Abstract
Plants store triacylglycerides in organelles called oil bodies, which are important fuel sources for germination. Oil bodies consist of a lipid core surrounded by an interfacial single layer membrane of phospholipids and proteins. Oleosins are highly conserved plant proteins that are important for oil body formation, solubilising the triacylglycerides, stabilising oil bodies, and playing a role in mobilising the fuel during the germination process. The domain structure of oleosins is well established, with N- and C-terminal domains that are hydrophilic flanking a long hydrophobic domain that is proposed to protrude into the triacylglyceride core of the oil body. However, beyond this general understanding, little molecular level detail on the structure is available and what is known is disputed. This lack of knowledge limits our understanding of oleosin function and concomitantly our ability to engineer them. Here, we review the state of play in the literature regarding oleosin structure and function, and provide some examples of how oleosins can be used in commercial settings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant seeds need a source of fuel to germinate. Once embryogenesis begins, the required chemical energy is released by catabolising fuel stores, which generally consist of starch, proteins, and fats (Waschatko et al. 2016). Triacylglycerides are glycerol esters of fatty acids and are a key energy storage molecule (Murphy 1993). However, because they are insoluble in water, plants store triacylglycerides in oil bodies, which are specialised organelles that provide easy access to the energy rich fats during the germination process.
The membrane of oil bodies comprises a phospholipid monolayer embedded with proteins (Yatsu and Jacks 1972; Fang et al. 2014; Kanazawa et al. 2020), which together envelop the stored triacylglycerides (Fig. 1a). The main protein component of oil bodies are oleosins (Tzen et al. 1993). Although the 3D structure of oleosins is largely unknown, they are predicted to fold into unique structure (Fig. 1b) that contains a central hydrophobic domain flanked by two hydrophilic terminal domains (an N-terminal domain and a C-terminal domain) (Pons et al. 2005). The terminal domain rest on the oil body membrane atop the phospholipid heads, while the central hydrophobic domain (H-domain) extends beyond the membrane and is embedded into the triacylglycerides of the oil body (Jolivet et al. 2017). Within the H-domain is an unusual proline knot motif that is predicted to form a hairpin turn (Fig. 1b) that is conserved across all oleosin sequences (Huang 1996).
a Schematic representation of the oil body structure (triacylglyceride core, phospholipids and proteins of the interfacial membrane). b. Schematic representation of predicted oleosin structure with the proline knot highlighted. c. Cryo-SEM images of the outer endosperm tissues of coconut adjacent to the testa. Magnified to show one cell. Bulbous looking spheres are the oil bodies indicated by arrows. Reproduced from Dave et al. (2019), with permission from Elsevier. d. Cryo-SEM images of the outer endosperm tissues of coconut adjacent to the testa. Multiple cells are in view, arrows show cells that have no oil bodies. Reproduced from Dave et al. (2019), with permission from Elsevier
First reported in the late 1980s (Murphy and Cummins 1989), oleosins are vital to the structural integrity of the oil body (Murphy 2001). Due to their amphiphilic nature, however, they are difficult to study and this has slowed research efforts. It has also resulted in conflicting results, especially in the context of the predicted structure (Li et al. 1992, Millichip et al. 1996). Overall, little is known about oleosin structure or function—knowledge gaps that if addressed may inform new applications for oleosins in industry. Here, we will focus on the current state of knowledge for oleosin structure and function with a brief overview of their biosynthesis and examples of how they are being developed for commercial use.
Oil bodies: composition and structure
Oil bodies are small (0.5–2 µm) lipid-based intracellular organelles in plants (Fig. 1c) (Tzen et al. 1993; Huang 1996; Shimada et al. 2008). They are found in high levels in certain tissues, such as in seeds, flowers, pollen, stamen, and fruits (Dave et al. 2019). Functioning as an energy reserve for seeds during germination and post-germinative growth, oil bodies are thought to provide an increased surface area for lipase action during triacylglyceride mobilisation after dormancy or during germination (Huang 1992). Not all cells contain the same number of oil bodies—some can be oil body rich, while others have none (Fig. 1d shows coconut cells, highlighted by arrows, without oil bodies).
Oil bodies consist of a lipid core surrounded by an interfacial membrane of phospholipids and proteins (Dave et al. 2019). The lipid core is largely composed of triacylglycerides, but some bioactive compounds can be found (e.g., vitamin E, carotenoids and phytosterols) (Acevedo-Fani et al. 2020). The phospholipid fraction of the interfacial membrane predominantly contains phosphatidylcholine, which accounts for ~ 50% of the total phospholipids. Minor fractions of other phospholipids can also be found, such as phosphatidylserine, phosphatidylethanolamine and phosphatidylinositol (Huang 1992; Payne et al. 2014). The protein fraction consists of membrane- specific proteins: oleosins, caleosins and steroleosins. The most abundant structural proteins of oil bodies are oleosins, covering most of the oil body’s surface (e.g., in Arabidopsis thaliana oleosins make up 79% of the oil body proteins (Jolivet et al. 2004)). Generally, oil body composition is 94–98% (w/w) triacylglycerides, 0.6–2.0% (w/w) phospholipids, and 0.6–3.0% (w/w) protein (Tzen et al. 1993; Nikiforidis et al. 2014).
The general arrangement of the interfacial membrane components (phospholipids and proteins) is largely known. At the interface, the hydrophobic tails of acyl moieties of phospholipids interact with the lipid core and the hydrophilic head groups face the cytosol (Huang 1992; Tzen and Huang 1992). Oleosins are oriented with their terminal domains atop the phospholipid monolayer. It is thought that positive residues on the N- and C-terminal domains interact with the negatively charged phosphate groups of the phospholipids to support the interfacial structure of oil bodies (Ratnayake and Huang 1996; White et al. 2008). Functionally, oleosins are thought to stabilise oil bodies against coalescence inside plant cells through electrostatic repulsion and steric effects (Frandsen et al. 2001; Maurer et al. 2013). The saturated nature of the fatty acids in the phospholipid fraction may also increase the physical stability of oil bodies; the lack of double bonds allows the fatty acids to be fully extended, and closely packed, promoting a firm anchorage of the oleosins and strengthening the oil body interface (Payne et al. 2014). Recent evidence suggests that the interfacial proteins of coconut oil bodies are disulfide-linked (Dave et al. 2019), which may further contribute to the stability of oil bodies.
Biosynthesis of oleosins and oil bodies
Oleosins are synthesised via the usual protein synthetic machinery, although there are several features to note. The ribosome synthesising the oleosin polypeptide chain is transported to the endoplasmic reticulum via the co-translation synthesis pathway (Fig. 2a) (Hills et al. 1993, Beaudoin et al. 2000, Huang and Huang 2017). During translation, the signal recognition particle (Fig. 2a, in green) binds to the signal region within the developing polypeptide chain of the polypeptide/ribosome complex. The signal recognition particle then binds the signal recognition particle receptor attached to the endoplasmic reticulum membrane (Fig. 2a in orange). This mode of translation embeds the oleosin into the endoplasmic reticulum membrane (Loer and Herman 1993). There are two regions within the oleosin polypeptide chain that may bind the signal recognition particle: the first being a section of the H-domain found close to the N-terminal and the second is the proline knot motif (van Rooijen and Moloney 1995; Abell et al. 1997; Abell et al. 2002; Beaudoin and Napier 2002; Huang and Huang 2017).)
a Schematic of oleosin synthesis. The signal recognition particle (green) binds to the polypeptide as it is being transcribed. The signal recognition particle carries the transcription machinery to the signal recognition particle receptor (orange) found on the endoplasmic reticulum membrane. Transcription of the oleosin then finishes with the H-domain being deposited into the membrane of the endoplasmic reticulum. The endoplasmic reticulum expands as more triacylglycerides are synthesised and more oleosins are added, finally forming an oil body ready to bud off the endoplasmic reticulum. b. Predicted schematic of how oleosin breakdown could occur. Oleosin is first phosphorylated or ubiquitinated, this allows a protease to recognise the oleosin and begin the process of digesting the oleosin. This then allows room for lipase to bind and begin metabolising triacylglycerides
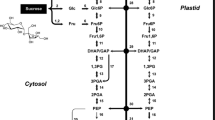
The mechanism through which the oil body forms is not fully understood, although the most promising hypothesis is the budding model. Following the budding model, oil bodies have three main steps in their formation: 1) fatty acid synthesis, 2) triacylglyceride assembly and 3) oil body budding (Song et al. 2017) (Fig. 2a). First the fatty acids are synthesised in a plastid using glycerol derived from photosynthesis. They are then moved to the endoplasmic reticulum and assembled into triacylglycerides. As triacylglycerides are being synthesised within the endoplasmic reticulum membrane, oleosins are being deposited into the same region creating a budding oil body that is then detached from the endoplasmic reticulum (Fig. 2a) (Wanner et al. 1981; Hsieh and Huang 2004). The mechanism of release is not understood and there is the question of how oleosins fold into their final structure in the oil body (Sarmiento et al. 1997, Song et al. 2017)—does this occur when deposited into the membrane or during the budding process? Whether oleosins play a role in forming the sharp curvature that facilitates detachment from the endoplasmic reticulum is an open question.
Although the budding model is more widely accepted, a second hypothesis for oil body formation is the post-encasement model. This model suggests that triacylglycerides build-up in the cytoplasm and are only encased in the oil body during the later stages of germination (Murphy 1993). This though does not seem likely as triacylglycerides are extremely hydrophobic and would not ‘linger’ in the hydrophilic environment of the cytoplasm. This theory also doesn’t explain how oleosins are deposited into the membrane, whereas in the budding model oleosins are targeted to the endoplasmic reticulum. These factors make the post-encasement model difficult to rationalise.
Decoding oleosin function
Stabilising oil bodies
It was established early on that oleosins stabilise oil bodies by preventing their coalescence or aggregation (Tzen and Huang 1992), which ensures a large surface area for lipase activity (Huang 1992). As noted, oleosins sit in the membrane of the oil body, with the two hydrophilic termini sitting atop of the phospholipid monolayer, and the H-domain nestled into the oil body. It is thought that both the N- and C-terminal domains and the H-domain play a role in stabilising the oil body.
Early studies demonstrated that oil bodies coalesce when the charge of the oleosins is neutralised, which could be due to the oleosin terminals dissociating from the phospholipids (Tzen et al. 1992). The removal of these domains results in the oil bodies coalescing and bursting, this leads to the idea that the two hydrophilic terminal domains brace the structure of the oil body membrane (Maurer et al. 2013). It is worth noting that most plant species have two different oleosin isoforms, which may have subtle differences in their function (Tzen et al. 1990). These isoforms have different molecular weights (e.g., in maize there is a 16 kDa lower, and an 18 kDa higher isoform), due to a longer C-terminus (Tzen et al. 1990). Both isoforms have been found in oil bodies together (Hsieh and Huang 2004) with the lower isoform being more effective in its function of stabilisation (Tzen et al. 1998).
The length of the H-domain, which is identical in both plant isoforms, is important for the size and stability of oil bodies. Peng et al. (2007) tested this hypothesis by reducing the length of the H-domain in an oleosin and examining how this affected the protein and oil body. The wild type H-domain is predicted to have 30 residues preceding the proline knot followed by a further 29 residues (30r-PK-29r). The study generated five truncated H-domain variants recombinantly: 18r-PK-29r, 18r-PK-17r,18r-PK-5r, 6r-PK-5r, and 0r-PK-0r (i.e., just the proline knot). The N- and C-terminal domains are retained. When compared to the wild type oleosin, the 18r-PK-29r and 18r-PK-17r variants are able to form normal sized, stable artificial oil bodies and prevented coalescence. This is consistent with another protein in the membrane of oil bodies, caleosin, which also has a very similar, but shorter H-domain (18r-PK-18r) (Peng et al. 2007). Oil bodies with the 18r-PK-5r, 6r-PK-5r, and 0r-PK-0r variants, however, show increasing susceptibility to coagulation, especially at elevated temperatures, leading to the conclusion that 18r-PK-17r is the shortest H-domain length required for oleosin stabilisation of oil bodies (Peng et al. 2007). Although this experiment would appear to show how unnecessarily long the oleosin H-domain is, finer truncations of the H-domain may reveal subtleties in the role of symmetry in the residues either side of the proline knot of the H-domain. Perhaps the longer tail and the proline knot play a role for the formation of the oil body in vivo.
Role in germination
Oil bodies have an important role in germination, particularly in the initial stages (Hsieh and Huang 2004; Purkrtova et al. 2008; Quettier and Eastmond 2009; Itabe 2010; Jolivet et al. 2013; Song et al. 2017). In general, oil bodies are metabolised by lipases and the glyoxysome, which is often found close to oil bodies (Hayashi et al. 2001). Lipases catalyse the stepwise hydrolysis of triacylglycerides to diacylglycerides, which is the first step in the gluconeogenic pathway (Lin et al. 1983; Wong and Schotz 2002) and the glyoxysome is a peroxisome and holds many of the enzymes that are involved in breaking down fatty acids to carbohydrates (Beevers 1979, 1980; Chapman and Trelease 1991). How lipase and the glyoxysome interact with the oil bodies is a mystery, however there is evedence that oleosins may be involved in these crucial interactions.
Oleosins are reported to be phosphorylated by a ‘serine, threonine, tyrosine protein kinase’ during germination (Fig. 2b) (Parthibane et al. 2012a, 2012b); Ramachandiran et al. 2018). In Arachis hypogaea (peanut), oleosin (OLE3) has been shown to be part of a complex of proteins that has duel monoacylglycerol acyltransferase and phospholipase A2 activities (Parthibane et al. 2012a, 2012b). The ‘serine, threonine, tyrosine protein kinase’ was shown to bind to peanut OLE3 and phosphorylated predominantly serine residues, particularly Ser18, which is not conserved across oleosins (Fig. 3a) (Parthibane et al. 2012a, 2012b). Similar studies in Arabidopsis thaliana OLE1 found that it was also phosphorylated by a ‘serine, threonine, tyrosine protein kinase’ on Thr166, which is again not conserved (Fig. 3a) (Ramachandiran et al. 2018). The role of phosphorylation may be to recruit proteins, such as proteases, to the oleosin, although these studies raise the possibility that the N- and C-terminal domains may have their own catalytic functions.
a Sequence alignment of oleosins. The green line signifies the approximate N-terminal, the orange line signifies the approximate H-domain, the purple line signifies the proline knot motif, and the pink line signifies the approximate C-terminal, Box one highlights Ser18, box two highlights Thr166. b. Oleosin models of OLE1 A. thaliana generated using AlphaFold, and models of almond, hemp, and sunflower oleosins generated using RaptorX. The proline knot motif of A. thaliana is highlighted with the conserved residues indicated
Thiol-proteases are reported to degrade oleosins from the oil body, allowing lipases access to the oil body for triacylglyceride digestion (Fig. 2b) (Sadeghipour and Bhatla 2002; Vandana and Bhatla 2006). Tracking the abundance of oleosins throughout germination using SDS-page analysis demonstrated that the lowest molecular weight sunflower oleosin disappears and this coincides with the increasing activity of a 65-kDa thiol-protease (Vandana and Bhatla 2006). Zymographic analysis demonstrated that the protease interacts with the oil body and that the protease could degrade oleosins when it was isolated with oil bodies. When the concentration of the protease is increased, all oleosins in the oil body were removed regardless of the isoform (Vandana and Bhatla 2006).
Similarly, oleosins, along with other proteins, may be removed from the oil body via the ubiquitination pathway (Fig. 2b) (Hsiao and Tzen 2011; Deruyffelaere et al. 2015). Removal of oleosins via ubiquitination is thought to occur as the first step in the germination process. Oleosins isolated from seeds during germination, and analysed by protein mass spectrometry, were found to contain ubiquitin. This was supported by immunological detection with antibodies against both oleosin and ubiquitin (Hsiao and Tzen 2011; Deruyffelaere et al. 2015). Higher isoforms of oleosins and caleosin (from Sesamum indicum (sesame) and Arabidopsis thaliana) were found to be ubiquitinated at the C-terminal regions (Hsiao and Tzen 2011; Deruyffelaere et al. 2015). There are lysine residues on the C-terminal domain of the higher isoform that could serve as a site for ubiquitination
A limited structural understanding of oleosins
The domain structure of oleosins is well established based on their amino acid sequence (Fig. 3a). The N- and C-terminal domains are hydrophilic, whereas the middle H-domain is hydrophobic (Huang 1992). The long H-domain, which spans around 68–74 residues, is thought to be the longest hydrophobic stretch of residues found in any protein (Huang 1992; Hsieh and Huang 2004; Huang and Huang 2015). Beyond the primary structure, where oleosin sequences have been accumulating as plant genome sequences are reported, the only structural information for oleosins comes from Fourier transform infrared spectroscopy and circular dichroism spectroscopy, which reports on the proteins secondary structure in the far-UV range (180–230 nm) and on protein tertiary structure in the near-UV range (260–320 nm). However, protein structures can now be predicted from the primary sequence using protein folding software, such as AlphaFold (Jumper et al. 2021) or RaptorX (Xu et al. 2021) with surprising accuracy—here we have considered the AlphaFold model of OLE1 from A. thaliana (Fig. 3b) and used RaptorX to generate oleosin structures from almond, hemp, and sunflower.
The N-terminal domain is roughly 40 residues in length and is predicted to contain both α-helices and β-sheets (Li et al. 1992, Li et al. 1993, Lacey et al. 1998). Despite these findings, the protein folding software AlphaFold predicts the N-terminal to be disordered (Fig. 3b). Similarly, the RaptorX models (Fig. 3b) of the N-terminal domain from almond, hemp, and sunflower oleosins were predicted to be largely disordered, although a short helix is predicted. The C-terminal domain, which is ~ 65 residues in length, is α-helical based on circular dichroism spectroscopy and Fourier transform infrared spectroscopy experiments (Li et al. 1992, Lacey et al. 1998). The C-terminal appears to have positively charged residues spaced periodically throughout the primary sequence (Fig. 3a). These positively charged residues are thought to be on the underside of the helix and interact with the negatively charged phospholipid heads to hold the terminal end to the oil body membrane (Li et al. 1992, Tzen et al. 1992, Lacey et al. 1998). The models from AlphaFold and RaptorX suggest that the C-terminal domain contains an α-helix, but is largely disordered. It may be that the N- and C-terminal domains have a unique fold, or that they require the interactions with phospholipids to correctly fold.
The secondary structure of the H-domain is a bone of contention. Early research carried out on rapeseed oleosins proposed that the hydrophobic domain was made up of antiparallel β-strands based on circular dichroism and Fourier transform infrared spectroscopy data (Li et al. 1992). However, this was quickly contested with contrasting evidence from circular dichroism experiments demonstrating largely α-helical content in the H-domain of sunflower oleosins (Millichip et al. 1996). These studies used different methods of protein purification leading to debate on which method best represents the in vivo structure (Beisson et al. 1996). However, there have been further reports since of α-helix content in safflower and sunflower seed oleosins (Lacey et al. 1998, Alexander et al. 2002) and β-strand content in rapeseed oleosins (Li et al. 2002). Whether these differences are due to species or differences in protein preparation remains to be seen.
Those who support the β-strand hypothesis suggest there are two β-strands, one going down from the N-terminus and one coming back up from the hairpin loop to the C-terminus, in an antiparallel arrangement (Li et al. 1992; Tzen et al. 1992; Huang 1996; Li et al. 2002). Li et al. (2002), also predicted that the β-sheets of separate oleosins will interact with each other via hydrogen bonds between the β-sheets. Those who support the α-helix hypothesis propose there are two antiparallel α-helices (Alexander et al. 2002). The α-helical model has the advantage of ensuring the partial charges on the peptide backbone are not exposed to the hydrophobic environment and instead form hydrogen bonds through the helix and that any hydrophilic sidechains can hydrogen bond in the inter-helical space securing the two helices together (Alexander et al. 2002). They further propose that two conserved residues Thr67 and Thr97 are in this inter-helical space and are conserved due to their role in holding the helices together (Alexander et al. 2002). However, a protein sequence alignment (Fig. 3a) shows that these residues are not highly conserved but there are usually threonine residues within the domain. The AlphaFold model suggests that the H- domain is α-helical. Others have generated their own models of oleosins with similar results to AlphaFold (Huang and Huang 2017).
Despite the inconsistencies in the secondary structure of the H-domain, most researchers agree on the importance of the proline knot motif that creates that hairpin turn (Tzen et al. 1992; Hsieh and Huang 2004). All the polypeptide chains of oleosins currently under study have the same three proline residues and one serine residue in the same position (PX5SPX3P) in the middle of the central hydrophobic chain (Hsieh and Huang 2004) (Fig. 3b). The proline knot also looks to be essential in inserting oleosins into the oil body during oil-body formation in the ER (as mentioned above). The conformation of the proline knot is noted in AlphaFold to be difficult to predict; this is likely due to its unusual protein sequence. It is becoming clear that the only way to determine the structure of oleosins is experimentally.
The translation of oleosins in industry
Despite the many gaps in our understanding of oleosin structure and function, oleosins have found application in commercial settings and we highlight some recent examples.
Oleosins have the potential to aid drug delivery. Cancer medications have a reputation of being non- specific (Schilsky 2010; La Thangue and Kerr 2011) and can be hydrophobic which makes drug delivery challenging. Hydrophobic medicines are not easily delivered by oral or intravenous methods due to poor solubility, instability, and low membrane permeability (Porter et al. 2007; Savjani et al. 2012; Cho et al. 2018). Some have exploited oleosin stabilised oil bodies to create easier and more effective methods for delivering cancer medications to their intended site. Here, oleosins are a part of an artificial oil body which holds in its centre the hydrophobic medicine meant for treating the cancer (Chiang et al. 2018; Cho et al. 2018). Both Chiang et al. (2018) and Cho et al. (2018) also fused specific signalling proteins to the oleosins to target the oil body and drug to the cancer cells. Chiang et al. (2018) fused an epidermal growth factor receptor targeting motif to the N-terminal domain of the oleosin which targets cells with the epidermal growth factor receptor, commonly found in lung cancer cells. Cho et al. (2018) instead fused to the C-terminal domain of oleosins an immunoglobulin-binding protein, which binds antibodies that could target breast cancer cells. The artificial oil body contained carmustine, which is a hydrophobic cancer drug. Both methods exploited oleosins fused with ancillary proteins that target the oil body and its hydrophobic drug payload to specific cells.
Human fibroblast growth factor (hFGF) has been shown to aid in wound healing and hair growth (Jimenez and Rampy 1999, Braun et al. 2004, Jang 2005, Lin et al. 2015). However, it is difficult to express recombinantly and has poor thermal stability and poor transdermal absorption (Kovacs et al. 2006; Wang et al. 2007). The expression of oleosins fused with hFGF in plants has been suggested as a possible solution (Li et al. 2017, Cai et al. 2018). Studies on oil bodies with oleosins fused to hFGF9 and hFGF10 isolated from safflower seeds found that both proteins were still able to work effectively (Li et al. 2017, Cai et al. 2018). Mice treated with oil bodies containing oleosin-hFGF had improved wound healing and hair growth compared with just recombinant hFGF. The oleosin and oil body were able to stabilise the hFGF and therefore make its application more efficient. Similar studies have used human epidermal growth factor (hEGF), which has similar applications as hFGF (Mroczkowski and Ball 1990; Jahovic et al. 2004; Hee Na et al. 2006) and the harvested oil bodies can be directly given to the patient (Qiang et al. 2020).
Despite a limited understanding of the structure of oleosins, these examples clearly demonstrate an opportunity to revolutionise drug delivery systems. A deeper knowledge of the structure of oleosins will allow us to understand how oil bodies are formed and stabilised. This will inform future engineering efforts, to utilise oil body systems to their full potential.
References
Abell BM, High S, Moloney MM (2002) Membrane protein topology of oleosin is constrained by its long hydrophobic domain. J Biol Chem 277(10):8602–8610. https://doi.org/10.1074/jbc.M103712200
Abell BM, Holbrook LA, Abenes M, Murphy DJ, Hills MJ, Moloney MM (1997) Role of the proline knot motif in oleosin endoplasmic reticulum topology and oil body targeting. Plant Cell 9(8):1481–1493. https://doi.org/10.1105/tpc.9.8.1481
Acevedo-Fani A, Dave A, Singh H (2020). Nature-assembled structures for delivery of bioactive compounds and their potential in functional foods. Frontiers in Chemistry 8(844). https://doi.org/10.3389/fchem.2020.564021
Alexander LG, Sessions RB, Clarke AR, Tatham AS, Shewry PR, Napier JA (2002) Characterization and modelling of the hydrophobic domain of a sunflower oleosin. Planta 214(4):546–551. https://doi.org/10.1007/s004250100655
Beaudoin F, Napier JA (2002) Targeting and membrane-insertion of a sunflower oleosin in vitro and in Saccharomyces cerevisiae: the central hydrophobic domain contains more than one signal sequence, and directs oleosin insertion into the endoplasmic reticulum membrane using a signal anchor sequence mechanism. Planta 215(2):293–303. https://doi.org/10.1007/s00425-002-0737-1
Beaudoin F, Wilkinson BM, Stirling CJ, Napier JA (2000) In vivo targeting of a sunflower oil body protein in yeast secretory (sec) mutants. Plant J 23(2):159–170. https://doi.org/10.1046/j.1365-313x.2000.00769.x
Beevers H (1979) Microbodies in higher plants. Annu Rev Plant Physiol 30(1):159–193. https://doi.org/10.1146/annurev.pp.30.060179.001111
Beevers H (1980). The role of the glyoxylate cycle. Lipids: Structure and Function Elsevier: 117–130. https://doi.org/10.1016/B978-0-12-675404-9.50010-2
Beisson F, Ferte N, Noat G (1996). Oil-bodies from sunflower (Helianthus annuus L.) seeds. Biochemical Journal 317(Pt 3): 955. https://doi.org/10.1042/bj3170955
Braun S, Auf dem Keller U, Steiling H, Werner S (2004). Fibroblast growth factors in epithelial repair and cytoprotection. Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences 359(1445): 753–757. https://doi.org/10.1098/rstb.2004.1464.
Cai J, Wen R, Li W, Wang X, Tian H, Yi S, Zhang L, Li X, Jiang C, Li H (2018) Oil body bound oleosin-rhFGF9 fusion protein expressed in safflower (Carthamus tinctorius L.) stimulates hair growth and wound healing in mice. BMC Biotechnology 18(1):51. https://doi.org/10.1186/s12896-018-0433-2
Chapman KD, Trelease RN (1991) Acquisition of membrane lipids by differentiating glyoxysomes: role of lipid bodies. J Cell Biol 115(4):995–1007. https://doi.org/10.1083/jcb.115.4.995
Chiang C-J, Lin L-J, Wu C-P, Chen C-J, Chao Y-P (2018) Development of nanoscale oil bodies for targeted treatment of lung cancer. J Agric Food Chem 66(36):9438–9445. https://doi.org/10.1021/acs.jafc.8b02972
Cho H-Y, Lee T, Yoon J, Han Z, Rabie H, Lee K-B, Su WW, Choi J-W (2018) Magnetic oleosome as a functional lipophilic drug carrier for cancer therapy. ACS Appl Mater Interfaces 10(11):9301–9309. https://doi.org/10.1021/acsami.7b19255
Dave AC, Ye A, Singh H (2019) Structural and interfacial characteristics of oil bodies in coconuts (Cocos nucifera L.). Food Chem 276:129–139. https://doi.org/10.1016/j.foodchem.2018.09.125
Deruyffelaere C, Bouchez I, Morin H, Guillot A, Miquel M, Froissard M, Chardot T, D’Andrea S (2015) Ubiquitin-mediated proteasomal degradation of oleosins is involved in oil body mobilization during post-germinative seedling growth in Arabidopsis. Plant Cell Physiol 56(7):1374–1387. https://doi.org/10.1093/pcp/pcv056
Fang Y, Zhu R-L, Mishler BD (2014) Evolution of oleosin in land plants. PLoS ONE 9(8):e103806. https://doi.org/10.1093/pcp/pcv056
Frandsen GI, Mundy J, Tzen JTC (2001) Oil bodies and their associated proteins, oleosin and caleosin. Physiol Plant 112(3):301–307. https://doi.org/10.1034/j.1399-3054.2001.1120301.x
Hayashi Y, Hayashi M, Hayashi H, Hara-Nishimura I, Nishimura M (2001) Direct interaction between glyoxysomes and lipid bodies in cotyledons of the Arabidopsis thaliana ped1 mutant. Protoplasma 218(1–2):83–94. https://doi.org/10.1007/BF01288364
Hee Na D, Seok Youn Y, Bok Lee I, Ji Park E, Jeon Park C, Choon Lee K (2006) Effect of molecular size of PEGylated recombinant human epidermal growth factor on the biological activity and stability in rat wound tissue. Pharm Dev Technol 11(4):513–519. https://doi.org/10.1080/10837450600941053
Hills MJ, Watson MD, Murphy DJ (1993) Targeting of oleosins to the oil bodies of oilseed rape (Brassica napus L.). Planta 189(1):24–29. https://doi.org/10.1007/BF00201339
Hsiao ES, Tzen JT (2011) Ubiquitination of oleosin-H and caleosin in sesame oil bodies after seed germination. Plant Physiol Biochem 49(1):77–81. https://doi.org/10.1016/j.plaphy.2010.10.001
Hsieh K, Huang AH (2004) Endoplasmic reticulum, oleosins, and oils in seeds and tapetum cells. Plant Physiol 136(3):3427–3434. https://doi.org/10.1104/pp.104.051060
Huang A (1996) Oleosins and oil bodies in seeds and other organs. Plant Physiol 110(4):1055. https://doi.org/10.1104/pp.110.4.1055
Huang AHC (1992) Oil bodies and oleosins in seeds. Annu Rev Plant Physiol Plant Mol Biol 43(1):177–200. https://doi.org/10.1146/annurev.pp.43.060192.001141
Huang C-Y, Huang AH (2017) Unique motifs and length of hairpin in oleosin target the cytosolic side of endoplasmic reticulum and budding lipid droplet. Plant Physiol 174(4):2248–2260. https://doi.org/10.1104/pp.17.00366
Huang M-D, Huang AH (2015) Bioinformatics reveal five lineages of oleosins and the mechanism of lineage evolution related to structure/function from green algae to seed plants. Plant Physiol 169(1):453–470. https://doi.org/10.1104/pp.15.00634
Itabe H (2010) Intracellular lipid droplet-associated proteins: unique members and their biological functions foreword. Biol Pharm Bull 33(3):341–341. https://doi.org/10.1248/bpb.33.341
Jahovic N, Güzel E, Arbak S, Yeğen BÇ (2004) The healing-promoting effect of saliva on skin burn is mediated by epidermal growth factor (EGF): role of the neutrophils. Burns 30(6):531–538. https://doi.org/10.1016/j.burns.2004.02.007
Jang J-H (2005) Stimulation of human hair growth by the recombinant human keratinocyte growth factor-2 (KGF-2). Biotech Lett 27(11):749–752. https://doi.org/10.1007/s10529-005-5624-y
Jimenez PA, Rampy MA (1999) Keratinocyte growth factor-2 accelerates wound healing in incisional wounds. J Surg Res 81(2):238–242. https://doi.org/10.1006/jsre.1998.5501
Jolivet P, Acevedo F, Boulard C, d’Andréa S, Faure JD, Kohli A, Nesi N, Valot B, Chardot T (2013) Crop seed oil bodies: from challenges in protein identification to an emerging picture of the oil body proteome. Proteomics 13(12–13):1836–1849. https://doi.org/10.1002/pmic.201200431
Jolivet P, Aymé L, Giuliani A, Wien F, Chardot T, Gohon Y (2017) Structural proteomics: Topology and relative accessibility of plant lipid droplet associated proteins. J Proteomics 169:87–98. https://doi.org/10.1016/j.jprot.2017.09.005
Jolivet P, Roux E, d’Andrea S, Davanture M, Negroni L, Zivy M, Chardot T (2004) Protein composition of oil bodies in Arabidopsis thaliana ecotype WS. Plant Physiol Biochem 42(6):501–509. https://doi.org/10.1016/j.plaphy.2004.04.006
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera- Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589. https://doi.org/10.1038/s41586-021-03819-2
Kanazawa T, Morinaka H, Ebine K, Shimada TL, Ishida S, Minamino N, Yamaguchi K, Shigenobu S, Kohchi T, Nakano A, Ueda T (2020) The liverwort oil body is formed by redirection of the secretory pathway. Nat Commun 11(1):6152. https://doi.org/10.1038/s41467-020-19978-1
Kovacs D, Cota C, Cardinali G, Aspite N, Bolasco G, Amantea A, Torrisi MR, Picardo M (2006) Expression of keratinocyte growth factor and its receptor in clear cell acanthoma. Exp Dermatol 15(10):762–768. https://doi.org/10.1111/j.1600-0625.2006.00459.x
La Thangue NB, Kerr DJ (2011) Predictive biomarkers: a paradigm shift towards personalized cancer medicine. Nat Rev Clin Oncol 8(10):587–596. https://doi.org/10.1038/nrclinonc.2011.121
Lacey DJ, Wellner N, Beaudoin F, Napier JA, Shewry PR (1998) Secondary structure of oleosins in oil bodies isolated from seeds of safflower (Carthamus tinctorius L.) and sunflower (Helianthus annuus L.). Biochemical Journal 334(2):469–477. https://doi.org/10.1042/bj3340469
Li M, Keddie J, Smith L, Clark D, Murphy D (1993) Expression and characterization of the N-terminal domain of an oleosin protein from sunflower. J Biol Chem 268(23):17504–17512
Li M, Murphy DJ, Lee K-HK, Wilson R, Smith LJ, Clark DC, Sung J-Y (2002) Purification and structural characterization of the central hydrophobic domain of oleosin. J Biol Chem 277(40):37888–37895. https://doi.org/10.1074/jbc.M202721200
Li M, Smith L, Clark D, Wilson R, Murphy D (1992) Secondary structures of a new class of lipid body proteins from oilseeds. J Biol Chem 267(12):8245–8253
Li W, Yang J, Cai J, Wang H, Tian H, Huang J, Qiang W, Zhang L, Li H, Li X (2017) Oil body- bound oleosin-rhFGF-10: a novel drug delivery system that improves skin penetration to accelerate wound healing and hair growth in mice. Int J Mol Sci 18(10):2177. https://doi.org/10.3390/ijms18102177
Lin W-H, Xiang L-J, Shi H-X, Zhang J, Jiang L-P, Cai P-T, Lin Z-L, Lin B-B, Huang Y, Zhang H-L (2015). Fibroblast growth factors stimulate hair growth through β-catenin and Shh expression in C57BL/6 mice. BioMed Research International 2015. https://doi.org/10.1155/2015/730139
Lin Y-H, Wimer LT, Huang AH (1983) Lipase in the lipid bodies of corn scutella during seedling growth. Plant Physiol 73(2):460–463. https://doi.org/10.1104/pp.73.2.460
Loer DS, Herman EM (1993) Cotranslational integration of soybean (Glycine max) oil body membrane protein oleosin into microsomal membranes. Plant Physiol 101(3):993–998. https://doi.org/10.1104/pp.101.3.993
Maurer S, Waschatko G, Schach D, Zielbauer BI, Dahl J, Weidner T, Bonn M, Vilgis TA (2013) The role of intact oleosin for stabilization and function of oleosomes. J Phys Chem B 117(44):13872–13883. https://doi.org/10.1021/jp403893n
Millichip M, Tatham AS, Jackson F, Griffiths G, Shewry PR, Stobart AK (1996) Purification and characterization of oil-bodies (oleosomes) and oil-body boundary proteins (oleosins) from the developing cotyledons of sunflower (Helianthus annuus L.). Biochemical Journal 314(1):333–337. https://doi.org/10.1042/bj3140333
Mroczkowski B, Ball R (1990). Epidermal growth factor: Biology and properties of its gene and protein precursor. Growth Factors, Differentiation Factors, and Cytokines, Springer: 18-30. https://doi.org/10.1007/978-3-642-74856-1_2
Murphy DJ (1993) Structure, function and biogenesis of storage lipid bodies and oleosins in plants. Prog Lipid Res 32(3):247–280. https://doi.org/10.1016/0163-7827(93)90009-l
Murphy DJ (2001) The biogenesis and functions of lipid bodies in animals, plants and microorganisms. Prog Lipid Res 40(5):325–438. https://doi.org/10.1016/s0163-7827(01)00013-3
Murphy DJ, Cummins I (1989) Seed oil-bodies: isolation, composition and role of oil-body apolipoproteins. Phytochemistry 28(8):2063–2069
Nikiforidis CV, Matsakidou A, Kiosseoglou V (2014) Composition, properties and potential food applications of natural emulsions and cream materials based on oil bodies. RSC Adv 4(48):25067–25078. https://doi.org/10.1039/C4RA00903G
Parthibane V, Iyappan R, Vijayakumar A, Venkateshwari V, Rajasekharan R (2012a) Serine/threonine/tyrosine protein kinase phosphorylates oleosin, a regulator of lipid metabolic functions. Plant Physiol 159(1):95–104. https://doi.org/10.1104/pp.112.197194
Parthibane V, Rajakumari S, Venkateshwari V, Iyappan R, Rajasekharan R (2012b) Oleosin is bifunctional enzyme that has both monoacylglycerol acyltransferase and phospholipase activities. J Biol Chem 287(3):1946–1954. https://doi.org/10.1074/jbc.M111.309955
Payne G, Lad M, Foster T, Khosla A, Gray D (2014) Composition and properties of the surface of oil bodies recovered from Echium plantagineum. Colloids Surf, B 116:88–92
Peng C-C, Lee VS, Lin M-Y, Huang H-Y, Tzen JT (2007) Minimizing the central hydrophobic domain in oleosin for the constitution of artificial oil bodies. J Agric Food Chem 55(14):5604–5610. https://doi.org/10.1021/jf070977o
Pons L, Chéry C, Mrabet N, Schohn H, Lapicque F, Guéant J-L (2005) Purification and cloning of two high molecular mass isoforms of peanut seed oleosin encoded by cDNAs of equal sizes. Plant Physiol Biochem 43(7):659–668. https://doi.org/10.1016/j.plaphy.2005.06.002
Porter CJ, Trevaskis NL, Charman WN (2007) Lipids and lipid-based formulations: optimizing the oral delivery of lipophilic drugs. Nat Rev Drug Discovery 6(3):231–248. https://doi.org/10.1038/nrd2197
Purkrtova Z, Jolivet P, Miquel M, Chardot T (2008) Structure and function of seed lipid body- associated proteins. CR Biol 331(10):746–754. https://doi.org/10.1016/j.crvi.2008.07.016
Qiang W, Gao T, Lan X, Guo J, Noman M, Li Y, Guo Y, Kong J, Li H, Du L (2020) Molecular pharming of the recombinant protein hEGF-hEGF concatenated with oleosin using Transgenic Arabidopsis. Genes 11(9):959. https://doi.org/10.3390/genes11090959
Quettier A-L, Eastmond PJ (2009) Storage oil hydrolysis during early seedling growth. Plant Physiol Biochem 47(6):485–490. https://doi.org/10.1016/j.plaphy.2008.12.005
Ramachandiran I, Vijayakumar A, Ramya V, Rajasekharan R (2018) Arabidopsis serine/threonine/tyrosine protein kinase phosphorylates oil body proteins that regulate oil content in the seeds. Sci Rep 8(1):1154. https://doi.org/10.1038/s41598-018-19311-3
Ratnayake C, Huang AHC (1996) Oleosins and oil bodies in plant seeds have postulated structures. Biochem J 317:956–958
Sadeghipour HR, Bhatla SC (2002) Differential sensitivity of oleosins to proteolysis during oil body mobilization in sunflower seedlings. Plant Cell Physiol 43(10):1117–1126. https://doi.org/10.1093/pcp/pcf142
Sarmiento C, Ross JH, Herman E, Murphy DJ (1997). Expression and subcellular targeting of a soybean oleosin in transgenic rapeseed. Implications for the mechanism of oil‐body formation in seeds. The Plant Journal 11(4): 783–796. https://doi.org/10.1046/j.1365-313x.1997.11040783.x
Savjani KT, Gajjar AK, Savjani JK (2012) Drug solubility: importance and enhancement techniques. International Scholarly Research Notices 2012:195727. https://doi.org/10.5402/2012/195727
Schilsky RL (2010) Personalized medicine in oncology: the future is now. Nat Rev Drug Discovery 9(5):363–366. https://doi.org/10.1038/nrd3181
Shimada TL, Shimada T, Takahashi H, Fukao Y, Hara-Nishimura I (2008) A novel role for oleosins in freezing tolerance of oilseeds in Arabidopsis thaliana. Plant J 55(5):798–809. https://doi.org/10.1111/j.1365-313X.2008.03553.x
Song Y, Wang X-D, Rose RJ (2017) Oil body biogenesis and biotechnology in legume seeds. Plant Cell Rep 36(10):1519–1532. https://doi.org/10.1007/s00299-017-2201-5
Tzen J, Huang A (1992) Surface structure and properties of plant seed oil bodies. J Cell Biol 117(2):327–335. https://doi.org/10.1083/jcb.117.2.327
Tzen J, Lie G, Huang A (1992) Characterization of the charged components and their topology on the surface of plant seed oil bodies. J Biol Chem 267(22):15626–15634
Tzen JT, Cao Y, Laurent P, Ratnayake C, Huang AH (1993) Lipids, proteins, and structure of seed oil bodies from diverse species. Plant Physiol 101(1):267–276. https://doi.org/10.1104/pp.101.1.267
Tzen JT, Chuang RL, Chen JC, Wu LS (1998) Coexistence of both oleosin isoforms on the surface of seed oil bodies and their individual stabilization to the organelles. The Journal of Biochemistry 123(2):318–323. https://doi.org/10.1093/oxfordjournals.jbchem.a021939
Tzen JT, Lai Y-K, Chan K-L, Huang AH (1990) Oleosin isoforms of high and low molecular weights are present in the oil bodies of diverse seed species. Plant Physiol 94(3):1282–1289. https://doi.org/10.1104/pp.94.3.1282
van Rooijen GJ, Moloney MM (1995) Structural requirements of oleosin domains for subcellular targeting to the oil body. Plant Physiol 109(4):1353–1361. https://doi.org/10.1104/pp.109.4.1353
Vandana S, Bhatla S (2006) Evidence for the probable oil body association of a thiol-protease, leading to oleosin degradation in sunflower seedling cotyledons. Plant Physiol Biochem 44(11–12):714–723. https://doi.org/10.1016/j.plaphy.2006.09.022
Wang Y, Yuan S, Wang P, Liu X, Zhan D, Zhang Z (2007) Expression, purification, and characterization of recombinant human keratinocyte growth factor-2 in Pichia pastoris. J Biotechnol 132(1):44–48. https://doi.org/10.1016/j.jbiotec.2007.08.024
Wanner G, Formanek H, Theimer R (1981) The ontogeny of lipid bodies (spherosomes) in plant cells. Planta 151(2):109–123. https://doi.org/10.1007/BF00387812
Waschatko G, Billecke N, Schwendy S, Jaurich H, Bonn M, Vilgis TA, Parekh SH (2016) Label- free in situ imaging of oil body dynamics and chemistry in germination. J R Soc Interface 13(123):20160677. https://doi.org/10.1098/rsif.2016.0677
White DA, Fisk ID, Mitchell JR, Wolf B, Hill SE, Gray DA (2008) Sunflower-seed oil body emulsions: Rheology and stability assessment of a natural emulsion. Food Hydrocolloids 22(7):1224–1232. https://doi.org/10.1016/j.foodhyd.2007.07.004
Wong H, Schotz MC (2002) The lipase gene family. J Lipid Res 43(7):993–999. https://doi.org/10.1194/jlr.r200007-jlr200
Xu J, McPartlon M, Li J (2021) Improved protein structure prediction by deep learning irrespective of co-evolution information. Nature Machine Intelligence 3(7):601–609. https://doi.org/10.1038/s42256-021-00348-5
Yatsu L, Jacks T (1972) Spherosome membranes: half unit-membranes. Plant Physiol 49(6):937–943. https://doi.org/10.1104/pp.49.6.937
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Board, A.J., Crowther, J.M., Acevedo-Fani, A. et al. How plants solubilise seed fats: revisiting oleosin structure and function to inform commercial applications. Biophys Rev 14, 257–266 (2022). https://doi.org/10.1007/s12551-021-00923-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-021-00923-5