Abstract
Plant diseases induce visible modifications on leaves with the advance of infection and colonization, thus altering their spectral reflectance pattern. In this study, we evaluated the visible spectral region of symptomatic leaves of five plant diseases: soybean rust (SBR), Calonectria leaf blight (CLB), wheat leaf blast (WLB), Nicotiana tabacum-Xylella fastidiosa (NtXf), and potato late blight (PLB). Ten spectral indices were calculated from the RGB channels (red, green, and blue) of images of leaves varying in percent severity, which were obtained under controlled lighting and homogeneous background. Image processing was automated for background removal and pixel-level index calculation. Each index was averaged across pixels at the leaf level. We found high levels of correlation between leaf severity and the majority of the spectral indices. The most highly associated spectral indices were overall hue index, visible atmospherically resistant index, normalized green red difference index, primary colors hue index, and soil color index. The leaf-level mean value of each of the ten indices and digital numbers on the RGB channels were gathered and used to train boosted regression tree models for predicting the leaf severity of each disease. Models for SBR, CLB, and WLB achieved high prediction accuracies (>97%) on the testing dataset (20% of the original dataset). Models for NtXf and PLB had prediction accuracies below 90%. The performance of each model may be directly related to the symptomatology of each disease. The method can be automated if the images are obtained under controlled light and homogeneous background, but improvements should be made in the method for using field- or greenhouse-acquired images which would require similar conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plant diseases induce visible modifications (symptoms or signs) on leaves with the advance of pathogenesis, thus altering their spectral reflectance pattern. The assessment of such spectra on plants, either visually or aided by image analysis software, has long been used in the field of phytopathometry for several purposes including disease detection, classification, and quantification (Barbedo 2016a; Bock et al. 2020).
The simplest image analysis-based approach used to determine the percent of visibly diseased (symptomatic) areas is to determine the total leaf area of an image and manually segment it via binarization into diseased and non-diseased areas (Lamari 2008; Bock et al. 2009). These areas are individually measured and the diseased:total (total = diseased + healthy) area ratio gives the proportion (or percentage) of the diseased area. This task is commonly performed by means of several image analysis software capable of thresholding images (Del Ponte et al. 2017). The method is considered the best approximation to determining the actual value of severity (Bock et al. 2009, 2016). To speed up a laborious and time-consuming manual annotation process at a leaf-by-leaf basis, programs have been written to automate the thresholding via batch processing of images (Bock et al. 2009; Stewart and McDonald 2014; Stewart et al. 2016; Karisto et al. 2017) that are available in commercial or open-source software such as Assess (Lamari 2008), ImageJ (Schneider et al. 2012), Quant (Vale et al. 2003), or custom-made ones (Barbedo 2016a). More recently, artificial intelligence algorithms have been proposed to predict categories of severity on individual leaves, not directly the percent values (Bock et al. 2020). The latter case has been possible using convolutional neural networks for semantic segmentation, a method that allows prediction of the binary classes at the pixel-level, and then the percent area affected can be obtained (Liang et al. 2019; Esgario et al. 2020; Gonçalves et al. 2020).
Regardless of the method, the images used in those studies are obtained using portable cameras or flatbed scanners (Barbedo 2016a). The latter is considered ideal given the uniform conditions during image acquisition, but the leaves should be detached and destroyed. While cameras can take in-field images, the issues related to nonstandard image acquisition arise during image processing, including the presence of complex target backgrounds that should be removed during processing (Barbedo 2016a).
The simplest and cheapest options of devices can simply capture the visible light from the visible electromagnetic spectrum (380 to 750 nm) and produce images from three spectral bands, red (564–580 nm), green (534–545 nm), and blue (420–440 nm), hence the RGB images, in which each pixel color is a “mixture” of the three bands. More sophisticated and currently more expensive cameras/sensors allow capturing more wavelengths in other zones of the electromagnetic spectrum, such as the red-edge (~700 nm) and the near-infrared (750–900 nm) bands. These images are called multispectral images. Hyperspectral imaging is performed with sensors that capture several bands (250–2500 nm) with fine wavelength resolution within the electromagnetic spectrum (Hagen and Kudenov 2013; Mahlein 2016; Barbedo 2016a; Bock et al. 2020).
The spectral indices are used to summarize the data of those several bands into a single value per pixel. These spectral indices are calculated by a mathematical formulation of two or more bands. The most common spectral index used to evaluate crop health is the Normalized Difference Vegetation Index (NDVI), which is calculated from the red and infrared bands (Rouse et al. 1974). In fact, the use of spectral indices has become routine in studies focusing on detection and quantification of plant diseases, but in the majority of cases using UAV- or aircraft-mounted multi- or hyperspectral imaging at field level (Mahlein 2016; Barbedo 2016a; Bock et al. 2020). RGB imaging has already been used for disease severity prediction in the field (Sugiura et al. 2016), but it has the disadvantages of being very sensitive to variations on conditions during image acquisition, such as lighting and shades, which may be the reason for its low accuracy and lack of use at the field. However, RGB imaging can be more useful for research purposes in which images are obtained under controlled conditions. For example, an automated procedure for research purposes, based on image thresholding, was developed to segment leaf images obtained by a flatbed scanner and measure the severity (and other traits) of more than 20,000 wheat leaves with Septoria leaf blight using the ImageJ software (Karisto et al. 2017).
The majority of studies regarding the prediction of disease severity on leaves using RGB images focused on evaluating machine learning algorithms to segment leaf images and calculate severity from the ratios between areas (Barbedo 2016b; Liang et al. 2019; Esgario et al. 2020; Gonçalves et al. 2020). To our knowledge, no previous work has studied the relationship between RGB-based spectral indices (RGBSI) and disease severity at the leaf level. We hypothesize that different RGBSIs, depending on the symptomatic features, are good candidates for predictors of disease severity at the leaf level since they are a function of the RGB bands. To test this hypothesis, we gathered image datasets of symptomatic leaves from five diseases. We studied the correlation between the disease severity and red, green, and blue channels intensity as well as ten RGBSIs extracted from the images. Furthermore, we use all RGBSI and the RGB data as predictors (13 predictors) in a boosted regression trees (BRT) model to predict the disease severity for each one of the datasets.
Material and methods
Image databases
We gathered five image datasets of different foliar diseases of different plants (Table 1), all previously annotated for percent area affected: soybean rust (Phakopsora pachyrhizi), Calonectria leaf blight (Calonectria pteridis) of eucalypts, Nicotiana tabacum-Xylella fastidiosa, wheat leaf blast (Pyricularia oryzae), and potato late blight (Phytophthora infestans growth and sporulation in leaf disks).
Diseased leaves were obtained from artificially inoculated plants (or leaf discs of 15-mm diameter for the case of Potato late blight) or from field surveys (for soybean rust). Image digitization was done under controlled conditions of luminosity (24-bit pixel depth) using either a flatbed scanner or stereo microscope-coupled camera device (for the potato late blight). The actual or reference severity of each image was measured manually using specific software (Table 1) for thresholding images and segmenting images into symptomatic and asymptomatic areas. Hence, disease severity was calculated by the percent leaf area presenting disease symptoms or pathogen growth and sporulation.
Overall approach
Our approach (Fig. 1) consisted of processing leaf images of each dataset sequentially in an automated way to obtain the mean value of RGB data (digital number on the red, green, and blue channels) and RGBSI for each image. As each image is loaded, the number of pixels is decreased to reduce computational cost. Next, the image’s background is removed using a color threshold, and the RGBSI is calculated for each pixel. The mean values of each index and the red, green, and blue intensities are aggregated (averaged) from all leaf pixels. After obtaining the image-derived data, we studied the relationship between the indices and the actual severity. Furthermore, we split the datasets into training and testing. Boosted regression trees models were trained for each disease, and we evaluated their prediction accuracy using the testing dataset (Fig. 1).
Digital image processing
Image manipulation
Images were loaded sequentially into the R software (R Core Team 2020) using the function stack() from the R package raster (Robert 2020) and further converted to a raster object. To speed up computations, the raster resolution was decreased in a ratio following an aggregation factor, which is the number of cells/pixels in each direction (horizontally and vertically) that is aggregated into one single cell/pixel. Aggregation factors of 4 for 96 dpi, 7 for 300 dpi, and 10 for 600 dpi images were used so the images ended up with ~6 dpi after aggregation (Table 2).
Leaf background removal
The background of the leaf was removed using a mask obtained from the original image. To classify the image into leaf (or leaf disc) and background, the masks were created by setting a threshold value on the digital number (DN) of the blue channel, for SBR, CLB, WLB, and NtXf images, and on the DN of the red channel for PLB images. The choice of the channel for thresholding was based on evaluating the channels that displayed the higher contrast between leaf and background. After manually evaluating different values of thresholds, a value was chosen for each dataset (Table 2). The low value in the threshold for PLB is due to the background color being black, and consequently leading to DN values to zero, differently from the other datasets with brighter backgrounds. The function fieldMask() from the R package FIELDimageR (Matias et al. 2020) was used to create the masks. Finally, the background was removed using the masks and the original image into the function mask() from the R package raster.
RGB-based spectral indices
Because the images were obtained under constant and controlled light conditions, we did not transform the DN values into reflectance since they would be proportional under such conditions. Therefore, the spectral indices were calculated directly from the DN of each respective channel. Ten RGBSI (Table 3) were calculated for each image at the pixel level. For this purpose, the function fieldIndex() from the FIELDimageR package was used. This function extracts the DN of each channel at the pixel and calculates different spectral indices. Since the fieldIndex() function does not have a built-in equation to calculate the grayscale values, it was calculated directly from the red, green, and blue DN values extracted from the raster object. Finally, the mean value of each RGBSI was calculated by averaging the values among all leaf pixels. The mean DN values of red, blue, and green channels were also calculated for each image.
All steps described here were automated to process all images sequentially. After the images were processed, the mean values of the RGBSI, RGB DNs, and actual severity data were put all together into a single dataset (one for each disease) for further data analysis and modeling.
RGB intensity distribution
Because DN on the red, green, and blue channels could be extracted from images, we selected two representative leaves of each one of the five datasets, one leaf with low severity and other with high severity. The purpose was to depict the distribution of the pixel intensity (given by the DN) of each channel on two different levels of severity. Distribution functions were constructed and plotted for visualization.
Correlation analysis
The relationship between the RGB-based spectral indices and disease severity was evaluated using Spearman’s rank correlation coefficient. This method was chosen due to its robustness and because we could not assume normality in the multivariate distribution. The cor.test() function from the R software was used to estimate the correlation parameters and associated P values.
Boosted regression trees
The main objective here was to fit a model to predict disease severity at the leaf level using the mean values of spectral indices and red, green, and blue mean DNs from each image as predictors, which added up to 13 predictors. We used boosted regression trees (BRT) models to build a predictive model linking these 13 predictors to disease severity. The BRTs were chosen due to their high predictive power for either regression or classification problems. The boosting algorithm is similar to the random forest method. The random forest method fits individual trees for random subsets of the original dataset. However, in boosting, the trees are fitted sequentially, in which each tree is fitted focusing on the data that were poorly modeled by the previous trees (James et al. 2013).
In the BRT algorithm some hyperparameters are important: (1) The learning rate or the shrinkage parameter (λ) defines the rate the method learns from the errors of the previous trees; (2) the interaction depth (d), which is the number of splits in the trees. This is because the trees are constructed using only a subset of d predictor variables; (3) the optimal number of trees (B), which needs to be estimated because a large number of trees might cause overfitting; (4) the minimum number of observations (MNO) in the terminal nodes of the trees; and (5) the bag fraction (bf), which defines the fraction of the training subset of the observations randomly picked to propose the next tree.
The computational procedure was done using the R software, where the BRT models were fitted using the function gbm() from the gbm package from R (Greenwell et al. 2020). We randomly split the datasets into training and validation data, in which 80% of the observations were used for model training. Predictions were performed using the R function predict() with the validation dataset.
Prediction accuracy metrics
The concordance between the BRT predictions and the actual/reference severity measured from each image, two continuous variables, was evaluated using Lin’s concordance correlation coefficient (CCC) analysis (Lin 1989). Lin’s CCC is given by ρc = r Cb, where r is the product-moment correlation coefficient between actual (ya) and predicted (yp) severity. Cb is the overall bias. It indicates the level of disagreement between the best fitting line (regression line between ya and yp) and the perfect agreement line (intercept 0 and slope 1). Cb is calculated from two bias parameters: the location shift parameter (u) and the scale shift (υ). u > 0 indicates that the regression between ya and yp produces a slope equal to 1 and an intercept different to zero. On the other hand, υ ≠ 1 indicates that the slope is different from 1 (Lin 1989; Madden et al. 2007). While r is a measure of precision, Cb is a measure of accuracy. In addition, we calculated the root mean square error (RMSE) between the actual and predicted severity as a measure of concordance. Lin’s CCC parameters were estimated using the function CCC() from the DescTools (Signorell 2020) package, while the RMSE was calculated using the function RMSE() from the caret package (Kuhn 2020).
Model selection
We tested a total of 192 combinations of the hyperparameters: shrinkage (λ = {0.001, 0.01, 0.1, 0.3}), interaction depth (d = {0.001, 0.01, 0.1, 0.3}), MNO (MNO = {5, 10, 15}), and bag fractions (bf = {0.5, 0.65, 0.8, 1}). The best models were chosen based on the calculated value of Lin’s CCC, ρc (Lin 1989), between the estimated values of severity for the validation dataset (remaining 20% of the original dataset).
Code availability and reproducibility
All analyses were performed using the R software. Scripts used for data analysis, including image processing, correlation analysis, and BRT model fitting were documented using Rmarkdown, and a website containing it can be freely accessed at git.io/JqUZw.
Results
Distribution of RGB intensity on images
Leaf images with low and high disease severity, defined arbitrarily for each disease, showed distinguishable patterns of the DN (pixel intensity, PI) distributions on the RGB channels. RGB images used to represent low and high severity levels are depicted in Supplementary figure 1. The low severity images showed lower PI on the blue and red channels than in the green channel (Fig. 2). However, a shift (increase) in the PI distribution of the red channel was noticed with the high severity category. For CLB, the mean PI value of the red channel of the high severity was much higher than in the green channel (Fig. 2B). Higher values were also observed on the green channel PIs for SBR, compared to the low-severity images (Fig. 2A). The variance of PI distributions on the green channel of high-severity images was quite higher for all diseases. There was a higher mean and variance of the PI distributions of the blue channel on high-severity images of CLB, NtXf, WLB, and mainly of PLB (Fig. 2).
Distribution of DN values or pixel intensity on red, blue, and green channels for low and high severity of representative images of soybean rust (low: 2.13%; high: 55.77%), Calonectria leaf blight (low: 1.74%; high: 77.46%), Nicotiana tabacum-Xylella fastidiosa (low: 2.64%; high: 75.82%), wheat leaf blast (low: 0.28%; high: 77.08%), and potato late blight (low: 0%; high: 82.74%) image datasets, depicted in each row, respectively. The images used for each category are displayed in Supplementary figure 1
RGBSI and severity
The majority of the RGBSI as well as DN values on the red, green, and blue channels were significantly associated with disease severity of all diseases (Fig. 3). The highest (absolute) values of Spearman’s rank correlation coefficient (r) were obtained between HUE and SBR (r = 0.995), HUE and CLB (r = 0.982), HUE and NtXf (r = 0.694), GLI and WLB (r = −0.886), and BGI and PLB (r = 0.899). Some RGBSI and color channels were not significantly (P > 0.05) associated with severity: the green channels for CLB and NtXf, SCI, NGRDI, VARI, and HUE for PLB (Fig. 3). The GLI, NGRDI, and VARI indices presented negative r values for all diseases. HUE is the index that seems to be directly correlated to the increase in disease severity on leaves with r values higher than 0.69, besides for PLB. Scatter plots with the relationship between severity and each variable (RGB channels and RGBSI) were generated (Supplementary figures 2 to 6). The relationship between severity and image attributes is quite variable. Some cases depict a linear relationship, such as HUE and SBR or CLB severity, while in other cases there is a nonlinear relationship, such as NGRDI for SBR, WLB, or CLB severity. We also notice different levels of within predictor correlations (Supplementary figure 7), mainly between NGRI and VARI, BGI and BI, or HUE and SCI, for instance.
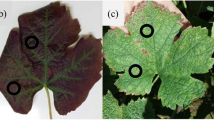
Representative leaves with the most correlated RGBSI calculated for each pixel are presented in Fig. 4 (mid column). It is possible to visualize a difference in RGBSI values comparing symptomatic and asymptomatic areas (Fig. 4). The distribution of reference severity values varied from 0.04 to 89.6% (median: 20.10%) for SBR, 0.8 to 95.4% (median: 9.8%) for CLB, 0.4 to 83.9% (median: 19.0%) for NtXf, 0 to 99.9% (median: 49.6%) for WLB, and 0 to 88.8% (median: 52.1%) for PLB (Fig. 4 right column).
Background-removed leaf images (first column), images with RGB-based indices more correlated to the severity (second column), and histogram of actual severity/sporulation distribution of each image dataset (third column). Each row corresponds to the following datasets sequentially: soybean rust, Nicotiana tabacum-Xylella fastidiosa, Calonectria leaf blight, wheat leaf blast, and potato late blight
BRT model selection
The models with higher accuracy (CCC) among the 192 models tested for each dataset have their hyperparameters shown in Table 4. Shrinkage (λ) or the learning rate was either 0.01, 0.1, or 0.3, with the lower value in the models for SBR and WLB. The models for SBR, CLB, NtXf, and CLB had lower splits in trees (d); on the other hand, the model with the longest trees was the WLB model, with 5 splits. All the models had MNO = 5. The model with the highest number of trees was the model for WLB (4988 trees), while the fewest was for NtXf (19 trees). The best five models, among all 192 models tested for each disease, with the lowest Lin’s concordance correlation coefficient (CCC) for predicting severity on the test dataset are shown in Supplementary Table 1.
Prediction accuracy
All 192 models were evaluated for their accuracy in predicting leaf-level severity of the testing dataset. The model with higher accuracy was the model for SBR (Table 5), which displayed ρc = 0.995. This was also the one with a lower prediction error (RMSE = 1.92 p.p.). In fact, the models for SBR, CLB, and WLB had very good overall performance, with low prediction errors (RMSE < 8.0 p.p.) and high accuracy (ρc > 0.97). Models for NtXf and PLB had the poorest performances among all the models. Both models had prediction errors higher than 8 p.p. and overall accuracy (ρc) lower than 0.9 (Table 5). Model precision values (Pearson’s r) were directly related to the RMSE, and consequently, the most precise model was SBR’s and the least precise model was PLB’s.
The most biased models were those for NtXf and PLB. These models tended to overestimate severity at low levels of severity and underestimate at high levels of severity (Fig. 5E). On the other hand, models for SBR, CLB, and WLB were the most unbiased models with regression curves very close to the perfect agreement and bias components (υ and ) very close to the ideal values (Fig. 5A, B, and C; Table 5).
Scatter plot for the relationship between actual and boosted regression trees model predictions of percent severity for (A) soybean rust, (B) Calonectria leaf blight, (C) wheat leaf blast, (D) growth and sporulation of potato late blight on leaf discs, and (E) Nicotiana tabacum-Xylella fastidiosa. The red solid line represents the linear model produced from the regression between actual and predicted values. The black dashed line represents a perfect agreement between the x- and y-axis. The CCC (concordance correlation coefficient) and RMSE (root mean square error) represent accuracy metrics. The closer the CCC to one, the more accurate and the closer the RMSE to zero, the better
Predictor’s relative influence
The BRT models assembled used all 13 predictor variables. However, the relative influence (RI) of each predictor varied among models (Fig. 6). The RI is a scaled measure (0 to 100%) of the contribution of each variable in decreasing the mean squared error loss function. The most influential variable for SBR, CLB, and NtXf was HUE, with relative influence of 87.8%, 94.0%, and 45.0%. For WLB, VARI had the highest contribution (28.4%), while for PLB, BGI had the highest RI (39.9%). Overall, predictors with high RI values also had a high Spearman’s correlation coefficient (Supplementary figure 8). The BRT model with the lower number of predictors with RI > 1% was the CLB model (2 predictors), while the BRT model for WLB had 9 predictors with RI > 1%. The number of predictors with RI > 1% was three for SBR, seven for NtXf and PLB, and nine for WLB (Fig. 6).
Discussion
In this study, a strong association was found between whole leaf percent severity determined by image analysis (assumed as actual severity) and RGBSI and the red, green, and blue DNs from digital images obtained under controlled conditions. The gathering of these variables, 13 in total, into boosted regression trees models allowed us to obtain accurate predictions of leaf severity for most disease datasets. The RGBSI that presented the highest absolute values of Spearman’s rank correlation |r| to severity were HUE, VARI, NGRDI, HI, and SCI, which presented |r| values >0.8 for at least two diseases. Consequently, these predictors exerted high relative influence values on the assembled BRT models for predicting severity. The DNs on the RGB channels were also correlated with severity, mainly the blue and red channels.
The prediction accuracy of the BRT models assembled here ranged from 0.86 (NtXf) to 0.99 (SBR). The variation in Lin’s concordance values (CCC) values is likely due to the differences among the characteristics of the symptoms for each respective disease, which reflects directly on their chromatic patterns. For SBR, CLB, and WLB, predictions by the BRT models were highly accurate (CCC >0.97). These diseases produce visibly contrasting symptomatic areas (necrosis or chlorosis) on leaves, which drastically affects the RGB distribution, with a major increase on the DN values on the red channel (Fig. 2A, B, and D). However, even with the same characteristics of necrosis and the same increase on the DN on the red channels, the accuracy of the predictions by the BRT model for the Xylella symptoms on tobacco plants was no higher than 0.87.
Different from the others, the PLB dataset was composed of leaf discs with growth and sporulation of P. infestans (Supplementary figure 1-J). As the severity increases, the leaf disc becomes covered with a white mycelial mass, which may explain the distribution of the DN values on RGB channels that are completely different from the other diseases (Fig. 2E). For PLB, the major shift was observed on the blue channel as severity increased (Fig. 2E), which was reflected in its correlation to severity (Fig. 3). Another characteristic observed is that in some leaves, as the severity increases, they become yellowish and some necrotic areas might appear, a pattern not observed among all leaves.
Among all diseases studied here, standard area diagrams (SADs) were developed and validated for assessing severity on SBR, WLB, and NtXf, and hence accuracy of visual estimates of severity was available for comparison with our results (Rios et al. 2013; Franceschi et al. 2020; Pereira et al. 2020). No such data are currently available for CLB and PLB, but experiments are underway. The accuracy of the unaided estimates of severity (meaning that raters were blindly assessing severity) ranged from 0.8 to 0.84 for SBR and WLB, while ranging from 0.63 to 0.78 among different groups that evaluated NtXf. The training with the SAD used as an aid during the assessment resulted in an increase in accuracy to 0.96 for SBR and WLB and 0.82 and 0.89 for NtXf (Rios et al. 2013, Franceschi et al. 2020, Pereira et al. 2020). Our predictions are as high or more accurate than the accuracy of visual estimates obtained using SADs for these diseases for an “average” rater, meaning that the system is equivalent to a “best rater” estimating visually.
Accuracy of our prediction and that obtained by visually assessing NtXf with the aid of a SAD were never higher than 0.9 on average among the four groups of raters. A small percentage of raters were able to achieve accuracy higher than 0.9. Two hypotheses can be drawn. First, NtXf symptoms depict visual features that are not clearly perceived, or overly perceived, during visual inspections. For example, NtXf symptoms are composed of small numerous necrotic punctuations over the leaf (Pereira et al. 2020) that appear to be difficult to visualize or distinguish from the asymptomatic tissue. Other leaves tend to become yellowish even with low levels of severity, maybe because of the X. fastidiosa systemic infection. The yellowing leaves might affect the perception of diseased areas. The same phenomenon may affect the visible spectral pattern of the “asymptomatic” leaf tissue. The second hypothesis is a low accuracy of the reference severity (the assumed actual severity), which is manually measured using color thresholding software. These hypotheses can be interrelated with the first since the symptomatic and asymptomatic areas are identified visually in all cases. The same set of actual severity is used to train our models and to validate results of visual accuracy.
The SBR dataset has been used recently to train convolutional neural network (CNN) architecture for the semantic segmentation of images and prediction of disease severity (Gonçalves et al. 2020). Differently from our work, their algorithms were trained for classifying each pixel of an image into symptomatic, asymptomatic, or image background, and then severity was calculated using diseased:total leaf area ratio. The authors reported prediction accuracies (CCC) ranging from 0.93 to 0.97. The advantage of their method, which is not possible using our approach, is the possibility to apply in field-acquired images that present a more complex background and uncontrolled light. In fact, in our method, images should be acquired under controlled conditions of luminosity and homogeneous backgrounds, preferentially with a color that is different from the leaf and disease symptoms (e.g., white or blue). Our method is more applicable when conditions for image acquisition can be as controlled as possible. However, the use of a spectralon (reference reflectance spectra) could be used in the field to correct for light sources (Sankaran et al. 2012; Gold et al. 2019). Another possible solution is to use a portable flatbed scanner, which enables lighting control and homogeneous background conditions (Zhang et al. 2019). Due to the models assembled here being disease-specific and empirically derived, specific training will be needed for other datasets or when new images obtained under similar conditions are included in the dataset. Other limitations include variations in the nutritional condition or occurrence of other abiotic stress on leaves that might impact heterogeneity of the healthy tissue spectra. Coinfected leaves with two or more different pathogens might display unpredicted spectral responses. As previously mentioned, the light source during image acquisition must be the same for images. We also recommend that the same device should be used for acquiring all images since each piece of equipment has a specific hardware configuration for capturing light on RGB channels. Another disadvantage of our method, mainly when compared to segmentation methods, which make use of hand-made masks from images diseased leaves, is the difficulty to make visual inspections on images to identify problems in symptom recognition (or quantification), differently from using binary mask for segmentation which enables us to make post-processing comparison of the resulting masks to the original masks.
Future research should focus on developing methods and devices for fast leaf digitization either in-field or in-house with controlled lighting conditions. Devices can be coupled with software for automated image processing (background removal) and online severity prediction. Moreover, other regions in the electromagnetic spectrum should be evaluated, such as near-infrared, infrared, or thermal regions, as for evaluating the relationship between leaf disease severity and other spectral indices, which has been currently implemented at the whole plot level in the field (Mahlein 2016).
Data availability
The datasets generated during and/or analyzed during the current study are available in the Open Science Framework repository (https://doi.org/10.17605/OSF.IO/CYHA7).
References
Barbedo JGA (2016a) A review on the main challenges in automatic plant disease identification based on visible range images. Biosystems Engineering 144:52–60
Barbedo JGA (2016b) A novel algorithm for semi-automatic segmentation of plant leaf disease symptoms using digital image processing. Tropical Plant Pathology 41:210–224
Bock CH, Cook AZ, Parker PE, Gottwald TR (2009) Automated image analysis of the severity of foliar citrus canker symptoms. Plant Disease 93:660–665
Bock C, Chiang KS, Del Ponte EM (2016) Accuracy of plant specimen disease severity estimates: concepts, history, methods, ramifications and challenges for the future. CAB Reviews 11:1–21
Bock CH, Barbedo JGA, Del Ponte EM, Bohnenkamp D, Mahlein A-K (2020) From visual estimates to fully automated sensor-based measurements of plant disease severity: status and challenges for improving accuracy. Phytopathology Research 2:9
Del Ponte EM, Pethybridge SJ, Bock CH, Michereff SJ, Machado FJ, Spolti P (2017) Standard area diagrams for aiding severity estimation: scientometrics, pathosystems, and methodological trends in the last 25 years. Phytopathology® 107:1161–1174
Escadafal R, Belghith A, Moussa HB (1994) Indices spectraux pour la dégradation des milieux naturels en Tunisie aride. 6ème Symp. Int. Mesures Physiques et Signatures En Télédétection
Esgario JGM, Krohling RA, Ventura JA (2020) Deep learning for classification and severity estimation of coffee leaf biotic stress. Computers and Electronics in Agriculture 169:105162
Franceschi VT, Alves KS, Mazaro SM, Godoy CV, Duarte HSS, Ponte EMD (2020) A new standard area diagram set for assessment of severity of soybean rust improves accuracy of estimates and optimizes resource use. Plant Pathology 69:495–505
Gitelson AA, Kaufman YJ, Stark R, Rundquist D (2002) Novel algorithms for remote estimation of vegetation fraction. Remote Sensing of the Environment 80:76–87
Gold KM, Townsend PA, Larson ER, Herrmann I, Gevens AJ (2019) Contact Reflectance spectroscopy for rapid, accurate, and nondestructive Phytophthora infestans clonal lineage discrimination. Phytopathology® 110:851–862
Gonçalves J de P, Pinto F de A de C, Queiroz DM de, Villar FM de M, Barbedo JGA, Ponte EMD (2020) Deep learning models for semantic segmentation and automatic estimation of severity of foliar symptoms caused by diseases or pests. OSF Preprints July 9. https://doi.org/10.31219/osf.io/wdb79.
Greenwell B, Boehmke B, Cunningham J, Developers GBM (2020) gbm: generalized boosted regression models. R package version 2.1.8. https://CRAN.R-project.org/package=gbm.
Hagen NA, Kudenov MW (2013) Review of snapshot spectral imaging technologies. Optical Engineering 52:090901
James G, Witten D, Hastie T, Tibshirani R (2013) An introduction to statistical learning: with applications in R. Springer, New York
Karisto P, Hund A, Yu K, Anderegg J, Walter A, Mascher F, BA MD, Mikaberidze A (2017) Ranking quantitative resistance to Septoria tritici blotch in elite wheat cultivars using automated image analysis. Phytopathology® 108:568–581
Kuhn M (2020) caret: Classification and Regression Training. R package version 6.0-86. https://CRAN.R-project.org/package=caret.
Lamari L (2008) Assess 2.0: image analysis software for disease quantification. The American Phytopathological Society, Saint Paul, MN
Liang Q, Xiang S, Hu Y, Coppola G, Zhang D, Sun W (2019) PD2SE-Net: computer-assisted plant disease diagnosis and severity estimation network. Computers and Electronics in Agriculture 157:518–529
Lin LI-K (1989) A concordance correlation coefficient to evaluate reproducibility. Biometrics 45:255–268
Louhaichi M, Borman MM, Johnson DE (2001) Spatially located platform and aerial photography for documentation of grazing impacts on wheat. Geocarto International 16:65–70
Madden LV, Hughes G, van den Bosch F (2007) The Study Of Plant Disease Epidemics. APS Press, St. Paul
Mahlein A-K (2016) Plant disease detection by imaging sensors—parallels and specific demands for precision agriculture and plant phenotyping. Plant Disease 100:241–251
Mathieu R, Pouget M, Cervelle B, Escadafal R (1998) Relationships between satellite-based radiometric indices simulated using laboratory reflectance data and typic soil color of an arid environment. Remote Sensing of Environment 66:17–28
Matias FI, Caraza-Harter MV, Endelman JB (2020) FIELDimageR: an R package to analyze orthomosaic images from agricultural field trials. The Plant Phenome Journal 3:e20005
Pereira WEL, de Andrade SMP, Del Ponte EM, Esteves MB, Canale MC, Takita MA, Coletta-Filho HD, De Souza AA (2020) Severity assessment in the Nicotiana tabacum-Xylella fastidiosa subsp. pauca pathosystem: design and interlaboratory validation of a standard area diagram set. Tropical Plant Pathology 45:710–722
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Richardson AJ, Wiegand CL (1977) Distinguishing vegetation from soil background information. Photogrammetric Engineering and Remote Sensing 43:1541–1552
Rios JA, Debona D, Duarte HSS, Rodrigues FA (2013) Development and validation of a standard area diagram set to assess blast severity on wheat leaves. European Journal of Plant Pathology 136:603–611
Robert JH (2020) raster: Geographic Data Analysis and Modeling. R package version 3.3-13. https://CRAN.R-project.org/package=caret.
Rouse J, Haas RH, Schell JA, Deering DW, others (1974) Monitoring vegetation systems in the Great Plains with ERTS. NASA special publication 351:309
Sankaran S, Ehsani R, Inch SA, Ploetz RC (2012) Evaluation of visible-near infrared reflectance spectra of avocado leaves as a non-destructive sensing tool for detection of laurel wilt. Plant Disease 96:1683–1689
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to ImageJ: 25 years of image analysis. National Methods 9:671–675
Signorell A (2020) DescTools: Tools for descriptive statistics. R package version 0.99.38. https://cran.r-project.org/package=DescTools.
Stewart EL, McDonald BA (2014) Measuring quantitative virulence in the wheat pathogen Zymoseptoria tritici using high-throughput automated image analysis. Phytopathology® 104:985–992
Stewart EL, Hagerty CH, Mikaberidze A, Mundt CC, Zhong Z, BA MD (2016) An improved method for measuring quantitative resistance to the wheat pathogen Zymoseptoria tritici using high-throughput automated image analysis. Phytopathology®, 106:782–788
Sugiura R, Tsuda S, Tamiya S, Itoh A, Nishiwaki K, Murakami N, Shibuya Y, Hirafuji M, Nuske S (2016) Field phenotyping system for the assessment of potato late blight resistance using RGB imagery from an unmanned aerial vehicle. Biosystem Engineering 148:1–10
Tucker CJ (1979) Red and photographic infrared linear combinations for monitoring vegetation. Remote Sensing of the Environment 8:127–150
Vale FXR, Fernandes Filho EI, Liberato JR (2003) QUANT. A software for plant disease severity assessment. In: Close R, Braithwaite M, Havery I (eds) Proceedings of the 8th International Congress of Plant Pathology, New Zealand, 8th. Sydney, NSW, Australia
Zarco-Tejada PJ, Berjón A, López-Lozano R, Miller JR, Martín P, Cachorro V, González MR, de Frutos A (2005) Assessing vineyard condition with hyperspectral indices: leaf and canopy reflectance simulation in a row-structured discontinuous canopy. Remote Sensing of the Environment 99:271–287
Zhang L, Wang L, Wang J, Song Z, Rehman TU, Bureetes T, Ma D, Chen Z, Neemo S, Jin J (2019) Leaf scanner: a portable and low-cost multispectral corn leaf scanning device for precise phenotyping. Computers and Electronics in Agriculture 167:105069
Acknowledgements
The first and second authors are thankful to the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES) for their scholarship. The third author is thankful to the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for providing scholarships. The fourth author received a research fellowship from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq)/Brazil.
Funding
This study was financed in part by the “Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil” (CAPES)—Finance Code 001.
Author information
Authors and Affiliations
Contributions
KSA conceived the idea, conducted data analysis, and wrote the manuscript; MG, JPA, MFQ planned, coordinated, and performed the experiments for obtaining the images; and ESGM, RFA, and EMD supervised the project and revised the manuscript. All authors provided feedback and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 1503 kb)
Rights and permissions
About this article
Cite this article
Alves, K.S., Guimarães, M., Ascari, J.P. et al. RGB-based phenotyping of foliar disease severity under controlled conditions. Trop. plant pathol. 47, 105–117 (2022). https://doi.org/10.1007/s40858-021-00448-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-021-00448-y