Abstract
Multidrug-resistant (MDR) bacteria are considered a health threat worldwide, and this problem is set to increase over the decades. The ESKAPE, a group of six pathogens including Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter spp. is the major source of concern due to their high death incidence and nosocomial acquired infection. Host defence peptides (HDPs) are a class of ribosomally synthesised peptides that have shown promising results in combating MDR, including the ESKAPE group, in- and outside bacterial biofilms. However, their poor pharmacokinetics in physiological mediums may impede HDPs from becoming viable clinical candidates. To circumvent this problem, chemical engineering of HDPs has been seen as an emergent approach to not only improve their pharmacokinetics but also their efficacy against pathogens. In this review, we explore several chemical modifications of HDPs that have shown promising results, especially against ESKAPE pathogens, and provide an overview of the current findings with respect to each modification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Chemically engineered HDPs are a promising tool to combat MDR bacteria. |
Not all currently available chemical modifications to HDPs result in better efficacy against MDR bacteria, while displaying low off-target side effects against host cells. |
Using multiple chemical modifications to one HDP simultaneously can add a synergistic effect against MDR bacteria, which offers significant clinical potential. |
1 Introduction
Antimicrobial resistance (AMR) refers to the ability of a microorganism to resist and adapt to antimicrobials for its survival [1]. This is considered one of the biggest health threats globally, taking a significant toll on both economic and health aspects. For human health, it is estimated that ten million people will be at risk from bacterial infection by 2050, which will surpass the mortality of cancer [1]. In the finance context, 100 trillion USD of the global economy may be lost over the next three decades as a result of AMR [1]. Currently, the rate at which microbes are acquiring resistance is overtaking the development of new drugs that reach clinics [1, 2]. This is one of the main factors for research and development (R&D) discouragement by pharmaceutical companies, since it leads to low returns on investment, given that by the time of patent expiration, the antibiotic may no longer offer a protective effect [1].
Multidrug-resistant (MDR) bacteria are a class of microorganisms that present resistance to more than one previously sensitive antibiotic [3]. Many strategies are employed by pathogens to escape antibiotic efficacy, such as altering their porins to become less permeable to antibiotics, overexpressing antibiotic efflux pump proteins and the production of enzymes capable of breaking down key structures of antibiotics [2]. Particularly, the most concerning MDR pathogenic agents, due to their high mortality and hospital incidence, are the ESKAPE group, which stands for six bacterial species: Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter spp. [4, 5]. S. aureus, for example, is a well-known pathogen that can circumvent the action of antibiotics not only through gene-induced resistance (e.g. mecA) but also through well-documented biofilm-forming activity in catheters and implants [6,7,8], presenting overall mortality that ranges between 15 and 50% in septic cases [9]. Vancomycin-resistant E. faecium is widely spread in several countries, present in 47% of blood culture isolates in Australia and responsible for 30% of acquired bacteraemia [10]. Several clinical isolates of K. pneumoniae have been shown to produce extended-spectrum β-lactamases and carbapenemases, resorting to toxic antibiotics for its treatment such as polymyxin and aminoglycosides [11, 12]. In the case of A. baumannii, the pathogen has been shown to develop resistance to conventional antibiotics more rapidly than other bacterial species, such as K. pneumoniae and P. aeruginosa. In addition to its fast adaptability to microbial drugs, A. baumannii has shown great adaptability to colonise and produce extremely antibiotic-resistant biofilms on various surfaces, especially in catheters, which contributes to it being the most serious bacterial-related problem worldwide [13]. Lastly, bacteria belonging to Enterobacter spp. have been typically a major concern to neonates and patients in the intensive care unit, specifically those requiring ventilation assistance [12].
Many strategies have been developed and are developing for the treatments of MDR clinically, for example, synthesising drugs that inhibit proteins produced by resistance genes, creating new compounds from existing classes of antibiotics (e.g. β-lactams, fluoroquinolones, macrolides), and introducing new antibiotics such as natural host defence peptides (HDPs) [14]. However, many of these approaches have failed to be approved by the Food and Drug Administration (FDA), or even if approved, they have been clinically limited due to displaying off-target side effects, such as nephrotoxicity. In addition, another problem that many large pharmaceutical companies face when attempting to release new drugs targeting specific enzymes is cross-resistance to existing classes of antibiotics. This impedes the development because in vitro or systemic assays may show that these new antibiotics can induce resistance to previously conventional antibiotics, which hinders FDA approval [14]. It is worth noting that, while many antibiotics have been recently approved or are in the late stages of clinical trials showing promising results against many species of bacteria, a lot of them may still fail to provide full benefits to biofilm-forming bacteria, especially in patients with catheters or certain types of implants. Therefore, this component should also be evaluated when assessing the overall efficacy of new antibiotics [15].
Being considered a new class of antibiotics, HDPs might provide innovative ‘weapons’ to combat MDR bacteria while presenting low cytotoxicity and high specificity towards both Gram-positive or Gram-negative bacteria and can even be effective against pathogens in biofilms. However, HDPs may have several limitations regarding their pharmacokinetics, demonstrating poor bioavailability in serum or in other tissues. This review aims to discuss how HDPs, different from traditional antibiotics, can be used to combat MDR bacteria in both planktonic and biofilm bacteria, and highlights what chemical approaches have been developed to improve the activity of HDPs, thus presenting better clinical outcomes.
2 The Application of Host Defence Peptides in Treating MDR Bacteria
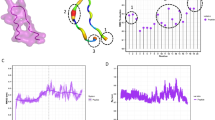
HDPs normally refer to cationic amphipathic short-sequence peptides produced by all six life kingdoms. They play roles in protecting the host against pathogens by direct killing infectious agents or ramping up the immune system's defence by various means [16, 17]. HDPs typically adopt secondary structures, such as alpha helices and beta sheets, but they can also exert their effect in a linear form. To establish bacterial death, HDPs have an overall positive charge that attracts the negatively charged phospholipids highly present on both Gram-positive and Gram-negative bacteria, mostly including phosphatidylserine and phosphatidylglycerol [18, 19]. However, in the case of Gram-positive bacteria, lipoteichoic acids play a more significant role in the binding of HDPs than phospholipids alone [19] (Fig. 1). Liposaccharides (LPS) found in Gram-negative bacteria are highly anionic molecules that largely increase the electronegativity potential of the outer membrane. The negative charge of LPS interacts with the positive charge of cationic HDPs through electrostatic interactions, which is an important mechanism for HDPs binding to the bacterial membrane [19] (Fig. 1A).Many studies have shown the potential for different HDPs to bind to LPS, leading to the accumulation of peptides on the bacterial membrane [20]. This has also been confirmed by the studies in endotoxemic mice, where the binding of HDP to LPS improved survival outcomes of the mice [20]. However, it is worth noting that although many HDPs present bactericidal activities, they may not prevent endotoxemia or even possess low to no affinity for binding to LPS [19, 20]. The HDP DNS-PMAP23, for example, has been shown to directly kill Escherichia coli strains with minimal involvement of LPS. The study demonstrated that the peptide does not bind or react with either the O antigen or the outer core of the bacterial cell surface, and that the phospholipids present on the bacterial membrane play a more significant role in the non-specific interaction than LPS alone.
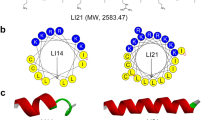
Schematic illustration of host defence peptides (HDPs) mechanism of action. A Differences between the linear and cyclic HDPs. The cyclic peptide display side chains that point outwards to its planar macrocycle, increasing the contact area with their target ligands, whereas side chains of linear peptide do not fully engage with their ligands in a three-dimensional plane. B Common targets of HDPs once translocated to bacteria cytoplasm. C Mechanism of action of HDPs on the bacterial membrane.
The electrostatic interaction between HDPs and the bacterial membrane disrupts the membrane fluidity, resulting in increased permeability and loss of cytoplasmic content, and eventually causes cell death [19, 21, 22]. It is noteworthy that the bactericidal efficacy of HDPs can be augmented through synergistic action when used in combination with conventional antibiotics, even in bacterial strains that are resistant to the antibiotic alone. For instance, Bip-P-113 has been demonstrated to have a synergistic effect when combined with vancomycin, even in strains carrying vancomycin-resistant genes [23].
Several mechanisms of membrane disruption caused by HDPs have been proposed, such as the barrel-stave, carpet and toroidal models [19, 22] (Fig. 1B, C). Moreover, some HDPs may also have intracellular targets, such as DNA- and RNA-associated proteins or ribosomes, and exert bactericidal effect via modulating different biological pathways [22]. For example, the lasso peptides MccJ25 and Cap have been shown to interact with the secondary channel of the RNA polymerase from E. coli, blocking the trigger loop of the enzyme necessary to add nucleotides to the growing chain of RNA [24]. Several amphibian HDPs, such as caerin, aurein, magainin and so on, have been shown to inhibit the activity of bacterial ATPase [25, 26]. Another mechanism of intracellular HDP is shown by the peptidomimetic L-27-11. This peptide binds to the lipopolysaccharide transporter, specifically the LptD, which immobilises the transport of this molecule onto the outer membrane of the bacteria, causing cell death [27].
Although HDPs have many limitations that may hinder their viability to become antimicrobial drugs (see Sect. 4 below), a major advantage of these peptides over conventional antibiotics is their potential to circumvent the development of antimicrobial resistance. Two possible reasons have been identified to address this phenomenon, the first being the non-specific target of HDPs, since they target electronegative molecules on the outer membrane of bacteria such as phospholipids or lipoteichoic acid, as mentioned previously [19, 22, 28]. Secondly, apart from their non-specific interaction, HDPs may target multiple cellular targets, thus requiring bacteria to undergo several mutations simultaneously to resist these peptides [28,29,30], which points to difficulties at the evolutionary level.
3 HDPs for Biofilms Formed by MDR Bacteria
Another major challenge is that MDR bacteria can sometimes form biofilms. Biofilms refer to the complex communities of microbes that may be found attached to a surface in methicillin-resistant Staphylococcus aureus (MRSA), A. baumanii, P. aeruginosa, S. aureus and some other bacterial infection [31]. Most pathogens and non-pathogens within microbial biofilms develop together as microbial communities [32], which can tolerate environmental stresses such as starvation and desiccation, to become more resilient to antibiotics [33]. Clinical treatment of biofilms is highly challenging as they are notoriously recalcitrant to antimicrobial therapy. Skin infection can result from multi-bacterial infection that leads to the formation of biofilms [34,35,36]. Besides, patients with indwelling medical devices such as catheters, pacemakers, joint prostheses, dentures, prosthetic heart valves and contact lenses are at a higher risk of developing nosocomial infections due to the potential for bacterial colonisation [37, 38] and biofilm formation on these surfaces [39, 40].
As described above, HDPs have shown broad-spectrum activity against many strains of Gram-positive and Gram-negative bacteria, including drug-resistant strains and fungi. In addition, HDPs exhibit synergy with conventional antibiotics and neutralise toxins in animal models. Some HDPs also confer anti-biofilm activity. The human cathelicidin peptide LL-37 was demonstrated to inhibit biofilm formation at a concentration 16-fold lower than its minimum inhibitory concentration (MIC) against planktonic bacteria [41]. LL-37 was subsequently shown to possess anti-biofilm activity against urinary tract isolates of S. aureus and E. coli at 1/32 to 1/2 MIC. Synthetic cathelicidin-derived anti-biofilm peptides (such as innate defence regulator 1018, DJK-5 and DJK-6) have been developed, which exhibit broad spectrum activity against the biofilms formed by MDR organisms [42]. A single 4 h treatment with hypromellose ointment containing SAAP-148 completely eradicated acute and established biofilm-associated infections with methicillin-resistant S. aureus and MDR A. baumannii from wounded ex vivo human skin and murine skin models [43]. Recently, we demonstrated that caerin 1.1/1.9 peptides isolated from the skin secretions of the Australian amphibian tree frog were able to inhibit the formation of biofilms of MRSA and A. baumannii with an in vitro model (unpublished data).
4 Shortcomings Hamper HDP Clinical Application
Unlike conventional antibiotics, which have an affinity for specific pathogen(s), HDPs target non-specific molecules of bacteria, reducing the likelihood of the bacteria developing resistance [18, 44, 45]. Several HDPs have proven to be efficient in the clinical setting, with some examples being commercialised, such as polymyxin B, gramicidin and daptomycin [29]. Surprisingly, the majority of HDPs in the market are classified as non-ribosomal peptides [29], which are most frequently found in bacteria and fungi, due to possessing features that confer them better structure stability, such as naturally non-proteogenic amino acids (e.g. hydroxyphenyl, dihydroxyphenyl-glycine or (4R)-4- [(E)-2-butenyl]-4-methyl-L-threonine), a macrocyclic structure, or other post-translation modifications (e.g. lipidation, glycosylation or N-methylation) [46]. Although these organisms have been shown to provide a rich source of natural HDP products, more than 93% of the HDPs discovered so far have been found in life forms other than prokaryotes and protists [47]. To date, whilst many HDPs are being trialled, no ribosomal HDP derived from organisms other than bacteria or fungi has been approved by the FDA for treating infections [29]. This shows the opportunities and challenges in commercialising ribosomal HDPs from other life forms.
There are many reasons associated with the failed attempts at making HDPs viable clinical products. The significant deficiency in their intrinsic pharmacokinetics is the major limitation of these peptides. Poor thermostability and a chemical structure propitious for targeting by proteases, especially in serum, largely reduces their efficacy at disrupting bacterial membranes [29]. For example, caerin 1.9, a peptide with promising pre-clinical results for anti-cancer and bactericidal activity [48, 49], has a short half-life of 1.16 h in male Sprague–Dawley rats [50]. Consequently, many of these peptides are only effective at higher concentrations (micromolar) in in vivo tests rather than the expected lower concentrations (nanomolar) . Whilst this concentration still offers a beneficial therapeutic value, it might cause off-target side effects, such as high haemolytic activity, nephrotoxicity and, potentially, neurotoxicity [29, 51], hindering their progress in clinical trials and approval by regulatory agencies [29]. This explains why most of HDPs with therapeutic potential are often prepared into a gel or cream formula, and are applied topically rather than being administered systemically [29].
In addition, it is worth noting that the development of bacterial resistance against certain HDPs in vitro has been observed [52, 53]. Shireen et al. [54] demonstrated that S. aureus (ATCC 29213) treated with manganin 2 and gramicidin D developed resistance by changing phospholipid composition and membrane rigidity, although only after several linage passages. Other mechanisms of resistance to HDPs have also been shown in S. aureus. For example, S. aureus has been shown to produce proteinases against LL-37, which decreased the activity of this HDP [55].
To address the previously described limitations of HPDs, various approaches have been developed to create a new platform for products that are safer, more effective and less susceptible to bacterial resistance. One of these approaches is chemically engineered HDPs, which involves modifying the structures of HDPs using various chemical methods. This approach holds promise in overcoming the challenges associated with natural HDPs and may pave the way for the development of novel antimicrobial agents with improved characteristics.
5 Chemical Engineering Host Defence Peptides for Better Anti-MDR Bacterial Activity
5.1 Engineering of Cationic HDPs with Helical Amphipathicity
As previously described, HDPs typically adopt secondary structures such as alpha helices or beta sheets, but can also be extended [56]. However, HDPs do not display these geometrical conformations in aqueous solution and usually undergo conformational change upon interacting with bacterial membranes [56,57,58]. The secondary structure of these HDPs is thought to be essential for their antimicrobial activity [44, 59], although this hypothesis has been disputed [57]. It is important to note that chemical modifications made to HDPs could potentially disturb or alter their secondary conformation, as reported in previous studies [60,61,62,63], which in turn may impact their effectiveness. Hence, when considering any chemical modifications to HDPs, it is crucial to take this into account and exercise caution.
To be classified as an HDP with a high therapeutic index, which means possessing potent bactericidal activity with low toxicity towards mammalian cells, an optimal balance between cationic and hydrophobic residues is necessary, in addition to the formation of their secondary structure [64]. A hydrophobic shift in peptide composition results in HDP inactivation and increased aggregation, whereas an increase in cationic charge leads to higher bactericidal activity at the expense of greater toxicity towards red blood cells [64]. To achieve optimal anti-bacterial activity, engineered HDPs or de novo designed antimicrobial peptides (AMPs) must possess specific chemical and structural characteristics, including an appropriate net charge, a polar and non-polar face, good amphipathicity and the ability to adopt an alpha-helical conformation upon interaction with bacterial membranes, as well as low toxicity.
Most known HDPs possess a cationic charge that ranges between +2 and +9 [65]. A higher charge, however, is not necessarily correlated with better antimicrobial effect. A slight increase in charge in the net charge of magainin 2 by replacement of some amino acids has been shown to improve bactericidal activity, but higher charges have been shown to decrease concomitantly antimicrobial activity and increase haemolytic activity [66]. It is generally recognised that a charge between + 4 and + 6 for most HDPs might be optimal in terms of balancing antimicrobial activity and haemolysis. To achieve this goal, the replacement of non-polar amino acids for polar ones, especially at the polar face of the peptide, can be done by typically substituting with Arg residue (Table 1). Arginine is prevalent in many HDPs, and it has been shown to aid membrane thinning upon insertion and formation of hydrogen bonds with water molecules [67].
It is worth noting that the modification of the D1 peptide, which previously had a net charge of +6 and was subsequently engineered to display a net charge of +11, has significantly increased its therapeutic index [68]. Specifically, replacing the hydrophobic amino acid in the midpoint of the helix with a cationic amino acid (Val to Lys) has not only improved the peptide’s bactericidal activity but also reduced its selectivity towards red blood cells. Recent studies have demonstrated the same effect of introducing a hydrophobic residue ‘break’ on the non-polar face [69, 70]. Thus, it can be inferred that the net charge of an engineered HDP may not be the sole determinant factor that researchers should focus on when attempting to optimise its activity.
In addition to the charged and polar face of the peptide, ideally, an optimal bactericidal HDP would have a hydrophobic face on the other side of the helix, making it amphipathic. Indeed, amphiphilicity has been shown to not only improve antimicrobial activity, but also enhances engineered peptide’s affinity to the membrane of red blood cells (RBCs) [71,72,73]. While the cationic side of the peptide drives and interacts with the molecule to the non-specific negatively charged molecules (LPS, LTA and anionic phospholipids), the hydrophobic face is responsible for the insertion of the peptide to the lipid monolayer [72].
Thus, when rationalising a peptide design, it is essential to consider the peptide’s length and functionalisation in various physiological conditions, such as high salt concentration, blood and serum. Optimising the peptide length can potentially reduce haemolytic activity [74, 75] (Table 1). However, if the haemolytic activity remains unchanged despite decreasing the peptide length, shorter amino acid chains can be advantageous due to their low production cost, making commercialisation more feasible [76].
5.2 PEGylation
Although many HDPs can demonstrate efficacy in vitro, they often fail to provide therapeutic efficacy in serum because of the ‘hostile’ environment caused by systemic circulation. In addition, most therapeutic peptides or proteins can have an overall hydrophobic profile, which affects their solubility in blood. In light of these problems, PEGylation of HDPs has been proposed as a strategy to enhance pharmacokinetic properties in serum, which is to conjugate polyethylene glycol (PEG) with various numbers of monomers to selected amino acids of therapeutic proteins or peptides [77]. PEG molecules could introduce many physiochemical proprieties to HDPs. Firstly, this polymer contains a backbone chain that oscillates between carbons and oxygen molecules (C–O–C), which presents an overall hydrophilic profile. Covalent attachment of this molecule therefore adds a polar component to the HDP, which then becomes more soluble in blood [78,79,80]. Secondly, PEGylation, particularly with high-molecular weight PEG molecules, can decrease the rate of renal excretion [81,82,83]. Also, due to PEG possessing a low immunogenic profile, the molecule can physically shield epitopes of the conjugated HDPs, making them less susceptible to becoming antibody targets [77, 84, 85].
However, it is worth noting that the sizes (molecular weights) and the structures (the functional groups on the branches) of PEGs and the choice of PEGylation site(s) on the HDP sequence might influence the interaction with the targets, consequently impacting the bioactivity [86,87,88]. Nevertheless, the overall benefit of improved pharmacokinetics in vivo normally outweighs this issue, as the PEGylated peptides circulate longer in blood [89,90,91]. Based on the advantages explained above, several attempts have been made to PEGylated HDPs to overcome the limitations described. As expected, most of the studies seeking to PEGylate HDPs have shown decreased bactericidal activity in vitro compared with their non-PEGylated counterparts [60, 86, 88, 92,93,94,95,96,97] (Table 2). To explain this phenomenon, some studies have suggested that PEGylation offers a physical barrier between the peptide and the bacterial membrane, which diminishes electrostatic interactions and thus reduce pore formation [88, 92]. Notwithstanding, all studies previously described, except for [95], have demonstrated a decreased haemolytic activity in vitro upon PEGylation of their respective HDPs.
In the context of reduced HDP bioactivity against bacteria, the results of these studies should be interpreted carefully. Although the bioactivity of PEGylated peptides was decreased compared with their non-PEGylated cognates, a better way to assess in vitro bioactivity that can be translated to in vivo assays is to incubate PEGylated and non-PEGylated HDPs in serum prior to measuring their MIC50 [86, 92]. The practicality of this hypothesis was shown by a previous study, in which P. aeruginosa PAO1 was inoculated in neutropenic mice in a sepsis model and treated intravenously with PEGylated SET-M33L [98]. Although PEGylated peptides show, in general, less activity in vitro, the PEGylates SET-M33L peptide promoted 20% higher survival rates in these mice when compared with the wild peptide type.
PEGylation has already proven to be a potent chemical modification to improve biomolecules, and although this technology has already been in use for many decades, very few studies have validated its therapeutic value in HDP in vivo for bacterial infection. It is worth mentioning that many successful PEGylated biomolecules that have been on the market, such as IFN-α2, have previously shown higher bioactivity in vivo, but very limited activity in vitro, than their non-PEGylated counterparts. PEGylated IFN-α has retained only 7% of its activity in vitro; however, the in vivo in clinical response induced a much stronger effect than non-PEGylated cognate protein [89, 99]. In our opinion, PEGylated HDPs can possibly be promising agents in the combat of MDR should more research and clinical trials validate this hypothesis.
5.3 Halogenation
Halogenation of peptides is one chemical modification that can increase the hydrophobicity and lipophilicity of peptides [100] and has been shown to either improve or decrease the antimicrobial effects of some HDPs [101, 102]. It includes the covalently binding of halogens to selected amino acids of the HPDs. A halogenated tetrapeptide scaffold varying at one residue (Xaa: Lys, Gly, Ala or Phe) has been engineered and shown low MIC values, especially for strains containing extended spectrum β-lactamase–carbapenemase-producing isolates [102]. Gram-negative bacteria have been shown to be more affected by bulky hydrophobic residues (Xaa Phe) when incorporated with CF3 halogen, although haemolytic activity has increased. However, using di-CF3 with Phe has decreased the overall activity of the peptide. Halogenation of tetrapeptide (CF3) with Phe has shown to have the lowest MIC values for all ESKAPE pathogen isolates containing multiple resistant genes, including MRSA.
5.4 Cyclisation of HDPs
As previously described, HDPs may present strong action against bacteria in vitro, but their susceptibility to proteolytic cleavage, rapid excretion in the kidneys and low oral bioavailability have impeded many peptides from becoming viable candidates for the biopharmaceutical industry. Many peptides usually present low penetrance to cell membranes because even short peptides far violate the ‘Lipinski rule of five’, which hinders the interaction with intracellular targets and thus affects oral uptake [103, 104]. The amide bonds in a linear peptide are more susceptible to being targeted by peptidase, particularly in the intestinal tract, which further reduces its oral availability [105]. Cyclisation of peptides has been considered as an approach to circumvent these problems (Fig. 1A). There are several methods of cyclisation, including head-to-tail, head-to-side chain and disulphide cyclisation. Head-to-tail cyclisation connects the two ends of the peptide, the N-terminus to the C-terminus, while head-to-side chain cyclisation connects the N-terminus to a side chain, such as a cysteine. Disulphide cyclisation occurs when two cysteines in the peptide backbone form a disulphide bond [106].
Many naturally cyclised HDPs have been discovered, and some of them have even been approved by regulatory agencies as antibiotics, such as the polymyxins, daptomycin, gramicidin and tyrothricin (Table 3) [29, 107,108,109]. Structural rigidity provided by cyclisation decreases the number of rotatable bonds in the peptide, which chemically constrains the peptide backbone and alters its stereochemistry, resulting in the new spatial geometry of these peptides, can often preclude the action of peptidases. In addition, the structural constraint in cyclised peptides improves their affinity with target ligand, due to fewer conformations that the peptide can achieve [104, 110]. In the case of head-to-tail ligation (i.e. ligation of the C- to N-terminus), fewer ionisable groups are encountered in the molecule, which improves the peptide’s polar surface area (PSA). Thus, the high structural stability and target affinity of the cyclised peptides in turn improves oral bioavailability [110, 111].
It is known that cyclisation of HDPs also exposes their side chains outwards, which may provide several benefits [104]. Exposure of these side chains throughout the peptide better distributes the polarity across the molecule, making peptides more amphiphilic [112]. In addition, side chain exposure also improves the electrostatic and membrane interaction of HDPs with several microbes, including those with resistant genes against conventional antibiotics, which consequently enhances the bioactivity (Table 3) [112, 113]. A study has produced a library of short-sequence peptoids, and compared the MIC50 values between their linear and head-to-tail cyclised structure [113]. It has been found that the cyclisation of these peptoids has, overall, decreased their MIC50, with some leading to seven to eight-fold lower than their linear counterparts, while also decreasing haemolytic activity. Cyclisation of antimicrobial peptides reduces their ability to bind to and permeate through phosphatidylcholine and cholesterol, which are major components of the outer leaflet of red blood cell membranes. This decrease in binding and permeating activity leads to a reduction in haematolytic activity [114].
To certain peptides, cyclisation via disulphide bonds may provide better benefits than head-to-tail cyclisation. Depending on the nature and sequence of the HDPs, cyclisation via cysteine amino acids can be a more facile, cheaper and quicker method [115]. Cysteine knotting in acyclic Kb1 confers equal protection against peptidases and only a slight reduction in thermostability when compared with their linear and wild head-to-tail cycle peptide type [116]. This shows that cystine bonding may play a more significant role in the structure and stability of the cyclotide rather than ligation of C- to N- terminus. In addition, the authors also produced mutants of those cyclotides with significant less cysteines. Their results showed that the number of cysteines was correlated with their susceptibility of being denatured by denaturants, such as urea, which demonstrate the importance of disulphide bonds in the structure stability of peptides. Another study has also shown the superiority of cysteine cyclisation in the case of lfcinB (Table 3) [117]. Nguyen et al. [117] has created a mutant of this peptide with two terminal cysteines cyclised via disulphide bonds, and this modified HDP has displayed a better antimicrobial activity and serum stability than its backbone joined cognates.
5.5 Non-canonical Amino Acids
Non-canonical amino acids (ncAAs) are a class of amino acids that differ from the 20 standard amino acids commonly found in proteins; they are carried by modified tRNAs during peptide synthesis. Some of these ncAAs, however, can be found in many natural HDPs, and usually are the results of metabolic intermediates of canonical amino acids [118]. The incorporation of ncAAs can improve the physiochemical characteristic of antimicrobial peptides to become more lethal to pathogens, reduce the likelihood of resistance development, and improve their bioavailability [118, 119]. Growing research interest has been advocated to either producing synthetic peptides through incorporating ncAAs into already known HDPs, or using it as a scaffold to mimic HDPs (peptidomimetics). This is because the balance between the overall charge and hydrophobicity of the HDP highly impacts the anti-bacterial and haemolytic activity [120], two characteristics that can be improved through the incorporation of ncAAs [113, 118], as described in section 5.1. The types of ncAAs that have been widely investigated in engineering HDPs chemically mainly include D-amino acids and poly-N-substituted glycines.
5.5.1 D-Amino Acid Substitution
Although amino-acids have an sp3 hybridised α-carbon, which gives them a chiral centre and the potential for mirror image isomers, the D-stereoisomer of amino acids is rarely found in proteins in living organisms [121]. The incorporation of these amino acids into peptides typically does not involve ribosomes themselves, and they are instead incorporated into the growing peptide chain through other synthetic mechanisms. Whilst rarely found in nature, there are many reasons why organisms may wish to include D-amino acids in the composition of peptides and proteins. The stereochemistry of these amino acids protects them against enzymes and proteinases that would otherwise degrade their peptide chain [122]. Bacteria, for example, incorporate D-amino acids in their peptidoglycan to become less likely to interact with proteinases, thus becoming more adaptable to certain environments [122]. The difficulty in cleaving peptides with D-amino acids that are stereoisomers may be because most of the peptidases and proteinases produced by bacteria are specific to L-amino acids. As a result, it may be challenging for bacteria to adapt and produce D-stereoisomer peptidases [119]. For example, polymyxin B is the product of the fermentation of Paenibacillus polymyxa, which is partly composed of aminobutyric acid and other D-amino acids in its residues [108]. The biosynthesis of penicillin G for example, which contains a metabolite of D-Valine, involves the joining and modification of three base amino acids by three enzymes: δ-(L-a-aminoadipyl)-L-cysteinyl-D-valine synthetase, isopenicillin-N synthase and acyl-CoA–isopenicillin-N acyltransferase [123].
It is not a coincidence that most of the market-approved antimicrobial peptides contain D-amino acids in their composition, such as polymyxins, daptomycin and gramicidin [29, 108, 122]. In light of the success and proven ability of D-amino acid hitting the clinics, many other HDPs have been modified to include D-amino acids in their composition for better stability and decreased toxicity towards mammalian cells, in addition to improving their bactericidal effect and potentially diminished chances of pathogens acquiring resistance [119, 122, 124,125,126,127] (Table 4). For example, the process of d-enantiomerisation of the peptide RR has demonstrated a 32-fold decrease in its MIC value against clinical and bacteria from MDR clinical isolates, without compromising on its haemolytic value (Table 4) [128]. In addition, the incorporation of D-amino acids might circumvent MRSA peptidases that have been observed to degrade HDPs such as LL-37 [55, 119, 122].
5.5.2 The Use of poly-N-Substituted Glycines
Poly-N-substituted glycines (peptoids) are amino acids whose side chains are bound to the amide group rather than the α-carbon [129], making them less susceptible to proteolytic degradation [129,130,131]. Although fully synthesised peptoids present their side chain in their amide group, they are still able to form certain secondary structures, such as helices, and have bactericidal effects and low haemolytic activity [101, 129, 132]. Interestingly, biodistribution and pharmacokinetics studies have shown that although peptoids have poor oral bioavailability, they have shown prolonged retention in the gastrointestinal tract compared to non-peptoid peptides [131]. Peptoids have displayed high bactericidal activity against a series of resistant bacteria. Low MIC values were identified for cyclic peptoids, for example, the peptoids constructed based on N-(3-aminopropyl)glycine (Nap), N-(2,2-diphenylethyl)glycine (Ndp) and N-(1-naphthylmethyl)glycine, showing a MIC value of 3.9, 7.8 and 7.8 μg/mL for MRSA, P. aeruginosa and A. baumannii respectively in vitro, while presenting low haematolytic activity (> 250 μg/mL) [133]. In addition, the cyclic peptoids of (NapNdp)3 have shown a lower rate of inducing resistance in S. aureus compared to MRSA [113] (see Table 4).
A study has constructed a peptoid using poly-N-substituted (Xaa Lys and Phe) halogenated at the phenyl ring of Phe, and demonstrated that chlorinated and brominated peptoids have lower MIC values for meticilin-resistant S. aureus and Staphylococcus epidermidis [101]. It is noteworthy that although the addition of iodine atoms has shown the lowest MIC values for most of the peptoids, full iodisation of them led to lower bactericidal activity, probably due to having reached a hydrophobicity threshold to exert an antimicrobial effect. Furthermore, high hydrophobicity is associated with aggregation, which could be a sign of a low safety profile [134, 135]. Halogenation can be used as a tool to improve the hydrophobicity of a peptoid, but but an optimal balance between hydrophobicity and cationic properties is necessary for optimal bacterial activity and reduced haemolytic effects [64]. Therefore, the choice and number of halogen atoms should be considered carefully to achieve the desired overall antimicrobial and haemolytic activity.
5.6 Polymerisation of HDPs
HDP polymers have two main components that contribute to their binding and penetration of the bacterial membrane, which are cationic and hydrophobic motifs on the polymer chain and network [136]. The cationic motifs facilitate the adsorption of the polymer onto the bacterial membrane, while the hydrophobic monomers aid in the insertion of the polymer into the lipid bilayer and disruption of the membrane [136]. A variety of monomers can be incorporated into the polymer chain. Cationic monomers, such as those containing ammonium or iminium groups, have been shown to exhibit low minimum inhibitory concentrations (MICs; < 4 mg/L for S. aureus) and haemolytic activity (HC50 > 500 mg/L) [136, 137]. To increase the hydrophobicity of the peptide polymer, different hydrophobic groups such as alkyl or cyclic groups have also been investigated [136]. However, the polymerisation of a high number of hydrophobic monomers has been found to increase haemolytic activity [64]. Therefore, caution should be exercised in balancing cationic and hydrophobic groups in these polymers. It should also be noted that the final HDP polymer product is not a defined peptide with a specific sequence of monomers, but a statistical approximation of a sequence due to the random nature of the polymerisation process.
A promising technique for synthesising HDP polymers is ring-opening polymerisation (ROP) [138]. This process uses small cyclic monomers as the building blocks, where the reaction is initiated by the addition of an initiator molecule, such as an alkali metal, that interacts with the cyclic monomer. The cyclic monomer undergoes nucleophilic attack by the initiator molecule, which opens up the ring. This generates a linear cationic intermediate, which can then attack other cyclic monomers, causing the ring to open and therefore perpetuating the process. This process can be repeated with different monomers to generate a polymer chain [138]. ROP has been demonstrated to be a facile and inexpensive method for synthesising hydrophobic and cationic monomers into polymers, particularly in comparison to solid-phase synthesis of peptides [138]. Using this technique, a previous study converted cationic and hydrophobic amino acids into their cyclic forms and then polymerised them to create a polymeric HDP [139]. This engineered HDP exhibited antimicrobial effects, demonstrating low MIC values (< 4 μg/mL) against S. aureus. However, its haemolytic activity was relatively high (HC50 = 16 μg/mL).
Expanding this concept, Lam et al. have synthesised a star-shaped peptide polymer composed of a PAMAM dendrimer core coupled with polymerised lysine and valine monomers, which they have termed as structurally nano-engineering antimicrobial peptide polymers (SNAPPs) [140]. This chimeric molecule has been shown to inhibit many ESKAPE pathogens that contain MDR genes, such as MRSA, colistin (also known as polymyxin E)-resistant MDR-P. aeruginosa and A. baumannii, MDR-E. faecalis and K. pneumoniae in the submicromolar concentration range, outperforming many other well-known HDPs, such as magainin II, ovispirin and melittin [140, 141]. In addition to the surprising bactericidal efficacy of SNAPPs against ESKAPE pathogens, they have displayed low haemolytic activity (HC50 > 1000 μg/mL) and low cytotoxicity against mammalian cell lines [140, 141]. An in vivo mouse assay using peritoneal injection demonstrated the effectiveness of this treatment in eliminating A. baumannii and improving survival rates in mice with bacterial infections, with a 100% survival rate compared to 20% in the control group. SNAPPs also outperformed imipenem in eliminating the infection [140].
Overall, HDP polymers are a relatively new area of research and are attracting more attention due to the promising results seen in vitro and in vivo [64, 136, 137, 140, 141]. However, some challenges of this approach may pose a hurdle in obtaining approval from health agencies. The products of the polymerisation process are diverse, which only gives a statistical approximation of the sequence, leading to difficulty in identifying the active component(s) and potential variations in composition between different batches.
6 Conclusions and Future Directions
New bactericidal and/or bacteriostatic compounds are needed to combat the growing antibiotic resistance, particularly in ESKAPE pathogens and biofilms. HDPs have been shown to be potential candidates, but their clinical application, especially those of ribosomally synthesised peptides, is limited by their suboptimal pharmacokinetics and off-site effects. It appears that modifying the structures of these peptides to enhance their clinical potential may require the use of multiple chemical techniques in combination. As demonstrated by the modification of the K4L7W peptide, the combination of cyclisation and D-amino acid substitution showed a synergistic effect, resulting in improved bactericidal activity, haemolytic activity and thermostability compared with the use of either technique alone [142]. On the other hand, peptide polymers are typically inexpensive to produce and can yield large amounts of products including the desired molecule(s), thus still facing the hurdle of uncertainty in the chemical formula. It is also important to consider the complication in production caused by multiple steps of purification, the identification of bioactive peptides and the adverse effects caused by non-active residues. In addition, the financial cost of using multiple techniques in combination might pose affordability issues for wider communities, particularly for underdeveloped countries. The development of chemically engineered HDPs has opened a new front in combating multidrug-resistant bacteria, which warrants further research.
References
O'Neill J. Tackling drug-resistant infections globally: final report and recommendations. London: Government of the United Kingdom; 2016. https://apo.org.au/node/63983.
Blair JMA, Webber MA, Baylay AJ, Ogbolu DO, Piddock LJV. Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol. 2015;13(1):42–51. https://doi.org/10.1038/nrmicro3380.
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–81. https://doi.org/10.1111/j.1469-0691.2011.03570.x.
Schultz F, Anywar G, Tang H, Chassagne F, Lyles JT, Garbe L-A, et al. Targeting ESKAPE pathogens with anti-infective medicinal plants from the Greater Mpigi region in Uganda. Sci Rep. 2020;10(1):11935. https://doi.org/10.1038/s41598-020-67572-8.
Santajit S, Indrawattana N. Mechanisms of antimicrobial resistance in ESKAPE pathogens. BioMed Res Int. 2016. https://doi.org/10.1155/2016/2475067.
Messias MCF, Mecatti GC, Priolli DG, de Oliveira CP. Plasmalogen lipids: functional mechanism and their involvement in gastrointestinal cancer. Lipids Health Dis. 2018;17(1):41. https://doi.org/10.1186/s12944-018-0685-9.
Teterycz D, Ferry T, Lew D, Stern R, Assal M, Hoffmeyer P, et al. Outcome of orthopedic implant infections due to different staphylococci. Int J Infect Dis. 2010;14(10):e913–8. https://doi.org/10.1016/j.ijid.2010.05.014.
Pendleton JN, Gorman SP, Gilmore BF. Clinical relevance of the ESKAPE pathogens. Expert Rev Anti Infect Ther. 2013;11(3):297–308. https://doi.org/10.1586/eri.13.12.
Hal SJv, Jensen SO, Vaska VL, Espedido BA, Paterson DL, Gosbell IB. Predictors of mortality in Staphylococcus aureus bacteremia. Clin Microbiol Rev. 2012;25(2):362–86. https://doi.org/10.1128/CMR.05022-11.
Safety ACo, Care QiH. AURA 2016: first Australian report on antimicrobial use and resistance in human health. Berlin: Australian Commission on Safety and Quality in Health Care; 2016.
Tzouvelekis LS, Markogiannakis A, Psichogiou M, Tassios PT, Daikos GL. Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: an evolving crisis of global dimensions. Clin Microbiol Rev. 2012;25(4):682–707. https://doi.org/10.1128/CMR.05035-11.
Oliveira DMPD, Forde BM, Kidd TJ, Harris PNA, Schembri MA, Beatson SA, et al. Antimicrobial resistance in ESKAPE pathogens. Clin Microbiol Rev. 2020;33(3):e00181-e219. https://doi.org/10.1128/CMR.00181-19.
Gedefie A, Demsis W, Ashagrie M, Kassa Y, Tesfaye M, Tilahun M, et al. Acinetobacter baumannii biofilm formation and its role in disease pathogenesis: a review. Infect Drug Resist. 2021;14:3711–9. https://doi.org/10.2147/idr.S332051.
Terreni M, Taccani M, Pregnolato M. New antibiotics for multidrug-resistant bacterial strains: latest research developments and future perspectives. Molecules. 2021. https://doi.org/10.3390/molecules26092671.
Sharma D, Misba L, Khan AU. Antibiotics versus biofilm: an emerging battleground in microbial communities. Antimicrob Resist Infect Control. 2019;8(1):76. https://doi.org/10.1186/s13756-019-0533-3.
Hancock REW, Haney EF, Gill EE. The immunology of host defence peptides: beyond antimicrobial activity. Nat Rev Immunol. 2016;16(5):321–34. https://doi.org/10.1038/nri.2016.29.
Wang G, Li X, Wang Z. APD3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2016;44(D1):D1087–93. https://doi.org/10.1093/nar/gkv1278.
Mahlapuu M, Håkansson J, Ringstad L, Björn C. Antimicrobial peptides: an emerging category of therapeutic agents. Front Cell Infect Microbiol. 2016;6:194. https://doi.org/10.3389/fcimb.2016.00194.
Savini F, Loffredo MR, Troiano C, Bobone S, Malanovic N, Eichmann TO, et al. Binding of an antimicrobial peptide to bacterial cells: Interaction with different species, strains and cellular components. Biochim Biophys Acta Biomembr. 2020;1862(8):183291. https://doi.org/10.1016/j.bbamem.2020.183291.
Brandenburg K, Heinbockel L, Correa W, Lohner K. Peptides with dual mode of action: killing bacteria and preventing endotoxin-induced sepsis. Biochim Biophys Acta Biomembr. 2016;1858(5):971–9. https://doi.org/10.1016/j.bbamem.2016.01.011.
Steiner H, Andreu D, Merrifield RB. Binding and action of cecropin and cecropin analogues: antibacterial peptides from insects. Biochim Biophys Acta Biomembr. 1988;939(2):260–6. https://doi.org/10.1016/0005-2736(88)90069-7.
Benfield AH, Henriques ST. Mode-of-action of antimicrobial peptides: membrane disruption vs. intracellular mechanisms. Front Med Technol. 2020. https://doi.org/10.3389/fmedt.2020.610997.
Wu C-L, Hsueh J-Y, Yip B-S, Chih Y-H, Peng K-L, Cheng J-W. Antimicrobial peptides display strong synergy with vancomycin against vancomycin-resistant E. faecium, S. aureus, and wild-type E. coli. Int J Mol Sci. 2020;21(13):4578.
Braffman NR, Piscotta FJ, Hauver J, Campbell EA, Link AJ, Darst SA. Structural mechanism of transcription inhibition by lasso peptides microcin J25 and capistruin. Proc Natl Acad Sci USA. 2019;116(4):1273–8. https://doi.org/10.1073/pnas.1817352116.
Laughlin TF, Ahmad Z. Inhibition of Escherichia coli ATP synthase by amphibian antimicrobial peptides. Int J Biol Macromol. 2010;46(3):367–74. https://doi.org/10.1016/j.ijbiomac.2010.01.015.
Santos P, Gordillo A, Osses L, Salazar L-M, Soto C-Y. Effect of antimicrobial peptides on ATPase activity and proton pumping in plasma membrane vesicles obtained from mycobacteria. Peptides. 2012;36(1):121–8. https://doi.org/10.1016/j.peptides.2012.04.018.
Andolina G, Bencze L-C, Zerbe K, Müller M, Steinmann J, Kocherla H, et al. A peptidomimetic antibiotic interacts with the periplasmic domain of LPTD from Pseudomonas aeruginosa. ACS Chem Biol. 2018;13(3):666–75. https://doi.org/10.1021/acschembio.7b00822.
Mahlapuu M, Håkansson J, Ringstad L, Björn C. Antimicrobial peptides: an emerging category of therapeutic agents. Front Cell Infect Microbiol. 2016. https://doi.org/10.3389/fcimb.2016.00194.
Dijksteel GS, Ulrich MMW, Middelkoop E, Boekema BKHL. Review: lessons learned from clinical trials using antimicrobial peptides (AMPs). Front Microbiol. 2021. https://doi.org/10.3389/fmicb.2021.616979.
Nguyen LT, Haney EF, Vogel HJ. The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 2011;29(9):464–72. https://doi.org/10.1016/j.tibtech.2011.05.001.
Olsen I. Biofilm-specific antibiotic tolerance and resistance. Eur J Clin Microbiol Infect Dis. 2015;34(5):877–86. https://doi.org/10.1007/s10096-015-2323-z.
Waters CM, Bassler BL. Quorum sensing: cell-to-cell communication in bacteria. Annu Rev Cell Dev Biol. 2005;21:319–46. https://doi.org/10.1146/annurev.cellbio.21.012704.131001.
Carvalho G, Forestier C, Mathias JD. Antibiotic resilience: a necessary concept to complement antibiotic resistance? Proc Biol Sci. 1916;2019(286):20192408. https://doi.org/10.1098/rspb.2019.2408.
Percival SL, Emanuel C, Cutting KF, Williams DW. Microbiology of the skin and the role of biofilms in infection. Int Wound J. 2012;9(1):14–32. https://doi.org/10.1111/j.1742-481X.2011.00836.x.
Brandwein M, Steinberg D, Meshner S. Microbial biofilms and the human skin microbiome. NPJ Biofilms Microbiomes. 2016;2:3. https://doi.org/10.1038/s41522-016-0004-z.
Kwiecinski J, Kahlmeter G, Jin T. Biofilm formation by Staphylococcus aureus isolates from skin and soft tissue infections. Curr Microbiol. 2015;70(5):698–703. https://doi.org/10.1007/s00284-014-0770-x.
Wu H, Moser C, Wang HZ, Hoiby N, Song ZJ. Strategies for combating bacterial biofilm infections. Int J Oral Sci. 2015;7(1):1–7. https://doi.org/10.1038/ijos.2014.65.
Piozzi A, Francolini I, Occhiaperti L, Di Rosa R, Ruggeri V, Donelli G. Polyurethanes loaded with antibiotics: influence of polymer-antibiotic interactions on in vitro activity against Staphylococcus epidermidis. J Chemother. 2004;16(5):446–52. https://doi.org/10.1179/joc.2004.16.5.446.
Mirzaei R, Mohammadzadeh R, Alikhani MY, Shokri Moghadam M, Karampoor S, Kazemi S, et al. The biofilm-associated bacterial infections unrelated to indwelling devices. IUBMB Life. 2020;72(7):1271–85. https://doi.org/10.1002/iub.2266.
Wu H, Moser C, Wang HZ, Høiby N, Song ZJ. Strategies for combating bacterial biofilm infections. Int J Oral Sci. 2015;7(1):1–7. https://doi.org/10.1038/ijos.2014.65.
Pletzer D, Coleman SR, Hancock RE. Anti-biofilm peptides as a new weapon in antimicrobial warfare. Curr Opin Microbiol. 2016;33:35–40. https://doi.org/10.1016/j.mib.2016.05.016.
Chung PY, Khanum R. Antimicrobial peptides as potential anti-biofilm agents against multidrug-resistant bacteria. J Microbiol Immunol Infect. 2017;50(4):405–10. https://doi.org/10.1016/j.jmii.2016.12.005.
de Breij A, Riool M, Cordfunke RA, Malanovic N, de Boer L, Koning RI, et al. The antimicrobial peptide SAAP-148 combats drug-resistant bacteria and biofilms. Sci Transl Med. 2018. https://doi.org/10.1126/scitranslmed.aan4044.
Fjell CD, Hiss JA, Hancock REW, Schneider G. Designing antimicrobial peptides: form follows function. Nat Rev Drug Disc. 2012;11(1):37–51. https://doi.org/10.1038/nrd3591.
Chen S, Zhang P, Xiao L, Liu Y, Wu K, Ni G, et al. Caerin 1.1 and 1.9 Peptides from Australian tree frog inhibit antibiotic-resistant bacteria growth in a murine skin infection model. Microbiol Spectr. 2021;9(1): e0005121. https://doi.org/10.1128/Spectrum.00051-21.
Schwarzer D, Finking R, Marahiel MA. Nonribosomal peptides: from genes to products. Nat Prod Rep. 2003;20(3):275–87. https://doi.org/10.1039/B111145K.
Rončević T, Puizina J, Tossi A. Antimicrobial peptides as anti-infective agents in pre-post-antibiotic era? Int Jo Mol Sci. 2019;20(22):5713.
Chen S, Zhang P, Xiao L, Liu Y, Wu K, Ni G, et al. Caerin 1.1 and 19. peptides from Australian tree frog inhibit antibiotic-resistant bacteria growth in a murine skin infection model. Microbiol Spectr. 2021;9(1):e00051-e121. https://doi.org/10.1128/Spectrum.00051-21.
Ni G, Chen S, Chen M, Wu J, Yang B, Yuan J, et al. Host-defense peptides caerin 1.1 and 1.9 stimulate TNF-alpha-dependent apoptotic signals in human cervical cancer HeLa cells. Front Cell Dev Biol. 2020. https://doi.org/10.3389/fcell.2020.00676.
Yang X, Li J, Chen S, Xiao L, Cao D, Wu X, et al. Preclinical pharmacokinetics, biodistribution, and acute toxicity evaluation of caerin 1.9 peptide in Sprague Dawley rats. Evid Based Complement Alternat Med. 2022. https://doi.org/10.1155/2022/9869293.
Falagas ME, Kasiakou SK. Toxicity of polymyxins: a systematic review of the evidence from old and recent studies. Crit Care. 2006;10(1):R27. https://doi.org/10.1186/cc3995.
El Shazely B, Yu G, Johnston PR, Rolff J. Resistance evolution against antimicrobial peptides in Staphylococcus aureus alters pharmacodynamics beyond the MIC. Front Microbiol. 2020. https://doi.org/10.3389/fmicb.2020.00103.
Assoni L, Milani B, Carvalho MR, Nepomuceno LN, Waz NT, Guerra MES, et al. Resistance mechanisms to antimicrobial peptides in Gram-positive bacteria. Front Microbiol. 2020. https://doi.org/10.3389/fmicb.2020.593215.
Shireen T, Singh M, Das T, Mukhopadhyay K. Differential adaptive responses of Staphylococcus aureus to in vitro selection with different antimicrobial peptides. Antimicrob Agents Chemother. 2013;57(10):5134-7. https://doi.org/10.1128/AAC.00780-13.
Sieprawska-Lupa M, Mydel P, Krawczyk K, Wójcik K, Puklo M, Lupa B, et al. Degradation of human antimicrobial peptide LL-37 by Staphylococcus aureus-derived proteinases. Antimicrob Agents Chemother. 2004;48(12):4673–9. https://doi.org/10.1128/AAC.48.12.4673-4679.2004.
Liang Y, Zhang X, Yuan Y, Bao Y, Xiong M. Role and modulation of the secondary structure of antimicrobial peptides to improve selectivity. Biomater Sci. 2020;8(24):6858–66. https://doi.org/10.1039/D0BM00801J.
Haney EF, Straus SK, Hancock REW. Reassessing the host defense peptide landscape. Front Chem. 2019. https://doi.org/10.3389/fchem.2019.00043.
Hwang PM, Vogel HJ. Structure-function relationships of antimicrobial peptides. Biochem Cell Biol. 1998;76(2–3):235–46. https://doi.org/10.1139/o98-026%M9923692.
Deslouches B, Montelaro RC, Urish KL, Di YP. Engineered cationic antimicrobial peptides (eCAPs) to combat multidrug-resistant bacteria. Pharmaceutics. 2020;12(6):501.
Dennison SR, Reddy SM, Morton LHG, Harris F, Badiani K, Phoenix DA. PEGylation enhances the antibacterial and therapeutic potential of amphibian host defence peptides. Biochim Biophys Acta Biomembr. 2022;1864(1):183806. https://doi.org/10.1016/j.bbamem.2021.183806.
Phoenix DA, Harris F, Mura M, Dennison SR. The increasing role of phosphatidylethanolamine as a lipid receptor in the action of host defence peptides. Prog Lipid Res. 2015;59:26–37. https://doi.org/10.1016/j.plipres.2015.02.003.
Lv S, Wang J, You R, Liu S, Ding Y, Hadianamrei R, et al. Highly selective performance of rationally designed antimicrobial peptides based on ponericin-W1. Biomater Sci. 2022;10(17):4848–65. https://doi.org/10.1039/d2bm00744d.
Timmons PB, O’Flynn D, Conlon JM, Hewage CM. Structural and positional studies of the antimicrobial peptide brevinin-1BYa in membrane-mimetic environments. J Pept Sci. 2019;25(11): e3208. https://doi.org/10.1002/psc.3208.
Etayash H, Hancock REW. Host defense peptide-mimicking polymers and polymeric-brush-tethered host defense peptides: recent developments, limitations, and potential success. Pharmaceutics. 2021;13(11):1820.
Haney EF, Hancock REW. Peptide design for antimicrobial and immunomodulatory applications. Peptide Sci. 2013;100(6):572–83. https://doi.org/10.1002/bip.22250.
Dathe M, Nikolenko H, Meyer J, Beyermann M, Bienert M. Optimization of the antimicrobial activity of magainin peptides by modification of charge. FEBS Lett. 2001;501(2–3):146–50. https://doi.org/10.1016/S0014-5793(01)02648-5.
Chan DI, Prenner EJ, Vogel HJ. Tryptophan- and arginine-rich antimicrobial peptides: structures and mechanisms of action. Biochim Biophys Acta. 2006;1758(9):1184–202. https://doi.org/10.1016/j.bbamem.2006.04.006.
Jiang Z, Vasil AI, Gera L, Vasil ML, Hodges RS. Rational design of α-helical antimicrobial peptides to target Gram-negative pathogens, Acinetobacter baumannii and Pseudomonas aeruginosa: utilization of charge, ‘specificity determinants’, total hydrophobicity, hydrophobe type and location as design parameters to improve the therapeutic ratio. Chem Biol Drug Des. 2011;77(4):225–40. https://doi.org/10.1111/j.1747-0285.2011.01086.x.
Jiang Z, Vasil AI, Vasil ML, Hodges RS. “Specificity determinants” improve therapeutic indices of two antimicrobial peptides Piscidin 1 and Dermaseptin S4 against the Gram-negative pathogens Acinetobacter baumannii and Pseudomonas aeruginosa. Pharmaceuticals. 2014;7(4):366–91.
Zhang S-K, Song J-W, Gong F, Li S-B, Chang H-Y, Xie H-M, et al. Design of an α-helical antimicrobial peptide with improved cell-selective and potent anti-biofilm activity. Sci Rep. 2016;6(1):27394. https://doi.org/10.1038/srep27394.
Papo N, Shai Y. Can we predict biological activity of antimicrobial peptides from their interactions with model phospholipid membranes? Peptides. 2003;24(11):1693–703. https://doi.org/10.1016/j.peptides.2003.09.013.
Hollmann A, Martínez M, Noguera ME, Augusto MT, Disalvo A, Santos NC, et al. Role of amphipathicity and hydrophobicity in the balance between hemolysis and peptide–membrane interactions of three related antimicrobial peptides. Colloids Surf B Biointerfaces. 2016;141:528–36. https://doi.org/10.1016/j.colsurfb.2016.02.003.
Ma Z, Yang J, Han J, Gao L, Liu H, Lu Z, et al. Insights into the antimicrobial activity and cytotoxicity of engineered α-helical peptide amphiphiles. J Med Chem. 2016;59(24):10946–62. https://doi.org/10.1021/acs.jmedchem.6b00922.
Akbari R, Hakemi Vala M, Hashemi A, Aghazadeh H, Sabatier J-M, Pooshang BK. Action mechanism of melittin-derived antimicrobial peptides, MDP1 and MDP2, de novo designed against multidrug resistant bacteria. Amino Acids. 2018;50(9):1231–43. https://doi.org/10.1007/s00726-018-2596-5.
Jin L, Bai X, Luan N, Yao H, Zhang Z, Liu W, et al. A designed tryptophan- and lysine/arginine-rich antimicrobial peptide with therapeutic potential for clinical antibiotic-resistant Candida albicans Vaginitis. J Med Chem. 2016;59(5):1791–9. https://doi.org/10.1021/acs.jmedchem.5b01264.
Deslouches B, Steckbeck JD, Craigo JK, Doi Y, Mietzner TA, Montelaro RC. Rational design of engineered cationic antimicrobial peptides consisting exclusively of arginine and tryptophan, and their activity against multidrug-resistant pathogens. Antimicrob Agents Chemother. 2013;57(6):2511–21. https://doi.org/10.1128/AAC.02218-12.
Hermanson GT. Chapter 18—PEGylation and synthetic polymer modification. In: Hermanson GT, editor. Bioconjugate techniques. 3rd ed. Boston: Academic Press; 2013. p. 787–838.
Turecek PL, Bossard MJ, Schoetens F, Ivens IA. PEGylation of biopharmaceuticals: a review of chemistry and nonclinical safety information of approved drugs. J Pharm Sci. 2016;105(2):460–75. https://doi.org/10.1016/j.xphs.2015.11.015.
Veronese FM, Mero A. The impact of PEGylation on biological therapies. BioDrugs. 2008;22(5):315–29. https://doi.org/10.2165/00063030-200822050-00004.
Morgenstern J, Baumann P, Brunner C, Hubbuch J. Effect of PEG molecular weight and PEGylation degree on the physical stability of PEGylated lysozyme. Int J Pharm. 2017;519(1):408–17. https://doi.org/10.1016/j.ijpharm.2017.01.040.
Du B, Jiang X, Huang Y, Li S, Lin JC, Yu M, et al. Tailoring kidney transport of organic dyes with low-molecular-weight PEGylation. Bioconjug Chem. 2020;31(2):241–7. https://doi.org/10.1021/acs.bioconjchem.9b00707.
Alvarez HM, So OY, Hsieh S, Shinsky-Bjorde N, Ma H, Song Y, et al. Effects of PEGylation and immune complex formation on the pharmacokinetics and biodistribution of recombinant interleukin 10 in mice. Drug Metab Dispos. 2012;40(2):360–73. https://doi.org/10.1124/dmd.111.042531.
Cai Y, Zhang Z, Fan K, Zhang J, Shen W, Li M, et al. Pharmacokinetics, tissue distribution, excretion, and antiviral activity of pegylated recombinant human consensus interferon-α variant in monkeys, rats and guinea pigs. Regul Pept. 2012;173(1–3):74–81. https://doi.org/10.1016/j.regpep.2011.09.008.
Zheng J-C, Lei N, He Q-C, Hu W, Jin J-G, Meng Y, et al. PEGylation is effective in reducing immunogenicity, immunotoxicity, and hepatotoxicity of α-momorcharin in vivo. Immunopharmacol Immunotox. 2012;34(5):866–73. https://doi.org/10.3109/08923973.2012.666979.
de Bourayne M, Meunier S, Bitoun S, Correia E, Mariette X, Nozach H, et al. Pegylation reduces the uptake of certolizumab pegol by dendritic cells and epitope presentation to T-Cells. Front Immunol. 2022. https://doi.org/10.3389/fimmu.2022.808606.
Gong Y, Andina D, Nahar S, Leroux JC, Gauthier MA. Releasable and traceless PEGylation of arginine-rich antimicrobial peptides. Chem Sci. 2017;8(5):4082–6. https://doi.org/10.1039/C7SC00770A.
Fang Y, Xue J, Gao S, Lu A, Yang D, Jiang H, et al. Cleavable PEGylation: a strategy for overcoming the “PEG dilemma” in efficient drug delivery. Drug Deliv. 2017;24(2):22–32. https://doi.org/10.1080/10717544.2017.1388451.
Han E, Lee H. Effects of PEGylation on the binding interaction of Magainin 2 and Tachyplesin I with lipid bilayer surface. Langmuir. 2013;29(46):14214–21. https://doi.org/10.1021/la4036985.
Kusano H, Akiba J, Ogasawara S, Sanada S, Yasumoto M, Nakayama M, et al. Pegylated interferon-α2a inhibits proliferation of human liver cancer cells in vitro and in vivo. PLoS ONE. 2013;8(12):e83195. https://doi.org/10.1371/journal.pone.0083195.
Podobnik B, Helk B, Smilović V, Škrajnar Š, Fidler K, Jevševar S, et al. Conjugation of polyPEG to interferon alpha extends serum half-life while maintaining low viscosity of the conjugate. Bioconjug Chem. 2015;26(3):452–9. https://doi.org/10.1021/bc500523t.
Cavallazzi Sebold B, Ni G, Li J, Li H, Liu X, Wang T. PEGylated IL-10: clinical development in cancer immunotherapy, where to go? Curr Oncol Rep. 2022. https://doi.org/10.1007/s11912-022-01355-4.
Singh S, Papareddy P, Mörgelin M, Schmidtchen A, Malmsten M. Effects of PEGylation on membrane and lipopolysaccharide interactions of host defense peptides. Biomacromol. 2014;15(4):1337–45. https://doi.org/10.1021/bm401884e.
Imura Y, Nishida M, Ogawa Y, Takakura Y, Matsuzaki K. Action mechanism of tachyplesin I and effects of PEGylation. Biochim Biophys Acta Biomembr. 2007;1768(5):1160–9. https://doi.org/10.1016/j.bbamem.2007.01.005.
Morris CJ, Beck K, Fox MA, Ulaeto D, Clark GC, Gumbleton M. Pegylation of antimicrobial peptides maintains the active peptide conformation, model membrane interactions, and antimicrobial activity while improving lung tissue biocompatibility following airway delivery. Antimicrob Agents Chemother. 2012;56(6):3298–308. https://doi.org/10.1128/AAC.06335-11.
da Silva APG, Unks D, Lyu S-c, Ma J, Zbozien-Pacamaj R, Chen X, et al. In vitro and in vivo antimicrobial activity of granulysin-derived peptides against Vibrio cholerae. J Antimicrob Chemother. 2008;61(5):1103–9. https://doi.org/10.1093/jac/dkn058.
Yu W, Wang J, Wang Z, Li L, Li W, Song J, et al. PEGylation of the antimicrobial peptide PG-1: a link between propensity for nanostructuring and capacity of the antitrypsin hydrolytic Ability. J Med Chem. 2021;64(14):10469–81. https://doi.org/10.1021/acs.jmedchem.1c00879.
Kaur N, Dilawari R, Kaur A, Sahni G, Rishi P. Recombinant expression, purification and PEGylation of Paneth cell peptide (cryptdin-2) with value added attributes against Staphylococcus aureus. Sci Rep. 2020;10(1):12164. https://doi.org/10.1038/s41598-020-69039-2.
Brunetti J, Falciani C, Roscia G, Pollini S, Bindi S, Scali S, et al. In vitro and in vivo efficacy, toxicity, bio-distribution and resistance selection of a novel antibacterial drug candidate. Sci Rep. 2016;6:26077. https://doi.org/10.1038/srep26077.
Carrat F, Bani-Sadr F, Pol S, Rosenthal E, Lunel-Fabiani F, Benzekri A, et al. Pegylated interferon alfa-2b vs standard interferon alfa-2b, plus ribavirin, for chronic hepatitis C in HIV-infected patients: a randomized controlled trial. JAMA. 2004;292(23):2839–48. https://doi.org/10.1001/jama.292.23.2839.
Gentry CL, Egleton RD, Gillespie T, Abbruscato TJ, Bechowski HB, Hruby VJ, et al. The effect of halogenation on blood–brain barrier permeability of a novel peptide drug. Peptides. 1999;20(10):1229–38. https://doi.org/10.1016/S0196-9781(99)00127-8.
Molchanova N, Nielsen JE, Sørensen KB, Prabhala BK, Hansen PR, Lund R, et al. Halogenation as a tool to tune antimicrobial activity of peptoids. Sci Rep. 2020;10(1):14805. https://doi.org/10.1038/s41598-020-71771-8.
Paulsen MH, Karlsen EA, Ausbacher D, Anderssen T, Bayer A, Ochtrop P, et al. An amphipathic cyclic tetrapeptide scaffold containing halogenated β2,2-amino acids with activity against multiresistant bacteria. J Peptide Sci. 2018;24(10): e3117. https://doi.org/10.1002/psc.3117.
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings1PII of original article. Adv Drug Deliv Rev. 2001;46(1):3–26. https://doi.org/10.1016/S0169-409X(00)00129-0.
Empting M. CHAPTER 1 an introduction to cyclic peptides. In: Cyclic peptides: from bioorganic synthesis to applications. The Royal Society of Chemistry; 2018. p. 1-14.
Haines DJ, Swan CHJ, Green JRB, Woodley JF. Mucosal peptide hydrolase and brush-border marker enzyme activities in three regions of the small intestine of rats with experimental uraemia. Clin Sci. 1990;79(6):663–8. https://doi.org/10.1042/cs0790663.
White CJ, Yudin AK. Contemporary strategies for peptide macrocyclization. Nat Chem. 2011;3(7):509–24. https://doi.org/10.1038/nchem.1062.
Wenzel M, Rautenbach M, Vosloo JA, Siersma T, Aisenbrey CHM, Zaitseva E, et al. The multifaceted antibacterial mechanisms of the pioneering peptide antibiotics tyrocidine and gramicidin S. MBio. 2018;9(5):e00802-e818. https://doi.org/10.1128/mBio.00802-18.
Avedissian SN, Liu J, Rhodes NJ, Lee A, Pais GM, Hauser AR, et al. A review of the clinical pharmacokinetics of polymyxin B. Antibiotics. 2019. https://doi.org/10.3390/antibiotics8010031.
Micklefield J. Daptomycin structure and mechanism of action revealed. Chem Biol. 2004;11(7):887–8. https://doi.org/10.1016/j.chembiol.2004.07.001.
Marsault E, Peterson ML. Macrocycles are great cycles: applications, opportunities, and challenges of synthetic macrocycles in drug discovery. J Med Chem. 2011;54(7):1961–2004. https://doi.org/10.1021/jm1012374.
Joo SH. Cyclic peptides as therapeutic agents and biochemical tools. Biomol Ther. 2012;20(1):19–26. https://doi.org/10.4062/biomolther.2012.20.1.019.
Shin SBY, Yoo B, Todaro LJ, Kirshenbaum K. Cyclic peptoids. J Am Chem Soc. 2007;129(11):3218–25. https://doi.org/10.1021/ja066960o.
Huang ML, Shin SB, Benson MA, Torres VJ, Kirshenbaum K. A comparison of linear and cyclic peptoid oligomers as potent antimicrobial agents. ChemMedChem. 2012;7(1):114–22. https://doi.org/10.1002/cmdc.201100358.
Unger T, Oren Z, Shai Y. The effect of cyclization of magainin 2 and melittin analogues on structure, function, and model membrane interactions: implication to their mode of action. Biochemistry. 2001;40(21):6388–97. https://doi.org/10.1021/bi0026066.
Jia X, Kwon S, Wang CA, Huang YH, Chan LY, Tan CC, et al. Semienzymatic cyclization of disulfide-rich peptides using Sortase A. J Biol Chem. 2014;289(10):6627–38. https://doi.org/10.1074/jbc.M113.539262.
Colgrave ML, Craik DJ. Thermal, chemical, and enzymatic stability of the cyclotide kalata b1: the importance of the cyclic cystine knot. Biochemistry. 2004;43(20):5965–75. https://doi.org/10.1021/bi049711q.
Nguyen LT, Chau JK, Perry NA, de Boer L, Zaat SAJ, Vogel HJ. Serum stabilities of short tryptophan- and arginine-rich antimicrobial peptide analogs. PLoS ONE. 2010;5(9):e12684. https://doi.org/10.1371/journal.pone.0012684.
Du Y, Li L, Zheng Y, Liu J, Gong J, Qiu Z, et al. Incorporation of non-canonical amino acids into antimicrobial peptides: advances, challenges, and perspectives. Appl Environ Microbiol. 2022. https://doi.org/10.1128/aem.01617-22.
Kapil S, Sharma V. d-Amino acids in antimicrobial peptides: a potential approach to treat and combat antimicrobial resistance. Can J Microbiol. 2021;67(2):119–37. https://doi.org/10.1139/cjm-2020-0142%M32783775.
Etayash H, Hancock REW. Host defense peptide-mimicking polymers and polymeric-brush-tethered host defense peptides: recent developments, limitations, and potential success. Pharmaceutics. 2021. https://doi.org/10.3390/pharmaceutics13111820.
Bruice PY. Organic chemistry. 7 International. Prentice Hall; 2014.
Cava F, Lam H, de Pedro MA, Waldor MK. Emerging knowledge of regulatory roles of D-amino acids in bacteria. Cell Mol Life Sci. 2011;68(5):817–31. https://doi.org/10.1007/s00018-010-0571-8.
Peñalva MA, Rowlands RT, Turner G. The optimization of penicillin biosynthesis in fungi. Trends Biotechnol. 1998;16(11):483–9. https://doi.org/10.1016/S0167-7799(98)01229-3.
Lee J-K, Park Y. All d-Lysine Analogues of the antimicrobial peptide HPA3NT3-A2 increased serum stability and without drug resistance. Int J Mol Sci. 2020;21(16):5632.
Zhong C, Zhu N, Zhu Y, Liu T, Gou S, Xie J, et al. Antimicrobial peptides conjugated with fatty acids on the side chain of D-amino acid promises antimicrobial potency against multidrug-resistant bacteria. Eur J Pharm Sci. 2020;141:105123. https://doi.org/10.1016/j.ejps.2019.105123.
Loffredo MR, Ghosh A, Harmouche N, Casciaro B, Luca V, Bortolotti A, et al. Membrane perturbing activities and structural properties of the frog-skin derived peptide Esculentin-1a(1–21)NH(2) and its diastereomer Esc(1–21)-1c: correlation with their antipseudomonal and cytotoxic activity. Biochim Biophys Acta Biomembr. 2017;1859(12):2327–39. https://doi.org/10.1016/j.bbamem.2017.09.009.
Li H, Anuwongcharoen N, Malik AA, Prachayasittikul V, Wikberg JE, Nantasenamat C. Roles of d-amino acids on the bioactivity of host defense peptides. Int J Mol Sci. 2016. https://doi.org/10.3390/ijms17071023.
Mohamed MF, Brezden A, Mohammad H, Chmielewski J, Seleem MN. A short D-enantiomeric antimicrobial peptide with potent immunomodulatory and antibiofilm activity against multidrug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii. Sci Rep. 2017;7(1):6953. https://doi.org/10.1038/s41598-017-07440-0.
Chongsiriwatana NP, Patch JA, Czyzewski AM, Dohm MT, Ivankin A, Gidalevitz D, et al. Peptoids that mimic the structure, function, and mechanism of helical antimicrobial peptides. Proc Natl Acad Sci USA. 2008;105(8):2794–9. https://doi.org/10.1073/pnas.0708254105.
Miller SM, Simon RJ, Ng S, Zuckermann RN, Kerr JM, Moos WH. Comparison of the proteolytic susceptibilities of homologous L-amino acid, D-amino acid, and N-substituted glycine peptide and peptoid oligomers. Drug Develop Res. 1995;35(1):20–32. https://doi.org/10.1002/ddr.430350105.
Seo J, Ren G, Liu H, Miao Z, Park M, Wang Y, et al. In vivo biodistribution and small animal PET of 64Cu-labeled antimicrobial peptoids. Bioconjug Chem. 2012;23(5):1069–79. https://doi.org/10.1021/bc300091d.
Bicker KL, Cobb SL. Recent advances in the development of anti-infective peptoids. Chem Commun. 2020;56(76):11158–68. https://doi.org/10.1039/D0CC04704J.
Andreev K, Martynowycz MW, Ivankin A, Huang ML, Kuzmenko I, Meron M, et al. Cyclization improves membrane permeation by antimicrobial peptoids. Langmuir. 2016;32(48):12905–13. https://doi.org/10.1021/acs.langmuir.6b03477.
Jiang L, Cao S, Cheung PP, Zheng X, Leung CWT, Peng Q, et al. Real-time monitoring of hydrophobic aggregation reveals a critical role of cooperativity in hydrophobic effect. Nat Commun. 2017;8:15639. https://doi.org/10.1038/ncomms15639.
Ratanji KD, Derrick JP, Dearman RJ, Kimber I. Immunogenicity of therapeutic proteins: influence of aggregation. J Immunotoxicol. 2014;11(2):99–109. https://doi.org/10.3109/1547691x.2013.821564.
Yang Y, Cai Z, Huang Z, Tang X, Zhang X. Antimicrobial cationic polymers: from structural design to functional control. Polymer J. 2018;50(1):33–44. https://doi.org/10.1038/pj.2017.72.
Ng VWL, Tan JPK, Leong J, Voo ZX, Hedrick JL, Yang YY. Antimicrobial polycarbonates: investigating the impact of nitrogen-containing heterocycles as quaternizing agents. Macromolecules. 2014;47(4):1285–91. https://doi.org/10.1021/ma402641p.
Zhou C, Qi X, Li P, Chen WN, Mouad L, Chang MW, et al. High potency and broad-spectrum antimicrobial peptides synthesized via ring-opening polymerization of α-aminoacid-N-carboxyanhydrides. Biomacromol. 2010;11(1):60–7. https://doi.org/10.1021/bm900896h.
Song A, Walker SG, Parker KA, Sampson NS. Antibacterial studies of cationic polymers with alternating, random, and uniform backbones. ACS Chem Biol. 2011;6(6):590–9. https://doi.org/10.1021/cb100413w.
Lam SJ, O’Brien-Simpson NM, Pantarat N, Sulistio A, Wong EHH, Chen Y-Y, et al. Combating multidrug-resistant Gram-negative bacteria with structurally nanoengineered antimicrobial peptide polymers. Nat Microbiol. 2016;1(11):16162. https://doi.org/10.1038/nmicrobiol.2016.162.
Li W, Hadjigol S, Mazo AR, Holden J, Lenzo J, Shirbin SJ, et al. Star-peptide polymers are multi-drug-resistant Gram-positive bacteria killers. ACS Appl Mater Interfaces. 2022;14(22):25025–41. https://doi.org/10.1021/acsami.1c23734.
Oren Z, Shai Y. Cyclization of a cytolytic amphipathic α-helical peptide and its diastereomer: effect on structure, interaction with model membranes, and biological function. Biochemistry. 2000;39(20):6103–14. https://doi.org/10.1021/bi992408i.
Fox MA, Thwaite JE, Ulaeto DO, Atkins TP, Atkins HS. Design and characterization of novel hybrid antimicrobial peptides based on cecropin A, LL-37 and magainin II. Peptides. 2012;33(2):197–205. https://doi.org/10.1016/j.peptides.2012.01.013.
Li T, Yang N, Teng D, Mao R, Hao Y, Wang X, et al. C-terminal mini-PEGylation of a marine peptide N6 had potent antibacterial and anti-inflammatory properties against Escherichia coli and Salmonella strains in vitro and in vivo. BMC Microbiol. 2022;22(1):128. https://doi.org/10.1186/s12866-022-02534-w.
Etayash H, Pletzer D, Kumar P, Straus SK, Hancock REW. Cyclic derivative of host-defense peptide IDR-1018 improves proteolytic stability, suppresses inflammation, and enhances in vivo activity. J Med Chem. 2020;63(17):9228–36. https://doi.org/10.1021/acs.jmedchem.0c00303.
Vernen F, Harvey PJ, Dias SA, Veiga AS, Huang YH, Craik DJ, et al. Characterization of tachyplesin peptides and their cyclized analogues to improve antimicrobial and anticancer properties. Int J Mol Sci. 2019. https://doi.org/10.3390/ijms20174184.
Qiu S, Zhu R, Zhao Y, An X, Jia F, Peng J, et al. Antimicrobial activity and stability of protonectin with D-amino acid substitutions. J Pept Sci. 2017;23(5):392–402. https://doi.org/10.1002/psc.2989.
Li Y, Liu T, Liu Y, Tan Z, Ju Y, Yang Y, et al. Antimicrobial activity, membrane interaction and stability of the D-amino acid substituted analogs of antimicrobial peptide W3R6. J Photochem Photobiol B Biol. 2019;200:111645. https://doi.org/10.1016/j.jphotobiol.2019.111645.
Jia F, Wang J, Peng J, Zhao P, Kong Z, Wang K, et al. D-amino acid substitution enhances the stability of antimicrobial peptide polybia-CP. Acta Biochim Biophys Sin. 2017;49(10):916–25. https://doi.org/10.1093/abbs/gmx091.
Acknowledgments
We thank Professor Abigail Elizur for her valuable advice and support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This study was supported by Deng Feng project of Foshan First People’s Hospital (2019A008), National Science Foundation of Guangdong province (2020A1515010855) and National Natural Science Foundation of China (31971355).
Conflict of Interest
The authors declare no conflict of interest.
Patient Consent to Participate/Publish
Not applicable.
Code Availability
Not applicable.
Availability of Data and Material
All relevant data and material are provided in the article.
Ethics Approval
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author Contributions
Conceptualisation, B.C., J.L., X.L. and T.W. B.L.; methodology, B.C. and T.W. Validation, B.C., X.L. and T.W. Formal analysis, B.C. Investigation, B.C. Resources, B.C., X.L. and T.W. Writing – original draft preparation, B.C., X.L. and T.W. Writing – review and editing, B.C., Q.F., H.L., G.N., X.L. and T.W. Visualisation, B.C. and T.W. Supervision, G.N. and T.W. Project administration, G.N., X.L. and T.W. Funding acquisition, G.N., X.L. and T.W. All authors have read and agreed to the published version of the manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Cavallazzi Sebold, B., Li, J., Ni, G. et al. Going Beyond Host Defence Peptides: Horizons of Chemically Engineered Peptides for Multidrug-Resistant Bacteria. BioDrugs 37, 607–623 (2023). https://doi.org/10.1007/s40259-023-00608-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40259-023-00608-3