Abstract
Sustainable agriculture relies on the use of herbicides to preserve soil carbon and minimize disturbance to the soil structure. Glyphosate and atrazine, widespread and frequently used herbicides in South America, can affect soil microbial populations involved in nutrient recycling. In this work, the effect of commercial glyphosate and atrazine on denitrifying and diazotrophic populations has been compared. A soil without a history of previous herbicide application was incubated with one or several doses of herbicide, which was monitored along the experiment, and the microbial rate of denitrification and N2 fixation, the abundance of specific genes nirS, nirK, nosZ, nifH and the community structure of diazotrophs were analyzed. One dose of glyphosate or atrazine increased by 55% and 54%, respectively, the rate of N2 fixation and significantly reduced the rate of N2O production by incomplete denitrifiers. Long time exposure to glyphosate increased the abundance of nirK, nosZ, and nifH genes, but atrazine significantly reduced the nosZ gene density. Remarkably, diazotrophs belonging to the Bradyrhizobium genus, predominant in this soil, constituted a resilient population that became enriched after incubation with glyphosate or atrazine. Therefore, short and long-exposure to glyphosate and atrazine modifies the performance and survival of diazotrophs and denitrifiers in soil impacting the N biogeochemical cycle and the soil quality.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The implementation of intensive agricultural practices to improve the crop yield has led to an increased reliance on herbicides for weed control. With the rise of direct sowing to preserve soil quality, herbicides like glyphosate and atrazine are among the main agrochemicals used in the last decades worldwide. Glyphosate is a non-selective herbicide that severely injures any living plant tissue. It is widely used against annual and perennial weeds due to its efficiency on weeds elimination and to the availability of glyphosate-resistant varieties of soybean, cotton, canola, and maize (Annett et al. 2014; Duke and Powles 2008; Van Stempvoort et al. 2014). Around 20% of the agricultural top soils of the European Union countries contain glyphosate, and 42% contain aminomethylphosphonic acid (AMPA), its main metabolite (Silva et al. 2018). Atrazine is widely used in American and Asian countries to control broad-leaved weeds and grasses. Atrazine is used in the US, but with a restricted approval that will be reviewed by 2035 (Erickson 2020). Although formerly used across Europe, due to its long-term persistence in the environment and toxicity for wildlife, an EU-wide ban was set in 2004, but atrazine is still used in some countries (Nödler et al. 2013). In South America, glyphosate is the most used herbicide for no-till crop systems, and atrazine, though with a more restricted use, remains allowed for most of the countries (Camargo et al. 2020).
The extensive use of herbicides has raised concern about its toxic effects over soil microbial populations and functionality, particularly after long exposure since repeated applications are very common due to the increased emergence of herbicide-resistant weeds. The main target organism of herbicides are weeds, but soil microbial communities are exposed to considerable amounts of herbicides that may affect its structure and activity (Van Bruggen et al. 2018). Glyphosate inhibits the aromatic amino acid biosynthesis in shikimate pathway (Boocock and Coggins 1983), a biochemical route of active growing plants and some microorganisms (Sviridov et al. 2015). Also, indirect effects on other soil organisms by altering the gut microbiota have been reported (Motta et al. 2018). Atrazine, a systemic herbicide applied mostly to the soil for season-long weed control, inhibits photosynthesis at the level of photosystem II. Despite the selectivity of atrazine to photosynthetic organisms, various direct toxic effects on nontarget organisms have been reviewed recently (Singh et al. 2018) and indirect effects due to disturbance on the composition of intestinal microbiota in amphibians have been reported (Zhao et al. 2022).
The whole soil microbiota seems not to experience severe adverse effects after application of glyphosate or atrazine irrespective of the history of exposure of soil to these herbicides. Minimal changes in active soil microbial communities have been observed after a single application of glyphosate, atrazine or the surfactants that usually are included in commercial products containing herbicides (Banks et al. 2014). A single application of glyphosate to soils with or without history of exposure to the herbicide had minor effects on the structure and substrate utilization capability of soil bacterial communities (Zabaloy et al. 2012), the diversity, abundance, and physiological profile of soil bacteria (Allegrini et al. 2015), or respiration rates and microbial community composition (Lane et al. 2012). Atrazine exposure showed ambiguous effects on microbial community structure and activity. The C mineralization rate increased but the diversity of carbon source utilization by microbial soil biomass was reduced after atrazine application (Mahia et al. 2008; Yang et al. 2021). Transient adverse effects on the richness of the soil bacterial community were observed after atrazine application (Moretto et al. 2017) and increasing amounts of Actinobacteria were reported after long-term application (Liu et al. 2020). Atrazine and glyphosate have been applied together resulting in a stimulation on soil C and N mineralization rate (Haney et al. 2002), whereby it has been suggested that glyphosate may mitigate the transient adverse effect of atrazine when used together for weed control in genetically modified corn cropping (Bonfleur et al. 2015).
The uncertain effects of these herbicides on the whole soil microbial community is, probably due to several factors. It may be considered that soil herbicides concentration in field experiments decay due to the runoff, absorption by plants and degradation by soil microorganisms, which even can be adapted after successive applications. On the other hand, the great diversity and functional redundancy of the whole soil microbiota facilitates the substitution of inhibited by unaffected microorganisms, but when more specific functions of soil microbiota are affected, the recovering of the soil activity may be not so evident or fast. In addition, a stimulating effect has been endorsed to the release of limiting nutrients (mainly N or P) derived from herbicide degradation (Mijangos et al. 2009; Zabaloy et al. 2012). The system is even more complex in field applications, because after the breakdown of dead plants, the input of nutrients into the soil favors microbial growth. Therefore, the agricultural management and environmental factors determine the response of soil microbial community and activity to the herbicides in the field (Nguyen et al. 2016).
Despite the relevance of N cycle for soil function, the response to herbicides of microbial populations involved in N transformations has been scarcely explored. Only the response of ammonia oxidizing bacteria and archaea to glyphosate (Allegrini et al. 2017; Zabaloy et al. 2017) or atrazine (Hernandez et al. 2011) have been studied. Therefore, in this work we compared the effect of glyphosate and atrazine over the two main bacterial populations involved in the transformation of gaseous N in bare soil. Denitrifying and diazotrophic populations of a non-previously exposed soil were evaluated in microcosm incubations after short and long-term exposure to glyphosate or atrazine. The determination of denitrifying and N-fixation rates, the abundance of nifH, nirS, nirK, nosZ genes, and the composition of the bacterial diazotrophic community were determined during the soil incubation. Repeated doses of herbicide were applied to study the effect of long-term exposure on soil microbiota and the herbicide concentration was monitored during the incubation.
Materials and methods
Site description, sampling, and physicochemical parameters of soils
Three agricultural soils from Uruguay with no history of exposure to glyphosate or atrazine were chosen for this study. Soil A was sampled from Centro Regional Sur (Facultad de Agronomía) located at Progreso, Canelones (34°36′S and 56°13′W), soil B was sampled from Paso de la Laguna (Experimental Field of Instituto Nacional de Investigación Agropecuaria) located at Treinta y Tres (32°55′S and 54°50′W), and soil C was sampled from Servicio Seroterápico (Facultad de Medicina) located at Canelones (34°38′S and 55°55′W). The soils A and B were used for forestation and natural pasture management, respectively. The soil C was close to a field that only in the two previous years was used for agriculture in a rotation management with sunflower-ryegrass- sunflower-canola. Soils A and B were used in this study for preliminary determinations of half-life time of glyphosate and atrazine, and soil C was employed for the whole incubation experiment to determine the effect of herbicides on microbial communities.
Sampling was conducted during spring 2014. Five soil cores (0–15 cm depth) separated at least by 5 m were collected and pooled to make a composite sample from each site. Soil samples were analyzed for physicochemical properties: pH was measured in water (1:2.5 w/v), organic C and N content were measured by the wet oxidation method and Kjeldahl analysis, respectively, P was quantified after citric acid extraction, and K was extracted with ammonium acetate 1N at pH 7.0. Table 1 shows the classification and physicochemical properties of these soils.
The water content of soil C was 28%. After humidity determination, a fraction of the soil samples was air-dried at room temperature, sieved (< 2 mm) and stored at room temperature until to set up the incubation assays.
Preliminary experiment to estimate the glyphosate and atrazine degradation rate in soil
The persistence of herbicides in soil was estimated in a preliminary assay to determine the time between successive applications of herbicides. The concentration of glyphosate and atrazine after a first application was analyzed in fresh soils A and B, respectively. Microcosms assays were made in 125 mL flasks with fresh soil (10 g dry weight) and herbicides were added in solution from stock commercial formulations: glyphosate (540 g active ingredient L−1 Roundup Full II®) and atrazine (3.3 mg L−1 aqueous solution 90%, Novazina 90 GD®). The concentration of herbicides (1.5 kg atrazine ha−1 and 1.67 kg glyphosate ha−1) was adjusted assuming that the herbicides penetrate to a depth of 10 cm down the soil and that the apparent density of soil was 1.30 g cm−3. For both applications the volume of the added solution of herbicide was adjusted to reach the humidity of the soil when it was sampled. The incubation was done at 20 °C in the dark.
The half-life time estimated for glyphosate was 9 days and for atrazine was 46 days, assuming linear degradation rate from the initial herbicide application. Considering these results, the experiment to study the effect of several applications of herbicide on soil microbiota was designed. Glyphosate was applied every 9 days and atrazine was applied every 49 days to soil C; thus, the microorganisms would be exposed to an herbicide concentration in the range of the initial dose.
Effect of application of glyphosate and atrazine on soil microbial populations: Experimental design
A microcosms assay was carried out as described above with fresh soil C to study the effect of herbicides amendment on soil diazotrophic and denitrifying populations. The soil was distributed in flasks and sterile deionized water (control), or sterile solutions of glyphosate or atrazine were added to each flask. Three sets of flasks (control, glyphosate, and atrazine) were incubated aerobically in the dark at 20 °C.
The effect of the first application of herbicides was analyzed by comparing the abundance and activity of bacterial populations at t0 (immediately after the application of each herbicide) and at t1 (after the previously estimated half-time life for each herbicide: 9 days for glyphosate and 46 days for atrazine). The effect of long-term exposure to the herbicides was analyzed at t2, after periodical applications of glyphosate (6 applications and 49 days of incubation) or atrazine (4 applications and 268 days of incubation). Three replicates were destructively sampled at each sampling time (t0, t1 and t2) for each herbicide incubation set with the respective replicates of control soil incubation without herbicide.
Determination of the residual concentration of glyphosate and atrazine in soil
The concentration of herbicide was determined as described below in triplicated flasks at days 0, 9, 32 and 49 for glyphosate-amended soil, and at days 0, 46, 90, 135 and 268 for atrazine-amended soil. Three replicates were sacrificed at each sampling time to quantify the residual concentration of herbicide.
The residual glyphosate was quantified in soil by ELISA (Abraxis®) according to the manufacturer's instructions. The extraction was made with the complete content of soil in each flask, which was shaken with 25 mL of 1 M NaOH for 30 min. The suspension was transferred into a 50 mL plastic centrifuge tube and centrifuged at 5000 rpm for 15 min. The supernatant was diluted with ultrapure water (1:100 v/v), neutralized with HCl and filtered through a 0.2 µm membrane cellulose acetate filter.
Atrazine was extracted from soil as described by Amadori et al. (2013). The complete content of the flask was shaken with 15 mL acetonitrile for 30 min, then the suspension was transferred into a 50 mL plastic centrifuge tube and centrifuged at 4000 rpm for 15 min. This step was carried out three times. The extract solution (45 mL) was concentrated by rota-evaporation (200 rpm, 40 °C) to 1 mL in acetonitrile, diluted with ultrapure water (1:1 v/v), filtered through a 0.2 µm membrane filter and stored at − 20 °C. Atrazine was quantified by HPLC as described by Bellini et al. (2014).
Characterization of microbial populations
Denitrification and diazotrophic potential activity
The denitrification potential activity was measured in triplicate according to D’Heane et al. (2003), replacing glucose by 1.5 mM potassium acetate, 0.9 mM sodium succinate and 2.0 mM methanol, and adding 4.5 mM KNO3 (final concentrations). The assays were performed under anaerobic conditions (N2 atmosphere) in 60 mL vials by mixing 5 g dry soil with 10 mL of sterile water. The vials were sealed with butyl rubber stoppers and aluminum caps and incubated in the dark with continuous shaking, at 20 °C. Vials without and with acetylene (10% v/v into the headspace) were incubated to determine incomplete and total denitrification rate, respectively (Yoshinari et al. 1977). Gas samples were taken with a gas tight syringe from the headspace every hour, between 3.5 h and 7.5 h of incubation. The N2O concentration was immediately measured in a gas chromatograph (GC-2014 Shimadzu) equipped with an electron capture detector and two packed columns Porapack Q, 80/100 mesh 6 ft × 1/8 inch. The operating conditions were as follows: carrier gas N2 (30 mL min−1), injector temperature 90 °C, column and oven temperature 40 °C and detector temperature 250 °C. The denitrification potential activity was calculated from the slope of the N2O production curve and expressed as the rate of N2O production (nmol N2O.g dry soil−1.h−1). The measures of the headspace of abiotic control vials incubated with sterilized soil (with and without acetylene) confirmed that the N2O production was a biological process.
The diazotrophic potential activity was measured in triplicate using the acetylene reduction assay (ARA) (Hardy et al. 1968). The assays were performed in 25 mL vials by mixing 3 g dry soil with 10 mL of sterile water. The flasks were amended with a mixture of three different carbon sources (glucose, sodium citrate and sodium malate) at 2 mM final concentration. Vials were supplemented with acetylene (15% v/v into the headspace) and incubated in the dark with continuous shaking at 20 °C. The gas from the headspace was sampled after 48 h of incubation and ethylene was measured with a gas chromatograph (SRI 8610) equipped with a ionization flame detector and a packed column Porapack R, 80/100 mesh 6 ft × 1/8 inch. The operating conditions were as follows: carrier gas N2 (54 mL min−1), column oven temperature 45 °C and hydrogen flow 15 mL.min−1. The diazotrophic potential activity was calculated as the ethylene produced after 48 h of incubation.
DNA extraction and abundance of nifH, nirS, nirK, and nosZ genes in soil microcosms
DNA was extracted from 0.45 g of soil microcosms in duplicated samples. Mo Bio PowerSoil™ DNA Isolation kits were used according to the manufacturer’s protocol. The quality of the extracted DNA was verified on an agarose 1.5% gel and stored at − 20 °C until use.
The abundance of nifH, nirS, nirK, and nosZ genes was estimated by Real Time PCR (qPCR). Quantification of genes was performed with the following primers: nifH PolF/PolR (Poly et al. 2001), nirS cd3aF/R3Cd (Throback et al. 2004), nirK 876/R3Cu (Henry et al. 2004) and nosZ 2F/2R (Henry et al. 2006). Genes were amplified using the Rotor-Gene SYBR Green PCR Master mix (QIAGEN®, Hilden, Germany) in a Rotor-Gene® 6000, model 5-Plex (CORBETT Research, Sidney). All samples were amplified by duplicate and a standard curve (with triplicate determinations) was included in each run. Triplicates of negative controls without DNA template were included in each run. The standard preparation, thermal cycles and reaction conditions for the quantification were as described previously for nifH gene (Ferrando and Fernandez-Scavino 2015) and for nirS, nirK and nosZ genes (Bellini et al. 2018). DNA samples were amplified in 10 μL reaction volumes containing 1 μL of diluted (one or two tenfold) template DNA, 0.5 μM of each primer, and 5 μL of Rotor-Gene SYBR Green PCR Master mix. The thermal cycle consisted of an initial step at 95 °C for 5 min followed by 35 cycles of 95 °C for 5 s and 60 °C for 10 s. The fluorescence signal was measured once per cycle after the annealing-elongation step via the addition of one step at 80 °C for 1 s. A melting curve was obtained after each amplification by increasing temperature from 60 °C to 94 °C at a rate of 1 °C s−1 in order to verify the specificity of amplification. All DNA extractions were performed in the same soil and results were expressed as the amount of gene copies per g of dry soil.
Diazotrophic community structure by nifH pyrosequencing
DNA extracted from soils was analyzed at Mr DNA Molecular Research (TX, USA) for nifH barcoded pyrosequencing. The amplicon sequencing procedure was described by Dowd et al. (2008) with nifH specific primers PolR and PolF (Poly et al. 2001). A HotStarTaq Plus Master Mix Kit (Qiagen, Valencia, CA) was used and PCR conditions were: first step of 3 min at 94◦C, 28 cycles of amplification (94◦C for 30 s; 53◦C for 40 s and 72◦C for 1 min); final elongation at 72◦C for 5 min. Several amplicon products from each sample were mixed in equal concentrations and purified using Agencourt Ampure beads (Agencourt Bioscience Corporation, MA, USA). Samples were sequenced utilizing Roche 454 FLX titanium instruments following the manufacturer’s guidelines.
The nifH gene sequencing data were processed according to Ferrando and Fernández-Scavino (2015). The data were initially split in libraries followed by a de novo search for chimeras with userarch61 and QIIME (Caporaso et al. 2010). The split and filtered libraries were then screened for frame shifts using the FrameBot tool (Fish et al. 2013) with an identity cutoff of 0.4 and a length cutoff of 80 amino acids. The frame-corrected nucleotide sequences were then clustered (0.03 distance) using uclust (QIIME). An OTU table was constructed with OTUs with more than 10 counts. After standardization to the sample with the lower number of sequences, the data were subjected to further analyses. The software Analytic Rarefaction 1.3 (http://strata.uga.edu/software/index.html) developed by Steven Holland (Supplementary Information Fig. S1) was used with the original data set to construct the rarefaction curves.
The nifH sequences were deposited in the NCBI database with the Project number PRJNA884411 and BioSample accession numbers SAMN31023775 to SAMN31023784.
Statistical analysis
The differences between treatments were analyzed by one-way ANOVA followed by Tuckey’s test to establish the significance of the differences among means (p < 0.05) for each variable in study. The data were log10 transformed in order to generate a normal distribution of residues and homogeneity of variance. All analyses were performed with the software InfoStat/Professional Version 2016.
Results
Residual concentrations of glyphosate and atrazine in soil microcosms
The glyphosate and atrazine concentrations were monitored during the soil incubation to assess the range of herbicide concentration across the experiment. The concentrations of glyphosate and atrazine in soil microcosms were measured immediately after the first application (t0), after the estimated half-life time following the first application (t1), at least once before the following applications, and finally, at the end of the incubation (t2), that corresponded to 9 days after the 6th application of glyphosate or 133 days after the 4th application of atrazine.
Both herbicides were degraded in soil C with a half-life time of 6.7 days for glyphosate and 79 days for atrazine (Fig. 1). Glyphosate concentration was reduced to 33% of the initial value after 9 days from the first application (Fig. 1a), indicating that at t1 soil microbiota was exposed to a range of glyphosate concentration between 3.7 and 11.1 mg kg−1 dry soil. The long-term exposure (t2), after 49 days and six applications, corresponded to a range of glyphosate concentration between the minimal residual value measured (3.7 mg kg−1) and the highest residual value measured plus the new dose of glyphosate added (16.8 mg kg−1 by day 32). Atrazine concentration was reduced to 71% of the initial value by day 49, after the first application (Fig. 1b), indicating that at t1 soil microbiota was exposed to a concentration range between 0.68 and 0.97 mg atrazine kg−1 dry soil. During the long-term exposure, atrazine concentration was in the range of 0.45 to 2.50 mg kg−1 and soil was sampled at day 268 (t2), corresponding to the lowest concentration in soil.
Glyphosate (a) and atrazine (b) concentration in incubated soil microcosms after successive applications of the herbicides. Each point represents the mean concentration measured in triplicated microcosms ± standard deviation. Repeated applications of herbicides are indicated by arrows. Sampling times for microbiological analysis were: t0 (immediately after the first application), t1 (before the second application) and, t2 that corresponds to 9 days after the last application of glyphosate or 133 days after the last application of atrazine
The frequency of herbicides application, though not representative of their use in the field, simulates the extreme situation of permanent exposure of soil microbiota to the herbicide.
Effect of glyphosate and atrazine on the activity and abundance of soil denitrifying populations
In order to determine the effect of the herbicides on the activity of denitrifying populations, the N2O production rate was measured in microcosms assays with or without acetylene to determine the Potential Total Denitrification Rate (PTDR) or the Potential Incomplete Denitrification Rate (PIDR), respectively. The PTDR, measured by the acetylene blockage method, includes the activity of microorganisms able to produce N2 as well as microorganisms able to produce N2O, due to the absence of the nosZ gene, responsible for the last step of denitrification. Denitrification rates were also measured in parallel in control microcosms incubated without herbicide amendment. The effect of short (t1) and long-term exposure (t2) to glyphosate or atrazine on the denitrification rate is shown in Fig. 2.
Potential total denitrification rate (a, b) and potential incomplete denitrification rate (c, d) in soil incubated with glyphosate (a, c) or atrazine (b, d) after one (t1) or several (t2) applications of the herbicide. Incubated soils with herbicide and without herbicide (non-exposed) were measured at each sampling time. Error bars represent standard deviation of three replicated microcosms. Different letters indicate significant differences between treatments (p < 0.05)
PTDR and PIDR were in the same order of magnitude for soils exposed and unexposed to both herbicides, though significant differences could be observed for certain conditions. PTDR was not significantly affected by short or long-term exposure to glyphosate (Fig. 2a). However, PIDR of soil after the first glyphosate application was significantly lower (17%) than PIDR of unexposed soil (Fig. 2c). This effect was transient, since after six applications of glyphosate (t2), PIDR was similar in control and exposed soils, suggesting that incomplete denitrifiers may have been adapted to glyphosate.
Conversely, atrazine was consistently inhibitory for the denitrifiers activity. The PTDR in control and exposed soils were similar after the first application but decreased significantly in soil exposed for long time to atrazine (Fig. 2b). At t2, the PTDR decreased by 12% in soil incubated with atrazine compared to control soil. The PIDR decreased significantly since the first exposure to atrazine, decaying by 17% at t1 and by 14% at t2 (after 268 days of incubation with atrazine), indicating that the inhibitory effect persisted but did not increase with time.
Therefore, atrazine showed a stronger inhibitory effect than glyphosate on denitrifiers activity. Also, incomplete denitrifiers seemed to be more affected than the whole denitrifying community after exposure to the herbicides.
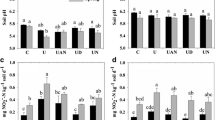
The effect of herbicides on soil microbiota was also analyzed through the quantification of specific genes involved in the bacterial denitrification pathway. The abundance of nirS and nirK genes was not negatively affected by soil incubations with glyphosate or atrazine (Fig. 3a–d). Moreover, soil incubated for long-term with glyphosate showed a significant increase of nirK gene from 2.2 × 107 to 4.7 × 107 copies g−1 dry soil (Fig. 3c).
Abundance of genes involved in the denitrification pathway in soils unexposed and exposed to glyphosate (left) or atrazine (right) after one (t1) or several (t2) applications. Abundance of nirS (a, b), nirK (c, d) and nosZ (e, f) genes were represented. Error bars represent standard deviation of two determinations. Different letters indicate significant differences among treatments for each gene (p < 0.05)
On the contrary, the nosZ gene abundance was affected after long-time incubation with both herbicides when compared with incubated soils without herbicide but showing opposite effects (Fig. 3e, f). The nosZ gene abundance increased significantly (4.0 × 106 to 8.1 × 106 copies. g−1 dry soil) after incubation with glyphosate, whereas decreased significantly (6.2 × 106 to 4.8 × 106 copies. g−1 dry soil) after atrazine incubation.
Thereby, except by a decline in the nosZ gene density after long-exposure to atrazine, the abundance of key genes of soil bacterial denitrifying populations remained unchanged or even increased after long-time incubation with glyphosate and atrazine.
Effect of glyphosate and atrazine on the activity and abundance of soil diazotrophic population
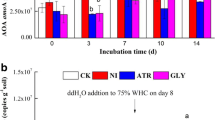
Glyphosate stimulated significantly N2 fixation activity with rates that increased by 55% and 166% compared to the controls, at t1 and t2, respectively (Fig. 4a). Also, the nifH gene density (5.6 × 106 copies. g−1 dry soil) was significantly higher than in control soil (2.6 × 106 copies.g−1 dry soil) at t2 (Fig. 5a). However, the increase of diazotrophic activity at t1 could not be explained by the abundance of nifH genes, which was lower in soil with glyphosate compared to control soil.
The exposure to atrazine significantly changed the rate of N2 fixation when compared to the respective control soil, showing a sharp increase (54%) after the first exposure and a great decline (47%) of the rate at t2 (Fig. 4b). The abundance of nifH genes was not significantly affected by the exposure to atrazine (Fig. 5b).
Therefore, though the density of the diazotrophic population seemed to be rarely affected by glyphosate and atrazine, the rate for N2 fixation was strongly impacted after exposure to herbicides, but mostly showing a stimulating effect.
Effect of glyphosate and atrazine on the diversity and composition of soil diazotrophic community
The affiliation and relative abundance of nifH gene sequences were analyzed to compare the composition of the diazotrophic bacterial community in exposed and unexposed soils. The unincubated soil showed sequences belonging to Alpha, Beta, Gamma and Deltaproteobacteria, with predominance of the Alphaproteobacteria, mainly associated to different species of the genus Bradyrhizobium. Bacteria from the groups Firmicutes, Cyanobacteria, Planctomycetes and Verrucomicrobia were found as minor members of the diazotrophic community in this soil (Fig. 6).
Heat map showing the relative proportion of nifH gene sequences retrieved from soils before incubation (t0a and t0b) and incubated with glyphosate or atrazine after the first herbicide application (t1 + G or t1 + A) or several herbicide applications (t2 + G or t2 + A), respectively. The control soils incubated without herbicides at t1 (t1 -G or t1 -A) and t2 (t2 -G or t2 -A) are also shown. The OTU number and the affiliation to the respective closest NifH protein sequence are shown
After incubation with glyphosate, a noticeable increase of the relative abundance of OTU 364 was observed (Fig. 6), which has a high level of similarity to the nifH sequence of Bradyrhizobium elkanii. The increase was more evident when compared with the incubated control after the first exposure (t1,). Other Alphaproteobacteria were unaffected by glyphosate at t1. Soil incubated with higher amounts of glyphosate for a long time (t2) seemed to increase the relative proportion of other members of the genus Bradyrhizobium (OTUs 8 and 360) compared to control soil. Conversely, the relative proportion of a few diazotrophs decreased after incubation with glyphosate. Short and long-term exposure affected Geobacter-like (OTU 390) and Verrucomicrobia-like (OTU 297) diazotrophs, whereas certain Betaproteobacteria (OTU 237), Firmicutes (OTU 260), Nostocales (OTU 176) and Planctomycetes (OTU 410) decreased only after long incubation with glyphosate.
The incubation with atrazine also increased the relative proportion of Bradyrhizobium elkanii-like diazotrophs (OTU 364). Alpha and Betaproteobacteria were mainly not affected by atrazine, but adverse effects were observed for Bacillus and certain members of Nostocales (OTUs 260 and 419, respectively). The long time of incubation required to test the effect of repeated additions of atrazine seemed to have a great influence on the diazotrophs distribution, since incubated soils at t2 with and without atrazine showed similar proportions for several OTUs. At t2, two members of the Firmicutes (OTU 260 and 200) and one of the Order Nostocales (OTU 405) seemed to be favored by the long-term exposure to atrazine.
Discussion
The current use of herbicides has increased substantially due to the expansion of intensive agriculture and pressure to obtain higher crop yields. Soil microbial diversity and functionality may be threatened by this continuous use of herbicides. Although designed to have selective toxicity over vegetal cells, herbicides (or other compounds like surfactants that are commonly included in its formulations) may have adverse effects on the soil microbiota. An additional concern has been raised recently since it was hypothesized that the selection pressure for glyphosate resistance could increase antibiotic resistant populations in soil microbiota (Hertel et al. 2022; Van Bruggen et al. 2018). Furthermore, the impact of herbicides on microbial populations involved in the transformations of limiting nutrients, such as N, could have serious consequences for sustainable agriculture.
In this work, we examine the response towards glyphosate and atrazine of the microbial populations of denitrifiers and diazotrophs, responsible for the exchange of N between soil and the atmosphere. A soil without history of herbicide application was incubated with and without glyphosate or atrazine under controlled conditions. After one (short-time exposure, t1) or several (long-time exposure, t2) doses of commercial herbicides, the activity, abundance, and diversity of diazotrophic and denitrifying soil populations were examined. The herbicides were applied repeatedly to the soil microcosms to maintain their concentration in the range of use in the field and their concentrations were monitored along the long-term incubation experiment.
Both herbicides were degraded during soil incubation. The rate of herbicide degradation depends on the soil type, temperature, and moisture, as well as the soil microbiota, the presence of plants and stubble, and the history of previous herbicide application. Therefore, it is not surprising to find different biodegradation rates reported for glyphosate and atrazine (Mueller et al. 2017; Muskus et al. 2019; Ngigi et al. 2011). The decay of glyphosate to 50% of the initial concentration was generally shorter than 10 days for different soils around the world (Al-Rajab and Schiavon 2010; Yang et al. 2015). Consistently, the half-life time of glyphosate after the first exposure was 6.7 days in our experiment. The half-life of atrazine was 79 days, which agrees with the estimate of 60 days reported for different soils (Ghadiri et al. 1984; Krutz et al. 2007). Considering this degradation, the periodical applications of herbicides to the microcosm assay allowed to keep the glyphosate and atrazine concentration within their range of use in the field simulating a long-time exposure to herbicides.
The effect of glyphosate and atrazine on a specific group of bacteria has been reported by a limited number of studies. Glyphosate increased the viable bacterial counts of Pseudomonas (Gimsing et al. 2004) and Gram-negative phospholipids fatty acids (PLFA) markers in soil (Weaver et al. 2007), though other authors observed negative effects on rhizospheric bacteria able to promote plant growth, such as Burkholderia spp., Pseudomonas spp., or Rhizobium spp. (Arango et al. 2014; Druille et al. 2016; Lorch et al. 2021; Zobiole et al. 2010). Atrazine and glyphosate showed adverse effects on the growth of axenic cultures of nitrogen fixing bacteria such as Azospirillum brasilense and Rhizobium sp., or bacteria employed in biological control, such as Bacillus subtilis and Bacillus thuringiensis (De Farias et al. 2021).
Among the microbial populations involved in the biogeochemical cycle of N, nitrifying bacteria and archaea have been the most studied in terms of their sensitivity to glyphosate or atrazine. Different responses have been observed for the abundance of both groups since high doses of glyphosate increased the nitrifying bacteria, whereas the density of nitrifying archaea remained unchanged (Zabaloy et al. 2017). Also, repeated applications of high doses of glyphosate (49 mg kg−1 soil) cause a shift in the community structure of ammonia-oxidizing bacteria of soils with and without history of exposure to the herbicide (Allegrini et al. 2017). On the other hand, simazine, a herbicide with a similar chemical structure than atrazine, also differentiated among ammonia oxidizing archaea and bacteria since only bacterial DGGE patterns were affected by the herbicide in high doses (Hernandez et al. 2011). In the present work, differential responses to the herbicides were also found for denitrifying populations (Table 2). The total denitrification rate decayed only after long exposure to atrazine (12%), but incomplete denitrification rate decreased after short and long exposure to atrazine (17% and 14%, respectively) or after the first exposure to glyphosate (17%). Since the measurement of total denitrification rate comprises the activity of both, complete and incomplete denitrifiers, the higher effect observed for the N2O rate production may be attributed to this process is carried out for a subgroup within microbial denitrifiers. Incomplete denitrification, performed by bacteria lacking the nosZ gene and by fungi, is a relevant environmental process since it contributes to N losses from soil and to the emission of N2O, a greenhouse gas. Fungal N2O production is comparable to the bacterial production in soils of diverse agroecosystems, including conventional farming, integrated crop and livestock systems, organic farming systems (Chen et al. 2014; Mothapo et al. 2013) or grasslands (Zhong et al. 2022), but fungal denitrification is higher than bacterial denitrification manly at acidic soils with complex C substrates like lignocellulose (Chen et al. 2015). In addition, agricultural practices such as organic fertilization or reduced tillage, which are frequently accompanied by the use of herbicides, increase denitrification rates and the contribution to N2O emission by fungi (Bosch et al. 2022; Wei et al. 2014). The application of glyphosate and propanil reduced N2O emissions in soils amended with rice straw or chitin, so the combination of these herbicides could be a useful N2O emission mitigation strategy in soils amended with organic fertilizers (Kyaw and Toyota 2007). It is also appropriate to mention that the nosZ gene analyzed in this work belongs to the Clade I present in many denitrifying bacteria, mainly Proteobacteria, while the microorganisms that possess the nosZ gene of clade II comprise multiple bacterial phyla for which their contribution to the N2O reduction remains still unclear (Shapleigh 2013).
Therefore, the observed decay of the N2O production rate after glyphosate and, mainly atrazine exposure, might have an environmental positive effect to prevent N losses from soil and reduce the emission of a greenhouse gas. Our results also suggest that incomplete denitrifiers could be suitable indicators to determine the inhibitory effects of these herbicides.
The abundance of denitrification specific genes was significantly affected by long-time exposure to both herbicides (Table 3). The incubations with glyphosate increased the densities of nirK and nosZ genes (from 2.2 × 107 to 4.7 × 107 and from 4.0 × 106 to 8.1 × 106 copies. g−1 dry soil, respectively) after incubation with glyphosate, but atrazine caused a significant decrease of nosZ gene density (from 6.2 × 106 to 4.8 × 106 copies. g−1 dry soil). Other authors reported that one dose of glyphosate or atrazine did not affect the density of nosZ gene (Zhang et al. 2018), or even one elevated dose of glyphosate (21.6 mg kg−1) did not produce noticeable changes in the abundance of denitrification genes in a sugarcane cropped soil (Das et al. 2022). Thus, our results showed that long-time exposure to herbicides affected differentially the abundance of denitrifying bacteria since two of the main populations that contributed to N2 losses increased after glyphosate exposure but a decrease of the population able to reduce N2O to N2 was observed after incubation with atrazine.
Although herbicides are frequently employed in agricultural managements that include inoculation of seeds with plant growth promoting bacteria, commonly diazotrophs, the response of diazotrophs to herbicides has been scarcely explored. The N2 fixation rate was markedly enhanced after one or several applications of glyphosate (55% and 166%, respectively) and the nifH gene abundance raised from 2.6 × 106 to 5.6 × 106 copies g−1 dry soil after long-time exposure (Table 2 and Table 3). This consistent stimulation of diazotrophic bacteria by the application of commercial glyphosate may be due to its degradation and consequent release of metabolites that are nutrients for nitrogenase activity and growth of diazotrophic bacteria. Conversely to our results, four annual applications of glyphosate (either 384 or 1440 g active ingredient ha−1) significantly decreased the abundance of culturable free-living diazotrophs in a temperate grassland soil (Druille et al. 2016).
The effect of atrazine on the N2 fixation rate depended on the time of exposure to the herbicide, showing a significant improvement (54%) after the first dose but a strong reduction of the diazotrophic activity (47% lower than the control soil) after long-time exposure (Table 2). These changes in diazotrophic activity occurred without significant changes in nifH gene density, suggesting that atrazine has reversible effects on the activity of diazotrophs or that the community composition of diazotrophs shifted during the long-time towards diazotrophs with lower tolerance to herbicides.
The taxonomic affiliation of nifH gene sequences revealed that this soil harbored representative members of Alpha, Beta, Gamma, and Delta Proteobacteria, as well as members of the orders Firmicutes, Nostocales, Planctomycetales and Verrucomicrobiales. The incubation with glyphosate seems to reduce the relative proportion of only certain members within the main taxonomic groups, such as Dechloromonas within Betaproteobacteria or Bacillus within Firmicutes, but Alphaproteobacteria, remained predominant with nifH sequences associated to different species of the genus Bradyrhizobium. Although the same trend was observed for soil incubated with atrazine, the effect would be less evident because the long-time of incubation also seems to affect control soil without atrazine. The most interesting outcome observed in bacterial community composition was that the proportion of diazotrophs associated to Bradyrhizobium (within Alphaproteobacteria) and to Paraburkholderia-Burkholderia (within Betaproteobacteria) would be enriched after soil incubation with glyphosate or atrazine. The persistence of Bradyrhizobium may be explained by the ability of bacteria belonging to this genus to degrade glyphosate and atrazine (Hernández-Guijarro et al. 2021; Vercellino and Gómez 2013). Although it has been reported that formulated products based on glyphosate were toxic to the strain Bradyrhizobium sp BR 3901 (Madureira-Barroso et al. 2020), it has been proposed that the effects of glyphosate on microbes of agricultural soils are minor and transient (Duke 2021). It is also interesting to note that most of the Bradyrhizobium strains recently isolated from peanut-nodules were denitrifiers, according to the genomic analysis, but only one third of this population could reduce N2O and had strong preference for N2O over NO−3 as electron acceptor (Gao et al. 2021).
Overall, the results of the present work evidenced that the application of one dose of glyphosate or atrazine does not stimulate the activity of microbial populations that can deprive the soil of N, moreover, decreases the N2O emission and stimulates N2 fixation. Long- time exposure to glyphosate increases the populations of denitrifiers and diazotrophs, and consequently accelerates the turnover of N in soil, and enhances noticeably the N2 fixation. Atrazine reduces one of the main populations able to transform N2O in N2, causing environmental detrimental effects, and decreases the rate of denitrification and N2 fixation, showing an opposite effect than glyphosate on the N turnover in soil. Remarkably, bacteria belonging to the Bradyrhizobium genus, the predominant diazotroph genus in this soil, seems to be a resilient population that became enriched after incubation with glyphosate or atrazine. These results contribute to implementing more sustainable agricultural practices combining herbicides application with plant growth promoting bacteria inoculation. (Sequencing data are available at the link: http://www.ncbi.nlm.nih.gov/bioproject/884411).
References
Allegrini M, Zabaloy MC, del Gómez EV (2015) Ecotoxicological assessment of soil microbial community tolerance to glyphosate. Sci Total Environ 533:60–68. https://doi.org/10.1016/j.scitotenv.2015.06.096
Allegrini M, del Gomez EV, Zabaloy MC (2017) Repeated glyphosate exposure induces shifts in nitrifying communities and metabolism of phenylpropanoids. Soil Biol Biochem 105:206–215. https://doi.org/10.1016/j.soilbio.2016.11.024
Al-Rajab AJ, Schiavon M (2010) Degradation of 14C-glyphosate and aminomethylphosphonic acid (AMPA) in three agricultural soils. J Environ Sci 22:1374–1380. https://doi.org/10.1016/s1001-0742(09)60264-3
Amadori MF (2013) Extraction method for the determination of atrazine, deethylatrazine, and deisopropylatrazine in agricultural soil using factorial design. J Braz Chem Soc 24:483–491
Annett R, Habibi HR, Hontela A (2014) Impact of glyphosate and glyphosate- based herbicides on the freshwater environment. J Appl Toxicol 34(5):458–479
Arango L, Buddrus-Schiemann K, Opelt K et al (2014) Effects of glyphosate on the bacterial community associated with roots of transgenic roundup ready® soybean. Eur J Soil Biol 63:41–48. https://doi.org/10.1016/j.ejsobi.2014.05.005
Banks ML, Kennedy AC, Kremer RJ, Eivazi F (2014) Soil microbial community response to surfactants and herbicides in two soils. Appl Soil Ecol 74:12–20. https://doi.org/10.1016/j.apsoil.2013.08.018
Bellini MI, Pinelli L, Dos Santos ME, Fernández Scavino A (2014) Bacterial consortia from raw water and sludges from water potabilization plants are able to degrade atrazine. Int Biodeterior Biodegrad 90:131–139
Bellini MI, Kumaresan D, Tarlera S, Murrell C, Fernández Scavino A (2018) Identification of active denitrifiers by DNA-stable isotope probing and amplicon sequencing reveals betaproteobacteria as responsible for attenuation of nitrate contamination in a low impacted aquifer. FEMS Microbiol Ecol 94:181–193
Bonfleur EJ, Tornisielo VL, Regitano JB, Lavorenti A (2015) The effects of glyphosate and atrazine mixture on soil microbial population and subsequent impacts on their fate in a tropical soil. Water Air Soil Pollut Focus 226:21. https://doi.org/10.1007/s11270-014-2190-8
Boocock MR, Coggins JR (1983) Kinetics of 5-enolpyruvylshikimate-3-phosphate synthase inhibition by glyphosate. FEBS Lett 154:127–133. https://doi.org/10.1016/0014-5793(83)80888-6
Bösch Y, Jones CM, Finlay R et al (2022) Minimizing tillage modifies fungal denitrifier communities, increases denitrification rates and enhances the genetic potential for fungal, relative to bacterial, denitrification. Soil Biol Biochem 170:108718. https://doi.org/10.1016/j.soilbio.2022.108718
Camargo E, Zapiola M, Avila L, Garcia M, Plaza G, Gazziero D, Hoyos V (2020) Current situation regarding herbicide regulation and public perception in South America. Weed Sci. https://doi.org/10.1017/wsc.2020.14
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Chen Y, Zhou W, Li Y et al (2014) Nitrite reductase genes as functional markers to investigate diversity of denitrifying bacteria during agricultural waste composting. Appl Microbiol Biotechnol 98:4233–4243. https://doi.org/10.1007/s00253-014-5514-0
Chen H, Mothapo NV, Shi W et al (2015) Soil moisture and pH control relative contributions of fungi and bacteria to N2O production. Microb Ecol 69:180–191. https://doi.org/10.1007/s00248-014-0488-0
da Barbosa CN, Hébert M-P, Fugère V et al (2022) A glyphosate-based herbicide cross-selects for antibiotic resistance genes in bacterioplankton communities. mSystems 7:e0148221. https://doi.org/10.1128/msystems.01482-21
D’Heane K, Moreels E, De Neve S, Chaves Daguilar B, Boeckx P (2003) Soil properties influencing the denitrification potential of Flemish agricultural soils. Bio Fertil Soils 38:358–366
Das S, Wang W, Reeves S et al (2022) Non-target impacts of pesticides on soil N transformations, abundances of nitrifying and denitrifying genes, and nitrous oxide emissions. Sci Total Environ 844:157043. https://doi.org/10.1016/j.scitotenv.2022.157043
De Farias DIOA, Leite RC, Ribeiro EA et al (2021) Glyphosate and atrazine inhibit growth of Azospirillum brasilense, Bacillus subtilis, Bacillus thuringiensis, Chromobacterium subtsugae and Saccharopolyspora spinosa. Agron Colomb 39:64–71. https://doi.org/10.15446/agron.colomb.v39n1.89870
Dowd SE, Sun Y, Wolcott RD et al (2008) Bacterial tag-encoded FLX amplicon pyrosequencing (bTEFAP) for microbiome studies: bacterial diversity in the ileum of newly weaned Salmonella infected pigs. Foodborne Pathog Dis 5:459–472
Druille M, García-Parisi PA, Golluscio RA et al (2016) Repeated annual glyphosate applications may impair beneficial soil microorganisms in temperate grassland. Agric Ecosyst Environ 230:184–190. https://doi.org/10.1016/j.agee.2016.06.011
Duke SO (2021) Glyphosate: uses other than in glyphosate-resistant crops, mode of action, degradation in plants, and effects on non-target plants and agricultural microbes. Rev Environ Contam Toxicol 255:1–65. https://doi.org/10.1007/398_2020_53
Duke SO, Powles SB (2008) Glyphosate: a once-in-a-century herbicide. Pest Manag Sci 64(4):319–325
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Erickson BE (2020) US EPA reapproves atrazine. https://www.epa.gov/ingredients-used-pesticide-products/atrazine. Accessed 6 Oct 2022
Ferrando L, Fernández Scavino A (2015) Strong shift in the diazotrophic endophytic bacterial community inhabiting rice (Oryza sativa) plants after flooding. FEMS Microbiol Ecol 91:fiv104. https://doi.org/10.1093/femsec/fiv104
Fish JA, Chai B, Wang Q et al (2013) FunGene: the functional gene pipeline and repository. Front Microbiol 4:291. https://doi.org/10.3389/fmicb.2013.00291
Gao Y, Mania D, Mousavi SA et al (2021) Competition for electrons favors N2O reduction in denitrifying Bradyrhizobium isolates. Environ Microbiol 23:2244–2259. https://doi.org/10.1111/1462-2920.15404
Ghadiri H, Shea PJ, Wicks GA, Haderlie LC (1984) Atrazine dissipation in conventional-till and no-till sorghum. J Environ Qual 13:549–552. https://doi.org/10.2134/jeq1984.00472425001300040008x
Gimsing AL, Borggaard OK, Jacobsen OS, Aamand J, Sørensen J (2004) Chemical and microbiological soil characteristics controlling glyphosate mineralisation in Danish surface soils. Appl Soil Ecol 27:233–242
Haney R, Senseman S, Krutz L, Hons F (2002) Soil carbon and nitrogen mineralization as affected by atrazine and glyphosate. Biol Fertil Soils 35:35–40. https://doi.org/10.1007/s00374-001-0437-1
Hardy RWF, Holsten RD, Jackson EK, Burns RC (1968) The acetylene-ethylene assay for N2 fixation: laboratory and field evaluation. Plant Physiol 43:1185–1207
Henry SE, Baudoi JC, López-Gutierrez F, Martin-Laurent A, Brauman PL (2004) Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. J Microbiol Meth 59:327–335
Henry S, Bru D, Stres B, Hallet S, Philippot L (2006) Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16SrRNA, narG, nirK, and nosZ genes in soils. Appl Environ Microb 72:5181–5189
Hernández M, Jia Z, Conrad R, Seeger M (2011) Simazine application inhibits nitrification and changes the ammonia-oxidizing bacterial communities in a fertilized agricultural soil. FEMS Microbiol Ecol 78:511–519. https://doi.org/10.1111/j.1574-6941.2011.01180.x
Hernández Guijarro K, De Gerónimo E, Erijman L (2021) Glyphosate biodegradation potential in soil based on glycine oxidase gene (thiO) from Bradyrhizobium. Curr Microbiol 78:1991–2000. https://doi.org/10.1007/s00284-021-02467-z
Hertel R, Schome K, Mittelstat MJ et al (2022) Characterization of glyphosate-resistant Burkholderia anthina and Burkholderia cenocepacia isolates from a commercial Roundup® solution. Environ Micob Rep 14:70–84. https://doi.org/10.1111/1758-2229.13022
Krutz LJ, Zablotowicz RM, Reddy KN, Koger CH III, Weaver MA (2007) Enhanced degradation of atrazine under field conditions correlates with a loss of weed control in the glasshouse. Pest Manag Sci 63:23–31
Kyaw KM, Toyota K (2007) Suppression of nitrous oxide production by the herbicides glyphosate and propanil in soils supplied with organic matter. Soil Sci Plant Nutr 53:441–447
Lane M, Lorenz N, Saxena J et al (2012) The effect of glyphosate on soil microbial activity, microbial community structure, and soil potassium. Pedobiologia 55:335–342. https://doi.org/10.1016/j.pedobi.2012.08.001
Liu Y, Fan X, Zhang T et al (2020) Effects of the long-term application of atrazine on soil enzyme activity and bacterial community structure in farmlands in China. Environ Pollut 262:114264. https://doi.org/10.1016/j.envpol.2020.114264
Lorch M, Agaras B, García-Parisi P et al (2021) Repeated annual application of glyphosate reduces the abundance and alters the community structure of soil culturable pseudomonads in a temperate grassland. Agric Ecosyst Environ 319:107503. https://doi.org/10.1016/j.agee.2021.107503
Madureira Barroso G, dos Santos JB, de Oliveira IT et al (2020) Tolerance of Bradyrhizobium sp. BR 3901 to herbicides and their ability to use these pesticides as a nutritional source. Ecol Indic. https://doi.org/10.1016/j.ecolind.2020.106783
Mahía J, Cabaneiro A, Carballas T et al (2008) Microbial biomass and C mineralization in agricultural soils as affected by atrazine addition. Biol Fertil Soils. https://doi.org/10.1007/s00374-008-0318
Mijangos I, Becerril JM, Albizu I et al (2009) Effects of glyphosate on rhizosphere soil microbial communities under two different plant compositions by cultivation-dependent and -independent methodologies. Soil Biol Biochem 41:505–513. https://doi.org/10.1016/j.soilbio.2008.12.009
Moretto JAS, Altarugio LM, Andrade PA et al (2017) Changes in bacterial community after application of three different herbicides. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnx113
Mothapo N, Chen H, Cubeta MA et al (2013) Phylogenetic, taxonomic and functional diversity of fungal denitrifiers and associated N2O production efficacy. Soil Biol Biochem 83:160–175. https://doi.org/10.1016/j.soilbio.2015.02.001
Motta EVS, Raymann K, Moran NA (2018) Glyphosate perturbs the gut microbiota of honey bees. Proc Natl Acad Sci U S A 115:10305–10310. https://doi.org/10.1073/pnas.1803880115
Mueller TC, Parker ET, Steckel L et al (2017) Enhanced atrazine degradation is widespread across the United States. Pest Manag Sci 73:1953–1961. https://doi.org/10.1002/ps.4566
Muskus AM, Krauss M, Miltner A et al (2019) Effect of temperature, pH and total organic carbon variations on microbial turnover of 13C315N-glyphosate in agricultural soil. Sci Total Environ 658:697–707. https://doi.org/10.1016/j.scitotenv.2018.12.195
Ngigi A, Dörfler U, Scherb H et al (2011) Effect of fluctuating soil humidity on in situ bioavailability and degradation of atrazine. Chemosphere 84:369–375. https://doi.org/10.1016/j.chemosphere.2011.03.068
Nguyen DB, Rose MT, Rose TJ et al (2016) Impact of glyphosate on soil microbial biomass and respiration: a meta-analysis. Soil Biol Biochem 92:50–57
Nödler K, Licha T, Voutsa D (2013) Twenty years later – atrazine concentrations in selected coastal waters of the Mediterranean and the Baltic Sea. Mar Pollut Bull. https://doi.org/10.1016/j.marpolbul.2013.02.018
Poly F, Monrozier LJ, Bally R (2001) Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Res Microbiol 152:95–103
Shapleigh JP (2013) The prokaryotes: prokaryotic physiology and biochemistry. In: Rosenberg E, Delong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes. Springer, Berlin, Heidelberg, pp 405–425
Silva V, Montanarella L, Jones A et al (2018) Distribution of glyphosate and aminomethylphosphonic acid (AMPA) in agricultural topsoils of the European Union. Sci Total Environ 621:1352–1359. https://doi.org/10.1016/j.scitotenv.2017.10.093
Singh S, Kumar V, Chauhan A et al (2018) Toxicity, degradation and analysis of the herbicide atrazine. Environ Chem Lett 16:211–237. https://doi.org/10.1007/s10311-017-0665-8
Sviridov AV, Shushkova TV, Ermakova IT et al (2015) Microbial degradation of glyphosate herbicides (review). Appl Biochem Microbiol 51:188–195. https://doi.org/10.1134/S0003683815020209
Throbäck IN, Enwall K, Javis A, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK, and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
Van Bruggen AHC, He MM, Shin K et al (2018) Environmental and health effects of the herbicide glyphosate. Sci Total Environ 616–617:255–268. https://doi.org/10.1016/j.scitotenv.2017.10.309
Van Stempvoort DR, Roy JW, Brown SJ, Bickerton G (2014) Residues of the herbicide glyphosate in riparian groundwater in urban catchments. Chemosphere 95:455–463
Vercellino M, Gómez MA (2013) Denitrifying capacity of rhizobial strains of Argentine soils and herbicide sensitivity. Ann Microbiol 63:1563–1570. https://doi.org/10.1007/s13213-013-0619-8
Weaver MA, Krutz LJ, Zablotowicz RM, Reddy KN (2007) Effects of glyphosate on soil microbial communities and its mineralization in a Mississippi soil. Pest Manag Sci 63:388–393. https://doi.org/10.1002/ps.1351
Wei W, Isobe K, Shiratori Y et al (2014) N2O emission from cropland field soil through fungal denitrification after surface applications of organic fertilizer. Soil Biol Biochem 69:157–167. https://doi.org/10.1016/j.soilbio.2013.10.044
Yang X, Wang F, Bento CPM, Meng L et al (2015) Decay characteristics and erosion-related transport of glyphosate in Chinese loess soil under field conditions. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2015.05.082
Yang F, Gao M, Lu H et al (2021) Effects of atrazine on chernozem microbial communities evaluated by traditional detection and modern sequencing technology. Microorganisms. https://doi.org/10.3390/microorganisms9091832
Yoshinari T, Hynes R, Knowles R (1977) Acetylene inhibition of nitrous oxide reduction and measurement of denitrification and nitrogen fixation in soil. Soil Biol Biochem 9:177–183
Zabaloy MC, Gómez E, Garland JL, Gómez MA (2012) Assessment of microbial community function and structure in soil microcosms exposed to glyphosate. Appl Soil Ecol 61:333–339. https://doi.org/10.1016/j.apsoil.2011.12.004
Zabaloy MC, Allegrini M, Tebbe DA et al (2017) Nitrifying bacteria and archaea withstanding glyphosate in fertilized soil microcosms. Appl Soil Ecol 117–118:88–95. https://doi.org/10.1016/j.apsoil.2017.04.012
Zhang M, Wang W, Tang L et al (2018) Effects of nitrification inhibitor and herbicides on nitrification, nitrite and nitrate consumptions and nitrous oxide emission in an Australian sugarcane soil. Biol Fertil Soils 54:697–706. https://doi.org/10.1007/s00374-018-1293-6
Zhao Q, Huang M, Yin J et al (2022) Atrazine exposure and recovery alter the intestinal structure, bacterial composition and intestinal metabolites of male Pelophylax nigromaculatus. Sci Total Environ 818:151701. https://doi.org/10.1016/j.scitotenv.2021.151701
Zhong L, Qing J, Liu M, Cai X et al (2022) Fungi and archaea control soil N2O production potential in Chinese grasslands rather than Bacteria. Front Microb. https://doi.org/10.3389/fmicb.2022.844663
Zobiole LHS, Oliveira RS, Kremer RJ et al (2010) Effect of glyphosate on symbiotic N2 fixation and nickel concentration in glyphosate-resistant soybeans. Appl Soil Ecol 44:176–180. https://doi.org/10.1016/j.apsoil.2009.12.003
Acknowledgements
The authors are very grateful to the MSc. Nadia Martin who did excellent technical work by performing many analyses and by contributing with enthusiasm to the accomplishment of this project. The authors are grateful to MSc. Lucas Martinez Arocena, who contributed to the analysis of the residual atrazine concentration.
Funding
This study was funded by the I + D Grant number 815 of the Comisión Sectorial de Investigación Científica (CISC) of the Universidad de la República (Udelar), Uruguay. All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ferrando, L., Bellini, M.I. & Fernández-Scavino, A. Differential response of denitrifying and diazotrophic soil populations to short and long-term exposure of glyphosate and atrazine. Environmental Sustainability 6, 229–242 (2023). https://doi.org/10.1007/s42398-023-00270-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42398-023-00270-z