Abstract
Aim
The objective of this study was to develop a physiologically based pharmacokinetic (PBPK) model for sinogliatin (HMS-5552, dorzagliatin) by integrating allometric scaling (AS), in vitro to in vivo exploration (IVIVE), and steady-state concentration–mean residence time (Css-MRT) methods and to provide mechanistic insight into its pharmacokinetic properties in humans.
Methods
Human major pharmacokinetic parameters were analyzed using AS, IVIVE, and Css-MRT methods with available preclinical in vitro and in vivo data to understand sinogliatin drug metabolism and pharmacokinetic (DMPK) characteristics and underlying mechanisms. On this basis, an initial mechanistic PBPK model of sinogliatin was developed. The initial PBPK model was verified using observed data from a single ascending dose (SAD) study and further optimized with various strategies. The final model was validated by simulating sinogliatin pharmacokinetics under a fed condition. The validated model was applied to support a clinical drug–drug interaction (DDI) study design and to evaluate the effects of intrinsic (hepatic cirrhosis, genetic) factors on drug exposure.
Results
The two-species scaling method using rat and dog data (TS-rat,dog) was the best AS method in predicting human systemic clearance in the central compartment (CL). The IVIVE method confirmed that sinogliatin was predominantly metabolized by cytochrome P450 (CYP) 3A4. The Css-MRT method suggested dog pharmacokinetic profiles were more similar to human pharmacokinetic profiles. The estimated CL using the AS and IVIVE approaches was within 1.5-fold of that observed. The Css-MRT method in dogs also provided acceptable prediction of human pharmacokinetic characteristics. For the PBPK approach, the 90% confidence intervals (CIs) of the simulated maximum concentration (Cmax), CL, and area under the plasma concentration–time curve (AUC) of sinogliatin were within those observed and the 90% CI of simulated time to Cmax (tmax) was closed to that observed for a dose range of 5–50 mg in the SAD study. The final PBPK model was validated by simulating sinogliatin pharmacokinetics with food. The 90% CIs of the simulated Cmax, CL, and AUC values for sinogliatin were within those observed and the 90% CI of the simulated tmax was partially within that observed for the dose range of 25–200 mg in the multiple ascending dose (MAD) study. This PBPK model selected a final clinical DDI study design with itraconazole from four potential designs and also evaluated the effects of intrinsic (hepatic cirrhosis, genetic) factors on drug exposure.
Conclusions
Sinogliatin pharmacokinetic properties were mechanistically understood by integrating all four methods and a mechanistic PBPK model was successfully developed and validated using clinical data. This PBPK model was applied to support the development of sinogliatin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. We provide an effective physiologically based pharmacokinetic (PBPK) modeling strategy based on mechanistic insight into human drug metabolism and pharmacokinetic properties from preclinical in vitro and in vivo data using allometric scaling (AS), in vitro to in vivo exploration (IVIVE), and steady-state concentration–mean residence time (Css-MRT) methods. |
2. The PBPK model of sinogliatin provides a useful tool to predict human pharmacokinetics and to evaluate the effects of extrinsic (e.g., drug–drug interactions) and intrinsic (e.g., hepatic cirrhosis, genetic) factors on drug exposure. |
3. We provide a methodology for learning and confirming preclinical and clinical data and for obtaining insightful pharmacokinetic understanding by integrating four methods for first-in-human research. |
1 Introduction
Diabetes mellitus (DM) is a metabolic disorder syndrome that represents an increasing threat to public health [1]. Type 2 DM (T2DM) accounts for more than 90% of the diabetic patient population, typically characterized by a combination of insulin resistance and β cell dysfunction. Although several classes of therapies for T2DM are available for clinical use, the need for novel therapies still remains to be met in order to improve the effectiveness of glycemic control; so far, current therapies can control glucose levels well in only 60% of T2DM patients [2]. Accumulating results from recent studies, including reports from several clinical studies, have demonstrated that small-molecule glucokinase activators (GKAs) may be able to meet such need [3].
Glucokinase plays a key role in glucose homeostasis as it initiates the first step. Glucokinase has also been suggested to act as a ‘glucose sensor’ in a glucose-dependent way that facilitates glucose homeostasis in the pancreas, liver, ventromedial hypothalamus, and gastrointestinal tract [4]. In pancreatic β cells, glucokinase was the central rate-limited enzyme controlling the threshold for glucose-stimulated insulin release (GSIR). In hepatic cells, glucokinase was the key enzyme for glucose uptake and hepatic glycogen production [5]. Therefore, GKAs represented a promising opportunity for T2DM treatment.
So far, no GKAs have been marketed anywhere in the world, although several GKAs are under preclinical or clinical development. Sinogliatin (HMS-5552, dorzagliatin) is a first-in-class novel GKA discovered a decade after the first GKA was published in 2003 [6]. Phase I and II clinical studies were completed by 2016. The results showed that sinogliatin can effectively control both fasting and postprandial glucose in healthy volunteers as well as T2DM patients with normal hepatic and renal function and was well-tolerated [7]. Meanwhile, additional clinical pharmacology studies followed by phase III studies in a broader T2DM patient population will be carried out to further evaluate its pharmacokinetic and PD profiles and to guide its use in the clinic in the future. In order to guide the study design and dose selection of clinical pharmacology studies and to fully understand the potential pharmacokinetic and safety profiles of sinogliatin in the broader population during its late-stage development, a mechanistic physiologically based pharmacokinetic (PBPK) model is required to accurately predict sinogliatin pharmacokinetics in different populations under various conditions.
Therefore, a mechanistic PBPK model was developed to fulfill these requirements mentioned. First, we summarized the sinogliatin in vitro ADME (absorption, distribution, metabolism, and excretion) data and pharmacokinetic profiles in rats, dogs, and monkeys (as shown in Table 1). Allometric scaling (AS), in vitro to in vivo exploration (IVIVE), steady-state concentration (Css)–mean residence time (MRT) (Css-MRT) methods were used to fully understand its drug metabolism and pharmacokinetic (DMPK) characteristics and underlying mechanisms in humans. Furthermore, these three methods were utilized to ensure the data quality for PBPK model development. On top of this, a mechanistic PBPK model was developed, optimized with human pharmacokinetic data from a single ascending dose (SAD) study in healthy subjects, and validated with sinogliatin pharmacokinetic data under fed conditions from a multiple ascending dose (MAD) study in T2DM patients.
2 Methods
2.1 Overall Strategy
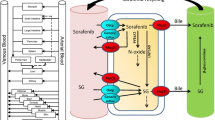
The overall strategy of the mechanistic understanding process and PBPK model construction is illustrated in Fig. 1. AS, IVIVE, and Css-MRT methods provided an understanding of sinogliatin DMPK characteristics based on preclinical in vitro and in vivo data. A PBPK model was developed using SimCYP® software (version 16.0; Certara, Sheffield, UK) by integrating the substrate-specific and system-specific parameters. The model was optimized with various methods based on human pharmacokinetic data from the SAD study in healthy subjects, and was validated by simulating human pharmacokinetic data under fed conditions from the MAD study in T2DM patients. Finally, the validated PBPK model provided verification of input data and understandings and evaluated the effect of extrinsic (drug–drug interactions [DDIs]) and intrinsic (hepatic cirrhosis, genetic) factors on drug exposure.
Overall strategy of mechanistic understanding process and physiologically based pharmacokinetic model development. *The Css-MRT approach has an insignificant contribution to sinogliatin PBPK development. AS allometric scaling, CL systemic clearance in central compartment, CLiv clearance after intravenous administration, Css-MRT steady-state concentration–mean residence time, DDI drug–drug interaction, fm metabolism fraction, IVIVE in vitro to in vivo exploration, MAD multiple ascending dose, PBPK physiologically based pharmacokinetic, PK pharmacokinetic, SAD single ascending dose, Vss steady-state distribution volume
2.2 Preclinical and Clinical Data Sources
Preclinical data used in this study were from the sinogliatin preclinical pharmacokinetics report no. 27; clinical data used in this study were from reports HMS0101-No.32, CPRCL-192-HMS5552/ PKR-PI [7].
2.2.1 Preclinical In Vitro Data
For absorption, dissolution profiles were collected from phosphate buffer of pH 1.2, 4.5, and 6.8; apparent permeability (Papp) was measured in the MDCK (Madin-Darby canine kidney)-2 cell line system. For distribution, the blood-to-plasma ratio (B/P) and unbound fraction in plasma (fu,p) in rat, dog, and humans were determined. For metabolism, the metabolic stability of sinogliatin in both the microsomal system (human and animal microsomes were purchased from XenoTech, LLC, Lenexa, KS, USA) and human recombinant cytochrome P450 (CYP) 3A4 (rhCYP3A4) enzyme system (BD Biosciences, Woburn, MA, USA) were determined. The values obtained from these in vitro studies are summarized in Table 2. For transporters, the efflux effect of P-glycoprotein (P-gp) on sinogliatin and the inhibition effects of sinogliatin on organic anion transporting polypeptide (OATP) 1B1and OATP1B3 were determined; the results are summarized in Electronic Supplementary Material Table 1.
2.2.2 Preclinical In Vivo Data
Pharmacokinetic parameters of sinogliatin in rat, dog, and monkey after single intravenous and oral dose administration are shown in Table 1. The plasma/tissue partition coefficient (Kp) of various tissues were determined in rat at 2, 6, 12, and 36 h post a single oral dose. The sinogliatin mass balance was assessed in rat after single intravenous and oral dose administration of 14C-sinogliatin; plasma, bile, urine, and fecal samples were collected to determine the recovery of total radioactivity and the elimination pathway.
2.2.3 Clinical Pharmacokinetic Study
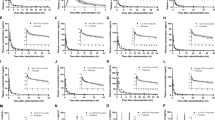
In the SAD study, 60 healthy subjects were randomly assigned to one of six sinogliatin dose cohorts (5, 10, 15, 25, 35, and 50 mg) or placebo with a sinogliatin:placebo ratio of 4:1. Sinogliatin was administered under a fasted state and plasma samples were collected up to 72 h post-dose. The primary pharmacokinetic parameters and the time–concentration profiles are shown in Table 3 and Fig. 2a, respectively.
Simulated (lines) and observed (points) mean plasma concentration–time curves of various dosages in a fasted condition using Model C and b fed condition using Model C. Lower limit of quantification of observed data was 1 μg/L. Conc. concentration, Obs clinical observed data, Sim physiologically based pharmacokinetic model simulated results
In the MAD study, 53 Chinese T2DM patients with fasting plasma glucose in the range of 7.0–11.1 mmol/L and with normal liver and kidney function were randomly assigned to one of five sinogliatin dose cohorts (25, 50, 100, 150, and 200 mg) or placebo with a sinogliatin:placebo ratio of 4:1. The effect of a standard T2DM diet on sinogliatin pharmacokinetics was measured after both single-dose administration and under steady state. Plasma samples were collected up to 72 h post-dose. The primary pharmacokinetic parameters and the time–concentration profiles are shown in Table 3 and Fig. 2b, respectively.
2.3 Understanding Preclinical Data Using Allometric Scaling (AS), In Vitro to In Vivo Exploration (IVIVE), and Steady-State Concentration–Mean Residence Time (C ss-MRT) Methods
In this study, AS, IVIVE, and Css-MRT methods were utilized to estimate human pharmacokinetics based on preclinical in vitro and in vivo data. The purpose of this was to aid understanding of the DMPK properties and underlying mechanisms of sinogliatin, and also for ensuring the quality of the data that will be used in later PBPK model development.
2.3.1 AS Calculation of Sinogliatin Human Systemic Clearance in the Central Compartment (CL) and Steady-State Distribution Volume
The intravenous systemic clearance in the central compartment (CL) (CLiv), body weight (BW) and fu,p of rat and dog were used to predict human CL using various AS methods. Equations for the AS methods are summarized in Table 4 and Electronic Supplementary Material Table 2. The Øie–Tozer method was selected to estimate the sinogliatin steady-state distribution volume (Vss) in humans [8] and the equations for this method are listed in Table 4.
2.3.2 IVIVE Prediction of Sinogliatin Human CL
An in vitro metabolic experiment showed that sinogliatin was predominantly metabolized by CYP3A4 and ketoconazole can 100% inhibit sinogliatin metabolism in human microsomes. Therefore, IVIVE extrapolation of human hepatic CL (CLh) from intrinsic CL (CLint) in the rhCYP3A4 enzyme system (CLint,rhCYP3A4) and microsomal system (CLint,microsome) was performed. Hepatic intrinsic CLint (CLint,hepatic) was scaled from rhCYP3A4 enzyme system using Eq. 1:
where MPPGL is mg protein per liver weight.
The parameters and source data used for extrapolation are summarized in Table 2. CLint,hepatic was extrapolated from microsomal experiments of rat, dog, and humans using Eq. 2:
The parameters used and the source or calculating equations are summarized in Electronic Supplementary Material Table 3.
The well-stirred model [9] was applied to calculate CLh (Eq. 3):
where Qliver is the blood flow of the liver.
The parameters and source data are summarized in Table 2 and Electronic Supplementary Material Tables 3 and 4. For both the rhCYP3A4 and microsomal systems, human total CLiv was obtained by adding observed human renal CL.
2.3.3 Estimation of Oral Plasma Concentration–Time Profiles Using the C ss-MRT Method
The Css-MRT method [10] was used to estimate human oral plasma pharmacokinetic profiles by normalizing in vivo pharmacokinetic profiles of rat, dog, and monkey. Superimposition of the pharmacokinetic profile of each species was obtained by dividing the plasma concentration by the Css (y-axis) and time divided by its corresponding MRT (x-axis). The normalized curves of various species were plotted together to fit the best pharmacokinetic model using ADAPT 5 package [31]. The human Css and MRT was multiplied by the y-axis and x-axis of the best-fitted normalized curve, we obtained human intravenous pharmacokinetic profiles and estimated human intravenous pharmacokinetic parameters. We then developed the oral pharmacokinetic model of animals to estimate the absorption parameters (absorption rate constant [ka] and absorbed fraction [Fa]). Human absorption parameters were assumed as average fitted values of animal absorption parameters. Finally, the full set of estimated human oral pharmacokinetic parameters was used to predict human oral pharmacokinetic profiles.
2.4 Sinogliatin Human Physiologically Based Pharmacokinetic (PBPK) Model Development
2.4.1 Model Development
The SimCYP® population-based simulator was used to develop the sinogliatin PBPK model. The drug-specific parameters are summarized in Table 2. Systemic parameters were kept as the default setting in the SimCYP® package. In the absorption part, we selected the advanced dissolution, absorption, and metabolism (ADAM) model, which consisted of nine compartments to mimic gastrointestinal tract physiology [11]. The immediate-release dissolution profiles obtained from the in vitro dissolution assay was input to simulate the dissolution kinetics. The human jejunum effective permeability (Peff), value was calculated from the in vitro MDCK experimental Papp. Propranolol and cimetidine were used as reference drugs to evaluate the system scalar. A full PBPK model was applied to the distribution part. Briefly, the full PBPK model defined mathematical compartments for different tissues linked by the circulatory system [12]. The volume of distribution was predicted using the method of Rodgers and Rowland [13]. In the elimination part, three different models were used to predict human clearance: (i) in vivo CLiv calculated from the two-species scaling method using rat and dog data (TS-rat,dog) (Model A); (ii) in vitro CYP3A4-mediated CLint values (Model B); and (iii) in vitro CYP3A4-mediated maximum rate of the metabolite formation (Vmax) and Michaelis-Menten constant (Km) (Model C). Observed human renal clearance were added to all three models to obtain the total clearance.
2.4.2 PBPK Model Simulation
Simulation was based on the SimCYP® virtual population database of ‘Chinese healthy volunteers’ (n = 594) provided by Pfizer Ltd (Sandwich, UK). The pharmacokinetics for sinogliatin 15 mg were simulated using three models separately, and the sinogliatin pharmacokinetics for six dose groups (5, 10, 15, 25, 35, 50 mg) were simulated using the final mechanistic model. The simulated pharmacokinetic parameters and profiles were compared with data observed in a clinical study to assess the acceptance of the model estimation.
2.4.3 Model Evaluation Criteria and Optimization Methods
The 90% confidence interval (CI) of the observation and simulation was used as the criterion to determine the simulation acceptance and accuracy of the PBPK model. Key pharmacokinetic parameters for assessing the goodness of the PBPK model are maximum concentration (Cmax), time to Cmax (tmax), CL, and area under the plasma concentration–time curve (AUC), as defined in the European Medicines Agency (EMA) guideline published on 21 July 2016 [14]. The model was refined and optimized based on the acceptance of these parameters.
For model optimization, the following steps may be taken:
-
(i)
Sensitivity analysis: evaluate the impact of the input parameters of interest on the model simulation within a ten-fold range around the experimental values.
-
(ii)
Parameter estimation: the most sensitive parameter will then be estimated by fitting the observed data.
-
(iii)
Experimental confirmation: if the estimated value in step 2 is significantly different from the experimental value, a supplementary experiment may be considered, especially when the original experimental design is problematic.
2.5 Model Validation
The food effect trial simulation was performed using the final mechanistic model based on the actual trial design. The only exception was that the actual clinical food effect study was carried out in patients with a relatively healthy status. This simulation was based on the hypothesis that there was no pharmacokinetic difference between healthy volunteers and T2DM patients since (i) some other clinical studies of sinogliatin have shown that there was no significant pharmacokinetic difference in the two populations in the dosage range of 25–50 mg; and (ii) the liver and renal function of all T2DM patients who participated in the clinical study were normal. The geometric means values and 90% CIs of key pharmacokinetic parameters from ten simulated trials were compared with the observed clinical data to validate the final PBPK model.
2.6 Model Application
2.6.1 Clinical Drug–Drug Interaction (DDI) Study Design
Four potential clinical DDI study designs were simulated with the inhibitor itraconazole: (A) multiple administration of itraconazole for 14 days, single administration of sinogliatin on the eighth day (D8); (B) multiple administration of itraconazole for 8 days, single administration of sinogliatin on D8; (C) multiple administration of itraconazole for 14 days, single administration of sinogliatin on the fourth day (D4); and (D) multiple administration of itraconazole for 8 days, single administration of sinogliatin on D4. For each design, the ratio of AUC (AUCR) with and without inhibitor (AUCR = AUCwith inhibitor/AUCwithout inhibitor) and the ratio of Cmax (CmaxR) with and without inhibitor (CmaxR = Cmax,with inhibitor/Cmax,without inhibitor) were compared to select the optimal clinical DDI trial design.
2.6.2 Drug Exposure Evaluation in Hepatic Cirrhosis Patients
Pharmacokinetic simulation in hepatic cirrhosis patients (Sim-Cirrhosis CP) was performed in three populations: Sim-Cirrhosis CP-A (mild), Sim-Cirrhosis CP-B (moderate), and Sim-Cirrhosis CP-C (serve). The simulated AUC and Cmax values of each population were compared with those of the healthy population to support clinical dose adjustment.
2.6.3 Impact of Cytochrome P450 (CYP) 3A4 Alleles on Hepatic Metabolism
The impact of CYP3A4 alleles on metabolism was evaluated for three CYP3A4 alleles: wild-type (CYP3A4*1) and variant-type (CYP3A4*16 and CYP3A4*18). Different CYP abundances [15] were input in the default Chinese healthy population, as shown in Electronic Supplementary Material Table 5. Simulated CL and AUC values were used to evaluate the inter-individual variability of impact of CYP3A4 alleles on hepatic metabolism.
3 Results
3.1 Understanding Drug Metabolism and Pharmacokinetic Characteristics Using the AS, IVIVE, and Css-MRT Method
The TS-rat,dog method was selected as the best AS method based on a unified strategy [16]. The estimated human CLiv and CL/F values were 10.4 and 11.7 L/h, respectively. The estimated CLiv values of other AS methods are listed in Electronic Supplementary Material Table 2. The Vss was estimated to be 108 L using the Øie–Tozer method, as shown in Table 4. The estimated CL/F and Vss values were 0.98- and 0.99-fold of the observed data, respectively.
Human CL and CL/F values were estimated to be 11.6 and 13.1 L/h, respectively, using the IVIVE method based on enzyme kinetic parameters of rhCYP3A4 (see details listed in Table 2) and the estimated CL/F was 1.10-fold of the observed data. The estimation results using microsomal data were not acceptable since the scaling factor between observed and predicted data (Fsc) of rat and dog were 37 and 148, far exceeding the default accepted criterion of 5. Details are listed in Electronic Supplementary Material Tables 3 and 4.
Css-MRT-normalized pharmacokinetic profiles of rat, dog, and monkey are plotted together in Fig. 3a. The normalized pharmacokinetic profiles of rat and monkey were in good superposition, but were different from that of dog, indicating the interspecies metabolism processes are different. Therefore, human oral pharmacokinetic parameters were estimated using a normalized rat–monkey pharmacokinetic profile and dog pharmacokinetic profile separately, and the results are summarized in Table 5. Simulated human oral pharmacokinetic profiles using dog data were within two-fold of observed profiles, providing acceptable estimation as shown in Fig. 3b. Human profiles using rat–monkey data provided relative poor estimation, as shown in Fig. 3c. The estimation difference depended on preclinical interspecies difference. An in vitro metabolism experiment demonstrated that the metabolic product types of humans are the same as those of dog, but different from those of rat or monkey.
a Normalized concentration–time curves by Css and MRT for rat, dog, and monkey at the lowest intravenous dose level. Simulated (lines) and observed (points) mean plasma concentration-time curves using data for b dog and c rat and monkey. Lower limit of quantification of observed data was 1 μg/L. Conc. concentration, Css steady-state concentration, MRT mean residence time, Obs clinical observed data, Sim physiologically based pharmacokinetic model simulated results
3.2 Sinogliatin PBPK Model Development and Evaluation
The final PBPK model was developed with the three different types of methods described previously as Model A, Model B, and Model C. Simulated oral pharmacokinetic parameters for the 15 mg single dose are shown in Table 6 and simulated oral pharmacokinetic profiles are presented in Fig. 4. A virtual Chinese population of 1000 subjects was generated for each simulation. All three models capture the observed data well. However, Model A could not simulate the effect of DDIs, hepatic cirrhosis, and genetic factors on drug exposure with apparent CLiv and Model B cannot predict non-linear enzymatic saturation kinetics in DDI simulation and CYP3A4 polymorphism evaluation. Hence, Model C was selected as the final model to estimate the oral pharmacokinetic parameters and profiles of various doses, as shown in Table 3 and Fig. 2a. For doses from 5 to 50 mg, 90% CIs of simulated CL, Cmax, and AUC values were within those observed in the healthy population, indicating the final model could accurately simulate the metabolism and absorption extent. The 90% CI of the simulated tmax was not the same but close to that of observation, indicating that the simulated absorption rate of the current PBPK model was faster than observed. Overall, the simulation results demonstrate that the sinogliatin PBPK model could capture the observed pharmacokinetic characteristics well in healthy populations.
Simulated (lines) and observed (points) mean plasma concentration–time curves of the sinogliatin 15 mg dosage. The black line represents the mean for the total virtual population. The dashed lines represent the 5th and 95th percentiles for the total virtual population. CI confidence interval, Conc. concentration, Obs clinical observed data
3.3 Sinogliatin PBPK Model Food Effect Simulation and Validation
The pharmacokinetic parameters and profiles of sinogliatin with food were simulated as shown in Table 3 and Fig. 2b. A virtual Chinese population of 1000 subjects was generated for each simulation. The 90% CIs of simulated CL, Cmax, and AUC values were within that observed and the 90% CI of the simulated tmax was partially within that observed, indicating that the final model performed a good simulation of food effect. The simulated profiles also showed that the model could well-fitted the pharmacokinetic characteristics.
3.4 Model Application
3.4.1 Clinical DDI Study Design
For each potential DDI design, the simulated AUC with itraconazole was higher than that without itraconazole. The AUCR of designs A, C, and D were insignificantly different while the AUCR of design B was half of that of designs A, C and D. This indicated that design B had weaker inhibition of CYP3A4. The CmaxR of sinogliatin presented a similar trend (unpublished data). Design D was selected as the final clinical DDI design. A virtual Chinese population of 1000 subjects was generated for each simulation.
3.4.2 Drug Exposure Evaluation in Hepatic Cirrhosis Patients
The simulated AUC values in three Sim-Cirrhosis-CP populations were higher than that in the Sim-Healthy (simulation in healthy patients) population. The simulated AUC in Sim-Cirrhosis-C was 2-fold and 1.5-fold higher than that in Sim-Cirrhosis-A and in Sim-Cirrhosis-B, respectively. The simulated Cmax presented a similar trend (unpublished data). A virtual population of 1000 subjects was generated for each simulation.
3.4.3 Impact of CYP3A4 Alleles on Hepatic Metabolism
The simulated results for different CYP3A4 alleles are shown in Electronic Supplementary Material Table 5. After oral administration of sinogliatin 15 mg, the AUC values in CYP3A4*1, CYP3A4*16, and CYP3A4*18 carriers were 1.09-fold, 0.65-fold, and 1.86-fold of that in CYP3A4 carriers. A virtual population of 1000 subjects was generated for each simulation.
4 Discussion
The overall strategy of the mechanistic understanding process could be summarized as follows:
-
1.
AS, IVIVE, and Css-MRT methods provided understanding of mechanistic DMPKs to support PBPK model development.
-
2.
Based on this knowledge and understanding, a PBPK model was developed by integrating physiologic and drug-specific parameters. The PBPK model was then optimized and validated using the observed clinical data from a SAD study and food effect (MAD) study, respectively.
-
3.
The final PBPK model could not only provide a knowledge base to learn and confirm input data and understanding but also evaluate the effect of extrinsic (DDI) and intrinsic (hepatic cirrhosis, genetic) factors on drug exposure.
AS, IVIVE, and Css-MRT provided mechanistic understanding to support sinogliatin PBPK model development; furthermore, according to the learning exercises of this case study, we proposed an effective strategy for PBPK model construction. For sinogliatin, the AS method provided the human pharmacokinetic parameters of CLiv and Vss. TS-rat,dog was selected as the best AS method according to a unified selection strategy [16]. Using this method we showed that sinogliatin is primarily eliminated through the liver with a high excretion ratio. Additionally, optimal CLiv and Vss values were fixed in the initial PBPK model in order to achieve acceptable elimination prediction; thus, we could focus on absorption optimization of the PBPK model. The IVIVE strategy verified in vitro metabolic data and mechanistically extrapolated in vivo CLh. We understood that sinogliatin was mainly metabolized by CYP3A4 and judged that in vitro microsomal data (rat and dog) were not able to be ascertained using this method. Finally, the in vitro Vmax and Km of rhCYP3A4 were input into the final PBPK model to capture the metabolism properties of sinogliatin based on the IVIVE concept. The Css-MRT approach provided human oral pharmacokinetic profiles of sinogliatin for different dosages. We showed the preclinical interspecies difference using this method: the dog metabolic process was more similar to that of human than that of rat and monkey. This approach had an insignificant contribution to sinogliatin PBPK development; however, for the first-in-human (FIH) studies it is recommended that an animal PBPK model be built to support human PBPK model construction [17,18,19]. Thus, knowledge on interspecies difference was essential to selecting the optimal animal species for constructing the preclinical PBPK model. Furthermore, based on these learning exercises in the sinogliatin case study, we proposed an effective strategy for PBPK model construction. Firstly, simple methods (e.g., AS, IVIVE, and Css-MRT) were used to understand the DMPK properties. This knowledge was then used to support PBPK model construction: the AS method provided human CLiv and Vss. These parameters were fixed in the initial PBPK model, allowing confidence to be gained regarding metabolism estimation which then made it possible to focus on absorption optimization. The IVIVE strategy verified the in vitro metabolic data, and confirmed the predominant CYP enzyme involved in in vivo metabolism and the corresponding fraction (fm). The IVIVE strategy ensured that the PBPK model performed good metabolism prediction using enzyme-level data. The Css-MRT approach provided the knowledge on interspecies difference that enabled selection of the optimal species to construct the preclinical PBPK model in some FIH studies.
Development of a successful prospective PBPK model strongly depends on the input data quality: the more accurate the input parameters, the better prediction the model can provide. Therefore, the primary effort in building a reliable PBPK model should be focused on improving the quality of the input parameters. For absorption, in dissolution testing, the in vitro dissolution profile of sinogliatin was acquired in phosphate buffer via immediate release since sinogliatin is a high-solubility compound with an aqueous phase solubility of 0.42–0.67 mg/mL. However, a 2016 SimCYP® absorption workshop recommended an in vitro dissolution study design, especially for poor-solubility compounds, and the use of a bio-equivalent median (such as simulated gastric fluid [SGF], fasted state simulated intestinal fluid [FaSSIF], fed state simulated intestinal fluid [FeSSIF]) and optimal experimental apparatus (such as USP (United States Pharmacopeia)-2 dissolution model and transfer experiment model) to capture the dissolution rate and precipitation time [20]. In the permeability study, the Caco-2 and/or MDCK-2 cell model should be validated with reference drugs to calibrate the difference between experimental and SimCYP® plug-in values. In the case of sinogliatin, the raw MDCK Papp of 7.72 (10–6 cm/s) could not be used in the model since the reference drugs used to validate the MDCK-2 cell model were unavailable. As a consequence, the absorption extent and rate (Cmax and tmax) could not be accurately estimated in the initial PBPK model (Model E). For metabolism, in the microsomal or recombinant human enzyme incubation system, the substrate metabolism fraction should be greater than 20% and the substrate concentration should be far less than the Km in order to accurately measure the ke [21]. In our case, sinogliatin microsomal data were not accepted because the substrate loss fraction in the rat and dog microsomal incubation systems were far less than 20%. Besides, the IVIVE estimated Fsc of rat and dog were 37 and 148, far exceeding the default accepted criterion of 5, which also proved that microsomal data were unavailable. The substrate metabolism fraction in the rhCYP3A4 incubation system was nearly 85%, achieving the ke calculation requirement. Hence, the enzyme kinetic parameters of rhCYP3A4 could be utilized for PBPK model development.
As a learning-confirming process, the initial PBPK model needs to be continuously refined and optimized. The optimization and validation process of the current PBPK model can be divided into four stages. In stage 1: the primary optimization effort was focused on obtaining accurate elimination estimation. The CLiv estimated by the TS-rat,dog method was put into the PBPK model to acquire acceptable elimination estimation results.
In stage 2, optimization effort was focused on absorption using sensitivity analysis, parameter estimation, and experiment confirmation with the following three steps: step 1, sensitivity analysis was performed to identify the input parameters with the most impact on the model outputs [22]; step 2, by fitting the observed data, the values for the most sensitive parameters were estimated; and step 3, additional experiments were carried out in standard experimental condition to optimize input data. For sinogliatin, the initial PBPK model could not capture the characteristics of absorption (Model E). Sensitivity analysis clearly revealed that the MDCK Papp was the most sensitive parameter affecting sinogliatin absorption. When the MDCK Papp changed ten-fold from that of the raw data value of 7.72 (10–6 cm/s), from 0.772 to 77.2 (10–6 cm/s), the Cmax and tmax output varied from 47.6 to 458 μg/L and 6.10 to 0.752 h, respectively. By fitting the observed data of ten healthy subjects receiving the 50 mg dose, the MDCK Papp was estimated to be 14.33 (10–6 cm/s). Updating the model with the estimated Papp (Model D), the simulation was optimized, with the output results of tmax being 1.17-fold that of the observed data. However, the prediction of Cmax was poor, being 0.58-fold of the observed value. At the same time, an in vitro MDCK-2 cell experiment was carried out with SimCYP® plug-in reference drugs cimetidine and propranolol used as positive controls. The new experimental MDCK Papp of 11.64 (10–6 cm/s) was then used to further update the PBPK model. The final simulated output of Cmax and tmax were 0.99-fold and 0.46-fold of the observed data, respectively. Considering the model application was focused on evaluating the extrinsic and intrinsic effect on the drug exposure (Cmax and AUC), good prediction for Cmax was essential for the current model. Finally, the experimental MDCK Papp was input to provide estimation of absorption. In stage 3, the elimination model was optimized using enzyme-level data for mechanistic insight into the elimination. The CLint,hepatic of rhCYP3A4 was input into the PBPK model. To further evaluate the non-linear enzymatic saturation kinetics in DDI simulation and evaluation of the enzyme polymorphism, the Vmax and Km of rhCYP3A4 were used in the final model. Finally, in stage 4, the optimized PBPK model was validated using observations from the clinical food effect trial. The validated results showed that the mechanistic PBPK model could capture pharmacokinetic properties of sinogliatin well in both a fasted and fed condition.
After the optimization efforts mentioned above, though much improved, some limitations still existed in the model. The 90% CIs of the estimated tmax in fasted and fed status were not within those of observed data. This may be caused by the in vitro–in vivo estimated deviation of dissolution. In vitro dissolution profiles of sinogliatin were obtained from phosphoric buffer with different pH values of 6.8, 4.5, and 1.2. The dissolution profile of pH 6.8 was input in the current model to simulate in vivo solubility since this pH value was closed to endogenous intestinal pH. However, the in vitro dissolution studies revealed that the dissolution capability of sinogliatin decreased significantly in lower pH. Hence, the model-input dissolution profile (pH 6.8) will predict the in vivo dissolution process faster with lower gastrointestinal pH (such as in the gastric tract and pyloric tract) by overlooking the precipitation time. The same limitation existed in tmax prediction of fed status. While the estimated tmax in fed status was delayed compared with that in fasted status, the gastric emptying time of food effect was built-in in the software. Since the model application was focused on evaluating the extrinsic and intrinsic effect of the drug exposure (Cmax and AUC), the under prediction of tmax was acceptable in the current model. The other limitation of the current model is the neglect of the contribution of membrane transporters. For an investigational drug mainly excreted through metabolism such as sinogliatin, the US Food and Drug Administration (FDA) recommend that it is essential to evaluate in vitro whether the drug is a substrate of P-gp, OATP1B1, or OATP1B3 based on the applicable decision tree. P-gp is expressed in the gastrointestinal tract, liver, and kidney, and has a role in limiting oral bioavailability. The in vitro efflux study of P-gp indicated sinogliatin might be a P-gp substrate. However, the contribution of P-gp in sinogliatin absorption is not anticipated to be significant, since sinogliatin is a drug with high solubility and permeability and the intestinal absorption is not a rate-limited step. It is appropriate to exempt such drugs from in vivo evaluation with a P-gp inhibitor based on the FDA guidance [23]. In vitro OATP1B1/1B3 studies also showed that sinogliatin has no inhibited effect on these transporters. Consequently, the contribution of OATP in the sinogliatin elimination process was not considered in the current model.
The PBPK model of sinogliatin provided the following positive impacts on drug development:
Provided a knowledge base to learn and confirm the data and understanding PBPK simulated results provided judgement of input data quality to help decide whether supplementary experiments were needed. Based on simulated results of Model E and sensitivity results, we judged that MDCK Papp was the most sensitive parameter for estimation of model absorption. In addition, the raw in vitro MDCK study design was defective. Hence, we decided to undertake an in vitro supplementary MDCK experiment to optimize this key input parameter. On the other hand, model-simulated results also verified knowledge and understanding provided by other methods. For sinogliatin, PBPK simulated results confirmed knowledge provided by the IVIVE method: by assuming the fm of CYP3A4 to be 100%, the systemic CL could be characterized well via the Vmax and Km of rhCYP3A4.
Clinical DDI study design. The PBPK model provided good quantitative prediction of DDIs via CYP3A4 in prior publications [24, 25]. In our study, design D was selected as the final clinical DDI study design after taking into account clinical safety and the extent of CYP3A4 inhibition, as follows: (i) shortened administration period of itraconazole compared with design A and C to prevent itraconazole-induced hepatotoxicity [26]; (ii) it acquired the maximum inhibition extent of hepatic CYP3A4 for sinogliatin; compared with design B, design D extended the sinogliatin–itraconazole interaction time on the premise that the CYP3A4 inhibition extent of D4 was insignificant compared with that of D8; and (iii) based on the increase in AUC and Cmax due to enzyme inhibition, the sinogliatin dosage was reduced in the clinical DDI trial to prevent drug-induced hypoglycemia. Furthermore, DDIs with erythromycin and rifampicin, as a moderate inhibitor and stimulator of CYP3A4, respectively, were also simulated to support further clinical study design (unpublished data)
Drug exposure evaluation in hepatic cirrhosis patients The PBPK approach was widely used for clinical pharmacokinetic predictions in liver cirrhosis patients, especially for CYP3A-metabolized drugs [27, 28]. For sinogliatin, exposure increases in the Sim-Cirrhosis-A, B, or C population were evaluated to support clinical dosage adjustment. A similar simulation was performed in two SimCYP® virtual chronic kidney disease populations: Sim-RenalGFR-30-60 and Sim-Renal GFR-less-30. However, the clinical dosage adjustment in patients with renal impairment should consider not just renal function but other concurrent factors such as the physiological changes caused by hepatic and kidney interactions and the possible altered activities of efflux transporters and hydrolytic enzyme(s) [29].
Impact of CYP3A4 alleles on hepatic metabolism The expression and catalytic activity of CYP3A are highly variable among individuals, and this variability is partially attributable to genetic factors. CYP3A4*16 and CYP3A4*18 were detected in East Asians with high allele frequency [15, 30]. Thus, the impact of these alleles on the pharmacokinetics of CYP3A4-metabolized drugs should be evaluated. The PBPK approach provided stratified analysis of in vivo metabolism simulation based on the phenotype of CYP3A4. Sinogliatin drug exposure of CYP3A4*18 carriers was 2.9-fold and 1.7-fold higher than that of CYP3A4*16 and wt-CYP3A4*1 carriers, indicating that CYP3A4 polymorphism may cause inter-individual variation of metabolism. However, the impacts of CYP3A4 alleles on catalytic activities was substrate-dependent, and hence the simulated results need further confirmation by clinical data.
5 Conclusion
In this study, we proposed an effective PBPK development strategy based on understanding of mechanistic pharmacokinetics. AS, IVIVE, and Css-MRT methods provide mechanistic insight into human DMPK properties from preclinical in vitro and in vivo data. This knowledge was used to support PBPK model development. The PBPK model was simulated and verified the pharmacokinetics of the SAD study. Various approaches were used to optimize the model. The final model was validated using food effect data from the MAD study. The validated PBPK model provided positive impacts on the drug development of sinogliatin: it was use for the selection of the final clinical DDI study design and evaluated the effects of intrinsic (hepatic cirrhosis, genetic) factors on drug exposure. Our study has three main implications: (i) provides an effective strategy for PBPK development based on mechanistic understandings provided by AS, IVIVE, and Css-MRT; (ii) a PBPK model was developed to simulate the effects of extrinsic and intrinsic factors on drug exposure to support clinical study design; and (iii) provides a methodology of learning and confirms preclinical and clinical data by integrating four methods for the FIH research.
Abbreviations
- AS:
-
Allometric scaling
- AUC:
-
Area under the curve
- BP:
-
Blood plasma ratio
- BW:
-
Body weight
- CI:
-
90% confidence interval
- CL:
-
Systemic clearance in central compartment
- CL/F:
-
Clearance after oral administration
- CLh:
-
Hepatic clearance
- CLint:
-
Intrinsic clearance
- CLiv:
-
Clearance after intravenous administration
- C max :
-
Peak concentration
- CLr:
-
Renal clearance
- C ss-MRT:
-
Steady-state concentration- mean residence time
- DDI:
-
Drug–drug interactions
- ER:
-
Excretion ratio
- F a :
-
Absorbed fraction
- F :
-
Bioavailability
- FASSIF:
-
Fasted state simulated intestinal fluid
- FESSIF:
-
Fed state simulated intestinal fluid
- FIH:
-
First in human
- F sc :
-
Scaling factor between observed and predicted data
- GK:
-
Glucokinase
- GKA:
-
Glucokinase activator
- H :
-
Hematocrit
- HLM:
-
Human liver microsomes
- IVIVE:
-
In vitro to in vivo exploration
- K a :
-
Absorption rate constant
- K m :
-
Michelis–Menten constant
- MPPGL:
-
mg protein per liver weight
- MRT:
-
Mean residence time
- M.W:
-
Molecular weight
- OATP:
-
Organic anion transporting polypeptide
- Obs:
-
Observed values
- P app :
-
Apparent permeability coefficient
- P eff :
-
Jejunum effective permeability
- P-gp:
-
P-Glycoprotein
- PK:
-
Pharmacokinetics
- PBPK:
-
Physiologically based pharmacokinetic
- rhCYP:
-
Recombinant human cytochrome P450 enzyme
- SAS:
-
Simple allometric scaling method
- Sim:
-
Simulated values
- SGF:
-
Simulated gastric fluid
- SSS:
-
Single species scaling method
- T2DM:
-
Type 2 diabetic patients
- T max :
-
Time at the peak concentration occurs
- V c :
-
Distribution volume of central compartment
- TS:
-
Two species scaling method
- V d :
-
Distribution volume of peripheral compartment
- V max :
-
Enzyme maximum rate of metabolite formation
- V ss :
-
Steady-state distribution volume
References
Olokoba AB, Obateru OA, Olokoba LB. Type 2 diabetes mellitus: a review of current trends. Oman Med J. 2012;27(4):269–73. https://doi.org/10.5001/omj.2012.68.
Inzucchi SE, Bergenstal RM, Buse JB, Diamant M, Ferrannini E, Nauck M, et al. Management of hyperglycemia in type 2 diabetes, 2015: a patient-centered approach: update to a position statement of the American Diabetes Association and the European Association for the Study of Diabetes. Diabetes Care. 2015;38(1):140–9. https://doi.org/10.2337/dc14-2441.
Grewal AS, Sekhon BS, Lather V. Recent updates on glucokinase activators for the treatment of type 2 diabetes mellitus. Mini Rev Med Chem. 2014;14(7):585–602.
Filipski KJ, Pfefferkorn JA. A patent review of glucokinase activators and disruptors of the glucokinase–glucokinase regulatory protein interaction: 2011-2014. Expert Opin Ther Pat. 2014;24(8):875–91. https://doi.org/10.1517/13543776.2014.918957.
Anderka O, Boyken J, Aschenbach U, Batzer A, Boscheinen O, Schmoll D. Biophysical characterization of the interaction between hepatic glucokinase and its regulatory protein: impact of physiological and pharmacological effectors. J Biol Chem. 2008;283(46):31333–40. https://doi.org/10.1074/jbc.M805434200.
Grimsby J, Sarabu R, Corbett WL, Haynes NE, Bizzarro FT, Coffey JW, et al. Allosteric activators of glucokinase: potential role in diabetes therapy. Science. 2003;301(5631):370–3. https://doi.org/10.1126/science.1084073.
Xu H, Sheng L, Chen W, Yuan F, Yang M, Li H, et al. Safety, tolerability, pharmacokinetics, and pharmacodynamics of novel glucokinase activator HMS5552: results from a first-in-human single ascending dose study. Drug Des Dev Ther. 2016;10:1619–26. https://doi.org/10.2147/dddt.s105021.
Oie S, Tozer TN. Effect of altered plasma protein binding on apparent volume of distribution. J Pharm Sci. 1979;68(9):1203–5.
Perrier D, Gibaldi M. Clearance and biologic half-life as indices of intrinsic hepatic metabolism. J Pharmacol Exp Ther. 1974;191(1):17–24.
Van den Bergh A, Sinha V, Gilissen R, Straetemans R, Wuyts K, Morrison D, et al. Prediction of human oral plasma concentration-time profiles using preclinical data: comparative evaluation of prediction approaches in early pharmaceutical discovery. Clin Pharmacokinet. 2011;50(8):505–17. https://doi.org/10.2165/11587230-000000000-00000.
Rostami-Hodjegan A. Physiologically based pharmacokinetics joined with in vitro-in vivo extrapolation of ADME: a marriage under the arch of systems pharmacology. Clin Pharmacol Ther. 2012;92(1):50–61. https://doi.org/10.1038/clpt.2012.65.
Xia B, Heimbach T, Lin TH, He H, Wang Y, Tan E. Novel physiologically based pharmacokinetic modeling of patupilone for human pharmacokinetic predictions. Cancer Chemother Pharmacol. 2012;69(6):1567–82. https://doi.org/10.1007/s00280-012-1863-5.
Rodgers T, Rowland M. Mechanistic approaches to volume of distribution predictions: understanding the processes. Pharm Res. 2007;24(5):918–33. https://doi.org/10.1007/s11095-006-9210-3.
Guideline on the qualification and reporting of physiologically based pharmacokinetic (PBPK) modelling and simulation. European Medicines Agency. CHMP458101.2016. http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2016/07/WC500211315.pdf. Accessed 20 Mar 2018.
Maekawa K, Harakawa N, Yoshimura T, Kim SR, Fujimura Y, Aohara F, et al. CYP3A4*16 and CYP3A4*18 alleles found in East Asians exhibit differential catalytic activities for seven CYP3A4 substrate drugs. Drug Metab Dispos. 2010;38(12):2100–4. https://doi.org/10.1124/dmd.110.034140.
Liu D, Song H, Song L, Liu Y, Cao Y, Jiang J, et al. A unified strategy in selection of the best allometric scaling methods to predict human clearance based on drug disposition pathway. Xenobiotica. 2016;46(12):1105–11. https://doi.org/10.1080/00498254.2016.1205761.
Gao ZW, Zhu YT, Yu MM, Zan B, Liu J, Zhang YF, et al. Preclinical pharmacokinetics of TPN729MA, a novel PDE5 inhibitor, and prediction of its human pharmacokinetics using a PBPK model. Acta Pharmacol Sin. 2015;36(12):1528–36. https://doi.org/10.1038/aps.2015.118.
Zhuang X, Lu C. PBPK modeling and simulation in drug research and development. Acta Pharm Sin B. 2016;6(5):430–40. https://doi.org/10.1016/j.apsb.2016.04.004.
Liu F, Zhuang X, Yang C, Li Z, Xiong S, Zhang Z, et al. Characterization of preclinical in vitro and in vivo ADME properties and prediction of human PK using a physiologically based pharmacokinetic model for YQA-14, a new dopamine D3 receptor antagonist candidate for treatment of drug addiction. Biopharm Drug Dispos. 2014;35(5):296–307. https://doi.org/10.1002/bdd.1897.
Fotaki N, Gray V, Krämer J, Diaz D, Flanagan T, Grove G. Dissolution highlights from the 2015 AAPS Annual Meeting in Orlando. Dissolut Technol. 2016;23(2):42–7. https://doi.org/10.14227/dt230216p42.
Barter ZE, Bayliss MK, Beaune PH, Boobis AR, Carlile DJ, Edwards RJ, et al. Scaling factors for the extrapolation of in vivo metabolic drug clearance from in vitro data: reaching a consensus on values of human microsomal protein and hepatocellularity per gram of liver. Curr Drug Metab. 2007;8(1):33–45.
Gabrielsson JL, Groth T. An extended physiological pharmacokinetic model of methadone disposition in the rat: validation and sensitivity analysis. J Pharmacokinet Biopharm. 1988;16(2):183–201.
Clinical drug interaction studies: study design, data analysis, and clinical implications guidance for industry. US Food and Drug Administration, CDER; 2017. https://www.fda.gov/downloads/drugs/guidancecomplianceregulatoryinformation/guidances/ucm292362.pdf. Accessed 20 Mar 2018.
Mano Y, Sugiyama Y, Ito K. Use of a physiologically based pharmacokinetic model for quantitative prediction of drug–drug interactions via CYP3A4 and estimation of the intestinal availability of CYP3A4 substrates. J Pharmaceut Sci. 2015;104(9):3183–93. https://doi.org/10.1002/jps.24495.
Vieira ML, Zhao P, Berglund EG, Reynolds KS, Zhang L, Lesko LJ, et al. Predicting drug interaction potential with a physiologically based pharmacokinetic model: a case study of telithromycin, a time-dependent CYP3A inhibitor. Clin Pharmacol Ther. 2012;91(4):700–8. https://doi.org/10.1038/clpt.2011.305.
Pettit NN, Pisano J, Weber S, Ridgway J. Hepatic failure in a patient receiving itraconazole for pulmonary histoplasmosis—case report and literature review. Am J Ther. 2016;23(5):e1215–21. https://doi.org/10.1097/mjt.0000000000000313.
Edginton AN, Willmann S. Physiology-based simulations of a pathological condition: prediction of pharmacokinetics in patients with liver cirrhosis. Clin Pharmacokinet. 2008;47(11):743–52. https://doi.org/10.2165/00003088-200847110-00005.
Johnson TN, Boussery K, Rowland-Yeo K, Tucker GT, Rostami-Hodjegan A. A semi-mechanistic model to predict the effects of liver cirrhosis on drug clearance. Clin Pharmacokinet. 2010;49(3):189–206. https://doi.org/10.2165/11318160-000000000-00000.
Zhao P, Vieira Mde L, Grillo JA, Song P, Wu TC, Zheng JH, et al. Evaluation of exposure change of nonrenally eliminated drugs in patients with chronic kidney disease using physiologically based pharmacokinetic modeling and simulation. J Clin Pharmacol. 2012;52(1 Suppl):91s–108s. https://doi.org/10.1177/0091270011415528.
Fukushima-Uesaka H, Saito Y, Watanabe H, Shiseki K, Saeki M, Nakamura T, et al. Haplotypes of CYP3A4 and their close linkage with CYP3A5 haplotypes in a Japanese population. Hum Mutat. 2004;23(1):100. https://doi.org/10.1002/humu.9210.
D’Argenio DZ, Schumitzky A, Wang X. ADAPT 5 user’s guide: pharmacokinetic/pharmacodynamic systems analysis software. Los Angeles: Biomed Simulations Resource; 2009. https://bmsr.usc.edu/software/adapt/citations/
Barter ZE, Tucker GT, Rowland-Yeo K. Differences in cytochrome p450-mediated pharmacokinetics between Chinese and Caucasian populations predicted by mechanistic physiologically based pharmacokinetic modelling. Clin Pharmacokinet. 2013;52(12):1085–100. https://doi.org/10.1007/s40262-013-0089-y.
Crewe HK, Barter ZE, Yeo KR, Rostami-Hodjegan A. Are there differences in the catalytic activity per unit enzyme of recombinantly expressed and human liver microsomal cytochrome P450 2C9? A systematic investigation into inter-system extrapolation factors. Biopharm Drug Dispos. 2011;32(6):303–18. https://doi.org/10.1002/bdd.760.
Brown RP, Delp MD, Lindstedt SL, Rhomberg LR, Beliles RP. Physiological parameter values for physiologically based pharmacokinetic models. Toxicol Ind Health. 1997;13(4):407–84. https://doi.org/10.1177/074823379701300401.
Davies B, Morris T. Physiological parameters in laboratory animals and humans. Pharm Res. 1993;10(7):1093–5.
Acknowledgements
We acknowledge Dr Bo Liu for his comments for the SimCYP® software technology support.
Funding
This study was supported by the National Natural Science Foundation of China (Nos. 81403013 and 81403015) and the ‘13th Five-Year’ National Major New Drug Projects (Nos. 2017ZX09101001-002-001 and 2017ZX09304031-001).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Yi Zhang, Shuang Ren, and Li Chen are employees of Hua Medicine (Shanghai) Ltd., the company developing sinogliatin. Ling Song, Ji Jiang, Dongyang Liu, Xijing Chen, and Pei Hu declare no conflicts of interest relevant to the contents of this manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Song, L., Zhang, Y., Jiang, J. et al. Development of a Physiologically Based Pharmacokinetic Model for Sinogliatin, a First-in-Class Glucokinase Activator, by Integrating Allometric Scaling, In Vitro to In Vivo Exploration and Steady-State Concentration–Mean Residence Time Methods: Mechanistic Understanding of its Pharmacokinetics. Clin Pharmacokinet 57, 1307–1323 (2018). https://doi.org/10.1007/s40262-018-0631-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-018-0631-z