fundação de Amparo à Pesquisa do Estado de São Paulo-FAPESP (process 2017/06399-3).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Julio Santos.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
de Menezes, C.L.A., Boscolo, M., da Silva, R. et al. Fungal endo and exochitinase production, characterization, and application for Candida biofilm removal. Braz J Microbiol 55, 2267–2277 (2024). https://doi.org/10.1007/s42770-024-01432-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-024-01432-9