Abstract
Salt threatens rice cultivation in many countries. Hence, breeding new varieties with high salt tolerance is important.
A panel of 2,391 rice accessions from the 3 K Rice Genome Project was selected to evaluate salt tolerance via the standard evaluation score (SES) in hydroponics under 60 mM NaCl at the seedling stage. Three sub-population panels including 1,332, 628, and 386 accessions from the original 2,391 ones were constructed based on low relatedness revealed by a phylogenetic tree generated by Archaeopteryx Tree. A genome-wide association study (GWAS) was conducted on the entire and sub-population panels using SES data and a selection of 5, 10, 20, and 40% of SNPs selected from the original 1,011,601 SNPs by filtering minor allele frequency > 5% and missing rate < 5%. To perform GWAS, three methods implemented in three different software packages were utilized.
Using the integration of GWAS programs, a total of four QTLs associated with SES scores were identified in different panels. Some QTLs co-located with previously detected QTL-related traits. qSES1.1 was detected in three panels, qSES1.3 and qSES2.1 in two panels, and qSES3.1 in one panel through GWAS by all three methods used and selected SNPs. These four QTLs were selected to detect candidate genes. Combining gene-based association study plus haplotype analysis in the entire population and the three sub-populations let us shortlist three candidate genes, viz. LOC_Os01g23640 and LOC_Os01g23680 for qSES1.1, and LOC_Os01g71240 for qSES1.3 region affecting salt tolerance. The identified QTLs and candidate genes provided useful materials and genetic information for future functional characterization and genetic improvement of salt tolerance in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is a staple food for more than half of the population in the world. In 2019, it was cultivated in more than 115 countries, of which Asia’s share is approximately 86% and 90% of the world's total area harvested and its production, respectively. The majority of the Asian rice cultivated area regions are South (44.3%) and South-East Asia (31.7%) in which soil salinization is a big problem (Sharma et al. 1999; Vinod et al. 2013; FAO 2021). Although rice – at the species level – is considered moderately sensitive to salinity with a salinity threshold estimated at 3 dS m−1, rice varieties are often classified from highly susceptible to highly tolerant according to the modified standard evaluation score (IRRI 2013). Therefore, rice varieties are often classified as salt-sensitive or salt-tolerant (Maas and Hoffman 1977; Munns and Tester 2008; Reddy et al. 2017).
Salt tolerance in rice is a quantitative character and is controlled by multiple genes (Munns 2005; Reddy et al. 2017; Liu et al. 2019). Therefore, it is important to identify quantitative trait loci (QTL) or genes for salt tolerance to get more genetic information in breeding programs to improve salt tolerance. It is no doubt that genetic and molecular studies on the salt tolerance of rice contribute to success in transgenic and breeding crop varieties. For example, the rice varieties to which Saltol – (SKC1, shoot K content 1), a major QTL for salt tolerance – has been transferred, showed improved salt tolerance by accumulating less Na+ and relatively more K+, thus maintaining a lower Na+/K+ ratio in leaves (Rahman et al. 2016; Tiwari et al. 2016). A large number of genes for salt tolerance in rice have been cloned from QTL studies such as SKC1, OsCCC1, OsNHX1 (Ren et al. 2005; Kong et al. 2011; Almeida et al. 2017; Reddy et al. 2017), among others. However, it is unlikely to reveal the whole genetic variation for the traits due to the limited number of parents. It is also difficult to develop rice elite varieties with high salt tolerance because of a lack of understanding of the mechanisms of salt tolerance in rice.
In recent decades, more than one hundred genes and a lot of QTLs conferring salinity tolerance traits were identified. Koyama et al. (2001) detected 11 QTLs related to Na+ uptake, K+ uptake, Na+:K+ ratio, Na+ concentration, and K+ concentration in the shoot tissue. Using an F2:3 population derived from a cross between Tarommahalli and Khazar varieties, Sabouri and Sabouri (2008) detected nine QTLs for root length, root dry weight, and ion exchanges under salt stress on different chromosomes. Thomson et al. (2010) detected 17 QTLs for seedling height, shoot Na+ concentration, shoot K+ concentration, shoot Na+:K+ ratio, root K+ concentration, root Na+:K+ ratio, SES score, and leaf chlorophyll content by using 140 IR29/Pokkali recombinant inbred lines (RILs). However, a majority of these QTLs were mapped using bi-parental linkage mapping populations (Koyama et al. 2001; Sabouri and Sabouri 2008; Thomson et al. 2010; Yamamoto et al. 2012; Reddy et al. 2017). This method shows high statistical power due to using many individuals sharing identical genotypes at a given locus; however, it has a low resolution because of the limited number of recombination events in the development of the population, and it only includes a few parts of the allelic variation available at the species level. Indeed, the resolution depends on the sample size and type of mapping population. The more advanced the population and the larger its size, the more recombination events could be captured.
Recently, a lot of QTL, QTN (quantitative trait nucleotide) or candidate genes have been identified by GWAS with high resolution due to the long recombination histories of natural populations. Many QTLs/candidate genes related to salt tolerance in rice have been detected by this method (Shi et al. 2017; Batayeva et al. 2018; Naveed et al. 2018; Liu et al. 2019). Shi et al. 2017 identified 11 loci containing 22 significant salt tolerance associated SNPs at the seed germination stage by using 478 rice accessions, and one candidate gene associated with vigor index under salt stress in rice. With 203 temperate japonica rice accessions, Batayeva et al. (2018) detected a total of 26 QTLs for nine measured traits, and six candidate genes were identified for five promising QTLs related to salt tolerance at the seedling stage in rice. Naveed et al. (2018) detected 20 QTN associated with 11 traits related to salinity tolerance in rice at germination and seedling stages, then 22 candidate genes for nine important QTNs were found by using 208 rice accessions. Liu et al. (2019) detected 15 promising candidates significantly associated with the target traits under saline conditions, including five known genes and 10 novel genes. These results provided valuable resources and genetic information for molecular dissection of salt tolerance to improve rice salt tolerance in future breeding programs.
Many methods implemented by software programs – such as TASSEL, PLINK, GAPIT, FaST-LMM, BLINK, and so on (Yu et al. 2006; Bradbury et al. 2007; Lippert et al. 2011; Huang et al. 2019) – have been released for GWAS analysis using different statistical models and different algorithms to reduce the confounders of GWAS, such as population structure and cryptic relatedness, as well as the run time and memory footprint in the analysis. TASSEL and GAPIT are often used in plant genome analysis, however, their running takes much time and memory footprint, thus their use has been often restricted to the analysis of relatively small population sizes and genotypic datasets (Yu et al. 2006; Bradbury et al. 2007, Guo et al. 2021). FaST-LMM could resolve that problem and is able to analyze a large sample size (Lippert et al. 2011). It has been reported that different GWAS programs could produce dissimilar detected QTLs results. Also, different sample subsets of the dataset such as the number of accessions and proportion of original genome data used in GWAS could lead to divergent results regarding detected QTLs (Yan et al. 2019). Regarding salt tolerance trait in rice, most of QTL/QTN detected through GWAS previously resulted from a relatively limited number of genotypes, such as less than 500. To date, no QTL for salt tolerance of rice detected from a large population such as containing 1,000 or 2,000 accessions has been reported. On the other hand, most of the detected QTLs for salt tolerance identified in previous studies were under high salinity while rice could be cultivated under moderate saline condition only, since salinities higher than 6.6 dS m−1 causes 50% yield losses or more (Zeng and Shannon 2000; Phan et al. 2023). Consequently, when the salinity level in the paddy field is higher than 6.6 dS m−1 (equivalent to 60 mM NaCl), rice is often not cultivated due to its very low grain yield. Therefore, the objective of the present research was to identify new QTLs and candidate genes related to salt tolerance of rice through GWAS and to compare the effectiveness of different GWAS analysis methods detected QTLs/candidate genes.
Material and methods
Plant growth
We selected 2,391 rice accessions from the 3,000 Rice Genomes Project (3K R.G.P. 2014) collected from 75 countries and belonging to nine variety types, viz. aromatic, aus, temperate japonica, tropical japonica, subtropical japonica, indica-1A, indica-1B, indica-2, and indica-3; and the admixture including japonica-admixed, indica-admixed, and admixed (Supplementary Table S1). The hydroponic experiment was carried out in a phytotron at Université catholique de Louvain (UCLouvain), Belgium, from March to April 2019. The seeds were sown directly in holes on extruded polystyrene plates floating on 40L tanks, 74.5 cm length × 54.5 cm width × 10.0 cm height. Four tanks with 630 holes each were used. Eight seeds of the same genotype were sown in each hole. The experimental design was laid out by an augmented randomized complete block design (Federer and Raghavarao 1975), and the number of blocks was 4 as the number of the tanks. Among the 2,391 accessions, 2,379 appeared only once (i.e. 8 seeds in one hole), thus 594–595 accessions per tank. The 12 remaining accessions were replicated once in each of the four tanks. The 23–24 remaining holes in each tank were filled by three unsequenced genotypes to calculate the effect of the tanks. The 40L tanks were filled with the Yoshida solution (Yoshida et al. 1976) which was renewed 7, 10, 14, and 17 days after the treatment. NaCl was applied at a concentration of 60 mM starting from the sowing time. The pH was adjusted daily from 5.0 to 5.5 with KOH 2 M or HCl 1 M. One week after sowing, five uniform plants per hole were maintained for evaluation. The climatic conditions in the phytotron were maintained at 30 °C/25 °C day/night, 85%-95% relative humidity, 12 h photoperiod, and 210 µmol m−2 s−1 photon flux density at the top of the tanks. The tanks were interchanged twice a week.
Phenotyping
SES (standard evaluation score) was determined after three weeks of treatment as described by IRRI (2013), with score 1 being the most salt tolerant, and score 9 being the most salt susceptible as below:
-
Score 1: Growth and tillering nearly normal
-
Score 3: Growth nearly normal but there is some reduction in tillering, some leaves whitish and rolled
-
Score 5: Growth and tillering reduced; most leaves whitish and rolled; only a few elongating leaves
-
Score 7: Growth completely ceases; most leaves dry; some plants dying
-
Score 9: Almost all plants dead or dying
-
The collected data of SES of each treatment were adjusted by augmentedRCBD package in R software (Aravind et al. 2020) based on the check-varieties.
SNP selection
A total of 1,011,601 GWAS SNPs (Single Nucleotide Polymorphisms) were sequenced by 3 K RGP and downloaded from the Rice SNP-Seek Database [https://snp-seek.irri.org, (Mansueto et al. 2016)]. We selected the SNPs with a minor allele frequency of > 5% and missing rates of < 5%, leading to 588,792 SNPs. Subsequently, we randomly selected subsets of 5, 10, 20, and 40% of these SNPs independently, leading to 29,316, 58,968, 118,065, and 235,475 SNPs, respectively.
Principal components analysis of genotypic data
Principal components analysis (PCA) of the 588,792 SNPs with phenotypic data was done with the default setting by PLINK 1.9 (Purcell et al. 2007). Then, the eigenvalue of the top three components was selected as covariate data.
Linkage disequilibrium decay
We analyzed linkage disequilibrium (LD) decay in the whole panel of 2,391 accessions. Random selection of 20% of the 588,792 selected SNPs was used to calculate LD decay. We calculated r2 as an estimation of LD using PLINK software version 1.9 (Purcell et al. 2007). The syntax was “–r2 –ld-window-kb 1000 –ld-window 9999 –ld-window-r2 0”. Marker pairs were grouped into bins of 1 kb and the average r2 value of each bin was calculated. The LD decay rate was measured as the distance at which the average r2 dropped to half of its maximum value (Huang et al. 2010; Shi et al. 2017).
Genome-wide association study
The three methods implemented following the software programs of GWAS were applied to compare the results.
The first method used was Trait Analysis by aSSociation, Evolution and Linkage (TASSEL) with a Mixed Linear Model (MLM or Linear Mixed Model – LMM – as in literature), using TASSEL software 5 (Zhang et al. 2010).
The second method was BLINK (Bayesian-information and Linkage-disequilibrium Iteratively Nested Keyway) by BLINK C version. It uses multi-locus model for testing markers across the genome and conducts two fixed-effect models iteratively. One model tests marker one by one with multiple associated markers fitted as covariates to account for the population while another model selects the covariate markers to directly control spurious association instead of kinship, unmasking the confounding effect between the testing marker and kinship (Huang et al. 2019).
The third method was FaST-LMM with a Factored Spectrally Transformed Linear Mixed Model, by FaST-LMM software (Lippert et al. 2011). We first randomly selected 2.5% of the 588,792 SNPs obtained for measuring genetic similarities between the accessions.
Each method was carried out with 5, 10, 20, and 40% of the total number of SNPs (588,792) across four accession panels. The entire panel included 2,391 accessions. Three subpanels including 1,332, 628, and 386 accessions were constructed based on low relatedness criteria revealed by the phylogenetic tree using Archaeopteryx Tree image which was generated in TASSEL 5.2.57 by using the neighbor-joining cladogram function of 2,391 accessions with 5,902 SNPs (Supplement Fig.S1 and Table S1). In the phylogenetic tree of 2,391 accessions, a pair of tips represent two accessions closely related to each other. Manually randomly removing one tip and keeping another tip of each pair in the phylogenetic tree of 2,391 accessions resulted in a subpanel including 1,332 accessions. Continuing randomly removing one of two related accessions in the tree allowed us to select successively new subpanels of 628 and 386 accessions. The panels of 2,391, 1,332, 628, and 386 accessions were collected from 75, 68, 62, and 47 countries, respectively (Supplementary Table S1). The threshold was set by suggestive p-value at 1/number of selected SNPs, and led to 3.4 × 10–5, 1.7 × 10–5, 8.5 × 10–6, and 4.2 × 10–6 in the selected SNPs at 5, 10, 20 and 40%, respectively as described in a previous study (Mao et al. 2022). The false discovery rate was estimated by Benjamini-Hochberg (BH) to adjust probabilities of detected associations by “p.adjust” function in R (Benjamini and Hochberg 1995). A significant threshold of 10% FDR was used to identify QTLs. The LD decay rate was measured as a distance at which the average r2 dropped to half of its maximum value (Huang et al. 2010; Shi et al. 2017). All the neighbor SNPs that (1) showed p ≤ 3.4 × 10–5, 1.7 × 10–5, 8.5 × 10–6, and 4.2 × 10–6 in the GWAS analysis using 5%, 10%, 20%, and 40% of the total of SNPs, respectively and (2) were within the linkage disequilibrium (LD) decay rate were considered as a single QTL. The SNP with the minimum p in one promising QTL was considered as a lead SNP of the QTL. Manhattan and Q-Q plots were created by the R package “GAPIT” using GWAS results (Zhang et al. 2010).
Haplotype analysis and candidate gene identification
Detecting candidate genes in 3 K RGP were described by Wang et al. (2017). Among all the detected QTLs, we focused on the most important QTLs to identify candidate genes by gene-based association analysis. A QTL was classified as “important” when it had been identified by all three methods of GWAS. We identified candidate genes for each important QTL through the following steps. Firstly, we identified all genes located inside the important QTL region (± 140 kb from lead SNP) from The Rice Annotation Project Database RAP-DB [https://rapdb.dna.affrc.go.jp/viewer/gbrowse/irgsp1/] and from the Rice Genome Browser [http://rice.uga.edu/cgi-bin/gbrowse/rice/, (Kawahara et al. 2013)]. Secondly, all available SNPs inside these QTLs were searched from the 32 million SNPs (32,064,217 SNPs) data set generated from 3 K RGP in the Rice SNP-Seek Database (https://snp-seek.irri.org), in order to estimate the position of the QTLs more accurately. Thirdly, all SNPs with an allele frequency less than 0.05 or a missing rate over 5% were removed and the remaining high-quality SNPs were used to analyze GWAS by the multi-locus GWAS analysis [mrMLM package in R software (Zhang 2019)]. Finally, we selected all genes harboring SNPs – excluding transposon/retrotransposon or expressed protein – with a p-value ≤ 0.001 (Naveed et al. 2018) for SES and searched for significant differences among major haplotypes (containing more than 10 accessions per haplotype) of the four panels through ANOVA and post hoc Duncan test. The genes with significance level p < 0.05 in all four tested panels were considered to be promising candidate genes of the target traits.
A confirmatory experiment with a panel of 1,332 accessions chosen above was conducted to confirm the results of QTLs detected in the main experiment.
Results
PCA of genotypic data, and LD decay
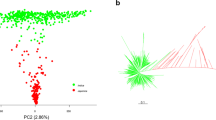
The PCA of genotypic data of the four panels with 2,391, 1,332, 628, and 386 accessions showed similar patterns, and the accessions were classified into four main subgroups: indica, japonica, aromatic, and aus (Fig. 1). The admixed accessions spread out on the whole the plot. It means that the selected subpanels fitted the genotypic diversity of the entire panel.
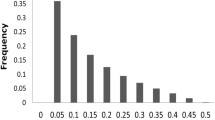
LD decay of the panels was analyzed quickly with 1 kb bin. The results indicated that the LD decay in the panel with 386 accessions was slightly faster than that of the three other panels. The LD decayed to half its maximum value within around 263 kb for the panel with 386 accessions and approximately 280 kb for the three other ones (Fig. 2). Thus, the neighboring SNPs within 280 kb should be considered as a single QTL.
SES of the different panels and subspecies
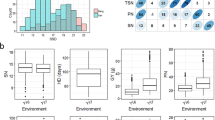
There was a wide range of variation in SES scores among the tested accessions, and the distribution did not differ between panels. The largest proportion of accessions had a SES score of 5, followed by 3, 7, and few accessions had a score of 1 or 9. The mean score was 4.61, 4.59, 4.64, and 4.64 for the panels containing 2,391, 1,332, 628, and 386 accessions, respectively. The ANOVA test showed that the mean of SES was not significantly different between the panels but well between the subspecies. Among them, japonica appeared as the most salt tolerant with the lowest SES, followed by aromatic, admixed, indica, and finally aus. (Fig. 3).
Average of SES score in four subspecies and admixed accessions in the entire panel with 2,391 accessions, and the three panels of 1,332, 628, and 386 accessions of rice grown in Yoshida et al. (1976) solution under 60 mM NaCl. Means ± SE. Bars with the same lowercase letter are not significantly different from each other according to Tukey’s test at the 5% level between the panels and subspecies
Detection of QTLs by different programs of GWAS analysis
We analyzed GWAS by three methods implemented by software programs (TASSEL, BLINK, and FaST-LMM) across four panels (2,391, 1,332, 628, and 386 accessions) and different numbers of SNPs (5, 10, 20, and 40% of the total of 588.792 SNPs). A total of 17 QTLs were detected in one or several of the four panels through GWAS by one or several of the three programs used and the selected SNPs (Fig. 4, Supplementary Table S3).
Among the three GWAS analysis programs tested, FaST-LMM and BLINK showed advantages in terms of runtime duration and size of samples used (number of accessions and number of SNPs). TASSEL could not accomplish GWAS analysis for 2,391 accessions due to the memory footprint of the computer used (processor AMD Ryzen 5 3500U with Radeon Vega Mobile Gfx 2.1 GHz, RAM 8 GB, hard disk 512 GB SSD) but FaST-LMM and BLINK could. These two software programs also provided a greater number of SNPs passing through the threshold than TASSEL. The obtained results showed that the greatest number of QTLs was detected by BLINK (14), followed by FaST-LMM (10), and finally TASSEL (4). Among them, 3 QTLs were detected by both FaST-LMM and BLINK. There were four QTLs identified by all three programs: qSES1.1, qSES1.3, qSES2.1, and qSES3.1 (Fig. 5 and Table 1). These four QTLs were considered the most important QTLs and were retained for further study. All these most important QTLs were detected in both main and confirmatory experiments.
Effect of the number of SNPs and the size of the panel used
The number of SNPs contained in the genotypic dataset and the number of accessions are two main components of sample size that influence the feasibility of GWAS runnings. In the present study, BLINK and FaST-LMM could use up to 40% of the total number of SNPs in all four panels, whereas TASSEL could complete the GWAS analysis with only up to 20, 10, and 5% of the total number of SNPs in the panels of 386, 628, and 1,332 accessions, respectively. The effect of different numbers of SNPs on detected QTLs was not clear. By setting the threshold of significant SNPs at p ≤ 3.4 × 10–5, 1.7 × 10–5, 8.5 × 10–6, and 4.2 × 10–6 in the GWAS analysis using 5%, 10%, 20%, and 40% of the total of SNPs, respectively, the detected QTLs did not depend on the selected SNPs, except for the analysis of 628 accessions in BLINK (Supplementary Table S3 and Fig. S2).
Among the four panels, we identified 3, 9, 6, and 9 QTLs related to SES in the panel containing 386, 628, 1332, and 2391 accessions, respectively. Among them, two QTLs, viz. qSES1.1 and qSES10.1, were detected in three out of the four panels (Table 2). There were some QTLs detected by the three methods in each subpanel: qSES3.1 in the panel of 386 accessions, qSES1.3 in the panel of 628 accessions, and qSES2.1 in the panel of 1332 accessions. One of three, three of nine, and two of nine QTLs were detected by two of the three methods in subpanel of 386, 628, and 2,391 accessions, respectively, and the remaining QTLs were detected by only one method (Supplementary Fig. S3).
QTL verification assay
A confirmatory experiment with 1,332 accessions was conducted to confirm the QTLs detected in the main experiment. The GWAS was analyzed by using BLINK with 5%, 10%, 20%, and 40% of the SNPs and with the three subpanels as in the main experiment. A total of 26 QTLs was detected in this check (Supplementary Table S2b). Among them, eight QTLs were in the same regions as those detected in the main experiment. All of the four most important QTLs were identified in both experiments.
Candidate genes for important QTLs
We selected all available SNPs inside the four most important QTLs, viz. qSES1.1, qSES1.3, qSES2.1, and qSES3.1, then re-ran a GWAS of these QTLs (gene-based analysis) with high-quality SNPs inside the QTLs to confirm the significant SNPs by the multi-locus GWAS analysis [mrMLM package in R software (Zhang 2019)]. Then, with the resolution of 280 kb for all identified important QTLs, we were able to narrow down a relatively small number of candidate genes for each QTL. Thus, we selected 66 genes – excluding transposon/retrotransposon, hypothetical protein, and expressed protein – in these four most important QTLs (Supplementary Table S4b). The GWAS results allowed us to choose 13 genes matching the selection: the genes harboring SNPs with the lowest p-value that passed the threshold of -log10p-value ≥ 3 to test significant phenotypic differences between haplotypes (Fig.S4, Table S5). Further haplotype analysis of the 13 genes detected three promising candidates significantly associated with the target traits (Table 3, 4, Fig. 6 and Supplementary Table S5): LOC_Os01g23640 (accession AK068083) and LOC_Os01g23680 (accession AK066580) for qSES1.3, and LOC_Os01g71240 (gene OsACA6, accession AK070064) for qSES1.11. Among them, LOC_Os01g71240 (OsACA6) encodes a calcium-transporting ATPase, plasma membrane-type.
Haplotype analysis of targeted genes LOC_Os01g23640, LOC_Os01g23680, and LOC_Os01g71240 for SES scores. The letter on the histogram (a, b, and c) indicated multiple comparison results at the significant level of 0.05 in the entire panel. The value on the histogram was the number of accessions of each haplotype
Discussion
To date, many QTLs related to salt tolerance have been mapped by using bi-parental linkage mapping populations and GWAS (Koyama et al. 2001; Sabouri and Sabouri 2008; Shi et al. 2017; Batayeva et al. 2018). They have resulted from GWAS analysis of small-size populations containing 191, 219, 295, or 478 accessions by EMMAX, GAPIT, or TASSEL (Shi et al. 2017; Batayeva et al. 2018; Naveed et al. 2018; Li et al. 2019). However, it is very difficult to compare the positions of the QTLs because they were detected on different panels and with different markers. Indeed, our results indicated that the QTLs detected by GWAS analysis were influenced by the number of accessions, number of SNPs/markers, and methods of analysis. First, in this research, among the 17 QTLs detected in one or several of the four panels through GWAS analysis using different ranges of SNPs selection by one or several methods, only two of them were detected in three of four panels and none was in all four panels, although these panels showed a similar population diversity as revealed by the PCA plot of genotyping data. The detected QTLs in the present study differed from those detected through GWAS previously by Shi et al. (2017) and Batayeva et al. (2018). Second, only 4 of the 17 detected QTLs were identified with all three methods, while 3 additional QTLs were detected with both FaST-LMM and BLINK. Yan et al. (2019) also reported that either different methods or amounts of input data often resulted in dissimilar associations with the target traits. Therefore, validating the results of GWAS by different methods is also recommended because each method uses a different algorithm for GWAS.
Four important QTLs, viz. qSES1.1, qSES1.3, qSES2.1, and qSES3.1, were identified in this study by all three methods used. To check the results, we also conducted a confirmatory experiment with 1,332 accessions and also identified these same important QTLs in this experiment (Supplementary Table S2b). It means that these QTLs were reliable. qSES1.1 co-located with a previously detected QTL related to Na+ uptake, K+ uptake, and the ratio of Na+/K+ in the rice tissues under salt stress (Koyama et al. (2001)), while the others were novel and were not published in Rice Genome Annotation Project (RGAP)—Rice Genome Annotation (Osa1) Release 7 [http://rice.uga.edu/cgi-bin/gbrowse/rice/, (Kawahara et al. 2013)], and Q-TARO: QTL Annotation Rice Online Database [http://qtaro.abr.affrc.go.jp/ (Yamamoto et al. 2012)].
Among the three retained candidate genes, LOC_Os01g71240 (OsACA6 gene, accession AK070064) encodes a calcium-transporting ATPase and has been reported to be related to abiotic stress such – as cold tolerance – in rice (Singh et al. 2014; Yamada et al. 2014). It has been well-documented that NaCl treatment caused an imbalance of Na+/Ca2+ homeostasis (Hu and Schmidhalter 2005; Munns and Tester 2008; Wang et al. 2012). The upregulation of the Ca2+-ATPase transcript level in NaCl treatment might lead to increased capacity of Ca2+-ATPase and help in lowering the cytosolic calcium level and thereby might be involved in the maintenance of Ca2+ homeostasis (Singh et al. 2014).
The identified QTLs and candidate genes provided useful genetic information for future functional characterization to improve salt tolerance in rice. Nevertheless, further study for the validation of these candidate genes should be considered. Based on the finding of these candidate genes, we chose some materials with expected high salt tolerance (SES score at 1 or 3, Supplementary Table S6). In that shortlist, Nona Bokra is a well-known highly salt-tolerant cultivar (Lutts et al. 1996). The list also contained some cultivars such as Taichung 65 or some Vietnamese cultivars such as Nep hoa vang, Nep ngau, X 21, and X 23. Noticeably, the highly salt-tolerant cultivar Pokkali is not on this list. It may suggest the existence of different genetic mechanisms controlling salt tolerance between the varieties.
Conclusion
We conducted GWAS analysis for SES by using different panels and numbers of SNPs by different methods of GWAS analysis. We propose to use TASSEL for panels with a low number of accessions and use BLINK or FaST-LMM for all the population sizes. We identified four important QTLs related to the salt tolerance of rice at the seedling stage. They included one QTL that co-located with a previously detected one and three other novel QTLs. Gene-based association analysis and haplotype analysis allowed us to detect three candidate genes, viz. LOC_Os01g23640 and LOC_Os23680 in qSES1.1, and LOC_Os01g71240 in qSES1.3 for the association with salt tolerance in rice. This detection provides useful germplasm and genetic information for the future improvement of salt tolerance in rice.
Abbreviations
- Chr.:
-
Chromosome
- GWAS:
-
Genome-wide association study
- LD:
-
Linkage disequilibrium
- SNP:
-
Single nucleotide polymorphism
- PCA:
-
Principal components analysis
- QTL:
-
Quantitative trait loci
- QTN:
-
Quantitative trait nucleotide
- SES:
-
Standard evaluation score
References
3K R.G.P. (2014) The 3,000 rice genomes project. GigaScience 3:3–7. https://doi.org/10.1186/2047-217X-3-7
Almeida DM, Gregorio GB, Oliveira MM, Saibo NJM (2017) Five novel transcription factors as potential regulators of OsNHX1 gene expression in a salt tolerant rice genotype. Plant Mol Biol 93:61–77. https://doi.org/10.1007/s11103-016-0547-7
Aravind J, Mukesh Sankar S, Wankhede DP, Kaur V (2020). augmentedRCBD: Analysis of augmented randomised complete block designs. R package version 0.1.6.9000, https://aravindj.github.io/augmentedRCBD/https://cran.r-project.org/package=augmentedRCBD
Batayeva D, Labaco B, Ye C, Li X, Usenbekov B, Rysbekova A, Dyuskalieva G, Vergara G, Reinke R, Leung H (2018) Genome-wide association study of seedling stage salinity tolerance in temperate japonica rice germplasm. BMC Genet 19:2. https://doi.org/10.1186/s12863-017-0590-7
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Stat Soc: Ser B (methodol) 57(1):289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
FAO (2021) FAOSTAT Database. http://www.fao.org/faostat/en/#data/QC (accessed January 6, 2021).
Federer WT, Raghavarao D (1975) On Augmented Designs. Biometrics 31:29–35. https://doi.org/10.2307/2529707
Guo Z, Zhou S, Wang S, Li WX, Du H, Xu Y (2021) Identification of major QTL for waterlogging tolerance in maize using genome-wide association study and bulked sample analysis. J Appl Genetics 62:405–418. https://doi.org/10.1007/s13353-021-00629-0
Hu Y, Schmidhalter U (2005) Drought and salinity: A comparison of their effects on mineral nutrition of plants. J Plant Nutr Soil Sci 168:541–549. https://doi.org/10.1002/jpln.200420516
Huang M, Liu X, Zhou Y, Summers RM, Zhang Z (2019) BLINK: a package for the next level of genome-wide association studies with both individuals and markers in the millions. GigaScience 8:giy154. https://doi.org/10.1093/gigascience/giy154
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z, Li M, Fan D, Guo Y, Wang A, Wang L, Deng L, Li W, Lu Y, Weng Q, Liu K, Huang T, Zhou T, Jing Y, Li W, Lin Z, Buckler ES, Qian Q, Zhang Q-F, Li J, Han B (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967. https://doi.org/10.1038/ng.695
IRRI (2013) Standard Evaluation System (SES) for Rice. p38. 5th ed. The International Rice Research Institute (IRRI), Los Baños, Philippines, 1099 Manila, Philippines
Kawahara Y, Bastide M, Hamilton J, Kanamori H, Mccombie W, Ouyang S, Schwartz D, Tanaka T, Wu J, Zhou S, Childs K, Davidson R, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee S, Kim J, Numa H, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6:1–10. https://doi.org/10.1186/1939-8433-6-4
Kong X-Q, Gao X-H, Sun W, An J, Zhao Y-X, Zhang H (2011) Cloning and functional characterization of a cation–chloride cotransporter gene OsCCC1. Plant Mol Biol 75:567–578. https://doi.org/10.1007/s11103-011-9744-6
Koyama ML, Levesley A, Koebner RMD, Flowers TJ, Yeo AR (2001) Quantitative trait loci for component physiological traits determining salt tolerance in rice. Plant Physiol 125:406–422. https://doi.org/10.1104/pp.125.1.406
Li N, Zheng H, Cui J, Wang J, Liu H, Sun J, Liu T, Zhao H, Lai Y, Zou D (2019) Genome-wide association study and candidate gene analysis of alkalinity tolerance in japonica rice germplasm at the seedling stage. Rice 12:24. https://doi.org/10.1186/s12284-019-0285-y
Lippert C, Listgarten J, Liu Y, Kadie CM, Davidson RI, Heckerman D (2011) FaST linear mixed models for genome-wide association studies. Nat Methods 8:833
Liu C, Chen K, Zhao X, Wang X, Shen C, Zhu Y, Dai M, Qiu X, Yang R, Xing D, Pang Y, Xu J (2019) Identification of genes for salt tolerance and yield-related traits in rice plants grown hydroponically and under saline field conditions by genome-wide association study. Rice 12:88. https://doi.org/10.1186/s12284-019-0349-z
Lutts S, Kinet JM, Bouharmont J (1996) Effects of salt stress on growth, mineral nutrition and proline accumulation in relation to osmotic adjustment in rice (Oryza sativa L.) cultivars differing in salinity resistance. Plant Growth Regul 19:207–218. https://doi.org/10.1007/BF00037793
Maas EV, Hoffman GJ (1977) Crop salt tolerance–current assessment. J Irrig Drain Div 103:115–134
Mansueto L, Fuentes RR, Borja FN, Detras J, Abriol-Santos JM, Chebotarov D, Sanciangco M, Palis K, Copetti D, Poliakov A, Dubchak I, Solovyev V, Wing RA, Hamilton RS, Mauleon R, McNally KL, Alexandrov N (2016) Rice SNP-seek database update: new SNPs, indels, and queries. Nucleic Acids Res 45:D1075–D1081. https://doi.org/10.1093/nar/gkw1135
Mao F, Wu D, Lu F, Yi X, Gu Y, Liu B, Liu F, Tang T, Shi J, Zhao X, Liu L, Ji L (2022) QTL mapping and candidate gene analysis of low temperature germination in rice (Oryza sativa L.) using a genome wide association study. PeerJ 10:e13407. https://doi.org/10.7717/peerj.13407
Munns R (2005) Genes and Salt Tolerance. New Phytol 167:645–663
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Naveed SA, Zhang F, Zhang J, Zheng T-Q, Meng L-J, Pang Y-L, Xu J-L, Li Z-K (2018) Identification of QTN and candidate genes for salinity tolerance at the germination and seedling stages in rice by genome-wide association analyses. Sci Rep 8:6505. https://doi.org/10.1038/s41598-018-24946-3
Phan NTH, Heymans A, Bonnave M, Lutts S, Pham CV, Bertin P (2023) Nitrogen use efficiency of rice cultivars (Oryza sativa L.) under salt stress and low nitrogen conditions. J Plant Growth Regul 42:1789–1803. https://doi.org/10.1007/s00344-022-10660-y
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Human Genet 81:559–575. https://doi.org/10.1086/519795
Rahman M, Thomson M, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail A (2016) Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization. Ann Bot 117(6):1083–1097. https://doi.org/10.1093/aob/mcw030
Reddy INBL, Kim BK, Yoon IS, Kim KH, Kwon TR (2017) Salt tolerance in rice: focus on mechanisms and approaches. Rice Sci 24:123–144. https://doi.org/10.1016/j.rsci.2016.09.004
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146. https://doi.org/10.1038/ng1643
Sabouri H, Sabouri A (2008) New evidence of QTLs attributed to salinity tolerance in rice. Afr J Biotech 7:4376–4383
Sharma AR, Singh DP (1999) Rice. In: Smith DL, Hamel C (eds) Crop yield: physiology and processes. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 109–168
Shi Y, Gao L, Wu Z, Zhang X, Wang M, Zhang C, Zhang F, Zhou Y, Li Z (2017) Genome-wide association study of salt tolerance at the seed germination stage in rice. BMC Plant Biol 17:1–11. https://doi.org/10.1186/s12870-017-1044-0
Singh A, Kanwar P, Yadav A, Mishra M, Jha S, Baranwal V, Pandey A, Kapoor S, Tyagi A, Pandey G (2014) Genome-wide expressional and functional analysis of calcium transport elements during abiotic stress and development in rice. FEBS J 281:894–915. https://doi.org/10.1111/febs.12656
Thomson MJ, de Ocampo M, Egdane J, Rahman MA, Sajise AG, Adorada DL, Tumimbang-Raiz E, Blumwald E, Seraj ZI, Singh RK, Gregorio GB, Ismail AM (2010) Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 3:148–160. https://doi.org/10.1007/s12284-010-9053-8
Tiwari S, Krishnamurthy SL, Kumar V, Singh B, Rao AR, Mithra SVA, Rai V, Singh AK, Singh NK (2016) Mapping QTLs for salt tolerance in rice (Oryza sativa L.) by bulked segregant analysis of recombinant inbred lines using 50K SNP chip. PLOS ONE 11:e0153610. https://doi.org/10.1371/journal.pone.0153610
Vinod KK, Krishnan SG, Babu NN, Nagarajan M, Singh AK (2013) Improving salt tolerance in rice: Looking beyond the conventional. In: Ahmad P, Azooz MM, Prasad MNV (eds) Salt Stress in Plants. Springer, New York, New York, NY, pp 219–260
Wang H, Zhang M, Guo R, Shi D, Liu B, Lin X, Yang C (2012) Effects of salt stress on ion balance and nitrogen metabolism of old and young leaves in rice (Oryza sativa L.). BMC Plant Biology 12:194. https://doi.org/10.1186/1471-2229-12-194
Wang X, Pang Y, Zhang J, Wu Z, Chen K, Ali J, Ye G, Xu J, Li Z (2017) Genome-wide and gene-based association mapping for rice eating and cooking characteristics and protein content. Sci Rep 7:17203. https://doi.org/10.1038/s41598-017-17347-5
Yamada N, Theerawitaya C, Cha-um S, Kirdmanee C, Takabe T (2014) Expression and functional analysis of putative vacuolar Ca2+-transporters (CAXs and ACAs) in roots of salt tolerant and sensitive rice cultivars. Protoplasma 251:1067–1075. https://doi.org/10.1007/s00709-014-0615-2
Yamamoto E, Yonemaru J, Yamamoto T, Yano M (2012) OGRO: The overview of functionally characterized genes in rice online database. Rice 5:26
Yan NA, Burbridge C, Shi J, Liu J, Kusalik A (2019) Effects of input data quantity on genome-wide association studies (GWAS). Int J Data Min Bioinform 22:19. https://doi.org/10.1504/IJDMB.2019.099286
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Routine procedure for growing rice plants in culture solution. Laboratory Manual for Physiological Studies of Rice. The International Rice Research Institute, pp 61–66
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. https://doi.org/10.1038/ng1702
Zeng L, Shannon MC (2000) Salinity effects on seedling growth and yield components of rice. Crop Sci 40:996–1003. https://doi.org/10.2135/cropsci2000.404996x
Zhang Y-M (2019) Editorial: The applications of new multi-locus GWAS methodologies in the genetic dissection of complex traits. Front Plant Sci 9:100. https://doi.org/10.3389/fpls.2019.00100
Zhang Z, Ersoz E, Lai C-Q, Todhunter RJ, Tiwari HK, Gore MA, Bradbury PJ, Yu J, Arnett DK, Ordovas JM, Buckler ES (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355–360. https://doi.org/10.1038/ng.546
Funding
This research received support from the Belgian Académie de Recherche et d’Enseignement Supérieur – Commission de la Coopération au Développement (ARES-CCD: www.ares-ac.be) and from Vietnam National University of Agriculture (T2021-01–04). The rice accessions were provided by IRRI to UCLouvain under SMTA2018-0139 and SMTA2018-0140 signed between these organizations.
Author information
Authors and Affiliations
Contributions
Conceptualization: Pierre Bertin, Cuong Van Pham, and Nhung Thi Hong Phan; experimental conducting: Nhung Thi Hong Phan and Pierre Bertin; data collection: Nhung Thi Hong Phan; formal analysis: Nhung Thi Hong Phan, Hanh Thi Tang; Loc Van Nguyen, Long Viet Nguyen; funding acquisition: Pierre Bertin, Cuong Van Pham, and Nhung Thi Hong Phan; supervision: Pierre Bertin and Cuong Van Pham; writing – original draft: Nhung Thi Hong Phan and Hanh Thi Tang; writing – review and editing: Nhung Thi Hong Phan and Pierre Bertin.
Corresponding author
Ethics declarations
Ethics approval
This work did not involve the studies of human or animal subjects.
Consent to participate
Not applicable.
Consent for publication
All authors agreed on the publication.
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Communicated by Izabela Pawłowicz
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Phan, N.T.H., Van Pham, C., Tang, H.T. et al. Integration of genome-wide association studies reveal loci associated with salt tolerance score of rice at the seedling stage. J Appl Genetics 64, 603–614 (2023). https://doi.org/10.1007/s13353-023-00775-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-023-00775-7