Abstract
The present study investigated the prevalence of pathogenic organisms (Salmonella spp, Vibrio cholerae, and Shigella spp) and their correlation to the abundance of faecal indicator organisms in water and riverbed sediments in the Apies River, South Africa. In all, 558 water and sediment samples were collected from 10 sites in the river (May 2013–February 2014) and analysed through culture and molecular (real-time PCR) techniques. Concentrations of faecal indicator organisms in sediments reached 1.39 × 105 (±standard deviation) CFU/100 mL. All three pathogens were detected in water and sediments. Pathogens were mostly detected in sediments at sites influenced either by wastewater treatment works or by informal settlements. During the wet and dry seasons (water column), a strong positive correlation was observed between E. coli and all pathogens; C. perfringens only correlated with V. cholerae. Within sediments, strong positive correlations were only observed between E. coli and Salmonella spp, E. coli and V. cholerae (dry season); E. coli and V. cholerae and E. coli and Shigella spp (wet season). No correlation was observed between sediments C. perfringens counts and all the pathogens. Thus, sediments of the Apies River harbour pathogenic organisms. Correlation between E. coli and pathogenic organisms in the sediments suggests that E. coli could also be an indicator of pathogens’ presence. However, the lack of a correlation between E. coli and some pathogens in sediments and between C. perfringens and all the pathogens highlights the need to investigate for more indicators of pathogens’ presence in this complex matrix.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The world’s water ecosystem continues to suffer severe deterioration due to microbial pollution from urban run-off (Crowther et al. 2002; Tyrrel and Quinton 2003; Reeves et al. 2004; Signor 2005) agricultural farms (Monis and Thompson 2003; Carey et al. 2004) or more localised sources like wastewater treatment works (WWTWs) (Abraham 2011; Teklehaimanot et al. 2014). Irrespective of the source, once in the aquatic environment, the survival of these microbial pollutants depends on a number of factors, amongst which is the attachment to suspended sediment particles (Gao et al. 2011). The attached bacteria, together with the sediment particle, may then become bigger, resulting in heavier complexes that can potentially settle out onto the riverbed. Several studies carried out on the role played by riverbed sediments as reservoirs of indicator bacteria, including human pathogens, have reported higher microbial counts in the sediments than in the water column (Jamieson et al. 2004; Characklis et al. 2005; Fries et al. 2008).
Chase et al. (2012) reported that levels of faecal indicator bacteria (FIB) in sediments could be up to 100-fold higher when compared to the water column concentration. Bacteria in riverbed sediments can be shielded from the killing effect of UV light coming from the sun (Craig et al. 2004; Stapleton et al. 2007; Koirala et al. 2008) by suspended particles. Sediments have also been found to enhance bacterial survival by providing a hiding place for the bacteria against predation from protozoa (Decamp and Warren 2000; Jamieson et al. 2005a, b). Furthermore, sediments often contain high concentrations of nutrients and other soluble organic matter essential for bacterial growth (Jamieson et al. 2005a, b; Garzio-hadzick et al. 2010). Apart from traditional indicator bacteria like E. coli, pathogenic bacteria, such as Vibrio cholerae, Salmonella spp and Shigella spp have also been reported in riverbed sediments (Santhiya et al. 2011; Vignesh et al. 2014).
Natural turbulences and/or human recreational activities could lead to the resuspension of microorganisms found in the bottom sediments (Craig et al. 2004; Budillon et al. 2006; Pandey et al. 2011). Several studies have reported higher levels of FIB in the water column of water bodies that have been associated with resuspension of riverbed sediments following disturbance of the bed sediments (Pianetti et al. 2004; Ibekwe and Papiernik 2010; Gonzalez et al. 2012) suggesting riverbed sediments could serve as an important source of microorganisms within the water column (Kinzelman and McLellan 2009; Korajkic et al. 2011). The resuspension from sediments could therefore represent a potential health hazard for populations using such untreated water for recreational purposes (Gao et al. 2011) as well as for other household activities—especially where treated pipe-borne water is not available.
South Africa’s water resources are negatively impacted by low average annual precipitation. Uneven distribution of surface and groundwater as a result of climate and geography makes the country water scarce (Molobela and Sinha 2011). The country entirely depends on surface water resources for most of its urban, industrial and irrigation water requirements (Basson 2011). Aquatic ecosystems and water resources on which most rural communities depend for domestic, recreational and agricultural uses are still being subject to heavy microbial pollution from different sources resulting in severe environmental, health and economic damage (Basson 2011). Several studies carried out in South Africa have focused on the quality of surface water (Kinge and Mbewe 2010; Chigor and Okoh 2012; Seanego and Moyo 2013; Sibanda and Okoh 2013; Teklehaimanot et al. 2014), and there is little information on the microbial quality of riverbed sediments. Like many countries in the world, South Africa’s aquatic ecosystem monitoring bodies have not taken microbial sediment quality into consideration in the development and modification of their water quality guidelines (Republic of South Africa, Department of Environmental Affairs 2012). It has been shown recently that the sediments of the Apies River, a widely used river in the Gauteng Province of South Africa, were heavily polluted with E. coli (Abia et al. 2015b). The sediments were found to contain as high as 105 times higher concentrations of E. coli than the water column. However, given the complex nature of the sediment matrix within the riverbed, choosing an appropriate indicator of faecal pollution, and possibly of pathogenic organisms (POs), could be an essential step for the successful monitoring of the microbial quality of bed sediments within the aquatic ecosystem. Based on this, another recent study carried out on the Apies River suggested that E. coli alone, though recognised as a good indicator of microbial quality within water bodies, was not sufficient to predict the microbial quality of sediments within these ecosystems (Abia et al. 2015b). The authors of this study concluded by recommending that in order to have a better picture of both recent and past effects of microbial pollution within riverbed sediments, it was necessary to check for the presence of both E. coli and other potential indicators of faecal pollution such as Clostridium perfringens within the sediments (Abia et al. 2015a). Despite these findings regarding the microbial quality (presence of E. coli and C. perfringens) of sediments in South African water bodies, there is still no data on the possible presence of POs within the riverbed sediments. Such information could improve the understanding of the possible contribution of sediments to the overall microbial load of water catchments thus providing the necessary preliminary evidence needed to motivate the need for increased allocation of funds for water resource protection initiatives within developing countries.
With the main aim of filling up this paucity of information regarding microbial sediment quality, the present study was carried out with objectives: (1) to investigate the extent to which riverbed sediments and water in the Apies River are contaminated with pathogenic microorganisms (Salmonella spp, V. cholerae and Shigella spp) and (2) to investigate any correlation between the abundance of indicator organisms (E. coli and C. perfringens) and the presence of the pathogens in the sediments and the overlaying water. The study was conducted from May to August 2013 (dry season) and January to February 2014 (wet season). The river is situated in the Gauteng Province of South Africa.
Materials and methods
Study site
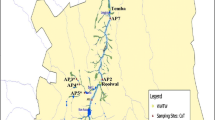
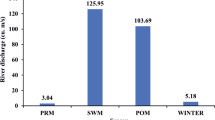
The present study was conducted on the Apies River, which is one of the main watercourses within the Gauteng Province of South Africa (Fig. 1). The study site and characteristics of the river as well as the various land uses around the river have been described in previous studies (Abia et al. 2015a, b). During the dry season, most parts of the river are fully accessible by communities that use its water for activities such as laundry and bathing. During the wet season, the river usually overflows its banks, especially after heavy rainfall (attached picture, Fig. 1). During both seasons, the water and sediments of Apies River are sources of spiritual cleansing and empowerment to many traditional healers and other religious groups. Important to note is the presence of four WWTWs—Daspoort, Rooiwal, Babelegi and Temba along the river that discharge their effluents directly into the river system. These WWTWs account for about 80 % of the total river discharge during the dry season (Venter 2007). Also, parts of the river’s course pass through informal settlements which completely lack sanitary facilities, thus making the river the only point of waste disposal in these areas.
Sample collection and processing
Sampling was conducted in the dry (May to August 2013) and the wet (January to February 2014) seasons. A total of 10 sampling sites were chosen for the study. In order to identify the possible sources of the pollution observed within the Apies River, study sites were selected based on the various land uses around them. This was done following an initial field visit to the river. Sites DAS, AP1, AP2, AP6, AP7, AP8 and AP9 were all located on the main Apies River. Sites AP3, AP4 and AP5 are tributaries to the Apies River. Water and sediment samples were collected during 14 sampling rounds in each season and analysed for E. coli, C. perfringens, V. cholerae, Salmonella spp and Shigella spp, using culture and molecular techniques. Site AP9 was only sampled during 13 rounds as the sampling site was not accessible on one of the rounds due to heavy rains that caused flooding of the river beyond it banks. Water samples were collected using clean sterile 1 L plastic containers following standard procedures (APHA/AWWA/WEF 2001). A sterile polypropylene autoclavable scooper was used to scoop the sediments samples from the top 5 cm of the riverbed. All samples were collected approximately 1 m (or further when possible) away from the river bank. This was the approximate distance the users would go during activities such as laundry or bathing. The collected sediments were transferred into sterile 100 mL polypropylene containers with lids. Both water and sediments were collected in triplicates at each sampling point and transported to the laboratory in a cooler box containing ice packs. All samples were processed and analysed within 6 h upon arrival to the laboratory. Prior to analysis, microorganisms were separated from the sediment samples using the water displacement approach as previously described (Abia et al. 2015c). The supernatant from resuspended sediment samples was then used for the enumeration of the FIOs and for enrichment in appropriate media depending on the pathogen of interest. For the water samples, after inverting the sampling bottle several times to allow for proper mixing of the water, the samples were used directly for culture and enrichment in appropriate media.
Enumeration of indicator organisms
Escherichia coli was enumerated using the Colilert® 18/Quanti-tray® 2000 defined substrate method and confirmed by real-time PCR as previously described (Abia et al. 2015b). C. perfringens was enumerated using the pour plate technique on D-cycloserine supplemented TSC agar and confirmed molecularly as described by Abia et al. (2015a).
Identification of pathogens
Sample enrichment and DNA extraction
Equal volume (50 mL) of water (50 mL of supernatant in the case of the sediment samples) was added to equal volume of double-strength broth (Environment Agency 2002). Selenite broth was used for Salmonella spp (Chitanand et al. 2008), peptone water for Shigella spp (Theron et al. 2001) and alkaline peptone water for V. cholerae (Nandi et al. 2000; Goel et al. 2005; Akoachere et al. 2013) and incubated at 35.0 ± 0.5 °C for 24 h. All culture media were purchased from Merck (South Africa). After incubation, 1 mL of the overnight broth culture was transferred into a centrifuge tube and spun at 13,000 g for 3 min. DNA was extracted from the harvested cells using the Instagene™ Matrix (Bio-Rad Laboratories, South Africa) following the manufacturer’s instruction. The supernatant from the tubes was then used as a source of template DNA for the real-time PCR reactions.
Primers and real-time PCR conditions
Genes targeted for the identification of the various pathogens were the outer membrane protein (ompW) and the cholera toxin (ctxAB) genes for V. cholerae, the invasive gene A (invA) for Salmonella spp and the invasive plasmid antigen H (ipaH) gene for Shigella spp (Table 1).
PCR reactions were carried out on a Corbett Life Science Rotor-Gene 6000 Cycler (Qiagen, Hilden, Germany). Amplification reactions were performed in a total volume of 20 µl consisting of 10 µl of a 2× SensiFAST™ HRM Mix (final concentration, 1×) (Bioline GmbH, Germany), 1 µl (final concentration, 1 µM) of each primer (Forward and Reverse) 5 µl of DNA template and 3 µl of nuclease-free water (1 µl of nuclease-free water in the case of V. cholerae). The PCR conditions for V. cholerae were optimised as described by le Roux and van Blerk (2011). This included an initial activation at 95 °C for 10 min followed by 50 cycles involving denaturation at 95 °C (10 s), annealing at 64 °C (15 s), extension at 72 °C (25 s) and acquiring after each cycle and a final extension at 72 °C for 5 min. Melting was done by ramping from 72 to 90 °C, with a 0.1 °C rise at each step, a pre-melt hold for 90 s on the first step followed by a hold for 2 s on the next steps. The same conditions were applied for Salmonella and Shigella. However, the PCR for Shigella was semi-nested as described by Theron et al. (2001). Melt curve analysis of the PCR product was carried out using the Rotor-Gene Q Series Software version 6.1 (Qiagen, Hilden, Germany) to detect the presence of the genes of interest for each organism. Samples for the POs were analysed in duplicate.
All PCR reactions included a positive (genomic DNA of a reference strain) and a negative control (made of the PCR reaction mixture without the template DNA). Reference strains used were Salmonella ser Typhimurium (ATCC® 14028) (American Type Culture Collection, Manassas, VA, USA), Shigella flexneri (ATCC® 12022) and V. cholerae O1 (NCTC 5941) (National Collection of Type Cultures, London, UK). All positive strains were obtained from the microbiology laboratory of the Natural Resources and the Environment Department of the CSIR (Council for Scientific and Industrial Research), Pretoria, South Africa.
Statistical analysis
Data analysis was performed using Microsoft Excel and SPSS statistical analysis software version 20 (IBM Corporation, Armonk, New York, USA). Concentrations of microorganisms were expressed as Log10 Geometric mean. For the calculation of the geometric means, all MPN values of E. coli below the detection limit were assumed to be 1. The Spearman’s rank correlation was used to determine the correlation between the abundance of each FIO and the presence of the various pathogens. For the pathogens, a sampling point was considered positive for any of the organism of interest when both duplicate samples from that site were positive at any given time. The one-way analysis of variance (ANOVA) was used to check for any statistical differences between data sets. All analyses were undertaken at a level of significance of α = 0.05.
Results and discussion
Abundance of indicator organisms and prevalence of pathogenic organisms in water and sediments
The importance of sediments as a reservoir and as a possible source of FIOs and POs within the water column in aquatic ecosystems has been studied for many years and still represents an issue of concern in the present day (Labelle et al. 1980; Burton et al. 1987; Santhiya et al. 2011; Walters et al. 2014). This is particularly a major problem in most developing countries that still rely on untreated water from polluted surface water bodies for their personal, domestic and recreational activities.
In the current study, the mean concentration of E. coli in the water ranged between 3.80 and 7.03 × 102 ± SD (Standard deviation) MPN/100 mL during the dry season and between 1.65 × 101 and 4.05 × 103 ± SD MPN/100 mL in the wet season as previously described by (Abia et al. 2015a). Sediment concentrations ranged between 3.90 and 1.47 × 103 ± SD MPN/100 mL and between 3.65 × 102 and 3.37 × 104 ± SD MPN/100 mL in the dry and the wet season, respectively. For C. perfringens, mean water concentrations for the dry season and wet season ranged from 4.18 × 102 to 1.72 × 104 ± SD CFU/100 mL and from 4.70 × 102 to 7.32 × 103 ± SD CFU/100 mL, respectively. The sediment concentrations ranged from 2.86 × 103 to 6.02 × 104 ± SD CFU/100 mL and 2.72 × 104 to 1.39 × 105 ± SD CFU/100 mL for the dry and wet season, respectively. There was a statistically significant difference (p < 0.05) between the mean water and the mean sediment counts for both FIOs and during both seasons. The wet season also recorded an overall statistically significant higher count (p < 0.05) than the dry season for each FIO. The pathogens were detected on a presence/absence basis targeting the ompW and ctxAB of V. cholerae, invA gene of Salmonella spp and the ipaH of Shigella. The overall prevalence of all the pathogens is shown in Table 2. The most detected pathogen during the entire sampling period (water and sediment) was V. cholerae (58.8 %) while Shigella spp recorded the lowest prevalence (10.1 %). Site AP1 recorded the highest prevalence of V. cholerae (52/56; 92.9 %), while site AP2 recorded the highest prevalence for Salmonella spp (24/56; 42.7 %), and site AP8 recorded the highest prevalence for Shigella spp (28/56; 49.6 %). Figure 2 shows the contribution of water and sediments to the overall observed prevalence of each pathogen at the individual sampling sites during the dry and the wet season. The pathogens were more detected at sampling sites during the wet season (January–February 2014) than during the dry season (May–August 2013) (Fig. 2). None of the pathogens was detected at site DAS and AP5 during the dry season.
All the sites that recorded the highest abundance of FIOs and prevalence of the POs were located along the main river course. The sites on the main river were either characterised by informal settlements (AP1) or located downstream a WWTW (DAS, AP2) or had a combination of both (AP8). The highest prevalence for V. cholerae was at site AP1 (Table 2) which is characterised by an informal settlement. Site AP8 is located downstream from two WWTWs (Babelegi and Temba) and also around agricultural areas; this site also demonstrated a similarly high V. cholerae prevalence as was observed at site AP1. These sites that recorded high prevalence of the POs equally recorded high abundance of the FIOs. The negative impact of informal settlements (Paulse et al. 2009; Khan and Khan 2012; Ndlovu et al. 2015) and agriculture (Kay et al. 2008; Páll et al. 2013) on the microbial quality of surface water bodies has been reported. However, considering that none of the V. cholerae isolated from the sediments and water of the Apies River carried the gene for cholera toxin production (ctx), it could be unlikely that this high prevalence (58.8 %) was solely due to human influence. Nonpathogenic strains of V. cholerae are widely distributed in the aquatic environment (Finkelstein 1996). Studies have shown that V. cholerae forms an integral part of the aquatic ecosystem and is usually associated with zooplanktons (copepods) that can contain up to 105 V. cholerae cells on their carapace and in their gut (Rawlings et al. 2007; de Magny et al. 2011; Kirschner et al. 2011). Although the ctx gene of V. cholerae was not isolated in this study, the detection of the ompW still represents a possible health threat as it has been shown that environmental stains of V. cholerae could contain other virulence factors that would allow them to induce infection under appropriate conditions within the intestines of rabbits (Faruque et al. 2004). Similarly, Bag et al. (2008) demonstrated that non-O1 and non-O139 strains of V. cholerae isolated from surface water in India showed haemolytic activity when exposed to human erythrocytes. Thus, these strains might have the potential of causing disease once in the human system, especially in immune-compromised individuals.
Sites AP2 and AP8 that recorded highest prevalence of Salmonella and Shigella, respectively, are located downstream from WWTWs. This site (AP2) also recorded the highest concentration of both FIOs in the sediments. Site AP2 is located downstream from the Rooiwal WWTW which has been reported to function above its operational capacity (South African Department of Water Affairs 2012). Teklehaimanot et al. (2014) earlier reported high prevalence of Salmonella spp in effluents from three WWTWs in South Africa and their respective receiving water bodies. However, just like V. cholerae, Salmonella spp have been found to survive in the environment for long periods and have also been found to infect other domestic animals and birds (Jeong et al. 2010). Some water amphibians and reptiles have also been reported to be carriers of Salmonella spp (CDC 2013) although humans are the only known natural hosts and reservoir for some strains like Salmonella enterica serovar Typhi (S. Typhi) (Kaur and Jain 2012). Considering that the detection of this pathogen was done through the identification of the invA gene which is highly specific for all members of the genus Salmonella, it would be impossible to determine whether the genes detected were of animal or human origin. Nevertheless, studies have reported human infections originating from Salmonella spp of animal origin (Olsvik et al. 1985; Zhao et al. 2003; Hendriksen et al. 2004). Thus, the detection of Salmonella in the sediments and water of the Apies river still represents a health threat if these waters are used untreated by surrounding communities.
Unlike V. cholerae and Samonella spp, the ipaH gene of Shigella spp was only detected at sites located on the main river course. Samples from the tributaries (AP3, AP4 and AP5) were all negative for the ipaH gene. Again, these tributary sites had little or no direct human influence suggesting that human activities around the Apies River may contribute to the pollution observed in the river. Shigella has been reported in other hosts such as monkeys, rabbits, calves, piglets and even chickens (Jiang et al. 2005; Pan et al. 2006; Shi et al. 2014). However, several studies using animal models have demonstrated that Shigella acquired through the oral route in these animals would hardly result in a disease condition in immuno-competent hosts due to immune clearance by the animals’ defence systems. The organism could only cause disease in immuno-suppressed experimental animals (Rabbani et al. 1995; Jeong et al. 2010; Mostowy et al. 2013; Shi et al. 2014). In humans, this is not the case as the organism can bypass the human immune system and invade the large intestine thus establishing an infection (Ashida et al. 2011). Therefore, considering that the faecal–oral route is the main mode of transmission of Shigella and that the organism has minimal ability of evading the immune system of non-human hosts, it is likely that the presence of this pathogen in the aquatic environment could be predominantly of human origin. Shi et al. (2014) also demonstrated that Shigella of human and chicken origins shared similar pathogenicity and that there was the possibility of human–poultry cross-infection. This means that even if the ipaH genes of Shigella isolated in the current study were of animal origin, they could still imply a possible threat to human. When compared to Salmonella (>105 CFU) and V. cholerae (~104 CFU) the infective dose of Shigella is relatively very low (<10–100 CFU) (Kothary and Babu 2001; Lemarchand et al. 2004; WHO 2005; Al-bashan 2012). Thus, the presence of Shigella in the aquatic environment even at low prevalence still represents a threat of public health importance, especially in areas where such surface water is used for drinking without prior treatment (Mulamattathil et al. 2014). Studies have also shown that Shigella is quickly inactivated when out of its host and does not survive for longer than 7 days in the environment when compared to other enteric pathogens (Mcfeters et al. 1974; Islam et al. 2001; Al-bashan 2012). This could therefore imply that the presence of the organisms in the environment may be indicative of recent faecal pollution. However, this could not be confirmed in this study considering that the detection of pathogens was carried out using PCR and not isolation of viable cells through culture. Although the ipaH gene detected in the present study is also carried by enteroinvasive E. coli strains (EIEC) (Theron et al. 2001), Shigella and EIEC are both causative agents of bacillary dysentery in humans (Hsu et al. 2010) and thus the presence of the ipaH gene in riverbed sediments is still indicative of possible negative health implications to users of the untreated river water.
The overall (water and sediments) seasonal pattern in the detection of all the three pathogens (V. cholerae, Salmonella spp and Shigella spp) was similar to that observed with the FIOs, with the wet season recording higher prevalence than the dry season. This observation strengthens the role played by seasonal variation on pollution in the aquatic environment. The higher prevalence of POs in the river system during the summer (wet) months observed in this study contradicts those of Kinge and Mbewe (2010). In their study, the authors recorded higher prevalence of Shigella spp during the winter (dry months) compared to the summer months in river catchments of the Northwest Province of South Africa. On the other hand, findings of this study agree with those of Mulamattathil et al. (2014), who reported a higher prevalence of Shigella in surface water in the Mafikeng area in the Northwest Province of South Africa during the summer months. However, results of the studies by Kinge and Mbewe (2010) and by Mulamattathil et al. (2014) were only based on surface water and did not include the detection of the pathogens in the bed sediments in these catchments. As opposed to the FIOs, the pathogens were more detected in the water column than in the riverbed sediments. However, a strong positive correlation (p < 0.01) was observed between the water prevalence and the sediment prevalence for all three pathogens. This indicates that there could be vertical movement of the organisms between the water column and the sediments with the sediments either acting as a sink for or a source (or both) of microorganisms within the aquatic environment.
Correlation between indicator organisms and pathogens in water and sediments
While several studies have investigated the correlation between FIOs and POs within the water column in aquatic environments, data demonstrating strong correlation between FIOs and pathogens in the sediments are rare (Weaver and Sinton 2009). Also, long debates surrounding the use of the term “indicator organisms” and the ambiguous use of the term “microbial indicator” led to the reclassification of indicator organism into process indicators (demonstrates how efficient a treatment process is—e.g. total coliforms), faecal indicators (indicate faecal contamination—e.g. E. coli) and index and model organisms (indicative of pathogen presence and behaviour, respectively—e.g. F-RNA coliphages) (Ashbolt et al. 2001).
In the present study, we investigated whether the abundance of E. coli and C. perfringens correlated with the detection of V. cholerae, Salmonella spp and Shigella spp, especially within the sediments. The nonparametric Spearman rank correlation coefficient was used to establish any relationship between the FIOs and the POs during the dry season (Table 3) and during the wet season (Table 4). During both seasons within the water column, a strong positive correlation was observed between E. coli and each of the pathogens while C. perfringens only correlated with V. cholerae (Tables 3, 4). Within the sediments, strong correlations were only observed between E. coli and Salmonella spp, E. coli and V. cholerae (dry season) and E. coli and V. cholerae and E. coli and Shigella spp (wet season) (Tables 3, 4). These results corroborate with findings of Abdallah et al. (2005). In their study, the authors reported that faecal coliforms were strongly correlated to Salmonella and V. cholerae in beach sand of the Gaza Strip. On the other hand, no correlation was observed between C. perfringens and any of the pathogens within the sediments. This could be due to the ability of C. perfringens to survive for very long periods in the environment even after a pollution event has occurred (Gemmell and Schmidt 2013; Shibata et al. 2004). Most bacterial pathogens survive for shorter periods in the environment than C. perfringens (WHO 2008). In a study by Tyagi and Chopra (2006), the authors also reported a lack of correlation between C. perfringens and other POs and suggested that while C. perfringens is not a good indicator of bacterial pathogens in the aquatic environment, it is a good index organism for viruses and some parasites that survive in the environment for longer periods.
Within the water column, however, E coli demonstrated a strong positive correlation with all three pathogens during both seasons while C. perfringens only correlated with V. cholerae. V. cholerae has been reported to survive longer in the environment than many other bacterial pathogens like Salmonella (Djaouda et al. 2013). This could possibly explain its correlation with C. perfringens as observed in our study. In a recent study by Gemmell and Schmidt (2013) on the microbiological quality of the Msunduzi River in KwaZulu-Natal, South Africa, the authors reported that E. coli was a suitable indicator of POs, especially Salmonella spp in river water. The level of detection of Salmonella dropped with a corresponding drop in the E. coli counts within the river. In their study, they also reported the lack of correlation between C. perfringens and pathogenic bacteria. Several other studies have reported a correlation between E. coli and other pathogenic bacteria (Abdallah et al. 2005; Devane et al. 2014; Leclerc et al. 2001).
Thus, the findings of the present study together with previous studies suggest that although E. coli is considered a faecal indicator organism, it could also be a good indicator of other bacterial pathogens, especially in the water column. Sediments represent a more complex environment than the water column in terms of microbial diversity and chemical composition. The fact that E. coli only correlated with some organisms in the sediments during the dry and wet season further emphasises the need for better indicators of pathogens in the sediments. The strong correlation between E. coli and the pathogens supports the classification of E. coli as a good index organism for pathogens like Salmonella. However, there exist no data on suitable index organisms of V. cholerae and Shigella spp in the environment. Also, the notion of index organisms should be studied extensively within a given environment before it is fully used.
Conclusion
The water from the Apies river is unsafe for use for recreation and if untreated, for any household use. The sediments of the Apies River are polluted with indicator and pathogenic bacteria. There is a strong correlation between E. coli and pathogenic bacteria (V. cholerae, Salmonella and Shigella) in the water column suggesting that E. Cole is not only a good indicator of faecal pollution, but could also be a good indicator for the presence of other pathogenic bacteria. However, the lack of correlation between E. coli and some pathogens in the sediments depending on the season as observed in the present study highlights the need to investigate for more indicators that could better indicate the presence of pathogens in this complex matrix. The lack of correlation between C. perfringens and the pathogens does not eliminate its suitability as an indicator of long-term faecal pollution indicator. Although no toxigenic V. cholerae strain was detected in this study, due to the high percentage of V. cholerae detected we recommend that further research be carried out to investigate if the environmental strains of V. cholerae in sediments and water of the Apies River carry other virulence genes that could allow them to initiate infection under appropriate conditions. It would also be necessary to investigate how long these organisms can survive in the sediments of the Apies River and if human-induced or increased flow conditions may cause their resuspension.
References
Abdallah SA, Elmanama AA, Fahd MI, Afifi S (2005) Microbiological beach sand quality in the Gaza Strip in comparison to seawater. Polish J Environ Stud 14:841–850
Abia ALK, Ubomba-Jaswa E, du Preez M, Momba MNB (2015a) Riverbed sediments in the Apies River, South Africa: recommending the use of both Clostridium perfringens and Escherichia coli as indicators of faecal pollution. J Soils Sediments 15:2412–2424. doi:10.1007/s11368-015-1209-0
Abia ALK, Ubomba-Jaswa E, Momba MNB (2015b) Impact of seasonal variation on Escherichia coli concentrations in the riverbed sediments in the Apies River, South Africa. Sci Total Environ 537:462–469. doi:10.1016/j.scitotenv.2015.07.132
Abia LKA, Ubomba-Jaswa E, Ssemakalu CC, Momba MNB (2015c) Development of a rapid approach for the enumeration of Escherichia coli in riverbed sediment: case study, the Apies River, South Africa. J Soils Sediments 15:2425–2432. doi:10.1007/s11368-015-1081-y
Abraham W-R (2011) Megacities as sources for pathogenic bacteria in rivers and their fate downstream. Int J Microbiol. doi:10.1155/2011/798292
Akoachere J-FT, Masalla T, Njom H (2013) Multi-drug resistant toxigenic Vibrio cholerae O1 is persistent in water sources in New Bell-Douala, Cameroon. BMC Infect Dis 13:366. doi:10.1186/1471-2334-13-366
Al-bashan MM (2012) Influences of some physico-chimical stress conditions on the survivality and resistibility of Shigella flexneri and Shigella sonnei. Glob Vet 9:706–716. doi:10.5829/idosi.gv.2012.9.6.52224
APHA/AWWA/WEF (2001) American Public Health Association, American Water Works Association, Water Environment Federation (APHA-AWWA-WEF) .2001. Standard methods for the examination of water and wastewater, 22nd edn. Washington
Ashbolt N, Grabow W, Snozzi M (2001) Indicators of microbial water quality. In: Fewtrell L, Barthram J (eds) water qual. IWA Publishing, London, pp 289–316
Ashida H, Ogawa M, Mimuro H et al (2011) Shigella are versatile mucosal pathogens that circumvent the host innate immune system. Curr Opin Immunol 23:448–455. doi:10.1016/j.coi.2011.06.001
Bag PKP, Bhowmik P, Hajra TK et al (2008) Putative virulence traits and pathogenicity of Vibrio cholerae non-o1, non-o139 isolates from surface waters in Kolkata, India. Appl Environ Microbiol 74:5635–5644. doi:10.1128/AEM.00029-08
Basson MS (2011) Water development in South Africa. UN-Water Int. Conf. Water Green Econ. Pract. Towar. Rio + 20. pp 1–12
Budillon F, Vicinanza D, Ferrante V, Iorio M (2006) Sediment transport and deposition during extreme sea storm events at the Salerno Bay (Tyrrhenian Sea): comparison of field data with numerical model results. Nat Hazards Earth Syst Sci 6:839–852. doi:10.5194/nhess-6-839-2006
Burton GAJ, Gunnison D, Lanza GR (1987) Survial of pathogenic bacteria in various freshwater sediments. Appl Environ Microbiol 53:633–638
Carey CM, Lee H, Trevors JT (2004) Biology, persistence and detection of Cryptosporidium parvum and Cryptosporidium hominis oocyst. Water Res 38:818–862. doi:10.1016/j.watres.2003.10.012
CDC (2013) Turtles and other reptiles are risky pets. http://www.cdc.gov/Features/SalmonellaFrogTurtle/. Accessed 31 Mar 2015
Characklis GW, Dilts MJ, Simmons OD et al (2005) Microbial partitioning to settleable particles in stormwater. Water Res 39:1773–1782. doi:10.1016/j.watres.2005.03.004
Chase E, Hunting J, Staley C, Harwood VJ (2012) Microbial source tracking to identify human and ruminant sources of faecal pollution in an ephemeral Florida river. J Appl Microbiol 113:1396–1406. doi:10.1111/jam.12007
Chigor VN, Okoh AI (2012) Quantitative detection and characterization of human adenoviruses in the buffalo river in the Eastern Cape province of South Africa. Food Environ Virol 4:198–208. doi:10.1007/s12560-012-9090-0
Chitanand MP, Gyananath G, Lade HS (2008) Bacterial assessment of ground water: a case study of Nanded city. J Environ Biol 29:315–318
Craig DL, Fallowfield HJ, Cromar NJ (2004) Use of microcosms to determine persistence of Escherichia coli in recreational coastal water and sediment and validation with in situ measurements. J Appl Microbiol 96:922–930. doi:10.1111/j.1365-2672.2004.02243.x
Crowther J, Kay D, Wyer MD (2002) Faecal-indicator concentrations in waters draining lowland pastoral catchments in the UK: relationships with land use and farming practices. Water Res 36:1725–1734. doi:10.1016/S0043-1354(01)00394-3
de Magny GC, Mozumder PK, Grim CJ et al (2011) Role of zooplankton diversity in Vibrio cholerae population dynamics and in the incidence of cholera in the Bangladesh Sundarbans. Appl Environ Microbiol 77:6125–6132. doi:10.1128/AEM.01472-10
Decamp O, Warren A (2000) Investigation of Escherichia coli removal in various designs of subsurface flow wetlands used for wastewater treatment. Ecol Eng 14:293–299
Department of Environmental Affairs Republic of South Africa (2012) South African water quality guidelines for coastal marine waters. Volume 2: Guielines for recreational use
Devane ML, Moriarty EM, Wood D et al (2014) The impact of major earthquakes and subsequent sewage discharges on the microbial quality of water and sediments in an urban river. Sci Total Environ 485–486:666–680. doi:10.1016/j.scitotenv.2014.03.027
Djaouda M, Gaké B, Ebang Menye D et al (2013) Survival and Growth of Vibrio cholerae, Escherichia coli, and Salmonella spp. in well water used for drinking purposes in Garoua (North Cameroon). Int J Bacteriol. doi:10.1155/2013/127179
Agency Environment (2002) The Microbiology of Drinking Water (2002)—Part 10—Methods for the isolation of Yersinia, Vibrio and Campylobacter by selective enrichment. Methods exam Waters Assoc Mater 10:14–17
Faruque SM, Chowdhury N, Kamruzzaman M et al (2004) Genetic diversity and virulence potential of environmental Vibrio cholerae population in a cholera-endemic area. Proc Natl Acad Sci 101:2123–2128. doi:10.1073/pnas.0308485100
Finkelstein RA (1996) Cholera, Vibrio cholerae O1 and O139, and other pathogenic vibrios. In: Baron S (ed) Medical microbiolology, 4th edn. University of Texas, Galveston
Fries JS, Characklis GW, Noble RT (2008) Sediment—water exchange of Vibrio sp. and fecal indicator bacteria: implications for persistence and transport in the Neuse River Estuary, North Carolina. USA Water Res 42:941–950. doi:10.1016/j.watres.2007.09.006
Gao G, Falconer R, Lin B (2011) Numerical modelling sediment-bacteria interaction processes in the Severn Estuary. J Water Resour Prot 3:22–31. doi:10.4236/jwarp.2011.31003
Garzio-hadzick A, Shelton DR, Hill RL et al (2010) Survival of manure-borne E. coli in streambed sediment: effects of temperature and sediment properties. Water Res 44:2753–2762. doi:10.1016/j.watres.2010.02.011
Gemmell ME, Schmidt S (2013) Is the microbiological quality of the Msunduzi River (KwaZulu-Natal, South Africa) suitable for domestic, recreational, and agricultural purposes? Environ Sci Pollut Res 20:6551–6562. doi:10.1007/s11356-013-1710-1
Goel AK, Tamrakar AK, Nema V et al (2005) Detection of viable toxigenic Vibrio cholerae from environmental water sources by direct cell duplex PCR assay. World J Microbiol Biotechnol 21:973–976. doi:10.1007/s11274-004-7317-4
Gonzalez RA, Conn KE, Crosswell JR, Noble RT (2012) Application of empirical predictive modeling using conventional and alternative fecal indicator bacteria in eastern North Carolina waters. Water Res 46:5871–5882. doi:10.1016/j.watres.2012.07.050
Hendriksen SWM, Orsel K, Wagenaar JA, Miko A (2004) Animal-to-human transmission of Salmonella Typhimurium DT104A variant. Emerg Infect Dis 10:2225–2227
Hsu B-M, Wu S-F, Huang S-W et al (2010) Differentiation and identification of Shigella spp. and enteroinvasive Escherichia coli in environmental waters by a molecular method and biochemical test. Water Res 44:949–955. doi:10.1016/j.watres.2009.10.004
Ibekwe A, Papiernik S (2010) Quantification of persistence of Escherichia coli O157: H7 in contrasting soils. Int J Microbiol. doi:10.1155/2011/421379
Islam MS, Hossain MA, Khan SI et al (2001) Survival of Shigella dysenteriae type 1 on fomites. J Heal Popul Nutr 19:177–182
Jamieson R, Gordon R, Joy D, Lee H (2004) Assessing microbial pollution of rural surface waters a review of current watershed scale modeling approaches. Agric Water Manag 70:1–17. doi:10.1016/j.agwat.2004.05.006
Jamieson RC, Joy DM, Lee H et al (2005a) Resuspension of sediment-associated Escherichia coli in a natural stream. J Environ Qual 34:581. doi:10.2134/jeq2005.0581
Jamieson RC, Joy DM, Lee H et al (2005b) Transport and deposition of sediment-associated Escherichia coli in natural streams. Water Res 39:2665–2675. doi:10.1016/j.watres.2005.04.040
Jeong K, Zhang Q, Nunnari J, Tzipori S (2010) A piglet model of acute gastroenteritis induced by Shigella dysenteriae Type 1. J Infect Dis 201:903–911. doi:10.1086/650995
Jiang J, Wang P, Pan G, Kang L (2005) Isolation and identification of rabbits Shigella dysenteriae in a large scale warren. J Anhui Agric Sci 33:1666–1667
Kaur J, Jain SK (2012) Role of antigens and virulence factors of Salmonella enterica serovar Typhi in its pathogenesis. Microbiol Res 167:199–210. doi:10.1016/j.micres.2011.08.001
Kay D, Crowther J, Fewtrell L et al (2008) Quantification and control of microbial pollution from agriculture: a new policy challenge? Environ Sci Policy 11:171–184. doi:10.1016/j.envsci.2007.10.009
Khan S, Khan W (2012) Isolation and identification of bacterial pollutants from the Berg and Plankenburg Rivers in the Western Cape, South Africa. Water SA 38:819–824
Kinge CW, Mbewe M (2010) Characterisation of Shigella spesies isolated from river catchments in the North West Province of South Africa. S Afr J Sci 106:211. doi:10.4102/sajs.v106i11/12.211
Kinzelman JL, McLellan SL (2009) Success of science-based best management practices in reducing swimming bans—a case study from Racine, Wisconsin, USA. Aquat Ecosyst Health Manag 12:187–196. doi:10.1080/14634980902907466
Kirschner AKT, Schauer S, Steinberger B et al (2011) Interaction of Vibrio cholerae non-O1/non-O139 with copepods, cladocerans and competing bacteria in the large Alkaline Lake Neusiedler See, Austria. Microb Ecol 61:496–506. doi:10.1007/s00248-010-9764-9
Koirala SR, Gentry RW, Perfect E et al (2008) Temporal variation and persistence of bacteria in streams. J Environ Qual 37:1559–1566. doi:10.2134/jeq2007.0310
Korajkic A, Brownell MJ, Harwood VJ (2011) Investigation of human sewage pollution and pathogen analysis at Florida Gulf coast beaches. J Appl Microbiol 110:174–183. doi:10.1111/j.1365-2672.2010.04869.x
Kothary MH, Babu US (2001) Infective dose of foodborne pathogens in volunteers: a review. J Food Saf 21:49–73. doi:10.1111/j.1745-4565.2001.tb00307.x
Labelle RL, Gerba CP, Goyal SM et al (1980) Relationships between environmental factors, bacterial indicators, and the occurrence of enteric viruses in estuarine sediments. Appl Environ Microbiol 39:588–596
le Roux WJ, van Blerk GN (2011) The use of a high resolution melt real-time polymerase chain reaction (PCR) assay for the environmental monitoring of Vibrio cholerae. African J Microbiol Res 5:3520–3526. doi:10.5897/AJMR11.695
Leclerc H, Mossel DA, Edberg SC, Struijk CB (2001) Advances in the bacteriology of the coliform group: their suitability as markers of microbial water safety. Annu Rev Microbiol 55:201–234. doi:10.1146/annurev.micro.55.1.201
Lemarchand K, Masson L, Brousseau R (2004) Molecular biology and DNA microarray technology for microbial quality monitoring of water. Crit Rev Microbiol 30:145–172. doi:10.1080/10408410490435142
Malorny B, Hoorfar J, Bunge C, Helmuth R (2003) Multicenter validation of the analytical accuracy of Salmonella PCR: towards an international standard. Appl Environ Microbiol 69:290–296. doi:10.1128/AEM.69.1.290-296.2003
Mcfeters GA, Bissonnette GK, Jezeski JJ et al (1974) Comparative survival of indicator bacteria and enteric pathogens in well water. Appl Mcrobiol 27:823–829
Molobela IP, Sinha P (2011) Management of water resources in South Africa: a review. African J Environ Sci Technol 5:993–1002. doi:10.5897/AJEST11.136
Monis PT, Thompson RCA (2003) Cryptosporidium and Giardia-zoonoses: fact or fiction? Infect Genet Evol 3:233–244. doi:10.1016/j.meegid.2003.08.003
Mostowy S, Boucontet L, Moya MJM et al (2013) The zebrafish as a new model for the in vivo study of Shigella flexneri interaction with phagocytes and bacterial autophagy. PLoS Pathog 9:e1003588. doi:10.1371/journal.ppat.1003588
Mulamattathil SG, Bezuidenhout C, Mbewe M, Ateba CN (2014) Isolation of environmental bacteria from surface and drinking water in Mafikeng, South Africa, and characterization using their antibiotic resistance profiles. J Pathog. doi:10.1155/2014/371208
Nandi B, Nandy RK, Mukhopadhyay S et al (2000) Rapid method for species-specific identification of Vibrio cholerae using primers targeted to the gene of outer membrane protein OmpW. J Clin Microbiol 38:4145–4151
Ndlovu T, Le Roux M, Khan W, Khan S (2015) Co-detection of virulent Escherichia coli genes in surface water sources. PLoS ONE 10:e0116808. doi:10.1371/journal.pone.0116808
Olsvik O, Sørum H, Birkness K et al (1985) Plasmid characterization of Salmonella typhimurium transmitted from animals to humans. J Clin Microbiol 22:336–338
Páll E, Niculae M, Kiss T et al (2013) Human impact on the microbiological water quality of the rivers. J Med Microbiol 62:1635–1640. doi:10.1099/jmm.0.055749-0
Pan B, Wang W, Xie Y et al (2006) Detection, serology classification and drug susceptibility of Shigella from experimental monkeys. Guangxi Agric Sci 37:331–332
Pandey PK, Soupir ML, Rehmann CR (2011) A model for predicting resuspension of Escherichia coli from streambed sediments. Water Res 46:115–126. doi:10.1016/j.watres.2011.10.019
Paulse AN, Jackson VA, Khan W (2009) Comparison of microbial contamination at various sites along the Plankenburg—and Diep Rivers, Western Cape, South Africa. Water SA 35:469–478
Pianetti A, Bruscolini F, Sabatini L, Colantoni P (2004) Microbial characteristics of marine sediments in bathing area along Pesaro-Gabicce coast (Italy): a preliminary study. J Appl Microbiol 97:682–689. doi:10.1111/j.1365-2672.2004.02352.x
Rabbani GH, Albert MJ, Rahman H et al (1995) Development of an improved animal model of shigellosis in the adult rabbit by colonic infection with Shigella flexneri 2a. Infect Immun 63:4350–4357
Rawlings TK, Ruiz GM, Colwell RR (2007) Association of Vibrio cholerae O1 El Tor and O139 Bengal with the copepods Acartia tonsa and Eurytemora affinis. Appl Environ Microbiol 73:7926–7933. doi:10.1128/AEM.01238-07
Reeves RL, Grant SB, Mrse RD et al (2004) Scaling and management of fecal indicator bacteria in runoff from a coastal urban watershed in Southern California. Environ Sci Technol 38:2637–2648. doi:10.1021/es034797g
Santhiya G, Lakshumanan C, Selvin J, Asha D (2011) Microbiological analysis of seawater and sediments in urban shorelines: occurrence of heavy metals resistance bacteria on Chennai beaches, Bay of Bengal. Microchem J 99:197–202. doi:10.1016/j.microc.2011.05.004
Seanego KG, Moyo NAG (2013) The effect of sewage effluent on the physico-chemical and biological characteristics of the Sand River, Limpopo, South Africa. Phys Chem Earth 66:75–82. doi:10.1016/j.pce.2013.08.008
Shi R, Yang X, Chen L et al (2014) Pathogenicity of Shigella in chickens. PLoS ONE 9:e100264. doi:10.1371/journal.pone.0100264
Shibata T, Solo-Gabriele HM, Fleming LE, Elmir S (2004) Monitoring marine recreational water quality using multiple microbial indicators in an urban tropical environment. Water Res 38:3119–3131. doi:10.1016/j.watres.2004.04.044
Sibanda T, Okoh AI (2013) Real-time PCR quantitative assessment of hepatitis A virus, rotaviruses and enteroviruses in the Tyume River located in the Eastern Cape Province, South Africa. Water SA 39:295–304
Signor RS (2005) Quantifying the impact of runoff events on microbiological contaminant concentrations entering surface drinking source waters. J Water Health 3:453–468. doi:10.2166/wh.2005.052
South African Department of Water Affairs (DWA) (2012) Green drop progress report. Department of Water Affairs, Pretoria
Stapleton CM, Wyer MD, Kay D et al (2007) Fate and transport of particles in estuaries, vols. I, II, III, IV Environment agency science report SC000002/SR1-4
Teklehaimanot GZ, Coetzee MAA, Momba MNB (2014) Faecal pollution loads in the wastewater effluents and receiving water bodies: a potential threat to the health of Sedibeng and Soshanguve communities, South Africa. Environ Sci Pollut Res 21:9589–9603. doi:10.1007/s11356-014-2980-y
Theron J, Morar D, du Preez M et al (2001) A sensitive seminested PCR method for the detection of Shigella in spiked environmental water samples. Water Res 35:869–874
Tyagi V, Chopra A (2006) Alternative microbial indicators of faecal pollution: current perspective. Iran J Environ Heal Sci Eng 3:205–216
Tyrrel SF, Quinton JN (2003) Overland flow transport of pathogens from agricultural land receiving faecal wastes. J Appl Microbiol 94(Suppl):87S–93S. doi:10.1046/j.1365-2672.94.s1.10.x
Venter A (2007) Prioritization of river basins in the Tshwane area with reference to faecal coliform bacteria for the purpose of the identification of candidate wetlands for rehabilitation. University of the Witwatersrand, Johannesburg
Vignesh S, Dahms H, Emmanuel KV et al (2014) Physicochemical parameters aid microbial community? A case study from marine recreational beaches, Southern India. Environ Monit Assess 186:1875–1887. doi:10.1007/s10661-013-3501-z
Walters E, Schwarzwälder K, Rutschmann P et al (2014) Influence of resuspension on the fate of fecal indicator bacteria in large-scale flumes mimicking an oligotrophic river. Water Res 48:466–477. doi:10.1016/j.watres.2013.10.002
Weaver L, Sinton L (2009) Deposition and survival of enteric microbes in aquatic sediments—a brief review. Institute of Environmental Science & Research Ltd, Christchurch
WHO (2005) Guidelines for the control of shigellosis, including epidemics due to Shigella dysenteriae type 1. WHO Press, Geneva
WHO (2008) WHO guidelines for drinking-water quality, Vol 1, 3rd edn. Recommendations. doi: 10.1016/S1462-0758(00)00006-6
Zhao S, Qaiyumi S, Friedman S et al (2003) Characterization of Salmonella enterica serotype Newport isolated from humans and food animals. J Clin Microbiol 41:5366–5371. doi:10.1128/JCM.41.12.5366-5371.2003
Acknowledgments
This work received funding from the Tshwane University of Technology (TUT), Water Research commission (WRC), South Africa (Grant numbers K5/2169 and K5/2147) and the National Research Foundation (NRF). Opinions expressed and conclusions arrived at, are those of the authors and are not necessarily to be attributed to the TUT, WRC or NRF.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Abbaspour.
Rights and permissions
About this article
Cite this article
Abia, A.L.K., Ubomba-Jaswa, E. & Momba, M.N.B. Prevalence of pathogenic microorganisms and their correlation with the abundance of indicator organisms in riverbed sediments. Int. J. Environ. Sci. Technol. 13, 2905–2916 (2016). https://doi.org/10.1007/s13762-016-1116-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-016-1116-y