Abstract
Hepatocyte cell adhesion molecule (HEPACAM), a member of immunoglobulin superfamily, is an adhesion molecule. Although dysregulation of several adhesion molecules has been implicated in the progression of non-small cell lung cancer (NSCLC), the expression profile and functions of HEPACAM in NSCLC remains unknown. In this study, it was found that the expression of HEPACAM was downregulated in NSCLC tissues. Forced expression of HEPACAM in NSCLC cells inhibited the growth and migration of the cancer cells, while knocking down the expression of HEPACAM promoted cell growth, migration, and metastasis. In the molecular mechanism study, HEPACAM was found to be a negative regulator of beta-catenin/TCF signaling. Taken together, this study revealed the suppressive roles of HEPACAM in NSCLC and restoring the function of HEPACAM in NSCLC might be a promising strategy for the therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung cancer (NSCLC) is one of the most common malignancies in the world [1]. Migration and dissemination of the tumor cells is the main cause of NSCLC-related death [2]. Many cell adhesion molecules act as tumor suppressors for the ability to restrict cell migration as well as growth due to contact inhibition [3]. Understanding the functions of these cell adhesion molecules would facilitate the development of new therapeutic strategies.
Several cell adhesion molecules have shown suppressive roles in the progression of NSCLC, such as E-cadherin [3], cell adhesion molecule 1 (CADM1), and carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1) [4, 5]. HEPACAM1, which has been identified as a cell adhesion molecule recently, belongs to the immunoglobulin superfamily [6]. The functions of hepatocyte cell adhesion molecule (HEPACAM) in the tumorigenesis have attracted much attention. Mutations in HEPACAM gene and hypermethylation of the HEPACAM promoter region have been reported [7, 8]. Downregulation of HEPACAM was found in a variety of cancer, including hepatocellular carcinoma (HCC), breast cancer, and so on [9, 10]. Forced expression of HEPACAM in HCC cells inhibited the proliferation and colony formation of HepG2 cells [10]. Similar phenotype was also observed in breast cancer cell line MCF7 [9]. In addition, re-expression of HEPACAM in MCF7 cells induced cell cycle arrest and senescence by upregulating the expression of P21, P27, and P53 [9]. Also, HEPACAM inhibited the proliferation of renal cancer cells by blocking the translocation of protein kinase C (PKC) [11]. On the other hand, HEPACAM showed suppressive roles in the angiogenesis and promoted the differentiation of glioblastoma cells [12, 13]. The reports for the effects of HEPACAM on the cell motility were controversial [14]. It has been found that upregulation of HEPACAM in MCF7, and HepG2 cells promoted cell migration [15, 16]. However, overexpression of HEPACAM inhibited the migration of the prostate cancer cells through inhibiting androgen receptor nuclear translocation and ERK signaling [17]. Theses controversial studies suggested that the effects of HEPACAM on the migration of cancer cells were dependent on the cancer types.
Beta-catenin/TCF signaling has been reported to play an important role in the progression of lung cancer [18, 19]. In the absence of wnt ligand, beta-catenin was phosphorylated by glycogen synthase kinase 3 (GSK3) beta and degradated. Inactivation of GSK3 beta (shown by upregulation of ser9 phosphorylation) led to the accumulation of beta-catenin in cytoplasm and activation of beta-catenin/TCF signaling [20]. Although regulation of beta-catenin/TCF signaling by several adhesion molecules has been reported, whether HEPACAM would regulate beta-catenin/TCF signaling remains unknown. This study aimed to examine the expression pattern and biological functions of HEPACAM in non-small cell lung cancer and investigate the underlying molecular mechanisms.
Materials and methods
Cell lines and culture conditions
The human NSCLC cell lines (H520 and H460) and HEK293T cells were purchased from the American Type Culture Collection (ATCC). Cells were cultured in Dulbecco’s modified Eagle’s medium (Invitrogen) supplemented with 10 % fetal bovine serum (FBS). All cultures were maintained in a humidified atmosphere containing 5 % CO2 at 37 °C.
Patients and specimens
A total of 52 NSCLC tissues and paired noncancerous tissues were obtained from patients who received surgery for NSCLC at Wujin Hospital, Jiangsu University. The diagnosis of NSCLC was histologically confirmed by two pathologists and none of the patients received preoperative therapy. All of the patients agreed with this study. Tissues and paired noncancerous tissues were stored at −80 °C.
Plasmid construction and transfection
The coding sequence of HEPACAM was amplified by PCR and inserted into the expression vector pcDNA 3.1-myc. The HEPACAM expression vector and empty pcDNA3.1 were transfected into H520 and H460 cells using Lipofectamine 2000 reagent (Invitrogen) following the manufacturer’s instructions. After the selection with G418, the resistant cells were pooled and further confirmed the expression of exogenous HEPACAM using anti-myc antibody.
Real-time PCR analysis
Total RNA was extracted from NSCLC tissues and matched noncancerous tissues using TRIzol reagent (Invitrogen) according to the manufacture’s instruction. The isolated RNA was used for cDNA synthesis using the reverse transcription kit (Promega, Madison, WI) following the manufacturer’s instructions. Real-time PCR was performed using a Stratagene MAXP3000 PCR system and Brilliant Q-PCR Master Mixture. The primer pair used for amplification of the human HEPACAM gene was as follows: forward primer, 5′-TGTAGTGAAGTGGCAGCTGA-3′ and reverse primer, 5′-TCTCATGTGAGCAGTTCAAG-3′. As an internal standard, a fragment of human beta-actin was amplified by PCR using the following primers: forward primer, 5′-GATCATTGCTCCTCCTGAGC-3′ and reverse primer, 5′-ACTCCT GCTTGCTGATCCAC-3′.
Western blotting
Cells were lysed using an immunoprecipitation assay buffer, separated by 10 % SDS-PAGE, transferred to polyvinylidene difluoride membranes (Millipore, Bedford, MA) and probed with specific antibodies. The immunoreactive protein bands were visualized by an ECL kit (Pierce). Antibody to HEPACAM was purchased from Abcam, antibodies to beta-catenin, Cyclin D1, c-Myc and c-Jun were purchased from Cell Signaling Technology, and antibody to GAPDH was purchased from Santa Cruz Biotechnology.
RNAi of HEPACAM in NSCLC cells
RNA interference (RNAi) lentivirus particles (si con and si HEPACAM) were purchased from GeneChem (China). Cells were incubated with the lentivirus particles for 24 h and then selected with the medium containing puromycine.
Immunohistochemistry
Clinical NSCLC tissues and paired noncancerous tissues were fixed in formalin, embedded in paraffin, and cut in 5 μm-thick consecutive sections. After deparaffin and antigen recovery (in sodium citrate solution, pH 6.0, 20 min, 98 °C), the sections were washed thrice in 0.01 mol/l phosphate-buffered saline (PBS) (8 mmol/l Na2HPO4, 2 mmol/l NaH2PO4, and 150 mmol/l NaCl) for 5 min each, blocked for 1 h in 0.01 mol/l PBS containing 0.3 % Triton X-100 and 5 % BSA, followed by addition of anti-HEPACAM (1:100) antibody at 4 °C overnight. After brief washes with 0.01 mol/l PBS, sections were incubated with 0.01 mol/l PBS containing horseradish peroxidase-conjugated rabbit anti-goat IgG (1:500) for 2 h, followed by development with 0.003 % H2O2 and 0.03 % 3,30-diaminobenzidine in 0.05 mol/l Tris–HCl (pH 7.6). Immunohistochemistry for each sample was repeated thrice.
Cell growth assay
Crystal violet assay was performed to examine the effects of HEPACAM on the growth of NSCLC cells. Equal number of control cells and experimental cells were seeded in 12-well plates and cultured in medium supplemented with 10 % FBS at a density of 1000 cells/well. Medium was changed every other day. After 10 days of culture under the standard condition, the medium was removed and the cells were stained with 0.5 % crystal violet solution in 20 % methanol. After staining for 10 min, the fixed cells were washed with PBS and photographed. The colonies were solved using 1 % SDS solution. OD 600 nm was measured.
Cell migration assay
Cell migration assay was performed using a modified Boyden chamber. Cells (2 × 105) suspended in 0.1 ml medium containing 1 % FBS were placed in the upper chamber, and the lower chamber was loaded with 0.152 ml medium containing 10 % FBS. 6 h later, cells that migrated to the lower surface of filters were detected with traditional H&E staining. The migrated cells were counted under the inverted microscope. The experiments were repeated thrice.
In vivo metastasis assay
H520 cells overexpressing luciferase were established by stable transfection and selection with G418. The activity of luciferase was determined by luciferin (Xenogen) and an in vivo imaging system (Xenogen). The luciferase labeled H520/si con cells and H520/si HEPACAM cells (1 × 106 cells in 200 μl PBS) were injected into the left ventricle of the nude mice (six mice for each group). The metastatic lesions were monitored. Before mice were anesthetized with Forane (Abbott), luciferin (150 mg/kg intraperitoneally) was injected into the nude mice 10 min before imaging. The nude mice were placed into a light-tight chamber of the CCD camera system (Xenogen), and the photons emitted from the luciferase expressing cells within the animal were quantified for 1 min using the software program Living Image (Xenogen) as an overlay on Igor (Wavemetrics). This study was carried out according to the recommendations of the National Institutes of Health.
Results
Downregulation of HEPACAM in NSCLC samples
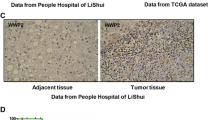
The matched normal and tumor tissues were used to examine the expression profile of HEPACAM transcripts in NSCLC tissues. The results showed that HEPACAM was expressed in almost all of the adjacent normal tissues. However, the mRNA level of HEPACAM was significantly downregulated in NSCLC tumor tissues (Fig. 1a). Moreover, the protein level of HEPACAM was decreased in tumor tissues shown by the immunohistochemistry staining and Western blot analysis (Fig. 1b,c). These observations suggested that the expression of HEPACAM was downregulated in NSCLC tissues.
The expression of HEPACAM was decreased in NSCLC tissues. a HEPACAM mRNA level in NSCLC samples and adjacent normal tissues. HEPACAM mRNA levels in 52 NSCLC samples and 52 paired adjacent normal tissues were examined using real-time PCR. The HEPACAM expression was normalized to that of beta-actin (left panel). Data was calculated from triplicates. In the right panel, the expression of HEPACAM was expressed as Log2([C]/[N]). C cancer tissues, N normal tissues. b The protein level of HEPACAM in NSCLC samples and paired adjacent normal tissues was examined by immunohistochemistry staining. c Western blotting analysis to examine the protein level of HEPACAM in NSCLC samples and paired adjacent normal tissues
HEPACAM inhibited the growth and migration of NSCLC cells
To further study the roles of HEPACAM in NSCLC, we stably transfected H520 and H460 cells with HEPACAM expression vector (myc-HEPACAM). Cells transfected with pcDNA3.1 plasmid served as control. After selection with the G418 for 2 weeks, cells were pooled and examined the expression of exogenously expressed HEPACAM (Fig. 2a). Cell migration assay using a Boyden chamber was performed to evaluate the function of HEPACAM in cell motility. Compared with the control cells, overexpression of HEPACAM in H520 and H460 cells inhibited cell migration remarkably (Fig. 2b). Also, colony formation assay was used to examine the effects of HEPACAM on the growth of NSCLC cells. Expression of HEPACAM inhibited the colony formation of H520 and H460 cells (Fig. 2c). In the next study, we knocked down the expression of HEPACAM in H520 and H460 cells using two independent targeting sequences (Fig. 3a). We investigated the effects of knocking down HEPACAM on the growth and migration of NSCLC cells. As shown in Fig. 3b, c, downregulation of HEPACAM promoted cell growth as well as cell migration of H520 and H460 cells (Fig. 3b, c). These data suggested that HEPACAM inhibited the growth and migration of NSCLC cells.
Forced expression of HEPACAM inhibited the growth and migration of H520 and H460 cells. a Forced expression of HEPACAM in H520 and H460 cells. H520 and H460 cells were transfected with pcDNA3.1 vector or the HEPACAM expression vector, and selected with G418. G418-resistant cells were pooled and confirmed the overexpression of HEPACAM by Western blot analysis. b Forced expression of HEPACAM impaired the migration of H520 and H460 cells in the migration assay using a Boyden chamber. c Colony formation assay to analyze the effects of HEPACAM on the growth of H520 and H460 cells by crystal violet assay. *P < 0.05; **P < 0.01
Knocking down the expression of HEPACAM promoted the growth and migration of H520 and H460 cells. a Downregulation of HEPACAM in H520 and H460 cells by RNA interference (RNAi). b Colony formation assay to examine the effects of knocking down HEPACAM on the growth of H520 and H460 cells by crystal violet staining. c Knocking down the expression of HEPACAM promoted the migration of H520 and H460 cells. *P < 0.05; **P < 0.01
HEPACAM negatively regulated beta-catenin/TCF signaling
To study the underlying molecule mechanism, we screened the effects of HEPACAM on various signaling pathways using reporter gene assay. Overexpression of HEPACAM was found to inhibit the activation of Topflash (reporter gene for beta-catenin/TCF signaling) induced by wnt3a (Fig. 4a). Moreover, knocking down the expression of HEPACAM promoted the inactivation of GSK3 beta (shown by the increase in the phosphorylation of Ser9 in GSK3 beta) (Fig. 4b). These observations suggested that knocking down the expression of HEPACAM activated beta-catenin/TCF signaling. In addition, downregulation of HEPACAM in H520 cells promoted the expression of several beta-catenin/TCF target genes, such as c-Jun, CyclinD1, and c-Myc (Fig. 4c). As shown in Fig. 4d, dominant negative beta-catenin (DN beta-catenin) abolished the promoting effects on cell growth induced by knocking down HEPACAM, suggesting HEPACAM regulated cell growth through modifying beta-catenin/TCF signaling (Fig. 4d). Taken together, HEPACAM negatively regulated beta-catenin/TCF signaling in NSCLC cells.
HEPACAM inhibited beta-catenin/TCF signaling. a Overexpression of HEPACAM inhibited the activity of Topflash reporter induced by wnt3a. b Knocking down the expression of HEPACAM promoted the phosphorylation of GSK 3beta. c HEPACAM inhibited the expression of target genes downstream beta-catenin/TCF signaling. d The inhibition of beta-catenin/TCF signaling by dominant negative beta-catenin (DN beta-catenin) abolished the growth advantage of H520 and H460 cells induced by downregulation of HEPACAM. *P < 0.05; **P < 0.01
Downregulating the expression of HEPACAM promoted the metastasis of H520 cells in vivo
To validate the effects of HEPACAM on the metastasis of NSCLC cells in vivo, H520 cells were labeled with luciferase gene and then knocked down the expression of HEPACAM. Cells were injected into the tail vein of nude mice, and the metastasis of these cells was monitored by an in vivo image system after the administration of luciferin (the substrate of luciferase). Knocking down the expression of HEPACAM resulted in an increase in metastasis signals (Fig. 5a, b), suggesting that downregulation of HEPACAM promoted the metastasis of NSCLC cells.
Knocking down the expression of HEPACAM promoted the metastasis of H520 cells in vivo. a Monitoring metastasis of H520/si con and H520/si HEPACAM cells on day 42. Images were obtained at the indicated time point after injection, respectively. b Mean photon counts of each group of mice were quantified and were displayed on day 42. Each bar represented the mean ± SD. *P < 0.05; **P < 0.01
Discussion
Up to date, five families of adhesion molecules have been found including immunoglobulin superfamily, caherin, selectin, integrins, and CD44. These adhesion molecules allow cells to communicate with the microenvironment. Several adhesion molecules have been reported as tumor suppressor. Therefore, downregulation of the adhesion molecules is a strategy which is taken by the tumor cells to promote metatasis. In this study, HEPACAM, a member of immunoglobulin family, was found to be downregulated in lung cancer and inhibits the growth and migration of NSCLC cells, which further emphasized the tumor suppressive roles of adhesion molecules.
Downregulation of HEPACAM was found in several cancer types, including HCC, bladder cancer, and breast cancer [8–10]. Hypermethylation of the HEPACAM promoter region was found in these malignancies [8]. Therefore, further analyzing the methylation status of HEPACAM promoter in the NSCLC samples will provide novel insight into its regulation. Although the inhibitory effects of HEPACAM on the cell growth were well recognized, the functions of HEPACAM in cell motility remained controversial. Re-expression of HEPACAM in HCC and breast cancer cells promoted the migration of cancer cells [15, 16], while HEPACAM inhibited the migration of prostate cancer cells in vivo [17]. Similarly, inhibition of cell migration by HEPACAM was observed in this study. These observations suggested that the functions of HEPACAM in cell migration were dependent on the context of the malignancies.
In this study, HEPACAM was found to negatively regulate the beta-catenin/TCF signaling. Several adhesion molecules were found to regulate beta-catenin/TCF signaling. E-cadherin, a member of the cadherin family, inhibited beta-catenin/TCF signaling by restricting beta-catenin on cell membrane [21]. However, integrin activated beta-catenin/TCF signaling through inactivating GSK3 beta [22]. In this study, knocking down the expression of HEPACAM inactivated GSK3beta (shown by upregulation of phosphorylated GSK3 beta) and increased the expression of several beta-catenin/TCF target genes. These observations suggested the negative regulation of beta-catenin/TCF signaling by HEPACAM. HEPACAM was reported to inhibit PKC signaling, androgen receptor signaling, and ERK signaling [17]. Therefore, restoration of HEPACAM in cancer cells might be a novel strategy for lung cancer therapy.
Although our study was indicative, further study is needed to find out the detailed mechanism for HEPACAM to regulate beta-catenin/TCF signaling. Also, investigations using HEPACAM knockout mice would further elucidate its functions in lung cancer.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65:5–29.
Lu S, Azada MC, Ou SH. Choroidal metastasis response to crizotinib in a ROS1-rearranged NSCLC patient. Lung Cancer. 2015;87:207–9.
Zhong K, Chen W, Xiao N, Zhao J. The clinicopathological significance and potential drug target of E-cadherin in NSCLC. Tumour Biol. 2015;36:6139–48.
Botling J, Edlund K, Lohr M, et al. Biomarker discovery in non-small cell lung cancer: integrating gene expression profiling, meta-analysis, and tissue microarray validation. Clin Cancer Res. 2013;19:194–204.
Fiori V, Magnani M, Cianfriglia M. The expression and modulation of CEACAM1 and tumor cell transformation. Ann Ist Super Sanita. 2012;48:161–71.
Barrallo-Gimeno A, Estevez R. GlialCAM, a glial cell adhesion molecule implicated in neurological disease. Adv Neurobiol. 2008;8:47–59.
Arnedo T, Lopez-Hernandez T, Jeworutzki E, et al. Functional analyses of mutations in HEPACAM causing megalencephalic leukoencephalopathy. Hum Mutat. 2014;35:1175–8.
Tao J, Liu Q, Wu X, et al. Identification of hypermethylation in hepatocyte cell adhesion molecule gene promoter region in bladder carcinoma. Int J Med Sci. 2013;10:1860–7.
Moh MC, Zhang T, Lee LH, Shen S. Expression of hepaCAM is downregulated in cancers and induces senescence-like growth arrest via a p53/p21-dependent pathway in human breast cancer cells. Carcinogenesis. 2008;29:2298–305.
Moh MC, Lee LH, Yang X, Shen S. HEPN1, a novel gene that is frequently down-regulated in hepatocellular carcinoma, suppresses cell growth and induces apoptosis in HepG2 cells. J Hepatol. 2003;39:580–6.
Tan B, Tan J, Du H, et al. HepaCAM inhibits clear cell renal carcinoma 786–0 cell proliferation via blocking PKCepsilon translocation from cytoplasm to plasma membrane. Mol Cell Biochem. 2014;391:95–102.
Lee LH, Moh MC, Zhang T, Shen S. The immunoglobulin-like cell adhesion molecule hepaCAM induces differentiation of human glioblastoma U373-MG cells. J Cell Biochem. 2009;107:1129–38.
Yang S, Wu X, Luo C, Pan C, Pu J. Expression and clinical significance of hepaCAM and VEGF in urothelial carcinoma. World J Urol. 2010;28:473–8.
Moh MC, Shen S. The roles of cell adhesion molecules in tumor suppression and cell migration: a new paradox. Cell Adhes Migr. 2009;3:334–6.
Zhang T, Moh MC, Lee LH, Shen S. The immunoglobulin-like cell adhesion molecule hepaCAM is cleaved in the human breast carcinoma MCF7 cells. Int J Oncol. 2010;37:155–65.
Chung Moh M, Hoon Lee L, Shen S. Cloning and characterization of hepaCAM, a novel Ig-like cell adhesion molecule suppressed in human hepatocellular carcinoma. J Hepatol. 2005;42:833–41.
Song X, Wang Y, Du H, et al. Overexpression of hepacam inhibits cell viability and motility through suppressing nucleus translocation of androgen receptor and ERK signaling in prostate cancer. Prostate. 2014;74:1023–33.
Togashi Y, Hayashi H, Terashima M, et al. Inhibition of beta-catenin enhances the anticancer effect of irreversible EGFR-TKI in EGFR-mutated non-small-cell lung cancer with a T790M mutation. J Thorac Oncol. 2015;10:93–101.
Fan C, Jiang G, Zhang X, et al. Zbed3 contributes to malignant phenotype of lung cancer via regulating beta-catenin and P120-catenin 1. Mol Carcinog. 2015;54 Suppl 1:E138–47.
Clevers H, Nusse R. Wnt/beta-catenin signaling and disease. Cell. 2012;149:1192–205.
Yan HB, Wang XF, Zhang Q, et al. Reduced expression of the chromatin remodeling gene ARID1A enhances gastric cancer cell migration and invasion via downregulation of E-cadherin transcription. Carcinogenesis. 2014;35:867–76.
Xie D, Yin D, Tong X, et al. Cyr61 is overexpressed in gliomas and involved in integrin-linked kinase-mediated Akt and beta-catenin-TCF/Lef signaling pathways. Cancer Res. 2004;64:1987–96.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Huanzhang Shao and Yinjie Gu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Shao, H., Gu, Y., Ding, J. et al. HEPACAM inhibited the growth and migration of cancer cells in the progression of non-small cell lung cancer. Tumor Biol. 37, 2621–2627 (2016). https://doi.org/10.1007/s13277-015-4084-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4084-9