Abstract
Genes encoding wheat low-molecular-weight glutenin subunits (LMW-GSs) that confer dough strength and extensibility were previously identified from Korean wheat cultivars. To improve low viscoelasticity of rice (Oryza sativa L.) dough caused by the lack of seed storage proteins comparable to wheat gluten, two genes, LMW03 and LMW28, encoding LMW-GSs are cloned from Korean wheat cultivar Jokyoung. The LMW genes are inserted into binary vectors under the control of the rice endosperm-specific Glu-B1 promoter. Transgenic rice plants expressing LMW03 or LMW28 in their seeds are generated using Agrobacterium-mediated transformation. The expression of recombinant wheat LMW-GS in the transgenic rice seeds was confirmed by SDS-PAGE and immunoblot analysis. Their accumulation in the endosperm and aleurone layers of rice seeds was observed through in situ immuno-hybridization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L.) and rice (Oryza sativa L.) are major cereal crops worldwide, representing important sources of human nutrition. Unlike rice dough, wheat dough is used in numerous food products including breads, pastas, noodles, couscous, and baked goods. The product diversity and quality of wheat dough are determined by the viscoelastic properties conferred by gluten proteins in wheat seed storage proteins (SSPs). Gluten proteins, comprising monomeric gliadins and polymeric glutenins, contribute to the extensibility and elasticity of wheat dough (Delcour et al. 2012). Based on their mobility in SDS-PAGE (sodium dodecyl sulfate-polyacrylamide gel electrophoresis), polymeric glutenins are divided into high-molecular-weight glutenin subunits (HMW-GSs, 70–90 kDa) and low-molecular-weight glutenin subunits (LMW-GSs, 20–45 kDa), which are linked together by inter-molecular disulfide bonds (Payne 1987). LMW-GSs (~ 60% of total glutenins) are crucial determinants of the processing qualities of wheat end-products and are more effective than HMW-GSs in some cases (Gupta et al. 1989; Cornish et al. 2001; Wang et al. 2016; D’Ovidio and Masci 2004).
LMW-GSs are encoded by genes (Glu-A3, Glu-B3, and Glu-D3) at the Glu-3 loci on the short arms of homoeologous group 1 chromosomes in hexaploid wheat, with 30–43 copies present in the wheat genome (D’Ovidio and Masci 2004; Gao et al. 2007; Lee et al. 2010, 2016; Gupta and Shepherd 1990). Based on their N-terminal sequences, LMW-GSs can be divided into LMW-m, LMW-s, and LMW-i types, indicating that the first amino acid residue of the mature protein is methionine, serine, and isoleucine, respectively (Lee et al. 2016; Rasheed et al. 2014; D’Ovidio and Masci 2004). LMW-s (35–45 kDa) proteins are the most abundant types of LMW-GS in all the genotypes; their N-terminal amino acid sequence is SHIPGL-. The LMW-m type of LMW-GS (30–40 kDa) contains various N-terminal sequences, including METSHI-, METSRI-, and METSCI-. The LMW-i types lack a specific N-terminal amino acid motif and their repetitive domain, ISQQQQ, is found directly after the signal peptide.
Gupta and Shepherd identified six, nine, and five alleles at the Glu-A3, Glu-B3, and Glu-D3 loci, respectively, in various hexaploid wheat cultivars (Gupta and Shepherd 1990). Several groups have also identified various allelic forms of LMW-GS, including four, three, and seven at the Glu-A3, Glu-B3, and Glu-D3 loci in wheat cultivar Xiaoyan 54 (Dong et al. 2010), one, five, and seven in Jokyoung (Lee et al. 2010; Beom et al. 2018), and one, two, and six in Keumkang, respectively (Lee et al. 2016). Using Aroona near-isogenic lines (NILs), Zhang et al. reported that various LMW-GS alleles confer different levels of strength and extensibility to wheat dough (Zhang et al. 2012; Rasheed et al. 2014), providing information to predict dough processing qualities in various wheat cultivars. The alleles at the Glu-A3 loci were ranked as Glu-A3d > Glu-A3b > Glu-A3c > Glu-A3f > Glu-A3a > Glu-A3e for dough strength and Glu-A3c > Glu-A3b = Glu-A3f > Glu-A3e for dough extensibility. The Glu-B3 alleles were ranked as follows: Glu-B3b = Glu-B3d = Glu-B3g > Glu-B3h > Glu-B3a > Glu-B3c for dough strength and Glu-B3i > Glu-B3f = Glu-B3g > Glu-B3h > Glu-B3a = Glu-B3b > Glu-B3d for dough extensibility. The information about the effects of most alleles at Glu-D3 is less complete, although some alleles were successfully ranked such as Glu-D3d = Glu-D3f > Glu-D3e > Glu-D3a = Glu-D3c = Glu-D3b for dough strength (Zhang et al. 2012).
Unlike wheat dough, rice dough has low viscoelasticity, due to the lack of SSPs with properties analogous to those of wheat gluten proteins. Rice SSPs include glutelins (60–80% of total SSPs), prolamins (20–30%), and globulins (2–8%), which are encoded by 15, 34, and 1 gene, respectively (Kawakatsu et al. 2008; Xu and Messing 2009; Yamagata and Tanaka 1986). Of these, prolamins, which are minor components of rice SSPs, are similar to α-/β-/γ-gliadins in wheat, γ-hordein in barley and γ-secalin in rye (Cameron-Mills and Brandt 1988; Kreis et al. 1985; Okita et al. 1985). Prolamins contain high levels of glutamine and low levels of lysine, histidine, cysteine, and methionine (Shyur et al. 1994), and they exhibit fewer elastic and cohesive properties than wheat glutenins (Koehler and Wieser 2013).
Allelic forms of HMW-GSs and LMW-GSs in various bread wheat cultivars contribute to their excellent bread-making quality. Studies have been undertaken to improve the dough extensibility and elasticity in wheat based on these quality scores (Rooke et al. 1999; Gadaleta et al. 2008; Popineau et al. 2001) and rye (Altpeter et al. 2004). Overexpression of the wheat HMW-GS, 1Dx5, resulted in improved end-product quality. Moreover, the bread-making qualities of rice were improved by adding wheat gluten to rice dough (Sivaramakrishnan et al. 2004; Shin 2009). In addition, many molecular breeding studies have been undertaken to improve the bread-making qualities of rice through genetic engineering, including the production of soybean β-conglycinin subunits in rice seeds to improve nutritional and processing properties (Motoyama et al. 2009) and the expression of wheat HMW-GSs such as 1Ax1, 1Bx7, 1Dx5, and 1Dy10 in rice seeds to improve dough-making qualities (Jeong et al. 2016; Park et al. 2014; Oszvald et al. 2007, 2013; Jo et al. 2017). However, improving the poor suitability of rice flour for various food products remains an important challenge in countries where rice is used as a staple crop.
In this study, to engineer wheat LMW-GS production in rice seeds, we cloned LMW03 and LMW28 from Korean wheat cultivar Jokyoung and placed them in expression cassettes under the control of the rice endosperm-specific Glu-B1 promoter (Qu et al. 2008). We introduced these cassettes into the genome of Korea rice cultivar Koami, with high amylose content, and selected individual transgenic rice lines expressing LMW03 and LMW28 by SDS-PAGE and immunoblot analysis. Finally, we observed the cellular localization of these proteins in rice seed endosperm by in situ immuno-hybridization.
Materials and methods
Vector construction and rice transformation
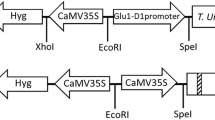
Previously, 43 genes encoding LMW-GSs from immature seeds of common Korean wheat cultivar Jokyoung (Triticum aestivum L. cv. Jokyoung) were cloned to the pGEM-T Easy vector (Promega) using LMW-GS gene-specific primers (Lee et al. 2010). Here, two of these genes, LMW03 (GenBank accession no. HQ619896) and LMW28 (no. HQ619899), were amplified by PCR using two primer pairs (for LMW03, forward primer 5′-AAAAAGCAGGCTATGAAGACCTTCCTCATCTTTGCCCTC-3′ and reverse primer 5′-AGAAAGCTGGGTTCAGTAGGCACCAACTCGGGTGC-3′, and for LMW28, forward primer 5′-AAAAAGCAGGCTATGAAGACCTTCCTCATCTTTGCTCTC-3′ and reverse primer 5′-AGAAAGCTGGGTTCAGTAGCCACCAACTCCAGTGCC-3′) containing an attB1 or attB2 for use in the Gateway cloning system. The amplified products were subcloned into the pDONR221 vector (Invitrogen, USA) and inserted into destination binary vector pMJ103 using the Gateway system (Karimi et al. 2002; Jo et al. 2017). The binary vector included the rice seed endosperm-specific Glu-B1 promoter, nopaline synthase (NOS) terminator and bialaphos resistance (Bar) gene as an herbicide-resistance marker (Fig. 3a). Using Agrobacterium tumefaciens (LBA4404)-mediated transformation, LMW03 and LMW28 were inserted into the genome of japonica-type Korean rice cultivar Koami as previously described (Kim et al. 2011; Jo et al. 2017).
Extraction of total seed storage proteins
To extract total rice SSPs, immature seeds were harvested from transgenic and non-transgenic rice plants at various stages of seed development, including 3, 5, 7, 10, 15, 20, 30, 40, and 50 days after flowering (DAF). Each frozen seed harvested from an individual line was ground in a bead beater (4.5 ms−1, 25 s, three repetitions, MP 24 × 4, FastPrep-24, MP) and combined with 350 µL of SDS-urea buffer (250 mM Tris–HCl, pH 6.8, 4% SDS, 8 M urea, 20% glycerol, and 5% β-mercaptoethanol) for 3 h at room temperature, followed by centrifugation at 15,000g for 10 min. The supernatant (containing total SSPs) was used for subsequent experiments, as previously described (Jo et al. 2017; Cho et al. 2016). Total SSP extraction for each sample was independently performed from three biological replicates and analyzed by SDS-PAGE and immunoblotting.
SDS-PAGE and immunoblotting
For SDS-PAGE, 10 µL of each total SSP sample was loaded onto a 12.5% SDS-PAGE gel, stained with Coomassie brilliant blue staining solution [0.1% (w/v) CBB R-250, 45% (v/v) methanol, and 45% (v/v) glacial acetic acid] for 3 h and destained in 10% (v/v) methanol and 10% (v/v) glacial acetic acid. The separated proteins on gels were blotted onto polyvinylidene difluoride membranes (PVDF; Bio-Rad) using a semi-dry transfer machine (Bio-Rad) for 2 h as previously described (Jo et al. 2017). Immunoblot analysis was performed using polyclonal-rat anti-LMW-GS primary antibody (diluted 1:2000) and horseradish-peroxidase (HRP)-conjugated anti-rat IgG secondary antibody (1:10,000, Promega, Madison, WI, USA). Target protein signals were visualized using a luminescence image analyzer (Chemiluminescence Fusion SL, Vilber Lourmat, Marne-la-Vallée, France).
In situ immuno-hybridization
Mature seeds from transgenic and non-transgenic rice plants were soaked overnight in distilled water and vertically sectioned with a razor blade. The sections were washed in distilled water, incubated in blocking solution [TBS, pH 6.8, containing 2% (w/v) skim milk] for 3 h and incubated overnight at 4 °C in blocking solution with polyclonal-rat anti-LMW-GS primary antibody (1:2000). The incubated sections were washed three times with washing solution (TBS, pH 6.8, containing 0.05% Tween-20) and incubated in washing solution with alkaline phosphatase (AP)-conjugated anti-rat IgG secondary antibody (1:10,000, Promega). Finally, the samples were washed three times in TBS with 0.05% Tween-20. Target proteins were visualized using the ProtoBlot®II AP system (Promega) and observed under a stereoscopic microscope (Leica M205C Microsystem, Leica, Heerbrugg, Switzerland) (Jo et al. 2017).
Phylogenetic tree analysis
The amino acid sequences of LMW03 (HQ619896) and LMW28 (HQ619899) from wheat cultivar Jokyoung were used as queries to align with the amino acid sequences of 45 LMW-GS alleles in various accessions in the micro-core collection (MCC) of Chinese wheat germplasm (Zhang et al. 2013). Multiple sequence alignment was then performed using Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/), and the phylogenetic tree was constructed with Dendroscope 3 software (ver. 3.5.7) as previously described (Cho et al. 2018).
Results and discussion
Structural characteristics of wheat low-molecular-weight glutenin subunits LMW03 and LMW28
Forty-three LMW-GS genes were previously cloned from the common Korean wheat cultivar Jokyoung, a hard white winter wheat variety that is widely used for both bread and noodle production in Korea, using LMW-GS-specific primers; these genes were classified into 13 haplotypes (Lee et al. 2010; Beom et al. 2018). Here, we cloned two of these genes, LMW03 and LMW28 (Fig. 1a), in a binary vector under the control of the endosperm-specific Glu-B1 promoter to produce transgenic rice plants overexpressing these proteins in seeds. The primary structures of LMW03 and LMW28 consist of a signal peptide, one N-terminal domain, one repetitive domain, and three C-terminal domains (I, II, and III). Their N-terminal amino acid sequences are METSHIPS for LMW03 and METSRVPG for LMW28 (Fig. 1a), indicating that they are m-type LMW-GSs. These proteins contain eight cysteine residues: one in the repetitive domain, five in C-terminal domain I, one in C-terminal domain II, and one in C-terminal domain III. According to the typical structures of LMW-GSs (D’Ovidio and Masci 2004; Kohler et al. 1993; Orsi et al. 2001), the second and fifth cysteines (Cys-166/-201 for LMW03 and Cys-179/-214 for LMW28), the third and fourth cysteines (Cys-174/-194 for LMW03 and Cys-187/-207 for LMW28), and the sixth and eighth cysteines (Cys-202/-311 for LMW03 and Cys-215/-326 for LMW28) are predicted to form intra-molecular disulfide bonds. The remaining (first and seventh) cysteines (Cys-65/-263 for LMW3 and Cys-46/-277 for LMW28) are thought to participate in inter-molecular disulfide bond formation (Fig. 1b).
Amino acid sequences and structures of low-molecular-weight glutenin subunits LMW03 and LMW28. a Amino acid sequence alignment of LMW03 and LMW28. b Structural analysis of LMW03 and LMW28. SP signal peptide, N- N-terminal region, C cysteine residues. Lines indicate intra-molecular disulfide bonds between two cysteines
To determine if LMW03 and 28 are expressed from Glu-3 genes on the short arms of chromosome 1A, 1B, and 1D in hexaploid wheat, we performed multiple sequence alignment with the amino acid sequences of 45 LMW-GS alleles whose genome positions have been determined in Chinese wheat germplasm (Zhang et al. 2013). As shown in Fig. 2, LMW03 is clustered into the Glu-B3 allele group, including B3-530a (JX877786), B3-530b (JF339179), and B3-530c (JX878003). LMW28 was classified into the Glu-D3 allele group including D3-522 (JX877918), D3-525 (JF339162), and D3-528 (JF339178), indicating that LMW03 and LMW28 are expressed from Glu-3 genes on chromosome 1B and 1D, respectively.
Furthermore, we previously reported that the LMW-GS alleles in Korean wheat cultivar Jokyoung include Glu-A3c, Glu-B3h, and Glu-D3a, as revealed by proteomic analysis (Beom et al. 2018). According to the ranking of LMW-GS alleles for dough strength and extensibility (Zhang et al. 2012), the effect of LMW-GS alleles in Jokyoung on end-product quality is believed to be good, although the results have not yet been reported.
Transgenic rice plants overexpressing wheat low-molecular-weight glutenin subunits
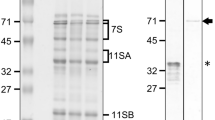
To produce transgenic rice plants overexpressing wheat low-molecular-weight glutenin subunits (LMW-GSs), we isolated two LMW-GS genes, LMW03 and LMW28, from Korean wheat cultivar Jokyoung using LMW-GS gene-specific primers and constructed binary vectors to express LMW03 and LMW28 under the control of the rice endosperm-specific Glu-B1 promoter, as shown in Fig. 3a. Using Agrobacterium-mediated transformation, four and five independent T0 transgenic rice lines overexpressing LMW03 and LMW28 in rice seeds, respectively, were produced. Among the transgenic rice lines that survived to the next generation (T1), we selected three representative lines (K11-1, K11-2, and K11-3) expressing LMW28 and three (K16-1, K06-2, and K06-3) expressing LMW03 from among the progeny (T4 generation) of the transgenic lines based on SDS-PAGE and immunoblot analysis (Fig. 3b). The intensity of a protein band at ~ 40 kDa (red arrow) on the SDS-PAGE gel seemed to be increased in seed protein extracts of the transgenic rice lines compared to wild-type rice. Indeed, immunoblot analysis showed the overexpression of LMW03 and LMW28 in seeds of the transgenic rice lines (Fig. 3c).
Binary vector construction and generation of transgenic rice plants expressing wheat LMW-GSs LMW03 and LMW28. a Binary vector used to generate transgenic rice plants expressing LMW03 and LMW28. The vector contains the Glu-B1 promoter, proteinase inhibitor II terminator (tPinII), cauliflower mosaic virus (CaMV) 35S promoter (p35S), and bialaphos resistance (Bar) gene as a herbicide-resistance marker and Nopaline synthase terminator (tNOS). b, c Expression analysis of total seed storage proteins (SSPs) in mature seeds of Koami (wild-type) and independent transgenic rice lines using SDS-PAGE and immunoblot analysis. Total SSPs (10 µL) were separated by SDS-PAGE (12.5%) and visualized by Coomassie brilliant blue (CBB) staining. Black arrows indicate rice SSPs including glutelin, prolamin, and globulin. Red and blue arrows indicate upregulated proteins in seeds of both transgenic rice lines. S/M size markers, WT wild type, T_LMW28 and 03 indicate transgenic rice lines expressing LMW28 and 03, respectively
In addition, we observed the intensity changes of one band at ~ 70 kDa (blue arrow), glutelin precursor at ~ 50 kDa, basic glutelin at ~ 30 kDa, acid glutelin at ~ 18 kDa, and prolamin at ~ 13 kDa on the SDS-PAGE gel in transgenic rice lines overexpressing LMW-GSs compared to wild type (Fig. 3b). Similar results were previously observed in transgenic rice seeds with suppressed expression of SSPs such as glutelin A, 13 kDa prolamin, and globulin. Two-DGE analysis coupled with LC-MS/MS showed that the individual suppression of glutelin, prolamin, and globulin expression is accompanied by the increased expression of ER chaperones, including binding protein (BiP) and protein disulfide isomerase-like (PDIL) (Cho et al. 2016; Lee et al. 2015; Kim et al. 2012). For example, Takaiwa et al. reported that foreign gene expression in rice seeds resulted in altered SSP expression (Takaiwa et al. 2015). The levels of Cys-poor 13-kDa prolamin increased in glutelin- and/or globulin-deficient transgenic rice seeds, whereas the levels of glutelins and ER chaperones increased in transgenic rice seeds with suppressed expression of 13-kDa prolamin (Kawakatsu et al. 2010). Heavy loading of recombinant proteins on the ER in often leads to ER stress, resulting in the production of ER chaperones accompanied by chalky and shriveled seeds (Qian et al. 2015; Wakasa et al. 2013). However, in the current study, transgenic rice seeds expressing LMW-GSs did not appear chalky or shriveled. These results indicate that the production of LMW-GSs and altered SSP expression in transgenic rice seeds (Fig. 3b) might cause ER stress, but it is not severe enough to alter seed phenotypes.
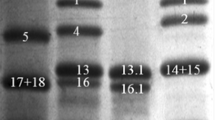
Furthermore, to observe where and when LMW-GSs accumulate in transgenic rice seeds, we observed their expression in seeds at different stages of development. The wheat LMW-GSs began to accumulate at 10 days after flowering (DAF) in immature seeds of transgenic rice and peaked at 30 DAF (Fig. 4a), which is similar to the expression pattern of rice glutelin, indicating the strict control of LMW-GS expression driven by the Glu-B1 promoter. To confirm the localization of the LMW-GSs in transgenic rice seeds, we performed in situ immuno-hybridization using primary antibodies specific for wheat LMW-GSs (Fig. 4b). The LMW-GSs accumulated in the endosperm and aleurone layer in transgenic rice seeds (Fig. 4b). A similar accumulation pattern was reported for transgenic rice seeds overexpressing HMW-GSs and ferritin under the control of the Glu-B1 promoter (Jo et al. 2017; Tosi et al. 2009).
Analysis of LMW-GS expression according to developmental stage and position in rice seeds. a LMW-GS expression was analyzed at 3–50 days after flowering in wild-type rice cultivar Koami (WT) and independent transgenic rice lines T_LMW03 and 28 overexpressing wheat LMW-GSs LMW03 and LMW28, respectively, by immunoblot analysis using polyclonal anti-LMW-GSs’ antibodies. b In situ immuno-hybridization in mature seeds. A magnified view of each red-boxed region in each line is shown. Scale bar: 1 mm
Conclusion
With the aim of improving the processing suitability of rice flour through the transgenic expression of viscoelastic proteins that confer high dough elasticity, we cloned wheat LMW-GS genes LMW03 and LMW28 from Korean wheat cultivar Jokyoung and generated transgenic rice seeds overexpressing wheat LMW03 and LMW28. The production of wheat LMW-GSs in rice seeds resulted in expression changes in rice seed storage proteins such as glutelin and prolamin. In situ immuno-hybridization demonstrated that the LMW-GSs accumulated in the endosperm of transgenic rice seeds. The study, as an important step for genetic engineering to develop the improved processing quality of rice dough, improves our understanding of the interaction between endogenous and recombinant seed proteins in rice.
References
Altpeter F, Popelka JC, Wieser H (2004) Stable expression of 1Dx5 and 1Dy10 high-molecular-weight glutenin subunit genes in transgenic rye drastically increases the polymeric glutelin fraction in rye flour. Plant Mol Biol 54(6):783–792. https://doi.org/10.1007/s11103-004-0122-5
Beom H-R, Kim JS, Jang Y-R, Lim S-H, Kim C-K, Lee CK, Lee J-Y (2018) Proteomic analysis of low-molecular-weight glutenin subunits and relationship with their genes in a common wheat variety. 3 Biotech 8(1):56
Cameron-Mills V, Brandt A (1988) A gamma-hordein gene. Plant Mol Biol (4):449–461. https://doi.org/10.1007/BF00039026
Cho K, Lee H-J, Jo Y-M, Lim S-H, Rakwal R, Lee J-Y, Kim Y-M (2016) RNA interference-mediated simultaneous suppression of seed storage proteins in rice grains. Front Plant Sci 7:1624
Cho K, Beom H-R, Jang Y-R, Altenbach S, Vensel WH, Simon-Buss A, Lim S-H, Kim MG, Kim JS, Lee J-Y (2018) Proteomic profiling and epitope analysis of the complex α-, γ-and ω-gliadin families in a commercial bread wheat. Front Plant Sci 9:818
Cornish GB, Bekes F, Allen H, Martin D (2001) Flour proteins linked to quality traits in an Australian doubled haploid wheat population. Aust J Agric Res 52(12):1339–1348
D’Ovidio R, Masci S (2004) The low-molecular-weight glutenin subunits of wheat gluten. J Cereal Sci 39(3):321–339
Delcour JA, Joye IJ, Pareyt B, Wilderjans E, Brijs K, Lagrain B (2012) Wheat gluten functionality as a quality determinant in cereal-based food products. Annu Rev Food Sci Technol 3:469–492
Dong L, Zhang X, Liu D, Fan H, Sun J, Zhang Z, Qin H, Li B, Hao S, Li Z (2010) New insights into the organization, recombination, expression and functional mechanism of low molecular weight glutenin subunit genes in bread wheat. PLoS One 5(10):e13548
Gadaleta A, Blechl AE, Nguyen S, Cardone MF, Ventura M, Quick JS, Blanco A (2008) Stably expressed d-genome-derived HMW glutenin subunit genes transformed into different durum wheat genotypes change dough mixing properties. Mol Breed 22(2):267–279. https://doi.org/10.1007/s11032-008-9172-8
Gao S, Gu YQ, Wu J, Coleman-Derr D, Huo N, Crossman C, Jia J, Zuo Q, Ren Z, Anderson OD, Kong X (2007) Rapid evolution and complex structural organization in genomic regions harboring multiple prolamin genes in the polyploid wheat genome. Plant Mol Biol 65(1–2):189–203. https://doi.org/10.1007/s11103-007-9208-1
Gupta RB, Shepherd K (1990) Two-step one-dimensional SDS-PAGE analysis of LMW subunits of glutelin. Theor Appl Genet 80(1):65–74
Gupta RB, Singh NK, Shepherd K (1989) The cumulative effect of allelic variation in LMW and HMW glutenin subunits on dough properties in the progeny of two bread wheats. Theor Appl Genet 77(1):57–64
Jeong N, Jeon S-H, Kim D-Y, Lee C, Ok H-C, Park K-D, Hong H-C, Lee S-S, Moon J-K, Park S-K (2016) Development of marker-free TaGlu-Ax1 transgenic rice harboring a wheat high-molecular-weight glutenin Subunit (HMW-GS) protein. J Life Sci 26(10):1121–1129
Jo Y-M, Cho K, Lee H-J, Lim S-H, Kim JS, Kim Y-M, Lee J-Y (2017) Cellular localization of wheat high molecular weight glutenin subunits in transgenic rice grain. Int J Mol Sci 18(11):2458
Karimi M, Inze D, Depicker A (2002) GATEWAY((TM)) vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7(5):193–195. https://doi.org/10.1016/S1360-1385(02)02251-3
Kawakatsu T, Yamamoto MP, Hirose S, Yano M, Takaiwa F (2008) Characterization of a new rice glutelin gene GluD-1 expressed in the starchy endosperm. J Exp Bot 59(15):4233–4245. https://doi.org/10.1093/jxb/ern265
Kawakatsu T, Hirose S, Yasuda H, Takaiwa F (2010) Reducing rice seed storage protein accumulation leads to changes in nutrient quality and storage organelle formation. Plant Physiol 154(4):1842–1854. https://doi.org/10.1104/pp.110.164343
Kim Y-M, Lee J-Y, Yoon U-H, Choi S-B, Ha S-H, Lim S-H (2011) New design of rice seed storage proteins. J Plant Biotechnol 38(4):263–271
Kim Y-M, Lee J-Y, Lee T, Lee Y-H, Kim S-H, Kang S-H, Yoon U-H, Ha S-H, Lim S-H (2012) The suppression of the glutelin storage protein gene in transgenic rice seeds results in a higher yield of recombinant protein. Plant Biotechnol Rep 6(4):347–353
Koehler P, Wieser H (2013) Chemistry of cereal grains. In: Gobbetti M, Gänzle M (eds) Handbook on sourdough biotechnology. Springer, Boston, pp 11–45. https://doi.org/10.1007/978-1-4614-5425-0_2
Kohler P, Belitz HD, Wieser H (1993) Disulphide bonds in wheat gluten: further cystine peptides from high molecular weight (HMW) and low molecular weight (LMW) subunits of glutenin and from gamma-gliadins. Z Lebensm Unters Forsch 196(3):239–247
Kreis M, Forde BG, Rahman S, Miflin BJ, Shewry PR (1985) Molecular evolution of the seed storage proteins of barley, rye and wheat. J Mol Biol 183(3):499–502
Lee J-Y, Kim Y-T, Kim B-M, Lee J-H, Lim S-H, Ha S-H, Ahn S-N, Nam M-H, Kim Y-M (2010) Cloning of low-molecular-weight glutenin subunit genes and identification of their protein products in common wheat (Triticum aestivum L.). Korean J Breed Sci 42(5):547–554
Lee HJ, Jo YM, Lee JY, Lim SH, Kim YM (2015) Lack of globulin synthesis during seed development alters accumulation of seed storage proteins in rice. Int J Mol Sci 16(7):14717–14736. https://doi.org/10.3390/ijms160714717
Lee JY, Beom HR, Altenbach SB, Lim SH, Kim YT, Kang CS, Yoon UH, Gupta R, Kim ST, Ahn SN, Kim YM (2016) Comprehensive identification of LMW-GS genes and their protein products in a common wheat variety. Funct Integr Genom 16(3):269–279. https://doi.org/10.1007/s10142-016-0482-3
Motoyama T, Maruyama N, Amari Y, Kobayashi K, Washida H, Higasa T, Takaiwa F, Utsumi S (2009) α′ Subunit of soybean β-conglycinin forms complex with rice glutelin via a disulphide bond in transgenic rice seeds. J Exp Bot 60(14):4015–4027
Okita TW, Cheesbrough V, Reeves CD (1985) Evolution and heterogeneity of the alpha-/beta-type and gamma-type gliadin DNA sequences. J Biol Chem 260(13):8203–8213
Orsi A, Sparvoli F, Ceriotti A (2001) Role of individual disulfide bonds in the structural maturation of a low molecular weight glutenin subunit. J Biol Chem 276(34):32322–32329
Oszvald M, Jenes B, Tömösközi S, Bekes F, Tamas L (2007) Expression of the 1D × 5 high molecular weight glutenin subunit protein in transgenic rice. Cereal Res Commun 35(4):1543–1549
Oszvald M, Balázs G, Pólya S, Tömösközi S, Békés F, Tamás L (2013) Wheat storage proteins in transgenic rice endosperm. J Agric Food Chem 61(31):7606–7614
Park S-K, Shin D, Hwang W-H, Hur Y-J, Kim T-H, Oh S-Y, Cho J-H, Han S-I, Lee S-S, Nam M-H (2014) Development of marker-free transgenic rice expressing the wheat storage protein, Glu-1Dy10, for increasing quality processing of bread and noodles. J Life Sci 24(6):618–625
Payne PI (1987) Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Annu Rev Plant Physiol 38(1):141–153
Popineau Y, Deshayes G, Lefebvre J, Fido R, Tatham AS, Shewry PR (2001) Prolamin aggregation, gluten viscoelasticity, and mixing properties of transgenic wheat lines expressing 1Ax and 1Dx high molecular weight glutenin subunit transgenes. J Agric Food Chem 49(1):395–401
Qian D, Tian L, Qu L (2015) Proteomic analysis of endoplasmic reticulum stress responses in rice seeds. Sci Rep 5:14255
Qu LQ, Xing YP, Liu WX, Xu XP, Song YR (2008) Expression pattern and activity of six glutelin gene promoters in transgenic rice. J Exp Bot 59(9):2417–2424. https://doi.org/10.1093/jxb/ern110
Rasheed A, Xia XC, Yan YM, Appels R, Mahmood T, He ZH (2014) Wheat seed storage proteins: advances in molecular genetics, diversity and breeding applications. J Cereal Sci 60(1):11–24. https://doi.org/10.1016/j.jcs.2014.01.020
Rooke L, Bekes F, Fido R, Barro F, Gras P, Tatham AS, Barcelo P, Lazzeri P, Shewry PR (1999) Overexpression of a gluten protein in transgenic wheat results in greatly increased dough strength. J Cereal Sci 30(2):115–120. doi:https://doi.org/10.1006/jcrs.1999.0265
Shin M (2009) Rice-processed food. Food Sci Ind 42:2–18
Shyur L-F, Wen T-N, Chen C-S (1994) Purification and characterization of rice prolamins. Bot Bull Acad Sin 35:65–71
Sivaramakrishnan HP, Senge B, Chattopadhyay PK (2004) Rheological properties of rice dough for making rice bread. J Food Eng 62(1):37–45. https://doi.org/10.1016/S0260-8774(03)00169-9
Takaiwa F, Wakasa Y, Takagi H, Hiroi T (2015) Rice seed for delivery of vaccines to gut mucosal immune tissues. Plant biotechnol J 13(8):1041–1055
Tosi P, Parker M, Gritsch CS, Carzaniga R, Martin B, Shewry PR (2009) Trafficking of storage proteins in developing grain of wheat. J Exp Bot 60(3):979–991
Wakasa Y, Yasuda H, Takaiwa F (2013) Secretory type of recombinant thioredoxin h induces ER stress in endosperm cells of transgenic rice. J Plant Physiol 170(2):202–210
Wang Y, Zhen S, Luo N, Han C, Lu X, Li X, Xia X, He Z, Yan Y (2016) Low molecular weight glutenin subunit gene Glu-B3h confers superior dough strength and breadmaking quality in wheat (Triticum aestivum L.). Sci Rep 6:27182
Xu JH, Messing J (2009) Amplification of prolamin storage protein genes in different subfamilies of the Poaceae. Theor Appl Genet 119(8):1397–1412. https://doi.org/10.1007/s00122-009-1143-x
Yamagata H, Tanaka K (1986) The site of synthesis and accumulation of rice storage proteins. Plant Cell Physiol 27(1):135–145
Zhang X, Jin H, Zhang Y, Liu D, Li G, Xia X, He Z, Zhang A (2012) Composition and functional analysis of low-molecular-weight glutenin alleles with Aroona near-isogenic lines of bread wheat. BMC Plant Biol 12:243. https://doi.org/10.1186/1471-2229-12-243
Zhang XF, Liu DC, Zhang JH, Jiang W, Luo GB, Yang WL, Sun JZ, Tong YP, Cui DQ, Zhang AM (2013) Novel insights into the composition, variation, organization, and expression of the low-molecular-weight glutenin subunit gene family in common wheat. J Exp Bot 64(7):2027–2040. https://doi.org/10.1093/jxb/ert070
Acknowledgements
This work was supported by grants from the National Institute of Agricultural Science (RDA PJ012458) and the Next-Generation BioGreen 21 Program (RDA PJ013149 and RDA PJ013159), South Korea. This study was also supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2016R1A6A3A11931182) to KC.
Author information
Authors and Affiliations
Contributions
YMJ designed and conducted the experiments; KC conducted the experiments and wrote the manuscript; SHL, JYK, and OH contributed critical reading and revision of the manuscript; JYL contributed scientific advice and corrected the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Cho, K., Jo, YM., Lim, SH. et al. Overexpressing wheat low-molecular-weight glutenin subunits in rice (Oryza sativa L. japonica cv. Koami) seeds. 3 Biotech 9, 49 (2019). https://doi.org/10.1007/s13205-019-1579-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-019-1579-x