Abstract
Biodegradation of endosulfan, a chlorinated cyclodiene insecticide, is generally accompanied by production of the more toxic and more persistent metabolite, endosulfan sulfate. Since our reported endosulfan degrader, Klebsiella pneumoniae KE-1, failed to degrade endosulfan sulfate, we tried to isolate an endosulfan sulfate degrader from endosulfan-polluted soils. Through repetitive enrichment and successive subculture using mineral salt medium containing endosulfan or endosulfan sulfate as the sole source of carbon and energy, we isolated a bacterium capable of degrading endosulfan sulfate as well as endosulfan. The bacterium KE-8 was identified as Klebsiella oxytoca from the results of 16S rDNA sequence analysis. In biodegradation assays with KE-8 using mineral salt medium containing endosulfan (150 mg l−1) or endosulfan sulfate (173 mg l−1), the biomass was rapidly increased to an optical density at 550 nm of 1.9 in 4 days and the degradation constants for α- and β-endosulfan, and endosulfan sulfate were 0.3084, 0.2983 and 0.2465 day−1, respectively. Analysis of the metabolites further suggested that K. oxytoca KE-8 has high potential as a biocatalyst for bioremediation of endosulfan and/or endosulfan sulfate.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Endosulfan (6,7,8,9,10,10-hexachloro-1,5,5a,6,9a-hexahydro-6,9-methano-2,3,4-benzodioxyanthiepin-3-oxide) is a chlorinated cyclodiene insecticide currently used throughout the world for the control of numerous pests in a wide variety of food and non-food crops. This broad-spectrum insecticide comprises two parent stereoisomers—α-endosulfan and β-endosulfan; the α to β ratio is about 2.33:4 (Goebel et al. 1982; Sutherland et al. 2002c), and both isomers are extremely toxic to aquatic organisms. In comparison to the more recalcitrant chlorinated cyclodiene insecticides, endosulfan is relatively labile in the environment (Maier-Bode 1968); reported half-life values of α-endosulfan range from 43 days to several months (Stewart and Cairns 1974; Rao and Murty 1980; Sethunathan et al. 2004), and endosulfan is easily degraded at alkaline pH (Sutherland et al. 2000) and/or photo-oxidized by UV light in aqueous medium (Knoevenagel and Himmelreich 1976). However, natural biodegradation alone is still insufficient to remove endosulfan and endosulfan derivatives in on-farm wastewater or during soil bioremediation.
Several intensive studies on the biodegradation of endosulfan in soils or in water environments have been conducted using pure- or mixed-cultures of microorganisms (Awasthi et al. 2003, 1997; Siddique et al. 2003; Lee et al. 2003; Sethunathan et al. 2004; Sutherland et al. 2002b, c; Kullman and Matsumura 1996) (Table 1). Since the major biotransformation products of endosulfan include endosulfan sulfate and endosulfan diol, and endosulfan sulfate is more toxic and more persistent than the parent endosulfan (Kaur et al. 1998), recent research has focused on degradation of endosulfan sulfate (Sutherland et al. 2002c; Sethunathan et al. 2004).
Recently, our group reported a potential endosulfan degrader, Klebsiella pneumoniae KE-1 (Kwon et al. 2002), and that the toxicity of endosulfan is closely associated with the oxidative stress via endosulfan-dependent reactive oxygen species generation (Sohn et al. 2004). Although K. pneumoniae KE-1 did not produce the toxic endosulfan sulfate during endosulfan degradation, this strain failed to degrade endosulfan sulfate. For practical applications, a bacterium capable of degrading endosulfan sulfate as well as endosulfan is necessary. In this study, we isolated an endosulfan sulfate degrader, Klebsiella oxytoca KE-8, from endosulfan-polluted soils through repetitive enrichment culture and successive subculture using mineral salt medium containing endosulfan or endosulfan sulfate as the sole source of carbon and energy. Biodegradation assays showed that K. oxytoca KE-8 has a high potential as a biocatalyst for bioremediation of endosulfan and/or endosulfan sulfate.
Materials and methods
Media and chemicals
The organic pesticide endosulfan and its metabolites, such as endosulfan sulfate, endosulfan diol, endosulfan lactone and endosulfan ether (Fig. 1) were purchased from Aldrich (Milwaukee, Wis.). Dichloromethane and acetone used in extraction and gas chromatography analysis were purchased from Junsei (Tokyo, Japan). The liquid mineral salt medium (MSM) used contained 1 g NH4NO3, 1 g K2HPO4, 0.5 g MgSO4·7H2O, 0.2 g CaCO3 in 1 l distilled water as previously reported (Kwon et al. 2002). Bacto agar (Difco, Detroit, Mich.) was added to the above MSM at a concentration of 1.5% (w/v) to prepare solid medium. Endosulfan and endosulfan sulfate were dissolved in methanol at a concentration of 10 g/l respectively and added to MSM at appropriate concentrations after sterilization.
Isolation of an endosulfan-, and endosulfan sulfate-degrading microorganism
Soil samples were collected from ginseng and pepper fields near Andong, Kyungpook, Korea. The field had generally received several applications of endosulfan in the summer months for 5 years. Soil samples (2 g) were suspended in 50 ml liquid MSM containing 150 mg l−1 endosulfan (endosulfan-MSM) in a 250 ml Erlenmeyer flask, and incubated at 30°C with shaking (70 rpm). After 7 days, 5 ml of each culture was re-inoculated into new endosulfan-MSM medium and further incubated at 30°C for 7 days. This subculture was repeated under the same culture conditions, and then an aliquot (0.2 ml) from each culture was applied to solid endosulfan-MSM for isolation of single colonies. After 4–5 days, a single colony in endosulfan-MSM was transferred to liquid endosulfan-MSM or methanol-MSM, respectively, to exclude methanol-dependent strains. To isolate an endosulfan sulfate degrader, two newly selected strains, together with the previously isolated nine bacterial strains capable of degrading endosulfan (Kwon et al. 2002), were cultured in liquid MSM containing 173 mg l−1 endosulfan sulfate. The strain named KE-8, which grows on endosulfan sulfate-MSM as well as endosulfan-MSM, has been deposited in the Korean Type Culture Collection under accession No. KCTC 10551BP.
Identification of the endosulfan degrader
To identify the KE-8 bacterium, a Vitek Gram-negative identification (+) kit (Biomerieux, Marcy L’Etoile, France) and a 16S rDNA sequence analysis were used. For sequence analysis, the same instrumentation and analytical techniques as previously reported (Kwon et al. 2002) were used. A 508 base segment of the 16S rDNA region was amplified with a forward primer, 9F [5′-GAGTTTGATCCTGGCTCAG; positions 9–27 (Escherichia coli 16S rDNA numbering)] and a reverse primer, 1542R (5′-AGAAAGGAGGTGATCCAGCC; positions 1542–1525) using an MJ Research (Waltham, Mass.) thermal cycler (Yoon et al. 1997). PCR was run for 35 cycles with the following thermal profile: denaturation at 94°C for 30 s, primer annealing at 55°C for 30 s and extension at 72°C for 45 s. The final cycle included extension for 10 min at 72°C. The PCR product was purified using a QIAquick PCR product purification kit (Qiagen, Hilden, Germany) and the purified 16S rDNA was sequenced using a cycle sequencing ready reaction kit, ABI Prism BigDye Terminator (Applied Biosystems, Foster City, Calif.) and an Applied Biosystems model 310 sequencer (Perkin Elmer, Boston, Mass.). The primer for the sequencing reaction was 536R (5′-GWATTACCGCGGCKGCTG-3′; positions: 536–519). The 16S partial sequence of isolated strain KE-8 was deposited in the GenBank database under accession no. AF440521.
Biodegradation of endosulfan and endosulfan sulfate
Pure culture experiments were performed in 250 ml flasks containing 50 ml MSM, supplemented with 150 mg l−1 endosulfan. After 5 days, the cells were collected by centrifugation at 8,000 g for 15 min, washed twice with 0.015 M phosphate buffer (pH 7.0), and suspended in MSM. The washed cells were inoculated into MSM containing endosulfan (154 mg l−1) or endosulfan sulfate (173 mg l−1) and cultured at 30°C in a rotary shaker (130 rpm) for 6 days. The initial pH was set to 7.2 to prevent chemical hydrolysis of endosulfan at alkaline pH; alkaline hydrolysis of endosulfan formed endosulfan diol, with approximately 10-fold increases in hydrolysis occurring with each increase in pH unit (Sutherland et al. 2000). Endosulfan and its metabolites, culture pH and biomass were determined at 1-day intervals. Endosulfan-free- or endosulfan sulfate-free MSM served as negative controls, and non-inoculated flasks were also run as a control. All data are represented as the mean value of triplicate experiments. To investigate the effect of pH on endosulfan degradation and cell growth, the initial pH of the MSM was adjusted with 1 N NaOH or 1 N HCl. The degradation constant (k1) was calculated using the following equation based on the first order kinetics derived from data from the exponential phase of degradation (Strotmann 2000):
where S0 is the initial substrate concentration, S is the remaining substrate concentration and t is the time in days.
Assay of endosulfan and its metabolites
Endosulfan and its metabolites were analyzed using a gas chromatography-ECD system as previously reported (Kwon et al. 2002).
Results
Isolation of endosulfan and endosulfan sulfate-degrading bacteria
From 30 soil samples, 16 different bacteria were isolated through repetitive enrichment culture and successive transfer using endosulfan-MSM. Among the different 16 strains, 14 showed more active growth in methanol-MSM medium compared to growth in endosulfan-MSM (results not shown). However, two strains, YS3-3 and KE-8, grew well in endosulfan-MSM, yielding an optical density at 550 nm of 0.6 in 7 days (Table 2), whereas these two strains had very poor growth in methanol-MSM. Strain KE-2 showed strong endosulfan degradation activity, and the average degradation rates for α-endosulfan and β-endosulfan were 13 and 2.4 mg l−1 day−1, respectively.
The two isolated bacterial strains, together with nine strains previously collected based on their ability to grow in endosulfan-MSM (Kwon et al. 2002), were transferred to solid endosulfan sulfate-MSM, and cultured at 30°C for 7 days. Among the 11 bacterial strains tested, only 1 bacterium, named KE-8, was able to grow on endosulfan sulfate-MSM. After several subcultures, KE-8 was finally selected as an endosulfan sulfate degrader. The previously selected endosulfan degrader, Klebsiella pneumoniae KE-1 (Kwon et al. 2002) failed to grow on endosulfan sulfate-MSM.
Identification of the endosulfan degrader
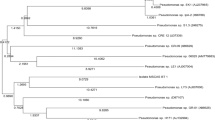
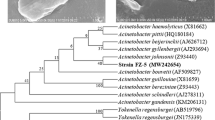
A partial 16S rDNA sequence was used to establish the identity of the isolated bacterial strain KE-8, the sequence being most similar to that of Klebsiella oxytoca type strain (ATCC 13182T; sequence similarity 98%). Thus, strain KE-8 was identified as Klebsiella oxytoca.
Biodegradation of endosulfan and endosulfan sulfate
K. oxytoca KE-8 was cultivated in endosulfan-MSM or endosulfan sulfate-MSM for 6 days. When endosulfan or endosulfan sulfate was omitted from the medium, no growth was observed and the pH remained at 7.2. In endosulfan-MSM, the cell density of KE-8 reached a maximum on day 4, and slightly decreased thereafter (Fig. 2a). The culture pH decreased to 5.6 from 7.2 on day 4 concomitant with cell density, and increased to 6.2 at the end of stationary phase. Endosulfan was degraded from 154 to 26 mg l−1 in 6 days (a decrease of 83%), and degradation of α-endosulfan was more evident than that of β-endosulfan; the degradation constants for α- and β-endosulfan were 0.3084 day−1 (R2=0.99) and 0.2983 day−1 (R2=0.962), respectively (Fig. 2b, d, e). A time- course of the metabolites showed that endosulfan diol and endosulfan lactone were major metabolites, their concentration increasing to 33 and 31 mg l−1 on days 4 and 5, respectively (Fig. 2c). At stationary phase, these concentrations were considerably decreased. Endosulfan sulfate was detected in the culture; the highest concentration was 11.8 mg l−1 on day 2, with the concentration slowly decreasing during the exponential and stationary phases. After 5 days, endosulfan sulfate was not detected. Other reported metabolites, such as endosulfan ether or endosulfan hydroxyether (Kwon et al. 2002; Siddique et al. 2003; Lee et al. 2003; Kullman and Matsumura 1996) were not detected. These results suggest that endosulfan sulfate is not the final metabolite in this culture, and that K. oxytoca KE-8 can degrade endosulfan sulfate as well as endosulfan.
Biodegradation of endosulfan by Klebsiella oxytoca KE-8. a Cell density (◊) of K. oxytoca KE-8 and culture pH (△) in endosulfan-MSM over 6 days. b Degradation of endosulfan isomers [α-endosulfan (▼), β-endosulfan (■), sum of α-endosulfan and β-endosulfan (●)] by K. oxytoca KE-8 in endosulfan-MSM. c Concentration changes of endosulfan diol (○), endosulfan lactone (▽) and endosulfan sulfate (□) in culture during endosulfan degradation. d, e Degradation constants for α-endosulfan (d) and β-endosulfan (e) based on first order kinetics from the data of the exponential phase of degradation
To investigate the degradation of endosulfan sulfate, K. oxytoca KE-8 was cultivated in MSM supplemented with endosulfan sulfate (173 mg l−1) as the sole source of carbon and energy. The cell density of KE-8 again reached a maximum on day 4 and afterwards slightly decreased (Fig. 3a). The culture pH decreased to 5.5 from 7.2 by day 4, concomitant with cell density, and increased to 6.1 at stationary phase. The increases in culture pH during stationary phase are a prominent characteristic of KE-8. Endosulfan sulfate was degraded from 173 to 61 mg l−1 over 6 days (a decrease of 64%). In a prolonged culture over 12 days, only 58.2±2.53 mg l−1 endosulfan sulfate remained (results not shown). About 94% of endosulfan sulfate metabolized was degraded in the first 4 days (degradation constant =0.2465 day−1, R2=0.971) (Fig. 3b). Analysis of metabolites showed that endosulfan lactone is a major metabolite of endosulfan sulfate. and that its concentration slowly increased, reached a maximum (44 mg l−1) on day 4, and then decreased rapidly. Only traces (<1 mg l−1) of endosulfan diol were detected, without concentration changes over 6 days, and endosulfan ether and endosulfan hydroxyether were not detected.
Biodegradation of endosulfan sulfate by K. oxytoca KE-8. a Changes of endosulfan sulfate concentration (●), culture pH (△), cell density (◊) and endosulfan lactone (▽) over 6 days cultivation in endosulfan sulfate-MSM. b The degradation constant for endosulfan sulfate based on the first order kinetics of data from the exponential phase of degradation
Discussion
Klebsiella oxytoca KE-8, a bacterium capable of degrading endosulfan sulfate as well as endosulfan, was isolated from the soil of a pepper field, Andong, Korea. The previously reported nine endosulfan degraders and a newly isolated endosulfan degrader YS3-3 did not grow on endosulfan sulfate-MSM. These results suggest that endosulfan sulfate is the most persistent metabolite of endosulfan for these strains. However, our results do not represent the degradation efficiency of endosulfan sulfate in the environment, because endosulfan sulfate was used as sole carbon and energy source in this study and our culture medium includes MgSO4 (0.05%). Supplementation of endosulfan sulfate as the sole sulfur source may be more effective in isolating endosulfan sulfate degraders, since it has a relatively reactive sulfite moiety (Sutherland et al. 2002c). In fact, an endosulfan-degrading Mycobacterium strain failed to metabolize endosulfan in sulfur-free medium containing 50 μM endosulfan upon supplementation of 200 μM MgSO4 (Sutherland et al. 2002b). Although degradation of endosulfan sulfate by this strain was not reported, the presence of other carbon sources or inorganic sulfate in the environment may affect the biodegradation activity or degradation efficiency.
In the biodegradation assay with K. oxytoca KE-8, the concentrations of endosulfan and endosulfan sulfate in the culture decreased rapidly with concomitant increases in biomass. During the exponential degradation phase, the specific degradation constants (and half-life t1/2) for α- and β-endosulfan, and endosulfan sulfate were 0.3084 day−1 (2.28 days), 0.2983 day−1 (2.41 days), 0.2465 day−1 (2.97 days), respectively (Figs. 2, 3). These results suggest that K. oxytoca KE-8 has similar degradation activity toward endosulfan isomers and endosulfan sulfate, and that K. oxytoca KE-8 could be successfully employed as a biocatalyst for bioremediation of endosulfan and endosulfan sulfate. This strain may provide a useful gene source to develop other strains for bioremediation (Sutherland et al. 2002a). The construction of a cosmid library from the KE-8 genome is in progress in our laboratory.
It is uncertain that endosulfan sulfate is converted to endosulfan diol as previously proposed (Verschueren 1999). In endosulfan sulfate-MSM culture, the concentration of endosulfan diol was maintained a very low level (<1 mg l−1), and endosulfan ether and endosulfan hydroxyether were not detected, whereas unidentified metabolites were detected by gas chromatography and HPLC. Therefore, we cannot exclude the possibility of direct desulfurization of endosulfan sulfate, or a novel degradation pathway in KE-8 as reported in enrichment culture of soil bacteria (Sutherland et al. 2002c). Further research on the metabolites of endosulfan sulfate, the degradation pathway of endosulfan sulfate, simultaneous degradation of endosulfan/endosulfan sulfate, and the effect of available sulfur compounds on degradation efficiency of endosulfan sulfate are necessary.
References
Awasthi N, Singh AK, Jain RK, Khangarot BS, Kumar A (2003) Degradation and detoxification of endosulfan isomers by a defined co-culture of two Bacillus strains. Appl Microbiol Biotechnol 62:279–283
Awasthi N, Manickam N, Kumar A (1997) Biodegradation of endosulfan by a bacterial coculture. Bull Environ Contam Toxicol 59:928–934
Goebel H, Gorbach S, Knauf W, Rimpau RH, Huttenbach H (1982) Properties, effect, residues and analytics of the insecticide endosulfan. Residue Rev 83:1–122
Kaur I, Mathur RP, Tandon SN, Dureja P (1998) Persistence of endosulfan (technical) in water and soil. Environ Technol 19:115–119
Knoevenagel K, Himmelreich R (1976) Degradation of compounds containing carbon atoms by photo-oxidation in the presence of water. Arch Environ Contam Toxicol 4:324–333
Kullman SW, Matsumura F (1996) Metabolic pathways utilized by Phanerochaete chrysosporium for degradation of the cyclodiene pesticide endosulfan. Appl Environ Microbiol 62:593–600
Kwon G-S, Kim J-E, Kim T-K, Sohn H-Y, Koh S-C, Shin K-S, Kim D-G (2002) Klebsiella pneumoniae KE-1 degrades endosulfan without formation of the toxic metabolite, endosulfan sulfate. FEMS Microbiol Lett 215:255–259
Lee S-E, Kim J-S, Kennedy IR, Park J-W, Kwon G-S, Koh S-C, Kim J-E (2003) Biotransformation of an organochlorine insecticide, endosulfan, by Anabaena species. J Agric Food Chem 51:1336–1340
Maier-Bode H (1968) Properties, effect, residues, and analytics of the insecticide endosulfan. Residue Rev 22:1–44
Rao DMR, Murty AS (1980) Persistence of endosulfan in soils. J Agric Food Chem 28:1099–1101
Sethunathan N, Megharaj M, Chen ZL, Williams BD, Lewis G, Naidu R (2004) Algal degradation of a known endocrine disrupting insecticide, α-endosulfan, and its metabolite, endosulfan sulfate, in liquid medium and soil. J Agric Food Chem 52:3030–3035
Siddique T, Okeke BC, Arshad M, Frankenberger WT Jr (2003) Biodegradation kinetics of endosulfan by Fusarium ventricosum and a Pandoraea species. J Agric Food Chem 51:8015–8019
Sohn H-Y, Kwon C-S, Kwon G-S, Lee J-B, Kim E (2004) Induction of oxidative stress by endosulfan and protective effect of lipid-soluble antioxidants against endosulfan-induced oxidative damage. Toxicol Lett 151:357–365
Stewart DKR, Cairns KG (1974) Endosulfan persistence in soil and uptake by potato tubers. J Agric Food Chem. 22:984–986
Strotmann U (2000) Futher development of the biological degradation test procedure considering respirometry. In: 1st Symposium on Biological Degradability, 1–10
Sutherland TD, Horne I, Harcourt RL, Russell RJ, Oakeshott JG (2002a) Gene cloning and molecular characterization of a two-enzyme system catalyzing the oxidative detoxification of β-endosulfan. Appl Environ Microbiol 68:6237–6245
Sutherland TD, Horne I, Harcourt RL, Russell RJ, Oakeshott JG (2002b) Isolation and characterization of a Mycobacterium strain that metabolizes the insecticide endosulfan. J Appl Microbiol 93:380–389
Sutherland TD, Horne I, Lacey MJ, Harcourt RL, Russell RJ, Oakeshott JG (2000) Enrichment of an endosulfan-degrading mixed bacterial culture. Appl Environ Microbiol 66:2822–2828
Sutherland TD, Weir KM, Lacey MJ, Horne I, Russell RJ, Oakeshott JG (2002c) Enrichment of a microbial culture capable of degrading endosulphate, the toxic metabolite of endosulfan. J Appl Microbiol 92:541–548
Verschueren K (1999) Handbook of environmental data of organic chemicals, 4th edn. Wiley, New York
Yoon JH, Lee ST, Kim SB, Kim WY, Goodfellow M, Park YH (1997) Restriction fragment length polymorphism analysis of PCR-amplified 16S ribosomal DNA for rapid identification of Saccharomonospora strains. Int J Syst Bacteriol 47:111–114
Acknowledgements
This work was supported by a grant from BioGreen21 program, Rural Development administration, Republic of Korea (1000520030096000).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kwon, GS., Sohn, HY., Shin, KS. et al. Biodegradation of the organochlorine insecticide, endosulfan, and the toxic metabolite, endosulfan sulfate, by Klebsiella oxytoca KE-8. Appl Microbiol Biotechnol 67, 845–850 (2005). https://doi.org/10.1007/s00253-004-1879-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1879-9