Abstract
Pepper (Capsicum annuum L.) is an economically important vegetable crop whose production and quality are severely reduced under adverse environmental stress conditions. The GATA transcription factors belonging to type IV zinc-finger proteins, play a significant role in regulating light morphogenesis, nitrate assimilation, and organ development in plants. However, the functional characteristics of GATA gene family during development and in response to environmental stresses have not yet been investigated in pepper. In this study, a total of 28 pepper GATA (CaGATA) genes were identified. To gain an overview of the CaGATAs, we analyzed their chromosomal distribution, gene structure, conservative domains, cis-elements, phylogeny, and evolutionary relationship. We divided 28 CaGATAs into four groups distributed on 10 chromosomes, and identified 7 paralogs in CaGATA family of pepper and 35 orthologous gene pairs between CaGATAs and Arabidopsis GATAs (AtGATAs). The results of promoter cis-element analysis and the quantitative real-time PCR (qRT-PCR) analysis revealed that CaGATA genes were involved in regulating the plant growth and development and the responses to various abiotic stresses and hormone treatments in pepper. Tissue-specific expression analysis showed that most CaGATA genes were preferentially expressed in flower buds, flowers, and leaves. Several CaGATA genes, especially CaGATA14, were significantly regulated under multiple abiotic stresses, and CaGATA21 and CaGATA27 were highly responsive to phytohormone treatments. Taken together, our results lay a foundation for the biological function analysis of GATA gene family in pepper.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the biological processes of plant seed germination, dormancy, senescence, and flower development and so on, different genes have different temporal and spatial expression characteristics regulated by transcription factors (TFs). TFs are key regulators to govern gene expression by specifically binding to cis-elements in the promoter region of the target gene, thus influencing or controlling many vital biological processes, such as cellular morphogenesis, signaling transduction, and environmental stress responses (Riechmann et al. 2000; Wray et al. 2003; Jin et al. 2015). Plants are always faced with the impact of abiotic stress in the common adverse environmental conditions, including drought, high soil salinity, heat, cold, and oxidative stress, which often lead to the slowdown of plant growth and development or even death, resulting in a reduction in crop production. Fortunately, plants have developed a series of protective mechanisms under stress condition in the long-term evolution, among which transcriptional regulations of gene expression play a vital role in a variety of developmental processes and adaptive stress responses to environmental stimuli in plants (Xu et al. 2007). GATA TFs are evolutionarily conserved transcription regulators existing in a wide range of eukaryotic organisms including fungi, plants, and metazoans. GATA genes are known for their DNA-binding preference to the consensus sequence (A/T)GATA(A/G), and they contain type IV zinc-finger motif with the consensus sequence CX2CX17–20CX2C (Teakle et al. 2002; Reyes et al. 2004). In animals, GATA proteins have two CX2CX17CX2C zinc-finger domains, but only the C-terminal zinc-finger is involved in DNA binding (Lowry and Atchley 2000). Most fungal GATA TFs contain only a single CX2CX17CX2C or CX2CX18CX2C domain (Scazzocchio 2000). A previous study of predicted 30 GATA family members in Arabidopsis and 29 in rice (Oryza sativa) shows that most plant GATA TFs only contain a single CX2CX18CX2C domain, but a few plant GATA TFs contain CX2CX20CX2C domain or two zinc-finger domains (Reyes et al. 2004; Bi et al. 2005).
The fundamental role of GATA TFs has been widely investigated in animals and fungi. GATA TFs have been reported to have important effects on development, differentiation, and cell proliferation in animals (Patient and McGhee 2002; Yang et al. 2020). In fungi, GATA TFs are involved in the regulation of nitrogen metabolism, circadian regulation, dimorphic transition and microsclerotium development, and siderophore production (Scazzocchio 2000; Song et al. 2020; Xin et al. 2020). In plants, GATA TFs play significant roles in the regulation of plant growth and development, such as flower development, shoot apical meristem, and leaf development (Zhang et al. 2013; Ding et al. 2015a; b). The initial interest in plant GATA TFs is due to the extensive regulation of light-response and circadian-regulation gene expression (Manfield et al. 2007; Shaikhali et al. 2012). Further studies have revealed the involvement of GATA TFs in the regulation of carbon and nitrogen metabolism, seed germination, cotyledon development, plant morphogenesis, and flower formation (Bi et al. 2005; Liu et al. 2005; Luo et al. 2010; Ding et al. 2015b; Bastakis et al. 2018). In addition, active participation in preventing photooxidative damage via tetrapyrrole biosynthesis (TPB) is found in Arabidopsis GATA TFs (Kobayashi and Masuda 2016). In recent years, Arabidopsis B-GATA TFs with a C-terminal LLM domain or an N-terminal HAN domain have been widely studied and described as regulators of plant growth and development, and hormones (Behringer et al. 2014; Behringer and Schwechheimer 2015; Klermund et al. 2016; Kobayashi et al. 2017). Generally, researchers have begun to study the function of GATAs in stress response, and hormone signaling, especially jasmonic acid and gibberellin signaling (Richter et al. 2010; 2013a; b), but there have been relatively few reports. However, there are few studies about the role of GATA TFs in regulating pepper development, let alone in response to abiotic stress and hormones.
As a solanaceous vegetable crop with a great economic value worldwide, pepper suffers from a variety of abiotic stresses in the changing environment, which have become the primary factors that can affect crop growth and development, quality, and yield (Wang et al. 2003). Understanding the mechanism of pepper growth and development and abiotic stress tolerance is essential for sustainable agriculture. Although the crucial roles of GATA TFs in model plants such as Arabidopsis and rice have been investigated, the characterization of the pepper GATA family and their responses to various stresses at the molecular level have not been well-known yet. In this study, we identified 28 CaGATA genes from pepper genome and investigated their chromosomal distribution, gene structure, conservative domains, cis-elements, and evolutionary relationship. Additionally, qRT-PCR analysis of GATA family revealed the tissue-specific expression functions of this family and its response to various abiotic stresses and hormone treatments. Our results shed light on the potential role of GATA TFs under abiotic stresses and hormone treatments in pepper.
Materials and methods
Genome-wide identification of GATA TFs in pepper
The hidden Markov model (HMM) profile of the GATA domain (PF00320) was downloaded from protein family (Pfam) database (http://pfam.xfam.org/) and used as queries to search the Pepper Genome Database (PGD, http://peppergenome.snu.ac.kr/, Zunla-1 genomes) to identify all the putative GATA genes of pepper by hmmer 3.2.1 software. All output genes with E-value ≤ 1e−5 were collected and confirmed by the Pfam and SMART (http://smart.embl-heidelberg.de/), and the incorrectly putative genes were eliminated. The non-redundant and identified genes were designated as pepper GATA genes (CaGATAs). The chromosomal location data, protein sequences, and genomic sequences of identified CaGATAs were derived from PGD, and their chromosomal locations were mapped onto chromosomes using TBtools software (Chen et al. 2020). The protein sequences were analyzed with EXPASY PROTOPARAM (http://web.expasy.org/ protparam/) to obtain the number of amino acids, molecular weight (Mw), and theoretical isoelectric point (pI). The subcellular locations were predicted using WoLF PSORT program (http://wolfpsort.org/), BUSCA (http://busca.biocomp.unibo.it/), and CELLO2GO (http://cello.life.nctu.edu.tw/cello2go/) (Horton et al. 2007; Yu et al. 2014; Savojardo et al. 2018).
Multiple sequence alignment and conserved domain analysis
The amino acid sequences of CaGATA proteins downloaded from PGD were aligned by using MEGA-X software and visualized by Jalview 2.11.1.0. The conserved domains within CaGATA proteins were searched and analyzed by Web CD-Search Tool (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), and then visualized by TBtools software.
Phylogenetic tree and gene duplication analysis
GATA proteins of A. thaliana (Arabidopsis thaliana) and O. sativa (Oryza sativa) were downloaded from the Arabidopsis Information Resource (https://www.arabidopsis.org/) and the Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/cgi-bin/ORF_infopage.cgi), respectively. The full amino acid sequences of GATA members from pepper, Arabidopsis and rice were aligned, and then, the phylogenetic tree was constructed by the maximum likelihood (ML) method with 1000 bootstrap tests, partial deletion, Jones-Taylor-Thornton (JTT) model by MEGA-X software. The whole-genome protein sequences of GATAs were compared between C. annuum (Capsicum annuum) and A. thaliana or within C. annuum for determining homology using BLASTP with an E-value ≤ 1 × 10−5, and then, MCScanX was employed to analyze the duplication events in these two species or in C. annuum alone. Finally, TBtools software was applied to display their relationship network.
Promoter sequence, gene structure, and conserved motif analysis
The 2000-bp genomic DNA sequence upstream of the start codon of each CaGATA gene was obtained from PGD, and the cis-elements were identified using the PlantCARE database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) (Lescot et al. 2002). The exon/intron structures of CaGATA genes were analyzed with GSDS 2.0 (http://gsds.cbi.pku.edu.cn/index.php) by aligning the cDNA sequences with the corresponding genomic sequences (Hu et al. 2015). The identification of CaGATA-conserved motifs was performed using the MEME program (http://meme-suite.org/tools/meme), and the identified motifs were confirmed by Pfam and SMART.
Plant growth conditions and treatments
In this study, the pepper line 17-03 was used for analysis of CaGATA gene expression. The seedlings were grown under natural light in a greenhouse with temperature regime of 24–28 °C day/20–25 °C night and relative humidity of approximate 70–80% until most of the fruits matured. The 13 tissues, namely, root, young stem, old stem, young leaf, mature leaf, old leaf, flower bud, flower, young fruit, expansion-stage fruit, mature green fruit, breaker-stage fruit, and red ripe–stage fruit, were collected from these plants to analyze the tissue-specific expression. To investigate the response to abiotic stresses and hormone treatments, the seedlings were grown in a growth chamber at 25 ± 2 °C with a photoperiod of 16-h light/8-h dark and light intensity of 200 μmol m−2 s−1 until reaching the age of 6–8 true leaves, and they were treated with drought, salinity (400 mM NaCl), cold (4 °C), heat (40 °C), methyl viologen (MV, 10 μM), ABA (100 μM), GA (100 μM), ethephon (ET, 200 μM), JA (100 μM), SA (100 μM), or IAA (100 μM), respectively. The third fully expanded leaf from the plant top was sampled from five or more seedlings in the control and treatment groups at 1 h, 3 h, 6 h, 12 h, 24 h, and 48 h after initiation of the treatments. All the samples were frozen in liquid nitrogen immediately and stored at − 80 °C until RNA isolation.
RNA extraction and qRT-PCR analysis
Total RNA was extracted with Trizol Reagent (Magen, Guangzhou, China), and the cDNA was synthesized using HiScript II 1st Strand cDNA Synthesis Kit (+ gDNA wiper) according to the manufacturer’s instructions (Vazyme, Nanjing, China). For qRT-PCR assay, primer pairs for pepper CaGATA genes were designed by Primer3 (http://primer3.ut.ee/) and their specificity of designed fragment was tested by SGN (https://solgenomics.net/). CaUBI-3 was used as the internal reference gene in pepper (Wan et al. 2011). The specific primer pairs of 23 CaGATA genes were designed and are listed in Table S4. The qRT-PCR reactions were performed with three technical replicates in 10-μl volumes containing 5-μL Fast SYBR™ Green Master Mix (2×) (Thermo Fisher, USA), 4-μL cDNA template (100 ng/μL), and 0.5 μL of each primer (10 μM). The PCR cycling conditions were as follows: pre-denaturation at 95 °C for 5 min, followed by 40 cycles of denaturation at 95 °C for 15 s, annealing at 58 °C for 20 s, and extension at 72 °C for 30 s. The relative expression level was calculated using the 2−ΔΔCT method (Schmittgen and Livak 2008), and then, the heatmaps were generated using Heml 1.0.3.7 software.
Results
Identification and characterization of CaGATA gene family in pepper
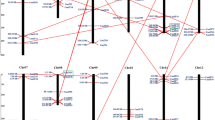
A total of 28 members of the GATA gene family were identified from PGD by the HMM analysis, and their uniqueness was verified. To determine their chromosomal distributions, all the CaGATA genes were mapped on the chromosomes according to their positions in the pepper genome database using the TBtools software. A total of 26 CaGATA genes were mapped onto 10 pepper chromosomes out of the total 12 chromosomes, except for chromosome 7 and chromosome 10, but the remaining 2 CaGATA genes were located on unanchored scaffolds. Chromosomes 5 contained the largest number of CaGATA genes, accounting for 17.86% of the total number. Three chromosomes, namely, chromosome 9, 11, and 12, each contained only one CaGATA gene: CaGATA26, CaGATA27, and CaGATA28, respectively (Fig. 1).
On the basis of their chromosomal locations, the 28 GATA genes were renamed CaGATA1-CaGATA28. Detailed information about each CaGATA gene is shown in Table 1. The open reading frame (ORF) lengths of CaGATA genes ranged from 360 to 1632 bp, and these CaGATA genes encoded 119–543 amino acids (aa). The molecular weights of these predicted CaGATA proteins ranged from approximate 13.27 to 60.32 kDa, and the theoretical isoelectric points ranged from 5.17 to 9.92. Of the 28 CaGATA proteins, 10 members shared the similar localization to nucleus; the rest 18 members were localized to more than one compartment.
Phylogenetic analysis and sequence alignment
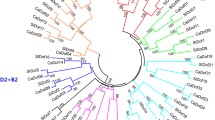
To better analyze the evolutionary relationships of CaGATA genes, an unrooted phylogenetic tree of 28 CaGATA proteins, 30 AtGATA proteins, and 29 OsGATA proteins was generated (Table S1 and Table S5). According to our cluster analysis combined with previous GATA gene study results in model plants, 87 GATA proteins were divided into four groups, namely, groups A, B, C, and D, containing 38, 29, 12, and 8 members, respectively (Fig. 2). Our cluster analysis results were largely similar to the previous phylogenetic analyses of GATA genes in Arabidopsis and rice (Reyes et al. 2004). Of these 87 proteins, 28 CaGATA proteins were distributed over all 4 groups, and the number of each group was 12, 9, 4, and 3, respectively. To further analyze the sequence features of the 28 CaGATA proteins, their amino acid sequences were aligned, and the results revealed that most CaGATA proteins contained only a single zinc-finger domain CX2CX18–20CX2C, and that other CaGATA proteins including CaGATA2 and CaGATA21 were found to lose some amino acids (Fig. 3). Group A, B, and D CaGATA proteins contained only a single CX2CX18CX2C domain, except CaGATA9 and CaGATA21. However, group C CaGATA proteins contained the integrated conservative domain CX2CX20CX2C.
An unrooted phylogenetic tree displaying the relationships of GATA genes in C. annuum, A. thaliana, and O. sativa. The 28 CaGATA proteins, 30 AtGATA proteins, and 29 OsGATA proteins were used to construct the tree. GATA proteins from different species are indicated by different symbols: C. annuum by solid circles, A. thaliana by solid squares, and O. sativa by solid triangles. The four different groups are indicated by different colors. Numbers at nodes represent bootstrap values based on 1000 replicates
Gene structure and conservative motif analysis
To better analyze the structure of CaGATA genes and the conserved motifs of CaGATA proteins, we constructed a phylogenetic tree of CaGATA genes. Groups A and B contained 2–3 CDSs, while groups C and D had 6–10 CDSs. Furthermore, CaGATA genes within the same group had analogous gene structures (Fig. 4). We identified 10 motifs (designated as motif 1 to 10) from the 28 CaGATA proteins with each gene containing motif 1 which was assigned to the ZnF_GATA according to annotations in the Pfam database (Fig. 4). Most CaGATA proteins within the same group contained similar motifs. For example, all the CaGATA proteins in group D contained conservative motif 4 representing ASXH in addition to motif 1 (Table S2 and Fig. S1). Taken together, these results revealed that different groups differed in gene structure significantly from each other.
A schematic diagram of the amino acid motifs of CaGATA proteins and gene structures of CaGATA genes in pepper. The positions of the sequences encoding the GATA motifs in CaGATA genes are shown. The phylogenetic tree displaying the relationships of CaGATA genes was constructed using MEGA-X according to the maximum likelihood (ML) method with 1000 bootstrap test replicates
Gene duplication of CaGATA genes
Gene families were generated through either tandem duplication or large-scale segmental duplication during evolution (Cannon et al. 2004). Therefore, we constructed comparative syntenic maps of pepper versus Arabidopsis. Thirty-five pairs of orthologous GATA genes were identified between CaGATAs and AtGATAs (Fig. 5 and Table S3). To analyze CaGATA gene duplication events, we identified 7 paralogous gene pairs located in genome of pepper, which were CaGATA3/CaGATA5, CaGATA6/CaGATA8, CaGATA7/CaGATA14, CaGATA11/CaGATA22, CaGATA20/CaGATA27, CaGATA3/CaGATA21, and CaGATA7/CaGATA25, respectively (Fig. S2). Interestingly, each paralogous gene pair was clustered together (Fig. 4), and all the GATA homologous gene pairs were located within the same group in the phylogenetic tree (Fig. 2).
Analysis of promoter sequences of CaGATA genes
To further explore the function and regulatory patterns of CaGATA genes, putative cis-regulatory elements in the 2000-bp genomic DNA sequence upstream of the start codon of each CaGATA gene were analyzed by using the PlantCARE database. Twenty major types of cis-elements were identified on the promoters of the CaGATA genes (Fig. 6 and Fig. S3). More than 24 types of light-responsive cis-elements, such as GT1-motif, Box4, TCT-motif, G-Box, and GATA-motif, were observed across almost all CaGATA genes. Light-responsive cis-elements accounted for the largest proportion of all the cis-elements on each gene promoter, followed by abiotic stress-responsive cis-elements including the cis-elements of MYB and MYC (Fig. S3 and Table S7) (Urao et al. 1993; Iwasaki et al. 1995; Abe et al. 1997; Agarwal et al. 2006; Onishi et al. 2006; Zhang et al. 2016b; Zhang et al. 2019). In addition, various cis-elements involved in hormone response (such as MeJA, abscisic acid, salicylic acid, gibberellin, and auxin), stress response (such as drought, low temperature, wound, anaerobic induction), defense and stress responsiveness, meristem expression, and circadian control, and so on, were also identified in promoter sequences of CaGATA genes. Predicted cis-elements suggested potential roles of pepper GATA genes in light, stress responses, phytohormone, and development.
Tissue-specific expression CaGATA genes in pepper
In order to examine the expression patterns of CaGATA genes during pepper growth and development, 23 genes were analyzed in 13 tissues, including root, young stem, old stem, young leaf, mature leaf, old leaf, flower bud, flower, young fruit, expansion-stage fruit, mature green fruit, breaker-stage fruit, and red ripe–stage fruit, using qRT-PCR (Table S4 and Table S6). A heat map was generated to show the expression profiles (Fig. 7). Twenty-three CaGATA genes were found to be expressed at different levels in these tissues. The relative expression of each gene in each tissue was compared with the expression level in the root. The results indicated that the relative expressions of genes were high in young tissue. For example, the relative expressions of genes in young stems were higher than in old stems, and those in young leaves were higher than those in mature leaves which in turn were higher than in old leaves. Similarly, those in flower bud were higher in flower. Interestingly, although the relative expressions of genes were high in young fruits, they were also relatively high in red ripe–stage fruits during fruit development, and they were even higher in red ripe fruits than in young fruits. Most CaGATA genes exhibited an extremely high expression in leaves, flowers, and flower buds, but a relatively low expression in fruits, especially in expansion-stage fruits, mature green fruits, and breaker-stage fruits. It was worth noting that CaGATA3, CaGATA13, CaGATA19, and CaGATA23 shared similar expression pattern with low levels in most tissues.
Expression profiles of 23 CaGATA genes in various pepper tissues by qRT-PCR. The relative expression level of each CaGATA gene in each tissue is compared with that in the root. The heat map is generated using Heml 1.0.3.7, and the bar on the right side of the heatmap indicates relative expression values
Expression profiles of CaGATA genes in response to abiotic stresses and hormone treatments
To understand the expression patterns of CaGATA genes under different environmental conditions, the expression pattern of 23 selected CaGATA genes was examined in response to various abiotic stresses and hormone treatments by qRT-PCR (Table S6). Heat map exhibited transcript level fold change in response to abiotic stresses and hormone treatments (Fig. 8 and Fig. 9). Cluster analysis results indicated that different expression levels of CaGATA genes were observed under various stresses (Fig. 8). Most CaGATA genes were significantly induced by oxidative stress (MV), but repressed by cold stress, salt stress, and heat stress, especially by drought stress, at different treatment time points. The expression levels of most genes were induced and reached a peak at the 12th hour of oxidative stress (MV), and then declined (Fig. S4a). More than half of the genes were upregulated at the 1st hour or the 3rd hour of salt treatment (Fig. S4b). But the expressions of CaGATA16 and CaGATA17 were not induced under salt stress; until at hour 48, they were downregulated (Fig. 8). Additionally, three CaGATA genes were significantly upregulated by heat stress, one of which, CaGATA4, was remarkably induced. Some genes were slightly induced, whereas four genes were significantly downregulated by heat stress, including CaGATA6, CaGATA19, CaGATA25, and CaGATA26 (Fig. S4c). Under cold stress, some genes were upregulated at certain time points, especially at hour 6, while 7 genes were downregulated (Fig. S4d). More than half of the genes were downregulated, and a few genes were slightly upregulated by drought stress (Fig. S4e). Interestingly, CaGATA14 was highly expressed under cold stress and drought stress (Fig. S4d and Fig. S4e).
The qPCR expression analysis of 23 CaGATA genes in response to abiotic stress. The pepper seedlings at the age of 6–8 true leaves were dealt with MV (10 μM), salinity (400 mM NaCl), heat (40 °C), cold (6 °C), or drought, respectively. The numbers on the right labels correspond to the duration of abiotic stresses treatments, i.e., 1 h, 3 h, 6 h, 12 h, 24 h, and 48 h respectively under each treatment. The relative expression level of each CaGATA gene under each abiotic stress at each point in time is compared with that under normal conditions at the corresponding time point. In MV treatment, water treatment was used as reference. The heat map was generated using Heml 1.0.3.7, and the bar on the right side of the heat map represents relative expression values
The qPCR expression analysis of CaGATA genes in response to hormone treatments. Pepper seedlings at the age of 6–8 true leaves were dealt with ABA (100 μM), GA (100 μM), ET (200 μM), JA (100 μM), SA (100 μM), or IAA (100 μM), respectively. The numbers on the right labels represent the time course of the hormone treatment, such as 1 h, 3 h, 6 h, 12 h, 24 h, and 48 h. The relative expression level of each CaGATA gene under each hormone treatment at each time point was compared with that sprayed with H2O at the corresponding time point. The heat map was generated using Heml 1.0.3.7, and the bar on the right side of the heat map represents relative expression values
In response to hormone treatments, most CaGATA genes were inhibited or not significantly upregulated after spraying different exogenous hormones. As shown in Fig. 9, more than half of CaGATA genes were lowly expressed or not induced under ABA treatment and ET treatment, similarly CaGATA4, CaGATA16, and CaGATA17 under GA treatment; CaGATA3, CaGATA6, CaGATA11, CaGATA15, CaGATA16, and CaGATA25 under SA treatment; and CaGATA3, CaGATA10, CaGATA11, CaGATA16, and CaGATA28 under JA treatment. CaGATA3, CaGATA4, CaGATA12, and CaGATA18 were not sensitive to IAA treatments. But some genes were upregulated at one or more time points, such as, CaGATA27 under ABA treatment; CaGATA1 under GA treatment; CaGATA21 under ET treatment; and CaGATA23 under SA, IAA, and JA treatments (Fig. 9). CaGATA7, CaGATA14, CaGATA21, CaGATA22, CaGATA23, and CaGATA27 responded to all hormone treatments (Fig. S5).
Discussion
Plant transcription factors, such as WRKY, MYB, bHLH, and zinc-finger, play a key role in governing gene regulation that mediates diverse biological processes in plant developmental processes, stress responses, and hormone signaling pathways (Franco-Zorrilla et al. 2014; Zhang et al. 2016a; Geng and Liu 2018; Li et al. 2019b; Liu et al. 2019). GATA proteins are defined as GATA TFs due to their specific binding to the consensus sequence (A/T) GATA (A/G) (Reyes et al. 2004). Although members of the GATA gene family have been characterized in model plants such as Arabidopsis and rice, a comprehensive analysis of this family has not been reported in the nutritionally and economically important pepper. Therefore, genome-wide characterization and expression analysis of GATA gene family in pepper will help us understand further CaGATA gene functions.
In this study, 28 CaGATA genes were identified and systematically analyzed based on the pepper genome. Consistent with GATA genes in Arabidopsis and rice, 28 CaGATA genes were classified into four subfamilies, designated as group A to D, of which group A contained the largest number of CaGATA genes (Reyes et al. 2004). Most CaGATA genes from same group shared the similar sequences length, pI, Mw, gene structure, and conserved motifs. All the GATA homologous gene pairs identified from pepper genomes or from pepper versus Arabidopsis genomes were tightly clustered together in the phylogenetic tree, indicating that these homologous gene pairs were more closely related mutually, and the topologies of the phylogenetic tree were consistent with the synteny analysis results. Variation in GATA family gene may be caused by gene or genome duplication events, which have been considered to be the primary source of genetic novelty in plant evolution (Yang et al. 2006; Yang et al. 2008; Gillis et al. 2009). The result exposed that the evolution of CaGATA genes was promoted by segmental duplication events.
Sequence determines structure, structure determines function. We found that group A, B, and D CaGATA proteins contained a single CX2CX18CX2C domain, while members of group C were the most peculiar as they possess CX2CX20CX2C domain that was relatively rare in the conserved domains of plant GATA genes (Reyes et al. 2004). Apart from ZnF_GATA domain, group C had TIFY and CCT domains, except for CaGATA2. The fully conserved TIFY domain and family previously known as ZIM was first named in Arabidopsis (Vanholme et al. 2007). Gradually, genome-wide identification of the TIFY gene family and their expression profiles in stress and phytohormone responses in a variety of species was carried out (Ye et al. 2009; Zhu et al. 2013; Zhao et al. 2016; Wang et al. 2017; Xia et al. 2017; He et al. 2020; Liu et al. 2020; Zhang et al. 2020b). The TIFY domain is widespread in JASMONATE ZIM-domain (JAZ) family proteins that mediate JA signaling and stress response (Chini et al. 2009; Bai et al. 2011; Sun et al. 2017). The CCT domain was first identified in the CONSTANS (CO) protein that plays an active role in sustaining its diurnal accumulation dynamics during Arabidopsis photoperiodic flowering (Suarez-Lopez et al. 2001; Gil et al. 2017). Beyond the flowering time, CCT domain-containing genes are associated with grain yield (Zhang et al. 2015). Moreover, recent studies show that they also associate with plant growth, development, photosynthesis, nutrition use efficiency, and stress tolerance (Morita et al. 2015; Liu et al. 2016; Yang et al. 2018; Zhang et al. 2018; Li et al. 2019a). Thus, CaGATA13, CaGATA23, and CaGATA24 might be involved in regulating plant development and tolerance or susceptibility of diverse stresses regulated by TIFY and CCT domains in pepper. Group D was the smallest with three GATA members, viz. CaGATA3, CaGATA5, and CaGATA21. They not only had the GATA motif but also contained ASXH domain. At present, the research on the ASXH domain have mainly focused on animals, which it behaves genetically as an enhancer of trithorax and polycomb group (Baskind et al. 2009; Fisher et al. 2010; Aravind and Iyer 2012). However, groups A and B only contained ZnF_GATA domain, except for CaGATA18 in group A that contained another domain, PRK13367, that is related to gallate dioxygenase. Therefore, these different highly conserved domains lead to the specificity and functional diversity of the CaGATA protein subfamily. Similar to the domain, CaGATA genes in the same subgroup had a similar number of exons. There were 2 to 3 CDSs in groups A and B, and 6 to 10 CDSs in groups C and D. In addition to light-responsive cis-elements and abiotic stress-responsive cis-elements (MYB and MYC cis-elements) found on the promoters of all CaGATA genes, there were great differences in cis-elements even in the same subgroup, such as abscisic acid-responsive cis-elements in CaGATA13 promoters in group C, but not in CaGATA23 promoters, and gibberellin-responsive cis-elements in CaGATA23 promoters, but not in CaGATA13 promoters. This might lead to functional differences in CaGATA genes in the same subgroup. To sum up, no matter in terms of promoter, conserved motif, gene structure, or domain, the GATA genes of pepper have undergone moderate divergence over the course of evolution.
The GATA gene family plays important roles in plant growth and development, such as flowering, floral organ development, shoot apical meristem, carbon assimilation in the leaf, and chloroplast development (Zhang et al. 2013; Ding et al. 2015a; b; An et al. 2020), and some genes in this family are highly expressed in green tissues (Chiang et al. 2012; Bastakis et al. 2018). For example, HANABA TARANU regulates the shoot apical meristem and leaf development in cucumber, or communicates with the meristem through the PNH, regulates floral organ development via JAG and BOP2, and maintains boundary morphology through CKX3 during flower development in Arabidopsis (Ding et al. 2015a; b). To evaluate the role of CaGATA TFs in the organ development of pepper, the tissue-specific expression patterns of the 23 CaGATA genes in 13 tissues of pepper were investigated by qRT-PCR analysis, and the results showed that most CaGATA genes were highly expressed in leaves, flower buds and flowers, and stems, especially in the young and green parts. And there were many light-responsive cis-elements identified in the promoters of CaGATA genes. These results implied that CaGATA genes might be related to light-regulated development, which could improve photosynthesis rate and crop yield through increasing chlorophyll content in green parts (Zhu et al. 2010). Interestingly, CaGATA genes in the same group exhibited similar expression patterns in different plant tissues, like CaGATA12 and CaGATA18, CaGATA22 and CaGATA28, CaGATA13 and CaGATA23, and CaGATA5 and CaGATA21. In the same tissues, the relative expression level of CaGATA genes in young parts was higher than in mature or old parts, but the expression patterns varied from tissue to tissue. During the period of fruit development, most CaGATA genes were highly expressed in both young fruits and red ripe–stage fruits, but the expression levels of some genes were higher in red ripe–stage fruits than in young fruits, such as CaGATA1, CaGATA6, and CaGATA7. It could be seen that some CaGATA genes might also regulate the development of pepper fruit, and then affect the fruit marketability and yield of pepper. However, the relative expression levels of CaGATA3, CaGATA13, and CaGATA23 were low in almost all tissues, but high in root, suggesting that these genes might be involved in root development or plant tolerance to stresses such as drought and salinity (Kiryushkin et al. 2019; Seo et al. 2020). The function of GATA genes during organ development in pepper might be similar to that in other crops. For example, the phylogenetic analysis displayed that CaGATA26 was closely related to GNC (AtGATA21) and CGA1 (AtGATA22). In addition, CaGATA26 exhibited the orthologous relationship with them. The paralog GATA TFs GNC and CGA1 play essential roles in the regulation of chloroplast development, photosynthetic efficiency, and carbon and nitrogen metabolism (Scheible et al. 2004; Bi et al. 2005; Hudson et al. 2011; Chiang et al. 2012). Chloroplasts are organelles converting light energy into chemical energy to maintain plant growth, and they widely exist in various organs of plants, especially in green parts, such as leaves and stems. Chloroplast development is also affected by light (Armstrong et al. 1995; Neuhaus and Emes 2000). In this study, CaGATA26 exhibited high expression levels in leave and stems and contained the larger number of light-responsive cis-elements than the other CaGATA genes. Our results also suggested that CaGATA26 might be involved in light response and chloroplast development, thereby affecting plant growth (Zhang et al. 2020a).
Furthermore, the results of transcriptional level analysis during floral development indicated that more than half of the CaGATA genes were highly expressed in flower bud and flower. Some studies have shown that GATA genes are involved in the regulation of flower formation. HANABA TARANU, a GATA transcription factor, functions as a key repressor that regulates floral development via regulatory networks involving GATA3 family genes, HANL2, GNC, and GNL (Zhao et al. 2004; Zhang et al. 2013). GNC and GNL also interact with MADS box transcription factor SOC1 to control the flowering time in Arabidopsis (Richter et al. 2013a). The most of CaGATA genes might regulate floral organ development, which is an important trait and would be useful for genetic improvement of high-yield breeding of crops (Wu et al. 2020). Therefore, transcription factor GATA can be used to further study the regulation of floral organ development in plants, so as to explore the expression regulation and metabolic pathway of genes related to floral organ development. These data would contribute to the further study of the functions of CaGATA genes during plant growth and development and provide useful information for genetic improvement of high-yield breeding of pepper.
At present, most GATA TFs are reported to be involved in the functional regulation of plant growth and development, but there have been few reports on the biological functions related to abiotic stress and hormone response, especially in pepper (Richter et al. 2010; Richter et al. 2013a). Even though a small number of GATA genes are reported to be involved in stress response, they are also associated with the regulation of plant growth and development. There is a cross-repressive interaction between the GATA factors GNC and GNL and the MADS box transcription factor SOC1 in flowering time control on the one side and greening and cold tolerance on the other (Richter et al. 2013a). Considering this, we analyzed the expression patterns of 23 CaGATA genes in response to various abiotic stresses and phytohormone treatments, including oxidation (MV), salinity, drought, heat, cold, ABA, GA, ET, JA, SA, or IAA. The comprehensive expression profiles of a large number of CaGATA genes are also essential for discovering novel regulators involved in abiotic stress responses and hormone signal transduction. In the present study, we found that three CaGATA genes with the cis-elements of abiotic stress-responsive and defense and stress responsiveness in their promoter regions, namely, CaGATA7, CaGATA14, and CaGATA21, were highly regulated by multiple abiotic stresses and hormone treatments, indicating that these genes might act as important regulators in stress and hormone responses. Besides, most CaGATA genes were significantly induced by oxidative stress, but repressed by cold stress, salt stress, and heat stress at different time points, especially drought stress. Different from other genes, GATA14 was strongly induced and significantly upregulated by cold and drought stress, and this gene played a positive role in response pathway of drought and cold stress. In various hormone responses, most CaGATA genes were suppressed or not significantly induced. Most of CaGATA genes were lowly expressed or not induced under ABA treatment. The expression level of CaGATA7, CaGATA11, CaGATA16, CaGATA22, and CaGATA23 decreased after ET treatment. CaGATA3, CaGATA4, CaGATA12, and CaGATA18 were not sensitive to IAA treatments. Of course, some genes were slightly upregulated at one or more time points after hormone treatment, such as CaGATA1, CaGATA3, CaGATA5, CaGATA10, CaGATA21, and CaGATA28 at hour 6 of GA treatment or hour 3 of ET treatment. Previous research has proved that two paralogous GATA genes in Arabidopsis, GNC and GNL/CGA1, are involved in the regulation of auxin and gibberellin signaling (Richter et al. 2013b). GATA26, which had the orthologous relationship with GNC and CGA1, also responded to GA treatment and IAA treatment. Our results showed that CaGATA genes might also function in the response to abiotic stresses and hormone treatments, which remains to be investigated.
Conclusions
We identified 28 GATA genes in the pepper genome and systematically analyzed their chromosome distribution, gene structure, conservative domains, cis-elements, and evolutionary mechanism. In addition, their expression patterns in different tissues and in response to various abiotic stresses and hormone treatments were analyzed by qRT-PCR. Members of the CaGATA gene family were preferentially expressed in flower buds, flowers, and leaves, and stems. Several CaGATA genes, especially CaGATA14, were significantly regulated by multiple abiotic stresses, and CaGATA21 and CaGATA27 were highly responsive to phytohormone treatments. These studies show that peper GATA family genes not only affect plant growth, but also respond to and adapt to external environmental factors, which has important practical significance. Our results will contribute to further study of the biological function of CaGATA genes in pepper, especially under abiotic stress.
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868. https://doi.org/10.1105/tpc.9.10.1859
Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, Zheng X, Zhu JK (2006) A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem 281:37636–37645. https://doi.org/10.1074/jbc.M605895200
An Y, Zhou Y, Han X et al (2020) The GATA transcription factor GNC plays an important role in photosynthesis and growth in poplar. J Exp Bot 71:1969–1984. https://doi.org/10.1093/jxb/erz564
Aravind L, Iyer LM (2012) The HARE-HTH and associated domains: novel modules in the coordination of epigenetic DNA and protein modifications. Cell Cycle 11:119–131. https://doi.org/10.4161/cc.11.1.18475
Armstrong GA, Runge S, Frick G, Sperling U, Apel K (1995) Identification of NADPH:protochlorophyllide oxidoreductases A and B: a branched pathway for light-dependent chlorophyll biosynthesis in Arabidopsis thaliana. Plant Physiol 108:1505–1517. https://doi.org/10.1104/pp.108.4.1505
Bai Y, Meng Y, Huang D, Qi Y, Chen M (2011) Origin and evolutionary analysis of the plant-specific TIFY transcription factor family. Genomics 98:128–136. https://doi.org/10.1016/j.ygeno.2011.05.002
Baskind HA, Na L, Ma Q, Patel MP, Geenen DL, Wang QT (2009) Functional conservation of Asxl2, a murine homolog for the drosophila enhancer of trithorax and polycomb group gene Asx. PLoS One 4:e4750. https://doi.org/10.1371/journal.pone.0004750
Bastakis E, Hedtke B, Klermund C, Grimm B, Schwechheimer C (2018) LLM-domain B-GATA transcription factors play multifaceted roles in controlling greening in Arabidopsis. Plant Cell 30:582–599. https://doi.org/10.1105/tpc.17.00947
Behringer C, Schwechheimer C (2015) B-GATA transcription factors-insights into their structure, regulation, and role in plant development. Front Plant Sci 6:90. https://doi.org/10.3389/fpls.2015.00090
Behringer C, Bastakis E, Ranftl QL, Mayer KF, Schwechheimer C (2014) Functional diversification within the family of B-GATA transcription factors through the leucine-leucine-methionine domain. Plant Physiol 166:293–305. https://doi.org/10.1104/pp.114.246660
Bi YM, Zhang Y, Signorelli T, Zhao R, Zhu T, Rothstein S (2005) Genetic analysis of Arabidopsis GATA transcription factor gene family reveals a nitrate-inducible member important for chlorophyll synthesis and glucose sensitivity. Plant J 44:680–692. https://doi.org/10.1111/j.1365-313X.2005.02568.x
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10. https://doi.org/10.1186/1471-2229-4-10
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools-an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chiang YH, Zubo YO et al (2012) Functional characterization of the GATA transcription factors GNC and CGA1 reveals their key role in chloroplast development, growth, and division in Arabidopsis. Plant Physiol 160:332–348. https://doi.org/10.1104/pp.112.198705
Chini A, Fonseca S, Chico JM, Fernandez-Calvo P, Solano R (2009) The ZIM domain mediates homo- and heteromeric interactions between Arabidopsis JAZ proteins. Plant J 59:77–87. https://doi.org/10.1111/j.1365-313X.2009.03852.x
Ding L et al (2015a) HANABA TARANU regulates the shoot apical meristem and leaf development in cucumber (Cucumis sativus L.). J Exp Bot 66:7075–7087. https://doi.org/10.1093/jxb/erv409
Ding L et al (2015b) HANABA TARANU (HAN) Bridges meristem and organ primordia boundaries through PINHEAD, JAGGED, BLADE-ON-PETIOLE2 and CYTOKININ OXIDASE 3 during flower development in Arabidopsis. PLoS Genet 11:e1005479. https://doi.org/10.1371/journal.pgen.1005479
Fisher CL et al (2010) Additional sex combs-like 1 belongs to the enhancer of trithorax and polycomb group and genetically interacts with Cbx2 in mice. Dev Biol 337:9–15. https://doi.org/10.1016/j.ydbio.2009.10.004
Franco-Zorrilla JM, Lopez-Vidriero I, Carrasco JL, Godoy M, Vera P, Solano R (2014) DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc Natl Acad Sci U S A 111:2367–2372. https://doi.org/10.1073/pnas.1316278111
Geng J, Liu JH (2018) The transcription factor CsbHLH18 of sweet orange functions in modulation of cold tolerance and homeostasis of reactive oxygen species by regulating the antioxidant gene. J Exp Bot 69:2677–2692. https://doi.org/10.1093/jxb/ery065
Gil KE et al (2017) Alternative splicing provides a proactive mechanism for the diurnal CONSTANS dynamics in Arabidopsis photoperiodic flowering. Plant J 89:128–140. https://doi.org/10.1111/tpj.13351
Gillis WQ, St John J, Bowerman B, Schneider SQ (2009) Whole genome duplications and expansion of the vertebrate GATA transcription factor gene family. BMC Evol Biol 9:207. https://doi.org/10.1186/1471-2148-9-207
He X et al (2020) Genome-wide identification and functional analysis of the TIFY gene family in the response to multiple stresses in Brassica napus L. BMC Genomics 21:736. https://doi.org/10.1186/s12864-020-07128-2
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:W585–W587. https://doi.org/10.1093/nar/gkm259
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Hudson D et al (2011) GNC and CGA1 Modulate chlorophyll biosynthesis and glutamate synthase (GLU1/Fd-GOGAT) expression in Arabidopsis. PLoS One 6:e26765. https://doi.org/10.1371/journal.pone.0026765
Iwasaki T, Yamaguchi-Shinozaki K, Shinozaki K (1995) Identification of a cis-regulatory region of a gene in Arabidopsis thaliana whose induction by dehydration is mediated by abscisic acid and requires protein synthesis. Mol Gen Genet 247:391–398. https://doi.org/10.1007/BF00293139
Jin J et al (2015) An Arabidopsis transcriptional regulatory map reveals distinct functional and evolutionary features of novel transcription factors. Mol Biol Evol 32:1767–1773. https://doi.org/10.1093/molbev/msv058
Kiryushkin AS, Ilina EL, Puchkova VA, Guseva ED, Pawlowski K, Demchenko KN (2019) Lateral root initiation in the parental root meristem of cucurbits: old players in a new position. Front Plant Sci 10:365. https://doi.org/10.3389/fpls.2019.00365
Klermund C, Ranftl QL, Diener J, Bastakis E, Richter R, Schwechheimer C (2016) LLM-domain B-GATA transcription factors promote stomatal development downstream of light signaling pathways in Arabidopsis thaliana hypocotyls. Plant Cell 28:646–660. https://doi.org/10.1105/tpc.15.00783
Kobayashi K, Masuda T (2016) Transcriptional regulation of tetrapyrrole biosynthesis in Arabidopsis thaliana. Front Plant Sci 7:1811. https://doi.org/10.3389/fpls.2016.01811
Kobayashi K et al (2017) Shoot removal induces chloroplast development in roots via cytokinin signaling. Plant Physiol 173:2340–2355. https://doi.org/10.1104/pp.16.01368
Lescot M et al (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Li M et al (2019a) Leaf variegation and impaired chloroplast development caused by a truncated CCT domain gene in albostrians barley. Plant Cell 31:1430–1445. https://doi.org/10.1105/tpc.19.00132
Li X, Guo C, Ahmad S, Wang Q, Yu J, Liu C, Guo Y (2019b) Systematic analysis of MYB family genes in potato and their multiple roles in development and stress responses. Biomolecules 9. https://doi.org/10.3390/biom9080317
Liu PP, Koizuka N, Martin RC, Nonogaki H (2005) The BME3 (Blue Micropylar End 3) GATA zinc finger transcription factor is a positive regulator of Arabidopsis seed germination. Plant J 44:960–971. https://doi.org/10.1111/j.1365-313X.2005.02588.x
Liu J, Shen J, Xu Y, Li X, Xiao J, Xiong L (2016) Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J Exp Bot 67:5785–5798. https://doi.org/10.1093/jxb/erw344
Liu Y et al (2019) A WRKY transcription factor PbrWRKY53 from Pyrus betulaefolia is involved in drought tolerance and AsA accumulation. Plant Biotechnol J 17:1770–1787. https://doi.org/10.1111/pbi.13099
Liu X, Zhao C, Yang L, Zhang Y, Wang Y, Fang Z, Lv H (2020) Genome-wide identification, expression profile of the TIFY gene family in Brassica oleracea var. capitata, and their divergent response to various pathogen infections and phytohormone treatments. Genes (Basel) 11. https://doi.org/10.3390/genes11020127
Lowry JA, Atchley WR (2000) Molecular evolution of the GATA family of transcription factors: conservation within the DNA-binding domain. J Mol Evol 50:103–115. https://doi.org/10.1007/s002399910012
Luo XM et al (2010) Integration of light- and brassinosteroid-signaling pathways by a GATA transcription factor in Arabidopsis. Dev Cell 19:872–883. https://doi.org/10.1016/j.devcel.2010.10.023
Manfield IW, Devlin PF, Jen CH, Westhead DR, Gilmartin PM (2007) Conservation, convergence, and divergence of light-responsive, circadian-regulated, and tissue-specific expression patterns during evolution of the arabidopsis GATA gene family. Plant Physiol 143:941–958. https://doi.org/10.1104/pp.106.090761
Morita R, Sugino M, Hatanaka T, Misoo S, Fukayama H (2015) CO2-responsive CONSTANS, CONSTANS-like, and time of chlorophyll a/b binding protein expression1 protein is a positive regulator of starch synthesis in vegetative organs of rice. Plant Physiol 167:1321–1331. https://doi.org/10.1104/pp.15.00021
Neuhaus HE, Emes MJ (2000) Nonphotosynthetic metabolism in plastids. Annu Rev Plant Physiol Plant Mol Biol 51:111–140. https://doi.org/10.1146/annurev.arplant.51.1.111
Onishi M, Tachi H, Kojima T, Shiraiwa M, Takahara H (2006) Molecular cloning and characterization of a novel salt-inducible gene encoding an acidic isoform of PR-5 protein in soybean (Glycine max [L.] Merr.). Plant Physiol Biochem 44:574–580. https://doi.org/10.1016/j.plaphy.2006.09.009
Patient RK, McGhee JD (2002) The GATA family (vertebrates and invertebrates). Curr Opin Genet Dev 12:416–422. https://doi.org/10.1016/s0959-437x(02)00319-2
Reyes JC, Muro-Pastor MI, Florencio FJ (2004) The GATA family of transcription factors in arabidopsis and rice. Plant Physiol 134:1718–1732. https://doi.org/10.1104/pp.103.037788
Richter R, Behringer C, Muller IK, Schwechheimer C (2010) The GATA-type transcription factors GNC and GNL/CGA1 repress gibberellin signaling downstream from DELLA proteins and PHYTOCHROME-INTERACTING FACTORS. Genes Dev 24:2093–2104. https://doi.org/10.1101/gad.594910
Richter R, Bastakis E, Schwechheimer C (2013a) Cross-repressive interactions between SOC1 and the GATAs GNC and GNL/CGA1 in the control of greening, cold tolerance, and flowering time in Arabidopsis. Plant Physiol 162:1992–2004. https://doi.org/10.1104/pp.113.219238
Richter R, Behringer C, Zourelidou M, Schwechheimer C (2013b) Convergence of auxin and gibberellin signaling on the regulation of the GATA transcription factors GNC and GNL in Arabidopsis thaliana. Proc Natl Acad Sci U S A 110:13192–13197. https://doi.org/10.1073/pnas.1304250110
Riechmann JL et al (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110. https://doi.org/10.1126/science.290.5499.2105
Savojardo C, Martelli PL, Fariselli P, Profiti G, Casadio R (2018) BUSCA: an integrative web server to predict subcellular localization of proteins. Nucleic Acids Res 46:W459–W466. https://doi.org/10.1093/nar/gky320
Scazzocchio C (2000) The fungal GATA factors. Curr Opin Microbiol 3:126–131. https://doi.org/10.1016/s1369-5274(00)00063-1
Scheible WR et al (2004) Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol 136:2483–2499. https://doi.org/10.1104/pp.104.047019
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108. https://doi.org/10.1038/nprot.2008.73
Seo DH, Seomun S, Choi YD, Jang G (2020) Root development and stress tolerance in rice: the key to improving stress tolerance without yield penalties. Int J Mol Sci 21. https://doi.org/10.3390/ijms21051807
Shaikhali J et al (2012) The CRYPTOCHROME1-dependent response to excess light is mediated through the transcriptional activators ZINC FINGER PROTEIN EXPRESSED IN INFLORESCENCE MERISTEM LIKE1 and ZML2 in Arabidopsis. Plant Cell 24:3009–3025. https://doi.org/10.1105/tpc.112.100099
Song X, Wang Y, Wang P, Pu G, Zou X (2020) GATA-type transcriptional factor Gat1 regulates nitrogen uptake and polymalic acid biosynthesis in polyextremotolerant fungus Aureobasidium pullulans. Environ Microbiol 22:229–242. https://doi.org/10.1111/1462-2920.14841
Suarez-Lopez P, Wheatley K, Robson F, Onouchi H, Valverde F, Coupland G (2001) CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410:1116–1120. https://doi.org/10.1038/35074138
Sun H et al (2017) The JASMONATE ZIM-domain gene family mediates JA signaling and stress response in cotton. Plant Cell Physiol 58:2139–2154. https://doi.org/10.1093/pcp/pcx148
Teakle GR, Manfield IW, Graham JF, Gilmartin PM (2002) Arabidopsis thaliana GATA factors: organisation, expression and DNA-binding characteristics. Plant Mol Biol 50:43–57. https://doi.org/10.1023/a:1016062325584
Urao T, Yamaguchi-Shinozaki K, Urao S, Shinozaki K (1993) An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5:1529–1539. https://doi.org/10.1105/tpc.5.11.1529
Vanholme B, Grunewald W, Bateman A, Kohchi T, Gheysen G (2007) The tify family previously known as ZIM. Trends Plant Sci 12:239–244. https://doi.org/10.1016/j.tplants.2007.04.004
Wan H et al (2011) Identification of reference genes for reverse transcription quantitative real-time PCR normalization in pepper (Capsicum annuum L.). Biochem Biophys Res Commun 416:24–30. https://doi.org/10.1016/j.bbrc.2011.10.105
Wang W, Vinocur B, Altman A (2003) Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 218:1–14. https://doi.org/10.1007/s00425-003-1105-5
Wang Y, Pan F, Chen D, Chu W, Liu H, Xiang Y (2017) Genome-wide identification and analysis of the Populus trichocarpa TIFY gene family. Plant Physiol Biochem 115:360–371. https://doi.org/10.1016/j.plaphy.2017.04.015
Wray GA, Hahn MW, Abouheif E, Balhoff JP, Pizer M, Rockman MV, Romano LA (2003) The evolution of transcriptional regulation in eukaryotes. Mol Biol Evol 20:1377–1419. https://doi.org/10.1093/molbev/msg140
Wu HM, Xie DJ, Tang ZS, Shi DQ, Yang WC (2020) PINOID regulates floral organ development by modulating auxin transport and interacts with MADS16 in rice. Plant Biotechnol J 18:1778–1795. https://doi.org/10.1111/pbi.13340
Xia W, Yu H, Cao P, Luo J, Wang N (2017) Identification of TIFY family genes and analysis of their expression profiles in response to phytohormone treatments and Melampsora larici-populina Infection in Poplar. Front Plant Sci 8:493. https://doi.org/10.3389/fpls.2017.00493
Xin C, Yang J, Mao Y, Chen W, Wang Z, Song Z (2020) GATA-type transcription factor MrNsdD regulates dimorphic transition, conidiation, virulence and microsclerotium formation in the entomopathogenic fungus Metarhizium rileyi. Microb Biotechnol. https://doi.org/10.1111/1751-7915.13581
Xu L, Yang L, Huang H (2007) Transcriptional, post-transcriptional and post-translational regulations of gene expression during leaf polarity formation. Cell Res 17:512–519. https://doi.org/10.1038/cr.2007.45
Yang X, Tuskan GA, Cheng MZ (2006) Divergence of the Dof gene families in poplar, Arabidopsis, and rice suggests multiple modes of gene evolution after duplication. Plant Physiol 142:820–830. https://doi.org/10.1104/pp.106.083642
Yang S, Zhang X, Yue JX, Tian D, Chen JQ (2008) Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol Gen Genomics 280:187–198. https://doi.org/10.1007/s00438-008-0355-0
Yang M et al (2018) Genome-wide association studies reveal the genetic basis of ionomic variation in rice. Plant Cell 30:2720–2740. https://doi.org/10.1105/tpc.18.00375
Yang L et al (2020) Methylation of a CGATA element inhibits binding and regulation by GATA-1. Nat Commun 11:2560. https://doi.org/10.1038/s41467-020-16388-1
Ye H, Du H, Tang N, Li X, Xiong L (2009) Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol Biol 71:291–305. https://doi.org/10.1007/s11103-009-9524-8
Yu CS, Cheng CW, Su WC, Chang KC, Huang SW, Hwang JK, Lu CH (2014) CELLO2GO: a Web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS One 9:e99368. https://doi.org/10.1371/journal.pone.0099368
Zhang X, Zhou Y, Ding L, Wu Z, Liu R, Meyerowitz EM (2013) Transcription repressor HANABA TARANU controls flower development by integrating the actions of multiple hormones, floral organ specification genes, and GATA3 family genes in Arabidopsis. Plant Cell 25:83–101. https://doi.org/10.1105/tpc.112.107854
Zhang J et al (2015) Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice. New Phytol 208:1056–1066. https://doi.org/10.1111/nph.13538
Zhang Y et al (2016a) An A20/AN1-type zinc finger protein modulates gibberellins and abscisic acid contents and increases sensitivity to abiotic stress in rice (Oryza sativa). J Exp Bot 67:315–326. https://doi.org/10.1093/jxb/erv464
Zhang Y et al (2016b) MYC cis-elements in PsMPT promoter is involved in chilling response of Paeonia suffruticosa. PLoS One 11:e0155780. https://doi.org/10.1371/journal.pone.0155780
Zhang Z et al (2018) The genetic architecture of nodal root number in maize. Plant J 93:1032–1044. https://doi.org/10.1111/tpj.13828
Zhang L, Song Z, Li F, Li X, Ji H, Yang S (2019) The specific MYB binding sites bound by TaMYB in the GAPCp2/3 promoters are involved in the drought stress response in wheat. BMC Plant Biol 19:366. https://doi.org/10.1186/s12870-019-1948-y
Zhang C et al (2020a) A GATA transcription factor from soybean (Glycine max) regulates chlorophyll biosynthesis and suppresses growth in the transgenic Arabidopsis thaliana. Plants (Basel) 9. https://doi.org/10.3390/plants9081036
Zhang X et al (2020b) Genome-wide identification of the Tify gene family and their expression profiles in response to biotic and abiotic stresses in tea plants (Camellia sinensis). Int J Mol Sci 21. https://doi.org/10.3390/ijms21218316
Zhao Y, Medrano L, Ohashi K, Fletcher JC, Yu H, Sakai H, Meyerowitz EM (2004) HANABA TARANU is a GATA transcription factor that regulates shoot apical meristem and flower development in Arabidopsis. Plant Cell 16:2586–2600. https://doi.org/10.1105/tpc.104.024869
Zhao G et al (2016) Genome-wide identification and functional analysis of the TIFY gene family in response to drought in cotton. Mol Gen Genomics 291:2173–2187. https://doi.org/10.1007/s00438-016-1248-2
Zhu XG, Long SP, Ort DR (2010) Improving photosynthetic efficiency for greater yield. Annu Rev Plant Biol 61:235–261. https://doi.org/10.1146/annurev-arplant-042809-112206
Zhu D, Bai X, Luo X, Chen Q, Cai H, Ji W, Zhu Y (2013) Identification of wild soybean (Glycine soja) TIFY family genes and their expression profiling analysis under bicarbonate stress. Plant Cell Rep 32:263–272. https://doi.org/10.1007/s00299-012-1360-7
Acknowledgements
Great gratitude goes to linguistics professor Ping Liu from Foreign Language College, Huazhong Agriculture University, Wuhan, China, for her work on English editing and language polishing.
Funding
This research was funded by National Key R&D Program of China, grant numbers 2016YFE0205500 and 2017YFD0101903; Natural Science Foundation of Hubei Province, grant number 2019CFB577; and China Agriculture Research System, grant number CARS-23-G28.
Author information
Authors and Affiliations
Contributions
Conceptualization, C.Y. and M.Y.; methodology, C.Y. and N.L.; software, C.Y., N.L. and S.G.; formal analysis, C.Y. and S.G.; writing—original draft preparation, C.Y. and N.L.; investigation, F.W.; resources, C.J. and Y.Y.; data curation, C.Y. and N.L.; writing—review and editing, C.Y., N.L., M.Y., and Y.Y.; project administration, C.J.; funding acquisition, M.Y. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by: Izabela Pawłowicz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Fig. S1:
The conservative domain analysis of CaGATA genes (PNG 271 kb)
Fig. S2:
Paralogous relationship analysis of CaGATA genes (PNG 416 kb)
Fig. S3:
The number of various cis-elements on the promoters of each CaGATA gene. Promoter sequences (-2000 bp) of 28 CaGATA genes were analyzed (PNG 387 kb)
Fig. S4:
Relative expression levels of CaGATA genes significantly induced by (a) MV, (b) Salt, (c) HT (Heat), (d) Cold, (e) Dr (Drought) (PNG 493 kb)
Fig. S5:
Relative expression levels of CaGATA genes significantly induced by (a) ABA, (b) GA, (c) ET, (d) SA, (e) IAA, (f) JA (PNG 441 kb)
ESM 1
(DOCX 30 kb)
ESM 2
(XLSX 77 kb)
Rights and permissions
About this article
Cite this article
Yu, C., Li, N., Yin, Y. et al. Genome-wide identification and function characterization of GATA transcription factors during development and in response to abiotic stresses and hormone treatments in pepper. J Appl Genetics 62, 265–280 (2021). https://doi.org/10.1007/s13353-021-00618-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-021-00618-3