Abstract
Purpose
Highly aggressive adult soft tissue sarcomas (STS), i.e., leiomyosarcomas (LMS) and undifferentiated pleomorphic sarcomas (UPS), present complex genomic anomalies and overall 5-year survival rates of 20 to 40%. Here, we aimed to identify new biomarkers that may be employed to improve the treatment of non-translocation STS patients. We validated 12 miRNAs implicated in tumor development using primary STS samples and selected miR-152 for further analysis in STS-derived cell lines.
Methods
59 primary STS samples (27 LMS and 32 UPS) and 10 matched normal control tissues were included in the study, as well as 3 STS-derived cell lines (HT1080, SW872 and SKLMS1) and a normal control mesenchymal cell line (hMSC). miRNA expression analyses were performed using a TaqMan microRNA Array platform and qRT-PCR (miR-152), respectively. The expression levels of the putative miR-152 targets MET and KIT were assessed using qRT-PCR and immunohistochemistry on tissue microarrays, respectively. In addition, various functional analyses were performed before and after miR-152 transfection into SKLMS1 cells.
Results
We found that 12 pre-selected miRNAs were down-regulated in primary STS tumor samples compared to its normal control samples. A statistically significant miR-152 down-regulation was found to be accompanied by high MET and KIT mRNA levels in both the primary samples and the STS-derived cell lines tested. miR-152 transfection in SKLMS1 cells led to a reduction in KIT and MET mRNA and protein levels which, in turn, was associated with a transient down-regulation of the PI3K/AKT pathway, a transient decrease in cell growth, and a transient increase in both apoptotic and S-phase cells.
Conclusions
Our data indicate that over-expression of MET and KIT in primary STS samples and its derived cell lines is associated with miR-152 down-regulation. This shift may play a role in STS development and, thus, may be used to identify patients at risk. The effect of MET down-regulation on downstream signaling pathways, such as the PI3K/AKT pathway, may provide a basis for the future design of novel STS treatment strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Malignant soft tissue sarcomas (STS) represent a heterogeneous group of mesenchymal tumors that comprise 1% of all adult cancers. The biological and pathological knowledge of these tumors is still incomplete. Around 50% of the STS patients have a life expectancy of less than 5 years [1], and the delineation of the metastatic potential of the tumors is primarily based on histopathologic parameters, including tumor grade, size and depth of local tissue infiltration. Leiomyosarcomas (LMS) exhibit several features of the smooth muscle lineage, including positive immunostaining for vimentin, pan-muscle actin, smooth muscle actin, desmin and h-caldesmon [2, 3]. The overall incidence of LMS increases with age (median age seventy years). Undifferentiated pleomorphic sarcoma (UPS) is characterized by the occurrence of spindle-shaped tumor cells that exhibit a diffuse pleomorphism [4], a fibroblastic or myofibroblastic appearance [5, 6] and vimentin positivity. UPS tumors tend to occur in the extremities and are mostly diagnosed late in adult life. Genetic studies have revealed similar recurrent DNA alterations in UPS and LMS tumors, including TP53 mutations, RB1 deletions and MDM2 and CDK4 amplifications [7–9]. The high aggressiveness of both histotypes and their notorious resistance to chemotherapy are considered responsible for their dismal 5-year overall survival rates, which range from 20 to 40%. Thus, a more profound knowledge of the mechanisms underlying these pathobiological features may be employed to improve the clinical management of these high-risk patients.
In a recent study a set of 67 genes was identified that can be used to predict metastasis in non-translocation STS cases [10]. It was found that the expression of these genes correlated with genome complexity and a poor prognosis. microRNAs (miRNAs) are non-coding RNAs that are involved in post-transcriptional gene expression regulation. As such, they may control important physiological processes such as cellular development, differentiation and signaling [11–14]. Anomalous miRNA expression has been found to be strongly correlated with malignancy [15] and in case of STS a strong association has been reported between miRNA expression, patient age and prognosis [16]. Previously, we identified 20 significantly deregulated miRNAs in both LMS and UPS [9]. Here, we aimed at identifying activated pathways that may be employed for differential STS diagnosis and treatment. We validated 12 cancer-related miRNAs and found that miR-152 was significantly down-regulated in non-translocation primary STS tumor samples (UPS and LMS) and STS-derived cell lines compared to controls. In addition, we show that ectopic miR-152 expression may reduce STS growth through down-regulation of its targets MET and KIT and, indirectly, through deregulation of the PI3K/AKT signaling pathway.
2 Materials and methods
2.1 Tumor specimens and cell lines
59 primary high-grade STS tumor samples (27 LMS and 32 UPS) were referred to the Rizzoli Orthopaedic Institute. Fresh and paraffin-embedded non necrotic tissues (≥ 90% viable tumor cells) were used and their diagnosis was established based on hematoxylin-eosin stained samples according to histopathological criteria. In Table 1 various clinicopathological data are listed, i.e., 32 males and 27 females with a mean age of 63 years; 33 patients developed metastases during a minimum follow-up period of 4 years. Ten non-oncologic tissues were also included as controls. All samples were treated in accordance with the authorizations issued by the Ethical Committee of the Rizzoli Orthopedic Institute and informed consent was obtained from all patients.
The human fibrosarcoma (HT1080), liposarcoma (SW872), leiomyosarcoma (SKLMS1) and normal mesenchymal (hMSC)-derived cell lines were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA; no. CCL-121™, HTB-92™, HTB-88™ and CRL-1486™, respectively) and they were routinely cultured in IMDM medium supplemented with 10% FBS, L-glutamine (2 mM), 100 U/ml penicillin and 100 μg/ml streptomycin (Invitrogen) at 37°C in a 5% CO2 humidified incubator.
2.2 RNA extraction
Total RNA was extracted from 59 STS specimens, 10 normal tissues, 3 STS-derived cell lines and a normal mesenchymal cell line using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol and stored at −80°C in RNAsecure reagent (Ambion, Inc., Austin, TX, USA) until use. The concentrations of the total RNAs were measured using a spectrophotometer, and their purity and quality were checked through denatured gel electrophoresis.
2.3 miRNA expression analysis in primary STS samples
Single stranded cDNA was generated from total RNA by reverse transcription using an Applied Biosystems TaqMan MicroRNA Multi Reverse Transcription Human RT Primer pool (cod. 4384791) following the manufacturer’s protocol. PCR pre-amplification of cDNA was performed using a TaqMan® PreAmp Master Mix Kit (Applied Biosystems, Life Technologies, CA, USA; cod. 4384267) on an Applied Biosystems 7900 Real-Time PCR system. The samples were incubated at 50°C for 2 min and at 94.5°C for 10 min, followed by 40 cycles of 97°C for 30 sec. and 59.7°C for 1 min. Next, miRNA expression levels were analyzed using an Applied Biosystems TaqMan® Custom MicroRNA Array (Applied Biosystems, Life Technologies, CA, USA; cod. 4449137), containing 12 selected (miRBase) miRNAs (miR-137, miR-21.5p, miR-100.5p, miR-409.5p, miR-152, miR-376a.3p, miR-487a, miR-1305, miR-212.3p, miR-892b, miR-223.3p, miR-138.5p.) and 2 small nucleolar RNAs (RNU44 and RNU48) as endogenous controls for data normalization. The array analyses were carried out according to the manufacturer’s protocol. Arrays were run in duplicate for each sample to allow technical variability assessment, and relative quantifications were carried out using the 2-∆∆CT method (Applied Biosystems, User Bulletin No. 2, P/N 4303859). Raw miRNA array data were analyzed using the SDS 2.3 software tool (Applied Biosystems). miRNA expression levels were normalized to the RNU44 and RNU48 median values, and the median ΔCT of normal tissues was used as calibrator. The ∆CT values were subjected to a Mann-Whitney test and corrected by false discovery rate (FDR) for multiple testing.
2.4 miRNA expression analysis in STS-derived cell lines
miR-152-specific reverse transcription and RT-PCR (TaqMan miRNA assay no. 000475; Applied Biosystems, Life Technologies, CA, USA) were carried out on RNA extracted from HT1080, SW872, SKLMS1and hMSC cells following the TaqMan MicroRNA Assay Protocol. Next, miR-152 expression levels were quantified using the 2-∆∆CT method with hMSC as calibrator, and the data were normalized to the endogenous reference RNU44 according to the Applied Biosystems protocol (TaqMan miRNA assay no. 4373384).
2.5 MET and KIT mRNA expression analysis in primary patient samples
The mRNA levels of the miR-152 targets MET and KIT were determined in 38 STS tumor samples and 10 normal tissues using the TaqMan Reverse Transcription Protocol (Applied Biosystems). Quantitative RT-PCR (qRT-PCR) was performed on an ABI7900 sequence detection system (Applied Biosystems). The MET and KIT expression levels were quantified using TaqMan Expression Assays (Hs01565584_m1 and Hs00174029_m1, respectively) according to the manufacturer’s protocol (Applied Biosystems). The MET and KIT expression levels were calculated using the 2-∆∆CT method and normalized to that of the housekeeping gene ACTB (Hs99999903_m1; TaqMan Expression Assays; Applied Biosystems). A pool of normal lymphocytes was used as calibrator.
2.6 Tissue microarray construction and immunohistochemistry
The MET and KIT protein expression patterns were evaluated by immunohistochemistry (IHC) on a tissue microarray (TMA) containing the 59 STS samples (see above) in duplicate for each patient. In order to select representative areas for TMA construction using a TMA Master system (3D Histech Ltd., Budapest, Hungary), hematoxylin and eosin (H&E)-stained slides were reviewed by a pathologist (P.P.). After antigen retrieval and blocking with a ready-to-use Universal Block mixture (catalog no. 71–00-61; KPL, Gaithersburg, MD, USA) for 20 min. at room temperature, TMAs were incubated at 4°C overnight with rabbit polyclonal anti-MET antibody C-28 (Cat. # sc-161, 1:300; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and rabbit monoclonal anti-KIT antibody D13A2 (1:50; Cell Signaling Technology, Danvers, MA, USA). As a negative control the antibodies were omitted during the primary incubations. Finally, IHC staining was performed using Dako LSAB + System-HRP followed by treatment with 3,3-diaminobenzidine (DAB) as chromogen (Dako, Milan, Italy). The TMAs were counterstained with Meyer’s hematoxylin.
According to the percentage of positive tumor cells, the samples were scored as 1 (< 50% positive cells) or 2 (≥ 50% positive cells). Staining intensities were scored as 0 (no visual staining), 1 (weak staining), 2 (moderate staining) or 3 (strong staining). The cut-off levels for the sum of the scores were applied as follows: 0–1 for negative, 2–3 for weak to moderate positivity in <50% of the tumor cells, and 4–5 for moderate to strong positivity in >50% of tumor cells. The latter was considered as protein over-expression. Non-oncologic tissues were used as controls.
2.7 Transfection of miR-152 precursor molecules
For each transfection, SKLMS1 cells were seeded at a density of 2.5 × 105 cells/well in 6-well plates. The transfections were performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). In each well both 50 nM miR-152 precursor (cod. PM12269; Ambion Inc., Austin, TX, USA) and negative control miRNA precursor (scramble) (cod. AM17110; Ambion) molecules were transfected into the cells. The transfection efficiencies were monitored after 12 h, 24 h and 48 h by flow cytometry (FACS Calibur, BD, San Jose, CA, USA) using Cy™3 and FAM™ dye-labeled Pre-miR Negative Controls (cod. AM17120; Ambion) and RT-PCR according to the TaqMan MicroRNA Assay Protocol (Applied Biosystems; miR-152 assay no. 000475).
2.8 Cell growth assay
The number of adherent, viable cells was determined microscopically using a Neubauer cell counting chamber. After 24 h, 48 h and 72 h of transfection, the cells were washed once in phosphate buffered saline (PBS) and harvested by trypsinization, after which viable cell numbers were determined using a Trypan Blue exclusion assay. Cell viability was defined as the percentage of cells that excluded 0.2% Trypan Blue.
2.9 Apoptosis assay
SKLMS1 cells were was transfected with miR-152 as described above and collected after 24 h, 48 h and 72 h. Next, the cells were washed twice in PBS, re-suspended in 0.1 ml binding buffer (10 mM HEPES, 140 mM NaCl, 5 mM CaCl2, pH 7.4), incubated for 20 min at room temperature with 2 μl annexin V-FITC supplemented with 10 μl propidium iodide (PI) (10 μg/ml) and, finally, analyzed by flow cytometry (FACS Calibur, BD, San Jose, CA, USA) using channel 1 for annexin V-FITC staining and channel 2 for PI staining. The experiments were repeated three times.
2.10 Cell cycle analysis
Cell cycle distributions were determined using propidium iodide (PI) staining in conjunction with flow cytometry. Briefly, after 24 h, 48 h and 72 h of transfection, cells were harvested by trypsinization and fixed with 70% ethanol. Next, the cells were stained for total DNA content with a solution containing 20 μg/ml PI, after which the cell cycle distribution was assessed using a FACScan flow cytometer (Becton Dickinson, San Jose, CA, USA).
2.11 MET and KIT mRNA expression analysis after miRNA transfection
Reverse transcription of mRNA extracted from SKLMS1 cells before and after miR-152 and scramble miRNA (see above) transfection was carried out using a High Capacity cDNA Archive kit (Applied Biosystems, Life Technologies, CA, USA) according to the manufacturer’s instructions. qRT-PCR was performed using an ABI 7900 sequence detection system (Applied Biosystems) according to the manufacturer’s protocol. The MET and KIT mRNA expression levels were calculated by the 2-∆∆CT method using TaqMan Expression Assays Hs01565584_m1 and Hs00174029_m1, respectively, and normalized to that of the housekeeping gene ACTB (Hs99999903_m1; TaqMan Expression Assays, Applied Biosystems).
2.12 Western blot analysis
According to standard procedures, 50 μg protein extracts were prepared from SKLMS1 cell lysates before transfection and 24 h after transfection and subjected to 12% SDS-PAGE. After nitrocellulose blot transfer, the resulting membranes were incubated with anti-MET antibody C-28 (Santa Cruz Biotechnology, Santa Cruz, CA, USA) (dilution 1:200), anti-KIT antibody D13A2 (Cell Signaling Technology, Danvers, MA, USA) (dilution 1:200), anti-PI3K antibody C73F8 (Cell Signaling Technology) (dilution 1:1000) anti p-AKT antibody Ser473-D9E (Cell Signaling Technology) (dilution 1:1000) and anti-AKT antibody 11E7 (Cell Signaling Technology) (dilution 1:1000). The resulting signals were visualized using an Immobilon Western Chemiluminescent HRP substrate (Millipore, Billerica, MA, USA), and quantified by densitometric analysis using a GS-800 imaging densitometer in conjunction with the Quantity One software tool (Bio-Rad, Hercules, CA, USA). A rabbit anti-Actin antibody (Sigma Chemical Co., St. Louis, MO, USA) was used as control.
2.13 Statistical analysis
A non-parametric Mann Whitney test was used to compare miRNA expression levels in unpaired samples. A χ2 test with a Fisher’s exact p value was used for frequency distribution of the variables. Cox regression univariate analysis was used to assess the influence of immunostaining score on the development of metastases. A Spearman’s rank coefficient (ρ) was used for non-parametric correlations. In vitro experiments were repeated three times and statistical differences were assessed by Student’ t-test. All statistical analyses were performed using SPSS software, with significance set at p < 0.05.
3 Results
3.1 Identification of MET as a putative STS-related miR-125 target
Based on our previous identification of 20 deregulated miRNAs common to LMS and UPS [9] (Supplementary Table 1), we set out to perform a miRNA target gene analysis using various bioinformatic tools (i.e., Targetscan Vert release 5.2, http://www.targetscan.org/, Pictar-Vert http://pictar.mdc-berlin.de/ and PITA, http://genie.weizmann.ac.il/pubs/mir07/index.html). Subsequently, we classified the putative target genes by DAVID (http://www.david.niaid.nih.gov) and, based on this, 12 miRNAs were selected (miR-137, miR-21.5p, miR-100, miR-152, miR-1305, miR-223.3p, miR-138.5p, miR-892b, miR409-5p, miR-376a-3p, miR-487a and miR-212.3p) of which the target genes were predominantly involved in oncogenesis and the regulation of argonaute (AGO) family proteins (Supplementary Table 2). The other miRNAs were excluded because they either presented with too few predicted target genes common to the three databases or because the target genes did not strictly relate to oncogenesis and/or miRNA biogenesis.
Through custom MicroRNA Array-based expression analysis (see Materials and methods) of the primary STS tumor samples and its corresponding normal tissue samples, we could confirm a lower expression in the STS tumor samples of miR-100.5p, miR-152, miR-376a.3p, miR-212.3p, miR-892b and miR-223.3p (Fig. 1), as also a minimal expression of miR-1305, with a statistically significant difference for miR-152 (Mann Whitney test, p = 0.028). In Table 2 all microRNA Array data are listed. Only miR-21.5 showed higher levels in the STS tumor samples than in the normal tissue samples, while undetectable levels were seen for miR-137, miR-409-5p, miR-487a and miR-138.5p.
Non-Gaussian distribution showing that miR-100 (a), miR-152 (b), miR-376a (c), miR-212.3p (d), miR-892b (e) and miR-223 (f) mRNA values were lower in primary STS tumors than in its corresponding normal tissues. A Mann Whitney test revealed a statistically significant difference between the two groups for miR-152
Based on our target gene analyses (see above and Supplementary Table 2), we selected KIT and MET as putative targets of miR-152 and, subsequently, verified their expression by qRT-PCR. By doing so, higher mRNA levels were found in the STS tumor samples than in the normal tissue samples (0.43 and 0.16 for KIT, p = 0.09; 6082.82 and 374.80 for MET; p = 0.05, respectively) (Fig. 2) with a significantly negative correlation between miR-152 and MET expression (ρ = 0.5, p = 0.003). No significant differences between the LMS and UPS histotypes were found, but when patients were divided into prognostic sub-groups, the MET mRNA levels were found to be significantly higher in metastatic than in non-metastatic patients (Mann Whitney test, p = 0.05) (Table 3).
3.2 MET protein expression in STS tumor samples corresponds to clinical parameters
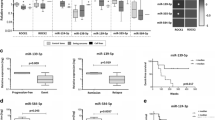
Using immunohistochemistry (IHC), we found that the MET protein was predominantly expressed in the cytoplasm of the STS tumor cells (Fig. 3), with over-expression corresponding to a uniform (> 50% positive cells) and moderate to strong immunoreactivity (score = 4–5) in 79% of the metastatic and 25% of the non-metastatic patients (χ2 = 11.5; p = 0.001). In the corresponding normal tissues MET was found to be focally and weakly expressed (score = 2). Based on the cytoplasmic intensity staining scores (1–5), univariate Cox’s analysis revealed that the risk of metastasis increased by 39.5% for each point increase in score (95%CI = 0.985–1.977, p = 0.06). When the clinical course of the patients exhibiting MET over-expression (score = 4–5) was matched with those with a low score (< 4), the 4-year disease-free survival rates were found to be 21% and 70%, respectively (Fisher’s exact test, p = 0.01). The KIT protein was found to be focally expressed in the majority of the STS tumor samples with a weak to moderate staining intensity (score = 2–3). No or minimal expression was seen in the corresponding normal tissues.
3.3 miR-152 is down-regulated in STS-derived cell lines
Next, we set out to analyze the miR-152 levels in the STS-derived cell lines HT1080, SW872 and SKLMS1. Although we found that all cell lines exhibited down-regulation of miR-152 expression compared to normal hSMC cells that were used as calibrator (Fig. 4), the SKLMS1 cells showed the lowest expression and were, therefore, considered as the best model for in vitro validation, representing the LMS histotype.
In order to subsequently investigate the putative role of miR-152 in LMS cell behavior, we transfected its pre-miRNA precursor into SKLMS1 cells. The transfection efficiency was validated by flow cytometry (Supplementary Fig. Suppl. 1a), and qRT-PCR analysis was used to confirm an increase in the miR-152 ΔCT value in transfected SKLMS1 cells compared to scramble (control) and non-transfected SKLMS1 cells. The highest efficacy was observed after 48 h (78.47%, at 24 h; 87.34% at 48 h and 76.54% at 72 h) concordant with the highest miR-152 expression (2^ -ΔCT of 1186 in transfected cells, 90 in scramble transfected cells and 91 in non-transfected cells) (Supplementary Fig. Suppl. 1b).
3.4 miR-152 decreases MET and KIT expression in SKLMS1 cells
After miR-152 transfection into SKLMS1 cells, the MET and KIT mRNA expression levels were found to be differentially down-regulated. Specifically, we found that ectopic miR-152 expression induced decreases in the MET mRNA levels of 16.7% at 24 h after transfection and 11.1% at 48 h after transfection, respectively, shifting back to control values at 72 h after transfection (Fig. 5a). The KIT mRNA levels were found to decrease to 47.5% at 72 h after transfection. Accordingly, Western blot analyses revealed a reduction in MET and KIT protein expression levels 24 h and 48 h after miR-152 transfection. This reduction was more pronounced for the MET protein (Fig. 5b). Among the cellular signaling cascades that can be affected by MET [17], we focused on the PI3K/AKT cascade. Through Western blot analyses, we observed reductions in the PI3K and p-AKT (phosphorylated at Ser 473) expression levels, with a maximum at 48 h after miR-152 transfection, and a slight increase in the total amount of AKT protein (Fig. 5c).
3.5 miR-152 decreases proliferation and increases apoptosis in SKLMS1 cells
By using a Neubauer cell counting chamber assay, we found that after ectopic miR-152 expression SKLMS1 cells exhibited a significant decrease in proliferation compared to non-transfected cells up to 48 h (27% at 24 h; p = 0.05 and 41% at 48 h; p = 0.007) and to a lesser extent at 72 h (18.7%) (Fig. 6a). Concomitantly, a gradual increase in apoptosis was seen compared to non-transfected cells up to 48 h (11.6% at 24 h; p = 0.08 and 39.6% at 48 h; p = 0.03), while at 72 h the apoptotic cell percentage shifted back to control values (Fig. 6b).
Ectopic miR-152 expression decreases cell proliferation (a) and increases apoptosis (b) up to 48 h after transfection, compared to a scramble control. (c) Cell cycle distributions revealing S phase increases at 24 h and 48 h after transfection. At 72 h after transfection the cell cycle distribution shifted back to control values. *p < 0.05
3.6 miR-152 blocks the cell cycle in the S phase in SKLMS1 cells
The effects of ectopic miR-152 expression on the cell cycle distribution of SKLMS1 cells were assessed by propidium iodide (PI) staining followed by flow cytometry. By doing so, we found that miR-152 transfected SKLMS1 cells exhibited increases in the percentages of S phase cells (Fig. 6c) at 24 h and 48 h after transfection compared to scramble miRNA transfected control SKLMS1 cells (from 15.46% to 23% and from 9% to 25.7%, respectively), while at 72 h after transfection the SKLMS1 cell cycle distribution shifted back to control values.
4 Discussion
miRNA expression deregulation has been reported in a variety of human cancers [18, 19], and some studies have indicated that miRNA deregulation may be employed for the diagnosis and prognosis of human sarcomas [16, 20–22]. On the basis of our previous work [9], we selected 12 cancer-related miRNAs deregulated in non-translocation sarcomas, i.e., LMS and UPS, to find common markers involved in STS development. Although TaqMan microarray-based analyses confirmed a down-regulation of these miRNAs in STS tumors compared to its corresponding normal tissues, this difference remained statistically significant only for miR-152 after qRT-PCR validation. Several factors may contribute to variations in results obtained by microarray and qRT-PCR-based expression analyses. The increased distance between the location of the PCR primers and the microarray probes on a given transcript may decrease the correlation between the two methods, as well as other parameters such as tissue type, sample preparation and platform-related deviations in data quality acquisition [23].
It has been reported that mir-152 may be involved in proliferation, invasion and angiogenesis, and its down-regulation has been observed in ovarian [24], prostate [25], bladder [26], endometrial [27, 28], pancreatic [29] and gastrointestinal [30, 31] cancers. In contrast, in neuroblastoma cells miR-152 has been found to be over-expressed compared to normal mature neurons [32]. As yet, few data are available on miR-152 in human sarcomas. A recent study indicated that plasma and tumor tissues from osteosarcoma patients may exhibit lower miR-152 expression levels than healthy controls, suggesting that miR-152 may serve as a biomarker for early tumor detection [33]. The currently known miR-152 targets include TNFRF6B, involved in hepatocarcinogenesis [34], DNMT1, involved in endometrial cancer [28], ADAM17, a metalloprotease, FGF2, a fibroblast growth factor, E2F3, a transcription factor, and MET and KIT [28, 35, 36]. MET and KIT are primarily involved in sarcoma development and progression [37, 38]. MET is a receptor tyrosine kinase that binds hepatocyte growth factor (HGF) and guides malignant progression by activating signaling pathways that promote cell motility, survival and proliferation [39]. KIT is also a receptor tyrosine kinase operating in cell signal transduction [40]. Several pathways act downstream of KIT to affect cell survival and proliferation and mutant forms of KIT are known to act as drivers of several tumors, including gastrointestinal stromal tumors (GIST) [41].
Here, we observed MET expression up-regulation in STS tumors, which was significantly inversely correlated with miR-152 expression. We also found that KIT expression up-regulation was less profound. KIT plays a predominant role in GIST (see above) and in subsets of acute myeloid leukemia and melanoma, and its mutation status may be predictive of the response to the tyrosine kinase inhibitor (TKI) imatinib [42]. Interestingly, we found that MET expression levels were significantly higher in patients who developed metastases during follow-up than in those that did not. These data, in conjunction with a strong and uniform MET expression in the majority of metastatic patients, underscore the clinical relevance of MET in sarcoma development and prognosis [38, 43]. The role of miR-152 in STS was further supported by our in vitro assays, indicating that ectopic miR-152 expression may lead to a reduction in KIT and MET expression, a transient decrease in cell growth, a transient increase in apoptosis and a transient cell cycle S-phase delay up to 48 h after transfection.
Our results are in agreement with previous reports showing that exogenous miR-152 expression may suppress the proliferation of CD133+ Hep3B cells through KIT down-regulation [36] and that miR-152 expression restoration in endometrial cancer cells may inhibit cell growth and induce cell cycle delay [24]. MET signaling has been reported to promote tumorigenicity in a variety of ways, including anomalous regulation of the PI3K/AKT pathway [44]. Alterations in this pathway are often encountered in cancer cells, favoring tumor cell survival and spread [45]. We found that exogenous expression of miR-152 in SKLMS1 cells led to a decrease in PI3K expression resulting from MET down-regulation, followed by down-regulation of pAKT (Ser473) expression. Therefore, targeting of MET by miR-152 may indirectly affect PI3K/AKT signaling, thereby decreasing STS proliferation and increasing STS apoptosis. Our results also suggest a prognostic role of MET in identifying STS patients at risk of developing metastases. In addition, our data suggests that miR-152 over-expression may sensitize STS cells to cytotoxic agents, thereby opening avenues for the design of novel therapeutic strategies. Additional in vivo studies will, however, be required for the concomitant administration of anti-neoplastic drugs with synthetic miRNA oligonucleotides.
References
P. Rutkowski, I. Lugowska, Follow-up in soft tissue sarcomas. Memory 7, 92–96 (2014)
M. Miettinen, J. F. Fetsch, Evaluation of biological potential of smooth muscle tumours. Histopathology 48, 97–105 (2006)
P. Picci, M. Manfrini, N. Fabbri, M. Gambarotti, D. Vanel, Atlas of Muscoloskeletal tumors and Tumorlike lesion (Springer, New York, 2014), pp. 311–313
C. D. M. Fletcher, J. A. Bridge, P. Hogendoorn, F. Mertens, WHO classification of tumours of soft tissue and bone, 4thEdition edn. (IARC Press, Lyon, 2013)
A. F. Nascimenàkto, C. P. Raut, Diagnosis and management of pleomorphic sarcomas (so-called "MFH") in adults. J. Surg. Oncol. 97, 330–339 (2008)
P. Dei Tos, Classification of pleomorphic sarcomas: where are we now? Histopathology 48, 51–62 (2006)
L. J. Helman, P. Meltzer, Mechanisms of sarcoma development. Nat. Rev. Cancer 3, 685–694 (2003)
T. Niini, L. Lahti, F. Michelacci, S. Ninomiya, C.M. Hattinger, M. Guled, T. Bohling, P. Picci, M. Serra, S. Knuutila, Array comparative genomic hybridization reveals frequent alterations of G1/S checkpoint genes in undifferentiated pleomorphic sarcoma of bone. Genes Chromosom. Cancer 50, 291–306 (2011)
M. Guled, L. Pazzaglia, I. Borze, N. Mosakhani, C. Novello, M. S. Benassi, S. Knuutila, Differentiating soft tissue leiomyosarcoma and undifferentiated pleomorphic sarcoma: a miRNA analysis. Genes Chromosom. Cancer 53, 693–702 (2014)
F. Chibon, P. Lagarde, S. Salas, G. Perot, V. Brouste, F. Tirode, C. Lucchesi, A. de Reynies, A. Kauffmann, B. Bui, P. Terrior, S. Bonvalot, A. LeCesne, D. Vince-Panchere, J. Y. Blay, F. Collin, L. Guillou, A. Leroux, J. M. Coindre, A. Aurias, Validated prediction of clinical outcome in sarcomas and multiple types of cancer on the basis of a gene expression signature related to genome complexity. Nat. Med. 16, 781–787 (2010)
S. L. Romero-Cordoba, I. Salido-Guadarrama, M. Rodriguez-Dorantes, A. Hidalgo-Miranda, miRNA biogenesis: biological impact in the development of cancer. Cancer Biol. Ther. 15, 1444–1455 (2014)
M. Vitiello, A. Tuccoli, L. Poliseno, Long non-coding RNAs in cancer: implications for personalized therapy. Cell. Oncol. 38, 17–28 (2015)
A. Ferraro, Altered primary chromatin structures and their implications in cancer development. Cell. Oncol. 39, 195–210 (2016)
V. Taucher, H. Mangge, J. Haybaeck, Non coding RNAs in pancreatic cancer: challenges and opportunities for clinical application. Cell. Oncol. 39, 295–318 (2016)
G. J. Nuovo, T. D. Schmittgen, Benign metastasizing leiomyoma of the lung: clinicopathologic, immunohistochemical, and micro-RNA analyses. Diagn. Mol. Pathol. 17, 145–150 (2008)
T. Greither, P. Würl, L. Grochola, G. Bond, M. Banche, M. Kappler, C. Lautenschläger, H. J. Holzhausen, S. Wach, A. W. Eckert, H. Taubert, Expression of microRNA 210 associates with poor survival and age of tumor onset of soft-tissue sarcoma patients. Int. J. Cancer 130, 1230–1235 (2012)
E. Gherardi, W. Birchmeier, C. Birchmeier, G. Vande Woude, Targeting MET in cancer: rationale and progress. Nat. Rev. Cancer 12, 89–103 (2012)
A. Esquela-Kerscher, F. J. Slack, Oncomirs - microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006)
M. V. Iorio, C. M. Croce, Causes and consequences of microRNA dysregulation. Cancer J. 8, 215–222 (2012)
S. Subramanian, W. O. Lui, C. H. Lee, I. Espinosa, T. O. Nielsen, M. C. Heinrich, C. L. Corless, A. Z. Fire, M. van de Rijn, MicroRNA expression signature of human sarcomas. Oncogene 27, 2015–2026 (2008)
L. S. Danielson, S. Menendez, C. S. Attolini, M. V. Guijarro, M. Bisogna, J. Wei, N. D. Socci, D. A. Levine, F. Michor, E. Hernando, A differentiation-based microRNA signature identifies leiomyosarcoma as a mesenchymal stem cell-related malignancy. Am. J. Pathol. 177, 908–917 (2010)
C. Zhou, W. Tan, H. Lv, F. Gao, J. Sun, Hypoxia-inducible microRNA-488 regulates apoptosis by targeting Bim in osteosarcoma. Cell. Oncol. 39, 463–471 (2016)
S. Morey, J. C. Ryan, F. M. Van Dolah, Microarray validation: factors influencing correlation between oligonucleotide microarrays and real-time PCR. Biol. Proc. Online 8, 175–193 (2006)
X. Zhou, F. Zhao, Z. N. Wang, Y. X. Song, H. Chang, Y. Chiang, H. M. Xu, Altered expression of miR-152 and miR-148a in ovarian cancer is related to cell proliferation. Oncol. Rep. 27, 447–454 (2012)
Z. Lichner, A. Fendler, C. Saleh, A. N. Nasser, D. Boles, S. Al-Haddad, P. Kupchak, M. Dharsee, P. S. Nuin, K. R. Evans, K. Jung, C. Stephan, N. E. Fleshner, G. M. Yousef, MicroRNA signature helps distinguish early from late biochemical failure in prostate cancer. Clin. Chem. 59, 1595–1603 (2013)
C. U. Köhler, O. Bryk, S. Meier, K. Lan, P. Rozynek, T. Brüning, H. U. Käfferlein, Analyses in human urothelial cells identify methylation of miR-152, miR-200b and miR-10a genes as candidate bladder cancer biomarkers. Biochem. Biophys. Res. Commun. 438, 48–53 (2013)
E. Hiroki, J. Akahira, F. Suzuki, S. Nagase, K. Ito, T. Suzuki, H. Sasano, N. Yaegashi, Changes in microRNA expression levels correlate with clinicopathological features and prognoses in endometrial serous adenocarcinomas. Cancer Sci. 101, 241–249 (2010)
T. Tsuruta, K. Kozaki, A. Uesugi, M. Furuta, A. Hirasawa, I. Imoto, N. Susumu, D. Aoki, J. Inazawa, miR-152 is a tumor suppressor microRNA that is silenced by DNA hypermethylation in endometrial cancer. Cancer Res. 71, 6450–6462 (2011)
M. Azizi, L. Teimoori-Toolabi, M. K. Arzanani, K. Azadmanesh, P. Fard-Esfahani, S. Zeinali, MicroRNA-148b and microRNA-152 reactivate tumor suppressor genes through suppression of DNA methyltransferase-1 gene in pancreatic cancer cell lines. Cancer Biol. Ther. 15, 419–427 (2014)
Y. Chen, Y. Song, Z. Wang, Z. Yue, H. Xu, C. Xing, Z. Liu, Altered expression of MiR-148a and MiR-152 in gastrointestinal cancers and its clinical significance. J. Gastrointest. Surg. 14, 1170–1179 (2010)
Y. X. Song, Z. Y. Yue, Z. N. Wang, Y. Y. Xu, Y. Luo, H. M. Xu, X. Zhang, L. Jiang, C. Z. Xing, Y. Zhang, MicroRNA-148b is frequently down-regulated in gastric cancer and acts as a tumor suppressor by inhibiting cell proliferation. Mol. Cancer 10, 1 (2011). doi:10.1186/1476-4598-10-1
D. Z. Liu, B. P. Ander, Y. Tian, B. Stamova, G. C. Jickling, R. R. Davis, F. R. Sharp, Integrated analysis of mRNA and microRNA expression in mature neurons, neural progenitor cells and neuroblastoma cells. Gene 495, 120–127 (2012)
N. G. Wang, D. C. Wang, B. Y. Tan, F. Wang, Z. N. Yuan, Down-regulation of microRNA152 is associated with the diagnosis and prognosis of patients with osteosarcoma. Int. J. Clin. Exp. Pathol. 8, 9314–9319 (2015)
Y. W. Dang, J. Zeng, R. Q. He, M. H. Rong, D. Z. Luo, G. Chen, Effects of miR-152 on cell growth inhibition, motility suppression and apoptosis induction in hepatocellular carcinoma cells. Asian Pac. J. Cancer Prev. 15, 4969–4976 (2014)
L. Li, Y. Y. Chen, S. Q. Li, C. Huang, Y. Z. Qin, Expression of miR-148/152 family as potential biomarkers in non-small-cell lung cancer. Med. Sci. Monit. 23, 1155–1161 (2015)
H. Huang, M. Hu, P. Li, C. Lu, M. Li, Mir-152 inhibits cell proliferation and colony formation of CD133+ liver cancer stem cells by targeting KIT. Tumor Biol. 36, 921–928 (2014)
T. Fukuda, E. Ichimura, T. Shinozaki, T. Sano, K. Kashiwabara, T. Oyama, T. Nakajima, T. Nakamura, Coexpression of HGF and c-met/HGF receptor in human bone and soft tissue tumors. Pathol. Int. 48, 757–762 (1998)
G. Lahat, P. Zhang, Q. S. Zhu, K. Torres, M. Ghadimi, K. D. Smith, W. L. Wang, A. J. Lazar, D. Lev, The expression of c-met pathway components in unclassified pleomorphic sarcoma/malignant fibrous histiocytoma (UPS/MFH): a tissue microarray study. Histopathology 59, 556–561 (2011)
L. Trusolino, A. Bertotti, P. M. Comoglio, MET signalling: principles and functions in development, organ regeneration and cancer. Nat. Rev. Mol. Cell Biol. 11, 834–848 (2010)
M. Miettinen, J. Lasota, KIT (CD117): a review on expression in normal and neoplastic tissues, and mutations and their clinicopathologic correlation. Appl. Immunohistochem. Mol. Morphol. 13, 205–220 (2005)
L. K. Ashman, R. Griffith, Therapeutic targeting of c-KIT in cancer. Expert Opin. Investig. Drugs 22, 103–115 (2013)
J. Noujaim, D. Gonzalez, K. Thway, R. L. Jones, I. Judson, P.(L576P) -KIT mutation in GIST: favourable prognosis and sensitive to imatinib? Cancer Biol. Ther. 17, 543–555 (2016)
E. D. Fleuren, M. H. Roeffen, W. P. Leenders, U. E. Flucke, M. Vlenterie, H. W. Schreuder, O. C. Boerman, W. T. van der Graaf, Y. M. Versleijen-Jonkers, Expression and clinical relevance of MET and ALK in Ewing sarcomas. Int. J. Cancer 133, 427–436 (2013)
M. Trovato, M. L. Torre, M. Ragonese, A. Simone, R. Scarfì, V. Barresi, G. Giuffrè, S. Benvenga, F. F. Angileri, G. Tuccari, F. Trimarchi, R. M. Ruggeri, S. Cannavò, HGF/c-met system targeting PI3K/AKT, STAT3/phosphorylated-STAT3 pathways in pituitary adenomas: an immunohistochemical characterization in view of targeted therapies. Endocrine 44, 735–743 (2013)
Y. Samuels, Z. Wang, A. Bardelli, N. Silliman, J. Ptak, S. Szabo, H. Yan, A. Gazdar, S. M. Powell, G. J. Riggins, J. K. Willson, S. Markowitz, K. W. Kinzler, B. Vogelstein, V. E. Velculescu, High frequency of mutations of the PIK3CA gene in human cancers. Science 304, 554 (2004)
Acknowledgments
The authors wish to thank Dr. Alba Balladelli for editing the paper and Ms. Cristina Ghinelli for her graphic work. This work was supported by 5‰ donation (Italy). Chiara Novello was supported by a FIRC Triennal fellowship “Mario e Valeria Rindi” (research project n°13748).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have read and approved the contents of this manuscript and declare that the work is original and has not been submitted or published elsewhere. The authors also state that there is no potential conflict of interest.
Ethical standards
The research protocol was approved by ethic committees of the Rizzoli Institute where the tumor samples were collected. All patients provided informed consent.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Pazzaglia, L., Novello, C., Conti, A. et al. miR-152 down-regulation is associated with MET up-regulation in leiomyosarcoma and undifferentiated pleomorphic sarcoma. Cell Oncol. 40, 77–88 (2017). https://doi.org/10.1007/s13402-016-0306-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-016-0306-4