Abstract
In pigeonpea, resistance against vascular wilt disease was assessed based on leaf images captured throughred-green–blue (RGB) and chlorophyll fluorescence imaging sensors. At leaf level, wilt response in RGB images was characterized by changes in pixel intensities in red, green, and blue channels leading to variation in texture. Texture analysis based on gray level co-occurrence matrix (GLCM) was able to explain variation pattern between resistance and susceptible genotypes. Extracted texture features particularly contrast and energy were significantly different between the two genotype groups. Training of a neural network model for contrast and energy feature enabled genotype prediction with 79–98% accuracy. Healthy leaf area estimated based on photosynthetic or quantum efficiency (Fv/Fm > 0.75 as healthy) in chlorophyll fluorescence images, indicated significant variation (p < 0.05) between genotype groups at 10–25 days after inoculation (dpi). In susceptible genotype, healthy area was observed to decrease in significant proportion over time as compared to resistant type. Resistant genotype was less sensitive to infection as healthy leaf area (Fv/Fm > 0.75) remained unaffected between 10-25dpi.At canopy level, although differences in pixel intensity (Fv/Fm > 0.75) were noted between inoculated and healthy (mock) particularly in susceptible types but differences between inoculated susceptible and resistant type were non-significant (p > 0.05). Although trained ML algorithms for leaf and canopy level images resulted low accuracy (41–54%) in genotype classification but with large number of images captured later than 15 dpi expected to increase in accuracy. A protocol to facilitate non-invasive imaging techniques in association with machine learning tools is proposed over the tedious, time consuming and error-prone conventional screening method.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pigeonpea (Cajanus cajan (L) Millsp.) is an important food legume grown in semi-arid tropical and subtropical farming systems under varied agro ecological systems of Asia and Africa. It provides high quality vegetable protein to human beings and one of the sources for animal feed and firewood. Globally, the area and production of pigeonpea is increased from 2.86 million hectares (mha) to 6.8 mha with the production increase from 1.96 million tons (mt) in1980s to 5.4 mton 2016-17 (FAO STAT, 2018). India contributes major share of world legume production as more than 72% of pigeonpea is produced in India (Indiastat.com, 2018–19). It is grown in about 4.5 mha but productivity level is low (729 kg/ha) due to various biotic and abiotic constraints (Nene, 1980). Vascular wilt (Fusarium udum) is economically the most important soil borne disease, causes 30–100% loss of grain yield (Nene & Kannaiyan, 1982; Upadhyay & Rai, 1992). Wilt symptom is well characterized by epinasty, followed by flaccidity, chlorosis, vascular browning and necrosis of the terminal leaflets (Agrios, 2005). Infection starts through roots and the pathogen colonizes profusely in xylem vessels. The disease is widely prevalent particularly in medium and late duration varieties in most of the regions (Ahlawat et al., 2005; Okiror, 2002). Management of vascular disease is very difficult as cultural, chemical and biological measures to control are generally ineffective (Nene 1980; Upadhyayand Rai 1992). The most effective control strategy is the use of resistant cultivars (Nene & Kannaiyan, 1982). However, genotypic improvement is impaired due to lack of precise evaluation method and moreover inheritance of wilt resistance is largely unknown (Jain & Reddy, 1995; Parupalli et al., 2017; Saxena et al., 2012; Singh et al., 2016). Resistance evaluation through sick-plot method is laborious and error prone as heterogeneous soil environment affects inoculum concentration (Nene & Kannaiyan, 1982). Soil temperature plays important role in infection process and symptom expression often being not uniform evaluation process becomes lengthy and indecisive. A quick, reliable, automatic, easy and nondestructive method of high-throughput phenotypic technologies are urgently requiringfor precise detection and phenotyping of resistance (Rousseau et al., 2013; West et al., 2003 and Bock et al., 2010).
Host-pathogeninteraction is captured in images have a high potential in accurate detection, identification and quantification of diseases on different scales prior to visual symptoms (Mahlein et al., 2013). For fast and accurate detection as well as assessment of host–pathogen interaction image processing techniques have shown lots of prospects (Cui et al., 2009; Kai et al., 2011). Transformation of RGB color images based on hue, saturation, and intensity color model enables object detection, recognition and estimation of different features (Rafael, 2018). Generation of color co-occurrence matrix (CCM) and image textures are useful to identify or classify level of host–pathogen interactions (Huang, 2007; Ha et al., 2017). Image texture provides information in the spatial arrangement of colors or intensities in an image and the most frequently used approaches to detect and classify symptoms (Al-Saddik et al., 2018).Specifically, RGB images have been used for plant disease identification (Pydipati et al., 2006) and evaluation of resistance (Diaz-Lago et al., 2003).
In addition to texture analysis, monitoring photosynthetic activity in leaves can rapidly assess early changes in photosynthetic properties (Maxwell & Johnson, 2000; Scholes & Rolfe, 2009).Chlorophyll fluorescence imaging(Chl-FI) is a non-invasive, non-destructive method provides wealth of information on the timing and location of pathogen development as well as to understand the regulation of photosynthesis from leaf to crop scale, allowing phenotyping of plants (Rolfe & Scholes, 2010; Pérez-Bueno et al., 2019). Responses of the photosynthetic machinery to biotic stress (caused by pathogens) based on standard Chl-FI parameters (Fv/Fm, ΦPSII, qP and NPQ) has been utilized for evaluation of quite a large number of host–pathogen interactions (Mahlein et al., 2013; Pérez-Bueno et al., 2019, Scholes & Rolfe, 2009; Simko et al. 2012) and resistance evaluation (Chaerleet al., 2007; Rousseau et al., 2013). Particularly, Fv/Fm parameter is used to diagnose and assess diseases since it is significantly correlated with visual severity of the pathogenic infection. For tobacco mosaic virus on tobacco leaves (Balachandran et al., 1994; Chaerle et al., 2007) and bacterial infection in bean (Rousseau et al., 2013) and rice (Sebela et al., 2018). Change in Fv/Fm parameter was used to presymptomatic diagnosis and disease assessment. Characteristics behaviour of Fv/Fm parameter for comparatively large number of diseases caused by fungal infection is summarized (Pérez-Bueno et al., 2019). Hemileia vastatrix on coffee plants (Honorato Júnior et al. 2015), downy mildew on lettuce leaves (Bauriegel et al. 2014) and grapevine (Csefalvay et al., 2009) powdery mildew and leaf blight in wheat (Kuckenberg et al., 2009; Rios et al., 2017), Rhizoctoniasolani in rice (Ghosh and Kanwar P, 2017), Botrytis cinerea in rice and tomato (Berger et al., 2004; Sekulska-Nalewajko et al., 2019), Pythium irregulare in ginseng (Ivanov & Bernards, 2016) and Rosellinia necatrix in avocado (Granum et al., 2015) where decrease in Fv/Fm value in infected tissue as compared to healthy has been well documented. However, Fv/Fm and other parameters do not always offer clear differences between healthy and infected tissues or do so at late stages of the disease (Pineda et al., 2018). Such uncertainty associated with Chl-FI data requires stringent statistical/mathematical solutions to enhance differences in response evaluation. Machine learning algorithms (ML) are powerful and efficient tools in automation of model building process and iteratively learn from noisy data to gain insights without explicit programming (Nichols et al., 2019). ML has been used to identify patterns/genes/proteins involved in plant-pathogen interactions (Sperschneider et al., 2016) and identification of plant diseases (Kaundal et al., 2006; Mokhtar et al., 2015; Calderón et al. 2013).
Characterization of wilt response in terms of leaf symptoms captured in RGB and/or Chl-FI images particularly associated stress parameters may facilitate phenotyping of pigeonpeagermplasms. Further, use of machine learning algorithms is likely to assure accuratephenotyping and facilitate possible automation of the process. Development of assessment protocol for wilt response in pigeonpea based on non-invasive imaging devices is anticipated for precise identification or classification of susceptible or resistant genotypes required in crop phenomics.
In the current communication, vascular wilt response captured in leaf and canopy images was assessed to develop a protocol for phenotyping resistance in pigeonpea. A protocol based on integrated RGB and Chlorophyll fluorescence imaging with machine learning tools was applied and proposed for resistance screening in pigeonpea.
Material and Methods
Pigeonpeaseedling preparation for evaluation of resistance
Pigeonpea genotypes consist of two susceptible and two resistant along with eight genotypes where wilt response not reported were considered for resistance evaluation (Nene and Kannaiyan, 1982). Seeds of total of 12 genotypes consists of: susceptible types ICP2376 (Acc No: ICP2-376) and Gulyallocal (Local landrace); resistant types Asha (Acc no: ICP88719) and Maruti (Acc no: ICP8863); six cultivars BDN711 (Acc no: BDN 2004–3), BDN708 (Acc no: BDN 711 × ICPL 20,096), BDN 716 (Acc no: BDN 2008–7), BSMR-736 (Local landrace), Pusa992 (Selection from ICPL 90,306) and Dharmaraj (Acc no: GRG-811); and two TS3R ( Acc no: Maruti-2) and TS3 (Local landrace) were collected from the Agriculture Research Station, Badnapura (Maharashtra) Agriculture Research Station, Kalaburgi (Karnataka) and Division of Genetics, ICAR-Indian Agricultural Research Institute, New Delhi. Seeds were surface sterilized in 1% NaClO solution for 1 min and washed with distilled water before sowing. For sowing of seeds, earthen pots (dia 0.16 cm) filled with sterilized cocopeat and sand mixture (50:50) was used and 10 seeds were sown in each pot. Total of 100 plants for each genotype (10 pots) were raised in the greenhouse (National Phytotron Facility) maintained at 28 °C ± 1.5 °C.
Inoculum preparation -Fusarium udum spore suspension
The pathogen was isolated from the wilt infested pigeonpea samples (stem) collected from Gulbarga (hotspot for fusarium wilt diseases), Karnataka, India. Pathogenicity test was carried out through seedling inoculation and reaffirmation of wilt symptoms. Purity of the isolate was established through single spore isolation and confirmation of the pathogen was based on the composition of large hooked conidia along with sickle-shaped macroconidia and elliptical microconidia (Type specimen Fusarium udum maintained in ITCC, Indian Agricultural Research Institute New Delhi). The isolate was multiplied in potato-dextrose-agar plates (28° ± 2 °C). Spore suspension in freshly sterilized distilled water was prepared from six-days old culture adjusting concentration to 3.4 × 104 spore/mL and stored in deep fridge for short term use.
Seedling inoculation
Thirty days old seedlings were inoculated under greenhouse conditions (National Phytotron Facility) with F udum spore suspension (3.4 × 104/ mL). A 5 mL of suspension was poured into each pot removing upper surface soil layer around the individual seedlings to facilitate direct contact with the root zone. For mock inoculation, seedlings were treated with distilled water. After inoculation, the seedlings were incubated in the greenhouse (day-night length 14–10 h with 400–450 µmol m−2 s−1, temperature 28° ± 2 °C Day-night; relative humidity 74–77%, moisture content in pot soil below field capacity).
Assessment of wilt severity index (WSI) for evaluation of resistance in pigeonpea genotypes.
Inoculated plants were observed dpily for the appearance of wilt symptoms. Twenty inoculated plants from each genotypes were randomly selected for scoring wilt severity. Individual plants were scored in 0–4 scale (Hervás et al., 1995); 0- no visible symptoms; 1- slight yellowing or pale color in leaves normally topside of the plants; 2- leaf yellowing and drooping; 3- leaf shedding and stunted plants; and 4-most of the leaves shedding and finally drying whole plants. Wilt severity index (WSI) was calculated at 5, 10, 15, 20 and 25 days based on the formula:
- Si:
-
symptom severity
- Ni:
-
number of plants with Si symptom
- Nt:
-
total number of plants
- G:
-
maximum rating scale.
Median time (days) for the development of 20% WSI was calculated for each genotypes and designated as WSI20. WSI at 25 days after inoculation (dpi) was considered as terminal wilt severity TWSI. Resistance component for WSI20 and TWSI for each genotypes was estimated (Parlevli et 1979; Poland et al., 2009).
Resistance component for WSI20 (RSI) and TWSI (MSI) was estimated:
where, X = test genotype; C = susceptible reference.
ICP2376 was used as susceptible check. Using RSI and MSI as components, relative resistance (R) level for the genotypes was estimated (Parlevli et al., 1979, Savary et al., 2012):
R is a dimensionless relative resistance coefficient which varies between 0 and 1: 0 ≤ R ≤ 1 in which 1 corresponds to the highest level of resistance, while 0 corresponds to maximum susceptibility. Genotypes were grouped based on k-means clustering as well as the membership of known resistant and susceptible genotypes included in the study.
Digital image acquisition of leaf symptoms for RGB image analysis
At 15 dpi, ten trifoliate leaves were collected from susceptible (ICP2376 and Gulyal local) and resistant (Asha and Maruti) genotype. Ten trifoliate leaves also were collected from the mock-inoculated plants from the corresponding genotypes. A total of 80 leaves (40 inoculated and 40 mock-inoculated) were considered for RGB images. Images were captured through a flatbed scanner (HP Scanner 1136 M) at 600 dpi, adjusted to 3500 X 2500 pixels and saved in JPEG format. Red, green and blue color channels were separated from the images and mean pixel intensity as Rmean, Gmean and Bmean were determined through MATLAB 2021a (Mathworks, Natick, MA). Several parameters like \({R}_{mean}\) + \({G}_{mean}+{B}_{mean}\), \({R}_{mean}\) / (\({R}_{mean}+{G}_{mean}+{B}_{mean}),\) \({G}_{mean}\) / (\({R}_{mean}+{G}_{mean}+{B}_{mean}),\) \({B}_{mean}\) / (\({R}_{mean}+{G}_{mean}+{B}_{mean}),\) \({G}_{mean}\) / \({R}_{mean},\) \({B}_{mean}\) / \({R}_{mean},\) \({(B}_{mean}\)-\({G}_{mean})/({B}_{mean}+{G}_{mean})\) were estimated to compare color intensity difference between the infected and healthy images. For texture analysis, Haralick features (Haralick et al., 1973) were extracted in Gray Level Co-occurrence Matrix (GLCM) for Contrast, Correlation, Energy and Homogeneity using glcm function in MATLAB (2021a). GLCM was used to check how often pairs of pixels with specified values and spatial orientation occur in the images. Contrast (K) measures the local variations in gray level from a pixel to its neighbour in an image where, i and j are indices referring to the location of pixel (p) in the GLCM (Al-Saddik et al., 2018). It shows texture fineness.
Correlation (R): measures the linear dependence of gray-level in a co-occurrence matrix or in other words, correlation intensity between neighboring pixels
\({\mu }_{i}\) and \({\mu }_{j}\) are the averages of row i and column j in a GLCM, respectively \({\sigma }_{i }and {\sigma }_{i}\) are the standard deviations of row i and column j in a GLCM, respectively.
Energy (E) known as an angular second moment, it is simply the sum of squared elements in the GLCM, It measures the uniformity in an image
Homogeneity (H) is a measure of closeness of a distribution of elements in the GLCM to the diagonal. Homogeneity is unity for diagonal GLCM. This is the case where all the pixels in the original image have the same value as their neighbor.
The extracted texture features in GLCM for susceptible and resistant genotypes were statistically compared using unpaired t-test (Welch two sample tests, SPSS24) to examine the significant difference between the two genotype groups. For extracted features, an artificial neural network (ANN) classifier (back propagation with input layer, hidden layers and output layer) was trained for identification of wilt response in the two genotype groups. For training the network, five case studies were made dividing the dataset into 90, 85, 80, 75 and 70%. After training the model, remaining data set 10, 15, 20, 25 and 30% were used as validation set for unbiased evaluation of the network. Further a test dataset was also used to examine an unbiased evaluation of the final network model and to observe the error rate in prediction. Finally, selection of hyper parameters (neuron layers) in the network configuration was chosen.
Chlorophyll fluorescence measurement for estimation of maximum photosynthetic efficiency or quantum efficiency parameter (Fv/Fm).
Preparation of materials for chlorophyll measurement
For chlorophyll measurement 20 pots (polypropylene dia 0.16 cm) from each of the 12 genotypes were inoculated (at 30 days) with F udum spore suspension along with equal number of control set (mock-inoculation with sterilized distilled water). All the pots were maintained in the chamber having uniform light and temperature (28° ± 2 °C). For image acquisition at leaf level, ten random trifoliate leaves were picked up at 0, 5, 10, 15, and 20 dpi. For canopy level image, top view of the whole plant was considered only at 15 dpi. For mock inoculation, equal number of leaves and plants from all the genotypes were maintained.
Chlorophyll fluorescence measurement, image acquisition and processing
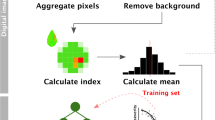
Chlorophyll fluorescence images were captured using Crop Reporter (Phenovasion Life Sciences, Wageningen, The Netherlands), a high-resolution multispectral imaging sensor installed at NanajiDeshmukh Plant Phenomics Center, IARI, New Delhi. Prior to image capture and acquisition, leaf and whole plant samples prepared were dark adapted for 15 min using chambers integrated with Scanalyzer 3D Phenotyping System (LemnaTec GmbH, Aachen, Germany). Later, time-lapse image of 24 frames per sample were captured within 1430 ms using sensor set up. The pixel intensity values of dark-adapted images were considered as F0 (minimum level of fluorescence). Subsequently, saturated pulse intensity (TF 800) of red-light flash with power LED 40 was used before capturing the fluorescence image. The maximum pixel intensity values from fluorescence images (Frame number 3 to 24) were considered as Fm (maximum level of fluorescence). The recorded images (1388 × 1038 spatial resolution) were processed using LemnaGrid software (LemnaTec GmbH). Raw RGB images (16-bit grey scale) were demosaiced using Adaptive Homogeneity Directed (AHD) algorithm to reconstruct a full color image from the incomplete output from the image sensor. The demosaiced pixels of these images were segregated into foreground and background pixels using normalized Fm intensity and Otsu thresholding filters. Edge noise was removed through erosion and dilatation steps before composing all parts identified as plant to one object. Grey calculators were used appropriately for calculating variable fluorescence value (Fv=Fm- F0), where the pixel intensity values of F0 was subtracted from pixel intensity values of Fm. Finally, maximum quantum yield of PSII photochemistry was derived as the ratio Fv/Fm and expressed as pixel-to-pixel information on the fluorescence image.
For comparison of wilt response between genotypes, proportion of healthy leaf area for each sample as a feature was estimated in HSV plane separately based on Fv/Fm values and apparent visual symptoms on leaves.
Experimental design and data analysis on maximum photosynthetic efficiency
At leaf level, wilt response in relation to maximum photosynthetic efficiency (derived from chlorophyll fluorescence images) in susceptible (ICP2376) and resistant (Maruti) genotypes was assessed. Photosynthetically healthy leaf area (Fv/Fm > 0.75) was estimated by image segmentation (on HSV color plane) selecting ten random leaves (each replication) to compare wilt response at 0–20 dpi. For comparison of healthy leaf area between the genotypes factorial ANOVA was performed using Generalized Linear Model (SPSS 24).
At canopy level (whole plant view), weighted pixel count was estimated as sum of the product between mid-point of interval (for Fv/Fm 0.70–0.80, 0.80–0.90 and 0.90–1.00) and pixel intensity divided by total pixel intensity. For estimation of weighted pixel count for each genotypes ten images from each of inoculated and mock samples were considered. Weighted pixel counts for the twelve genotypes were shown in tornado chart and for their comparison independent sample t-test was performed. Subsequently, relative pixel counts (%) for all the inoculated samples (genotypes) were fitted in polynomial curves. Polynomial curves (R2 = 0.98) were compared based on non-parametric Kolmogorov–Smirnov test (skewed data).
Image features and machine learning
Infected areas as features in Chlorophyll fluorescence images (leaf as well as whole plant view) were estimated by segmenting the images in HSV color plane based on Fv/Fm value comparing healthy image. Threshold value (Fv/Fm > 0.75) matched with healthy area and marked as brown (> 0.75), symptomatic as yellow (0.52 to < 0.75) and blue (< 0.52). The relative areas were estimated for all the 12 genotypes that are grouped under three categories (susceptible, tolerant and resistant based on resistance index estimates).
For classification of CFI images based on extracted features in each category of images (susceptible, tolerant and resistant), five machine learning algorithms (K-nearest neighbor, Support Vector Machine, Random Forest Classifier, Decision Tree Classifier and Naïve Bayes) were trained to classify the genotypes for developing prediction model. ML algorithms, trained with large collection of noisy data on relative leaf areas, were tested for their prediction accuracy after removing skewness and correlation from the dataset. To reduce skewness in the data points scalar transforms of relative leaf areas was performed by subtracting the mean and dividing by the standard deviation to shift the distribution to have a mean of zero and a standard deviation of one. To omit correlation, skewness, and outliers of the dataset PCA was performed.
Results
Wilt severity index (WSI) and grouping of genotypes
Typical wilt symptoms, pale yellowing and drooping of leaves, were noted in the inoculated plant as compared to mock inoculation. Time taken for expression of 20% wilt severity index (WSI20) indicated variations between the genotypes (Table 1 and Supplementary Fig. 1). The WSI20 was noted to vary from the minimum 8 to maximum 18 days. Terminal wilt severity (TWSI at 25 dpi) also shown variations between the genotypes as the minimum and maximum values were noted 9.3 and 97.5 respectively. Clustering based on the RSI and MSI, indicated genotypes could be grouped into three distinct as RSI and MSI were significant (p < 0.05) for three groups. Three groups were separated by the R values 0.71 and above, between 0.30 to 0.70and below 0.30. Based on cluster membership of the known resistance and susceptible genotypes three groups were designated as resistant (R values 0.71 and above), tolerant (between 0.30 to 0.70 tolerant) and susceptible (below 0.30 as susceptible).
It appeared that assessment of resistance based on leaf symptoms fairly correspond with the pattern of wilt response in the genotypes. Therefore, wilt response in pigeonpea genotypes based on the leaf symptoms can serve as the reference indicator for resistance evaluation. Otherwise, leaf symptoms captured in the image has the potential for phenotyping resistance.
RGB image analysis of reference genotypes
Typical wilt symptoms on leaves were visible within a week or two after inoculation as pale yellowing and drooping was distinct in comparison to mock-inoculated leaves (Fig. 1). In the inoculated samples mean pixel intensity particularly for red, green and blue color increased proportionately as compared to the samples of mock-inoculated leaves (Fig. 2a, b, c and d). Parameters in ratio estimated out of the mean pixel intensity did not show much difference to distinguish between inoculated and mock inoculated samples. Changes in pixel intensities of color channels had reflected changes in the gray values of inoculated and mock-inoculated groups. Differences in gray values were characterized by the changes in image texture. Spatial variation in pixel intensities in regions and tone based on gray level co-occurrence indicated significant difference between the resistant and susceptible genotypes. Texture features particularly contrast and energy had significant variation (p < 0.01) between resistant and susceptible groups (Table 2 and Fig. 3).Significant difference in feature patterns between the resistant and susceptible genotypes is a valuable indicator to develop neural network model to classify genotypes in terms of wilt severity or otherwise resistance.
Characteristics pale yellowing and flaccidity of leaves in susceptible pigeonpea genotype (ICP2376 and Gulyal local); slight yellow and normal looking leaves of resistant genotypes (Asha and Maruti) noted at 15 days after soil-inoculation (30 days seedlings inoculated with F udum spore suspension 3.4 × 104 and maintained in glasshouse 28° ± 1.5 °C) in comparison to the mock-inoculation (distilled water)
Comparison of mean pixel intensity in red, green and blue color channel in the infected leaf images of pigeonpea genotypes (a = ICP2376, b = Gulyal Local, c = Asha, d = Maruti) at 15 days after inoculation (30 days old seedlings inoculated with F udum spore suspension and maintained in glasshouse at 28° ± 1.5 °C)
Training ANN, with two hidden layers having 10 and 5 nodes in layer 1 and 2 respectively was observed to map the input layer with output labels fairly and classified the genotype groups with 79–98% accuracy (Table 3). Different sets for validation particularly with 10–25% sample data gave higher levels of accuracy (92–100%) but reduced when sample data increased to 30%. Test samples used for testing the model accuracy in prediction was observed to increase as the sample set increased up to 25% level. It indicated with large number of observation (images) could improve the training accuracy or otherwise increase model fitness for the better prediction or classification of genotypes.
It was evident that RGB images on leaves are useful to explain differential vascular wilt response in pigeonpea genotypes.
Maximum photosynthetic efficiency or quantum efficiency parameter (Fv/Fm) and classification of genotype groups
At leaf level, photosynthetic efficiency parameter (Fv/Fm) was compared to distinguish between infected and healthy leaf samples. Photosynthetic efficiency parameter in inoculated and mock samples indicated Fv/Fm ≥ 0.75 correspond with healthy area (segmented in HSV plane as yellow to brown area) and below 0.75 correspond to wilt infection (Fig. 4). Parameter Fv/Fm below 0.44 was observed to match with visible wilt symptoms (blue area segmented in HSV plane). Ratio between 0.48–0.74 corresponded with leaf portion not apparently showing any symptoms but photosystem II got altered due to infection. Quantum efficiency parameter (Fv/Fm ≥ 0.75) measured in terms of fluorescent yellow–brown area was higher in healthy leaf samples (mock-inoculated) and remained almost constant till 20 dpi in all the genotypes irrespective of resistance. In inoculated samples of susceptible genotype (ICP2376), the healthy area was observed to reduce over time (0–20 dpi) as compared to mock-inoculated samples (Fig. 4). In resistant genotype (Maruti) contrastingly healthy area in inoculated samples more or less remained unchanged over time. Comparison of mean healthy area in the leaf samples of susceptible and resistant genotypes indicated significant difference (p < 0.001) at 15–20 dpi although variation was not prominent at 10 dpi (Fig. 4). Significant interaction (p < 0.01) was recorded between genotype and day of observations (dpi). Resistant genotype appeared to be less affected in terms of photosynthetic efficiency as the reduction trend in healthy area was stabilized by 15–20 dpi in comparison to susceptible genotype where downward trend continued. It was evident that wilt response measured in terms of photosynthetic efficiency parameter could be used as index for making quantitative difference between the genotypes. Otherwise, quantum efficiency signals could be used as sensible indicator for discrimination of pigeonpea genotypes.
At leaf scale photosynthetic efficiency (Fv/Fm) in (a) susceptible (ICP2376) and resistant (Maruti) genotype at 0, 5, 10, 15 and 20 dpi (inoculated with spore suspension 3.4 × 104/mL) maintained in glasshouse (28° ± 1.5 °C), and (b) comparison of relative healthy leaf area (Fv/Fm > 0.75) estimated from the chlorophyll fluorescence images of susceptible and resistant genotypes
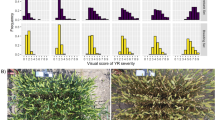
At canopy level, weighted pixel intensity estimated at 15 DPI showed significant difference (p < 0.05, independent sample t-test) between inoculated and mock samples for seven genotypes while in five genotypes did not show any difference (Fig. 5a). For inoculated samples of all the genotypes, plotting relative pixel count (%) for Fv/Fm parameter indicated difference between the genotypes as they got separated by two distinct peaks (Fig. 5b). For susceptible genotypes, peaks were observed mostly in the class interval 70–80 whereas in resistant and tolerant genotypes in 80–90 class intervals indicating tolerant and resistant genotypes had less sensitivity to the infection. Resistant genotypes observed to be less affected by wilt infection as compared to susceptible ones;somewhat similar trend was observed at leaf levelobservation. However, fitted polynomials (R2 = 0.98 with low RMSE) for the relative pixel count (%) curves were shown to be non-significant (p > 0.05, Kolmogorov–Smirnov test). It indicated at 15 dpi wilt response based on Fv/Fm parameter may not be sufficient for classification of genotypic behavior in pigeonpea. Non-significant trendin the existing data set (canopy level) assumed low predictability in pattern recognition for any statistical model desiredfor genotype classification.
Comparison of Chl-FI parameter at canopy scale a) weighted pixel intensity above Fv/Fm ≥ 0.75 (tornado chart) andb) quantum efficiency (Fv/Fm) curve in terms of relative pixel count (%) in inoculated (30 days old seedling subjected to soil-inoculation with 3.4 × 10 4 /mL) pigeonpea genotypes at 15 dpi
To predict a possible trend or pattern in the genotypes based on Fv/Fm parameter, dataset from leaf as well as canopy view images wereused to train five machine learning algorithms (https://github.com/SHUBHAJYOTIDAS/Image-based-high-throughput-phenotyping-for fusarium-wilt-resistance-in-pigeon-pea-Cajanus cajan). Training algorithms showed about 52–54% accuracy in classification of the genotypes except decision tree where performance was comparatively low (Table 4). Removing dimensionality and increasing interpretability without minimizing information in the dataset Naive Bayes algorithm had shown improved trend and clustering of data. Weights in the models were noted to configure the outputs in general but each individual prediction was characterized by low bias and high variance. A model with high variance indicated that the data set represented accurately but lead to overfitting or otherwise insufficiency in training data.
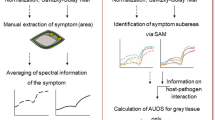
It became indicative that for better accuracy in genotype prediction or to improve performance in predictability requires large number of features from input data (images) from large number of genotypes. In addition, image capture of little advanced level of infection (later than 15 dpi) might generate wilt response pattern for genotypes classification. With possible improvement in methodology, an image-based protocol has been proposed for phenotyping pigeonepa germplasms (Fig. 6).
Discussion
Image based assessment for wilt resistance has been performed to predict pigeonpea genotypes. Vascular wilt response captured through RGB and Chl-FI techniques has appropriately detected features for genotype classification. Textural pattern in RGB images and photosynthetic efficiency parameter (Fv/Fm) from Chl-FI are the indicators to distinguish infected leaves from the healthy ones. Difference in texture features and Fv/Fmindex for healthy leaf area has been useful in classifying genotype in susceptible and resistant or tolerant types. Therefore, image based evaluation has an application potential in crop phenomics for identifying resistance in pigeonpea. However, higher accuracy in genotype classification or prediction could be achieved with further refinement of the assessment procedure.
Vascular wilt in pigeonpea is well characterized by pale yellowing and drooping exhibited in the leaves. Manifestation of yellowing and drooping is ascribed to relative increase in pixel intensities particularly blue and red channels. High pixel intensity in blue and red color channel is reported to be associated with plant stress indicating changes in stomatal conductance and chlorophyll degradation (Bock et al., 2010). In pigeonpea, characteristics increase in pixel intensities particularly in red and blue channels can be ascribed to plant stress causing changes in stomatal conductance and chlorophyll degradation. Fakrentrapp et al. (2019) has reported significant increase in intensities for blue and red colour channel in inoculated tomato plants than the mock inoculated ones. Ha et al. (2017) has used RGB images for detection of vascular wilt in radish based on texture analysis. Association of blue ratios B/BG or B/BR in vascular wilt is also reported and shown to be the best indicators of early-stage infection by Verticillium wilt in olive (Calderón et al. 2013). However, spectral indices like, Photochemical Reflectance Index (PRI), structural, chlorophyll and carotenoid indices are noted to detect only moderate to severe V. dahliae infection in olive. Al-Saddik et al (2018) has used a combination of spectral and textural data to identify and detect grapevine yellowing. Therefore, texture features from leaf images have potential in phenotyping pigeonpea genotypes as far as wilt response is concerned. Although hyperspectral images have not been considered but its possibility as an indicator needs to be explored.
Chl-FI parameter has indicated regulation of photosynthesis from leaf to crop scale can reflect wilt response in pigeonpea. Response in terms of Chl-FI parameters to biotic stress caused by pathogens has been utilized for evaluation of quite a large number of host–pathogen interactions (Mahlein et al., 2013; Perez-Buenoet al. 2019, Scholes & Rolfe, 2009; Simko et al., 2012; Rousseau et al., 2013; Chaerle et al., 2007). In the present study, Chl-FI data at leaf level has provided a relatively pure signal of wilt symptoms, which helps understanding their features to classify genotypes. However, differences in canopy level could not be ascertained universally for all the genotypes except susceptible genotypes. At canopy level signals influenced both by the plant structure and morphology could not extend wilt response for genotype classification. Large number of sample images and/or consideration of wilt response later than 15 dpi accuracy is expected to increase. Consideration of other standard Chl-FI parameters like ΦPSII, qP and NPQ in addition to Fv/Fm might increase the resolution for genotype classification. Chl-FI parameters do not always offer clear differences between healthy and infected tissues or do so at late stages of the disease (Pineda et al., 2018). Therefore, combinatorial imaging analysis with more parameters and use of ML algorithms offers operational decision-making process easier where large number of image samples may be involved in screening process (Calerdon et al. 2013).
To sum up, current finding makes some foregrounds to improve quantitative analysis for grouping of pigeonpea genotypes through imaging techniques. Genotype grouping taking features from large number of genotypes can be a possible way to improve prediction performance and thus making an automated process. Inclusion of other standard Chl-FI parameters as features may likely to generate more accurate mapping between image input and output. Therefore, combination of most sensitive indices or features at leaf and canopy levels may be a reasonable approach to make automation of phenotyping process.
RKB, PS and RA- Plant Pathology
DR, CV, SK-Phenomics
SD and SNM-Machine learning
KG- NIAPB
RSR- Genetics
Data availability
We authors agree to make data availability to publisher journal.
References
Agrios, G. N. (2005) Plant Pathology. Academic Press: PP. 522–534.
Ahlawat, I. P. S., Gangaiah, B., & Singh, I. P. (2005). Pigeonpea (Cajanuscajan) research in India—an overview. Indian Journal of Agricultural Sciences, 75, 309–320.
Al-Saddik, H., Laybros, A., & Billiot Band Cointault, F. (2018). Using image Texture and Spectral reflectance analysis to detect Yellowness and Esca in Grapevines at leaf-level. Remote Sens., 10, 618.
Baker, N. R., & Rosenqvist, E. (2004). Applications of chlorophyll fluorescence can improve crop production strategies: An examination of future possibilities. Journal of Experimental Botany, 55, 1607–1621. https://doi.org/10.1093/jxb/erh196
Baker, N. R. (2008). Chlorophyll Fluorescence: A probe of photosynthesis in vivo. – Annu. Rev. Plant Biol., 59, 89–113.
Balachandran, S., Osmond, C. B., & Daley, P. F. (1994). Diagnosis of the earliest strain-specific interactions between Tobacco mosaic virus and chloroplasts of tobacco leaves in vivo by means of chlorophyll fluorescence imaging. Plant Physiology, 104, 1059–1065. https://doi.org/10.1104/pp.104.3.1059
Bauriege, E., Brabandt, H., Gärber, U., & Herppich, W. B. (2014). Chlorophyll fluorescence imaging to facilitate breeding of Bremialactucae-resistant lettuce cultivars. Computers and Electronics in Agriculture, 105, 74–82. https://doi.org/10.1016/j.compag.2014.04.010
Berger, S., Papadopoulos, M., Schreiber, U., & Kaiser Wand Riots, T. (2004). Complex regulation of gene expression, photosynthesis and sugar levels by pathogen infection in tomato. Physiologia Plantarum, 122, 419–428. https://doi.org/10.1111/j.1399-3054.2004.00433.x
Bock, C. H., Poole, G. H., Parker, P. E., & Gottwald, T. R. (2010). Plant disease severity estimated visually, by digital Photography and image analysis, and by hyperspectral imaging. Critical Rev Plant Sci., 29, 59–107.
Calderón, R., Navas-Cortés, J. A., & Lucena, C. (2013). High-resolution airborne hyperspectral and thermal imagery for early detection of Verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sensing of Environment, 139, 231–245.
Chaerle, L., Hagenbeek, D., De-Bruyne, E., & DerStraetenD, V. (2007). Chlorophyll fluorescence imaging for disease –resistance screening of sugar beet. Plant Cell Tiss Organ Cult., 91, 97–106.
Clérivet, A., Déon, V., Alami, I., Lopez, F., Geiger, J. P., & Nicole, M. (2000). Tyloses and gels associated with cellulose accumulation in vessels are responses of plane tree seedlings (Platanusacerifolia) to the vascular fungus Ceratocystis fimbriata f. spplatani. Trees Struct. Funct., 15, 25–31.
Csefalvay, L., DiGaspero, G., Matous, K., Bellin, D., Ruperti, B., & Olejnickova, J. (2009). Pre-symptomatic detection of Plasmoparaviticolainfection in grapevine leaves using chlorophyll fluorescence imaging. European Journal of Plant Pathology, 125, 291–302. https://doi.org/10.1007/s10658-009-9482-7
Cui, D., Minzan, L., & Zhang, Q. (2009). Development of an optical sensor for crop leaf chlorophyll content detection. Computers and Electronics in Agriculture, 69, 171–176.
Diaz-Lago, J. E., Stuthman, D. D., & Leonard, K. J. (2003). Evaluation of components of partial resistance to oat crown rust using digital image analysis. Plant Disease, 87, 667–674. https://doi.org/10.1094/PDIS.2003.87.6.667
Dong, X., Ling, N., Wang, M., Shen, Q., & Guo, S. (2012). Fusaric acid is a crucial factor in the disturbance of leaf water imbalance in Fusarium-infected banana plants. Plant PhysiolBiochem, 60, 171–179.
Fakrentrapp J, Ria, F, Geilhausen M, Panassiti B (2019) Detection of Gray mold leaf infections prior to visual symptom appearance using a five-band Multispectral sensor. Front. Plant Sci., https://doi.org/10.3389/fpls.2019.00628
FAO STAT: https://www.fao.org/faostat (2018-19). Area, Production and Productivity of Pigeonpea world data
Fradin, E. F., & Thomma, B. P. H. J. (2006). Physiology and molecular aspects of Verticillium wilt diseases caused by V.dahliaeand V.albo- atrum. Molecular Plant Pathology, 7, 71–86.
Ghosh, S., & KanwarP, J. G. (2017). Alterations in rice chloroplast integrity, photosynthesis and metabolome associated with pathogenesis of Rhizoctoniasolani. Science and Reports, 7, 41610. https://doi.org/10.1038/srep41610
Granum, E., Pérez-Bueno, M. L., Calderón, C. E., Ramos, C., de Vicente, A., & Cazorla, F. M. (2015). Metabolic responses of avocado plants to stress induced by Rosellinianecatrixanalysed by fluorescence and thermal imaging. European Journal of Plant Pathology, 142, 625–632. https://doi.org/10.1007/s10658-015-0640-9
Ha, J. G., Moon, H., Kwak, J. T., Hassan, S. I., Dang, L. M., Lee, O. N., & Park, H. Y. (2017). J. Appl. Remote Sens., 11, 042621.
Haralick, R. M., Shanmugam, K., & Dinstein, I. (1973). Textural Features for Image Classification. IEEE Transactions on Systems, Man, and Cybernetics, 3, 610–621.
Hervás, A., Trapero-Casas, J. L., & Jiménez-Díaz, R. M. (1995). Induced resistance against Fusarium wilt of chickpea by nonpathogenic races of Fusarium oxysporum f. sp. ciceris and nonpathogenic isolates of F. oxysporum. Plant Disease, 79, 1110–1116.
HonoratoJunior, J., Zambolim, L., Duarte, H. S. S., Aucique-Pérez, C. E., & Rodrigues, F. A. (2015). Effects of epoxiconozale and pyraclostrobin fungicides in the infection process of Hemileiavastatrix on coffee leaves as determined by chlorophyll a fluorescence imaging. Journal of Phytopathology, 163, 968–977. https://doi.org/10.1111/jph.12399
Huang, K. Y. (2007). Application of artificial neural network for detecting Phalaenopsis seedling diseases using color and texture features. Computers and Electronics in Agriculture, 57, 3–11.
Indiastat.com: https://www.indiastat.com (2018–19.) Area, Production and Productivity, of Pigeonpea in India
Ivanov, D. A., & Bernards, M. A. (2016). Chlorophyll fluorescence imaging as a tool to monitor the progress of a root pathogen in a perennial plant. Planta, 243, 263–279. https://doi.org/10.1007/s00425-015-2427-9
Jain, K. C., & Reddy, M. V. (1995). Inheritance of resistance to Fusarium wilt in pigeonpea (Cajanuscajan (L.) Millsp.). Indian J Genet, 55, 434–437.
Kai S, Zhikun L, Hang S, Chunhong G (2011) A research of maize disease image recognition of corn based on BP networks, in IEEE Third International Conference on Measuring Technology and Mechatronics Automation, pp.246–249.
Kaundal, R., Kapoor, A. S., & Raghava, G. P. S. (2006). Machine learning techniques in disease forecasting: A case study on rice blast prediction. BMC Bioinformatics, 7, 485.
Kuckenberg, J., Tartachnyk, I., & Noga, G. (2009). Temporal and spatial changes of chlorophyll fluorescence as a basis for early and precise detection of leaf rust and powdery mildew infections in wheat leaves. Precision Agriculture, 10, 34–44. https://doi.org/10.1007/s11119-008-9082
Leng, Q., Qi, H., Miao, J., Zhu, W., & Su, G. (2015). One-class classification with extreme learning machine. Mathematical Problems Engineering, pp. 11.
Lorenzen, B. and Jensen, A. (1988). Reflectance of blue, green, red and near infrared radiation from wetland vegetation used in a model discriminating live and dead above ground biomass. https://doi.org/10.1111/j.1469-8137.1988.tb04173.x
Mahlein, A. K., Alisaac, E., Masri, A. A., Behmann, J., Dehne, H. W., & Erich-Christian, O. E. C. (2013). Comparison and combination of Thermal, Fluorescence, and Hyperspectral imaging for monitoring Fusarium head blight of Wheat on spikelet scale. Sensors, 19, 2281. https://doi.org/10.3390/s19102281
Maxwell, K., & Johnson, G. N. (2000). Chlorophyll fluorescence – a practical guide. Journal of Experimental Botany, 51, 659–668.
Mokhtar, U., Ali, M. A., Hassanien, A. E., & Hefny, H. (2015). Identifying two of tomatoes leaf viruses using support vector machine. Information Systems Design and Intelligent Applications, pp. 771–782.
Nene, Y. L., & Kannaiyan, J. (1982). Screening of pigeonpea for resistance to Fusarium wilt. Plant Disease, 66, 306–307.
Nene Y L(1980)Proceedings Consultants Group. Discussion on Resistance to soil borne diseases in Legumes, ICRISAT, India.167 pp.
Nichols, J. A., Hsien, W., Chan, H., & Baker, M. A. B. (2019). Machine learning: Applications of artificial intelligence to imaging and diagnosis. Biophysical Reviews, 11, 111–118.
Okiror, M. A. (2002). Genetics of wilt resistance in pigeonpea. Indian J Genet, 62, 218–220.
OxboroughK, B. N. R. (1997). Resolving chlorophyllafluorescence images of photosynthetic efficiency into photo-chemical and non-photochemical components-calculation of qPandFv’/Fm’ without measuring F0’. Photosynthesis Research, 54, 135–142.
Parlevliet, J. E. (1979). Components of resistance that reduce the rate of disease epidemic development. Annual Review of Phytopathology, 17, 203–232.
Parupalli, S., Saxena, R. K., Sameerkumar, C. V., Sharma, M., Singh, V. K., Vechalapu, S., Kavikishor, P. B., Saxena, K. B., & VarshneyRK,. (2017). Genetics of fusarium wilt resistance in pigeonpea as revealed by phenotyping of RILs S. Journal of Food Legumes, 30, 241–244.
Pérez-Bueno, M. L., Pineda, M., & Barón, M. (2019). Phenotyping Plant Responses to Biotic Stress by Chlorophyll Fluorescence Imaging. Frontiers in Plant Science, 10, 1135. https://doi.org/10.3389/fpls.2019.01135
Pineda, M., Pérez-Bueno, M. L., & Barón, M. (2018). Detection of bacterial infection in melon plants by classification methods based on imaging data. Frontiers in Plant Science, 9, 164. https://doi.org/10.3389/fpls.2018.00164
Poland, J. A., Balint-Kurti, P. J., Wisser, R. J., Pratt, R. C., & Nelson, R. J. (2009). Shades of gray: The world of quantitative disease resistance. Trends in Plant Science, 14, 21–29.
Pydipati R, Burks TF, Lee WS (2006) Identification of citrus disease using color texture features and discriminant analysis. Comput. Electron. Agric. 52 (1)
Rafael, G. (2018). Digital image processing. New York, NY: Pearson.
Rahman, M. A., Abdullah, H., & Vanhaecke, M. (1999). Histopathology of susceptible and resistant Capsicum annuumcultivars infected with Ralstonia solanacearum. Journal of Phytopathology, 147, 129–140.
Rios, J. A., Aucique-Pérez, C. E., Debona, D., Cruz Neto, L. B. M., Rios, V. S., & Rodrigues, F. A. (2017). Changes in leaf gas exchange, chlorophyll a fluorescence and antioxidant metabolism within wheat leaves infected by Bipolarissorokiniana. The Annals of Applied Biology, 170, 189–203. https://doi.org/10.1111/aab.1232
Rolfe, S. A & Scholes, J. D. (2010). Chlorophyll fluorescence imaging of plant-pathogen interactions. Protoplasma, 247, 163–175. https://doi.org/10.1007/s00709-010-0203-z
Rousseau, C., Belin, E., Bove, E., Rousseau, D., Fabre, F., & Berruyer, R. (2013). High throughput quantitative phenotyping of plant resistance using chlorophyll fluorescence image analysis. Plant Methods, 9, 17. https://doi.org/10.1186/1746-4811-9-17
Roy, P. S. (1989). Spectral reflectance characteristics of vegetation and their use in estimating productive potential. Proc. Indian Acad. Sci. (plant Sci.), 99, 59–81.
Santhanam, P., VanesseHP, AlbertI., Faino, L., & NurnbergerT and Thomma BP,. (2013). Evidence for functional diversification within a fungal NEP1-like protein family. Molecular Plant-Microbe Interactions, 26, 278–286.
Savary, S., Nelson, A., Willocquet, L., & PanggaI, A. J. (2012). Modelling and mapping potential epidemics of rice diseases globally. Crop Protection, 34, 6–17.
Saxena, K. B., Kumar, R. V., Saxena, R. K., Sharma, M., Srivastava, R. K., Sultana, R., Varshney, R. K., Vales, M. I., & Pande, S. (2012). Identification of dominant and recessive genes for resistance to Fusarium wilt in pigeonpea and their implication in breeding hybrids. Euphytica, 188, 221–227.
Scholes, J. D., & Rolfe, S. A. (2009). Chlorophyll fluorescence imaging as tool for understanding the impact of fungal diseases on plant performances: A Phenomics prospective. Functional Plant Biology, 36, 880–892.
Sebela, D., Quinones, C., Cruz, C. V., Ona, I., Olejnickova, J., & Jagadish, K. S. V. (2018). Chlorophyll fluorescence and reflectance-based non-invasive quantification of blast, bacterial blight and drought stresses in rice. Plant and Cell Physiology, 59, 30–43. https://doi.org/10.1093/pcp/pcx
Sekulska-Nalewajko, J., Kornas, A., Gocławski, J., Miszalski, Z., & Kuźniak, E. (2019). Spatial referencing of chlorophyll fluorescence images for quantitative assessment of infection propagation in leaves demonstrated on the ice plant: Botrytis cinereapathosystem. Plant Methods, 15, 18. https://doi.org/10.1186/s13007-019-0401-4
Simko, I., Jimenez-Berni, J. A., & Furbank, R. T. (2012). Detection of decay in fresh-cut lettuce using hyperspectral imaging and chlorophyll fluorescence imaging. Postharvest Biology and Technology, 106, 44–52. https://doi.org/10.1016/j.postharvbio.2015.04.007
Singh, D., Sinha, R. V. P., Singh, M. N., Singh, D. K., Kumar, R., & Singh, A. K. (2016). Genetics of Fusarium Wilt Resistance in Pigeonpea (Cajanuscajan) and Efficacy of Associated SSR Markers. Plant Pathology Journal, 32, 95–101.
Sperschneider, J., Gardiner, D. M., Dodds, P. N., Tini, F., & CovarelliL, S. K. B. (2016). EffectorP: Predicting fungal effector proteins from secretomes using machine learning. New Phytologist, 210, 743–761. https://doi.org/10.1111/nph.13794
Upadhyay, R. S., & Rai, B. (1992). Wilt disease of pigeonpea. In: Singh, U., Mukhopadyaya, U., Kumar, A., & Chaube, H. S. (Eds.). Plant Disease of International Importance. Prentice Hall, Engelwood cliffs New Jersey, pp. 388–404.
West, J. S., Bravo, C., Oberti, R., & Lemaire, D. (2003). The potential of optical canopy measurement for targeted control of field crop diseases. Annual Review of Phytopathology, 41, 593–614.
Zhou, B. J., Jia, P. S., Gao, F., & Guo, H. S. (2012). Molecular characterization and function analysis of a necrosis and ethylene-inducing, protein-encoding gene family from Verticillium dahliae. Molecular Plant-Microbe Interactions, 25, 964–975.
Acknowledgements
The authors gratefully acknowledge Head and Professor, Division of Plant Pathology, Joint Director and Director, ICAR-Indian Agricultural Research Institute New Delhi for overall support.
Funding
This work was funded by the ICAR-Indian Agricultural Research Institute New Delhi under the Post-Graduate Research activities and National Agricultural Science Fund (NASF) funded NanajiDeshmukh Plant Phenomics Center, ICAR-IARI, New Delhi.
RKB acknowledges the RGNFD fellowship received from UGC, Govt of India, New Delhi; For the pursuance of PhD degree at Indian Agricultural Research Institute, New-Delhi-110012.
Author information
Authors and Affiliations
Contributions
Conceptualization: PS and RKB; investigation: RKB, DR, SK and PS; writing—original draft preparation: PS, RKB, RD, SD and SNM; reviewing and editing: RKB, PS, DR, KG and RSR; machine learning- SD and SNM; supervision: RA, CV and PS.
Corresponding author
Ethics declarations
Disclosure of potential conflicts of interest
The authors declare that they have no conflict of interest.
Research involving Human Participants / Animals
Research work has been carried out in different pigeonpeagenotypesfor resistance phenotyping. There is no involvement of Humans participation or Animals present experimentaltrial purpose.
Informed consent
All authors have informed and had consent about reviewing and publishing in European Plant Pathology Journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12600_2022_993_MOESM1_ESM.docx
Supplementary file1 (DOCX 26 KB) Supplementary Fig. 1 Wilt severity index 20 (WSI20) of reference pigeonpea genotypes (susceptible – ICP2376 and Gulyal local; resistant- Asha and Maruti) in 30 days old seedlings (soil inoculation with F udum spore suspension 3.4 x104 /mL) maintained in glasshouse (28°±1.5°C).
Rights and permissions
About this article
Cite this article
Bannihatti, R.K., Sinha, P., Raju, D. et al. Image Based High throughput Phenotyping for Fusarium Wilt Resistance in Pigeon Pea (Cajanus cajan). Phytoparasitica 50, 1075–1090 (2022). https://doi.org/10.1007/s12600-022-00993-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-022-00993-5