Abstract
Sclerotinia stem rot (SSR), caused by Sclerotinia sclerotiorum, is a globally important, yield limiting disease of soybean. Progress has been made in our understanding of this pathosystem at the plant level, such as the key role of oxalic acid in disease development and the importance of cell wall-degrading enzymes and other secreted proteins. Unfortunately, advances have largely focused on the fungal side of this interaction and only provide glimpses into the plant mechanisms governing resistance to this pathogen. With the absence of commercially available resistant soybeans, chemical and cultural solutions are being used by farmers to manage SSR with limited success. Additional research is needed to identify S. sclerotiorum resistance mechanisms that can be exploited to improve genetic resistance in soybean and decrease reliance on spray regimes. Technologies such as transgenics and RNAi could be exploited to improve the level of resistance to S. sclerotiorum in soybean. This review offers insight into the hurdles of managing SSR at the plant level and potential solutions that might be adopted in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Sclerotinia sclerotiorum lifestyle characterization
Sclerotinia sclerotiorum (lib.) deBary has traditionally been considered a prototypical necrotrophic pathogen. Necrotrophic pathogens are characterized by nutrient acquisition from cells killed by a host of cell wall-degrading enzymes and toxins produced by the pathogen. Necrotrophs are further characterized by their wide host range and lack of complete resistance in their plant hosts. Partial, quantitative resistance of soybean to Sclerotinia stem rot (SSR) has been observed in field trials of commercial varieties (Conley et al. 2017) and through QTL mapping (Bastien et al. 2014; Vuong et al. 2008; Zhao et al. 2015). However, reliance on this type of quantitative resistance can result in limited control. This is in contrast to biotrophic pathogens which depend on living plant tissue for nutrient acquisition and propagation. Biotrophic fungi release effectors, which are small, secreted proteins that contribute to virulence by manipulating host physiology and suppressing plant basal defenses (Rafiqi et al. 2012). Resistance to biotrophic pathogens is associated with a strong hypersensitive (HR) response which can result in programmed cell death through the process of effector-triggered immunity (Raffaele and Kamoun 2012) after effectors (“avirulence proteins”; Bourras et al. 2016) are recognized (directly or indirectly) by immune receptors (“resistance proteins”) (Williams et al. 2016). It has been assumed that the presence of cell wall-degrading enzymes may render effectors less crucial in necrotrophic pathogens, and thus secreted proteins have been an understudied area of S. sclerotiorum research.
Recent studies indicate that establishment of SSR is more intricate than brute degradation of host tissues. Due to the complicated nature of S. sclerotiorum and host interactions, Kabbage et al. (2015) emphasized the importance of a more holistic view of this pathosystem that considers a hemi-biotrophic lifestyle of S. sclerotiorum. Contrary to the inelastic classification of S. sclerotiorum as strictly nectrotrophic, Kabbage et al. (2015) describe two phases of infection based on microscopic imaging. In the early stages, the fungus grows intracellularly without killing host cells, during which the pathogen is presumed to secrete pathogenicity factors, including oxalic acid (OA), that modulate the host response. Following the brief establishment stage, necrotrophy is initiated, and the fungus procures nutrients from dead host tissue. Transient biotrophic growth persists at the leading edge of lesions while cell death is initiated at the trailing end. This model argues that a more relevant measure of host-pathogen interactions is the damage caused to the host versus the pathogen lifestyle and the process leading to damage. This relationship is demonstrated in “damage-response” curves where the outcome of the interaction is the primary consideration.
Important S. sclerotiorum virulence factors
Oxalic acid functions to suppress host defenses and induce cell death
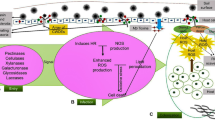
In support of the idea of an early biotrophic stage, virulence factors related to the suppression of host defenses have been characterized and are known to be important to S. sclerotiorum infection (Fig. 1). The most studied of these virulence factors, in S. sclerotiorum, is oxalic acid (OA) and the conjugate base, oxalate. Oxalate is a di-carboxylic acid and alters an array of plant physiological processes that facilitate fungal colonization in the host plant. Oxalic acid is not directly toxic but may have a more refined role as a signaling molecule that hijacks the host cell and provides a more conducive environment for fungal growth (Kim et al. 2008; Williams et al. 2011; Kabbage et al. 2013).
S. sclerotiorum virulence factors serve various regulatory functions. Oxalic acid (OA: orange lines), broadly, dampens host defense responses and suppresses autophagy at the leading edge of infection while enhancing reactive oxygen species (ROS), apoptosis, and the action of cell wall-degrading enzymes (CWDE) at the trailing edge of the lesion. Effector-like secreted proteins (purple lines) similarly function to evade host defense responses and induce cell death. Cell wall-degrading-enzymes (blue lines) are essential for degradation of infection surfaces, and endopolygalacturonases (PGs) are involved in the induction of programmed cell death via cytosolic Ca2+ signaling
Oxalate deficient S. sclerotiorum mutants were shown to be poorly pathogenic (Cessna et al. 2000; Dickman and Mitra 1992; Godoy et al. 1990; Kim et al. 2008; Liang et al. 2015a; Williams et al. 2011), and these mutations provoked morphological changes in the fungus in several studies (Rollins 2003; Rollins and Dickman 2001; Liang et al. 2015a). Studies with the OA mutants showed that in the early stages of infection, OA dampens the oxidative burst of the host plant by creating a reductive state, thus suppressing host defenses (Williams et al. 2011). These oxidative bursts are caused by the release of reactive oxygen species (ROS). Reactive oxygen species occur endogenously in cells as byproducts of the reduction of molecular oxygen (O2), but plants produce higher ROS levels under stressful conditions as a protective strategy (Kotchoni and Gachomo 2006; Mittler et al. 2011; Petrov and Van Breusegem 2012; Noctor et al. 2014; Sewelam et al. 2016; Xia et al. 2015; Wojtaszek 1997). Indeed, oxalate-deficient mutants of S. sclerotiorum appear to be actively recognized by the host plant, leading to classical defense responses that include an ROS burst and callose deposition (Williams et al. 2011). Processes involving recognition of pathogens by the host and evasion of such defenses by the pathogen have been largely described in response to biotrophs. Thus, the initial pathogenic phase of this well-established necrotroph, surprisingly, displays features akin to those observed during biotrophic interactions.
In contrast to this early phase, S. sclerotiorum was shown to elicit programmed cell death during disease development and concordantly increase ROS levels in the plant at the later stages of the infection process (Kim et al. 2008; Williams et al. 2011). ROS induction is associated with plant defense mechanisms involving HR to biotrophic pathogens. However, this response was shown to be exploited by pathogens with a predominantly necrotrophic lifestyles (Liu et al. 2012; Curtis and Wolpert 2002) including S. sclerotiorum (Kim et al. 2008). At later stages, when necrotic tissues are necessary for nutrient acquisition by S. sclerotiorum, OA flips the switch on ROS regulation and enhances its production to cause apoptosis-like programmed cell death in the host, as indicated by DNA laddering and TUNEL reactive cells (Kim et al. 2008). Recent work showed that this increase in ROS levels by S. sclerotiorum is achieved by hijacking host NADPH oxidases in soybean (Ranjan et al. 2018).
Additional proposed roles for OA have been reported including the acidification of the surrounding environment (Xu et al. 2015). This acidification enhances the activity of cell wall degrading enzymes and enzymes involved in signaling, such as the MAP-kinases associated with sclerotial development (Bateman and Beer 1965; Chen et al. 2004; Rollins and Dickman 2001). OA was also proposed to function as chelating agent, particularly to chelate calcium as evidenced by the presence of calcium oxalate crystals in the infection court (Uloth et al. 2015). Calcium is known to be an important component of cellular signaling and structure. In toto, this simple dicarboxylic acid is a key molecule with multifaceted roles in the pathogenic success of S. sclerotiorum. Thus, OA constitutes a prime target to achieve resistance to this pathogen, hence the importance of studying the genes and pathways involved in its biogenesis.
Genes related to OA production are known to be important for pathogenicity
Functional genetic studies have identified various genes related to OA synthesis and accumulation in S. sclerotiorum. For example, Ss-Oah1, an oxaloacetate acetylhydrolase (OAH)- encoding gene has been demonstrated to be essential for the accumulation of OA; S. sclerotiorum deletion Ss-oah1 mutants are unable to accumulate OA in culture or during plant infection (Liang et al. 2015a; Xu et al. 2015). Ss-Oah1 catalyzes the conversion of oxaloacetate to OA during the final step in OA synthesis, (Dutton and Evans 1996; Munir et al. 2001) and accumulation of Ss-Oah1 transcripts is dependent on the transcription factor, Ss-Pac1. Ss-Pac1 was previously observed to be necessary for neutral pH-induced OA accumulation (Rollins 2003), and four Pac C binding sites were identified in the Ss-oah1 promotor region (Liang et al. 2015a). Gene deletion mutants also lacked compound appressoria and were deficient in sclerotia development, however, subsequent T-DNA and targeted gene replacement Ss-oah1 mutants exhibited normal growth and sclerotial development in a pH lower than 5.5 compared to wildtype S. sclerotiorum (Xu et al. 2015), indicating the pH-dependency of Ss-Oah1. Upon challenging plant hosts, mutants did cause small lesions on tomato, Arabidopsis, and soybean leaves, but the hyphae gradually lost viability (Liang et al. 2015a). The development of even small lesions suggests that other pathogenicity factors, besides OA, contribute to initial infection and primary lesion development. Oxalate decarboxylase genes Ss-Odc1 and Ss-Odc2 have also been characterized (Liang et al. 2015b), and mutants of Ss-Odc2 were, similarly, hindered in their ability to form appressoria and lesions affecting virulence on soybean leaves. Wounding restored lesion development, indicating that the contribution of Ss-Odc2 to virulence was, likewise, incomplete.
Secreted proteins function to dampen the host immune response
Despite the importance of OA as a major virulence factor in S. sclerotiorum, recent studies have singled out other virulence components. While necrotrophic pathogens are thought to have fewer effectors due to an ephemeral interaction with their host plants compared to biotrophic pathogens, exploratory research has identified several secretory proteins with effector-like activity. In agreement with the early biotrophic trope of S. sclerotiorum’s lifestyle, the potential effectors identified may assist S. sclerotiorum in evading host detection and condition the plant for the impending necrotrophic stage.
As observed in other pathosystems, S. sclerotiorum secretory proteins may dampen the host immune response by the suppression of jasmonic/ethylene signaling pathway. Ss-Itl is one such effector candidate (Zhu et al. 2013). The RNAi silencing of Ss-Itl in S. sclerotiorum led to a strong defense response in Arabidopsis as evidenced by expression of marker genes involved in the JA/ET and SA signaling pathway, and the Ss-Itl-silenced version of S. sclerotiorum was unable to successfully invade Arabidopsis. The protein is also critical for normative physiology of S. sclerotiorum, with silenced strains having aberrant, detrimental hyphal and sclerotial structures. Another S. sclerotiorum secreted protein, Ss-Rhs1, was also found to be critical for virulence. The silencing of Ss-Rhs1 led to aberrant sclerotia and hindered the formation of compound appressoria on A. thaliana. Virulence of the silenced strain was reduced on A. thaliana and Brassica napus despite the continued production of OA, amylases, cellulases, proteases, and pectinases (Yu et al. 2017). Yet, another effector-like protein, Ss-Caf1, was identified by Xiao et al. (2014), and was shown to be important for virulence. The identified gene, Ss-Caf1, encodes a secretory protein candidate with a Ca2+ binding EF hand motif likely involved in compound appressorium formation and sclerotial development. Lesions on rapeseed failed to form when challenged with the mutant, despite greater quantities of OA being produced, likely due to impeded tissue penetration as hyphae lacked modification into the melanin-rich infection cushions known as compound appressoria. This protein also induces cell death within the host as observed following transient expression in Nicotiana benthamiana (Xiao et al. 2014). Similarly, the protein Ss-Ssvp1 was shown to induce plant cell death (Lyu et al. 2016). SsSsvp1 appears to induce cell death through the disruption of the mitochondrial respiration chain. Indeed, Bimolecular fluorescence complementation (BiFC) assays demonstrated an interaction between Ss-Ssvp1 and QCR8 (a subunit of the cytochrome b-c1 complex) that disrupted localization of QCR8 in the mitochondria. Ss-Cp1, a certato-platanin protein, was recently identified as yet another inducer of cell death when expressed in N. benthamiana. This secreted protein was found to interact with At-PR1 in the apoplast via yeast two-hybridization, Glutathione S-Transferase, Co-immunoprecipitation, and BiFC assays (Yang et al. 2018). Due to their role in virulence and modification of host plants, targeting secreted, effector-like proteins from S. sclerotiorum or elucidating and altering interacting host proteins may be an effective SSR control strategy.
Though functional studies describing secreted proteins from S. sclerotiorum are limited, genomics studies identified a large repertoire of potential secreted proteins that can serve as effectors (Amselem et al. 2011). Using bioinformatics approaches, Guyon et al. (2014), identified 78 effector candidates based on protein domains and motifs found in known fungal effectors, signatures of positive diversifying selection, and recent gene duplication events. Derbyshire et al. (2017) identified 70 effector candidates, only nine of which overlapped with Guyon et al., using a more complete genome derived from the optical map of the S. sclerotiorum genome combined with PacBio and Illumina sequencing. Twenty-two sequences contained predicted functional domains, and four have previous associations with effector-like activity in other fungi. Furthermore, RNAseq data by Derbyshire et al. (2018) identified 374 abundant sRNAs targeting immunity components in 10 host plants, demonstrating that S. sclerotiorum has another arsenal of virulence factors, besides effectors, that can be deployed to avoid detection or immune responses. While these results are intriguing, the importance and deployment of these effectors and sRNAs for invasion and host defense suppression by S. sclerotiorum will have to be elucidated through functional studies.
The brute force of cell-wall-degrading enzymes is vital for necrotrophic infection stages
Cell-wall-degrading-enzymes (CWDEs) were a principle focus of early studies in S. sclerotiorum pathogenicity. As a necrotrophic pathogen that requires plant cell death for growth and infection, S. sclerotiorum possesses an arsenal of enzymes to transverse host barriers, including cell walls. The S. sclerotiorum genome contains 106 carbohydrate active enzymes and affiliated proteins (CAZymes) which assist in the degradation of cell wall substrates including cellulose (20), hemicellulose (40), hemicellulose and pectin side chains (13), and pectin (33) (Amselem et al. 2011). Depolymerization of cellulose, hemicellulose, and pectin and subsequent degradation of plant structural components facilitates pathogenesis and access to mono and oligo-saccharides. Levels of pectinolytic and cellulolytic enzymes are correlated with disease severity and pathogenicity (Lumsden 1969; Prade et al. 1999; Willats et al. 2001). Using polysaccarhide depolymerases and glucosidases, the fungus converts cellulosic, pectinolytic, and hemicellulosic substrates into simple sugars for assimilation (Riou et al. 1991). Glycoside hydrolase activities accompany polysaccharidase enzymes to release monosaccharides from polymers of the cell wall.
Several hydrolases are produced which differ in the isoelectric point and molecular weight, thus increasing the efficiency of hydrolytic applications. Furthermore, these enzymes sometimes function in a nonspecific, multifunctional manner, targeting multiple proteins, and induction can often occur via multiple substrates. Cellulolytic and pectinolytic enzymes, for example, are induced by both cellulolytic or pectinolytic substrates, perhaps indicating a common regulatory system. In S. sclerotiorum, 69 enzymes involved in cellulose depolymerization and 24 hemicellulolytic enzymes have been identified through comparative genomics (Kubicek et al. 2014; Zhao et al. 2013). The deployment of such a vast number of enzymes facilitates infection in a wide range of polysaccharide cell wall compositions, thus facilitating its promiscuous ability to infect numerous hosts.
In turn, plants do combat enzymes to an extent, as indicated by the production of polygalacturonase-inhibiting proteins (PGIPs) which target the activity of the pectin-degrading enzymes, endopolygalacturonases (PGs) (De Lorenzo and Ferrari 2002). Endopolygalacturonase production by S. sclerotiorum induces cytosolic Ca2+ signaling and cell death which can be suppressed with administration of PGIP (Zuppini et al. 2005). These PGs are important pathogenicity determinants and S. sclerotiorum isolate screening has revealed a positive correlation between pectinase activity, using pectin agar medium, and the virulence of 25 isolates (Asoufi et al. 2007). Sclerotinia has been found to upregulate 11 genes related to pectin degradation during infection of sunflower. However, genes involved in cellulose and hemicellulose degradation are scarcer than those encoded by nearly all plant cell wall degrading pathogens (Amselem et al. 2011). This observation may be related to S. sclerotiorum’s propensity to infect via flowers, which are abundant in pectin, and, indeed Huzar-Novakowiski and Dorrance (2018) observed differential host-plant resistances when soybean lines were inoculated at the flower with ascospores versus at the petiole with mycelia. The activity of PGs and OA is synergistic, with OA acidifying the cellular environment (Cessna et al. 2000; Dutton and Evans 1996; Zhou and Boland 1999), thus facilitating PG activity and weakening pectic polymers (Kurian and Stelzig 1979). Cell wall-degrading enzymes have an important functional role in the necrotrophic nutrient acquisition ability of SSR and host counter-response to these CWDEs is a prospective area of plant defense research. Additionally, the comparatively high activity of pectinic versus cellulosic and hemicellulosic enzyme activity explains why S. sclerotiorum is an effective pathogen of dicots with relatively higher quantities of pectinic polysaccarhides compared to monocots which have primary cell walls characterized by a richness of hemicellulosic polysaccharides.
The genetics of quantitative resistance to SSR
Various QTL conferring resistance to SSR in soybean have been identified
In the absence of elicitors of strong resistance to the pathogen, polygenic alleles with minor effects are widely believed to contribute to resistance to S. sclerotiorum. Partially resistant soybean genotypes have been selected and identified (Bastien et al. 2014; Boland and Hall 1987; Grau et al. 1982; Han et al. 2008; Huynh et al. 2010; Iquira et al. 2015; Kim and Diers 2000; Li et al. 2010; McCaghey et al. 2017; Sebastian et al. 2010; Zhao et al. 2015). Overall, 103 quantitative trait loci (QTL) that contributed to resistance have been recorded in Soybase on 18 out of 20 chromosomes (Soybase.org 2010), however the contribution to klenducity, or disease escape mechanisms, versus physiological resistance has often not yet been elucidated. Three QTL were identified by Kim and Diers (2000) and 28 QTL were identified by Arahana et al. (2001) which individually explain 4–10% of the phenotypic variation between 153 and five, respectively, recombinant inbred lines compared to SSR-susceptible parent, ‘Williams 82.’ Additionally, Vuong et al. (2008) mapped four QTL for Sclerotinia stem rot resistance that each explained 5.5 to 12.1% of the phenotypic variance in Sclerotinia stem rot development, and Guo et al. (2008) identified seven QTLs which explained 6.0–15.7% of resistance phenotype differences in their populations. Identification of these loci provide an opportunity to use marker assisted selection (MAS) as a potential tool for the screening of lines resistant to SSR. However, such an approach presents practical challenges that must be overcome to deploy SSR resistance.
Polygenic resistance to SSR presents breeding challenges
While polygenic resistance (quantitative resistance) is thought be more durable than qualitative resistance; breeding using quantitative resistance is complicated. This includes the “drag” of deleterious and undesirable traits within and near QTL regions, existence of numerous QTL with minimal sole contribution to SSR resistance, and epistatic interactions that pose a challenge to heritability (Moellers et al. 2017). Furthermore, the genetics of physiological resistance to S. sclerotiorum are not well understood. Current ‘field tolerant’ soybean cultivars may be tolerant due to avoidance phenotypes such as flowering time and plant height or entangled environmental and genetic interactions. For example, Kim and Diers (2000) used Novartis S19–90 as a source of resistance in breeding lines and mapped three QTL that accounted for 8–10% of disease severity index (DSI) variability. However, two were associated with disease escape mechanisms of greater height, increased lodging, and later flowering date. These escape mechanisms make screening for physiological disease resistance in a field setting difficult. Furthermore, flowering time or canopy closure may differentially align with apothecial development in varied environments, thus impacting disease resistance across environments. In disease nurseries, screening is complicated by aggregated distributions of inoculum and differing favorable microenvironments for infection within a field which may result in differential disease pressure. The same study observed differential disease resistance and heritability between research environments with broad-sense heritability of DSI ranging from 0.30–0.71 across research sites within Michigan (Kim and Diers 2000). These heritabilities are lower than those found by Miklas and Grafton (1992), ranging from 0.50–0.77. Therefore, while resistance to SSR can be selected for in a field setting, there remains a persistent need to identify consistent, heritable resistance across environments.
Methods have been developed to screen for physiological resistance to SSR
To circumvent resistance conferred by escape mechanisms, breeders have mapped QTL and screened lines using inoculation methods that avoid selecting exclusively klendusity-associated traits (Arahana et al. 2001; Guo et al. 2008; Vuong et al. 2008). However, only few, partially resistant lines were identified. These inoculation methods include severing the stem of soybean and placing a mycelial plug on the cut stem (Vuong et al. 2004, 2008), inoculation of a cut petiole (Del Rio et al. 2001), or ascospore inoculation of flowers (Cline and Jacobsen 1983; Huzar-Novakowiski and Dorrance 2018; Pérès 1995; Rousseau et al. 2004). Floral bud inoculations with cotton pads dipped in mycelial suspensions have also been used with reproducible inoculations (Iquira et al. 2015). The results of these inoculation techniques for resistance differentiation may vary depending on the inoculation technique (Rousseau et al. 2004; Huzar-Novakowski and Dorrance 2018) and the isolate used (Willbur et al. 2017). Huzar-Novakowiski and Dorrance (2018) suggest that multiple screening methods are needed to successfully capture varied resistance mechanisms in selections. Similarly, Willbur et al. (2017) evaluated a panel of S. sclerotiorum isolates from the United States and observed differential aggressiveness on soybean cultivars, suggesting that multiple isolates are needed to screen for broad resistance. McCaghey et al. (2017), therefore, utilized both SSR field nurseries and greenhouse evaluations with several S. sclerotiorum isolates to identify recombinant inbred lines (RIL) with consistent, moderate to high levels of resistance. Inoculation methods are useful for capturing physiological resistance, but screening methods should be comprehensive to capture the quantitative nature of SSR resistance and differences in S. sclerotiorum pathogenicity.
Methods of evaluating lines in the field may include the infestation of fields with sclerotia collected from alternate hosts (Kim et al. 1999) or from naturally infested fields rotated with susceptible hosts such as sunflower (McCaghey et al. 2017). Infestation of soil and planting alternate hosts homogenizes inoculum and encourages more uniform infections in field trials. Lines may be evaluated using a disease severity index or DSI (Grau et al. 1982; Kim and Diers 2000) which differentiates between lateral branch and mainstem infections.
Modern soybean breeding has emphasized lateral branching, and mainstem infections are associated with the disruption of the mainstem vascular system, which results in severe yield loss. A system which uses both artificial inoculations and field evaluations (McCaghey et al. 2017) allows one to select for both physiological resistance and field tolerance in addition to favorable agronomic properties.
Biotechnology to control Sclerotinia stem rot
Genetic engineering can reduce SSR but is not commercially deployed
While traditional breeding for SSR resistance has identified lines ranging from field tolerance to high levels of quantitative resistance, traditional breeding is time intensive and has its challenges when dealing with alleles with additive effects as described above. Thus, the identification of specific genes and pathways that can be targeted via genetic engineering will be helpful to introgress SSR resistance into commercial soybean germplasm. Efforts to use transgenic approaches targeting SSR have not been widely explored. Enhanced resistance in soybean targeting OA for degradation has been observed (Cunha et al. 2010; Donaldson et al. 2001). Donaldson et al. (2001) first developed a transgenic soybean line which expressed an oxalate oxidase (OxO). This physically resilient wheat germin is conserved in monocots and is proposed to oxidize oxalic acid to carbon dioxide and hydrogen peroxide (Dumas et al. 1993; Lane et al. 1993). Oxalate oxidase serves multiple purposes (Donaldson et al. 2001). First, it can hinder the action of oxalate on cell walls. Second, it can remove oxalate, which suppresses host oxidative bursts (Cessna et al. 2000), from the cellular environment. Third, the production of hydrogen peroxide is known to both trigger programmed cell death involved in the HR response, and it mediates the cross-linking of cell wall glycoproteins (Levine et al. 1994). The transformation event occurred without obvious yield penalties and resistance in a field setting was equivalent to the most resistance inbred line tested by Cober et al. (2003). Cuhna et al. (2010) also targeted oxalate but with a transformation event using the decarboxylase gene (OxDC) which resulted in delayed lesion development. Oxalate decarboxylase converts oxalate into carbon dioxide and formate with no hydrogen peroxide production (Kesarwani et al. 2000), and transgenic lines had reduced lesion size by up to 96%. The supposed avoidance of ROS production may prevent programmed cell death related to fungal growth and colonization (Kim et al. 2008; Ranjan et al. 2018), though no comparative studies examine differences in resistance or the induction of programmed cell death in these lines. While lesion development is successfully impaired, these transgenic lines have yet to be exploited commercially.
The use of RNAseq to assess global gene expression during S. sclerotiorum infection of soybean is a promising tool to develop additional genetic engineering technologies for enhanced resistance. Expression data from soybean and other crop plants will provide valuable information on soybean susceptibility factors for gene editing or post transcriptional gene silencing and gene candidates for cis or transgenic plant development. RNAseq methods were used to evaluate the interaction of S. sclerotiorum with crop hosts such as, canola (Joshi et al. 2016; Girard et al. 2017; Seifbarghi et al. 2017), pea (Chang et al. 2018), and common bean (Oliveira et al. 2015).
RNA-interference strategies show promising results
More recently, RNA interference (RNAi) strategies have been experimentally developed as a defense strategy against S. sclerotiorum and other fungal pathogens. Through various studies in fungal pathogens, it is known that large and small double-stranded RNA (dsRNA) molecules can be transported within and between species for effective gene silencing (Wang et al. 2016). The phenomenon, which occurs endogenously in eukaryotic organisms for gene regulation and the control of viruses, has been exploited via host-induced gene silencing (HIGS) to target pathogenicity factors of biotrophic and mycotoxigenic fungal pathogens. These include, but are not limited to Verticillium dahliae (Song and Thomma 2018), Puccinia striiformis (Qi et al. 2017), Fusarium spp. (Chen et al. 2016; Cheng et al. 2015; Ghag et al. 2014; Hu et al. 2015; Koch et al. 2013), and Blumeria graminis (Nowara et al. 2010). Mycotoxin production has also been targeted for Aspergillus flavus in corn (though with off-target effects on kernals; Masanga et al. 2015) and peanut, with near 100% aflatoxin reduction in B1 and B2 chemotypes (Arias et al. 2015).
RNAi has also shown success for controlling necrotrophic pathogens. RNAi was shown to be an effective strategy for controlling Botrytis cinerea (Wang et al. 2016) through the targeting of both Dicer proteins, Bc-Dcl1 and Bc-Dcl2. Dicer proteins process dsRNA into small interfering RNA (siRNA) which are used by B. cinerea to interfere with host immunity genes. Similarly, dcl1 and dcl2 double disruption mutants of S.sclerotiorum exhibited pathogen debilitation and enhanced mycovirus susceptibility, indicating a functioning and important silencing machinery that can be targeted or utilized for silencing with foreign dsRNA (Mochama et al. 2018). These studies demonstrate that targeting single genes versus multiple, functionally redundant genes might improve RNAi silencing efficacy. Additionally, RNA silencing machinery genes such as argonaute proteins (which forms a RISC complex to guide sRNAs to targets) or RNA-dependent RNA polymerases can be potential targets. Andrade et al. (2016) also used RNAi to reduce virulence of S. sclerotiorum through the targeting of the structural gene, chitin synthase (Chs). RNAi can also be deployed to alter pathogen targets or plant genes otherwise exploited by pathogens. Recently, the silencing of NADPH oxidases involved in ROS production needed for S. sclerotiorum infection was undertaken by Ranjan et al. (2018). The introduction of dsRNA using a viral vector through the process of virus-induced gene silencing (VIGS), in this system, enhanced soybean resistance to S. sclerotiorum. RNAi technologies using host-induced silencing provide a promising and precisely-targeted method to enhance host resistance.
Furthermore, new technologies, such as spray-induced gene silencing (SIGS) provides the opportunity to apply dsRNA constructs in a spray form without the regulatory hurdles of bringing genetically engineered technologies to market. Sheet-like clay nanoparticles have been found to extend the integrity of dsRNA on plant surfaces past the five to seven days observed with naked dsRNA, and dsRNA can persist in the presence of clay nanoparticle for up to 30 days (Mitter et al. 2017). These exogenous dsRNA applications can be used in conjunction with RNAseq data for functional studies to understand the importance of various genes to S. sclerotiorum pathogenicity (McLoughlin et al. 2018), although virulence factors may differ in importance depending on the host, as is observed in the case of Ss-Oah1 which is more important for disease development on soybean than pea and faba bean (Xu et al. 2015). One can also speculate that a mycovirus-induced gene silencing vector can be used to silence fungal genes. Gene silencing and editing technologies such as RNAi provide opportunities for the specific, targeted control of SSR and would abate the use of chemical control.
Conclusions
As previously discussed, genetics are one of the most powerful tools to reduce the severity and incidence SSR in soybean. While progress has been made in the identification of resistant cultivars (Bastien et al. 2014; Boland and Hall 1987; Grau et al. 1982; Han et al. 2008; Huynh et al. 2010; Iquira et al. 2015; Kim and Diers 2000; Li et al. 2010; McCaghey et al. 2017; Sebastian et al. 2010; Zhao et al. 2015), a need persists to identify cultivars that sustain heritable resistance across environments (Kim and Diers 2000) and with multiple isolates of S. sclerotiorum (Willbur et al. 2017). Our understanding of the S. sclerotiorum pathosystem has been enhanced by better understanding the role of oxalic acid in immune suppression as well as the identification of specific CWDEs and secreted proteins implicated in pathogenic success. Transgenic and RNAi approaches provide a unique opportunity for the precise targeting of both susceptibility factors on the host side and virulence factors on the pathogen side. However, in the absence of commercially deployed biotechnological approaches in SSR management, soybean farmers across the globe will be left with incomplete management options.
Abbreviations
- BiFC:
-
Bimolecular fluorescence complementation
- CAZymes:
-
carbohydrate active enzymes
- Chs:
-
chitin synthase
- CWDE:
-
cell wall-degrading enzymes
- DSI:
-
disease severity index
- dsRNA:
-
double-stranded RNA
- HIGS:
-
host-induced gene silencing
- HR:
-
hypersensitive response
- MAS:
-
marker assisted selection
- OxDC:
-
oxalate decarboxylase
- OxO:
-
oxalate oxidase
- OA:
-
oxalic acid
- sRNA:
-
small RNA
- SSR:
-
Sclerotinia stem rot
- PG:
-
endopolygalacturonases
- OAH:
-
oxaloacetate acetylhydrolase
- PGIP:
-
polygalacturonase-inhibiting protein
- QTL:
-
quantitative trait loci
- RIL:
-
recombinant inbred lines
- RNAi:
-
RNA interference
- ROS:
-
reactive oxygen species
- siRNA:
-
small interfering RNA
- SIGS:
-
spray-induced gene silencing
- VIGS:
-
virus-induced gene silencing
References
Amselem J, Cuomo CA, van Kan JAL, Viaud M, Benito EP, Couloux A, Coutinho PM, de Vries RP, Dyer PS, Fillinger S, Fournier E, Gout L, Hahn M, Kohn L, Lapalu N, Plummer KM, Pradier J‐M, Quévillon E, Sharon A, Simon A, ten Have A, Tudzynski B, Tudzynski P, Wincker P, Andrew M, Anthouard V, Beever RE, Beffa R, Benoit I, Bouzid O, Brault B, Chen Z, Choquer M, Collémare J, Cotton P, Danchin EG, Da Silva C, Gautier A, Giraud C, Giraud T, Gonzalez C, Grossetete S, Güldener U, Henrissat B, Howlett B, Kodira C, Kretschmer M, Lappartient A, Leroch M, Levis C, Mauceli E, Neuvéglise C, Oeser B, Pearson M, Poulain J, Poussereau N, Quesneville H, Rascle C, Schumacher J, Ségurens B, Sexton A, Silva E, Sirven C, Soanes DM, Talbot NJ, Templeton M, Yandava C, Yarden O, Zeng Q, Rollins JA, Lebrun M‐H, Dickman M (2011) Genomic analysis of the necrotrophic fungal pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS Genetics 7:e1002230
Andrade CM, Tinoco MLP, Rieth AF, Maia FCO, Aragão FJL (2016) Host-induced gene silencing in the necrotrophic fungal pathogen Sclerotinia sclerotiorum. Plant Pathology 65:626–632
Arahana VS, Graef GL, Specht JE, Steadman JR, Eskridge KM (2001) Identification of QTLs for resistance to in soybean. Crop Science 41:180–188
Arias RS, Dang PM, Sobolev VS (2015) RNAi-mediated control of aflatoxins in peanut: method to analyze mycotoxin production and transgene expression in the peanut/Aspergillus pathosystem. Journal of Visualized Experiments: JoVE 106
Asoufi H, Hameed KM, Mahasneh A (2007) The cellulase and pectinase activities associated with the virulence of indigenous Sclerotinia sclerotiorum isolates in Jordan Valley. The Plant Pathology Journal 23:233–238
Bastien M, Sonah H, Belzile F (2014) Genome wide association mapping of resistance in soybean with a genotyping-by-sequencing approach. The Plant Genome 7:1–13
Bateman DF, Beer SV (1965) Simultaneous production and synergistic action of oxalic acid and polygalacturonase during pathogenesis by Sclerotium rolfsii. Phytopathology 55:204–211
Boland GJ, Hall R (1987) Evaluating soybean cultivars for resistance to Sclerotinia sclerotiorum under field conditions. Plant Disease 71:934–936
Bourras S, McNally KE, Müller MC, Wicker T, Keller B (2016) Avirulence genes in cereal powdery mildews: the gene-for-gene hypothesis 2.0. Frontiers in Plant Science 7:241
Cessna SG, Sears VE, Dickman MB, Low PS (2000) Oxalic acid, a pathogenicity factor for Sclerotinia sclerotiorum, suppresses the oxidative burst of the host plant. The Plant Cell 12:2191–2199
Chang HX, Sang H, Wang J, McPhee KE, Zhuang X, Porter LD, Chilvers MI (2018) Exploring the genetics of lesion and nodal resistance in pea (Pisum sativum L.) to Sclerotinia sclerotiorum using genome-wide association studies and RNA-Seq. Plant Direct 2:e00064
Chen C, Harel A, Gorovoits R, Yarden O, Dickman MB (2004) MAPK regulation of sclerotial development in Sclerotinia sclerotiorum is linked with pH and cAMP sensing. Molecular Plant-Microbe Interactions 17:404–413
Chen W, Kastner C, Nowara D, Oliveira-Garcia E, Rutten T, Zhao Y, Deising HB, Kumlehn J, Schweizer P (2016) Host-induced silencing of Fusarium culmorum genes protects wheat from infection. Journal of Experimental Botany 67:4979–4991
Cheng W, Song XS, Li HP, Cao LH, Sun K, Qiu XL, Xu YB, Yang P, Huang T, Zhang JB, Qu B (2015) Host-induced gene silencing of an essential chitin synthase gene confers durable resistance to Fusarium head blight and seedling blight in wheat. Plant Biotechnology Journal 13:1335–1345
Cline MN, Jacobsen BJ (1983) Methods for evaluating soybean cultivars for resistance to Sclerotinia sclerotiorum. Plant Disease 67:784–786
Cober ER, Rioux S, Rajcan I, Donaldson PA, Simmonds DH (2003) Partial resistance to white mold in a transgenic soybean line. Crop Science 43:92
Conley SP, Roth AC, Gaska JM, and Smith DL (2017) 2017 Wisconsin soybean performance trials. Departments of Plant Pathology and Agronomy, University of Wisconsin, Madison. Retrieved from coolbean.info: http://www.coolbean.info/library/documents/2017_Soybean_VT_FINAL.pdf
Cunha WG, Tinoco MLP, Pancoti HL, Ribeiro RE, Aragão FJL (2010) High resistance to Sclerotinia sclerotiorum in transgenic soybean plants transformed to express an oxalate decarboxylase gene. Plant Pathology 59:654–660
Curtis MJ, Wolpert TJ (2002) The oat mitochondrial permeability transition and its implication in victorin binding and induced cell death. Plant Journal 29:295–312
De Lorenzo G, Ferrari S (2002) Polygalacturonase-inhibiting proteins in defense against phytopathogenic fungi. Current Opinion in Plant Biology 5:295–299
Del Rio L, Kurtzweil NC, Grau CR (2001) Petiole inoculation as a tool to screen soybean germ plasm for resistance to Sclerotinia sclerotiorum. (Abstract) Phytopathology 91:S176
Derbyshire M, Denton-Giles M, Hegedus D, Seifbarghy S, Rollins J, van Kan J, Seidl MF, Faino L, Mbengue M, Navaud O, Raffaele S, Hammond-Kosack K, Heard S, Oliver R (2017) The complete genome sequence of the phytopathogenic fungus Sclerotinia sclerotiorum reveals insights into the genome architecture of broad host range pathogens. Genome Biology and Evolution 9:593–618
Derbyshire MC, Mbengue M, Barascud M, Navaud O, Raffaele S (2018) Small RNAs from the plant pathogenic fungus Sclerotinia sclerotiorum highlight candidate host target genes associated with quantitative disease resistance. bioRxiv 1:354076
Dickman MB, Mitra A (1992) Arabidopsis thaliana as a model for studying Sclerotinia sclerotiorum pathogenesis. Physiological and Molecular Plant Pathology 41:255–263
Donaldson PA, Anderson T, Lane BG, Davidson AL, Simmonds DH (2001) Soybean plants expressing an active oligomeric oxalate oxidase from the wheat gf-2.8 (germin) gene are resistant to the oxalate-secreting pathogen Sclerotina sclerotiorum. Physiological Molecular Plant Pathology 59:297–307
Dumas B, Sailland A, Cheviet JP, Freyssinet G, Pallett K (1993) Identification of barley oxalate oxidase as a germin-like protein. Comptes Rendus de l'Academie des Sciences. Serie III, Sciences de la Vie 316:793–798
Dutton MV, Evans CS (1996) Oxalate production by fungi: its role in pathogenicity and ecology in the soil environment. Canadian Journal of Microbiology 42:881–895
Ghag SB, Shekhawat UK, Ganapathi TR (2014) Host-induced post-transcriptional hairpin RNA-mediated gene silencing of vital fungal genes confers efficient resistance against fusarium wilt in banana. Plant Biotechnology Journal 12:541–553
Girard IJ, Tong C, Becker MG, Mao X, Huang J, de Kievit T, Fernando WD, Liu S, Belmonte MF (2017) RNA sequencing of Brassica napus reveals cellular redox control of Sclerotinia infection. Journal of Experimental Botany 68:5079–5091
Godoy G, Steadman JR, Dickman MB, Dam R (1990) Use of mutants to demonstrate the role of oxalic acid in pathogenicity of Sclerotinia sclerotiorum on Phaseolus vulgaris. Physiological and Molecular Plant Pathology 37:179–191
Grau CR, Radke VL, Gillespie FL (1982) Resistance of soybean cultivars to Sclerotinia sclerotiorum. Plant Disease 66:506–508
Guo X, Wang D, Gordon SG, Helliwell E, Smith T, Berry SA, Dorrance AE (2008) Genetic mapping of QTLs underlying partial resistance to in soybean PI 391589A and PI 391589B. Crop Science 48:1129–1139
Guyon K, Balagué C, Roby D, Raffaele S (2014) Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum. BMC Genomics 15:336
Han F, Katt M, Schuh W, and Webb DM (2008) QTL controlling Sclerotinia stem rot resistance in soybean. U.S. Patent 7250,552. Date issued: 18 September
Hu Z, Parekh U, Maruta N, Trusov Y, Botella JR (2015) Down-regulation of Fusarium oxysporum endogenous genes by host-delivered RNA interference enhances disease resistance. Frontiers in Chemistry 3:1
Huynh TT, Bastien M, Iquira E, Turcotte P, Belzile F (2010) Identification of QTLs associated with partial resistance to white mold in soybean using field-based inoculation. Crop Science 50:969–979
Huzar-Novakowiski J, Dorrance AE (2018) Ascospore inoculum density and characterization of components of partial resistance to Sclerotinia sclerotiorum in soybean. Plant Disease 102:1326–1333
Iquira E, Humira S, François B (2015) Association mapping of QTLs for Sclerotinia stem rot resistance in a collection of soybean plant introductions using a genotyping by sequencing (GBS) approach. BMC Plant Biology 15:5
Joshi RK, Megha S, Rahman MH, Basu U, Kav NN (2016) A global study of transcriptome dynamics in canola (Brassica napus L.) responsive to Sclerotinia sclerotiorum infection using RNA-Seq. Gene 590:57–67
Kabbage M, Williams B, Dickman MB (2013) Cell death control: the interplay of apoptosis and autophagy in the pathogenicity of Sclerotinia sclerotiorum. PLoS Pathogens 9:e1003287
Kabbage M, Yarden O, Dickman MB (2015) Pathogenic attributes of Sclerotinia sclerotiorum: switching from a biotrophic to necrotrophic lifestyle. Plant Science 233:53–60
Kesarwani M, Azam M, Natarajan K, Mehta A, Datta A (2000) Oxalate decarboxylase from Collybia velutipes molecular cloning and its overexpression to confer resistance to fungal infection in transgenic tobacco and tomato. Journal of Biological Chemistry 275:7230–7238
Kim HS, Diers BW (2000) Inheritance of partial resistance to sclerotinia stem rot in soybean research supported by the Michigan Agricultural Experiment Station and grants from the Michigan Soybean Promotion Committee. Crop Science 40:55–61
Kim HS, Sneller CH, Diers BW (1999) Evaluation of soybean cultivars for resistance to Sclerotinia stem rot in field environments. Crop Science 39:64
Kim KS, Min JY, Dickman MB (2008) Oxalic acid is an elicitor of plant programmed cell death during Sclerotinia sclerotiorum disease development. Molecular Plant-Microbe Interactions 21:605–612
Koch A, Kumar N, Weber L, Keller H, Imani J, Kogel KH (2013) Host-induced gene silencing of cytochrome P450 lanosterol C14α-demethylase–encoding genes confers strong resistance to fusarium species. Proceedings of the National Academy of Sciences 110:19324–19329
Kotchoni SO, Gachomo EW (2006) The reactive oxygen species network pathways: an essential prerequisite for perception of pathogen attack and the acquired disease resistance in plants. Journal of Biosciences 31:389–404
Kubicek CP, Starr TL, Glass NL (2014) Plant cell wall–degrading enzymes and their secretion in plant-pathogenic fungi. Annual Review of Phytopathology 52:427–451
Kurian P, Stelzig DA (1979) The synergistic role of oxalic acid and endopolygalacturonase in bean leaves infected by Cristulariella pyramidalis. Phytopathology 69:1301–1304
Lane BG, Dunwell JM, Ray JA, Schmitt MR, Cuming AC (1993) Germin, a protein marker of early plant development, is an oxalate oxidase. Journal of Biological Chemistry 268:12239–12242
Levine A, Tenhaken R, Dixon R, Lamb C (1994) H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell 79:583–593
Li D, Sun M, Han Y, Teng W, Li W (2010) Identification of QTL underlying soluble pigment content in soybean stems related to resistance to soybean white mold (Sclerotinia sclerotiorum). Euphytica 172:49–57
Liang HJ, Di YL, Li JL, Zhu FX (2015) Baseline sensitivity and control efficacy of fluazinam against Sclerotinia sclerotiorum. European Journal of Plant Pathology 142:691–699
Liang X, Liberti D, Li M, Kim YT, Hutchens A, Wilson R, Rollins JA (2015a) Oxaloacetate acetylhydrolase gene mutants of Sclerotinia sclerotiorum do not accumulate oxalic acid, but do produce limited lesions on host plants. Molecular Plant Pathology 16:559–571
Liang X, Moomaw EW, Rollins JA (2015b) Fungal oxalate decarboxylase activity contributes to Sclerotinia sclerotiorum early infection by affecting both compound appressoria development and function. Molecular Plant Pathology 16:825–836
Liu Z, Zhang Z, Faris JD, Oliver RP, Syme R, McDonald MC, McDonald BA, Solomon PS, Lu S, Shelver WL, Xu S, Friesen TL (2012) The cysteine rich necrotrophic effector SnTox1 produced by Stagonospora nodorum triggers susceptibility of wheat lines harboring Snn1. PLoS Pathogens 8:e1002467
Lumsden RD (1969) Sclerotinia sclerotiorum infection of bean and the production of cellulase. Phytopathology 59:653–657
Lyu X, Shen C, Fu Y, Xie J, Jiang D, Li G, Cheng J (2016) A small secreted virulence-related protein is essential for the necrotrophic interactions of Sclerotinia sclerotiorum with its host plants. PLoS Pathogens 12:e1005435
Masanga JO, Matheka JM, Omer RA, Ommeh SC, Monda EO, Alakonya AE (2015) Downregulation of transcription factor aflR in Aspergillus flavus confers reduction to aflatoxin accumulation in transgenic maize with alteration of host plant architecture. Plant Cell Reports 34:1379–1387
McCaghey MM, Willbur J, Ranjan A, Grau CR, Chapman S, Diers B, Groves C, Kabbage M, Smith DL (2017) Development and evaluation of Glycine max germplasm lines with quantitative resistance to Sclerotinia sclerotiorum. Frontiers in Plant Science 8:1495
McLoughlin AG, Wytinck N, Walker PL, Girard IJ, Rashid KY, Kievit T, Fernando WD, Whyard S, Belmonte MF (2018) Identification and application of exogenous dsRNA confers plant protection against Sclerotinia sclerotiorum and Botrytis cinerea. Scientific Reports 8:7320
Miklas PN, Grafton KF (1992) Inheritance of partial resistance to white mold in inbred populations of dry bean. Crop Science 32:943–948
Mitter N, Worrall EA, Robinson KE, Li P, Jain RG, Taochy C, Fletcher SJ, Carroll BJ, Lu GM, Xu ZP (2017) Clay nanosheets for topical delivery of RNAi for sustained protection against plant viruses. Nature Plants 3:16207
Mittler R, Vanderauwera S, Suzuki N, Miller G, Tognetti VB, Vandepoele K, Gollery M, Shulaev V, Van Breusegem F (2011) ROS signaling: the new wave? Trends in Plant Science 16:300–309
Mochama P, Jadhav P, Neupane A, Marzano SY (2018) Mycoviruses as triggers and targets of RNA silencing in white mold fungus Sclerotinia sclerotiorum. Viruses 10:214
Moellers TC, Singh A, Zhang J, Brungardt J, Kabbage M, Mueller DS, Grau CR, Ranjan A, Smith DL, Chowda-Reddy RV, Singh AK (2017) Main and epistatic loci studies in soybean for Sclerotinia sclerotiorum resistance reveal multiple modes of resistance in multi-environments. Scientific Reports 7:3554
Munir E, Yoon JJ, Tokimatsu T, Hattori T, Shimada M (2001) New role for glyoxylate cycle enzymes in wood-rotting basidiomycetes in relation to biosynthesis of oxalic acid. Journal of Wood Science 47:368–373
Noctor G, Mhamdi A, Foyer CH (2014) The roles of reactive oxygen metabolism in drought: not so cut and dried. Plant Physiology 164:1636–1648
Nowara D, Gay A, Lacomme C, Shaw J, Ridout C, Douchkov D, Hensel G, Kumlehn J, Schweizer P (2010) HIGS: host-induced gene silencing in the obligate biotrophic fungal pathogen Blumeria graminis. The Plant Cell 22:3130–3141
Oliveira MB, de Andrade RV, Grossi-de-Sá MF, Petrofeza S (2015) Analysis of genes that are differentially expressed during the Sclerotinia sclerotiorum–Phaseolus vulgaris interaction. Frontiers in Microbiology 6:1162
Pérès (1995) Sclerotinia du soja: Premières évaluations du comportment variétal. CETIOM-Oléoscope 26:28–29
Petrov VD, Van Breusegem F (2012) Hydrogen peroxide- a central hub for information flow in plant cells. AoB Plants 289:8735–8741
Prade RA, Zhan D, Ayoubi P, Mort AJ (1999) Pectins, pectinases and plant-microbe interactions. Biotechnology and Genetic Engineering Reviews 16:361–392
Qi T, Zhu X, Tan C, Liu P, Guo J, Kang Z, Guo J (2017) Host-induced gene silencing of an important pathogenicity factor PsCPK1 in Puccinia striiformis f. sp. tritici enhances resistance of wheat to stripe rust. Plant Biotechnology Journal 16:797–807
Raffaele S, Kamoun S (2012) Genome evolution in filamentous plant pathogens: why bigger can be better. Nature Reviews Microbiology 10:417–430
Rafiqi M, Ellis JG, Ludowici VA, Hardham AR, Dodds PN (2012) Challenges and progress towards understanding the role of effectors in plant–fungal interactions. Current Opinion in Plant Biology 15:477–482
Ranjan A, Jayaraman D, Grau C, Hill JH, Whitham SA, Ané JM, Smith DL, Kabbage M (2018) The pathogenic development of Sclerotinia sclerotiorum in soybean requires specific host NADPH oxidases. Molecular Plant Pathology 19:700–714
Riou C, Freyssinet G, Fevre M (1991) Production of cell wall-degrading enzymes by the phytopathogenic fungus Sclerotinia sclerotiorum. Applied Environmental Microbiology 57:1478–1484
Rollins JA (2003) The Sclerotinia sclerotiorum pac1 gene is required for sclerotial development and virulence. Molecular Plant-Microbe Interactions 16:785–795
Rollins JA, Dickman MB (2001) pH signaling in Sclerotinia sclerotiorum: identification of a pacC/RIM1 homolog. Applied and Environmental Microbiology 67:75–81
Rousseau G, Huynh Thanh T, Dostaler D, Rioux S (2004) Greenhouse and field assessments of resistance in soybean inoculated with sclerotia, mycelium, and ascospores of Sclerotinia sclerotiorum. Canadian Journal of Plant Science 84:615–623
Sebastian SA, Lu H, Han F, Kyle D, and Hedges BR (2010). Genetic loci associated with Sclerotinia tolerance in soybean. Unites States Patent 7,790,949 B2. Date issued: 7 September
Seifbarghi S, Borhan MH, Wei Y, Coutu C, Robinson SJ, Hegedus DD (2017) Changes in the Sclerotinia sclerotiorum transcriptome during infection of Brassica napus. BMC Genomics 18:266
Sewelam N, Kazan K, Schenk PM (2016) Global plant stress signaling: reactive oxygen species at the cross-road. Frontiers in Plant Science 7:187
Song Y, Thomma BP (2018) Host-induced gene silencing compromises Verticillium wilt in tomato and Arabidopsis. Molecular Plant Pathology 19:77–89
SoyBase (2010) the USDA-ARS soybean genetics and genomics database. http://soybase.org
Uloth MB, Clode PL, You MP, Barbett MJ (2015) Calcium oxalate crystals: an integral component of the Sclerotinia sclerotiorum/Brassica carinata pathosystem. PLoS One 10:e0122362
Vuong TD, Hoffman DD, Diers BW, Miller JF, Steadman JR, Hartman GL (2004) Evaluation of soybean, dry bean, and sunflower for resistance to Sclerotinia sclerotiorum. Crop Science 44:777–783
Vuong TD, Diers BW, Hartman GL (2008) Identification of QTL for resistance to Sclerotinia stem rot in soybean plant introduction 194639. Crop Science 48:2209
Wang M, Weiberg A, Lin FM, Thomma BP, Huang HD, Jin H (2016) Bidirectional cross-kingdom RNAi and fungal uptake of external RNAs confer plant protection. Nature Plants 2:16151
Willats WG, McCartney L, Mackie W, Knox JP (2001) Pectin: cell biology and prospects for functional analysis. Plant Molecular Biology 47:9–27
Willbur JF, Ding S, Marks ME, Lucas H, Grau CR, Groves CL, Kabbage M, Smith DL (2017) Comprehensive Sclerotinia stem rot screening of soybean germplasm requires multiple isolates of Sclerotinia sclerotiorum. Plant Disease 101:344–353
Williams B, Kabbage M, Kim HJ, Britt R, Dickman MB (2011) Tipping the balance: Sclerotinia sclerotiorum secreted oxalic acid suppresses host defenses by manipulating the host redox environment. PLoS Pathogens 7:e1002107
Williams SJ, Yin L, Foley G, Casey LW, Outram MA, Ericsson DJ, Lu J, Boden M, Dry IB, Kobe B (2016) Structure and function of the TIR domain from the grape NLR protein RPV1. Frontiers in Plant Science 7:1850
Wojtaszek P (1997) Mechanisms for the generation of reactive oxygen species in plant defense response. Acta Physiologiae Plantarum 19:581–589
Xia XJ, Zhou YH, Shi K, Zhou J, Foyer CH, Yu JQ (2015) Interplay between reactive oxygen species and hormones in the control of plant development and stress tolerance. Journal of Experimental Botany 66:2839–2856
Xiao X, Xie J, Cheng J, Li G, Yi X, Jiang D, Fu Y (2014) Novel secretory protein Ss-Caf1 of the plant-pathogenic fungus Sclerotinia sclerotiorum is required for host penetration and normal sclerotial development. Molecular Plant-Microbe Interactions 40:40–55
Xu L, Xiang M, White D, Chen W (2015) pH dependency of sclerotial development and pathogenicity revealed by using genetically defined oxalate-minus mutants of Sclerotinia sclerotiorum. Environmental Microbiology 17:2896–2909
Yang G, Tang L, Gong Y, Xie J, Fu Y, Jiang D, Li G, Collinge DB, Chen W, Cheng J (2018) A cerato-platanin protein SsCP1 targets plant PR1 and contributes to virulence of Sclerotinia sclerotiorum. New Phytologist 217:739–755
Yu Y, Xiao J, Zhu W, Yang Y, Mei J, Bi C, Qian W, Qing L, Tan W (2017) Ss-Rhs1, a secretory Rhs repeat-containing protein, is required for the virulence of Sclerotinia sclerotiorum. Molecular Plant Pathology 18:1052–1061
Zhao Z, Liu H, Wang C, Xu JR (2013) Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics 14:274
Zhao X, Han Y, Li Y, Liu D, Sun M, Zhao Y, Lv C, Li D, Yang Z, Huang L, Teng W (2015) Loci and candidate gene identification for resistance to Sclerotinia sclerotiorum in soybean (Glycine max L. Merr.) via association and linkage maps. The Plant Journal 82:245–255
Zhou T, Boland GJ (1999) Mycelial growth and production of oxalic acid by virulent and hypovirulent isolates of Sclerotinia sclerotiorum. Canadian Journal of Plant Pathology 21:93–99
Zhu W, Wei W, Fu Y, Cheng J, Xie J, Li G, Yi X, Kang Z, Dickman MB, Jiang D (2013) A secretory protein of necrotrophic fungus Sclerotinia sclerotiorum that suppresses host resistance. PLoS One 8:e53901
Zuppini A, Navazio L, Sella L, Castiglioni C, Favaron F, Mariani P (2005) An endopolygalacturonase from Sclerotinia sclerotiorum induces calcium-mediated signaling and programmed cell death in soybean cells. Molecular Plant-Microbe Interactions 18:849–855
Acknowledgements
We would like to thank the funding sources that make our work and inquiry possible including the Wisconsin Soybean Marketing Board (WSMB), the North Central Soybean Research Program (NCSRP), SciMed at the University of Wisconsin-Madison, and the Department of Plant Pathology at University of Wisconsin-Madison.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Leandro J. Dallagnol
Rights and permissions
About this article
Cite this article
McCaghey, M., Willbur, J., Smith, D.L. et al. The complexity of the Sclerotinia sclerotiorum pathosystem in soybean: virulence factors, resistance mechanisms, and their exploitation to control Sclerotinia stem rot. Trop. plant pathol. 44, 12–22 (2019). https://doi.org/10.1007/s40858-018-0259-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-018-0259-4