Abstract
Tamarind seeds are one of the major sources of different nutrients and minerals and are used as antioxidant components in various food products. Bioactive peptides are released by enzymatic hydrolysis of major food proteins followed by different separation techniques to isolate them from the mixture of other components. Further incorporation of bioactive peptides in various food products may lead to more diversification, fulfilling different nutritional requirements, and promoting health benefits. The objective of this study was to optimize the parameters for the formation of tamarind protein hydrolysates with the use of pepsin and trypsin. It was observed that pepsin and trypsin derived hydrolysate showed the maximum degree of hydrolysis and other bioactive characteristics (DPPH activity, reducing power, ABTS activity, and antidiabetic activity) under optimized conditions. High-performance liquid chromatography was performed for amino acid profiling on trypsin derived protein hydrolysate. From this study, pepsin and trypsin derived hydrolysate can be considered as a rich source of natural antioxidants for developing food formulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The protein-derived fragments, which provide health benefits to humans on their consumption, are termed as bioactive peptides (Bhandari et al. 2019). In recent years, a number of studies have been conducted to extract bioactive peptides from food proteins and expand this area for numerous health benefits. Also, food protein hydrolysates and peptides are considered favourable ingredients for the production of functional foods (Kanbargi et al. 2017), and digestion of these proteins can lead to the release of bio-functional peptides in the body (Sonawane and Arya 2017a, b). There are two methods for isolation of these peptides from food proteins- enzymatic hydrolysis and microbial fermentation. Following these methods, short-chain peptides are separated using different techniques like size-exclusion chromatography, electro-dialysis ultra-filtration, freeze-drying desalting, high-performance liquid chromatography, etc. (Bhandari et al. 2019).
Tamarindus indica L. is one of the most important leguminous tree species, and it is indigenous to tropical Africa and is grown widely in Asian countries. Today, India is the largest producer of tamarind seeds (100,000 tons) from 250,000 tons of tamarind (Singh et al. 2007). Tamarind seeds contain 7–10% of protein with a high amount of essential amino acids and are used as natural antioxidants in food products (Natukunda et al. 2016). Ultra-high performance analyses of tamarind seeds showed the presence of catechin, procyanidin B2, caffeic acid, ferulic acid, chloramphenicol, myricetin, morin, quercetin, apigenin and kaempferol (Razali et al. 2015).
The objective of the study and the present investigation were conducted on tamarind seeds to optimize the conditions for the production of its protein hydrolysates and determine different physiological activities and amino acid profiling by high-pressure liquid chromatography.
Materials and methodology
Sample preparation for protein hydrolysate
Good quality tamarind seeds were procured from the local market of Pune, Maharashtra, India. Sun-dried seeds were ground to form a powder and stored in airtight containers for further use.
Extraction and quantification of protein
Protein extraction was carried out as the method suggested by Kanbargi et al. (2016) with little modifications. 1.5 g of powdered sample was mixed with 20 ml of each solvent (distilled water, 50 mM Tris—HCl buffer pH 7.5, a mixture of 0.6 M NaCl and 0.1% HCl in 1:3 ratio and combination of acetic acid and acetone) separately and stirred on a magnetic stirrer for 2 h each followed by centrifugation at 8000 rpm for 25 min at 4 ºC. The supernatant collected was vacuum dried for 16 h at 40 ºC. Quantification of protein was done by the Lowry method, taking bovine serum albumin as standard. The sieve size used for powder separation was 70 mesh.
Optimization of process parameters for preparation of protein hydrolysate
A central composite RSM design of three variables was used for the preparation of protein hydrolysate. Independent variables included enzyme concentration (A), hydrolysis time (B) and solid–liquid ratio (C). Coded levels of independent variables are shown in Table 1. The levels of these variables were selected based on preliminary literature data. Response variables were the degree of hydrolysis, DPPH activity, reducing power, and antidiabetic activity.
Enzymatic hydrolysis
A solution of extracted protein from Tris HCl buffer was taken for enzymatic hydrolysis with enzymes pepsin (pH 2.5) and trypsin (pH 7.5) separately with each sample. The optimum temperature for both enzymes is 37 ºC. The concentration of protein in the solution was taken as 0.1 mg/mL, and the experiments were run according to the experimental design. After completion of the experiment, a solution was heated in boiling water to stop the action of enzymes, followed by centrifugation at 7000 RPM for 10 min with subsequent filtration.
Degree of hydrolysis
There are different methods through which the degree of hydrolysis can be determined, such as pH–stat, osmometry, soluble nitrogen content and tri-nitrobenzene sulfonic acid (TNBS) method on the basis of different principles (Nielsen et al. 2001). This property is considered as an indicator to screen the proteolysis along with a yield of hydrolysis (Guerard et al. 2002). Kanbargi et al. (2016) defined the degree of hydrolysis as the amount of protein soluble in 10% trichloro-acetic acid (TCA) to the total protein present in the sample. It can be measured by dissolving 500 µL of protein hydrolysate in 500 µL of 10% TCA followed by incubation for 30 min. After centrifugation at 300 rpm for 20 min, quantification of protein was done by the Lowry method of the supernatant to obtain soluble protein and protein hydrolysate to get total protein. It is calculated as:
DPPH activity
DPPH (2,2-diphenyl-1-picrylhydrazyl) activity shows the hydrogen donating efficiency of the component. In this assay, stable free radical DPPH reacts with antioxidants with weak hydrogen bonding resulting in discolouration of the molecule. It is a measure of compounds, which are radical scavengers (Moon and Shibamoto 2009). The colour changes from purple to yellow due to the formation of DPPH on the absorption of hydrogen radical from an antioxidant. DPPH shows a strong absorption band at 517 nm due to its odd electron and solution appears a deep violet colour, the absorption vanishes as the electron pairs off. The resulting decolorization is stoichiometric with respect to the number of electrons taken up (Kedare and Singh 2011). DPPH was taken as control with methanol as blank. The absorbance was measured at 517 nm in a UV–Visible spectrophotometer (Kanbargi et al. 2016). It is calculated as:
Reducing power
The reducing power is based on the transformation of Fe3+ to Fe2+. Ferric Reducing Antioxidant Power assay (FRAP) is based on reduction of a colorless Fe3+-TPTZ complex into intense blue Fe2+-TPTZ once it interacts with a potential antioxidant (Spiegel et al. 2020). Based on the electron transfer mechanism, reducing power was measured by the method suggested by Alam et al. (2013) taking 250 µL of sample and adding 250 µL of 0.2 M sodium- phosphate buffer to it by the addition of 250 µL of 1% potassium ferricyanide. The mixture was heated at 50 ºC for 20 min. Then, 250 µL of 10% trichloroacetic acid was added and shaken gradually. 50 µL of 0.1% FeCl3 was added to 250 µL of the above mixture, after which there was the addition of 200 µL of distilled water. The mixture was incubated for 10 min at room temperature and centrifuged. The absorbance was measured at 700 nm by taking ferrous sulphate as the standard. Here, the increase in absorbance of the reaction mixture implies an increase in the reducing power of the sample, thus antioxidant activity.
Antidiabetic activity
α-amylase inhibition assay was performed for estimation of the antidiabetic activity of the protein sample. The digestive enzyme (α-amylase) is responsible for hydrolyzing dietary starch to maltose, which breaks down to glucose, prior to absorption. Inhibition of the α-amylase should reduce the unfavourable high postprandial blood glucose peak in diabetics. α-amylase inhibition is also a useful target in obesity (Kumanan et al. 2010). 500 µL of sample was added to 500 µL of 0.02 M sodium phosphate buffer containing 0.5 mg/mL α-amylase solution (pH 6.9 with 0.006 M NaCl). The mixture was incubated for 10 min at 25 ºC. 500 µL of 1% starch solution made in 0.02 M sodium phosphate buffer and again kept for incubation for 10 min at 25 ºC, followed by the addition of 1 mL DNSA reagent (dinitro—salicylic acid) and incubation in boiling water bath for 5 min. The solution was allowed to cool down to room temperature with the addition of 10 mL of distilled water (Sathiavelu et al. 2013). The absorbance was taken at 540 nm, and the percentage of antidiabetic activity was calculated as:
Amino acid profiling
High-performance liquid chromatography analysis was performed on a Shimadzu Prominence HPLC system equipped with LC-20AT quaternary gradient pump, SPD-M20A diode array detector (DAD), CBM20A communication bus module, CTO-10AS VP column oven, Rheodyne injector, and Shimadzu LC solution (ver 1.21 SP1) software. A solution of amino acids standards from Sigma (AAS-18) solution containing Alanine, Arginine, Asparagine, Aspartic acid, Glutamic acid, Glycine, Histidine, Isoleucine, Tryptophan, Phenylalanine, Serine, Valine, Tyrosine, Lysine, Glutamine, Leucine, Threonine, Cystine and Methionine was prepared for the calibration curve. Further, the tamarind sample was prepared in Tris–HCl buffer, followed by enzymatic hydrolysis by trypsin at the optimized concentration following the same procedure to form a protein hydrolysate and kept in freezing conditions. The amino acid composition is determined by the HPLC method using phosphate buffer and acetonitrile as eluent with a flow rate of 1 mL/min using detectors Ex: 350 nm; Em: 450 nm. Injection volume was kept as 1 µL for standards and 5 µL for the sample, with a total run time of 50 min.
Statistical analysis
Statistical software package Design Expert ®Version 7.0 (Stat Ease, Inc., Minneapolis, MN, USA) was used for the analysis of the data. Significant differences between the conditions were evaluated by Duncan’s Test. The cluster analysis of data was performed using SPSS 16.0 software to get a comprehensive comparative overview of independent variables. The output obtained was plotted as a dendrogram, and interpretation of data was made accordingly.
Results and discussion
Protein extraction and optimization of processes parameters for preparation of protein hydrolysate
Lowry method was used for the quantification of protein in tamarind seeds after extraction from the mentioned solvents. It was concluded that Tris HCl buffer (42.3 mg protein/g sample) is the most suitable solvent as it resulted in maximum protein extraction than other solvents. Kanbargi et al. (2016) depicted similar results. Therefore, enzymatic hydrolysis was carried out using Tris HCl buffer using both enzymes, pepsin and trypsin.
The effect of different independent variables on the responses was studied considering both the enzymes. Because of the effect of variables, each response varied, and a different range in values of responses was observed. A quadratic model was selected for analysis of all responses. Experimental data fit the response equation with a high degree of significance (P < 0.05). A high value of the coefficient of determination (R2) of the predicted model for both designs was observed. However, lack-of-fit was not significant (P ≤ 0.05) for all responses with each independent variable value. Thus, the models were considered significant due to their subjective nature. For maximum desirability, the pepsin concentration was kept 5% for 2.5 hours of hydrolysis at solid-liquid ratio (1.5:x) of 20 and for trypsin, the trypsin concentration was kept 1.5% for 3 h of hydrolysis at solid–liquid ratio (1.5:x) of 20. The other parameters for analysis of variance are shown in Tables 2 and 3, including F-value, P-value and regression coefficients.
Degree of hydrolysis
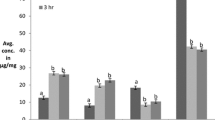
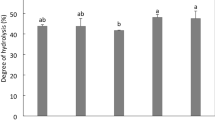
The degree of hydrolysis is defined as the proportion of cleaved peptide bonds in a protein hydrolysate. The hydrolysis of protein in tamarind protein hydrolysate with the use of pepsin was found to be 38–58% and with trypsin 58–79% approximately was found for a different set of independent variables. ANOVA for the quadratic model as fitted to experimental results (Tables 2 and 3) showed R2 values of 0.98 and 0.96 by pepsin and trypsin, respectively. Three of the parameters are solid: liquid, hydrolysis time and enzyme concentration, are the most significant variables for variation in the degree of hydrolysis at the linear level (P ≤ 0.05). Direct positive relation with the gradual increase was observed for variation of the degree of hydrolysis with two independent variables. The degree of hydrolysis increased with an increase in enzyme concentration and hydrolysis time provided for incubation (Figs. 1 and 2), which in turn, is in favour of previous studies reviewed, only if, conditions like pH and temperature are kept according to the type of enzyme used (Baharuddin et al. 2016). During the preparation of protein hydrolysate, it was reported earlier that if given a longer extraction time for peptides from Limonia acidissima, the degree of hydrolysis increased (Sonawane and Arya 2017a, b). Jian et al. (2016) reported the augmentation in the degree of hydrolysis with an increase in enzyme concentration. On the other hand, a decrease in response was observed with an increase in solid–liquid ratio, as less protein will be extracted with an increase in dilution of solvent. Similar results were reported by Lv et al. (2011) while extracting protein from grape seeds.
DPPH activity
DPPH radical scavenging activity varied from 51 to 70% and 55 to 75% in the case of pepsin and trypsin, respectively. An increase in DPPH radical scavenging activity was observed with an increase in enzyme concentration and hydrolysis time, and the increase was significant with P ≤ 0.05 and coefficient of regression of 0.93 and 0.96 for pepsin and trypsin (Tables 2 and 3). In protein hydrolysate, antioxidant activity is mainly dependent on amino acid composition (Chi et al. 2015). DPPH is a free radical and shows a colour-changing phenomenon when interacting with hydrogen donating substance (antioxidant). The degree of hydrolysis has a direct impact on DPPH radical scavenging activity as more of the bioactive peptides would be released with the increase in time and enzyme concentration and hence, the antioxidant activity (Mahmoud et al. 2017). Similar results were reported by Naqash and Nazeer, (2010), depicting the direct relationship between enzyme concentration and DPPH activity. Peptides isolated from trypsin showed more degree of hydrolysis and hence, antioxidant activity. On the contrary, solid–liquid ratio augmentation leads to a decrease in protein extraction leading to a direct influence on DPPH radical scavenging activity (Figs. 1 and 2).
Reducing power
The values varied from 270 to 291 mM of FeSO4/g of sample for pepsin and 370–389 mM of FeSO4/g of sample for trypsin. Lack of fitness was not significant, and hence, both the models for pepsin and trypsin were significant. Similar variation in reducing power was observed for both the experimental designs with the independent variables, as shown in Figs. 1 and 2. A study reported that protein hydrolyzed from Mustelus mustelus had lower reducing power (Bougatef et al. 2009). Another study reported that peptide hydrolysate decreased oxidation by binding with iron or chelating to form peroxidative iron (Zhu et al. 2006). Reducing power increased with increased enzyme concentration and hydrolysis time. Hydrolysis time being the significant (P ≤ 0.05) variable affecting reducing power. A significant decrease was also there due to an increase in the solid–liquid ratio, similar to the DPPH radical scavenging activity. The greater reducing power depicts that hydrolysate could donate electrons to free radicals, which prevents the propagation step (Dey and Dora 2014).
Antidiabetic activity
α-amylase inhibition assay was performed to calculate in vitro antidiabetic activity of the extracted peptides. Hydrolysates are able to inhibit α amylase enzyme to delay the rate of starch hydrolysis (Ngoh and Gan 2016). α-amylase is an enzyme that yields monosaccharides (glucose, maltose) from polysaccharides (glycogen, starch) (Chaudhari et al. 2013). The activity varied from 52 to 74% and 32–51% in pepsin and trypsin, respectively. Lack of fit was not significant, and thus, the model was significant. From the whole variable under study (Tables 2 and 3), hydrolysis time provided for incubation is the most significant variable affecting antidiabetic activity at the linear level (P ≤ 0.05). The reduction of glucose levels in the body due to inhibition by α amylase is an in vitro indicator of controlling diabetes. More time provided for incubation, longer is the interaction time, more release of the peptides, and the enzyme will get sufficient time to release shorter peptides influencing α amylase activity (Ngoh and Gan 2016). With an increase in the solid–liquid ratio, the activity decreases. In addition, not much significant variation was noted with the change in enzyme concentration, as shown in Figs. 1 and 2.
Numerical optimization
Based on the objective of the study, the numerical optimization was aimed to obtain a protein hydrolysate with maximum degree of hydrolysis, antioxidant and antidiabetic activities. Independent variables were set for the goal with values in a range. Results obtained for the optimization process are given in Tables 4 and 5. From the optimization value, the desirability obtained for the process is 0.97 and 0.94 for pepsin and trypsin, respectively. A difference was drawn between the predicted and actual values of the parameters to check the efficiency of the developed conditions and the effectiveness of the experiment.
Cluster analysis
The data obtained from determining different physiological functions of the bioactive peptides isolated from the preparation of protein hydrolysate by pepsin and trypsin were subjected to rescaled distance cluster analysis, and the results are shown in Figs. 3 and 4. It is evident from Fig. 3 that there was formation of two major clusters based on the hydrolysis time provided for incubation. The first cluster comprises of providing time for hydrolysis for protein hydrolysis between 2 and 2.84 h, while the second cluster comprises hydrolysis time of 1.16–1.5 h. It implies that among all the variables under study, hydrolysis time significantly affects the preparation of protein hydrolysate by pepsin. Similarly, in Fig. 4, two major clusters are formed, as there is not much difference in enzyme concentration. Therefore, the synergistic effect of all the variables on the responses was observed.
Amino acid profiling
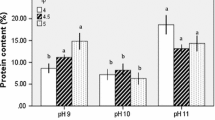
The maximum bioactivity was found for the tamarind sample on the action with trypsin enzyme, which was hence, chosen for determining amino acid composition by HPLC. The sample run with its retention time, area and height are presented in Fig. 5 and Table 6. From the Fig. 5 and Table 6, the composition of amino acids like aspartic acid, glutamic acid, asparagine, serine, histidine, glutamine, glycine, threonine, arginine, alanine, tyrosine, cystine, valine, methionine, phenylalanine, isoleucine, leucine, lysine from bioactive peptides isolated from tamarind seed protein was observed where asparagine showed maximum concentration and tryptophan was found to be absent in tamarind seeds.
Conclusion
Tris HCl buffer was used as the best solvent as it resulted in the maximum extraction of protein from tamarind seeds. Optimum conditions for the preparation of protein hydrolysate with the use of two enzymes, pepsin and trypsin, separately were obtained. The degree of hydrolysis was maximum in trypsin-derived hydrolysate. Cluster analysis data obtained from tamarind seed protein hydrolysate concluded that hydrolysis time was the significant variable affecting the responses majorly. It was observed that augmentation in responses was there with an increase in enzyme concentration and hydrolysis time, whereas a decrease was there with an increase in solid: liquid ratio due to more dilution leading to less interaction with enzyme and less protein extraction. In addition, amino acid profiling by HPLC was performed on trypsin-derived sample. Thus, more studies on tamarind protein hydrolysate for its extraction and purification can lead to the formation of functional foods for numerous human benefits.
Code availability
Not applicable.
References
Alam MN, Bristi NJ, Rafiquzzaman M (2013) Review on in vivo and in vitro methods evaluation of antioxidant activity. Saudi Pharm J 21(2):143–152. https://doi.org/10.1016/j.jsps.2012.05.002
Baharuddin SH, Mustahil NA, Abdullah AA, Sivapragasam M, Moniruzzaman M (2016) Ecotoxicity study of amino acid ionic liquids towards Danio rerio fish: effect of cations. Proc Eng 148:401–408. https://doi.org/10.1016/j.proeng.2016.06.431
Bhandari D, Rafiq S, Gat Y, Gat P, Waghmare R, Kumar V (2019) A review on bioactive peptides: physiological functions, bioavailability and safety. Int J Pept Res Ther. https://doi.org/10.1007/s10989-019-09823-5
Bougatef A, Hajji M, Balti R, Lassoued I, Triki-Ellouz Y, Nasri M (2009) Antioxidant and free radical-scavenging activities of smooth hound (Mustelus mustelus) muscle protein hydrolysates obtained by gastrointestinal proteases. Food Chem 114(4):1198–1205. https://doi.org/10.1016/j.foodchem.2008.10.075
Chaudhari MG, Joshi BB, Mistry KN (2013) In vitro antidiabetic and anti-inflammatory activity of stem bark of Bauhinia purpurea. Bull Pharm Med Sci (BOPAMS) 1(2)
Chi CF, Hu FY, Wang B, Li T, Ding GF (2015) Antioxidant and anticancer peptides from the protein hydrolysate of blood clam (Tegillarca granosa) muscle. J Func Foods 15:301–313. https://doi.org/10.1016/j.jff.2015.03.045
Dey SS, Dora KC (2014) Antioxidative activity of protein hydrolysate produced by alcalase hydrolysis from shrimp waste (Penaeus monodon and Penaeus indicus). J Food Sci Technol 51(3):449–457. https://doi.org/10.1007/s13197-011-0512-z
Guerard F, Guimas L, Binet A (2002) Production of tuna waste hydrolysates by a commercial neutral protease preparation. J Mol Catal B Enzym 19:489–498. https://doi.org/10.1016/S1381-1177(02)00203-5
Jian J, Tian QY, Hettinghouse A, Zhao S, Liu H, Wei J, Overkleeft HS (2016) Progranulin recruits HSP70 to β-glucocerebrosidase and is therapeutic against Gaucher disease. EBioMedicine 13:212–224. https://doi.org/10.1016/j.ebiom.2016.10.010
Kanbargi KD, Sonawane SK, Arya SS (2016) Functional and antioxidant activity of Ziziphus jujube seed protein hydrolysates. J Food Meas Character 10(2):226–235. https://doi.org/10.1007/s11694-015-9297-5
Kanbargi KD, Sonawane SK, Arya SS (2017) Encapsulation characteristics of protein hydrolysate extracted from Ziziphus jujube seed. Int J Food Prop 20(12):3215–3224. https://doi.org/10.1080/10942912.2017.1282516
Kedare SB, Singh RP (2011) Genesis and development of DPPH method of antioxidant assay. J Food Sci Technol 48(4):412–422. https://doi.org/10.1007/s13197-011-0251-1
Kumanan R, Manimaran S, Saleemulla K, Dhanabal SP, Nanjan MJ (2010) Screening of bark of Cinnamomum tamala (Lauraceae) by using α -amylase inhibition assay for anti-diabetic activity. Int J Pharm Biomed Res 1(2):69–72
Lv C, Jia X, Li M, Yang J, Zhao G (2011) Optimization of extraction process of crude protein from grape seeds by RSM. Food Sci Technol Res 17(5):437–445. https://doi.org/10.3136/fstr.17.437
Mahmoud A-H et al (2017) Angiotensin I-converting enzyme inhibitory activity and antioxidant capacity of bioactive peptides derived from enzymatic hydrolysis of buffalo milk proteins. Int Dairy J 66:91–98
Moon JK, Shibamoto T (2009) Antioxidant assays for plant and food components. J Agric Food Chem 57(5):1655–1666. https://doi.org/10.1021/jf803537k
Naqash SY, Nazeer RA (2010) Antioxidant activity of hydrolysates and peptide fractions of Nemipterus japonicus and Exocoetus volitans muscle. J Aquat Food Prod Technol 19(3–4):180–192. https://doi.org/10.1080/10498850.2010.506256
Natukunda S, Muyonga JH, Mukisa IM (2016) Effect of tamarind (Tamarindus indica L.) seed on antioxidant activity, phytocompounds, physicochemical characteristics, and sensory acceptability of enriched cookies and mango juice. Food Sci Nutr 4(4):494–507. https://doi.org/10.1002/fsn3.311
Ngoh YY, Gan CY (2016) Enzyme-assisted extraction and identification of antioxidative and α-amylase inhibitory peptides from Pinto beans (Phaseolus vulgaris cv. Pinto). Food Chem 190:331–337. https://doi.org/10.1016/j.foodchem.2015.05.120
Nielsen PM, Petersen D, Dambmann C (2001) Improved method for determining food protein degree of hydrolysis. J Food Sci 66(5):642–646. https://doi.org/10.1111/j.1365-2621.2001.tb04614.x
Razali N, Mat Junit S, Ariffin A, Ramli NS, Abdul Aziz A (2015) Polyphenols from the extract and fraction of T. indica seeds protected HepG2 cells against oxidative stress. BMC Complementary Altern Med 15:438. https://doi.org/10.1186/s12906-015-0963-2
Sathiavelu A, Sangeetha S, Archit R, Mythili S (2013) In vitro antidiabetic activity of aqueous extract of the medicinal plants Nigella sativa, Eugenia jambolana, Andrographis paniculata and Gymnema sylvestre. Int J Drug Dev Res 5(2):323–328
Singh D, Wangchu L, Moond SK (2007) Processed products of Tamarind. Nat Prod Rad 6(4):315–321
Sonawane SK, Arya SS (2017a) Bioactive L acidissima protein hydrolysates using Box-Behnken design. 3 Biotech 7(3):218. https://doi.org/10.1007/s13205-017-0862-y
Sonawane SK, Arya SS (2017b) Citrullus lanatus protein hydrolysate optimization for antioxidant potential. J Food Meas Charact 11(4):1834–1843. https://doi.org/10.1007/s11694-017-9565-7
Spiegel M, Kapusta K, Kołodziejczyk W, Saloni J, Żbikowska B, Hill GA, Sroka Z (2020) Antioxidant activity of selected phenolic acids–ferric reducing antioxidant power assay and QSAR analysis of the structural features. Molecules 25(13):3088
Zhu D, Lai Y, Shelat PB, Hu C, Sun GY, Lee JC (2006) Phospholipases A2 mediate amyloid-β peptide-induced mitochondrial dysfunction. J Neurosci 26(43):11111–11119. https://doi.org/10.1523/JNEUROSCI.3505-06.2006
Funding
Authors declare that no financial support is received for proposed work.
Author information
Authors and Affiliations
Contributions
DB and SS have drafted the manuscript and YG have guided and have conceded proof reading of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Authors have read and approved the manuscript and all are aware of its submission to 3 Biotech.
Consent for publication
Not applicable.
Human and animal rights
Author declares that proposed work does not contain any animal and human studies.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhandari, D., Shirsat, S. & Gat, Y. Bioactive characterization and optimization of Tamarindus indica seed protein hydrolysate: technology and application of natural enzymes. 3 Biotech 12, 338 (2022). https://doi.org/10.1007/s13205-022-03386-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03386-x