Abstract
Cross distance minimization algorithm (CDMA) is an iterative method for designing a hard margin linear SVM based on the nearest point pair between the convex hulls of two linearly separable data sets. In this paper, we propose a new version of CDMA with clear explanation of its linear time complexity. Using kernel function and quadratic cost, we extend the new CDMA to its kernel version, namely, the cross kernel distance minimization algorithm (CKDMA), which has the requirement of linear memory storage and the advantages over the CDMA including: (1) it is applicable in the non-linear case; (2) it allows violations to classify non-separable data sets. In terms of testing accuracy, training time, and number of support vectors, experimental results show that the CKDMA is very competitive with some well-known and powerful SVM methods such as nearest point algorithm (NPA), kernel Schlesinger-Kozinec (KSK) algorithm and sequential minimal optimization (SMO) algorithm implemented in LIBSVM2.9.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Classification is one of the most widely studied problems within the field of machine learning. As one of the commonest tasks in supervised learning, it is the process of finding a model or function that learns a mapping between the set of observations and their respective classes [1, 2]. As a successful classifier, support vector machines (SVMs) have attracted much interest in theory and practice [3–6].
Respectively, let \(X = \{x_i: i \in I\}\) and \(Y = \{y_j: j \in J\}\) denote two classes of data points in \(\mathbf{R}^n\) (\(\mathbf{R}\) stands for the set of all real numbers, n for dimension), and I and J are the corresponding sets of indices. If X and Y are linearly separable, the task of a SVM is to find the maximal margin hyperplane,
where (\(w^*\), \(b^*\)) is the unique solution of the following quadratic model:
It has been shown that \(M = 2/||w||\) is the maximal margin, and (\(w^*\), \(b^*\)) could be computed from the nearest point pair (\(x^*\), \(y^*\)) between the two convex hulls CH(X) and CH(Y) [7, 8],
where (\(x^*\), \(y^*\)) is the solution of the following nearest point problem (NPP),
Cross distance minimization algorithm (CDMA) [9] is an iterative algorithm for computing a hard margin SVM via the nearest point pair between CH(X) and CH(Y). The CDMA is simple to implement with fast convergence, but it works only on linearly separable data. In this paper, we first give a new version of the CDMA with clear explanation of its linear time complexity. Then we extend the new CDMA to solving the non-linear and non-separable problems, in a similar way to generalize the S-K algorithm [10].
This paper is organized as follows. Section 2 describes some related work to the CDMA. Section 3 presents a new version of the CDMA and introduces its kernel version. Section 4 provides experimental results and Sect. 5 concludes the paper.
2 Related work
2.1 Sequential minimal optimization algorithm
The basic idea of sequential minimal optimization (SMO) algorithm is to obtain an analytical solution for subproblems resulting from the working set decomposition [11]. The performance of SMO could be further improved by the maximal violating pair working set selection [12, 13]. LIBSVM [14] is a well-known SMO-type implementation, which solves a linear cost soft margin SVM below:
In Eq. (4), \(\xi _i\) and \(\zeta _j\) denote the slack variables corresponding to two classes.
2.2 Nearest point algorithm
The most similar approach to our work is the nearest point algorithm (NPA) by Keerthi et al. [7], which combines the Gilbert’s algorithm [15] and the Mitchell algorithm [16] to find the nearest points between the two convex hulls CH(X) and CH(Y). In order to deal with the non-linear and non-separable cases, the NPA solves a new nearest point problem that is equivalent to the following quadratic cost soft margin SVM [17] (QCSM-SVM),
Note that negative values of \({\xi _i}\) or \({\zeta _j}\) cannot occur at the optimal solution of QCSM-SVM because we may reset \(\xi _i = 0\) (\(\zeta _j = 0\)) to make the solution still feasible and also strictly decrease the total cost if \(\xi _i < 0\) (\(\zeta _j < 0\)). Thus, there is no need to include nonnegativity constraints on \(\xi _i\) and \(\zeta _j\).With a simple transformation, the QCSM-SVM model can be converted into an instance of hard margin SVM (Eq. 2) with the dimensionality of \(n + |I| + |J|\) by mapping \({x_i} \in X\) to \({x_i}^\prime \in X^\prime\) and \({y_j} \in Y\) to \(y_j^\prime \in Y^\prime\) as follows:
where \({e_k}\) denotes the \(n + \left| I \right| + \left| J \right|\) dimensional vector in which the kth component is one and all other components are zero. Accordingly, \(w'\) and \(b'\) can be computed as:
The kernel function transformed is given by
where K denotes the original kernel function, \({\delta _{i,j}}\) is one if \(i=j\) and zero otherwise. Thus, for any pair of \((x_i, y_j)\) the modified kernel function is easily computed.
2.3 Schlesinger-Kozinec algorithm
Another closely related approach is Schlesinger-Kozinec (SK) algorithm [10]. The original SK algorithm solves the NPP problem by computing an approximate point pair (\(x^*\), \(y^*\)) to construct a \(\varepsilon\)-optimal hyperplane \(w^* \cdot x + b^* = 0\), where \(w^* = x^* - y^*\) and \({b^*} = (\parallel {y^*}{\parallel ^2} - \parallel {x^*}{\parallel ^2})/2\). This algorithm starts from two arbitrary points \(x^* \in CH(X), y^* \in CH(Y)\) and stops when satisfying the \(\varepsilon\)-optimal criterion, namely,
where \({v_t} \in X \cup Y\), \(t = \mathop {\arg \min }\limits _{i \in I \cup J} m({v_i})\), and
The SK algorithm has been generalized to its kernel version (KSK) [10]. It should be noted that in the SK algorithm, if \({v_t} = {x_t}(t \in I)\) violates the \(\varepsilon\)-optimal criterion, \(x^*\) will be updated by the Kozinec rule \({x^{new}} = (1 - \lambda ){x^*} + \lambda {x_t}\), with \(y^*\) fixed and \(\lambda\) computed for minimizing the distance between \(x^{new}\) and \(y^*\). Otherwise, if \({v_t} = {y_t}(t \in J)\) violates the \(\varepsilon\)-optimal criterion, \(y^*\) will be updated by the Kozinec rule \({y^{new}} = (1 - \mu ){y^*} + \mu {y_t}\), with \(x^*\) fixed and \(\mu\) computed for minimizing the distance between \(y^{new}\) and \(x^*\). If \(\parallel {x^*} - {y^*}\parallel - m({v_t}) < \varepsilon\) holds, \({w^*} = {x^*} - {y^*}\) and \({b^*} = (\parallel {y^*}{\parallel ^2} - \parallel {x^*}{\parallel ^2})/2\) gives an \(\varepsilon\)-optimal solution, which coincides with the optimal hyperplane for \(\varepsilon = 0\).
3 Cross kernel distance minimization algorithm
3.1 Cross distance minimization algorithm
Let d stand for the Euclid distance function. In this subsection, we will depict a new version of the CDMA in Algorithm 2, where \(\varepsilon\) is called “precision parameter”, for controlling the convergence condition.
The original CDMA (OCDMA, see Algorithm 1) is designed to iteratively compute the nearest points of two convex hulls. If \((x^*, y^*)\) is a nearest point pair obtained by solving the NPP in (3), i.e., \(d(CH(X), CH(Y))=||x^*-y^*||\), then it can be proven that their perpendicular bisector is exactly the hard-margin SVM of X and Y [9].

The procedure of OCDMA includes two basic blocks. The steps 3–4 are to compute a nearest point pair \((x^*, y^*)\) between CH(X) and CH(Y). The steps 5–6 are to construct a SVM, \(f(x)=w^* \cdot x + b^*\).

Compared to the OCDMA, the new CDMA can reduce half an amount of distance computations needed in the case \(\lambda > 0\) by minimizing \(d(z, y^*)\) instead of \(d(x_2, y^*)\) and \(d(z, y^*)\) as well as \(d(x^*, z)\) instead of \(d(x^*, y_2)\) and \(d(x^*, z)\). The geometric meaning of z point is illustrated in Fig. 1. According to Theorem 5 in Ref. [9], if \(x_1 = x^*\) is not the nearest point from \(y^*\) to CH(X), there must exist another point \({x_2} \ne {x_1}\) in X such that \(d(z,{y^*}) < d({x^*},{y^*})\), where
The geometric meaning of point z: (a) if \(\lambda\) =1, then \(z=x_2\) is a point in X; (b) if \(0<\lambda <1\), then \(z=x_1+\lambda (x_2-x_1)\) is actually the vertical point from \(y^*\) to the line segment \(CH\{x_1, x_2\}\). If \(x_1\) is not the nearest point from \(y^*\) to CH(X), there must exist another point \(z=x_2\) or \(z=x_1+\lambda (x_2-x_1)\) such that \(d(z, y^*)<d(x^*, y^*)\)
Similarly, if \(y_1 = y^*\) is not the nearest point from \(x^*\) to CH(Y), there must exist another point \(y_2 \ne y_1\) in Y such that \(d({x^*},z) < d({x^*},{y^*})\), where
Therefore, if (\(x^*, y^*\)) is not the nearest point pair, the CDMA can always find a nearer one until it converges.
It has been shown that CDMA converges with a rough time complexity of \(O(I(\varepsilon ) \cdot (\left| X \right| + \left| Y \right| ))\), where \(I(\varepsilon )\) is guessed a constant as an open problem [9], representing the total number of running loops for CDMA to converge at \(\varepsilon\)-precision. In fact, \(I(\varepsilon )\) can be clearly bounded by a simple constant, namely, \(I(\varepsilon ) \le L/\varepsilon\), where L may be estimated as \(L = {\max _{x \in X,y \in Y}}\left\{ {\left\| {x - y} \right\| } \right\}\), because we cannot get a cross distance decrease less than \(\varepsilon\) in each running loop until satisfying the stopping condition for CDMA.
3.2 Cross kernel distance minimization algorithm
Based on the description mentioned above, now we generalize the CDMA to its kernel extension, namely, cross kernel distance minimization algorithm (CKDMA). Given a kernel function \(K({x_i},{y_j}) = \phi ({x_i}) \cdot \phi ({y_j})\), the objective of CKDMA is to minimize the kernel distance \(kd(\tilde{x},\tilde{y})\) between \(\tilde{x} \in CH\left( {\phi (X)} \right)\) and \(\tilde{y} \in CH\left( {\phi (Y)} \right)\),
For \({\tilde{x}^*} \in CH\left( {\phi (X)} \right)\) and \({\tilde{y}^*} \in CH\left( {\phi (Y)} \right)\), we can express them as,
If \({\tilde{x}^*}\) is not the nearest point from \({\tilde{y}^*}\) to \(CH\left( {\phi (X)} \right)\), we can find \({t_1} \in I\) to compute a new point \({\tilde{x}^{new}} = (1 - \lambda ){\tilde{x}^*} + \lambda {\tilde{x}_{{t_1}}}\), where \(\lambda = \min [1, (({\tilde{x}_{{t_1}}} - {\tilde{x}^*}) \cdot ({\tilde{y}^*} - {\tilde{x}^*}))/(({\tilde{x}_{{t_1}}} - {\tilde{x}^*}) \cdot ({\tilde{x}_{{t_1}}} - {\tilde{x}^*}))]\), such that \({\tilde{x}^{new}}\) is nearer to \({\tilde{y}^*}\) than \({\tilde{x}^*}\), i.e., \(\left\| {{{\tilde{x}}^{new}} - {{\tilde{y}}^*}} \right\| < \left\| {{{\tilde{x}}^*} - {{\tilde{y}}^*}} \right\|\).
Because \({\tilde{x}^{new}} = \sum \limits _{i \in I} {\alpha _i^{new}{{\tilde{x}}_i}} = (1 - \lambda )\sum \limits _{i \in I} {{\alpha _i}{{\tilde{x}}_i}} + \lambda {\tilde{x}_{{t_1}}} = \sum \limits _{i \in I} {[(1 - \lambda ){\alpha _i} + \lambda {\delta _{i,{t_1}}}]{{\tilde{x}}_i}}\), \({\tilde{x}^{new}}\) can be obtained by updating \({\alpha _i}\), namely,
where \({\delta _{i,j}}\) is one if \(i = j\) and zero otherwise.
Similarly, if \({\tilde{y}^*}\) is not the nearest point from \({\tilde{x}^*}\) to \(CH\left( {\phi (Y)} \right)\), we can also find a nearer point \({\tilde{y}^{new}} = (1 - \mu ){\tilde{y}^*} + \mu {\tilde{y}_{{t_2}}},{t_2} \in J\) with the adaptation rule below,
For convenience, we further introduce the following notations:
Using these notations to express the CDMA in terms of kernel inner product, it is easy to obtain the CKDMA described in Algorithm 2. Note that sqrt indicates the square root function.

In Step 6 of the CKDMA, \(A,C,{D_i},{F_j}\) are updated as follows:
And in Step 9, \(B, C, {E_i}, {G_j}\) are updated as follows:
Moreover, in Algorithm 2, \(d^*\) stands for the distance between the nearest points, and \(d^{new}\) for its updated value. If \(d^* - d^{new}\) is less than \(\varepsilon\), the CKDMA will return the result of a linear function in kernel space, namely, \(f(x) = {{\tilde{w}}^*} \cdot \phi (x) + {{\tilde{b}}^*}\), which can be further expressed as a kernel form below,
In addition, note that CKDMA can only solve separable problems in a kernel-induced feature space. After using Eq. (6) mentioned in Sect. 2.2, it can be directly applied to non-separable cases, for finding a generalized optimal hyperplane with the quadratic cost function.
Finally, it seems that the update rules of the CKDMA are very similar to that of the KSK algorithm [10]. However, there are at least two notable differences between them: (1) in each iteration the KSK algorithm updates only one of two points, either \(x^*\) or \(y^*\), whereas the CKDMA generally updates both of them; (2) Given \(x^*\) and \(y^*\), the two algorithms may pick different data points even in the same training set for updating the nearest pair. For example, suppose \(X = \{ {x_1},{x_2},{x_3}\}\) and \(Y = \{ {y_1}\}\). In Fig. 2a, the KSK algorithm will choose \(x_3 \in X\) to update \(x^*\) with \(x_{KSK}^{new} = (1 - \lambda ){x^*} + \lambda {x_3}\) . This is because
and \(m(x_3) < m(x_2)\) (see Sect. 2.3).
But from Fig. 2, the CKDMA will choose \(x_2 \in X\) to update \(x^*\) with \(x_{CKDMA}^{new} = (1 - \lambda ){x^*} + \lambda {x_2}\), because \(d({y^*},CH\{ {x^*},{x_2}\} ) < d({y^*},CH\{ {x^*},{x_3}\} )\) (see Sect. 3.1).
4 Experimental results
In this section, we conduct a series of numerical experiments to evaluate the classification performances of CKDMA on 17 data sets, which are listed with their code, size and dimensionality in Table 1. All these data sets are selected from the UCI repository [18], but Fourclass from THK96a [19], German.number and Heart from Statlog [20], Leukemia and Gisette from LIBSVM Data [21]. The experiments are divided into two parts. The first part compares the proposed method with KSK, NPA and LIBSVM2.9, and the second part shows the influence of precision parameter \(\varepsilon\) on CKDMA.
4.1 Comparison with other algorithms
We used 14 data sets in this experiment. The 14 data sets can be seen in Table 2. They are binary problems with features linearly scaled to [−1, 1]. Note that the “training + test” sets have been partitioned for the last four data sets, including mo1, mo2, mo3 and spl. For the sake of homogeneity in results’ presentation, we first merge the training and test sets of them. And then, we randomly split each of them into two halves for 10 times, one half for training, the other for testing. We report the average results on all the experiments, which were conducted with RBF kernel function on a Dell PC having I5-2400 3.10-GHz processors, 4-GB memory, and Windows 7.0 operation system. The two parameters, C and \(\gamma\) respectively from the candidate sets \(\{2^i|i=-4,-3,...,3,4\}\) and \(\{2^i|i=-7,-6,...,4,5\}\), were determined by 5-fold cross validation on the training sets. The precision parameter \(\varepsilon\) was chosen as \(10^{-5}\) for CKDMA, \(10^{-3}\) for KSK and NPA, and the default (i.e., \(10^{-3}\)) for LIBSVM 2.9. Note that, in experimental results, the bold values represent higher accuracies, less training time or less support vectors.
From Table 2, it can be seen that CKDMA obtains higher accuracies on six data sets than KSK, six data sets than NPA, and seven data sets than LIBSVM2.9, respectively. Even in the case of lower accuracies, CKDMA can achieve the same level of performance with other algorithms. Moreover, CKDMA generally takes less training time and produces less support vectors than KSK and NPA, but more than LIBSVM2.9.
In addition, we also conducted experiments on three large/high-dimensional data sets (i.e., a9a, leu and gis), with the results described in Table 3. Since it take too long cross validation time to choose C and \(\gamma\) for Libsvm 2.9, we directly set the default values, namely, \(C=1\) and \(\gamma =1/dim\) (0.00813 for a9a, 0.00014 for leu, and 0.00020 for gis). From Table 3, we can get that, on the selected large/high-dimensional data sets, the CKDMA is comparable to the other three classifiers.
Next, we further compare the statistical significance of CKDMA with KSK, NPA, LIBSVM. Referring to [22, 23], we use Bonferroni-Dunn test for comparison over all the selected data sets. Table 4 presents the average ranking of four algorithms, and Fig. 4 provides the corresponding critical difference (CD) diagram. The test results show there are no significant differences between CKDMA and the others.
4.2 Influence of precision parameter
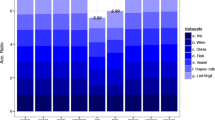
In this subsection we investigate influence of precision parameter \(\varepsilon\) on performance of CKDMA, with \(\varepsilon\) set from \(10^{-1}\) to \(10^{-6}\). We choose eight data sets of different dimensions (i.e., fou, mo1, hea, ger, bre, son, a9a and mus). For mo1 and a9a, we directly use the indicated training and testing sets for conducting experiments. For fou, hea, ger, bre, son and mus, we randomly split each of them into two halves only once, one half for training, the other for testing. The corresponding results are shown in Fig. 3, including testing accuracy, training time, and number of support vectors.
From Fig. 3, we can see that as \(\varepsilon\) gets smaller and smaller, testing accuracies generally tend to increase and become stable. In Fig. 3a, a vertical dashed line marks a good choice of \(10^{-5}\) for CKDMA because it gives rise to satisfactory performance with relatively reasonable training time (see Fig. 3b) and number of support vectors (see Fig. 3c).
5 Conclusions
In this paper, we have explained the linear time complexity for CDMA, and further proposed a new iterative algorithm, i.e., the cross kernel distance minimization algorithm (CKDMA), to train quadratic cost support vector machines using kernel functions. Over CDMA, CKDMA can take advantages to solve non-linear and non-separable problems, preserving the simplicity in geometric meaning. Moreover, it gives a novel way to understand the nature of support vector machines. Since the basic idea of CKDMA is to express the CDMA in kernel-induced feature space, it can be implemented in a similar way to KSK and NPA, but different from LIBSVM. Experimental results show that the CKDMA is very competitive with NPA, KSK and LIBSVM2.9 on the selected data sets. As future work, we would try to introduce large margin clustering [24] for rapid generation of supports vectors.
References
Webb D (2010) Efficient piecewise linear classifiers and applications. Ph. D. Dissertation, The Graduate School of Information Technology and Mathematical Sciences, University of Ballarat
Wang X, Ashfaq RAR, Fu A (2015) Fuzziness based sample categorization for classifier performance improvement. J Intell Fuzzy Syst 29(3):1185–1196
Ertekin S, Bottou L, Giles CL (2011) Nonconvex online support vector machines. IEEE Trans Pattern Anal Mach Intell 22(2):368–381
Wang X, He Q, Chen D, Yeung D (2005) A genetic algorithm for solving the inverse problem of support vector machines. Neurocomputing 68:225–238
Wang X, Lu S, Zhai J (2008) Fast fuzzy multi-category SVM based on support vector domain description. Int J Pattern Recognit Artif Intell 22(1):109–120
Xie J, Hone K, Xie W, Gao X, Shi Y, Liu X (2013) Extending twin support vector machine classifier for multi-category classification problems. Intell Data Anal 17(4):649–664
Keerthi SS, Shevade SK, Bhattacharyya C, Murthy KRK (2000) A fast iterative nearest point algorithm for support vector machine classifier design. IEEE Trans Neural Netw 11(1):124–136
Bennett KP, Bredensteiner EJ (1997) Geometry in learning. In: Geometry at work
Li Y, Liu B, Yang X, Fu Y, Li H (2011) Multiconlitron: a general piecewise linear classifier. IEEE Trans Neural Netw 22(2):267–289
Franc V, Hlaváč V (2003) An iterative algorithm learning the maximal margin classifier. Pattern Recogn 36(9):1985–1996
Platt JC (1999) Fast training of support vector machines using sequential minimal optimization. In: Sch\(\ddot{o}\)lkopf B, Burges C, Smola A (eds) Advances in kernel methods: support vector learning, MIT Press, Cambridge, p. 185–208
Keerthi SS, Shevade SK, Bhattacharyya C, Murthy KRK (2001) Improvements to Platt’s SMO algorithm for SVM classifier design. Neural Comput 13(3):637–649
Shevade SK, Keerthi SS, Bhattacharyya C, Murthy KRK (2000) Improvements to the SMO algorithm for SVM regression. IEEE Trans Neural Netw 11(5):1188–1194
Chang CC, Lin CJ (2001) Libsvm: a library for support vector machines (Online). http://www.csie.ntu.edu.tw/cjlin/libsvm. Accessed 10 June 2013
Gilbert EG (1966) Minimizing the quadratic form on a convex set. SIAM J Control 4:61–79
Mitchell BF, Dem’yanov VF, Malozemov VN (1974) Finding the point of a polyhedron closest to the origin. SIAM J Control 12(1):19–26
Friess TT, Harisson R (1998) Support vector neural networks: the kernel adatron with bias and softmargin, Univ. Sheffield, Dept. ACSE, Tech. Rep. ACSE-TR-752
Frank A, Asuncion A (2010) UCI machine learning repository (Online). http://archive.ics.uci.edu/ml. Accessed 23 Nov 2013
Ho TK, Kleinberg EM (1996) Building projectable classifiers of arbitrary complexity. In: Proceedings of the 13th international conference on pattern recognition, Vienna, p. 880–885
Brazdil P, Gama J (1999) Statlog datasets (Online). http://www.liacc.up.pt/ml/old/statlog/-datasets.html. Accessed 25 Apr 2014
Lin CJ Libsvm data (Online). https://www.csie.ntu.edu.tw/cjlin/libsvmtools/datasets/. Accessed 11 Aug 2015
Demšar J (2006) Statistical comparisons of classifiers over multiple data sets. J Mach Learn Res 7:1–30
Garca S, Herrera F (2009) An extenson on ’statistical comparisons of classifiers over multiple data sets’ for all pairwise comparisons. J Mach Learn Res 9:2677–2694
Xu L, Hu Q, Hung E, Chen B, Xu T, Liao C (2015) Large margin clustering on uncertain data by considering probability distribution similarity. Neurocomputing 158:81–89
Acknowledgments
This work was supported in part by the National Science Foundation of China under Grant 61175004, the Beijing Natural Science Foundation under Grant 4112009, the Program of Science Development of Beijing Municipal Education Commission under Program KM201010005012, and the Specialized Research Fund for the Doctoral Program of Higher Education of China under Grant 20121103110029.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Leng, Q. & Fu, Y. Cross kernel distance minimization for designing support vector machines. Int. J. Mach. Learn. & Cyber. 8, 1585–1593 (2017). https://doi.org/10.1007/s13042-016-0529-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13042-016-0529-8