Abstract
Agricultural practices change soil’s physical and chemical properties, therefore modifying soil microbial communities. Conservation tillage is widely used to improve the soil texture and nutrient status in the dryland regions of northern China. However, little is known about the influence of soil properties on microbes, in particular on the effect of conservation tillage on soil bacterial communities. Here, we studied the effect of a 5-year tillage treatment on soil properties and soil bacterial communities in the dryland regions of northern China using a high-throughput sequencing technology and quantitative PCR of 16S rRNA genes. We compared the changes in soil bacterial diversity, and composition was measured for conservation tillage, including zero plow or chisel plow, and for conventional tillage using plow. Our results show that conservation tillage increased the Simpson index by 378 % and exhibited significantly dissimilar polygenetic diversity, with r of 1, and taxonomic diversity, of r higher than 0.49, compared to conventional tillage. This finding demonstrates that conservation tillage modifies soil bacterial diversity. Chisel plow and zero tillage increase the abundance of the genus Bacillus, including 85 % of the phylum Firmicutes, and of Rhizobiales belonging to the Alphaproteobacteria. Overall conservation tillage increased the abundance of profitable functional bacteria species.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Agriculture is one of the most significant anthropogenic contributors to the alteration of soil characteristics including the physical, chemical, and biological properties of the soil (Kladivko 2001). These activities affect the relative abundance, diversity, and activities of the resident microbes. Tillage is the major agricultural practice that impacts the soil quality because the physical disturbance of the soil leads to alterations in soil water content, the mechanical composition of soil particles, and the degree of mixing of the crop residues within the soil matrix (Kladivko 2001). Soil microbiota plays important roles in the ecological processes that are facilitated by soil disturbance and contribute both directly and indirectly to crop growth and quality, nutrient cycling quality, and the sustainability of soil productivity (Roger-Estrade et al. 2010). The effect of tillage appears to correlate with the changes in soil structure that alter microbial diversity; following convention tillage, the soil is expected to exhibit differences in the community structure, diversity, and abundance of microbes compared with that following conservation tillage (Brussaard et al. 2007).
Numerous studies have focused on how microbial abundance and structure are affected by reduced tillage and have indicated that soil organic carbon is a majority factor modifying the soil microbial community (Ceja-Navarro et al. 2010; Pastorelli et al. 2013). However, a few studies have shown that conservation tillage enhances soil carbon sequestration, but these reports have described changes in microbial biomass and diversity in northern China that account for approximately 56 % of the Chinese land area, only using indirect techniques (Chen et al. 2009; Wang et al. 2008). These results do not clearly and accurately demonstrate how conservation tillage alters the soil microbial community. Therefore, to better understand the effect of conservation tillage on the soil microbial community, it would be valuable to obtain accurate information on soil microbes using a high-throughput sequencing technology to determine the correlation between the soil properties and the microbiome community.
In this study, we collected soil samples from three types of 5-year tillage treatments, specifically plow tillage, as an example of convention tillage, as well as zero tillage and chisel plow tillage, as examples of conservation tillage. Then, we compared the bacterial communities using high-throughput sequencing technology and quantitative PCR. We hypothesized that the different tillage treatments would target distinct soil microbiomes and different soil properties (such as soil texture and nutrient status) and, therefore, would likely influence different characteristics. Consequently, this study had three main objectives: (1) to study the influence of different tillage treatments under the same residue retention on soil physicochemical characteristics; (2) to analyze the effects of different tillage treatments on the enumeration, taxonomic distribution, and phylogenetic composition of soil bacterial communities; and (3) finally, to determine any correlations between the soil variables and changes in the abundance, diversity, and composition of soil bacteria under different tillage treatments.
2 Material and methods
2.1 Experimental site and tillage treatments
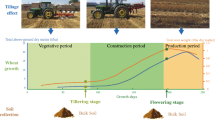
This study was performed at Northwest A&F University, Yangling, Shaanxi, China (34°17′N latitude, 108°04′E longitude at an elevation of 521 m above sea level) on the Eum-Orthrosols (Chinese soil taxonomy) soil, with a mean bulk density of 1.29 g/cm−3. The experimental area was in the central Shaanxi plain (Fig. 1), which belongs to zone 5 (Fig. 1) in the dryland regions of northern China (Wang et al. 2007). The mean annual precipitation is 633 mm, with an average yearly temperature of 13.2 °C. The soil was collected from a long-term trial that began in 2009. The experimental treatments combined different tillage methods on residue retention for croplands with a wheat-maize rotation. The experimental design included a randomized complete block with four replications. Winter wheat (cultivar Shaan mai -139) was planted on October 17, 2013, using wheat drills. Urea fertilizer (N > 46 %) was used as the source of the nitrogen, and phosphorus (P) fertilizer in the form of calcium phosphate (Ca2 (PO4)3 (P = 16 %) was equally applied in all of the treatments at a rate of 750 kg/ha at the time of soil preparation.
The study compared the effects of three different tillage treatments where sowing was carried out on the residues of the previous crop. The main tillage treatment characteristic (Fig. 1) included the following; plow tillage, where the soil was plowed up to 20–30 cm deep using a moldboard plow, followed by the rotavator for the final seedbed preparation; zero tillage, where the tillage treatments were performed to limit the soil disturbance, but to ensure germination we adopted a rotary tillage at 0–5 cm; and finally, chisel plow tillage, where fertilizers were first applied followed by the use of a chisel plow with a depth of 30–35 cm, and 40 cm between lines was used once.
2.2 Soil sampling and physicochemical analysis
Soil samples were collected at depths of 0–20 cm on May 6, 2014, 210 days after the wheat was sown. Each sampling site consisted of five subsamples collected from plots using a standard soil corer, away from plant roots. Samples were sieved through a 2-mm mesh to eliminate large rocks and roots. Each composite soil sample was homogenized and stored at 4 °C for less than 24 h before DNA extraction. We analyzed 12 soil samples (three tillage treatments with four replicate sampling sites per tillage).
The physical and chemical analyses of the soil were performed in the laboratory. Measurements of pH, soil organic carbon, total nitrogen, and soil texture were performed as described by Zhao et al. (2014). The soil concentrations of inorganic nitrogen (nitrate and ammonia) and dissolved organic carbon and nitrogen were determined using the procedure described by Berthrong et al. (2013). The soil volumetric water content was measured at sampling points (five times per sample, averaged) using a FieldScout TDR 100 Soil Moisture Meter with Case (Spectrum Technologies Inc., Plainfield, USA). The urease and invertase activities were assayed using 5-g soil samples containing an appropriate substrate (added to the soil) and incubated for 24 h (37 °C) at the optimal pH for each enzyme type, as described by Gu et al. (2009). The catalase activities were determined as described by Trasar-Cepeda et al. (1999).
2.3 DNA extraction, bacterial 16S gene PCR amplification, and Illumina sequencing
Microbial DNA was extracted from 3 g of fresh soil thrice (using a 1-g sample each time) using an E.Z.N.A.® Soil DNA kit (Omega Bio-Tek, Inc., Norcross, GA) according to the manufacturer’s instructions. The concentration and quality of the DNA were detected using a spectrophotometer (NanoDrop2000, Thermo Scientific, Wilmington, DE, USA). Primers pairs F515 (5′-GTG CCA GCM GCC GCG GTA A-3′) and R806 (5′-GGA CTA CHV GGG TWT CTA AT-3′) targeting the V4 region of the 16S rRNA gene were used for PCR (Peiffer et al. 2013). This primer set provided a comprehensive coverage with the highest taxonomical accuracy of the bacterial sequence. In addition, the reverse primer contained a 6-bp error-correcting barcode unique to each sample. The protocol for PCR amplification of the 16S rRNA gene was described previously (Caporaso et al. 2010). Each PCR product was subjected to pyrosequencing using the Illumina MiSeq platforms at Majorbio BioPharm Technology Co., Ltd., Shanghai, China.
FLASH software was used to merge the pairs of reads from the original DNA fragments (Caporaso et al. 2010). Further sequence analysis was performed using the USEARCH v5.2.32 to filter and eliminate noise from the data by clustering similar sequences with less than 3 % dissimilarity and the Quantitative Insights Into Microbial Ecology pipeline software to select 16S rRNA operational taxonomic units from the combining reads of clustered operational taxonomic units with 97 % similarity (Edgar 2010).
2.4 Quantitative PCR analyses
The relative abundances of 16S rRNA genes were measured for bacteria by quantitative PCR with the bacterial-specific primer pairs F515 (5′-GTG CCA GCM GCC GCG GTA A-3′) and R806 (5′-GGA CTA CHV GGG TWT CTA AT-3′) (Berthrong et al. 2013). The abundances of the bacterial 16S rRNA genes were quantified using standard curves generated from 10-fold serial dilutions of cloned full-length copies of the 16S rRNA gene. The 20 μL quantitative PCR reactions contained 10 μL EvaGreen 2× qPCR MasterMix (Applied Biological Materials Inc., Richmond, Canada), each primer (final concentration 0.3 μM), and bacterial, environmental, or standard DNA templates (2 μL per reaction). The quantitative PCR was carried out using a Bio-Rad C1000/CFX96 Thermocycler (Bio-Rad, Hercules, CA) using the following three-step thermocycler protocols: initial denaturation at 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s, 53 °C for 30 s, and 70 °C for 60 s. All quantitative PCR reactions were run in triplicate with each DNA template. The amplification efficiency of the quantitative PCR was 86–97 % (R 2 > 0.992).
2.5 Statistical and bioinformatics analysis
Alpha diversity was calculated using Mothur to estimate richness with the Chao1 estimator and the abundance-based estimators, while we estimated diversity using the Shannon diversity and Simpson diversity indices. For beta diversity, taxonomic and phylogenetic community comparisons were performed using Bray-Curtis and weighted and unweighted UniFrac distance matrices. Non-metric multi-dimensional scaling was used to illustrate the clustering of the different samples. Analysis of similarities was performed to test the significance of separation between different tillage treatments.
To study the relationship between the soil characteristics and the abundance of dominant phyla and the four classes of Proteobacteria, we used a redundancy analysis. Correlations between the soil bacterial community structure and soil characteristics were determined using Mantel tests with 999 permutations.
All statistical analyses and Spearman’s rank correlations between the abundant phyla and soil properties were calculated using the statistical package for the social sciences version 22.0 software packages (SPSS Inc.). Non-metric multi-dimensional scaling, analysis of similarities, redundancy analysis, and Mantel tests were performed using “vegan” packages in the R v3.20 statistical environment. A P < 0.05 was considered statistically significant.
2.6 Sequence accession numbers
The 16S rRNA gene sequences obtained in this study have been deposited in the National Center for Biotechnology Information Sequence Read Archive database with the accession number SRP058463.
3 Results and discussion
This study was focused on the response of soil bacterial communities to conservation tillage, which was considered to significantly improve soil texture, moisture, and nutrient levels in northern Chinese dryland regions (He et al. 2007; Wang et al. 2007). Here, we used ecological and statistical methods to determine the soil properties affected by conservation (chisel plow and zero tillage) and convention (plow) tillage treatments and the main factors contributing to altering soil bacterial (alpha and beta) diversity and community composition.
3.1 Tillage treatment contributes to differences in soil characteristics
The edaphic properties differed significantly between tillage treatments (Table 1). We found that chisel plow tillage and zero tillage soils had a significantly higher fraction of clay and a lower fraction of silt compared with plow tillage soils. Conservation tillage treatments increased the soil moisture and had a considerably significant relationship with the soil clay fraction (r = 0.726, P = 0.005), consistent with the finding of a previous study (Prakash et al. 2010). Chisel plow and zero tillage significantly increased the organic carbon content of the soil, which can be explained by the alterations in soil aggregation mediated by clay-size particles and an increase in the crop residues, leading to sequestration of soil carbon (Neumann et al. 2013). Based on those findings, we hypothesized that changes in soil properties shifted the distribution of the soil bacterial communities.
3.2 Soil properties modulate, soil bacterial alpha, and beta diversity
Statistically significant differences in the soil bacterial richness and diversity under different tillage treatments were observed using the Shannon and Simpson indices. Compared to the plow tillage treatment, the chisel plow and zero tillage treatments strongly increased the Simpson index by 378 % (average) and decreased the Shannon index by 6 % (average). The relationship between the bacterial diversity indices and abundance produced humped (Shannon H’, Fig. 2a) and U-shaped (Simpson’s index, Fig. 2a) patterns reflecting conservation and conversion tillage. At the vertex of both two patterns, we found that plow tillage (Fig. 2a), which may moderately perturb the microbiota and decrease the soil texture and moisture content, decreased in the competitive niche exclusion and selection mechanisms between populations. Such consequences could lead to an increase in the H’ value and a decrease in Simpson’s index (Lienhard et al. 2013). Despite the high similarity of the alpha diversity indices, the abundance of 16S rRNA genes in the zero tillage soils was greater than it was in chisel plow tillage (3.98 × 107 and 1.94 × 107 copies g·dry soil−1, respectively). This observation can be explained by similarities in the soil texture and moisture between the zero and chisel plow tillage. The soil texture and moisture are principle factors affecting soil pore connectivity, which leads to changes in bacterial diversity, and are significantly related to the Shannon (P = 0.03) and Simpson (P = 0.01) indices (Carson et al. 2010). The zero tillage, which caused less soil disturbance than the chisel plow tillage did, could provide a suitable environment for multiplying soil bacteria. Consequently, our study supports the view that conservation tillage treatments strongly affect soil texture and altered soil moisture content, causing changes in alpha diversity and the abundance of soil bacteria.
a Schematic representation based on the regression model of the relationship between the logarithm of 16S rRNA gene copy number and Shannon’ diversity index (H’ value) and Simpson’s diversity index. b Significant (P < 0.05) taxonomic and phylogenetic similarity of soil bacterial community under tillage treatments using analysis of similarities. Significance of taxonomic and phylogenetic similarity between pairwise tillage treatments (CPT and ZT, CPT and PT, ZT and PT) were tested. *0.01 < P < 0.05, ** P < 0.01, ns not significant, PT plow tillage, ZT zero tillage, CPT chisel plough tillage
The soil bacterial beta-diversity was analyzed using non-metric multidimensional scaling and analysis of similarities between both the taxonomic and phylogenetic measures. Our result indicated that the bacterial phylogenetic compositions (weighted UniFrac) under both conservation tillage treatments—chisel plow tillage (r = 1, P = 0.028) and zero tillage (r = 1, P = 0.026)—significantly and strongly differed from those under plow tillage (Fig. 2b). Furthermore, the soil bacterial phylogenetic composition (weighted UniFrac) and the phylogenetic membership (unweighted UniFrac) under chisel plow and zero tillage (r = 0.177 and 0.49, P = 0.064 and 0.031, respectively) showed a close similarity to plow tillage (Fig. 2b). Although the bacterial taxonomic composition under the three tillage treatments had a similar polygenetic composition, the zero tillage versus plow tillage (r = 0.99, P = 0.03) was much more different from the chisel plow versus plow tillage (r = 0.49, P = 0.05). The significance of this variation can be explained by changes in soil texture. A Mantel test revealed that the soil texture was significantly related to soil bacterial beta-diversity in regards to phylogenetic composition (sand, r = 0.451), membership (silt, r = 0.459; sand, r = 0.195), and taxonomic composition (silt, r = 0.340; sand, r = 0.340). The similar soil texture (clay and silt) fraction, which correlated with the differences in the soil moisture and nutrient concentrations, contributed to the analogous polygenetic and taxonomic composition between chisel plow tillage and zero tillage. The differences in silt and sand fraction under zero tillage suggest that it was a safer agriculture treatment to improve and preserve soil microbial diversity (Ceja-Navarro et al. 2010), leading to higher nutrient levels (organic carbon) than plow tillage was in our study. This can explain why larger differences in both bacterial polygenetic and taxonomic diversity were found between the zero tillage and plow tillage (Fig. 2b) than there were in other tillage strategies. Consequently, our findings suggest that conservation tillage alters the soil silt and sand fraction, thereby causing considerable changes in the soil nutrients and ultimately changing the soil bacterial beta-diversity.
3.3 Effects of different tillage protocols on soil bacterial taxonomic distribution
Our analysis of all the soil samples yielded 201,264 quality sequences in total and 9455−23,023 sequences per sample (mean = 16,772), and 97.3 % of the sequences were classified at the phylum level. We found that the phyla Proteobacteria, Actinobacteria, and Acidobacteria were the most abundant in the samples, which was consistent with several other studies (Nacke et al. 2011; Navarro-Noya et al. 2013). Furthermore, the phylum Firmicutes were highly abundant in the zero and chisel plow tillage soils (Fig. 3a). Significant differences in soil bacterial composition were observed between the conservation and convention tillage soils (Fig. 3a). In particular, high relative abundances of Firmicutes were observed in chisel plow and zero tillage soils, while Actinobacteria, Acidobacteria, Alphaproteobacteria, Betaproteobacteria, and Chloroflexi were richer in plow tillage soils.
a Relative abundance of the dominant bacterial phyla (Proteobacterial classes) in all soil samples combined and in each tillage treatments. Relative abundances are based on the proportional frequencies of those DNA sequences that could be classified. Different letters indicate significant differences (ANOVA, P < 0.05, Tukey’s HSD post-hoc analysis) among tillage treatments. b Spearman’s rank correlation coefficients showed that the soil physical-chemical characteristics were significant (P < 0.05) positively and negatively correlated with the relative abundance of the dominant bacterial phyla. *0.01 < P < 0.05, ** P < 0.01, ns not significant
To explain why some bacterial phyla are more abundant in some soils than others, the concept of copiotrophic versus oligotrophic bacteria has been proposed (Fierer et al. 2007). Firmicutes have been described as fast-growing copiotrophs that thrive in environments with high carbon availability (Rodrigues et al. 2013). Among the classes of Firmicutes, the relative abundance of Bacilli, which are primarily obligate aerobes (Zhao et al. 2014), was significantly higher in the chisel plow and zero tillage soils than it was in the plow tillage. The genus Bacillus, which belongs to the class Bacilli and possesses numerous growth-promoting characteristics and is adapted to local conditions (Abiala et al. 2015), was the most represented among the Firmicutes in chisel plow and zero tillage (85 % each). Navarro-Noya et al. (2013) reported that zero tillage increases the relative abundance of Firmicutes, but a study by Lienhard et al. (2013) found that convention tillage increases the abundance of Firmicutes due to their ability to produce endospores. In our study, the soil texture (silt/sand fraction) was significantly correlated with the abundance of Firmicutes (Fig. 3b) and was significantly higher in the chisel plow and zero tillage treatments than it was in the plow tillage treatment. The conservation tillage treatments produced soil with better ventilation and nutrient status than the convention tillage did, which may explain the higher relative abundance of Firmicutes in chisel plow and zero tillage. Despite the Proteobacteria having been described as fast-growing copiotrophs that are stimulated in carbon-rich environments (Jenkins et al. 2010), competition for nutrients with Firmicutes may explain the reduced numbers of Alphaproteobacteria and Betaproteobacteria we observed under the chisel plow and zero tillage compared to the plow tillage. In particular, our study found that more than 42 % of the Alphaproteobacteria were of the order Rhizobiales, which are regarded as heterotrophic, nitrogen-fixing organisms (Li et al. 2014). This result was significantly correlated with the changes in the soil texture and nutrient conditions (Fig. 3b) and may explain why the relative abundance of Rhizobiales was higher in the chisel plow and zero tillage (44 % and 46 %, respectively) than it was in plow tillage (42 %) soils. Actinobacteria and Acidobacteria are considered as oligotrophs (Pascault et al. 2013), and their similar soil textures and nutrient levels may explain their lower abundance in the chisel plow and zero tillage than in the plow tillage soils. Conservation tillage treatments alter soil texture fraction and improve the aeration and nutrient status, thereby increasing the relative abundance of profitable functional bacteria species.
Considering the entire bacterial community composition, our study revealed a significant correlation between soil properties and the dominant phyla identified (r = 0.518, P = 0.002; Fig. 4). In addition, the soil properties can explain the variation (81 %, Fig. 4) in bacterial composition between the conservation (chisel plow and zero tillage) and convention (plow) tillage. The soil texture (silt/sand fraction) had stronger effects (a longer arrow) on the composition of the bacterial communities (Fig. 4) than other properties did. Indeed, the soil texture was significantly (Fig. 3b) correlated with the relative abundances of the dominant bacterial phyla (class) including the Acidobacteria, Firmicutes, Actinobacteria, Chloroflexi, and Alphaproteobacteria. These results partially support the notion that soil texture is the strongest factor in structuring bacterial communities (Bach et al. 2010; Carson et al. 2010). Our study suggests that the soil texture fraction is the principal factor that plays an important role in determining soil nutrient status and moisture (Davinic et al. 2012; Li et al. 2014) and contributes to the differences in soil bacterial composition between conservation (chisel plow and zero tillage) and convention (plow) tillage.
Redundancy analysis of abundant phyla (Proteobacterial classes, diamond) and 11 soil characteristics (arrows) soil texture (clay/silt/san fraction), soil organic carbon (SOC), volumetric water content (VHC), dissolved organic nitrogen (DON), total nitrogen (TN), NH4-N, urease, invertase, and catalase of different soils from tillage treatments. PT plow tillage (circle), ZT zero tillage (square), CPT chisel plough tillage (triangle). *0.01 < P < 0.05, ** P < 0.01
4 Conclusions
In this study, we found that 5-year tillage treatments significantly changed the soil texture, moisture, and nutrient levels, as well as modified the soil bacterial diversity and community composition of soil from the drylands of China. Although a larger Shannon’s index value was found in plow tillage soil, a more diverse soil bacterial community was observed in the conservation (chisel plow and zero) tillage soils, which had abundant functional microorganisms (including Alphaproteobacteria/Rhizobiales and Firmicutes/Bacillus) and relatively large Simpson’s indexes. Most importantly, our results demonstrate that conservation tillage improved the soil texture fraction, which was the main factor that increased the soil moisture and improved the aeration and nutrient status, contributing to the changes in the soil bacterial community. We believe our findings in this study can enhance the understanding of the role of conservation tillage in altering soil bacterial community and contribute to building a stable and functional soil environment. Furthermore, this knowledge may contribute to producing sustainable crops in the dryland regions of northern China.
References
Abiala MA, Odebode AC, Hsu SF, Blackwood CB (2015) Phytobeneficial properties of bacteria isolated from the rhizosphere of maize in southwestern Nigerian soils. Appl Environ Microbiol. doi:10.1128/AEM.00570-15, AEM.00570-00515
Bach EM, Baer SG, Meyer CK, Six J (2010) Soil texture affects soil microbial and structural recovery during grassland restoration. Soil Biol Biochem 42(12):2182–2191. doi:10.1016/j.soilbio.2010.08.014
Berthrong ST, Buckley DH, Drinkwater LE (2013) Agricultural management and labile carbon additions affect soil microbial community structure and interact with carbon and nitrogen cycling. Microb Ecol 66(1):158–170. doi:10.1007/s00248-013-0225-0
Brussaard L, de Ruiter PC, Brown GG (2007) Soil biodiversity for agricultural sustainability. Agr Ecosyst Environ 121(3):233–244. doi:10.1016/j.agee.2006.12.013
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. doi:10.1038/nmeth.f.303
Carson JK, Gonzalez-Quinones V, Murphy DV, Hinz C, Shaw JA, Gleeson DB (2010) Low pore connectivity increases bacterial diversity in soil. Appl Environ Microbiol 76(12):3936–3942. doi:10.1128/AEM.03085-09
Ceja-Navarro JA, Rivera-Orduna FN, Patino-Zuniga L, Vila-Sanjurjo A, Crossa J, Govaerts B, Dendooven L (2010) Phylogenetic and multivariate analyses to determine the effects of different tillage and residue management practices on soil bacterial communities. Appl Environ Microbiol 76(11):3685–3691. doi:10.1128/AEM.02726-09
Chen H, Hou R, Gong Y, Li H, Fan M, Kuzyakov Y (2009) Effects of 11 years of conservation tillage on soil organic matter fractions in wheat monoculture in Loess Plateau of China. Soil Tillage Res 106(1):85–94
Davinic M, Fultz LM, Acosta-Martinez V, Calderon FJ, Cox SB, Dowd SE, Allen VG, Zak JC, Moore-Kucera J (2012) Pyrosequencing and mid-infrared spectroscopy reveal distinct aggregate stratification of soil bacterial communities and organic matter composition. Soil Biol Biochem 46:63–72. doi:10.1016/j.soilbio.2011.11.012
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461. doi:10.1093/bioinformatics/btq461
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364
Gu Y, Wang P, Kong CH (2009) Urease, invertase, dehydrogenase and polyphenoloxidase activities in paddy soil influenced by allelopathic rice variety. Eur J Soil Biol 45(5-6):436–441. doi:10.1016/j.ejsobi.2009.06.003
He J, Li H, Wang X, McHugh AD, Li W, Gao H, Kuhn NJ (2007) The adoption of annual subsoiling as conservation tillage in dryland maize and wheat cultivation in northern China. Soil Till Res 94(2):493–502. doi:10.1016/j.still.2006.10.005
Jenkins SN, Rushton SP, Lanyon CV, Whiteley AS, Waite IS, Brookes PC, Kemmitt S, Evershed RP, O’Donnell AG (2010) Taxon-specific responses of soil bacteria to the addition of low level C inputs. Soil Biol Biochem 42(9):1624–1631. doi:10.1016/j.soilbio.2010.06.002
Kladivko EJ (2001) Tillage systems and soil ecology. Soil Till Res 61(1-2):61–76. doi:10.1016/S0167-1987(01)00179-9
Li H, Ye DD, Wang XG, Settles ML, Wang J, Hao ZQ, Zhou LS, Dong P, Jiang Y, Ma ZS (2014) Soil bacterial communities of different natural forest types in Northeast China. Plant Soil 383(1-2):203–216. doi:10.1007/s11104-014-2165-y
Lienhard P, Terrat S, Prévost-Bouré NC, Nowak V, Régnier T, Sayphoummie S, Panyasiri K, Tivet F, Mathieu O, Levêque J, Maron P-A, Ranjard L (2013) Pyrosequencing evidences the impact of cropping on soil bacterial and fungal diversity in Laos tropical grassland. Agron Sustain Dev 34(2):525–533. doi:10.1007/s13593-013-0162-9
Nacke H, Thurmer A, Wollherr A, Will C, Hodac L, Herold N, Schoning I, Schrumpf M, Daniel R (2011) Pyrosequencing-based assessment of bacterial community structure along different management types in German forest and grassland soils. PLoS One 6(2):e17000. doi:10.1371/journal.pone.0017000
Navarro-Noya YE, Gomez-Acata S, Montoya-Ciriaco N, Rojas-Valdez A, Suarez-Arriaga MC, Valenzuela-Encinas C, Jimenez-Bueno N, Verhulst N, Govaerts B, Dendooven L (2013) Relative impacts of tillage, residue management and crop-rotation on soil bacterial communities in a semi-arid agroecosystem. Soil Biol Biochem 65:86–95. doi:10.1016/j.soilbio.2013.05.009
Neumann D, Heuer A, Hemkemeyer M, Martens R, Tebbe CC (2013) Response of microbial communities to long-term fertilization depends on their microhabitat. FEMS Microbiol Ecol 86(1):71–84. doi:10.1111/1574-6941.12092
Pascault N, Ranjard L, Kaisermann A, Bachar D, Christen R, Terrat S, Mathieu O, Leveque J, Mougel C, Henault C, Lemanceau P, Pean M, Boiry S, Fontaine S, Maron PA (2013) Stimulation of different functional groups of bacteria by various plant residues as a driver of soil priming effect. Ecosystems 16(5):810–822. doi:10.1007/s10021-013-9650-7
Pastorelli R, Vignozzi N, Landi S, Piccolo R, Orsini R, Seddaiu G, Roggero PP, Pagliai M (2013) Consequences on macroporosity and bacterial diversity of adopting a no-tillage farming system in a clayish soil of Central Italy. Soil Biol Biochem 66:78–93. doi:10.1016/j.soilbio.2013.06.015
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL, Buckler ES, Ley RE (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci U S A 110(16):6548–6553. doi:10.1073/pnas.1302837110
Prakash R, Singh D, Pathak NP (2010) The effect of soil texture in soil moisture retrieval for specular scattering at c-band. Prog Electromagnetics Res-Pier 108:177–204. doi:10.2528/pier10050403
Rodrigues JL, Pellizari VH, Mueller R, Baek K, Jesus Eda C, Paula FS, Mirza B, Hamaoui GS Jr, Tsai SM, Feigl B, Tiedje JM, Bohannan BJ, Nusslein K (2013) Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc Natl Acad Sci U S A 110(3):988–993. doi:10.1073/pnas.1220608110
Roger-Estrade J, Anger C, Bertrand M, Richard G (2010) Tillage and soil ecology: partners for sustainable agriculture. Soil Till Res 111(1):33–40. doi:10.1016/j.still.2010.08.010
Trasar-Cepeda C, Camiña F, Leirós MC, Gil-Sotres F (1999) An improved method to measure catalase activity in soils. Soil Biol Biochem 31(3):483–485. doi:10.1016/S0038-0717(98)00153-9
Wang Q, Bai Y, Gao H, He J, Chen H, Chesney R, Kuhn N, Li H (2008) Soil chemical properties and microbial biomass after 16 years of no-tillage farming on the Loess Plateau, China. Geoderma 144(3):502–508
Wang XB, Cai DX, Hoogmoed WB, Oenema O, Perdok UD (2007) Developments in conservation tillage in rainfed regions of North China. Soil Till Res 93(2):239–250. doi:10.1016/J.Still.2006.05.005
Zhao J, Zhang R, Xue C, Xun W, Sun L, Xu Y, Shen Q (2014) Pyrosequencing reveals contrasting soil bacterial diversity and community structure of two main winter wheat cropping systems in China. Microb Ecol 67(2):443–453. doi:10.1007/s00248-013-0322-0
Acknowledgments
The National Natural Science Foundation of China (Project Nos. 31571621 and 31471451) and the Fundamental Research Funds for the Central Universities supported this work. We are grateful to the Northwest A&F University for providing the experimental site. We also thank anonymous reviewers and the editors for the constructive comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Wang, Z., Liu, L., Chen, Q. et al. Conservation tillage increases soil bacterial diversity in the dryland of northern China. Agron. Sustain. Dev. 36, 28 (2016). https://doi.org/10.1007/s13593-016-0366-x
Accepted:
Published:
DOI: https://doi.org/10.1007/s13593-016-0366-x