Abstract
Background
The aberrant expression of stromal gene signatures in breast cancer has been widely studied. However, the association of stromal gene signatures with tumor immunity, progression, and clinical outcomes remains lacking.
Methods
Based on eight breast tumor stroma (BTS) transcriptomics datasets, we identified differentially expressed genes (DEGs) between BTS and normal breast stroma. Based on the DEGs, we identified dysregulated pathways and prognostic hub genes, hub oncogenes, hub protein kinases, and other key marker genes associated with breast cancer. Moreover, we compared the enrichment levels of stromal and immune signatures between breast cancer patients with bad and good clinical outcomes. We also investigated the association between tumor stroma-related genes and breast cancer progression.
Results
The DEGs included 782 upregulated and 276 downregulated genes in BTS versus normal breast stroma. The pathways significantly associated with the DEGs included cytokine–cytokine receptor interaction, chemokine signaling, T cell receptor signaling, cell adhesion molecules, focal adhesion, and extracellular matrix–receptor interaction. Protein–protein interaction network analysis identified the stromal hub genes with prognostic value in breast cancer, including two oncogenes (COL1A1 and IL21R), two protein kinases encoding genes (PRKACA and CSK), and a growth factor encoding gene (PLAU). Moreover, we observed that the patients with bad clinical outcomes were less enriched in stromal and antitumor immune signatures (CD8 + T cells and tumor-infiltrating lymphocytes) but more enriched in tumor cells and immunosuppressive signatures (MDSCs and CD4 + regulatory T cells) compared with the patients with good clinical outcomes. The ratios of CD8 + /CD4 + regulatory T cells were lower in the patients with bad clinical outcomes. Furthermore, we identified the tumor stroma-related genes, including MCM4, SPECC1, IMPA2, and AGO2, which were gradually upregulated through grade I, II, and III breast cancers. In contrast, COL14A1, ESR1, SLIT2, IGF1, CH25H, PRR5L, ABCA6, CEP126, IGDCC4, LHFP, MFAP3, PCSK5, RAB37, RBMS3, SETBP1, and TSPAN11 were gradually downregulated through grade I, II, and III breast cancers. It suggests that the expression of these stromal genes has an association with the progression of breast cancers. These progression-associated genes also displayed an expression association with recurrence-free survival in breast cancer patients.
Conclusions
This study identified tumor stroma-associated biomarkers correlated with deregulated pathways, tumor immunity, tumor progression, and clinical outcomes in breast cancer. Our findings provide new insights into the pathogenesis of breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
The tumor microenvironment (TME) consists of heterogeneous components including borders, blood vessels, lymph vessels, extracellular matrix (ECM), immune and/or inflammatory cells, secretomes, coding or non-coding RNAs, small organelles, tumor cells, and surrounding associated or regulatory cells [1]. Among the tumor-associated regulatory cells, stromal cells playing various crucial oncogenic roles in the TME. In the TME, heterogeneous stromal cells are associated with tumor growth, invasion, progression, and metastasis [2, 3]. These tumor-associated stromal cells promote various dysregulated biological functions including extracellular matrix remodeling, cellular migration, neoangiogenesis, and evasion of immunosurveillance through the production of several types of onco-regulators including cytokines, chemokines, matrix metalloproteinases (MMPs), extracellular matrix (ECM), and growth factors [4]. It is recently demonstrated that the tumor-associated stromal cells playing pivotal roles in the resistance of cancer therapy [5]. The oncogenic intrinsic properties of stromal cells substantially regulated the genotype and phenotype of surrounding cancer cells in the TME [6].
In the breast cancer TME, tumor-associated stromal cells are associated with cancer initiation, development, progression, angiogenesis, metastasis, recurrence, and therapeutic resistance [7]. Survival of breast cancer patients is correlated with stromal biology including the reorganization of the extracellular matrix (ECM) to promote cancer invasion and migration, changes in the phenotypes of stromal cell, variability in the stromal gene expression profiles, and changes in cellular signaling cascades to aid surrounding cancer cells [8]. Various parts of stromal compartments have crucial effects on breast cancer TME. The tumor-promoting intrinsic properties of the stroma are associated with the tumorigenesis of breast cancer [7]. Finak et al. reported that the expression of stromal gene signatures is correlated with the clinical outcomes of breast cancer patients [9]. Winslow et al. identified stromal gene signatures that are associated with clinical features in different types of molecular subtypes of breast cancer [10]. Altogether, these studies provide the clue that the stromal cells have substantial onco-regulatory roles in the TME of breast cancer.
Herein, we performed comprehensive bioinformatic analyses to identify molecular alterations in breast tumor stroma versus normal stroma. We identified differentially expressed genes (DEGs), hub genes from the interactions of DEGs, and regulatory transcription factors (TFs), and kinases and pathways associated with the DEGs. We also identified stromal genes having a significant link with recurrence-free survival in breast cancer patients. Moreover, we found certain tumor stromal genes, which were gradually dysregulated through the three different grades of breast cancer, and their dysregulation was associated with poor prognosis in patients.
Materials and methods
Data selection and pre-processing
We systematically searched for the gene expression omnibus (GEO) database using keywords “breast cancer,” “stroma,” and “tumor stroma”. Ultimately, we identified eight datasets: GSE9014 (sample size n = 123) [9], GSE83591 (n = 53) [11], GSE31192 (n = 17) [12], GSE26910 (n = 12) [13], GSE10797 (n = 33) [14], GSE8977 (n = 22) [15], GSE33692 (n = 22) [16], and GSE14548 (n = 34) [17] (Supplementary Table S1). We combined the eight datasets into a single dataset (including stromal data and excluding other data) using the NetworkAnalyst software [18]. The ComBat method was utilized to remove batch effects from the eight datasets [19] (effects of batch removal shown in Supplementary Fig. S1). Each dataset was normalized by base-2 log transformation or quantile normalization. The combined dataset included 240 tumor stroma and 76 normal stroma samples. We also downloaded the dataset of gene expression profiles (RSEM normalized) in TCGA breast cancer cohort (n = 1212) from the genomic data commons (GDC) data portal (https://portal.gdc.cancer.gov/). We further normalized the RSEM gene expression values by base-2 log transformation [20]. In addition, we used the clinical data of a breast tumor stroma cohort (GSE9014) to evaluate the survival time differences in breast cancer patients [9].
Identification of DEGs between breast tumor stroma and normal stroma by a meta-analysis
We employed the R package “limma” to identify the DEGs between BTS and normal stroma [21]. A meta-analysis of the eight datasets was performed using Cochran’s combination test [22]. The false discovery rate (FDR), calculated by the Benjamini–Hochberg method [23], was used to adjust for multiple tests. We selected the all DEGs with a threshold of absolute value with combined effect size (ES) > 0.41 and FDR < 0.05.
Pathway and functional enrichment analysis
We performed pathway enrichment analysis of the set of genes that were differentially expressed between BTS and normal stroma using the GSEA software [24]. The KEGG pathways [25] significantly associated with the set of genes upregulated and downregulated in BTS versus normal stroma were identified, respectively, using a threshold of FDR < 0.05.
Identification of TFs, protein kinases, and master transcriptional regulators (MTRs) that are significantly associated with the DEGs
To link gene expression signatures to upstream cell signaling networks, we used the eXpression2Kinases algorithm [26] to identify the upstream TFs and kinases that regulate the DEGs. In the eXpression2Kinases algorithm, we used a threshold of hypergeometric P value ≤ 0.05 for identifying upstream TFs and kinases. Besides, we utilized the Cytoscape plug-in iRegulon [27] to identify the MTRs for the upregulated and downregulated DEGs, with a threshold of the minimum normalized enrichment score (NES) > 3.0 which corresponds to an approximate FDR between 3 and 9%.
Protein–protein interactions (PPIs)
We constructed PPI networks of the DEGs using STRING (version v11 [28]). We input all DEGs into STRING. The rank of genes was identified by the Cytoscape plugin cytoHubba [29]. Hub nodes were identified using a threshold of medium interaction score ≥ 0.40 and we selected the degree of interaction ≥ 25 for identifying most interacted genes in the PPI. We identified hub genes, protein kinases, oncogenes, and tumor suppressor genes (TSGs) by comparing the hub nodes with TFs, protein kinases, oncogenes, and TSGs obtained from GSEA (https://www.gsea-msigdb.org/gsea/index.jsp). The online tool “Calculate and draw custom Venn diagrams” (http://bioinformatics.psb.ugent.be/webtools/Venn/) was used to identify common TF encoding genes, protein kinase encoding genes, oncogenes, and TSGs between different groups. We visualized the PPI networks using Cytoscape (version 3.6.1) [30].
Evaluation of immune scores, stromal scores, and tumor purity in stromal breast cancer subtypes
We utilized the “ESTIMATE” R package to calculate immune score representing the enrichment levels of immune cells, stromal score representing content of stromal cells, and tumor purity for each breast tumor sample [31] in (GSE9014 [9]). We compared immune scores, stromal scores, and tumor purity between the patients without disease recurrence and the patients with disease recurrence. We consider the Wilcoxon sum rank test P value ≤ 0.05 for identifying significant difference between both groups.
Quantification of the enrichment levels of immune and stromal signatures
We used the single-sample gene-set enrichment analysis (ssGSEA) to quantify the enrichment scores of immune and stromal signatures in tumors based on the expression levels of their marker genes [32]. We defined the ratio of immune signatures in a tumor sample as the ratio of the average expression levels of their marker genes. The immune and stromal signatures analyzed included B cells, CD8 + T cells, CD4 + regulatory T cells, macrophages, neutrophil, natural killer (NK) cells, tumor-infiltrating lymphocytes (TILs), regulatory T cells (Tregs), cytolytic activity, T cell activation, T cell exhaustion, T follicular helper cells (Tfh), M2 macrophages, tumor-associated macrophage (TAM), myeloid-derived suppressor cell (MDSC), endothelial cell, and cancer-associated fibroblasts (CAFs). Their marker genes are shown in Supplementary Table S2.
Survival analysis
We used the clinical data of a BTS cohort (GSE9014) which involved 53 breast cancer patients with clinical information available [9] (Supplementary Table S3) for survival analysis. We compared the recurrence-free survival (RFS) between breast cancer patients classified based on gene expression levels (expression levels > median versus expression levels < median). Kaplan–Meier survival curves were used to show the survival time differences, and the log-rank test was utilized to evaluate the significance of survival time differences between both groups. We used the function “survfit” in the R package “survival” to perform survival analysis and the function “coxph” in the R package “survival” for the univariate and multivariable Cox regression analyses [33].
Identification of DEGs between breast cancer patients with different tumor grades, clinical outcomes, and survival prognosis
In the GSE9014 database [9], we identified the DEGs between the breast cancer patients without and with disease recurrence and the DEGs among the breast cancer patients with different grades (grade I, grade II, and grade III) (Student’s t test, P < 0.05). We then identified the common genes between both groups of DEGs. We further analyzed the association of the expression of these common genes with the RFS of breast cancer patients. To identify the DEGs among the breast cancer patients with different grades, we utilized the R package “multcomp” [34].
Statistical and computational analysis
We used the two-tailed Student’s t test to compare two classes of normally distributed data, including gene expression levels and the ratios of immune signatures, and the one-tailed Mann–Whitney U test to compare two classes of data that were not normally distributed, including immune scores, stromal scores, tumor purity, and ssGSEA scores. The FDR evaluated by the Benjamini–Hochberg method [23] was used to adjust for multiple tests. We used the R package “ggplot2” to visualize the plots. For multiple probes of a single gene, we averaged the expression values of all probes into a single value by NetworkAnalyst [18]. The online tool “Calculate and draw custom Venn diagrams” (http://bioinformatics.psb.ugent.be/webtools/Venn/) was used to identify common genes between the different groups.
Results
Identification of DEGs between BTS and normal stroma
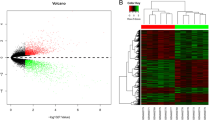
We identified 1058 DEGs between BTS and normal stroma. Among these DEGs, 782 were upregulated (Supplementary Table S4) and 276 were downregulated (Supplementary Table S5) in BTS. The top 25 upregulated (the highest ES) genes included COL10A1, SULF1, INHBA, NOX4, COMP, COL11A1, RAB31, IFI30, COL8A1, CTSB, LRRC15, SDC1, WISP1, LAMP5, LEF1, ASPN, MSR1, MNDA, SLAMF8, UNC5B, SLA, TYROBP, C3AR1, ITGAX, and COL8A2 (Table 1). Among them, the upregulation of COL11A1 and IFI30 was associated with a worse prognosis in breast cancer patients (Fig. 1A–C). In addition, the top 25 downregulated (the lowest ES) genes included FIGF, SPRY2, DLK1, SFRP1, TGFBR3, HLF, CD36, GPC3, LIFR, CAPN6, RELN, AKR1C3, CAV1, PLP1, MATN2, SDPR, SOCS2, ITM2A, LDB2, SYNM, EGFR, NACA, NOVA1, SPTBN1, and SEMA3G in the BTS (Table 1). Among these genes, the downregulation of SPRY2, CAV1, SOCS2, ITM2A, LDB2, and NACA in BTS was associated with worse prognosis of breast cancer patients (Fig. 1A, D–I).
Expression levels of eight prognostic genes in BTS and their associations with survival prognosis in breast cancer. A Expression of eight prognostic genes in BTS versus normal stroma. We investigated the survival of top 25 upregulated and top 25 downregulated genes. The upregulation of COL11A1 and IFI30 (upregulated in BTS) is associated with a worse prognosis (B, C). The downregulation of SPRY2, CAV1, SOCS2, ITM2A, LDB2, and NACA (downregulated in BTS) is associated with a worse prognosis (D–I)
Identifications of pathways significantly associated with the breast tumor stromal DEGs
GSEA [24] identified 82 KEGG pathways [25] significantly associated with the DEGs upregulated in BTS (Fig. 2A and Supplementary Table S6). Among them, the top 20 (the lowest FDR) pathways are displayed in Fig. 2A. These pathways were mainly involved in immune, stromal signatures, including cytokine–cytokine receptor interaction, Toll-like receptor signaling, antigen processing and presentation, chemokine signaling, T cell receptor signaling, B cell receptor signaling, natural killer cell-mediated cytotoxicity, leukocyte transendothelial migration, hematopoietic cell lineage, complement and coagulation cascades, Fc gamma R-mediated phagocytosis, Fc epsilon RI signaling pathway, NOD-like receptor signaling, Jak-STAT signaling pathway, cytosolic DNA-sensing, RIG-I-like receptor signaling, cell adhesion molecules (CAMs), focal adhesion, ECM–receptor interaction, regulation of actin cytoskeleton, adherens junction, tight junction, and gap junction. Moreover, many cancer-associated pathways were included in the 82 pathways, including MAPK signaling, TGF-beta, VEGF signaling, calcium signaling, mTOR signaling, and apoptosis. Besides, we identified 67 KEGG pathways [25] associated with the DEGs downregulated in BTS. The top 20 (the lowest FDR) pathways are displayed in Fig. 2B and Supplementary Table S7. The downregulated pathways are mainly associated with metabolism (propanoate metabolism, tryptophan metabolism, valine, leucine and isoleucine degradation, lysine degradation, beta-alanine metabolism, limonene and pinene degradation, arachidonic acid metabolism, butanoate metabolism, fatty acid metabolism, histidine metabolism, metabolism of xenobiotics by cytochrome P450, drug metabolism-cytochrome P450, pyruvate metabolism, glycerolipid metabolism, arginine and proline metabolism, steroid hormone biosynthesis, retinol metabolism, ascorbate and aldarate metabolism, linoleic acid metabolism, glycine, serine and threonine metabolism, alanine, aspartate and glutamate metabolism, ether lipid metabolism, glycolysis/gluconeogenesis, etc.), cancers (pathways in cancer, small cell lung cancer, endometrial cancer, non-small cell lung cancer, bladder cancer, thyroid cancer, pancreatic cancer, renal cell carcinoma, prostate cancer, acute myeloid leukemia, colorectal cancer, etc.), and cellular signaling and development (ribosome, PPAR signaling pathway, neurotrophin signaling pathway, insulin signaling pathway, spliceosome, p53 signaling pathway, Wnt signaling pathway, etc.). Altogether, our pathway analysis underlines that the stromal gene signatures are associated with the alteration of pathways that regulating tumor immunity, cellular signaling, metabolism, and cancers.
KEGG pathways are significantly associated with the upregulated and downregulated genes in BTS versus normal stroma identified by GSEA [24]. A Top 20 pathways significantly associated with the DEGs upregulated in BTS. B Top 20 pathways significantly associated with the DEGs downregulated in BTS. FDR: false discovery rate
Upstream TFs, kinases, and MTRs regulating the DEGs
The eXpression2Kinases algorithm identified 20 upstream TFs playing a significant regulatory role toward the DEGs (Supplementary Table S8). These TFs included IRF8, NFE2L2, TP63, RUNX1, SPI1, SMAD4, TRIM28, GATA2, AR, SUZ12, EGR1, KLF4, GATA1, RELA, TCF3, PPARD, RCOR1, TP53, SALL4, and NANOG. Interestingly, among the 20 upstream TFs, the genes encoding IRF8, PPARD, and RUNX1 were significantly upregulated in BTS and the gene encoding KLF4 was significantly downregulated in BTS (Supplementary Fig. S2A). IRF8 is a tumor suppressor involved in the regulation of the signaling of breast cancer cells [35]. Dysregulation of NFE2L2 is correlated with poor outcomes in breast cancer patients [36]. In the harsh metabolic conditions of the TME, PPARD promotes the survival of breast cancer cells [37]. In triple-negative breast cancer (TNBC), the expression of RUNX1 is correlated with the poor survival prognosis [38]. One of the isoforms of KLF4 is associated with the carcinogenesis of breast cancer [39]. Besides, we identified 116 upstream protein kinases, including CDK1, LYN, CSNK2A1, MAPK3, RPS6KA1, PRKACA, and TGFBR2 (Supplementary Table S9). Among these kinases, the genes encoding DYRK2, LYN, ERBB2, RPS6KA1, and PRKACA were significantly upregulated in BTS and the gene encoding PKD1, RPS6KA5, EGFR, and FOXO3 were significantly downregulated in BTS (Supplementary Fig. S2B). CSNK2A1 expression levels are significantly higher in a basal subtype of breast cancer [40].
MAPKs are associated with the downstream oncogenic signaling pathways in breast tumorigenesis [41]. Furthermore, we identified 13 MTRs, including IRF8, ETV7, STAT2, SPI1, IRF1, RELA, STAT1, IKZF1, RUNX1, IRF7, NFKB1, NKX3-2, and BATF, which were involved in the regulation of the genes upregulated in BTS (Fig. 3A and Supplementary Table S10). We also identified 11 MTRs targeting the genes downregulated in BTS, including HLF, FOXP2, JUND, NANOS1, RBBP9, FOS, TAF1, FOSL1, JUN, HAND1, and FOXJ3 (Fig. 3B and Supplementary Table S10). Interestingly, the genes encoding the MTRs RUNX1, IRF7, ETV7, STAT2, and IRF8 were upregulated in BTS (Fig. 3A), and the genes encoding the MTRs HLF and FOXJ3 were downregulated in BTS (Fig. 3B). Altogether, these results indicate that a number of TFs and protein kinases play significant roles in regulating the breast cancer stromal gene signatures and are associated with the pathogenesis of breast cancer.
Regulatory networks of the master transcriptional regulators (MTRs) and their targeted differentially expressed genes (DEGs) between BTS and normal stroma. A Regulatory network of the MTRs and their targeted upregulated genes in BTS. B Regulatory network of the MTRs and their targeted downregulated genes in BTS. In the center, green color octagon indicates MTRs, and purple color oval indicates DEGs
Identification of prognostic hub genes in breast tumor stroma
To identify the hub genes of the DEGs in BTS, we input all the DEGs into the STRING tool [28]. We identified 233 hub genes (degree ≥ 25), including 194 upregulated and 39 downregulated genes in BTS (Supplementary Table S11 and Supplementary Fig. S3). Finally, we displayed the top 50 hub genes (EGFR, TLR4, ITGAM, IL10, TLR2, CD86, IL1B, MMP9, ITGB2, TLR8, TLR7, ITGAX, MYC, CXCL10, TYROBP, CXCR4, IRF8, TLR3, CASP3, CTLA4, CSF1R, PLEK, LCP2, CD80, C3AR1, MYD88, IL10RA, PIK3R1, CYBB, SYK, SELL, FCGR2A, CXCL9, CCR7, CCR1, LYN, IRF7, CXCL1, PTGS2, RAC2, ERBB2, FCER1G, ISG15, HCK, CXCR3, CD4, IL7, FCGR2B, COL1A1, and OASL) in Fig. 4A. We found that the upregulation of hub genes MMP9, FCER1G, CD86, ITGAM, TLR2, and COL1A1 (upregulated DEGs in BTS) was significantly associated with poor RFS (Fig. 5A–F). These data indicate that the dysregulation of breast cancer stromal hub genes is likely to be associated with poor prognosis in breast cancer patients.
Protein–protein interactions of prognostic hub genes in BTS. A Protein–protein interaction network of top 50 hub genes. The hub genes in yellow are associated with poor prognosis in breast cancer. B Two prognostic hub oncogenes (IL21R and COL1A1) interact with other hub genes. C Interactions of two prognostic protein kinases encoding genes (PRKACA and CSK) with other hub genes. D Prognostic growth factor encoding gene PLAU is a hub gene that interacts with other stromal hub genes
Protein–protein interaction network analysis identifies prognostic hub genes in BTS. A–F High expression of the top hub genes is associated with poor prognosis in breast cancer. F–G The elevated expression of oncogenic hub genes COL1A1 and IL21R is associated with poor prognosis in breast cancer. H–I The elevated expression of protein kinases encoding genes (PRKACA and CSK) is correlated with poor prognosis in breast cancer. J The elevated expression of growth factor encoding gene PLAU is linked with poor prognosis in breast cancer. The survival analysis is performed in the breast cancer dataset GSE9014
Hub oncogenes, protein kinases encoding genes, and cytokines and growth factor-encoding genes are associated with poor survival prognosis in BTS
We identified the hub genes belonging to four gene families, including oncogenes, genes encoding protein kinases, genes encoding cytokines and growth factors, and tumor suppressor genes (Supplementary Fig. S3). We found 17 oncogenic hub genes, including CD74, CIITA, CLTC, COL1A1, ERBB2, FCGR2B, IL21R, MUC1, SYK (upregulated in BTS), and EGFR, EPS15, FOXO3, MET, MYC, PPARG, RPL22, ZBTB16 (downregulated in BTS). Also, we found 11 protein kinases encoding genes (CSF1R, CSK, EIF2AK2, HCK, LYN, PRKACA, RNASEL, SYK, and ERBB2 (upregulated in BTS) and EGFR and MET (downregulated in BTS), 20 cytokines and growth factors encoding genes (CCL11, CCL7, CMTM6, CXCL10, CXCL11, CXCL9, IL10, IL16, IL1B, IL1RN, IL7, OSM, PLAU, PMCH, and TNFSF4 (upregulated in BTS) and CAT, CCL27, CXCL1, CXCL2, and CXCL3 (downregulated in BTS), and 2 tumor suppressors genes (upregulated TNFAIP3 and downregulated PIK3R1).
We investigated the association of these hub genes with survival prognosis. Besides, we investigated the specific gene-family-centric PPI of prognostic hub oncogenes (IL21R and COL1A1) (Fig. 4B), hub protein kinase genes (PRKACA and CSK) (Fig. 4C), and hub cytokines-and-growth-factor genes (PLAU) (Fig. 4D) with other stromal hub genes. We revealed that these families of genes were interacted with other stromal hub genes (Fig. 4B–D), indicating their regulatory roles in the TME of breast cancer. Survival analysis revealed that the upregulation of two oncogenes (IL21R and COL1A1), two protein kinase genes (PRKACA and CSK), and a cytokine and growth factor gene (PLAU) is associated with shorter RFS in breast cancer patients (Fig. 5F–J). Altogether, these results indicate that the dysregulation of many tumor stroma-derived gene signatures is associated with unfavorable clinical outcomes in breast cancer patients.
Comparisons of immune and stromal signatures between breast cancer patients with good and bad clinical outcomes
We found that stromal scores were lower in the breast cancer patients with bad clinical outcomes (Wilcoxon sum rank test, P ≤ 0.05) (Fig. 6A). In contrast, tumor purity was higher in breast cancer patients with bad clinical outcomes (Fig. 6A). Interestingly, the enrichment scores (ssGSEA scores) of CD8 + T cells (P = 0.007), TILs (P = 0.03), and endothelial cells (P = 0.05) were lower in breast cancer patients with bad clinical outcomes than in those with good clinical outcomes (Fig. 6B). In contrast, MDSCs (P = 0.05) is more highly enriched in the breast cancer patients with bad clinical outcomes (Fig. 6B). The ratios of CD8 + /CD4 + regulatory T cells were lower in the breast cancer patients with bad clinical outcomes (Student’s t test, P = 4.5 × 10–05) (Fig. 6C). These results indicate that increased immune-promoting signatures are associated with better prognosis in breast cancer, while increased immunosuppressive signatures are associated with worse prognosis. This is consistent with the findings of previous studies [42,43,44,45,46].
Comparisons of immune and stromal signatures between breast cancer patients with good and bad clinical outcomes. A Comparisons of stromal scores and tumor purity between the bad and good clinical outcome groups of breast cancer patients. B The enrichment scores (ssGSEA scores) of CD8 + T cells, TILs, and endothelial cells are lower in the bad clinical outcome group. The enrichment scores of MDSCs is higher in the bad clinical outcome. C The ratios of CD8 + /CD4 + regulatory T cells are lower in the bad clinical outcome group
Stromal gene signatures significantly altered with the grades, clinical outcomes, and survival prognosis
We found 1955 DEGs among three grades (grade I, II, and III) of breast cancers (F-test, P < 0.05) (Fig. 7 and Supplementary Table S12). Besides, we found 1471 DEGs between the good clinical outcome (patients without disease recurrence) and bad clinical outcome (patients with disease recurrence) groups (Student’s t test, P < 0.05) (Fig. 7 and Supplementary Table S13). There were 124 common genes between both groups of DEGs (Fig. 7 and Supplementary Table S14). Furthermore, we found 20 of the 124 genes (MCM4, SPECC1, IMPA2, AGO2, COL14A1, ESR1, SLIT2, IGF1, CH25H, PRR5L, ABCA6, CEP126, IGDCC4, LHFP, MFAP3, PCSK5, RAB37, RBMS3, SETBP1, and TSPAN11) whose expression had a significant association with RFS prognosis (Fig. 7 and Supplementary Table S14).
20 genes which are significantly and gradually upregulated (MCM4, SPECC1, IMPA2, and AGO2) or downregulated (COL14A1, ESR1, SLIT2, IGF1, CH25H, PRR5L, ABCA6, CEP126, IGDCC4, LHFP, MFAP3, PCSK5, RAB37, RBMS3, SETBP1, and TSPAN11) through grade I, II, and III of breast cancers, deregulated in the bad clinical outcome group, and associated with poorer survival prognosis
MCM4, SPECC1, IMPA2, and AGO2 were gradually upregulated through grade I, II, and III of breast cancers, and their elevated expression was associated with the worse clinical outcomes RFS (Fig. 8A). In contrast, COL14A1, ESR1, SLIT2, IGF1, CH25H, PRR5L, ABCA6, CEP126, IGDCC4, LHFP, MFAP3, PCSK5, RAB37, RBMS3, SETBP1, and TSPAN11 were gradually downregulated through grade I, II, and III of breast cancers and their reduced expression were associated with worse clinical outcomes and RFS (4 genes are shown in Fig. 8B and 12 genes in Supplementary Fig. S4).
Stromal genes significantly altered among the grades (grade I, II, and III), clinical outcomes, and their association with survival prognosis. A Gradually upregulated genes among three grades and their association with bad clinical outcome. B Gradually downregulated genes among three grades and their association with bad clinical outcome (Only 4 genes are shown)
The univariate Cox regression analyses identified 23 genes (out of the 38 prognostic genes) were significant prognostic factors. These genes included COL11A1, CAV1, ITM2A, LDB2, CD86, TLR2, COL1A1, SPECC1, IMPA2, AGO2, COL14A1, SLIT2, IGF1, CH25H, PRR5L, ABCA6, CEP126, IGDCC4, LHFP, MFAP3, RAB37, SETBP1, and TSPAN11 (Supplementary Fig. S5). We further performed the multivariate Cox regression analysis with the expression levels of the 23 genes, breast cancer subtypes (including Her2 status, ER status, and PR status), and age being the predictor variables. We found that 19 prognostic genes (COL11A1, ITM2A, LDB2, CD86, TLR2, COL1A1, IMPA2, AGO2, COL14A1, SLIT2, CH25H, PRR5L, CEP126, IGDCC4, LHFP, MFAP3, RAB37, SETBP1, and TSPAN11), the three subtypes, and age were significant prognostic factors (Supplementary Fig. S6).
Analysis of the dysregulated stromal genes in TCGA BRCA cohort
We further analyzed the expression of the stromal prognostic upregulated and downregulated genes in the TCGA breast invasive carcinoma (BRCA) cohort. Interestingly, we found that the stromal upregulated prognostic genes COL11A1 and IFI30 were significantly upregulated in BRCA than in healthy tissue (Fig. 9A). Also, the downregulated stromal prognostic genes SPRY2, CAV1, SOCS2, ITM2A, LDB2, and NACA were significantly downregulated in BRCA (Fig. 9A). Moreover, we compared the expression levels of stromal prognostic hub genes (COL1A1, MMP9, CD86, FCER1G, ITGAM, and TLR2), oncogenes (COL1A1 and IL21R), protein kinases encoding genes (PRKACA and CSK), and chemokines and growth factors encoding gene (PLAU) between TCGA BRCA and healthy tissue. We found that the expression levels of COL1A1, MMP9, CD86, FCER1G, ITGAM, IL21R, CSK, and PLAU genes were consistently upregulated in BRCA (Fig. 9B). Only PRKACA and TLR2 were slightly downregulated in BRCA. Finally, we investigated the expression levels of prognostic stromal genes whose expression levels gradually altered among the three grades of tumor stroma and also altered between breast cancer patients with bad and good clinical outcomes. Interestingly, we found that MCM4, SPECC1, and AGO2 genes were upregulated, and ABCA6, CEP126, CH25H, COL14A1, IGDCC4, IGF1, MFAP3, PCSK5, RBMS3, SLIT2, TSPAN11, LHFP, and SETBP1 were downregulated in BRCA (Table 2). These results confirmed that the stromal transcriptomes contribute to breast tissue carcinogenesis.
Comparisons of the expression levels of stromal dysregulated prognostic genes between TCGA BRCA and healthy tissue. A Stromal upregulated prognostic genes (COL11A1 and IFI30) are upregulated in BRCA, and downregulated prognostic genes (SPRY2, CAV1, SOCS2, ITM2A, LDB2, and NACA) are downregulated in BRCA. B The stromal hub genes (MMP9, COL1A1, CD86, FCER1G, ITGAM, and TLR2), oncogenes (COL1A1 and IL21R), protein kinases encoding gene (CSK), and chemokines and growth factors encoding gene (PLAU) are significantly upregulated in BRCA. The Student’s t test P values and fold change are shown
Discussion
Since stromal cells control various types of tumor phenotypes, including tumor growth, invasion, progression, metastasis, and angiogenesis [2, 3], identification of novel molecular features in BTS is significant. In this study, by analyzing a combined dataset composed of eight breast tumor stromal transcriptomic datasets, we identified the deregulated stromal gene signatures and their associated cellular signaling pathways and PPIs, as well as their associations with antitumor immunosuppression, poor clinical outcomes, and tumor progression in breast cancer (Fig. 10). Our meta-analysis identified 782 upregulated and 276 downregulated stromal genes in BTS versus normal stroma. The previous studies have shown that COL10A1, COL11A1, NOX4, and COL8A1 were upregulated in the TME of breast cancer, and their upregulation was associated with the progression of aggressive breast cancers [17]. Overexpression of COL11A1 is associated with worse clinical outcomes, including overall survival and disease-free survival [47]. SULF1, elevated with the second-highest ES in the BTS, is associated with the remodeling of extracellular matrix during the progression of breast cancer [48]. Elevated expression of COMP was found in the epithelial and stromal cells of invasive breast carcinomas [49]. Another bioinformatics study revealed that IFI30, INHBA, and CTSB were upregulated in breast cancer [50]. A previous study showed that FIGF, SFRP1, and SPRY2 were consistently downregulated in multifocal invasive lobular breast tumors [51]. In breast cancer reactive stroma, ITM2A downregulation is associated with shorter survival of patients [13]. Matrix-producing stromal LDB2 has prognostic value in breast cancer [52]. Among the downregulated genes in BTS, many have been associated with breast cancer onset, invasion, progression, and metastasis [53,54,55,56,57,58]. It was also reported that the expression levels of DLK1 and CD36 were downregulated in breast cancer [53]. The type III TGF-beta receptor (TGFBR3), a tumor suppressor gene, is associated with breast cancer progression and metastasis [54]. Silencing of GPC3 expression is associated with the growth, invasion, and metastasis of MCF-7 human breast cancer cells [55]. Altogether, many of the aberrantly expressed genes in BTS have been associated with breast cancer pathogenesis and carcinogenesis.
Based on these dysregulated genes, we identified pathways upregulated and downregulated in BTS. The upregulated pathways were mainly involved in immune signatures, stromal signatures, oncogenic signatures, and metabolism. Many of these pathways were involved in cancer initiation, progression, angiogenesis, and metastasis in breast cancer [59,60,61]. For example, the group of cytokines and their receptors are associated with breast cancer growth and progression [59]. The ECM–receptor interaction pathway, a major stromal signaling pathway, is involved in breast cancer development [62]. The downregulated pathways were mainly associated with the alteration of metabolisms. In fact, deregulated cellular metabolisms in glucose, amino acid, and other nutrients, are major hallmarks of cancers [63]. The ribosome pathway is one of the major targeting pathways in cancer therapeutic [64]. As a tumor-suppressive signaling, the p53 signaling pathway is a crucial target in cancer therapy [65]. These results indicate that the transcriptional signatures of BTS are associated with the alteration of numerous cancer-associated pathways.
We identified 13 and 11 MTRs which regulate the upregulated stromal DEGs and downregulated stromal DEGs, respectively. Previous studies have shown that these MTRs are associated with breast carcinogenesis [13, 66,67,68,69]. For example, ETV7 plays a substantial role in the oncogenesis of breast tissue [67]. In nodal positive breast cancer patients, another MTR, STAT2 is associated with RFS [66]. RELA is elevated in the CAFs derived from Her2 + breast cancer tissue [68]. Another study confirmed the altered expression of JUND, FOS, and JUN in breast cancer tissue [69]. These results indicate that these TFs may control the BTS-associated transcriptions in the TME of breast cancer.
We also identified key hub genes, especially oncogenes, protein kinases encoding genes, cytokines and growth factors encoding genes in BTS based on the PPI network analysis. PPI networks indicate cellular signaling, communication, and crosstalk between the cells [70]. These hub genes included adverse prognostic factors, such as MMP9, FCER1G, CD86, ITGAM, TLR2, COL1A1, IL21R, PRKACA, CSK, PLAU, MYC, and RNASEL. In luminal A breast cancers, the matrix metalloproteinase gene MMP9 is associated with poor clinical outcomes [10]. In addition, a previous study have shown that ITGAM and TLR2 act as hub genes in breast cancer [71]. IL21R is highly expressed in breast cancer cells and is associated with proliferation, invasion, and migration of breast cancer cells [72]. The elevated levels of COL1A1 are linked with shorter survival and chemotherapy resistance [73]. Moody et al. reported that PRKACA mediated the therapy resistance in breast cancer [74]. Another prognostic stromal protein kinase, CSK, is associated with the cellular growth of hormone-independent breast cancer tissue [75]. In metastatic breast cancer, elevated PLAU is associated with poor survival prognosis [76]. However, we found that the upregulation of MYC (an oncogene downregulated in BTS) and downregulation of RNASEL (a protein kinase gene downregulated in BTS) were associated with worse survival in breast cancer patients. These data suggest that the deregulation of key molecules in PPI networks associated with stromal signatures contributes to the aggressive TME compartment in breast cancer.
The BTS-specific molecules and their associated pathways and interaction networks are potential prognostic biomarkers and therapeutic targets for breast cancer. For example, the upregulation of COL11A1 and IFI30 could indicates a worse prognosis, while the downregulation of SPRY2, CAV1, SOCS2, ITM2A, LDB2, and NACA could indicates a better prognosis in breast cancer. The small molecule inhibitors targeting the kinases upregulated in BTS, such as CSF1R, CSK, EIF2AK2, HCK, LYN, PRKACA, RNASEL, SYK, and ERBB2, could be effective in controlling breast tumor progression.
This study has several limitations. First, the findings were obtained by the bioinformatics analysis but lack of experimental validation. Second, the number of cancer samples with clinical data, such as survival time, tumor grade, and cancer subtypes, is limited in this study. Finally, we analyzed mRNA expression profiles which are not necessarily the same as protein expression profiles, due to some factors affecting the translation from mRNA level to protein level, such as post-translational modification. Therefore, to translate our findings into clinical applications, further experimental and clinical validation would be necessary.
Conclusions
Our data provide pivotal molecular insights into breast tumor stroma characterization, which may have substantial effects on the stroma-based treatment recommendations for breast cancer patients.
Data availability
The datasets (GSE9014 [9], GSE83591 (n = 53)[11], GSE31192 (n = 17) [12], GSE26910 (n = 12) [13], GSE10797 (n = 33) [14], GSE8977 (n = 22) [15], GSE33692 (n = 22) [16], and GSE14548 (n = 34) [17]) were downloaded from the NCBI GEO database (https://www.ncbi.nlm.nih.gov/geo/), the TCGA breast cancer dataset was downloaded from the website https://gdc-portal.nci.nih.gov/.
References
Guo S, Deng C-X. Effect of stromal cells in tumor microenvironment on metastasis initiation. Int J Biol Sci. 2018;14:2083–93. https://doi.org/10.7150/ijbs.25720.
Bussard KM, Mutkus L, Stumpf K, et al. Tumor-associated stromal cells as key contributors to the tumor microenvironment. Breast Cancer Res. 2016;18:84. https://doi.org/10.1186/s13058-016-0740-2.
Valkenburg KC, de Groot AE, Pienta KJ. Targeting the tumour stroma to improve cancer therapy. Nat Rev Clin Oncol. 2018;15:366–81. https://doi.org/10.1038/s41571-018-0007-1.
Hanahan D, Coussens LM. Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell. 2012;21:309–22. https://doi.org/10.1016/j.ccr.2012.02.022.
Choi H, Sheng J, Gao D, et al. Transcriptome analysis of individual stromal cell populations identifies stroma-tumor crosstalk in mouse lung cancer model. Cell Rep. 2015;10:1187–201. https://doi.org/10.1016/j.celrep.2015.01.040.
Pienta KJ, McGregor N, Axelrod R, Axelrod DE. Ecological therapy for cancer: defining tumors using an ecosystem paradigm suggests new opportunities for novel cancer treatments. Transl Oncol. 2008;1:158–64. https://doi.org/10.1593/tlo.08178.
Mao Y, Keller ET, Garfield DH, et al. Stromal cells in tumor microenvironment and breast cancer. Cancer Metastasis Rev. 2013;32:303–15. https://doi.org/10.1007/s10555-012-9415-3.
Conklin MW, Keely PJ. Why the stroma matters in breast cancer: insights into breast cancer patient outcomes through the examination of stromal biomarkers. Cell Adh Migr. 2012;6:249–60. https://doi.org/10.4161/cam.20567.
Finak G, Bertos N, Pepin F, et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat Med. 2008;14:518–27. https://doi.org/10.1038/nm1764.
Winslow S, Leandersson K, Edsjö A, Larsson C. Prognostic stromal gene signatures in breast cancer. Breast Cancer Res. 2015;17:23. https://doi.org/10.1186/s13058-015-0530-2.
Liu H, Dowdle JA, Khurshid S, et al. Discovery of stromal regulatory networks that suppress ras-sensitized epithelial cell proliferation. Dev Cell. 2017;41:392-407.e6. https://doi.org/10.1016/j.devcel.2017.04.024.
Harvell DME, Kim J, O’Brien J, et al. Genomic signatures of pregnancy-associated breast cancer epithelia and stroma and their regulation by estrogens and progesterone. Horm Cancer. 2013;4:140–53. https://doi.org/10.1007/s12672-013-0136-z.
Planche A, Bacac M, Provero P, et al. Identification of prognostic molecular features in the reactive stroma of human breast and prostate cancer. PLoS ONE. 2011;6: e18640. https://doi.org/10.1371/journal.pone.0018640.
Casey T, Bond J, Tighe S, et al. Molecular signatures suggest a major role for stromal cells in development of invasive breast cancer. Breast Cancer Res Treat. 2009;114:47–62. https://doi.org/10.1007/s10549-008-9982-8.
Karnoub AE, Dash AB, Vo AP, et al. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature. 2007;449:557–63. https://doi.org/10.1038/nature06188.
Knudsen ES, Ertel A, Davicioni E, et al. Progression of ductal carcinoma in situ to invasive breast cancer is associated with gene expression programs of EMT and myoepithelia. Breast Cancer Res Treat. 2012;133:1009–24. https://doi.org/10.1007/s10549-011-1894-3.
Ma X-J, Dahiya S, Richardson E, et al. Gene expression profiling of the tumor microenvironment during breast cancer progression. Breast Cancer Res. 2009;11:R7. https://doi.org/10.1186/bcr2222.
Xia J, Gill EE, Hancock REW. NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc. 2015;10:823–44. https://doi.org/10.1038/nprot.2015.052.
Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8:118–27. https://doi.org/10.1093/biostatistics/kxj037.
Koboldt DC, Fulton RS, McLellan MD, et al. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70. https://doi.org/10.1038/nature11412.
Ritchie ME, Phipson B, Wu D, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47–e47. https://doi.org/10.1093/nar/gkv007.
Cochran WG. The combination of estimates from different experiments. Biometrics. 1954;10:101–29. https://doi.org/10.2307/3001666.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc: Ser B (Methodol). 1995;57:289–300.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50. https://doi.org/10.1073/pnas.0506580102.
Kanehisa M, Furumichi M, Tanabe M, et al. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017;45:D353–61. https://doi.org/10.1093/nar/gkw1092.
Clarke DJB, Kuleshov MV, Schilder BM, et al. eXpression2Kinases (X2K) Web: linking expression signatures to upstream cell signaling networks. Nucleic Acids Res. 2018;46:W171–9. https://doi.org/10.1093/nar/gky458.
Janky R, Verfaillie A, Imrichová H, et al. iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLoS Comput Biol. 2014;10: e1003731. https://doi.org/10.1371/journal.pcbi.1003731.
Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47:D607–13. https://doi.org/10.1093/nar/gky1131.
Chin C-H, Chen S-H, Wu H-H, et al. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8:S11. https://doi.org/10.1186/1752-0509-8-S4-S11.
Shannon P, Markiel A, Ozier O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504. https://doi.org/10.1101/gr.1239303.
Yoshihara K, Shahmoradgoli M, Martínez E, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. https://doi.org/10.1038/ncomms3612.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7. https://doi.org/10.1186/1471-2105-14-7.
Terry Therneau. A package for survival analysis in R, August 23, 2021. https://cran.rproject.org/web/packages/survival/index.html.
Multiple Comparisons Using R. In: Routledge & CRC Press. https://www.routledge.com/Multiple-Comparisons-Using-R/Bretz-Hothorn-Westfall/p/book/9781584885740. Accessed 14 Feb 2021
Luo X, Xiong X, Shao Q, et al. The tumor suppressor interferon regulatory factor 8 inhibits β-catenin signaling in breast cancers, but is frequently silenced by promoter methylation. Oncotarget. 2017;8:48875–88. https://doi.org/10.18632/oncotarget.16511.
Wolf B, Goebel G, Hackl H, Fiegl H. Reduced mRNA expression levels of NFE2L2 are associated with poor outcome in breast cancer patients. BMC Cancer. 2016. https://doi.org/10.1186/s12885-016-2840-x.
Wang X, Wang G, Shi Y, et al. PPAR-delta promotes survival of breast cancer cells in harsh metabolic conditions. Oncogenesis. 2016;5:e232–e232. https://doi.org/10.1038/oncsis.2016.41.
Ferrari N, Mohammed ZMA, Nixon C, et al. Expression of RUNX1 correlates with poor patient prognosis in triple negative breast cancer. PLoS ONE. 2014. https://doi.org/10.1371/journal.pone.0100759.
Ferralli J, Chiquet-Ehrismann R, Degen M. KLF4α stimulates breast cancer cell proliferation by acting as a KLF4 antagonist. Oncotarget. 2016;7:45608–21. https://doi.org/10.18632/oncotarget.10058.
Kren BT, Unger GM, Abedin MJ, et al. Preclinical evaluation of cyclin dependent kinase 11 and casein kinase 2 survival kinases as RNA interference targets for triple negative breast cancer therapy. Breast Cancer Res. 2015;17:19. https://doi.org/10.1186/s13058-015-0524-0.
Feng Y, Spezia M, Huang S, et al. Breast cancer development and progression: risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis. 2018;5:77–106. https://doi.org/10.1016/j.gendis.2018.05.001.
Liu W, Ye H, Liu Y-F, et al. Transcriptome-derived stromal and immune scores infer clinical outcomes of patients with cancer. Oncol Lett. 2018;15:4351–7. https://doi.org/10.3892/ol.2018.7855.
Zhang S-C, Hu Z-Q, Long J-H, et al. clinical implications of tumor-infiltrating immune cells in breast cancer. J Cancer. 2019;10:6175–84. https://doi.org/10.7150/jca.35901.
Jin YW, Hu P. Tumor-infiltrating CD8 T cells predict clinical breast cancer outcomes in young women. Cancers (Basel). 2020;12:E1076. https://doi.org/10.3390/cancers12051076.
Ai L, Mu S, Wang Y, et al. Prognostic role of myeloid-derived suppressor cells in cancers: a systematic review and meta-analysis. BMC Cancer. 2018;18:1220. https://doi.org/10.1186/s12885-018-5086-y.
Huang Y, Ma C, Zhang Q, et al. CD4+ and CD8+ T cells have opposing roles in breast cancer progression and outcome. Oncotarget. 2015;6:17462–78. https://doi.org/10.18632/oncotarget.3958.
Jia D, Liu Z, Deng N, et al. A COL11A1-correlated pan-cancer gene signature of activated fibroblasts for the prioritization of therapeutic targets. Cancer Lett. 2016;382:203–14. https://doi.org/10.1016/j.canlet.2016.09.001.
Bret C, Moreaux J, Schved J-F, et al. SULFs in human neoplasia: implication as progression and prognosis factors. J Transl Med. 2011;9:72. https://doi.org/10.1186/1479-5876-9-72.
Grigoriadis A, Mackay A, Reis-Filho JS, et al. Establishment of the epithelial-specific transcriptome of normal and malignant human breast cells based on MPSS and array expression data. Breast Cancer Res. 2006;8:R56. https://doi.org/10.1186/bcr1604.
Xiang Y-J, Fu Q-Y, Ma Z-B, et al. Screening for candidate genes related to breast cancer with cDNA microarray analysis. Chronic Dis Transl Med. 2015;1:65–72. https://doi.org/10.1016/j.cdtm.2015.02.001.
Norton N, Advani PP, Serie DJ, et al. Assessment of tumor heterogeneity, as evidenced by gene expression profiles, pathway activation, and gene copy number, in patients with multifocal invasive lobular breast tumors. PLoS ONE. 2016;11: e0153411. https://doi.org/10.1371/journal.pone.0153411.
Winslow S, Lindquist KE, Edsjö A, Larsson C. The expression pattern of matrix-producing tumor stroma is of prognostic importance in breast cancer. BMC Cancer. 2016;16:841. https://doi.org/10.1186/s12885-016-2864-2.
Makoukji J, Makhoul NJ, Khalil M, et al. Gene expression profiling of breast cancer in Lebanese women. Sci Rep. 2016;6:36639. https://doi.org/10.1038/srep36639.
Dong M, How T, Kirkbride KC, et al. The type III TGF-beta receptor suppresses breast cancer progression. J Clin Invest. 2007;117:206–17. https://doi.org/10.1172/JCI29293.
Castillo LF, Tascón R, Lago Huvelle MA, et al. Glypican-3 induces a mesenchymal to epithelial transition in human breast cancer cells. Oncotarget. 2016;7:60133–54. https://doi.org/10.18632/oncotarget.11107.
Johnson RW, Finger EC, Olcina MM, et al. Induction of LIFR confers a dormancy phenotype in breast cancer cells disseminated to the bone marrow. Nat Cell Biol. 2016;18:1078–89. https://doi.org/10.1038/ncb3408.
Fu J, Khaybullin R, Zhang Y, et al. Gene expression profiling leads to discovery of correlation of matrix metalloproteinase 11 and heparanase 2 in breast cancer progression. BMC Cancer. 2015. https://doi.org/10.1186/s12885-015-1410-y.
Ferreira MA, Gamazon ER, Al-Ejeh F, et al. Genome-wide association and transcriptome studies identify target genes and risk loci for breast cancer. Nat Commun. 2019;10:1741. https://doi.org/10.1038/s41467-018-08053-5.
Esquivel-Velázquez M, Ostoa-Saloma P, Palacios-Arreola MI, et al. The role of cytokines in breast cancer development and progression. J Interferon Cytokine Res. 2015;35:1–16. https://doi.org/10.1089/jir.2014.0026.
Chow MT, Luster AD. Chemokines in cancer. Cancer Immunol Res. 2014;2:1125–31. https://doi.org/10.1158/2326-6066.CIR-14-0160.
Stacker SA, Achen MG. The VEGF signaling pathway in cancer: the road ahead. Chin J Cancer. 2013;32:297–302. https://doi.org/10.5732/cjc.012.10319.
Bao Y, Wang L, Shi L, et al. Transcriptome profiling revealed multiple genes and ECM-receptor interaction pathways that may be associated with breast cancer. Cell Mol Biol Lett. 2019;24:38. https://doi.org/10.1186/s11658-019-0162-0.
Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23:27–47. https://doi.org/10.1016/j.cmet.2015.12.006.
Catez F, Dalla Venezia N, Marcel V, et al. Ribosome biogenesis: an emerging druggable pathway for cancer therapeutics. Biochem Pharmacol. 2019;159:74–81. https://doi.org/10.1016/j.bcp.2018.11.014.
Stegh AH. Targeting the p53 signaling pathway in cancer therapy—the promises, challenges, and perils. Expert Opin Ther Targets. 2012;16:67–83. https://doi.org/10.1517/14728222.2011.643299.
Wu H-T, Liu J, Li G-W, et al. The transcriptional STAT3 is a potential target, whereas transcriptional STAT5A/5B/6 are new biomarkers for prognosis in human breast carcinoma. Oncotarget. 2017;8:36279–88. https://doi.org/10.18632/oncotarget.16748.
Mathot P, Grandin M, Devailly G, et al. DNA methylation signal has a major role in the response of human breast cancer cells to the microenvironment. Oncogenesis. 2017;6: e390. https://doi.org/10.1038/oncsis.2017.88.
Tchou J, Kossenkov AV, Chang L, et al. Human breast cancer associated fibroblasts exhibit subtype specific gene expression profiles. BMC Med Genomics. 2012;5:39. https://doi.org/10.1186/1755-8794-5-39.
Chen D, Yang H. Integrated analysis of differentially expressed genes in breast cancer pathogenesis. Oncol Lett. 2015;9:2560–6. https://doi.org/10.3892/ol.2015.3147.
Pawson T, Nash P. Protein–protein interactions define specificity in signal transduction. Genes Dev. 2000;14:1027–47. https://doi.org/10.1101/gad.14.9.1027.
He L, Wang D, Wei N, Guo Z. Identification of potential therapeutic targets using breast cancer stroma expression profiling. Transl Cancer Res. 2016. https://doi.org/10.21037/7804.
Wang L-N, Cui Y-X, Ruge F, Jiang WG. Interleukin 21 and its receptor play a role in proliferation, migration and invasion of breast cancer cells. Cancer Genomics Proteomics. 2015;12:211–21.
Liu J, Shen J-X, Wu H-T, et al. Collagen 1A1 (COL1A1) promotes metastasis of breast cancer and is a potential therapeutic target. Discov Med. 2018;25:211–23.
Moody SE, Schinzel AC, Singh S, et al. PRKACA mediates resistance to HER2-targeted therapy in breast cancer cells and restores anti-apoptotic signaling. Oncogene. 2015;34:2061–71. https://doi.org/10.1038/onc.2014.153.
Xiao T, Li W, Wang X, et al. Estrogen-regulated feedback loop limits the efficacy of estrogen receptor–targeted breast cancer therapy. Proc Natl Acad Sci USA. 2018;115:7869–78. https://doi.org/10.1073/pnas.1722617115.
Banys-Paluchowski M, Witzel I, Aktas B, et al. The prognostic relevance of urokinase-type plasminogen activator (uPA) in the blood of patients with metastatic breast cancer. Sci Rep. 2019;9:2318. https://doi.org/10.1038/s41598-018-37259-2.
Funding
This work was supported by China Pharmaceutical University (Grant numbers 3150120001 to XW).
Author information
Authors and Affiliations
Contributions
MNU performed data analyses, conceived the research, and prepare the manuscript. XW conceived the research, designed analysis strategies, and wrote the manuscript. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12282_2022_1332_MOESM2_ESM.xlsx
Supplementary file2: Table S1. Description of individual breast tumor stroma study included in the meta-analysis. GEO: Gene Expression Omnibu. (XLSX 11 KB)
12282_2022_1332_MOESM4_ESM.xlsx
Supplementary file4: Table S3. The patient's characteristics of GSE9014, including number of samples, tumor grade, hormone receptor status, lymph node status, and types of therapies. (XLSX 12 KB)
12282_2022_1332_MOESM9_ESM.xlsx
Supplementary file9: Table S8. Putative enriched transcription factors through Transcription Factor Enrichment Analysis (TFEA). (XLSX 9 KB)
12282_2022_1332_MOESM10_ESM.xlsx
Supplementary file10: Table S9. Ranked list of enriched kinases based on overlap between known kinase–substrate phosphorylation interactions and the proteins. (XLSX 13 KB)
12282_2022_1332_MOESM11_ESM.xlsx
Supplementary file11. Table S10. List of master transcription factors associated with the regulation of stromal DEGs. (XLSX 10 KB)
12282_2022_1332_MOESM13_ESM.xlsx
Supplementary file13: Table S12. List of 1955 genes which are significantly dysregulated (F-test, P < 0.05) among three grades. (XLSX 95 KB)
12282_2022_1332_MOESM14_ESM.xlsx
Supplementary file14: Table S13. List of 1471 genes which are significantly dysregulated (Student t-test, P < 0.05) between the bad clinical and good clinical outcome groups. (XLSX 84 KB)
12282_2022_1332_MOESM15_ESM.xlsx
Supplementary file15: Table S14. List of 124 genes which are are commonly found between the grades-basis (grade I, grade II, and grade III) and clinical outcome basis (bad versus good clinical outcome) analysis. (XLSX 14 KB)
About this article
Cite this article
Uddin, M.N., Wang, X. Identification of key tumor stroma-associated transcriptional signatures correlated with survival prognosis and tumor progression in breast cancer. Breast Cancer 29, 541–561 (2022). https://doi.org/10.1007/s12282-022-01332-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-022-01332-6