Abstract
The environmental impact of chlorinated pesticides, including endosulfan, is not only caused by their persistency in the ecosystem but also from their toxic effects on off-target living organisms. In this study, three different strains of microorganisms, namely Afipia genosp, Sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum that are capable of biodegrading endosulfan at low concentrations (100 µg/L) from a tea cultivation field were reported. The isolated microbial consortium biodegraded 59% of the total endosulfan (63% α-endosulfan, 57% β-endosulfan) at pH 6.5. The same consortium biodegraded 98% of the total endosulfan (96% of α-endosulfan, 97% of β-endosulfan) at pH 8.4. All endosulfan removal performances were observed for a period of 25 days and the experiments were conducted at 25 °C, which was a relatively lower temperature compared to other endosulfan biodegradation studies in the literature. Additional carbon source did not change the overall endosulfan removal. No endosulfan sulfate production was observed during the study.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Statement of Novelty
In this paper, we investigated biodegradation of endosulfan, a persistent, chlorinated pesticide by a three bacteria consortium, consists of Afipia genosp, Sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum for the first time in literature. We registered these microorganisms to Genbank with the ID numbers of KY827230, KY827094, and KY827087, respectively. We are also reporting the growth rates of the consortium, which we observed in the microcosms. These microorganisms may help to develop sustainable remediation methods for chlorinated compounds. In this paper we investigated the microbial fate of endosulfan at low concentrations (100 mg/L) at room temperature (20 °C to 25 °C), at two different pH values (6.5 and 8.4) in water. We defined experimental conditions to mimic groundwater towards an efficient in-situ or on-site bioremediation. Alkaline conditions are resulted in better endosulfan removal. We also showed that isolation and enrichment of endosulfan degraders without sulfur prevents production of more toxic by-product endosulfan sulfate.
Introduction
Endosulfan is a chlorinated pesticide and a known endocrine disruptor with a wide spectrum of application in agricultural production. The effects of endosulfan on organisms have been investigated since the 1970s and from the beginning of 2007 the use of endosulfan has been gradually banned in several countries because of its adverse effects on human health and environment [1]. The presence of endosulfan in the environment has the potential to change the native microbial population distribution [2]. The bioaccumulation of endosulfan in several non-target organisms was also reported in the literature [3]. Moreover, presence of endosulfan in different environmental reservoirs (air, water and soil), even in remote locations, was reported [4]. Due to its persistence in the environmental reservoirs, scientific studies on fate and transport of, endosulfan will be important in coming years.
Endosulfan and its isomers, which are transported by evaporation, infiltration and surface, run-off mainly causes non-point source pollution in the environment. Therefore, on-site or in-situ (bio)remediation treatment methods are among the most preferred engineering applications for the removal of endosulfan from soil and groundwater. Effective and economically feasible bioremediation can be achieved using native soil microorganisms. Therefore, improving the knowledge on related native microorganisms and determining their pollutant degradation rates is a scientific necessity.
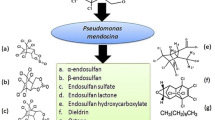
Commercially available technical-grade endosulfan (6,7,8,9,10,10-hekzakloro-1,5,5a,6,9,9a-hekzahidro-6,9-metano-2,3,4-benzo-dioxathiepin-3-oksit, CAS No. 115-29-7), has two isomers (α-endosulfan and β-endosulfan) in its composition, in a 7 to 3 ratio, respectively. Both of these isomers and some of their degradation by-products are toxic [5, 6]. Environmental fate studies indicate that β-endosulfan is more persistent in the environment than α-endosulfan [7]. The difference in the environmental fate of these isomers is determined by the difference in their vaporization, photo-decomposition and alkaline hydrolysis characteristics [8]. Although α-endosulfan has a higher volatility and rapid degradation rate compared to β-endosulfan, α-endosulfan is more toxic [7]. Endosulfan sulfate, a common endosulfan (bio)degradation by-product, is among the most toxic endosulfan compounds [4]. Some abiotic degradation of endosulfan is observed only under alkaline conditions [9], but it is not easy to distinguish this spontaneous abiotic hydrolysis from bacterial hydrolysis. Microorganisms dominate most of the endosulfan degradation in nature. A recent literature survey indicates more than 50 wild fungal, bacterial or actinomycetes species are capable of degrading endosulfan [10]. A list of these microbial species is given in Table 1. A majority of these species were isolated from soils exposed to endosulfan, using culture enrichment techniques [5, 11,12,13,14,15]. In most of these studies, endosulfan sulfate was observed as the by-product of the biodegradation process. The presence of endosulfan sulfate in environmental systems indicates (bio)degradation [16]. Endosulfan sulfate producing microorganisms are not useful for bioremediation applications since endosulfan sulfate is also toxic to organisms. Biodegradation pathways of endosulfan is reported in different studies in the literature (Fig. 1) [17] “Copyright, owned by [American Society of Microbiology]; used by permission.”

Reproduced with permission from [17]
Biodegradation pathway of endosulfan
The solubility of endosulfan in water is limited to 530 µg/L at 25 °C due to its hydrophobic nature [6]. Generally, observed endosulfan concentrations are lower than its maximum water solubility concentration, in natural water bodies. The solubility of endosulfan is enhanced in water with solvents in endosulfan biodegradation studies, according to the literature. Moreover, relatively higher temperatures, which may not be observed in field, are used in these biodegradation studies. In addition, the overall growth response of microorganisms and biological endosulfan degradation rates at low endosulfan concentrations is not known. Therefore, the investigation of microbial endosulfan degradation under natural conditions is of scientific interest.
In this study, we are reporting a consortium of microorganisms that can survive by biodegrading endosulfan at a low endosulfan concentration (100 µg/L) at room temperature (20 °C to 25 °C) without producing endosulfan sulfate as a by-product. These bacteria are adopted to relatively low pH environment but can also be efficient under alkaline conditions. Therefore, we believe that these microorganisms may contribute to efficient in-situ remediation of endosulfan contaminated sites. Overall endosulfan biodegradation rates of this consortium is also provided in this study.
Materials and Methods
Reactors and Experimental Set-ups Used in the Study
The biodegradation of endosulfan was monitored using 600 mL reaction volumes in 1L flasks. The reactors were stirred continuously (Labcon, 3100U) at a constant speed of 150 rpm. Four sets of experiments were designed. pH values were adjusted to pH 6.5 ± 0.5 or pH 8.4 ± 0.5. Biodegradation of endosulfan is also observed with or without the addition of glucose (4 mg/L) to the reactors, as the external carbon source. All experiments were conducted using two parallel reactors for each set. The reactors were covered with aluminum foil to prevent the photo-degradation of the endosulfan.
The reactors containing basic mineral medium were sterilized in an autoclave (NUVE OST32) at 121 °C under 1.06 bar pressure. Later endosulfan was added into the cooled solutions, using cold filter (0.22 µm filter) sterilization. Each reactor was inoculated with one-milliliter microbial consortium, prepared from slant agars in a biosafe cabin under sterile conditions.
Chemicals Used in the Experiments
All chemicals used in the experiments were analytical grade or higher. The endosulfan (%99.8 pure) was purchased from Bayer. The solution fed into the reactors was composed of K2HPO4 (25.50 mg), KH2PO4 (65.25 mg), Na2HPO4·7 H2O (100 mg), NH4Cl (195.10 mg), CaCl2 (82.50 mg), MgCl2·6H2O (67.5), FeCl3·6H2O (0.75 mg), Wolfe solution 10 ml [NaCl (1 g), MgCl2·6H2O (3 g), CoCl2 6H2O (0.1 g), MnCl2·4H2O (0.05 g), FeCl3·6H2O (0.1 g), ZnCl2·7H2O (0.1 g), CuCl2 (0.01 g)] per liter. To enhance the consumption of endosulfan as a sulfur and carbon source, all chemicals used in the experiments were selected in chlorinated forms. 0.1 N NaOH and 0.1 N HCl were used for pH adjustments of the reactors. Deionized water was used in all preparations and reactors.
Isolation and Identification of Microorganisms
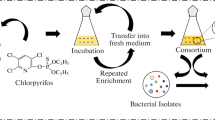
In order to isolate endosulfan degrading native soil bacteria, several soil samples from tea cultivation fields were collected. The use of any kind of pesticide is prohibited in tea cultivation fields in Turkey. Thus, none of the collected soil samples had any previously recorded endosulfan exposure history. The soil samples were taken from the top 10 cm of the soil with presterilized spoons and bags. Ten grams of the collected soil samples were screened for any debris or plant residuals under sterile conditions. The processed soil samples were mixed with a sterilized mineral solution, which contained 0.1 mg/L endosulfan in 150 mL volume in 250 mL flasks. After the slurry was incubated for 5 days on a shaker at 100 rpm, 10 mL supernatant was transferred to a fresh mineral solution under the same growing conditions and this first step of the enrichment process was repeated. In the second step, the microorganisms were transferred to a fresh solution every 24 h and this cycle was repeated 3 times. Then 50 mL of the microcosm was taken to a 100 mL flask to use in the experiments. The isolated bacterial consortium was stored on an endosulfan containing slant agar.
The isolated bacterial species were identified at Ankara University’s Biotechnology Center (Ankara, Turkey) using PCR technique. A 16S rRNA Gene DNA series analysis [18] was conducted. The specific primers used for 16S rRNA were the 27F (5′AGA-GTT-TGA-TCC-TGG-CTC-AG-3′) forward primer and the 1492R (5′GGT-TAC-CTT-GTT-ACG-ACT-T-3′) reverse primer. The classification of the strains was obtained by using the Evolutionary Relationship Three Neighbor-Joining method [19].
Analytic Techniques
The endosulfan isomers (α- and β-endosulfan) were analyzed using a gas chromatography instrument equipped with an electron capturing detector (Agilent Technologies 6890N GC/ECD). For the analysis, a HP-5 colon (30 m × 0.32 mm × 0.25 µm) and an injection volume of 2 µl was used. GC was used in splitless injection mode. High purity helium (He) was the carrier gas and nitrogen (N2) was used as make-up gas in the system. The detector temperature was set to 250 °C. The colon temperature was initially 60 °C and ramped to 280 °C at a rate of 20 °C per minute for 12 min [20]. The GC-ECD system was calibrated to detect the α and β isomers of endosulfan (R2 = 0.99). We used a 5-point calibration curve for the detection of endosulfan. Lowest observed level of the instrument was 1 μg/L. Lowest qualification concentration of the analysis was used as 5 μg/L.
Determination of Microbial Growth, Biodegradation Process and Kinetics Parameters
The growth of microorganisms in microcosms was monitored by observing the changes in optical density at 600 nm, using a UV–VIS digital spectrophotometer (Dr. Lange Cadas 200). The pH of the batch reactors were adjusted using HPLC grade 0.1N NaOH and 0.1N HCl. The pH measurements were made using an Orion pH-meter (81028NUWP ROSS Ultra Combination PH meter). The effect of external carbon sources and pH (6.5 and 8.4) on the biodegradation of endosulfan was also investigated during the biodegradation studies.
Endosulfan degradation kinetics was evaluated presuming first order reaction kinetics [21, 22]. such that;
where S0 is the initial substrate concentration (µg/L), S is the final substrate concentration (µg/L), k1 is the degradation constant (day−1), and t is time (day).
Results
Identification and Growth Kinetics of Endosulfan Degrading Bacterial Consortium
The three microorganisms in the endosulfan biodegrading consortium were identified as Afipia genosp species, sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum species with 100%, 100% and 99% precisions, respectively. The phylogenic tree presentation of the bacteria is given in Fig. 2. The gene sequences of Afipia genosp species, sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum were deposited in the GenBank database with the accession numbers of KY827230, KY827094, and KY827087, respectively.
In this study, no endosulfan sulfate formation was observed during the biodegradation process (Fig. 3). Endosulfan sulfate peaked at 24 min in the GC-Spectrum. According to Mukherjee et al. bacteria form endosulfan diol, a non-toxic by-product of endosulfan, during the biodegradation process. Endosulfan sulfate formation is usually observed during fungal biodegradation of endosulfan. The non-existence of endosulfan sulfate can also be related to using non-sulfur chemicals, which force the utilization of sulfur from endosulfan compound for microbial growth, in the experiments [7].
Afipia genosp species are rod-shaped, gram negative, oxidase positive microorganisms that produce gray-white, shiny, convex, and opaque 1.5 mm colonies on agar plates. The colonies on blood agar base were observed after 72 h incubation period at 32 °C [23]. No studies have been found in the literature on endosulfan biodegradation by the Afipia species. However, there are some studies on the biodegradation of polyaromatic hydrocarbons (PAH) by Afipia broomeae [24]. There are very limited scientific articles on the assimilation of sulfur by Afipia species [25].
The Sphingomonas species are defined as gram-negative, rod-shaped, chemoheterotrophic, strictly aerobic bacteria. They can metabolize a wide variety of carbon sources and survive in low concentrations of nutrients. Although the ability of the Sphingomonas sp. to biodegrade endosulfan is also reported for the first time in this study, these bacteria are known for their dehalogenation ability. Sphingomonas species are native to several environmental reservoirs such as soil and fresh water bodies. Sphingomonas species can biodegrade several anthropogenic chemicals, including chlorinated pesticides dibenzo-p-dioxine and dibenzofuron [26], carbofuran [27], hexachlorocyclohexane [28], polychlorofenol [29], 2,4-didlorobenzoic acid [30], dehydroabietic acid [8], as well as simple aromatic and polyaromatic hydrocarbons [31].
Methylobacterium species can utilize halogenated compounds for their carbon requirements. These gram-negative bacteria have the ability to use methane as well as other complex organic compounds as the sources of carbon and energy. These rod shaped pleomorphic, aerobic, facultative and methylotrophic organisms have the ability to survive in several natural environments (soil, dust, clean water, lake sediments, plants and rice fields) as well as in anthropogenic environments such as hospital surroundings [32]. Methylobacterium species are the only species that are related to biogeochemical sulfur cycle (Anesti 2004), in this study.
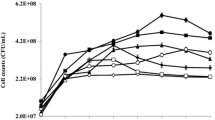
The disappearance of total endosulfan in the microcosms are related to the growth of microbial consortium under all pH values (pH 6.5 and 8.4) with or without the addition of glucose as the external carbon source. Microbial growth was observed as the time dependent optical density (OD) change, at 600 nm wavelength. The simultaneous change in endosulfan concentration and the optical density in the reactors are given in Fig. 4, for each experimental set.
a Removal of endosulfan and growth of microbial consortium at pH 6.5. b Removal of endosulfan and growth of microbial consortium in the presence of glucose at pH 6.5; c Removal of endosulfan and growth of microbial consortium at pH 8.4; d Removal of endosulfan and growth of microbial consortium in the presence of glucose at pH 8.4
Typical microbial growth phases (lag, exponential and stationary phases) were observed in all experiments. Shorter lag phases (≤ 5 days) were observed in the presence of external carbon source (glucose) in all reactors. Less than two-day lag phase was distinct for the reactors, where pH was low (pH 6.5). The reactors that were operated under acidic conditions (pH 6.5) had distinct exponential growth phases. In the alkaline reactors (pH 8.4), the transition from lag phase to exponential phase was diffused and exponential growth phase was extended to the 20th day or longer. The lag phase was observed in all reactors independent of carbon source and pH difference. Higher optical densities were observed in the reactors containing glucose.
The gradual increase in the microorganism numbers may indicate enzymatic adaptation need for endosulfan degradation. The study conducted by Goswami and Singh also reported a similar gradual biodegradation of endosulfan in the absence of an additional carbon source. In their study, when dextrose was used as the carbon source shorter lag phases were also observed in the reactors [33].
Biological Removal Rates of Endosulfan by the Isolated Consortium
The disappearance of endosulfan from the solution is statistically (p < 0.05) correlated to the increase in the optical density. This correlation is accepted as the biological removal of endosulfan by the isolated microbial consortium. No change in the endosulfan concentration was observed in control reactors in this study either. Control reactors were microcosms without microbial inoculation that were kept in the dark.
59% of the total endosulfan was biodegraded (63% α-endosulfan and 57% β-endosulfan) in the microcosms at pH 6.5 in 25 days. In the reactors at pH 8.4, 98% of the total endosulfan was biologically removed (96% α-endosulfan, 97% β-endosulfan) at the same time period. Presence of glucose as external carbon source did not change the overall removal efficiency (in percent) in the reactors. This observation also showed that endosulfan was the limiting substrate in the reactors.
Different reaction rate orders were investigated and it was concluded that first order reaction model fits best to the experimental results. Removal rate constants are given in Fig. 5. The degradation rate constants of α-endosulfan and β-endosulfan isomers were 0.111 day−1 and 0.116 day−1 at pH 6.5, respectively (Fig. 5a) The degradation rate constants of α-endosulfan and β-endosulfan isomers in the presence of glucose at pH 6.5 were 0.132 day−1 and 0.121 day−1, respectively (Fig. 5b). The degradation rate constants of α-endosulfan and β-endosulfan isomers at pH 8.4 were 0.203 day−1 and 0.237 day−1, respectively (Fig. 5c). The degradation constants of α-endosulfan and β-endosulfan isomers, in the presence of glucose at pH 8.4 were 0.182 day−1 and 0.244 day−1, respectively (Fig. 5d). First order removal rates of total endosulfan, at different pH values, in the presence or absence of glucose are calculated. According to these results, at pH 8.4 endosulfan removal rates are 0.221 (day−1) and 0.214 (day−1) in the absence and presence of glucose, respectively. At pH 6.5 the removal rates are calculated as 0.116 (day−1) and 0.125 (day−1) in the absence and presence of glucose, respectively. These results indicate that the removal rates are higher in high pH values. No significant removal rate changes is observed with the addition of carbon source.
Discussion
In this study, three native bacterial species, which can biodegrade endosulfan were isolated from soil. The results of this study show that these three native bacteria, namely Afipia genosp, Sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum are capable of growing by utilizing endosulfan isomers at low concentrations (100 µg/L). Soil samples collected from the tea cultivation fields did not have any known endosulfan exposure history, therefore ability of these soil microorganisms to utilize endosulfan require further research. Microorganisms need a carbon source for their metabolic activities and survival. Bioavailability of carbon determines the fate of bioremediation, in return. In this study, it was showed that endosulfan can be utilized and considered as carbon and sulfur source for the three endosulfan-degrading microorganisms of the consortium. Although endosulfan is utilized as a carbon and sulfur source by these three isolated bacteria, we believe that the success of endosulfan remediation application will be determined by the biological removal of sulfur from endosulfan molecule. Therefore, Methylobacterium sp. may be a key microorganism in successful bioremediation applications. The experimental design used in this study that eliminated the external sulfur in the solution, forced microorganisms to utilize endosulfan as a sulfur source. Since the results of this study did not show endosulfan sulfate formation at the end of the experiments, this consortium has the potential for bioremediation applications.
We observed slightly lower endosulfan removal rates compared to the other studies reported in the literature (Table 2). This can be related to a slow microbial growth rate since the growth rate is inversely related to the temperature of the microcosm. Temperature effect of on the endosulfan biodegradation and growth rate for these microorganisms will be evaluated separately.
According to the results of this study, it is also argued that a media without external sulfur sources must be preferred during the isolation of endosulfan degrading microorganisms [34]. Some studies in the literature used additional carbon sources (such as dextrose and glucose) in the mineral medium during biodegradation of endosulfan. In one of these studies, Bordetella sp. B9, a Gram negative bacteria, biodegraded 53% and 48% of α-endosulfan and, β-endosulfan with the addition of dextrose in 18 days, respectively. Without the additional carbon source, 80% of the α-endosulfan and 86% of the β-endosulfan were degraded [33]. Awasti et al. (1997) reported that addition of glucose to the mineral medium did not affect the degradation of endosulfan [35]. In a different study, Kumar and Philip used dextrose as an additional source of carbon with a mixed culture (Staphylococcus sp., Bacillus circulans I, Bacillus circulans II) for endosulfan biodegradation [36]. Following a four week incubation period, they concluded that endosulfan was degraded 71% in an aerobic environment and 76% in a facultative environment, whereas when the extra carbon source was used, degradation increased by 13% in the aerobic systems and 12% in the facultative anaerobic systems. Bhalerao and Puranik determined that the Aspergillus niger species is capable of degrading endosulfan in 12 days [37]. These relatively different observations indicate that presence of easily biodegradable carbon source may enhance endosulfan removal, depending on experimental design. Singh et al. reported that Achromobacter xylosoxidans species isolated from soil can use endosulfan as the sole source of sulfur and found that 94% of the α-endosulfan and 85% of the β-endosulfan were degraded. In this study, relatively lower percentages of α-endosulfan removal were observed. However, β-endosulfan removal performance of the bacterial consortium was found to be higher [38].
In this study, higher endosulfan removals percentages were obtained at higher initial pH values for both isomers. When Awasthi et al. investigated the effect of pH during degradation of endosulfan isomers in the soil, they noted that while there was no degradation of endosulfan isomers at pH 3, a slow degradation was observed at pH 5. Degradation was significant in pH range of 7.5 to 8.5 [39]. In the study conducted by Hussain et al. pH ranges between 4 and 10 were used for biodegradation and pH 6 was observed as the optimum value for biodegradation of both isomers [40]. The findings of this study support the conclusions those of the study by Awasthi [39]. Since our microcosms at pH 8.4 performed better. Therefore, the experimental findings of this study support the observations that better endosulfan removal performance can be achieved under alkaline environmental conditions.
Slight decrease in the pH of microcosms can be related to the accumulation of biodegradation by-products. The decrease in the pH value is related to the occurrence of HCl or to the microorganisms producing organic acid, as stated in the literature as well [21, 37, 41, 42].
First order removal rate constants of this study observed in the absence of glucose are compared to the values reported in the literature in Table 2. Literature survey shows different degradation rate constants for α-endosulfan and β-endosulfan isomers depending on the pH values. Endosulfan degradation constants observed in this study are parallel to the results given in the literature. For example, the results are comparable to the findings of Kwon et al. [21] and Kumar et al. [43]. In another study, two different isomer degradation rate constants were calculated for biphasic removal of endosulfan [44]. A number of endosulfan biodegradation studies including this study observed the removal of endosulfan from the solution in several phases.
The bacterial consortium of this study is adapted to a slightly acidic soil environment in the tea cultivation fields. Their short lag phase and distinct exponential phase are the proof of this observation. Endosulfan biodegradation performance of Afipia genosp, Sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum at higher pH values, which was investigated in this study, (with better overall removal percentages, due to additional abiotic degradation) and their ability to survive at low concentrations and temperature indicate the reliable use of this consortium in bioremediation applications.
Conclusions
The results of this study show that endosulfan-degrading microorganisms can be found in soils without known contaminant exposure history.
Afipia genosp, Sphingomonas yanoikuyae Q1 and Methylobacterium rhodesianum are capable of biodegrading endosulfan in acidic and alkaline conditions, which is reported in this study for the first time in literature.
Endosulfan at low concentrations can be utilized as carbon and sulfur source by this consortium. These bacteria can biodegrade endosulfan at 20 °C to 25 °C, which is a relatively lower temperature than other studies in the literature.
Endosulfan degradation rates of the bacterial consortium used in this study are comparable to other similar endosulfan biodegradation studies that exist in literature.
References
Gupta, P.K., Gupta, R.C.: Pharmacology, toxicology and degradation of endosulfan A review. Toxicology 13(2), 115–130 (1979)
Sun, X., Zhu, L., Wang, J., Wand, J., Su, B., Du, Z., Dua, P.: Effects of endosulfan on the populations of cultivable microorganisms and the diversity of bacterial community structure in Brunisolic soil. Water Air Soil Pollut. 228, 169 (2017)
Falkowska, L., Reindl, A.R., Szumiło, E., Kwaśniak, J., Staniszewska, M., Bełdowska, M., Lewandowska, A., Krause, I.: Mercury and chlorinated pesticides on the highest level of the food web as exemplified by herring from the Southern Baltic and African penguins from the zoo. Water Air Soil Pollut. 224(5), 1549 (2013)
Weber, J., Halsall, C.J., Muir, D., Teixeira, C., Small, J., Solomon, K., Hermanson, M., Hung, H., Bidleman, T.: Endosulfan, a global pesticide: a review of its fate in the environment and occurrence in the Arctic. Sci. Total Environ. 408(15), 2966–2984 (2010)
Bajaj, A., Pathak, A., Mudiam, M.R., Mayilraj, S., Manickam, N.: Isolation and characterization of a Pseudomonas sp. strain IITR01 capable of degrading -endosulfan and endosulfan sulfate. J. Appl. Microbiol. 109(6), 2135–2143 (2010)
ATSDR.: Toxicological profile for endosulfan. U.S. Department of Health and Human Services Public Health Service Agency For Toxic Substances and Disease Registry, 323 (2015)
Mukherjee, I., Mittal, A.: Bioremediation of endosulfan using Aspergillus and Cladosporium oxysporum. Bull. Environ. Contam. Toxicol. 75, 1034–1040 (2005)
Cotham, W.E.J., Bidleman, T.F.: Degradation of malathion, endosulfan and fenvelarate in seawater and sea water/sediment in microcosms. J. Agric. Food Chem. 37, 824–828 (1989)
Sutherland, T.D., Weir, K.M., Lacey, M.J., Horne, I., Russell, R.J., Oakeshott, J.G.: Enrichment of a microbial culture capable of degrading endosulphate, the toxic metabolite of endosulfan. J. Appl. Microbiol. 92, 541–548 (2002)
Shetty, P.K., Mitra, J., Murthy, N.B.K., Namitha, K.K., Savitha, K.N., Raghu, K.: Biodegradation of cyclodiene insecticide Endosulfan by Mucor thermohyalospora MTCC 1384. Curr. Sci. 79(9), 1381–1383 (2000)
Kaur, I., Mathur, R.P., Tandon, S.N.: Persistence of endosulfan (technical) in water and soil. Environ. Technol. 19, 115–119 (1998)
Taira, K., Hayase, N., Arimura, N., Yamashita, S., Miyazaki, T., Furukawa, K.: Cloning and nucleotide sequence of the 2,3-dihydroxybiphenyl dioxygenase gene from the PCB-degrading strain of Pseudomonas paucimobilis Q1. Biochemistry, 27, 3990–3996 (1998)
Kumar, M., Laksmi, C.V., Khanna, S.: Biodegradation and bioremediation of endosulfan contaminated soil. Biores. Technol. 99, 3116–3122 (2008)
Arshad, M., Hussain, S., Saleem, M.: Optimization of environmental parameters for biodegradation of alpha and beta endosulfan in soil slurry by Pseudomonas aeruginosa. J. Appl. Microbiol. 104(2), 364–370 (2008)
Elsaid, O.E.G., Abdelbagi, A.O., Elsheikhc, E.A.E.: Microbial degradation of endosulfan in carbon free media and selective media. Res. J. Agric. Biol. Sci. 6(3), 257–562 (2010)
Masoud, A.A., Abdel-Wahab Arafa, N.A., El-Bouraie, M.: Patterns and trends of the pesticide pollution of the shallow Nile Delta Aquifer (Egypt). Water Air Soil Pollut. 229, 148 (2018)
Sutherland, T.D., Horne, I., Lacey, M.J., Harcourt, R.L., Russell, R.J., Oakeshott, J.G.: Enrichment of an endosulfan-degrading mixed bacterial culture. Appl. Environ. Microbiol. 66, 2822–2828 (2000)
Stefan, R.I., Atlas, R.M.: Polymerase chain reaction. Appl. Environ. Microbiol. 45, 137–161 (1991)
Saitou, N., Nei, M.: The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987)
EPA Method 508.1., Determination of Chlorinated Pesticides, Herbicides, and Organohalides by Liquid-Solid Extraction and Electron Capture Gas Chromatography -Revision 2.0.38 pp
Kwon, G.S., Sohn, H.Y., Shin, K.S., Kim, E., Seo, B.: Biodegradation of the organochlorine insecticide, endosulfan, and the toxic metabolite, endosulfan sulfate, by Klebsiella oxytoca KE-8. Appl. Microbiol. Biotechnol. 67, 845–850 (2005)
Lee, J.B., Sohn, H.Y., Shin, K.S., Jo, M.S., Kim, J.E.,. Lee, S.W., Shin, J.W., Kum, E.J., Kwon, G.S.: Isolation of a soil bacterium capable of biodegradation and detoxification of endosulfan and endosulfan sulfate. J. Agric. Food Chem. 54, 8824–8828 (2006)
Bergey, D.H., Holt, J.G.: Bergey’s Manual of Systematic Bacteriology. Lippincott Williams & Wilkins, Philadelphia p. 787 (1994)
Bodour, A.A., Wang, J.M., Brusseau, M.L., Maier, R.M.: Temporal change in culturable phenanthrene degraders in response to long-term exposure to phenanthrene in a soil column system. Environ. Microbiol. 5, 888–895 (2003)
Moosvi, S.A., McDonald, I.R., Pearce, D.A., Kelly, D.P., Wood, A.P.: Molecular detection and isolation from Antarctica of methylotrophic bacteria able to grow with methylated sulfur compounds. Syst. Appl. Microbiol. 28, 541–554 (2005)
Wittich, R.M., Wilkes, H., Sinnwell, V., Francke, W., Fortnagel, P.: Metabolism of dibenzo-p-dioxin by Sphingomonas sp. Appl. Environ. Microbiol. strain RW1, 58, 1005–1010 (1992)
Mohn, W.W.: Bacteria obtained from a sequencing batch reactor that are capable of growth on dehydroabietic acid. Appl. Environ. Microbiol. 61, 2145–2150 (1995)
Imai, R., Nagata, Y., Fukuda, M., Takagi, M., Yano, K.: Molecular cloning of a Pseudomonas paucimobilis gene encoding a 17-kilodalton polypeptide that eliminates HCl molecules from gamma-hexachlorocyclohexane. J. Bacteriol. 173, 6811–6819 (1991)
Nohynek, L., Nurmiaho-Lassila, E., Suhonen, E., Busse, H., Mohammadi, M., Hantula, J.: Description of chlorophenol-degrading Pseudomonas sp. strains KF1T, KF3, and NKF1 as a new species of the genus Sphingomonas, Sphingomonas subarctica sp. nov.. Int. J. Syst. Evol. Microbiol. 46, 1042–1055 (1996)
Ka, J., Holben, W., Tiedje, J.: Genetic and phenotypic diversity of 2,4-dichlorophenoxyacetic acid (2,4-D)-degrading bacteria isolated from 2,4-D treated field soils. Appl. Environ. Microbiol. 60, 1106–1115 (1994)
Khan, A., Wang, R., Cao, W., Franklin, W., Cerniglia, C.: Reclassification of a polycyclic aromatic hydrocarbon-metabolizing bacterium, Beijerinckia sp. strain B1, as Sphingomonas yanoikuyae by fatty acid analysis, protein pattern analysis, DNADNA hybridization, and 16S ribosomal DNA sequencing. Int. J. Syst. Evol. Microbiol. 46, 466–469 (1996)
Green, P.N., Bousfield, I.J.: A taxonomic study of some gram-negative facultative methylotrophic bacteria. J. Gen. Microbiol. 128, 623–638 (1982)
Goswami, S., Singh, D.K.: Biodegradation of a and b endosulfan in broth medium and soil microcosm by bacterial strain Bordetella sp. Biodegradation B9, 20, 199–207 (2009)
Kataoka, R., Takagi, K.: Biodegradability and biodegradation pathways of endosulfan and endosulfan sulfate. Appl Microbiol Biotechnol. 97(8), 3285–3292 (2013)
Awasthi, N., Manickam, N., Kumar, A.: Biodegradation of endosulfan by a bacterial coculture. Bull. Environ. Contam. Toxicol. 59(6), 928–934 (1997)
Kumar, M., Philip, L.: Endosulfan mineralization by bacterial isolates and possible degradation pathway identification. Bioremediat. J. 10(4), 179–190 (2006)
Bhalerao, T.S., Puranik, P.R.: Biodegradation of organochlorine pesticide, endosulfan, by a fungal soil isolate, Aspergillus niger. Int. Biodeterior. Biodegrad. 59(4), 315–321 (2007)
Singh, N.S., Singh, D.K.: Biodegradation of endosulfan and endosulfan sulfate by Achromobacter xylosoxidans strain C8B in broth medium. Biodegradation 22(5), 845–857 (2011)
Awasthi, N., Ahuja, R., Kumar, A.: Factors influencing the degradation of soil-applied endosulfan isomers. Soil Biol. Biochem. 32, 1697–1705 (2000)
Hussain, S., Arshad, M., Saleem, M., Zahir, Z.A.: Screening of soil fungi for in vitro degradation of endosulfan. World J. Microbiol. Biotechnol. 23, 939–945 (2007)
Awasthi, N., Singh, A.K., Jain, R.K., Khangarot, B.S., Kumar, A.: Degradation and detoxification of endosulfan isomers by a defined co-culture of two Bacillus strains. Appl. Microbiol. Biotechnol. 62, 279–283 (2003)
Siddique, T., Okeke, B.C., Arshad, M., Frankenberger, W.T.J.: Biodegradation kinetics of endosulfan by Fusarium ventricosum and a Pandoraea species. J. Agric. Food Chem. 51(27), 8015–8019 (2003)
Kumar, K., Devi, S.S., Krishnamurthi, K., Kanade, G.S., Chakrabarti, T.: Enrichment and isolation of endosulfan Degrading and detoxifying bacteria. Chemosphere 68, 317–322 (2007)
Thangadurai, P., Suresh, S.: Biodegradation of endosulfan by soil bacterial cultures. Int. Biodeterior. Biodegrad. 94, 38–47 (2014)
Mukherjee, I., Gopal, M.: Degradation of beta-endosulfan by Aspergillus niger. Toxicol. Environ. Chem. 46, 217–221 (1994)
Katayama, A., Matsumura, F.: Degradation of organochlorine pesticides, particularly endosulfan, by Trichoderma harzianum. Environ. Toxicol. Chem. 12(6), 1059–1065 (1993)
Kullman, S., Matsumura, F.: Metabolic pathways utilized by Phanerochaete chrysosporium for degradation of the cyclodiene pesticide endosulfan, Appl, Environ, Microbiol,, 593–600 (1996)
Guha, A., Kumari, B., Bora, T.C., Deka, P.C., Roy, M.K.: Bioremediation of endosulfan by Micrococcus sp. Indian J. Environ. Health 42, 9–12 (2000)
Lee, S.E., Kim, J.S., Kennedy, I.R., Park, J.W., Kwon, G.S., Koh, S.C., Kim, J.E.: Biotransformation of an organochlorine insecticide, endosulfan by Anabaena species. J. Agric. Food Chem. 51(5), 1336–1340 (2003)
Park, B.S., Lee, S.E.: Biotransformation of β-endosulfan by Anabaena sp. PCC 7120. Agric. Chem. Biotechnol. 47(1), 38–41 (2004)
Sethunathan, N., Megharaj, M., Chen, Z.L., Williams, B.D., Lewis, G., Naidu, R.: Algal degradation of a known endocrine disrupting insecticide, α-endosulfan, and its metabolite, endosulfan sulfate, in liquid medium and soil. J. Agric. Food Chem. 52, 3030–3035 (2004)
Weir, K.M., Sutherland, T.D., Horne, I., Russell, R.J., Oakeshott, J.G.: A single monooxygenase, Ese, is involved in the metabolism of the organochlorides endosulfan and endosulfate in an Arthrobacter sp. Appl. Environ. Microbiol. 72, 3524–3530 (2006)
Kalyani, S., Sharma, J., Singh, S., Dureja, P.: Enrichment and isolation of endosulfan-degrading microorganism from tropical acid soil. J. Environ. Sci. Health B 44, 663–672 (2009)
Li, W., Dai, Y., Xue, B., Li, Y., Peng, X., Zhang, J., Yan, Y.: Biodegradation and detoxification of endosulfan in aqueous medium and soil by Achromobacter xylosoxidans strain CS5. J. Hazard. Mater. 167, 209–216 (2009)
Kong, L., Zhu, S., Zhu, L., Xie, H., Su, K., Yan, T., Wang, J., Wang, J., Wang, F., Sun, F.: Biodegradation of organochlorine pesticide endosulfan by bacterial strain Alcaligenes faecalis JBW4. J. Environ. Sci. 25(11), 2257–2264 (2013)
Narkhede, C.P., Patil, A.R., Koli, S., Suryawanshi, R., Wagh, N.D., Salunke, B.K., Patil, S.V.: Studies on endosulfan degradation by local isolate Pseudomonas aeruginosa. Biocatal. Agric. Biotechnol. 4(2), 259–265 (2015)
Ito, K., Kawashima, F., Takagi, K., Kataoka, R., Kotake, M., Kiyota, H., Yamazaki, K., Sakakibara, F., Okada, S.: Isolation of endosulfan sulfate-degrading Rhodococcus koreensis strain S1-1 from endosulfan contaminated soil and identification of a novel metabolite, endosulfan diol monosulfate. Biochem. Biophys. Res. Commun. 473(4), 1094–1099 (2016)
Chauhan, A., Pathak, A., Ewida, A.Y., Griffiths, Z., Stothard, P.: Whole genome sequence analysis of an Alachlor and Endosulfan degrading Pseudomonas strain W15Feb9B isolated from Ochlockonee River. Florida. Genomics Data 8, 134–138 (2016)
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bilgin, A., Sanin, S.L. Isolation and Identification of Endosulfan Degrading Native Bacterial Consortium from Agricultural Soils. Waste Biomass Valor 11, 3303–3313 (2020). https://doi.org/10.1007/s12649-019-00662-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-019-00662-5