Abstract
Probiotics become important bacteria in our daily life due to their benefit on human health. In this study, a subset of bacterial strains from children was isolated and evaluated for beneficial probiotic traits such as antimicrobial activity, bile and acid tolerance, and pathogenic cell adherence inhibition. The strain with the best antimicrobial activity was selected for further characterization on the basis of morphological, biochemical characteristics and gene sequence. This strain was Gram-positive, oxidase and catalase-negative, and it produced acids by fermenting sugar and starch as carbon sources. Additionally, it could only hydrolyze bile-esculin, but not red blood cells. The 16S rDNA gene sequence revealed that this strain was Enterococcus faecalis. Interestingly, this strain effectively inhibited a variety of pathogens by acid and bacteriocin production and was bile-tolerant, able to survive under acidic condition. In the safety assessments, E. faecalis MTC 1032 could adhere to host epithelial cells and evidently inhibited pathogenic cell adhesion as demonstrated by cell reduction over time of E. coli ATCC 25922 and S. typhimurium ATCC 13311 on Caco-2 cell line. In summary, it was clearly represented that E. faecalis MTC 1032 provided suitable properties and could be a candidate as a probiotic strain in food supplements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Probiotics are generally defined as live microbial supplements which provide health benefits to the host by bringing balance in the host intestinal microbial flora (Fuller 1989). Probiotics have been reported as therapeutic microorganisms in many cases such as constipation (Koebnick et al. 2003), cholesterolemia (Lye et al. 2009), hypertension (Lye et al. 2009), colorectal carcinoma (Osterlund et al. 2007), ulcerative colitis (Floch 2010), Crohn’s diseases, irritable bowel syndrome (Krammer et al. 2005), food allergies (del Giudice et al. 2010) and antibiotic induced diarrhea (Alam and Mushtaq 2009). Moreover, several metabolic compounds including butyric acid, lactic acid and other agents produced by probiotic strains have antimicrobial effects (Gu et al. 2008; Kaewsrichan et al. 2007; Nagpal et al. 2012). A number of research have reported antagonistic properties of probiotics against many common gastrointestinal pathogens, e.g. Helicobacter pylori (Mukai et al. 2002), Clostridium perfringens (Kim et al. 2007), Campylobacter jejuni (Chaveerach et al. 2004), Salmonella spp. (Casey et al. 2004), Escherichia coli (Miyazaki et al. 2010) and Listeria monocytogenes (Kramer 1977).
A number of requirements have been identified for strains to be potential probiotics. Probiotics must be non-pathogenic and are able to survive through the gastrointestinal tract condition (Guo et al. 2010; Noguchi et al. 2012). The required characteristics include resistance to gastric acid and bile tolerance (Fuller 1989). In addition, another important issue for probiotics is the ability to produce antimicrobial compounds, including enzymes, organic acids, hydrogen peroxide and bacteriocins (antimicrobial peptides) and other biologically active substances that can inhibit the activity or growth of enteric pathogens (Linaje et al. 2004; Rodríguez et al. 2012). The major probiotic strains include lactic acid bacteria (LAB) (Sanders 2000; Tinrat et al. 2011), but probiotic properties for certain Enterococci strains have also been reported (Franz et al. 2003; Linaje et al. 2004). E. faecium M-74 (Hlivak et al. 2005) and E. faecium SF68 (Canani et al. 2007) are well known as commercial probiotic strains. Additionally, E. faecium EK13 was used to reduce contamination of Salmonella enterica serovar Dusseldorf in gonobiotic Japanese quails by producing bacteriocins (Kühn et al. 2003). It is clearly shown that Enterococci used as the candidate for probiotic strain still hold a great importance. Therefore, this study was focused on identifying the new Enterococcus strain as a novel probiotic source. Beneficial probiotic traits were also extensively investigated for future development of human food supplements.
Materials and methods
Isolation of potential probiotic strains
Bacterial samples were collected from children feces and were transferred to phosphate buffer saline (pH 7.0) as an initial dilution. Then, the tenfold dilution series were prepared up to 10−4 before each dilution was cultivated onto brain heart infusion (BHI; Difco) agar plates and incubated at 37 °C for 24 h. After incubation, isolated colonies were selected and subcultured onto BHI plates before storage at − 80 °C in 30% glycerol BHI broth.
Preliminary screening for antimicrobial activity
Antimicrobial assay was performed on each isolated colonies. In brief, overnight culture of isolated strains was inoculated on the BHI plate as a lower layer. Next, overnight culture of selected indicator strains (Salmonella typhi and Salmonella typhimurium) was adjusted to yield approximately 1.0 × 108 CFU/ml. After that, an aliquot of Salmonella spp. culture was mixed into 0.75% BHI soft agar and overlaid as a top layer agar. Plates were incubated at 37 °C for 24 h and colonies that gave a surrounding clear zone were selected for further study.
Identification of the isolated strains
Primary identification was preformed based on biochemical and physiological characteristics of the isolated strains. The assays for oxidase activity and catalase activity were performed with 1% N,N,N′,N′-tetramethyl-p-phenylenediamine and 5% hydrogen peroxide, respectively. Pseudomonas aeruginosa ATCC 9027 was a positive control and Escherichia coli ATCC 25922 was a negative control in the oxidase test. Staphylococcus aureus ATCC 25923 and Enterococcus sp. were used as the positive and negative controls for the catalase test, respectively. The acid production was performed by using carbohydrate utilization such as d-glucose, d-fructose, lactose, mannitol, sorbitol, inulin and starch which were assayed by adding each of sugar in cystine tryptic agar (CTA) to a final concentration of 10 g/l (Tinrat et al. 2011).
In addition, species identification of the isolated strains was performed by 16S rDNA analysis. Briefly, overnight culture of isolated strains were transferred into TE buffer (10 mM Tris–HCl pH 8.0, 1 mM EDTA). The sample was boiled about 10–15 min and the supernatant was collected by centrifugation at 9500 rpm for 5–10 min. The 16S rDNA gene was amplified by using prokaryotic primers (5′-GCC TAA CAC ATGCAA GTC GA-3′: UFUL and 5′-CGT ATT ACC GCGGCT GCT GG-3′: URUL) (Phalakornkule and Tanasupawat 2006–2007; Hongpattarakere et al. 2008). The PCR reaction mixture consisted of 1 × PCR buffer (pH 8.8) for Taq DNA polymerase (Biolab), 2–5 μl of DNA template, 0.4 μM deoxynucleoside triphosphate (dNTPs), 0.4 μM of each primer and 1 unit of Taq polymerase in the final volume of 20 μl. The amplification program was as follows: initial denaturation at 95 °C for 5 min; 30 cycles of 95 °C for 5 min, 55 °C for 30 s, and 72 °C for 30 s; and a final extension step at 72 °C for 5 min. After amplification, PCR products were purified with ethanol precipitation and dissolved in Hi-Di Formamide before 16S rDNA sequencing by using a BigDye v.3.1 cycle sequencing kit (Applied Biosystems, California, USA) with the previous primers (Thompson et al. 1997). The nucleotide sequencing was determined with automated sequencing. Finally, the sequence was checked with in the National Center for Biotechnology Information (NCBI) with clearly known sequence by using BLAST program in GenBank (www.ncbi.nlm.nih.gov/blast).
Determination of antimicrobial activity
Antimicrobial activity of isolated strains was detected by agar spot test (Gu et al. 2008). The indicator strains were used in this study including Bacillus cereus ATCC 11778, Escherichia coli ATCC 25922, Staphylococcus aureus ATCC 25923, Enterococcus sp., Klebsiella pneumoniae, Shigella flexneri, Shigella sonnei, Salmonella typhi, Salmonella typhimurium ATCC 13311, Salmonella paratyphi A, Salmonella paratyphi B, Proteus vulgaris, Pseudomonas aeruginosa ATCC 9027, Salmonella enteritidis, Staphylococcus epidermidis, Lactobacillus plantarum ATCC 14917 and Lactobacillus salivarius MTC 1026. In brief, overnight isolated strains were spotted on BHI plates and incubated at 37 °C for 24 h. Then, the plates were overlaid with the culture of indicator strains and adjusted to yield approximately 1.0 × 108 CFU/ml in 0.75% BHI agar. Next, all tested plates were incubated at 37 °C for 24–48 h depending on indicator strains. Finally, inhibition zones around the spots were measured and presented as the average inhibition diameter in millimeters.
Assay of bacteriocin activity
The inhibitory activity of probiotic strains used various indicator strains; B. cereus ATCC 11778, E. coli ATCC 25922, S. aureus ATCC 25923, S. typhimurium ATCC 13311, P. aeruginosa ATCC 9027, L. plantarum ATCC 14917, L. casei ATCC 39392, L. sakei ATCC 15521, and L. salivarius MTC 1026. Inhibitory activity of probiotic strains was screened by agar well diffusion assay. Briefly, overnight culture of isolated strains were centrifuged at 9500 rpm for 20 min at 4 °C and the supernatant was concentrated up to fivefolds by speed vacuum at 4 °C. Then, catalase enzyme was added to concentrated supernatants before adjusted to pH 6.0 with 1 M NaOH and filtered through 0.2 μm pore size. The cultures of indicator strains were adjusted to yield approximately 1.0 × 108 CFU/ml in soft BHI agar (0.75%) before overlaid the base agar. Next, concentrated supernatants of isolated strains were dropped into punched wells before incubated at 37 °C for 1–2 day depending on indicator strains. The inhibition zones around the wells were measured and presented as the average inhibition diameter in millimeters (Tinrat et al. 2011).
Effect of temperature on bacterial survival
The isolated strains were cultivated in BHI broth and incubated at 37 °C for 24 h. The overnight cultures were adjusted to yield approximately 1.0 × 108 CFU/ml. Then, aliquot culture of isolated strains were transferred to BHI broth and incubated for 24 h at various temperatures; 37, 45, 55 and 65 °C. Finally, the survival cell numbers were detected at 0 and 24 h by colony plate count method compared with the starting point.
The in vitro survival in simulated gastric juice and small intestinal juice
An aliquot of each washed cell suspension was put in simulated gastric juices at varies pH; 2.0, 3.0, 4.0, 6.0 and 7.0 and then vigorously mixed for 10 s before incubated at 37 °C. After 0, 90 and 180 min, each culture was taken for determining the number of survival cells by plate count method compared with the starting point.
Moreover, the cell suspension was put in 0.15, 0.3 and 1.0% Oxagall bile salt juices, pH 8.0 and without Oxagall bile salt juices as the control and then vortex vigorously for 10 s before incubated at 37 °C. At 0, 90 and 180 min, each culture was taken for determining the number of survival cells by plate count method compared with the starting point (Kim and Austin 2008; Allameh et al. 2012).
Antibiotic susceptibility test
The antibiotic susceptibility test was studied by agar diffusion method which was used 3 groups of antibiotics (Xu et al. 2008). In group A, 5 antibiotics (penicillin, ampicillin piperacillin, ceftazidime, and bacitracin) were grouped by inhibiting cell wall synthesis. In group B, 6 antibiotic agents (gentamicin, streptomycin, tetracycline, erythromycin chloramphenicol and amikacin) were grouped by inhibiting protein synthesis. In group C, 3 antibiotic agents (nalidixic acid, ciprofloxacin and norfloxacin) were grouped by inhibiting nucleic acid synthesis. Then, overnight culture of the tested strain was adjusted to yield approximately 1.0 × 108 CFU/ml. After that, the suspension of tested strain was cultivated onto the surface of BHI plates. After about 10–15 min, sterile antibiotic discs were placed on the surface of these plates before incubated at 37 °C for 24–48 h. Finally, the inhibition zones around each of the antibiotic discs were determined the average inhibition diameter in millimeters.
Adhesion assay
Caco-2 cells were obtained from the American Type Culture Collection (ATCC®HTB-37™, USA). The cells were cultured in Dulbecco’s Modified Eagle’s minimal medium (DMEM) (JR Sciencific, CA) supplemented with 10% (v/v) heat-inactivated (30 min at 56 °C) fetal calf serum (JR Sciencific, CA) and 1% 100 U/ml penicillin and 100 μg/ml streptomycin (JR Sciencific, CA) at 37 °C in an atmosphere of 5% CO2/95% air. For adhesion assay, Caco-2 cells were washed twice with sterile PBS (pH 7.0) and transferred with 0.25% trypsin (JR Sciencific, CA) into a 24-well tissue culture plates (USA). Caco-2 cells were seeded at concentration of 105 cells/well and changed cell culture medium every other day. After 14-day incubation at previous condition, the cell medium culture was replaced by fresh non-supplement DMEM for 60 min before the adhesion assay and adhesion-inhibition assay (Tinrat et al. 2011).
Caco-2 monolayers in a 24-well tissue culture plate were washed twice with sterile PBS (pH 7.0). The bacterial suspension concentration of 109–1010 CFU/ml was added and coincubated with Caco-2 monolayers for 60 min (Moussavi and Adams 2010). Unattached bacteria were discharged by washing with sterile PBS (pH 7.0). Caco-2 cells and adherent bacteria were lysed and released by agitating with pipette in sterile PBS (pH 7.0). The number of viable adherent bacteria was determined by colony forming unit (CFU/ml) by plating method. L. plantarum ATCC 14917 as control was determined on MRS agar and Enterococcus faecalis MTC 1032 and pathogenic strains were determined on BHI agar.
Adhesion-inhibition assay
Various conditions were performed, including (1) Competition assay (Yu et al. 2011): E. faecalis MTC 1032 suspensions (109 CFU/ml) and pathogen suspension (109 CFU/ml) including E. coli ATCC 25922 and S. typhimurium ATCC 13311 were mixed in each well and coincubated with Caco-2 monolayers at 37 °C for 1 h, (2) Exclusion assay (Yu et al. 2011): E. faecalis MTC 1032 suspensions (109 CFU/ml) were first applied with Caco-2 monolayer in each well for 1 h and then pathogen suspensions (109 CFU/ml) were added in each well and then incubated for 1 h at 37 °C, (3) Displacement assay: pathogen suspensions (109 CFU/ml) were preincubated in each well for 1 h at 37 °C before adding E. faecalis MTC 1032 suspensions (109 CFU/ml) and incubated in previously described condition. L. plantarum ATCC14917 was used as the control strain. Caco-2 monolayers were washed with sterile PBS (pH 7.0) and were lysed by agitating with pipette in PBS (pH 7.0). The cell lysates were plated on selective medium to determine the number of viable bacterial cells by colony plate counting. MacConkey agar (Difco) was used for E. coli ATCC 25922, bismuth sulfite agar (Difco), xylose lysine deoxycolate agar (Difco) and billiant-green phenol red lactose agar (Difco) for S. typhimurium ATCC 13311 and MRS agar for Lactobacilli strains.
Results and discussion
Screening and identification of Enterococcus faecalis MTC 1032
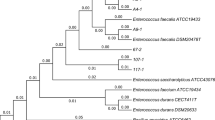
Approximately 60 isolated strains from children feces were selected to screen for potential probiotic strains. The result of antimicrobial activity screening assay showed that two out of sixty isolated strains could inhibit Salmonella spp. growth under aerobic condition. In this study, the isolated strain with the best inhibitory activity was selected to be identified and further evaluated for probiotic properties. This strain, so called MTC 1032, was oxidase and catalase-negative, Gram-positive, cocci shape. In addition, this strain could produce acids by fermenting d-glucose, d-fructose, lactose, mannitol, sorbitol, inulin and starch as carbon sources. It could only hydrolyze bile-esculin, but not red blood cells. This isolated strain was identified by PCR amplification with universal bacterial primers (UFUL and URUL) and the sequence of 16S rDNA was aligned with nucleotide–nucleotide BLAST (blastn) program of the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.hov). The result showed that the isolated strain had 99% homology with E. faecalis IMAU 10060 isolated from camel milk. Taken together, all phenotype characteristics suggested that this isolated strain could be E. faecalis.
Determination of antimicrobial activity and bacteriocin production
Inhibitory activity of the bacterial strain against 17 pathogenic strains was determined by agar spot test. The results showed that E. faecalis MTC 1032 could inhibit 10 out of 17 bacterial strains (Table 1). This inhibitory effect depended on many factors such as acid production and bacteriocin. This study also determined the effect of bacteriocin that has inhibitory effect against other strains. E. faecalis MTC 1032 supernatant could inhibit pathogens about 6 out of 9 strains (Table 2). The best inhibitory effect was against B. cereus ATCC 11778. L. plantarum ATCC 14917 was also tested as a probiotic control and it could broadly inhibit all pathogens.
Effect of temperature on bacterial survival
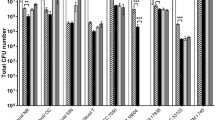
Heat-tolerant ability of probiotic bacteria is the basic requirement for applications in food industry. This property is necessary for probiotic product formulation and maintenance condition. It is of interest that the probiotic bacteria must be active while they are under human body temperature. Moreover, when probiotic bacteria are packed for commercial purpose, they must be stable under higher temperature condition. Therefore, the survival rate at various temperatures of the isolated strains (25, 37, 45 and 55 °C) were extensively studied (Fig. 1). The result showed that survival rate of the tested strain varied during incubation time. E. faecalis MTC 1032 could survive at 25, 37 and 45 °C. When compared with L. plantarum ATCC 14917, E. faecalis MTC 1032 could grow better than the control strain about 1 − log in all time periods. In addition, E. faecalis MTC 1032 grew at 55 °C for 8 h while the control strain could not.
Effect of temperature on the growth of E. faecalis MTC 1032 (a) and L. plantarum ATCC 14917 (b) after incubation at various temperature (25, 37, 45, and 55 °C). In bar graph, the turbidity of cells were showed As following; a: at 25 °C, b: at 37 °C, c: at 45 °C, and d: at 55 °C. And in Line graph, the number of survival cells (CFU/ml) were presented as following; filled triangle: at 25 °C, *at 37 °C, opened diamond: at 45 °C, and filled circle: at 55 °C. *Indicated statistically difference between groups as determined by paired t test
Effect of simulated gastric juices and simulated small intestinal transit on viability
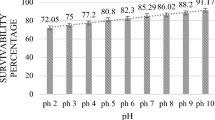
Another good feature of probiotics is ability to resist unsuitable conditions in the gastrointestinal tract. The survival rate during passage through the human gastrointestinal tract is a major challenge for effective delivery of these beneficial bacteria due to their fastidious characteristics in nature. Probiotic strain must survive during transition through the upper part of the gastrointestinal tract in order to act in the lower gastrointestinal section. Therefore, the isolated strain was cultured to determine the effect of pH on survival. The results showed that E. faecalis MTC 1032 was more sensitive at pH 2.0–3.0 than other conditions. The cell numbers were reduced about 5 − 6log CFU when comparing with the initial numbers. At pH 4.0 and 6.0, this strain could survive more than the control strain about 2 − log CFU/ml. It was demonstrated that the most favorable pH of this isolated strain was 7.0 (Fig. 2).
Effect of artificial gastric juices on the growth of E. faecalis MTC 1032 (a) and L. plantarum ATCC 14917 (b) during 180 min of various pH levels (pH 2.0, pH 3.0, pH 4.0, pH 6.0, and pH 7.0). In bar graph, the turbidity of cells were shown as following; A: at pH 2.0, B: at pH 3.0, c: at pH 4.0, d: at pH 6.0 and e: at pH 7.0. And in line graph, the number of survival cells (CFU/ml) were presented as following; opened triangle: at pH 2.0, filled diamond: at pH 3.0, *at pH 4.0, diamond circle: at pH 6.0 and diamond square: at pH 7.0. *Indicated statistically difference between groups as determined by paired t test
The effects of simulated small intestinal juices (0, 0.15, 0.3 and 1.0% bile salts) on the viability of E. faecalis MTC 1032 were also performed in this study. The result showed that this strain could survive at all tested bile salt concentrations more than the control strain about 1 − log CFU. The most favorable bile salt concentration of the isolated strain was at 0.3% (Fig. 3).
Effect of small intestinal juice on the growth of E. faecalis MTC 1032 (A) and L. plantarum ATCC 14917 (B) during 180 min of various bile salt concentrations (0, 0.15, 0.30, and 1.0% bile salt). In bar graph, the turbidity of cells were showed as following; a: at 0% bile salt, b: at 0.15% bile salt, c: at 0.30% bile salt, and d: 1.0% at bile salt. And in line graph, the number of survival cells (CFU/ml) were presented as following; filled triangle: at 0% bile salt, *at 0.15% bile salt, filled diamond: at 0.30% bile salt, and opened circle: at 1.0% bile salt. *Indicated statistically difference between groups as determined by paired t test
Antibiotic susceptibility test
One of the important properties of the probiotic bacteria is the ability to resist antibiotic drugs after receiving antimicrobial treatment in order to remain and being useful in the host. Therefore, antibiotic susceptibility test was performed by disc diffusion method. The results showed that piperacillin was the most sensitive antibiotic agent with 27-mm clear zone diameter. According to the CLSI guideline, E. faecalis MTC 1032 was found resistant to 3 antibiotics, including ampicillin, ciprofloxacin, and norfloxacin, respectively. In contrast, E. faecalis MTC 1032 was sensitive to chloramphenicol and tetracycline.
Inhibition of the pathogenic adhesion by E. faecalis MTC 1032 on human epithelial colorectal adenocarcinoma cells (Caco-2)
Adhesion of the probiotic bacteria to intestinal mucosa is an essential feature for colonization in the lower part of gastrointestinal tract. Therefore, adherence assay was carried out using the human intestinal cell line Caco-2, a well characterized cellular lineage established from human colonic adenocarcinoma. The results indicated that E. faecalis MTC 1032 could adhere on Caco-2 monolayers (Fig. 4). The adhesion cell numbers of this strain was 6.29 × 109 CFU/ml from the initial cell number of 2.62 × 1010 CFU/ml. In addition, E. faecalis MTC 1032 was determined inhibitory effect against pathogen adhesion on Caco-2 cells for antagonistic activity. The inhibition levels depended on pathogenic bacterial strains. the result showed that E. faecalis MTC 1032 could inhibit invasion of pathogens. The numbers of E. coli ATCC 25922 and S. typhimurium ATCC 13311 was significantly decreased while the adhesion cell number of E. faecalis MTC 1032 was almost unchanged and did not differ to the control strain. Moreover, it was demonstrated that E. faecalis MTC 1032 could inhibit the adhesion of E. coli ATCC 25922 more than S. typhimurium ATCC 13311 about 1 − log CFU/ml.
a Competition assay: effective adhesion of the E. faecalis MTC 1032 on Caco-2 cells comparing with L. plantarium ATCC 14917. In bar graph, the number of survival bacterial cells on adhesion-inhibition cells of E. faecalis MTC 1032 on E. coli ATCC 25922 were shown as following; A: the cell number of L. plantarum MTC 1026, B: the cell number of E. coli ATCC 25922 (coincubated with L. plantarium ATCC 14917), C: the cell number of E. faecalis MTC 1032 and D; the cell number of E. coli ATCC 25922 (coincubated with E. faecalis MTC 1032). And in line graph, the number of survival bacterial cells L. plantarium ATCC 14917 on S. typhimurium ATCC 13311 were presented as following; filled triangle: the cell number of L. plantarum ATCC 14917, opened diamond: adhesion rate of S. typhimurium ATCC 1311 (coincubated with L. plantarium ATCC 14917), x: E. faecalis MTC 1032 and ο: S. typhimurium ATCC 13311 (coincubated with E. faecalis MTC 1032). b Exclusion assay: effective adhesion of the E. faecalis MTC 1032 on Caco-2 cells comparing with L. plantarium ATCC 14917. In bar graph, the number of survival bacterial cells on adhesion-inhibition cells of E. faecalis MTC 1032 on E. coli ATCC 25922 were shown as following; A: the cell number of L. plantarum MTC 1026, B: the cell number of E. coli ATCC 25922 (coincubated with L. plantarium ATCC 14917), C: the cell number of E. faecalis MTC 1032 and D; the cell number E. coli ATCC 25922 (coincubated with E. faecalis MTC 1032). And in line graph, the number of survival bacterial cells L. plantarium ATCC on S. typhimurium ATCC 13311 were presented as following; filled triangle: the cell number of L. plantarum ATCC 14917, opened diamond: adhesion rate of S. typhimurium ATCC 1311 (coincubated with L. plantarium ATCC 14917), x: E. faecalis MTC 1032 and ο: S. typhimurium ATCC 13311(coincubated with E. faecalis MTC 1032). c Displacement assay: effective adhesion of the E. faecalis MTC 1032 on Caco-2 cells comparing with L. plantarium ATCC 14917. In bar graph, the number of survival bacterial cells on adhesion-inhibition cells of E.faecalis MTC 1032 on E. coli ATCC 25922 were shown as following; A: the cell number of L. plantarum MTC 1026, B: the cell number of E. coli ATCC 25922 (coincubated with L. plantarium ATCC 14917), C: the cell number of E. faecalis MTC 1032 and D; the cell number of E. coli ATCC 25922 (coincubated with E. faecalis MTC 1032). And in line graph, the number of survival bacterial cells L. plantarium ATCC 14917 on S. typhimurium ATCC 13311 were presented as following; filled triangle: the cell number of L. plantarum ATCC 14917, opened diamond: adhesion rate of S. typhimurium ATCC 1311 (coincubated with L. plantarium ATCC 14917),: E. faecalis MTC 1032 and ο: S. typhimurium ATCC 13311(coincubated with E. faecalis MTC 1032). *Indicated statistically difference between groups as determined by paired t test
Discussion
Probiotics are nonpathogenic microorganisms of human origin, when administered in enough amounts, confer a health benefit on the host and enable to prevent or improve health some diseases (Fuller 1986). Probiotics may be natural temporary composition of the resident intestinal microflora (Fuller 1989). Nowadays, probiotic strains are used not only in fermented foods but also in pharmaceutical preparations which must be recognized for their “generally recognized as safe (GRAS)” pattern (Fuller 1989). A major development in food industry was focused on using the probiotic strains for enhancing the value product (Jankovic et al. 2010). There is growing scientific evidence to support the concept that the maintenance of healthy gut microflora may provide protection against gastrointestinal disorders including gastrointestinal infections and inflammatory bowel diseases (Kligler and Cohrssen 2008). Although there are several probiotic strains on the market around the world, the need to find new strain probiotic bacteria are still ongoing. The probiotics include mostly lactobacilli (Sanders 2003) but many suitable probiotic candidates have also occurred among enterococci (Lauková et al. 2008). It has been reported that Enterococcus spp. can produce antimicrobial substances with an antagonistic effect against various pathogenic bacteria (Vahjen et al. 2002). Many of them also exert beneficially influence on host. In contrast to their positive properties, some enterococcal strains are suspected to have pathogenic properties for humans. Additionally, they are also resistant to several antibiotics. However, they have been observed among traits isolated from clinical and food sources. Pathogenic potential could only be associated with clinical strains, not food strains. Enterococci seem to possess both advantages and disadvantages. Therefore, it is an important to clearly study properties of Enterococci in dairy products and animal feed (Strompfová and Lauková 2007).
It has been reported that human normal flora could be an excellent source of probiotic microorganisms (Sathyabama et al. 2012). Therefore, the microorganisms from children fecal microbiota were selected for further characterization in this study. The selected bacteria was isolated and identified by phenotypic and genotypic methods for further characterization as a probiotic. It was found that E. faecalis MTC 1032 possessed the best characteristics of probiotics. Enterococci, particularly E. faecalis have been involved in the reduction or prevention of gastrointestinal tract infections (Franz et al. 2011). Enterococci belong to the group of LAB, known to produce lactic acid as the end product of sugar fermentation, and antimicrobials, which are active against many pathogens (Campos et al. 2006). Many strains of E. faecalis produce bacteriocins and have been reported on probiotic potential (Al Atya et al. 2015; Belguesmia et al. 2011). E. faecalis MTC 1032 was evaluated the characteristics as the main selective points for a potential probiotic strain. E. faecalis MTC 1032 could grow well up to 45–55 °C which was the optimum condition used in food processing. It had a maximum survivability at pH of 7 and could survive at pH 2.0–3.0 of simulated gastric juice and 1% bile salt of simulated small intestinal juice. Our data demonstrated that E. faecalis MTC 1032 had antimicrobial activity against many pathogenic strains. Moreover, this strain has reliable safety, because it was isolated from human and did not destroy red blood cells. E. faecalis MTC 1032 could adhere and inhibit the adhesion of pathogens including E. coli ATCC 25922 and S. typhimurium ATCC 13311 on Caco-2 monolayers. The similar results were reported previously (Jin et al. 2000; Lauková et al. 2008). Jin et al.’s reported (2000) that E. faecium 18C23 was able to inhibit the adhesion of E. coli K88ac about > 90% at a seed concentration of 109 CFU/ml to the small intestine mucus of piglets. Inhibitory substances such as lactic acid and bacteriocin were shown to be related with other studies (Carlos et al. 2009). It has been reported that E. faecium BFE 900 from black olives could produce a bacteriocin as called enterocin 900. It was antagonistic towards many microorganisms such as Lactobacillus sake, Clostridium butyricum and Listeria monocytogenes (Franz et al. 1996). The experimental report in 2007 (Poeta et al. 2007) indicated that 25 of the 140 enterococci displayed antimicrobial activity against L. monocytogenes and 33 isolates strains against Pediococcus pentosaceus, and different enterococcal species. In 2010, antibacterial activity of E. faecium IJ-06, E. faecium IJ-21, and E. faecium IJ-31 and E. faecalis IJ-11 as bacteriocin against Listeria monocytogenes ATCC 19115, and Bacillus subtilis ATCC 27142 (Poeta et al. 2007). Therefore, considering all presented data, E. faecalis MTC 1032 was an appropriate strain to be a candidate as a useful probiotic strain for applying in food products.
References
Al Atya AK, Drider-Hadiouche K, Ravallec R, Silvain A, Vachee A, Drider D (2015) Probiotic potential of Enterococcus faecalis strains isolated from meconium. Front Microbiol 6:227
Alam S, Mushtaq M (2009) Antibiotic associated diarrhea in children. Indian Pediatr 46:491–496
Allameh SK, Daud H, Yusoff FM, Saad CR, Ideris A (2012) Isolation, Identification and characterization of Leuconostoc mesenteroides as anew probiotic from intestine of snakehead fish (Channa striatus). Afr J Biotechnol 11:3810–3816
Belguesmia Y, Madi A, Sperandio D, Merieau A, Feuilloley M, Prévost H, Drider D, Connil N (2011) Growing insights into the safety of bacteriocins: the case of enterocin S37. Res Microbiol 162(2):159–163
Campos CA, Rodriguez O, Calo-Mata P, Prado M, Barros-Velazquez J (2006) Preliminary characterization of bacteriocins from Lactococcus lactis, Enterococcus faecium and Enterococcus mundtii strains solated from turbot (Psetta maxima). Food Res Int 39:356–364
Canani RB, Cirillo P, Terrin G et al (2007) Probiotics for treatment of acute diarrhoea in children: randomised clinical trial of five different preparations. BMJ 335(7615):340
Carlos AR, Santos J, Semedo-Lemsaddek T, Barreto-Crespo MT, Ten Reiro R (2009) Enterococci from artisanal dairy products show high levels of adaptability. Int J Food Microbiol 129:194–199
Casey PG, Casey GD, Gardiner GE et al (2004) Isolation and characterization of anti-Salmonella lactic acid bacteria from the porcine gastrointestinal tract. Lett Appl Microbiol 39:431–438
Chaveerach P, Lipman LJ, van Knapen F (2004) Antagonistic activities of several bacteria on in vitro growth of 10 strains of Campylobacter jejuni/coli. Int J Food Microbiol 90:43–50
Del Giudice MM, Leonardi S, Maiello N, Brunese FP (2010) Food allergy and probiotics in childhood. J Clin Gastroenterol 44:22–25
Floch MH (2010) Probiotic therapy for ulcerative colitis. J Clin Gastroenterol 44:237–238
Franz CM, Huch M, Abriouel H et al (2011) Enterococci as probiotics and their implications in food safety. Int J Food Microbiol 151:125–140
Franz CM, Schillinger U, Holzapfel WH (1996) Production and characterization of enterocin 900, a bacteriocin produced by Enterococcus faecium BFE 900 from black olives. Int J Food Microbiol 29:255–270
Franz CM, Stiles ME, Schleifer KH, Holzapfel WH (2003) Enterococci in foods—a conundrum for food safety. Int J Food Microbiol 88:105–122
Fuller R (1986) Probiotics. Soc Appl Bacteriol Symp Ser 15:1–7
Fuller R (1989) Probiotics in man and animals. J Appl Bacteriol 66:365–378
Gu RX, Yang ZQ, Li ZH et al (2008) Probiotic properties of lactic acid bacteria isolated from stool samples of longevous people in regions of Hotan, Xinjiang and Bama, Guangxi, China. Anaerobe 14:313–317
Guo XH, Kim JM, Nam HM et al (2010) Screening lactic acid bacteria fromswine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 16:321–326
Hlivak P, Odraska J, Ferencik M et al (2005) One-year application of probiotic strain Enterococcus faecium M-74 decreases serum cholesterol levels. Bratisl Lek Listy 106:67–72
Hongpattarakere T, Chanthachum S, Buntin N (2008) Screening of lactic acid bacteria from gastrointestinal tracts of marine fish for their potential use as probiotics. Songklanakarin J Sci Technol 30:141–148
Jankovic I, Sybesma W, Phothirath P et al (2010) Application of probiotics in food products—challenges and new approaches. Curr Opin Biotechnol 21:175–181
Jin LZ, Marquardt RR, Zhao X (2000) A strain of Enterococcus faecium (18C23) inhibits adhesion of enterotoxigenic Escherichia coli K88 to porcine small intestine mucus. Appl Environ Microbiol 66:4200–4204
Kaewsrichan J, Chandarajoti K, Kaewnopparat S et al (2007) Evaluation of lactobacilli containing suppository formulation for probiotic use. Mahidol Univ J Pharm Sci 34:1–8
Kim DH, Austin B (2008) Characterization of probiotic carnobacteria isolated from rainbow trout (Oncorhynchus mykiss) intestine. Lett Appl Microbiol 47:141–147
Kim PI, Jung MY, Chang YH et al (2007) Probiotic properties of Lactobacillus and Bifidobacterium strains isolated from porcine gastrointestinal tract. Appl Microbiol Biotechnol 74:1103–1111
Kligler B, Cohrssen A (2008) Probiotics. Am Fam Physician 78:1073–1078
Koebnick C, Wagner I, Leitzmann P et al (2003) Probiotic beverage containing Lactobacillus casei Shirota improves gastrointestinal symptoms in patients with chronic constipation. Can J Gastroenterol 17:655–659
Kramer J (1977) Inhibition of different serotypes of Listeria monocytogenes by enterocins in solid and liquid media. J Med Microbiol 10:367–372
Krammer HJ, Schlieger F, Harder H et al (2005) Probiotics as therapeutic agents in irritable bowel syndrome. Z Gastroenterol 43:467–471
Kühn I, Iversen A, Burman LG et al (2003) Comparison of Enterococcal populations in animals, humans, and the environment—a European study. Int J Food Microbiol 88:133–145
Lauková A, Simonová M, Strompfová V et al (2008) Potential of enterococci isolated from horses. Anaerobe 14:234–236
Linaje R, Coloma MD, Perez-Martínez G, Zuniga M (2004) Characterization of faecal enterococci from rabbits for the selection of probiotic strains. J Appl Microbiol 96:761–771
Lye HS, Kuan CY, Ewe JA et al (2009) The improvement of hypertension by probiotics: effects on cholesterol, diabetes, renin, and phytoestrogens. Int J Mol Sci 109:3755–3775
Miyazaki Y, Kamiya S, Hanawa T et al (2010) Effect of probiotic bacterial strains of Lactobacillus, Bifidobacterium, and Enterococcus on enteroaggregative Escherichia coli. J Infect Chemother 16:10–18
Moussavi M, Adams MC (2010) An in vitro study on bacterial growth interactions and intestinal epithelial cell adhesion characteristics of probiotic combinations. Curr Microbiol 60:327–335
Mukai T, Asasaka T, Sato E et al (2002) Inhibition of binding of Helicobacter pylori to the glycolipid receptors by probiotic Lactobacillus reuteri. FEMS Immunol Med Microbiol 32:105–110
Nagpal R, Kumar A, Kumar M et al (2012) Probiotics, their health benefits and applications for developing healthier foods: a review. FEMS Microbiol Lett 334:1–15
Noguchi S, Okada S, Sugiyama H et al (2012) Lactobacillus plantarum NRIC1832 enhances IL-10 production from CD4 + T cells in vitro. Biosci Biotechnol Biochem 76:1925–1931
Osterlund P, Ruotsalainen T, Korpela R et al (2007) Lactobacillus supplementation for diarrhoea related to chemotherapy of colorectal cancer: a randomised study. Br J Cancer 97:1028–1034
Phalakornkule C, Tanasupawat S (2006–2007) Characterization of lactic acid bacteria from traditional Thai fermented sausages. J Cult Collect 5:46–57
Poeta P, Costa D, Rojo-Bezares B et al (2007) Detection of antimicrobial activities and bacteriocin structural genes in faecal enterococci of wild animals. Microbiol Res 162:257–263
Rodríguez E, Arqués JL, Rodríguez R et al (2012) Antimicrobial properties of probiotic strains isolated from breast-fed infants. J Funct Foods 4:542–551
Sanders ME (2000) Considerations for use of probiotic bacteria to modulate human health. J Nutr 130:384–390
Sanders ME (2003) Probiotics: considerations for human health. Nutr Rev 61:91–99
Sathyabama S, Vijayabharathi R, Bruntha Devi P et al (2012) Screening for probiotic properties of strains isolated from feces of various human groups. J Microbiol 50: 603–12
Strompfová V, Lauková A (2007). In vitro study on bacteriocin of Enterococci associated with chickens. Anaer 13:228–237
Thompson JD, Gibson TJ, Plewniak F et al (1997) The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tinrat S, Saraya S, Chomnawang MT (2011) Isolation and characterization of Lactobacillus salivarius MTC 1026 as a potential probiotic. J Gen Appl Microbiol 57(6):365–378
Vahjen W, Jadamus A, Simon O (2002) Influence of a probiotic Enterococcus faecium strain on selected bacterial groups in the small intestine of growing turkey poults. Arch Tierernahr 56:419–429
Xu HY, Tian WH, Wan CX et al (2008) Antagonistic potential against pathogenic microorganisms and hydrogen peroxide production of indigenous lactobacilli isolated from vagina of Chinese pregnant women. Biomed Environ Sci 21:365–371
Yu Q, Wang Z, Yang Q (2011) Ability of Lactobacillus to inhibit enteric pathogenic bacteria adhesion on Caco-2 cells. World J Microbiol Biotechnol 27:881–886
Acknowledgements
I would like to thank the Office of the Higher Education Commission, Thailand for supporting the fund under the program of Scholarships for Frontier Research Network for the join Ph.D. Program and Thailand Toray Science Foundation. We would like to extend our thanks to the Thailand Research Fund and Faculty of Pharmacy, Mahidol University (IRG5780007) for financial support on publication. I also thank staffs of the Department of Microbiology, Faculty of Pharmacy, Mahidol University for clarifying several points in this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tinrat, S., Khuntayaporn, P., Thirapanmethee, K. et al. In vitro assessment of Enterococcus faecalis MTC 1032 as the potential probiotic in food supplements. J Food Sci Technol 55, 2384–2394 (2018). https://doi.org/10.1007/s13197-018-3155-5
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13197-018-3155-5