Abstract
Flowering time is of great significance for crop reproduction, yield, and regional adaptability, which is intricately regulated by various environmental cues and endogenous signals. Photoperiod is a key flowering regulating factor due to its relatively high concern in the same geography, while other factors change year over year. Photoperiodic flowering time is controlled by a complex genetic network, which is regulated by florigen. RICE FLOWERING LOCUS T 1 (RFT1) is the closest homologue of Heading date 3a (Hd3a), which is thought to encode a mobile flowering signal and promote floral transition under short-day (SD) conditions in rice. Recent molecular genomic advancement revealed how to identify the molecular nature of florigen, its function, expression, and how to diversify its regulatory pathway. Here, we summarized the recent understanding of flowering time regulation mainly focused on rice Hd3a, its molecular function, expression, and diversified photoperiodic regulatory pathways under short-day and long-day (LD) conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Rice is one of the most important cereal crops and has wide adaptability in diverse climatic regions ranging from 44° N to 35° S and 6 ft below to 2700 ft above sea level (Molla 2022), which traits and mechanisms confer this adaptability is a major question in rice biology. Heading date (or flowering time) is a crucial trait for the regional and seasonal adaptation of rice and influences grain yield. Rice is a facultative short-day (SD) plant; SD conditions promote, and long-day (LD) conditions inhibit flowering (Sohail et al. 2022). Flowering time is a dramatic change from the vegetative stage to the reproductive stage and is predominantly controlled by genetic pathways, integrated with internal and external cues (Wu et al. 2018; Zhang et al. 2019). Most plant species can initiate flowering for reproduction mainly depending on environmental changes in photoperiod and temperature (Song et al. 2015; Tan et al. 2016). Flowering time is vital for seasonal changes, regional adaptability, and often affluent to perceive (Zhang et al. 2017). Plants are generally classified into three categories regarding photoperiodic flowering initiation response; long-day plants (LDPs), initiate flowering when the day length becomes longer, short-day plants (SDPs), start flowering when the day length becomes shorter, while day-neutral plants (DNPs) are unaffected by photoperiodic fluctuations. The molecular mechanism of photoperiodic FT has been comprehensively examined in a model LDP (Arabidopsis thaliana) (Song et al. 2015) and a traditional SDP (Oryza sativa L.) (Tsuji et al. 2011; Shrestha et al. 2014). Flowering time regulation is controlled by several genes, which are suppressed/expressed in an association with environmental dynamics, i.e., photoperiod and temperature (Huan et al. 2018). Recent studies reported FLOWERING LOCUS T (FT) and Heading date 3a (Hd3a), as florigen in Arabidopsis and rice, respectively. These florigen moves in a form of mobile signals from the leaf to shoot apical meristem (SAM) and commence flowering (Goretti et al. 2017). More recently, the molecular mechanism of florigen function in shoot apical cells was revealed in rice. Hd3a florigen interacts with 14-3-3 proteins in the cytoplasm and forms a ternary complex with OsFD1 in the nucleus (Tsuji 2017). The ternary complex is known as the florigen activation complex (FAC), which activates OsMADS15, homologue of A. thaliana APETALA1 (AP1), which leads to flowering (Camoni et al. 2018). FD is a basic leucine zipper (bZIP)-containing transcription factor, and its loss-of-function mutants are late flowering (Taoka et al. 2011). These results indicate that 14-3-3 proteins act as intracellular receptors for florigen in shoot apical cells and offer new approaches to manipulate flowering in various crops and trees. Advancement in molecular genetics provides a clear concept of flowering time regulation mechanism in rice from molecular genetics, molecular biology, and comparative biology point of view. In this review article, we deliberate our understanding of flowering time, using the current findings of a strong candidate for rice mobile flowering signals, florigen, and incorporated the presence of complex layers of gene networks integrated with the synthesis of florigen protein and its subsequent transport and perception.
2 Rice florigen
Rice has two florigens, Heading date 3a (Hd3a) and RICE FLOWERING LOCUS T 1 (RFT1), which are crucial for FT in rice. Regulations of these florigens involve a complex genetic network, which is discussed below.
3 Hd3a protein as rice florigen
Florigen, a mobile signal that moves from leaves to SAM and induces flowering, has been elucidating since it was identified 70 years ago. Understanding the nature of mobile floral signals provides insight into the molecular mechanism of flowering induction (Brambilla and Fornara 2013). Recent research identified FT, encoding florigen in Arabidopsis, whereas Hd3a, an ortholog of FT, was reported in rice, which moves from leaves to SAM and causes flowering. These results showed that Hd3a is florigen in rice (Tamaki et al. 2007).
Hd3a was first identified as a quantity traits locus (QTL) that promotes flowering in rice under inductive short-day conditions (Yamamoto et al. 1998). The overexpression of Hd3a with cumulative promoter promoted flowering time, while RNA interference (RNAi) of Hd3a suppressed flowering in rice. (Monna et al. 2002).
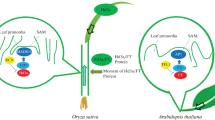
The size of Hd3a/FT is about 22 kDa and their structures are similar to mammalian phosphatidylethanolamine-binding proteins (PEBPs) or Raf kinase inhibitor proteins (RKIP) (Ahn et al. 2006). However, it is not clear whether Hd3a/FT binds or not with phosphatidylethanolamine and kinase inhibitor. Recent molecular studies revealed that FT has a strong downstream effect on the flowering promotion of the known photoperiodic flowering pathway in Arabidopsis (Kobayashi and Weigel 2007; Guo et al. 2020; Zong et al. 2020). Arabidopsis and rice flowering time are assisted by LD and SD, respectively (Liu et al. 2020). FT mainly activates floral activity in Arabidopsis that expresses in leaves vascular tissue and transfers to SAM to induce flowering (Kobayashi et al. 1999; Shim and Jang 2020). The interaction of FT with the bZIP transcription activation factor results in full activation of FLOWERING LOCUS T (FD) in SAM, this activation is necessary for floral promotion in rice (Fig. 1) (Abe et al. 2008; Peng et al. 2021).
Anatomical research showed that the product of FT is the main site for floral expression. If FT codes florigen and can act as a mobile floral agent via transcripts (mRNA) or protein, either of which interacts with FD in SAM. Now the question is whether mRNA or protein acts as a mobile agent for florigen activity (Zeevaart 2006). So the question was responded with rice as a model SD plant to elucidate the nature of florigen activity. Firstly, to determine the exact Hd3a transcription site and accumulation of mRNA. Using transgenic plants, the promoter activity of β-glucuronidase (GUS) was expressed in the presence of the Hd3a promoter in leaf blades with no expression in SAM. FT expression is tissue-specific in Arabidopsis, while this expression is promoted by FT in LD (Arabidopsis) and Hd3a in SD (rice) (Kobayashi and Weigel 2007). Hd3a mRNA stores more in leaf blade but lower in leaf sheet and much lower than ¼ in SAM under inductive SD conditions. Hd3a expression is specific to the vascular tissue of leaf blades. Hence, it is different from Hd3a mRNA, which transfers a significant amount from leaves to SAM (Fig. 1) (Jang et al. 2008).
Secondly, the tissue localization of Hd3a was studied from rice transgenic plants driven from the Hd3a promoter expressed as Hd3a-GFP fusion protein. Hd3a-GFP is located in the vascular tissue of leaf blades just below SAM where the node is localized. Thus, Hd3a-GFP protein is produced in the vascular tissue of the leaf blade and conveyed to the phloem where it is unloaded from vascular tissue into meristem tissue just below the SAM (Tamaki et al. 2007). Necessarily, the location of Hd3a-GFP was tested in other vascular tissues using different tissue-specific promoters conforming to the expression of Hd3a-GFP, revealing that Hd3a protein transmits from leaves and vasculature to SAM. This discussion revealed that Hd3a acts as a mobile flowering signal in rice and Hd3a/FT long-distance mobility is observed from rice proteome phloem sap (Aki et al. 2008). Hd3a mobility has probably not been detected because of photoperiod conditions, and sampling time was unsuitable for Hd3a detection. However, Hd3a/FT protein has been noticed in the phloem sap tissue of CmFTL2, an Hd3a/FT ortholog of Cucurbita maxim, which moves from graft union and induces flowering (Lin et al. 2007). Phloem sap acquired from the Brassica napus inflorescence also contains FT protein (Giavalisco et al. 2006). The above discussion provides additional information that Hd3a/FT is a florigen protein. Arabidopsis behavior also showed that FT is florigen. Phloem expresses FT-GFP and Hd3a-GFP transfer to SAM and promotes flowering but FT nuclear localization does not because of immobilization (Mathieu et al. 2007; Jaeger and Wigge 2007; Wu et al. 2020).
4 RFT1 is essential for flowering in rice
RFT1 is also the second florigen and the closest homolog of Hd3a and 13 FT-like proteins in rice. RFT1 is located on chromosome 6, 11.5 kb away from Hd3a. RFT1 is 91% similar to Hd3a and most likely produced by tandem duplication (Sun et al. 2014; Hori et al. 2016). Tissue-specific expression of RFT1 and Hd3a is the same under SD. Therefore, RFT1 may function with Hd3a as the second florigen in rice. The RFT1 involvement as florigen is examined from a detailed analysis of Hd3a suppression by RNAi rice plants. The flowering time of Hd3a RNAi plants was more than 30 days late, while RFT1 RNAi flowering time was normal under SD conditions, indicating that RFT1 did not promote flowering on its own. Both Hd3a and RFT1 were suppressed in double RNAi plants and did not flower earlier than 300 days of germination. Thus, RFT1/Hd3a is involved completely in the regulation of rice flowering time. The expression level of RFT1 is relatively low in wild rice as compared to Hd3a RNAi and is associated with the reproductive transition in SAM of rice plants. The upregulation of RFT1 is associated with an abundance of histone-3-acetylation (H3A) near the transition induction site (Tan et al. 2016; Zhu et al. 2017). The above discussion suggests that the expression of RFT1 enhances in the absence of Hd3a, and RFT1 is Hd3a complementary in function. Therefore, RFT1 acts as the second florigen in rice flowering time regulation under specific conditions (Nemoto et al. 2016).
5 Gene network of flowering regulation in rice
Recently, several flowering genes have been cloned, which play an important role in photoperiodic flowering regulation (Zhang et al. 2019). Rice photoperiodic flowering is monitored by two independent signaling pathways. The GIGANTEA-Heading date-1-Heading date 3a (OsGI–Hd1–Hd3a) pathway is evolutionarily associated with the Arabidopsis GIGANTEA-CONSTANS-FLOWERING LOCUS T (GI–CO–FT) pathway (Song et al. 2015). The evolutionarily unique pathway, Early heading date 1-Hd3a/Rice FT1 (Ehd1-Hd3a/RFT1), controlled photoperiodic flowering in rice but does not function in Arabidopsis due to the lack of an Ehd1 ortholog (Doi et al. 2004; Song et al. 2015). Heading date 3a (Hd3a) and RICE FLOWERING LOCUS T1 (RFT1) are two florigens, which regulate flowering time under short-day (SD) and long-day (LD) conditions in rice (Tsuji et al. 2013). Hd3a and RFT1expression are activated under SD and LD conditions, respectively (Chardon and Damerval 2005; Komiya et al. 2009). Hd3a/RFT1 is mainly regulated by Hd1 and Ehd1 under both SD and LD conditions (Gao et al. 2013). Ehd1 is upregulated by Early heading date 2 (Ehd2) (Matsubara et al. 2008), Ehd3 (Matsubara et al. 2011), Ehd4 (Gao et al. 2013), Oryza sativa GIGENTA (OsGI) (Park et al. 2008), Ghd8 (Yan et al. 2011), OsMADS51 (Ryu et al. 2009), OsMFT1 (Song et al. 2018), and OsMADS51 (Kim et al. 2007), while downregulated by CONSTANS-Like 4 (OsCOL4) (Lee et al. 2010), OsCOL9 (Liu et al. 2016), and OsCOL15 (Wu et al. 2018) under SD. In contrast, Ehd1 is upregulated by Ehd4 (Gao et al. 2013), OsMADS56, OsMADS51 (Ryu et al. 2009), Ghd8 (Yan et al. 2011), LEC2, and FUSCA3-Like 1 (OsLFL1) (Peng et al. 2008), and OsVIL2 (O. Sativa VIN3‐LIKE 2) (Hori et al. 2013), while downregulated by OsCOL4 (Wu et al. 2018), OsCOL9 (Liu et al. 2016), OsCOL10 (Tan et al. 2016), OsCOL13 (Sheng et al. 2016), OsCOL15 (Wu et al. 2018), OsCOL16 (Wu et al. 2017), DTH8 (Zhu et al. 2017), and OsELF3 (Yang et al. 2013) (Se13, OsTrx1 under LD conditions. Similarly, Hd1 is upregulated by OsGI under both SD and LD conditions (Fig. 2) (Hori et al. 2013).
6 Photoperiods regulate the molecular genetic pathways of rice flowering
It is clear from the molecular study of Hd3a/FT protein that the induction of flowering is mainly measured from the expression level of Hd3a/FT genes that promote rice flowering. Recent studies provide the molecular understanding of Hd3a photoperiod regulation and the important features of this day-length regulation are LD suppression and SD promotion of Hd3a (Fig. 2).
7 Hd3a promotion under SD
Induction of Hd3a expression is at its peak about 30 days before the floral transition occurs under SD conditions in rice (Komiya et al. 2008). The circadian peak of Hd3a activity and mRNA production correspond at dawn, referring to an important unit of Hd3a transcriptional regulation (Ishikawa et al. 2005; Tamaki et al. 2007). Hd3a expression is mainly regulated by Hd1 and Ehd1 (Izawa 2007). Hd1 was first reported as a major photoperiodic-sensitive QTL, it translates B-box zinc finger protein with C-terminal of CCT (CONSTANS, CONSTANS-LIKE, and TIMING OF CAB EXPRESSION1) Domain (Yano et al. 2000). During SD, the loss-of-function allele of Hd1 results in suppression of flowering time and decreases Hd3a mRNA production (Izawa et al. 2002; Ishikawa et al. 2005). Phylogenetic studies revealed that Hd1 is a CO ortholog of Arabidopsis, which is a key activator of FT (Izawa et al. 2003). Hd1 involvement in Hd3a transcription is not so coherent, but the differences in spatiotemporal pattern of Hd3a and Hd1 mRNA contribute to a more complex mechanism (Hayama et al. 2003). Hd1 mRNA expression level is maximum at midnight, while Hd3a mRNA level peaks after 4–6 h of Hd1 (Izawa et al. 2002; Shikawa et al. 2005).
OsGI is an ortholog of Arabidopsis thaliana GIGANTEA (GI) and a basic upstream regulator of Hd1 expression (Hayama et al. 2002). OsGI is a big protein existing both in the nucleus and cytoplasmic matrix of cells in rice. Inhibition of OsGI expression by RNAi causes a delay in flowering, which reduces mRNA cumulating under SD conditions (Hayama et al. 2003; Kim et al. 2007). The OsGI expression is on peaks at dusk, while Hd1 peaks at midnight, referring to the involvement of somewhat unknown mechanisms, which make the differences in the time of their expression (Sawa et al. 2007).
During circadian rhythm, the OsGI expression is regulated upstream of the circadian clock (Hayama et al. 2002; Mizoguchi et al. 2005). In Arabidopsis, circadian clock mutant GI has abnormal expression (Niwa et al. 2007), and GI itself is also considered a clock-associated protein (Locke et al. 2006; Zeilinger et al. 2006). Ehd1 is another major gene that regulates the expression of Hd3a. Ehd1 is identified from the cross of the T65 cultivar and Nipponbare (Doi et al. 2004). Ehd1 encodes B-type regulation response, which reduces affinity for the target DNA sequence. Mutation suppresses the expression of Ehd1, which reduces the expression of Hd3a under SD, reflecting that Ehd1 acts as an upstream regulator of Hd3a (Kim et al. 2007). Ehd1 expression peaks before and after dawn under SD conditions, and the expression of Hd3a is like Ehd1 in the absence of Hd1 functional allele (Doi et al. 2004).
Ehd1 expression is controlled by two factors, OsMADS51 and Ghd7. OsMADS51 is the only activator of Ehd1 under SD and LD conditions, while Ghd7 suppresses Ehd1 expression only under LD conditions (Fig. 2). OsMADS51 activity is also suppressed by upregulation of OsGI because RNAi suppression of OsGI decreased Ehd1 and OsMADS51 expression (Fig. 2). OsMADS51 and OsGI exhibited similar circadian rhythms, peaking at dusk under SD conditions (Lee and An 2015).
8 Hd3a suppression under LD
In many developmental stages, the Hd3a expression is low under LD conditions as compared to SD conditions. Hd1 also plays a role in the suppression of Hd3a under LD, while promoting it under SD conditions. Prior to Hd1 cloning, it was considered that mutant of Hd3a near-isogenic lines (NILs) delay flowering time under SD conditions and promote flowering under inductive LD, indicating that Hd1 suppress the flowering time under SD condition and day-length modify the expression level of Hd1 (Lin et al. 2000).
Photochromic signaling disturbs the day-length conversation of Hd1 function because this action was not noted in phytochrome deficient plants; the flowering of rice double mutant se5 and Hd1 (which lack both phytochrome A and B and also Hd1 gene), which is slightly later than single se5 mutant (lacks only phytochromes) (Izawa et al. 2002). Furthermore, the Hd3a transcript production is low in se5 and Hd1 as compared to the se5 mutant plant. The day length also modifies the function of Hd1, which was supported by manipulation of OsGI expression resulting in alteration in the expression of Hd3 and Hd1 expression (Hayama et al. 2003). Overexpression of OsMTS1 and OsGI delay flowering under both SD and LD, indicating that OsMTS1 and OsGI act as inhibitors of flowering under both conditions. In OsGI overexpression plants, the Hd1 mRNA expression was increased under LD and SD conditions, while mRNA of Hd3a and Hd1 is negatively associated with each other, revealing that Hd1 suppresses Hd3a expression when it is extremely induced by OsGI overexpression line. OsGI overexpression also promotes Hd3a expression through upregulation of Ehd1 of OsGI-OsMADS51 pathway, but this expression is probably not observed due to strong inhibitor activity of Hd1. A diurnal temporal expression of Hd3a and Hd1 showed a high level of Hd3a suppressed by Hd1 during the light period. Under SD conditions, the Hd1 expression probably increases at the start of dust, thus maintaining Hd1 as an activator. However, OsGI overexpression accumulates a high level of Hd1 and converts into a repressor during the light period. Under LD conditions, Hd1 expression is high during the day and converts Hd1 into a suppressor (Hayama et al. 2003). Therefore, the external coincidence model can be applied to rice; a circadian clock regulates external light signals intermediated by phytochrome, and Hd1 expression generate a specific response to day length (Izawa 2007).
The suppressor behavior of Hd1 is not noted in association with Arabidopsis thaliana CO, showing that no different mechanisms that deliver this function. A photoperiodic sensitive QTL Hd2 may better explain this mechanism. Hd2 was identified as QTL from crossing indicia cultivar Kasalath and japonica cultivar Nipponbare (Yamamoto et al. 1998). Hd2 required a functional allele of Hd1 to delay flowering time under SD (Lin et al. 2000). Furthermore, Hd2 is also necessary for Hd6, a photoperiodic sensitive QTL that delays flowering time under LD (Yamamoto et al. 2000).
Ehd1 expression suppresses under LD conditions, which results in low Hd3a expression. Gh7 is also necessary for Ehd1 suppression (Fig. 2). Ghd7 contains a CCT domain, which is about 60% similar to Hd1, but Hd7 does not have a zinc finger motif like Hd1. Ghd7 does not affect Ghd1 expression but intensely suppresses Ehd1 expression. The expression of Ghd7 is higher under inductive LD conditions in leaf vascular tissues and peaks significantly at dusk (Tsuji et al. 2013).
9 Genes contribute to Hd3a/FT signaling in SAM
The recognition of Hd3a/FT as florigen discloses the window for researchers to study the molecular mechanism of florigen in SAM. In this section, we will focus on FD, CEN/TFL1, MADS-box transcription factor (FUL/CAL/AP1), and MADS-box transcription factor (SOC1). The genes of above-mentioned categories are brief in below Table 1.
FD
– In rice SAM, the interaction of Hd3a with bZIB transcription factor, which is orthologous of Arabidopsis FD results in the transformation of the vegetative phase into the reproductive phase. In Arabidopsis thaliana, FD accumulates on the edges of SAM where flower primordial produced. Its initiation starts in the vegetative stage before the arrival of FT. FD induces floral meristem identity gene FRUITFULL (FUL) and its paralog APETALA1 (AP1) in the presence of the FT gene (Wigge et al. 2005; Abe et al. 2005). FD functions in not reported in rice. There are about seven FD orthologous (OsbZIP24, OsbZIP27, OsbZIP54, OsbZIP55, OsbZIP56, OsbZIP69, and OsbZIP77) in rice whose function is not identified till now (Table 1). Among them, the OsbZIP24 mRNA level is comparatively more in SAM. OsbZIP54 mRNA distribute in the whole plant, epically SAM where the other FD-like bZIB factors is below the detection level (Nijhawan et al. 2008). Still, it is essential to study if there is any unclear functional differentiation among FD proteins.
10 MADS-box transcription factors
The genic web acting downstream of FT in Arabidopsis involved MADS-box transcription factors. FT along with FD and LEAFY is essential for the induction of FUL and AP1 in SAM. (Abe et al. 2005; Michaels et al. 2005; Teper-Bamnolker and Samach 2005). The regulation of FT-dependent gene has also been detected in Arabidopsis leaves. The expressions of FUL and SEPALATA3 were promoted by overexpression of FT in leaves and decreased in fd and ft mutants (Teper-Bamnolker and Samach 2005). Therefore, both leaves and SAM share the same regulatory mechanisms. OsMADS14/RAP1B, an orthogonal of AP1 and OsMADS15 is upregulated by Hd3a in the leaves under SD (Komiya et al. 2008). This results shows that the similar regulatory mechanism involved in SAM and OsMADS14 is expressed in apical meristem during a reproductive stage (Giavalisco et al. 2006; Furutani et al. 2006) and its ectopic behavior quickly endorses rice flowering (Jeon et al. 2000). AP1 MADS-box proteins are serving as key mediators of Hd3a/FT activity in cereal crops flowering time. OsMADS14 also acts as an activator of Hd3a in leaves (Kim et al. 2007; Lee et al. 2007). MADS-box protein suppresses the overexpression of Constants1 (SOC1) in Arabidopsis. The latest study showed that FT mediates SOC1 activation downstream of CO. SOC1 interacts directly with AGL24-MADS-box proteins in the SAM tips, and the direct attachment of AGL24-SOC1 forms a complex with the promoter of AGL24 and SOC1 and upregulates their expression level (Liu et al. 2008), whereas the regulatory mechanism and expression site is distinct in rice. Inserted T-DNA of OsMADS50 reduces the transcription of Hd3a in the leaves, while RNAi inhibition of Hd3a or Hd3a and RFT1 do not cause any modifications in the expression of OsMADS50 (Table 1). Therefore, OsMADS50 functions upstream of Hd3a in rice leaves (Lee et al. 2004; Komiya et al. 2008).
CEN/TFL1
– CENTORADIALIS (CEN)/TERMINAL FLOWER 1 (TFL1) protein provides an interesting understanding of Hd3a/FT signals in SAM. CEN/TFL1 is homologous of Hd3a/FT, which encodes RKIP proteins, however, the CEN/TFL1 protein inhibits flowering. The TFL1 mRNA of Arabidopsis is restricted to the central inflorescence shoot meristem cells, and the TFL1 protein moves to the external cells. The suppression of flowering was comprehensively studied by Arabidopsis FT and TFL1 (Hanzawa et al. 2005; Ahn et al. 2006). Proteins carrying individual scums or alternatives to specific sections of FT and TFL1 are overexpressed in double ft tfl1 mutants to draw residues or regions, that give the specific activity of both proteins. The crystal structure of TFL1 and FT were matched with specific map regions. These analyses showed that the consequent Tyr85 in His88 and FT in TFL1 are vital for the contrasting function of FT and TFL1 (Hanzawa et al. 2005), and the section liable for the difference is confined in 14 amino acid segments of the external loop of proteins at C-termini.
– There are about four CEN/TFL1-like genes in the rice genome (Chardon et al. 2005). RICE CENTRORADIALS1 (RCN1), RCN2, and RCN3 can suppress flowering initiation and affect panicle structure when overexpressed (Table 1) (Nakagawa et al. 2002; Zhang et al. 2005). The RCN3 protein can be observed in the phloem sap of rice, which suggests that the protein signals are linked between different parts (Aki et al. 2008). An attractive assumption that explains the differences between CEN/TFL1 and Hd3a/FT controlling activity is that the external loop introduces different partners to CEN/TFL1 or Hd3a/FT to define the capabilities of the complex. Screening analysis of protein interaction using a precise external loop provides a deeper understanding of Hd3a/FT signals and promotes flower promotion.
11 Conclusions and perspective
Decades of research on flowering time expand our understanding of the molecular mechanism of flowering time regulation in rice. A molecular control network is becoming complex day by day due to novel genes and their regulatory mechanisms. Florigen is also referred to as a mobile flowering hormone because of its mobility and flowering activation nature. For flowering time regulation in rice, it is necessary to explore Hd3a/FT synthesis, identification, transport, receptor activity, and cellular signaling pathway. Recent advances confirmed that still very less is known about Hd3a trafficking activity, the direction of Hd3a in the phloem, how Hd3a in companion cells is transported into the sieve element system, and how Hd3a targets SAM after unloading from the upper end of the phloem are all open questions for discussion. Flowering time research has also identified many important genes as a resources for breeding. Natural variation and artificial selection of flowering genes that are/were vanished through the domestication process should not only contribute to exploiting the mechanism of flowering time gene regulation but also facilitate future breeding programs.
Data Availability
The data that support this study cannot be publicly shared due to privacy reasons and may be shared upon reasonable request to the corresponding author if appropriate.
References
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Sci 309:1052–1056
Abe M, Fujiwara M, Kurotani K, Yokoi S, Shimamoto K (2008) Identification of dynamin as an interactor of rice GIGANTEA by tandem affinity purification (TAP). Plant Cell Physiol 49:420–432
Ahn JH, Miller D, Winter VJ, Banfield MJ, Lee JH, Yoo SY, Henz SR, Brady RL, Weigel D (2006) A divergent external loop confers antagonistic activity on floral regulators FT and TFL1. EMBO J 25:605–614
Aki T, Shigyo M, Nakano R, Yoneyama T, Yanagisawa S (2008) Nano scale proteomics revealed the presence of regulatory proteins including three FT-like proteins in Phloem and Xylem saps from rice. Plant Cell Physiol 49:767–790
Bradley D, Ratcliffe O, Vincent C, Carpenter R, Coen E (1997) Inflorescence commitment and architecture in Arabidopsis. Science 275:80–83
Brambilla V, Fornara F (2013) Molecular control of flowering in response to day length in rice. J Integr Plant Bio 5:4–12
Camoni L, Visconti S, Aducci P, Marra M (2018) 14-3-3 Proteins in plant hormone signaling: doing several things at once. Front Plant Sci 9:297
Chardon F, Damerval C (2005) Phylogenomic analysis of the PEBP gene family in cereals. J Mol Evol 61:579–590
Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A (2004) Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 18:926–936
Gao H, Xiao MZ, Guilin F, Jun C, Mingna J, Yulong R, Weixun W, Zhou K, Sheng P, Zhou F, Jiang L, Wang J, Zhang X, Guo X, Wang J, Cheng Z, Wu C, Wang H, Wan JM (2013) Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice. PLoS Genet 9:1–12
Giavalisco P, Kapitza K, Kolasa A, Buhtz A, Kehr J (2006) Towards the proteome of Brassica napus phloem sap. Proteom 6:896–909
Goretti D, Martignago D, Landini M, Brambilla V, Gomez AJ, Gnesutta N, Galbiati F, Collani S, Takagi H, Terauchi R, Mantovani R, Fornara F (2017) Transcriptional and post-transcriptional mechanisms limit Heading Date 1 (Hd1) function to adapt rice to high latitudes. PLoS Genet 13:1006530
Gu Q, Ferrandiz C, Yanofsky MF, Martienssen R (1998) The FRUITFULL MADS-box gene mediates cell differentiation during Arabidopsis fruit development. Dev 125:1509–1517
Guo T, Mu Q, Wang J, Vanous AE, Onogi A, Iwata H, Yu J (2020) Dynamic effects of interacting genes underlying rice flowering-time phenotypic plasticity and global adaptation. Genome Res 30:673–683
Hayama R, Izawa T, Shimamoto K (2002) Isolation of rice genes possibly involved in the photoperiodic control of flowering by a fluorescent differential display method. Plant Cell Physiol 43:494–504
Hayama R, Yokoi S, Tamaki S, Yano M, Shimamoto K (2003) Adaptation of photoperiodic control pathways produces short day flowering in rice. Nature 422:719–722
Hori K, Matsubara K, Yano M (2016) Genetic control of flowering time in rice: integration of Mendelian genetics and genomics. Theor Appl Genet 129:2241–2252
Huan Z, Shanshan Z, Tianzhen L, Chunming W, Zhijun C, Xin Z, Liping C, Peike S, Maohong C, Chaonan L, Jiachang W, Zhe Z, Juntao C, Liang Z, Cailin L, Xiuping GZ, Jiulin W, Jie W, Ling J, Chuanyin W, Jianmin W (2018) Delayed heading date1 interacts with OSHAP5C/D, delays flowering time and enhances yield in rice. Plant Biotec J 17:531–539
Ishikawa R, Tamaki S, Yokoi S, Inagaki N, Shinomura T, Takano M, Shimamoto K (2005) Suppression of the floral activator Hd3a is the principal cause of the night break effect in rice. Plant Cell 17:3326–3336
Izawa T (2007) Day length measurements by rice plants in photoperiodic short-day flowering. Int Rev Cytol 256:191–222
Izawa T, Oikawa T, Sugiyama N, Tanisaka T, Yano M, Shimamoto K (2002) Phytochrome mediates the external light signal to repress FT orthologs in photoperiodic flowering of rice. Genes Dev 16:2006–2020
Izawa T, Takahashi Y, Yano M (2003) Comparative biology comes into bloom: genomic and genetic comparison of flowering pathways in rice and Arabidopsis. Curr Opin Plant Biol 6:113–120
Jaeger KE, Wigge PA (2007) FT protein acts as a long-range signal in Arabidopsis. Cur Biol 17:1050–1054
Jang S, Marchal V, Panigrahi KCS, Wenkel S, Soppe W, Deng XW, Valverde F, Coupland G (2008) Arabidopsis COP1 shapes the temporal pattern of CO accumulation conferring a photoperiodic flowering response. EMBO J 27:1277–1288
Kim SL, Lee S, Kim HJ, Nam HG, An G (2007) OsMADS51 is a short day flowering promoter that functions upstream of Ehd1, OsMADS14, and Hd3a. Plant Physiol 145:1484–1494
Kobayashi Y, Weigel D (2007) Move on up, it’s time for change-mobile signals controlling photoperiod-dependent flowering. Genes Dev 21:2371–2384
Kobayashi Y, Kaya H, Goto K, Iwabuchi M, Araki T (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1972
Kojima S, Takahashi Y, Kobayashi Y, Monna L, Sasaki T, Araki T, Yano M (2002) Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short day conditions. Plant Cell Physiol 243:1096–1105
Komiya R, Yokoi S, Shimamoto K (2009) A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Dev 136:3443–3450
Kyozuka J, Kobayashi T, Morita M, Shimamoto K (2000) Spatially and temporally regulated expression of rice MADS box genes with similarity to Arabidopsis class A, B and C genes. Plant Cell Physiol 41:710–718
Lee, An (2015) Regulation of flowering time in rice. J Plant Biol 58:353–360
Lee SY, Kim J, Han JJ, Han MJ, An GH (2004) Functional analyses of the flowering time gene OsMADS50, the putative SUPPRESSOR OF OVEREXPRESSION OF CO 1/AGAMOUS-LIKE 20 (SOC1/AGL20) ortholog in rice. Plant J 38:754–764
Lee YS, Dong HJ, Dong YL, Jakyung Y, Gynheung A (2010) OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB. Plant J 63:18–30
Lin HX, Yamamoto T, Sasaki T, Yano M (2000) Characterization and detection of epistatic interactions of 3 QTLs, Hd1, Hd2, and Hd3, controlling heading date in rice using nearly isogenic lines. Theor Appl Genet 101:1021–1028
Lin MK, Belanger H, Lee YJ, Varkonyi GE, Taoka KI, Miura E, Xoconostle CB, Gendler K, Jorgensene RA, Phinney B, Lough TJ, Lucas WJ (2007) FLOWERING LOCUS T protein may act as the long-distance florigenic signal in the cucurbits. Plant Cell 19:1488–1506
Liu H, Gu F, Dong S, Liu W, Wang H, Chen Z, Wang J (2016) Constans-like 9 (COL9) delays the flowering time in Oryza sativa by repressing the ehd1 pathway. Biochem Biophys Res Commun 479:173–178
Liu J, Gong J, Wei X, Yang S, Huang X, Li C, Zhou X (2020) Dominance complementation of Hd1 and Ghd8 contributes to extremely late flowering in two rice hybrids. Mol Breed 40:76
Locke JCW, Kozma BL, Gould PD, Feher B, Kevei E, Nagy F, Turner MS, Hall A, Millar AJ (2006) Experimental validation of a predicted feedback loop in the multi-oscillator clock of Arabidopsis thaliana. Mol Syst Biol 2:59
Mandel MA, Gustafson BC, Savidge B, Yanofsky MF (1992) Molecular characterization of the Arabidopsis floral homeotic gene APETALA1. Nature 360:273–277
Mathieu J, Warthmann N, Küttner F, Schmid M (2007) Export of FT protein from phloem companion cells is sufficient for floral induction in Arabidopsis. Cur Biol 17:1055–1060
Matsubara K, Yamanouchi U, Wang ZX, Minobe Y, Izawa T, Yano M (2008) Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1. Plant Physiol 148:1425–1435
Matsubara K, Yamanouchi U, Nonoue Y, Sugimoto K, Yano M (2011) Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J 66:603–612
Michaels SD, Himelblau E, Kim SY, Schomburg FM, Amasino RM (2005) Integration of flowering signals in winter-annual Arabidopsis. Plant Physiol 137:149–156
Mizoguchi T, Wright L, Fujiwara S, Cremer F, Lee K, Onouchi H, Mouradov A, Fowler S, Kamada H, Putterill J, Coupland G (2005) Distinct roles of GIGANTEA in promoting flowering and regulating circadian rhythms in Arabidopsis. Plant Cell 17:2255–2270
Molla AK (2022) Flowering time and photoperiod sensitivity in rice: Key players and their interactions identified. Plant Cell 34:3489–3490
Monna L, Lin HX, Kojima S, Sasaki T, Yano M (2002) Genetic dissection of a genomic region for a quantitative trait locus, Hd3, into two loci, Hd3a and Hd3b, controlling heading date in rice. Theor Appl Genet 104:772–780
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750
Nemoto Y, Nonoue Y, Yano M, Izawa T (2016) Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. Plant J 86:221–233
Nijhawan A, Jain M, Tyagi AK, Khurana JP (2008) Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol 146:333–350
Niwa Y, Ito S, Nakamichi N, Mizoguchi T, Niinuma K, Yamashino T, Mizuno T (2007) Genetic linkages of the circadian clock-associated genes, TOC1, CCA1 and LHY, in the photoperiodic control of flowering time in Arabidopsis thaliana. Plant Cell Physiol 48:925–937
Park SJ, Song LK, Shinyoung L, Byoung IJ, Hai LP, Sung HP, Chu MK, Choong HR, Su HP, Yuan HX, Joseph C, Gynheung A, Chang DH (2008) Rice Indeterminate 1 (OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod. Plant J 56:1018–1029
Peng LT, Shi ZY, Li L, Shen GZ, Zhang JL (2008) Overexpression of transcription factor OsLFL1 delays flowering time in Oryza sativa. J Plant Physiol 165:876–885
Peng Q, Zhu C, Liu T, Zhan S, Feng S, Wu C (2021) Phosphorylation of OsFD1 by OsCIPK3 promotes the formation of RFT1-containing florigen activation complex for long-day flowering in rice. Mol Plant 14:1135–1148
Ryu CH, Lee S, Cho LH, Kim SL, Lee YS, Choi SC, Jeong HJ, Yi J, Park SJ, Han CD, An G (2009) OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plant Cell Environ 32:1412–1427
Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz SZ, Yanofsky MF, Coupland G (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288:1613–1616
Sawa M, Nusinow DA, Kay SA, Imaizumi T (2007) FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Sci 318:261–265
Sheng P, Wu F, Tan J, Zhang H, Ma W, Chen L, Wang J, Wang J, Zhu S, Guo X, Wang J, Zhang X, Cheng Z, Bao Y, Wu C, Liu X, Wan J (2016) A CONSTANS-liketranscriptional activator, OsCOL13, functions as a negative regulator of flowering downstream of OsphyB and upstream of Ehd1 in rice. Plant Mol Biol 92:209–222
Shikawa R, Tamaki S, Yokoi S, Inagaki N, Shinomura T, Takano M, Shimamoto K (2005) Suppression of the floral activator Hd3a is the principal cause of the night break effect in rice. Plant Cell 17:3326–3336
Shim JS, Jang G (2020) Environmental signal-dependent regulation of flowering time in Rice. Int J Mol Sci 21:6155
Shrestha R, Gomez AJ, Brambilla V, Fornara F (2014) Molecular control of seasonal flowering in rice, Arabidopsis and temperate cereals. Ann Bot 114:1445–1458
Sohail A, Shah L, Liu L, Islam A, Yang Z, Yang Q, Anis GB, Xu P, Khan RM, Li J et al (2022) Mapping and validation of qHD7b: major heading-date QTL functions mainly under long-day conditions. Plants 11:2288
Song YH, Shim JS, Kinmonth SHA, Imaizumi T (2015) Photoperiodic flowering: time measurement mechanisms in leaves. Annu Rev Plant Biol 66:441–464
Song S, Wang G, Hu Y, Liu H, Bai X, Qin R (2018) OsMFT1 is regulated by Ghd7 and OsLFL1 to increase spikelets per panicle and delay heading date in rice. J Exp Bot 69:4283–4293
Sun C, Chen D, Fang J, Wang P, Deng X, Chu C (2014) Understanding the genetic and epigenetic architecture in complex network of rice flowering pathways. Protein Cell 5:889–898
Tamaki S, Matsuo S, Wong HL, Yokoi S, Shimamoto K (2007) Hd3a protein is a mobile flowering signal in rice. Science 316:1033–1036
Tan J, Jin M, Wang J, Wu F, Sheng P, Cheng Z, Wang J, Zheng X, Chen L, Wang M, Zhu S, Guo X, Zhang X, Liu X, Wang C, Wang H, Wu C, Wan J (2016) OsCOL10, a CONSTANS-like gene, functions as a flowering time repressor downstream of Ghd7 in rice. Plant Cell Physiol 57:798–812
Taoka KI, Izuru O, Tsuji H, Furuita K, Hayashi K, Yanase T, Yamaguchi M, Nakashima C, Purwestri YA, Tamaki S (2011) 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476:332–335
Teper-Bamnolker P, Samach A (2005) The flowering integrator FT regulates SEPALLATA3 and FRUITFULL accumulation in Arabidopsis leaves. Plant Cell 17:2661–2675
Tsuji H (2017) Molecular function of florigen. Breed Sci 67:327–332
Tsuji H, Taoka K, Shimamoto K (2011) Regulation of flowering in rice: two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol 14:45–52
Tsuji H, Taoka K, Shimamoto K (2013) Florigen in rice: complex gene network for florigen transcription, florigen activation complex, and multiple functions. Curr Opin Plant Biol 16:228–235
Wu W, Zheng XM, Chen D, Zhang Y, Ma W, Zhang H, Sun L, Yang Z, Zhao C, Zhan X, Shen X, Yu P, Fu Y, Zhu S, Cao L, Chenga S (2017) OsCOL16, encoding a CONSTANS-like protein, represses flowering by upregulating Ghd7 expression in rice. Plant Sci 260:60–69
Wu W, Zhang Y, Zhang M, Zhan X, Cao L (2018) The rice constans-like protein OsCOlL5 suppresses flowering by promoting Ghd7 and repressing RID1. Biochem Biophys Res Commun 495:1349
Wu CC, Wei FJ, Chiou WY, Tsai YC, Wu HP, Gotarkar D, Hsing YIC (2020) Studies of rice Hd1 haplotypes worldwide reveal adaptation of flowering time to different environments. PLoS ONE 15:e0239028
Yamaguchi A, Kobayashi Y, Goto K, Abe M, Araki T (2005) TWIN SISTER OF FT (TSF) acts as a floral pathway integrator redundantly with FT. Plant Cell Physiol 46:1175–1189
Yamamoto T, Kuboki Y, Lin SY, Sasaki T, Yano M (1998) Fine mapping of quantitative trait loci Hd-1, Hd-2 and Hd-3, controlling heading date of rice, as single Mendelian factors. Theor Appl Genet 97:37–44
Yamamoto T, Lin HX, Sasaki T, Yano M (2000) Identification of heading date quantitative trait locus Hd6 and characterization of its epistatic interactions with Hd2 in rice using advanced backcross progeny. Genetics 154:885–891
Yan WH, Wang P, Chen HX, Zhou HJ, Li QP, Wang CR, Ding ZH, Zhang YS, Yu SB, Xing YZ, Zhang QF (2011) A Major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant 4:319–330
Yang Y, Peng Q, Chen GX, Li XH, Wu CY (2013) OsELF3 is involved in circadian clock regulation for promoting flowering under long-day conditions in Rice. Mol Plant 1:202–215
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2483
Zeevaart JAD (2006) Florigen coming of age after 70 years. Plant Cell 18:1783–1789
Zeilinger MN, Farre EM, Taylor SR, Kay SA, Doyle FJ (2006) A novel computational model of the circadian clock in Arabidopsis that incorporates PRR7 and PRR9. Mol Syst Biol 2:58
Zhang SH, Hu WJ, Wang LP, Lin CF, Cong B, Sun CR, Luo D (2005) TFL1/CEN-like genes control intercalary meristem activity and phase transition in rice. Plant Sci 168:1393–1408
Zhang Z, Hu W, Shen G, Liu H, Hu Y, Zhou X, Liu T (2017) Alternative functions of Hd1 in repressing or promoting heading are determined by Ghd7 status under long-day conditions. Sci Rep 7:5388
Zhang H, Zhu S, Liu T, Wang C, Cheng Z, Zhang X, Chen L, Sheng P, Cai M, Li C, Wang J, Zhang Z, Chai J, Zhou L, Lei C, Guo X, Wang J, Wang J, Jiang L, Wu C, Wan J (2019) DELAYED HEADING DATE1 interacts with OsHAP5C/D, delays flowering time and enhances yield in rice. Plant Biot J 17:531–539
Zhu S, Wang J, Cai M, Zhang H, Wu F, Xu Y, Li C, Cheng Z, Zhang X, Guo X, Sheng P, Wu M, Wang J, Lei C, Wang J, Zhao Z, Wu C, Wang H, Wan J (2017) The OsHAPL1-DTH8-Hd1 complex functions as the transcription regulator to repress heading date in rice. J Exp Bot 68:553–568
Zong W, Ren D, Huang M, Sun K, Feng J, Zhao J, Guo J (2020) Strong photoperiod sensitivity is controlled by cooperation and competition among Hd1, Ghd7 and DTH8 in rice heading. New Phytol 229:1635–1649
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sohail, A. Genetic and signaling pathways of flowering regulation in rice (Oryza sativa L.). Braz. J. Bot 46, 599–608 (2023). https://doi.org/10.1007/s40415-023-00910-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-023-00910-y