Abstract
Ductal carcinoma in situ (DCIS) is a precursor lesion that can gives rise to invasive breast cancer (IBC). It has been proposed that both the nature of the lesion and the tumor microenvironment play key roles in progression to IBC. Here, laser capture microdissected tissue from pure DCIS and pure IBC were employed to define key gene expression profiles in either the epithelial or stromal compartment associated with disease progression. Each tissue had distinct gene expression profiles, and a DCIS/IBC classifier accurately distinguished DCIS versus IBC in multiple independent data sets. However, contrary to other studies that profiled DCIS associated with invasive disease, we found that the most significant alterations in gene expression were observed in the epithelial compartment rather than in the stroma. In particular, genes associated with epithelial-to-mesenchymal transition and myoepithelial cell-specific genes were enriched in invasive cancer relative to pure DCIS. Such alterations in transcript levels were associated with all subtypes of breast cancer, but were particularly indicative of poor outcome in ER-negative breast cancer. Together, these studies indicate that lesion-specific differences in gene expression associated with invasive phenotype are particularly relevant in the progression of DCIS to invasive breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is a clinically heterogeneous and complex disease, encompassing a wide variety of pathological entities with variable clinical behavior. It is now widely acknowledged that accumulation of genetic anomalies contributes to the acquisition of an increasingly aggressive, invasive, or therapy-resistant tumor phenotype. In contrast to invasive breast cancer (IBC), the heterogeneity of the breast cancer precursor lesion ductal carcinoma in situ (DCIS) is not well investigated.
Due to enhanced screening methodologies, an increasing number of women are diagnosed with DCIS, a non-obligate precursor to IBC [1]. Left untreated, up to 53% of DCIS cases will progress to IBC [2]. Traditionally, estimating the risk of DCIS recurrence or progression to IBC and subsequent clinical management decisions have been based on evaluation of standard clinical–pathological features. Nuclear grade, the presence of necrosis, and size of the lesion have all been linked to increased risk of DCIS recurrence [3, 4]. In addition, numerous biomarkers have been investigated for better risk stratification of patients with DCIS. Elevated Ki-67 levels, p53 mutations, and HER2 amplification are known to be associated with a higher nuclear grade and necrosis [5, 6]. Although, such indicators provide some valuable information about clinical course, they are often not able to accurately predict the prognosis for the individual patient and guide proper and effective treatment. As a result, many DCIS patients are over-treated, while others progress to invasive carcinoma despite treatment. Therefore, there is currently significant interest in identifying molecular events driving invasive progression, not only for use in determining at which point the lesion is most likely to progress to malignancy, but also in hopes of identifying new molecular targets that could halt progression at these early stages.
The development of new technologies has offered the opportunity to explore the molecular complexity of human cancer. Multiple studies have investigated various clinical and biological aspects of the disease, including tumor classification, prediction of prognosis, and response to different treatments. For example, whole gene expression profiling of IBC lead to identification of molecularly distinct subtypes with different prognosis, driving oncogenic changes, and treatment modalities [7, 8]. While initially used largely for academic purposes, genome-wide analysis technologies are becoming more reliable and have been shown to be powerful diagnostic tools. In this context, several multi-gene assays (e.g., MammaPrint, 21-gene recurrence score, HOXB13/IL17BR ratio) have been developed for diagnostic purposes and are now commercially available [9–11].
In contrast to IBC, the molecular complexity and diversity of DCIS is much less well characterized. A few studies have investigated genetic, epigenetic, and gene expression changes in breast tissue from early stages of breast cancer progression including atypical ductal hyperplasia (ADH) and DCIS [12–14]. The majority of these studies reported that most dramatic changes occur at the transition from normal breast tissue to ADH, and relatively minor differences are observed between DCIS and IBCs. Surprisingly, in those studies, when transcriptional profiles of neoplastic epithelial cells and stromal cells were compared between DCIS and IBC, more robust changes in gene expression were observed in the stroma. When DCIS and IBC coexist in the same lesion, their gene expression profiles are very similar but the transcriptome of pure DCIS is largely uncharacterized. This is due to the difficulty in obtaining fresh DCIS tissue for research studies. Most DCIS do not form mass lesions allowing for straightforward tissue collection, and the entire DCIS lesion has to be microscopically evaluated to exclude the presence of invasive breast carcinoma. The objective of this study was to compare gene expression profiles of microdissected epithelial and stromal components of pure DCIS and IBC, and to determine the significance of alterations in gene expression related to the clinical behavior of breast cancer.
Materials and methods
Study population
DCIS and IBC cases were obtained from the tumor bank at Thomas Jefferson University. DCIS was classified as low, intermediate, and high nuclear grade [15]. Nottingham classification was used to determine histological grade of IBC [16]. Estrogen, progesterone receptors, and HER2 expressions were determined by immunohistochemistry using CAP guidelines for stain interpretation [17]. The nuclear grade and hormone receptor status of the lesions are depicted in Table 1. DCIS patients were treated by excision of the lesion with a negative margin and did not receive adjuvant hormonal or radiation therapy. DCIS and IBC were matched for estrogen receptor, progesterone receptor, and HER2 status. The study was approved by the Thomas Jefferson University Institutional Review Board.
LCM and RNA isolation and amplification
Neoplastic epithelial cells and tumor stroma were laser capture microdissected as previously described using Leica LCM system (Leica Microsystems, Germany) [18]. Total RNA was extracted using the Qiagen microRNA kit. Quality of RNA was assessed using the Agilent 2100 Bioanalyzer. Total RNA was amplified using the NuGEN™ WT-Ovation™ FFPE RNA Amplification System V2. First-strand synthesis of cDNA was performed using a unique first-strand DNA/RNA chimeric primer mix, resulting in cDNA/mRNA hybrid molecules. Following fragmentation of the mRNA component of the cDNA/mRNA molecules, second-strand synthesis was performed and double-stranded cDNA was formed with a unique DNA/RNA heteroduplex at one end. In the final amplification step, RNA within the heteroduplex was degraded using RNaseH, and replication of the resultant single-stranded cDNA was achieved through DNA/RNA chimeric primer binding and DNA polymerase enzymatic activity. The amplified single-stranded cDNA was purified for accurate quantitation of the cDNA and to ensure optimal performance during the fragmentation and labeling process. The single-stranded cDNA was assessed using spectrophotometric methods in combination with the Agilent Bioanalyzer. The appropriate amount of amplified single-stranded cDNA was fragmented and labeled using the Encore™ cDNA Biotin Module. The enzymatically and chemically fragmented product (50–100 nt) was labeled via the attachment of biotinylated nucleotides onto the 3′-end of the fragmented cDNA.
Microarray hybridization
The resultant fragmented and labeled cDNA was added to the hybridization cocktail in accordance with the NuGEN™ guidelines for hybridization onto Affymetrix GeneChip® arrays. Following the hybridization for 16–18 h at 45°C in an Affymetrix GeneChip® Hybridization Oven 640, the array was washed and stained on the GeneChip® Fluidics Station 450 using the appropriate fluidics script, before being inserted into the Affymetrix autoloader carousel and scanned using the GeneChip® Scanner 3000.
Microarray data pre-processing
Raw intensity files from the Affymetrix Human Exon 1.0 ST GeneChip arrays were processed using Affymetrix Expression Console version 1.1. Gene-level expression measurements were computed using the iterPLIER algorithm on the “core” probesets and exported with annotation release 32, dated June 23, 2011. Additional array preprocessing was performed in Matlab version 7.11.0 (R2010b), where expression values were converted to iterPLIER + 16 by adding 16, and then transformed to log2 scale. Before analysis, data was filtered to exclude probesets with no gene annotation, and genes with multiple probesets were handled by averaging their rows together and scaling by the probeset with the largest standard deviation. All subsequent analysis was performed in Matlab unless otherwise noted. Data are deposited at GEO as data set GSE33692.
Unsupervised learning
Genes with variance in the top 25th percentile were used for both principal component analysis and hierarchical clustering, to observe patterns in the microarray data. Principal component analysis was performed across samples and the second, third, and fourth components were plotted as a function of the first component to observe natural separations that may be associated with sample tissue characteristics. Hierarchical clustering was performed on both genes and samples, using Pearson’s correlation distance metric and average linkage.
Differential expression analysis
Differential gene expression analysis was performed on paired epithelial versus stromal samples within IBC and DCIS cases separately, using a paired t test. Differential gene expression analysis for IBC versus DCIS samples was performed within the epithelial samples and stromal samples, to identify genes associated with progression within each of these compartments, using a two-sample t test with unequal variance. 10,000 sample permutations were performed to obtain better P value estimations for the two-sample t tests. To account for multiple testing, P values were adjusted based on the estimated false discovery rate (FDR), using the procedure introduced by Storey [19]. The FDR employed and rationales are consistent with prior studies and analyses [20–23]. For increased statistical power, genes with variance below the 50th percentile or expressed below the 25th percentile in over 90% of the samples being compared were assigned an adjusted P value of 1 and filtered out prior to FDR estimation [24]. After this filtering step was performed, an alternative method for differential gene expression analysis, statistical analysis of microarrays (SAM) [25], was applied and used to obtain an improved assessment of the FDR, estimated by randomly permuting samples in the dataset.
Comparisons among independent datasets
The set of genes differentially expressed in the epithelium (Storey FDR < 40%) and the set of genes differentially expressed in the stroma (Storey FDR < 40%) were used to evaluate gene expression profiles in independent microarray datasets. Series matrix files and annotation for gene expression datasets GSE3893, GSE14548, and GSE26304 were downloaded from the Gene Expression Omnibus. Datasets were imported into Matlab and the epithelial and stromal differential expression lists were mapped to each dataset based on gene symbol. Genes with multiple probesets were handled by averaging their rows together and scaling by the probeset with the largest standard deviation. For the epithelial and stromal differential gene sets, psuedo-expression signatures in each GEO dataset were defined by median-centering gene expression profiles, multiplying the median-centered profiles for downregulated genes by −1, and taking the average over all genes. These psuedo-expression signatures were used to rank breast tissue samples from low-to-high expression and observe phenotypic trends as a function of expression gradient. The distribution of IBC samples along the expression gradient was assessed for significance using the Kolmogorov–Smirnov test.
Gene set enrichment analysis
A gene set database was compiled from previous studies, including epithelial-to-mesenchymal transition (EMT) genes reported in Table S1 in Taube et al. [26]; myoepithelial-specific genes reported in Table 3A in Grigoriadis et al. [27]; invasion-associated genes in Table 1 in Kim et al. [28]; progression-specific genes in Schuetz et al. [29], DCIS/invasive discriminating genes in Table 2 in Hanneman et al. [30]; and DCIS/invasive transition genes from Table 1 in Ma et al. [12]. The gene set enrichment analysis (GSEA) software tool [31] was used to identify similarities between expression profiles in our microarray dataset and the previously reported disease progression/invasion gene sets. In addition, functional enrichment analysis of the topmost differentially expressed genes in the epithelial compartment (FDR < 40% and FDR < 10%) was performed using the database for annotation, visualization and integrated discovery (DAVID) [32].
Retrospective meta-analysis of clinical outcome in breast cancer patients
A microarray dataset that was previously compiled from the public repositories Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) [33] and ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) [34] was used to evaluate the epithelial IBC versus DCIS expression signature, along with myoepithelial and EMT-specific subsets, in the context of clinical samples [35]. This dataset includes 2,254 breast cancer cases, of which 1,740 were identified as ER-positive and 514 were identified as ER-negative, based on ESR1 mRNA expression [35]. Samples were analyzed in separate groups based on ER status. Differential expression of the averaged gene signature magnitude among these sample groups was evaluated using two-tailed t test. Kaplan–Meier analysis was used to evaluate survival trends within sample groups, including 936 with relapse free survival time (790 ER-pos., 146 ER-neg.), and 488 with overall survival time (358 ER-pos., 130 ER-neg.). The log-rank test was used to evaluate differences in survival curves for high versus low signature-expressing populations.
Results
Preparation of microdissected tissue
To directly investigate the epithelial and stromal compartments, clinical specimens of DCIS and IBC were used (Table 1). All DCIS lesions evaluated did not progress on to IBC over the follow up period (minimum 8 years). A total of three normal tissues, nine DCIS, and ten IBC cases were subjected to laser capture microdissection (LCM). LCM was preformed on 14-μm thick sections of frozen material stained with hematoxylin and eosin. Representative images of the tissues and the dissected region are provided in Fig. 1A. RNA was isolated from each tissue and subjected to hybridization on Affymetrix gene chips. Initially, normalized RNA expression data was used to evaluate the relatively purity of the epithelial and stromal compartments (Fig. 1B). These data demonstrated the expected enrichment of epithelial-specific markers (e.g., E-cadherin and epithelial cell adhesion molecule) in the DCIS and IBC epithelial microdissected compartments relative to the stromal specimens and confirms that all tissues were enriched via the microdissection.
DCIS and IBC and associated stroma exhibit distinct gene expression profiles
To determine in an unbiased fashion whether the gene expression observed in the DCIS, IBC, and associated stromal compartments were distinct, principle component analysis was performed on the top 25% variably expressed genes. The resulting first principle component of the gene expression data clearly differentiated the epithelial and stromal components in DCIS and IBC (Fig. 2a). The fourth principle component differentiated the DCIS from IBC (Fig. 2a). Thus, gene expression patterns in each of the four tissues analyzed are distinct. In addition, the fourth principal component associated with DCIS to IBC transition demonstrated a stronger separation between samples in the epithelial compartment. To further evaluate relational differences in gene expression, the top 25% of variably expressed genes were used to perform hierarchical clustering (Supplementary Table 1). As shown in Fig. 2b, the epithelial and stromal compartments were partitioned into two highly distinct clusters. In addition, within both the epithelial and stromal tissue compartments, DCIS, IBC, and normal breast tissue formed disease state-specific clusters. Importantly, this categorization of the tissue of origin was present irrespective of commonly employed clinical–pathological features such as grade, ER, and Her2 status.
Intrinsic differences in gene expression between DCIS and IBC. a Principal component analysis demonstrates that epithelial and stromal tumor compartments are distinct between both DCIS and IBC. b Clustering analyses of the top 25% of variably expressed genes further demonstrated that the gene expression profiles differentiate DCIS and IBC irrespective of tumor grade, ER/PR, and HER2 status
To quantify the predominant gene expression changes within epithelial and stromal compartment of DCIS versus IBC, statistical tests for differential expression were applied. Using an ad-hoc cutoff of >1.5 absolute fold change and P < 0.01, there were 223 distinct genes in the epithelial compartment as compared to 151 in the stromal compartment (Fig. 3a, Supplementary Table 2). After correcting for multiple hypothesis testing by limiting the Storey FDR to 40%, the majority of genes in the DCIS versus IBC compartment remained significant, while only 22 stromal genes were retained (Fig. 3b, Supplementary Table 3), similar results were observed using the significance analysis of microarrays with a 10% FDR (Supplementary Fig. 1). Use of the more stringent criteria of a Storey FDR of <10% resulted in the retention of differentially expressed genes only in the epithelial compartment (Supplementary Fig. 2). Combined, these analyses showed that a majority of significant changes in mRNA expression occur in the epithelial compartment, as opposed to the tumor-associated stroma. Functional enrichment analysis of gene ontology biological process terms revealed that the genes up regulated with FDR < 40% in IBC were involved in biological processes such us adhesion (P = 1.2E-18), extracellular matrix organization (P = 2.5E-11), collagen fibril organization (P = 2.5E-10), skeletal system development (P = 2.2E-9), extracellular structure organization (P = 7.1E-9), blood vessel development (P = 5.3E-6), vasculature development (P = 6.7E-6), skin development (P = 8.4E-6) and immune response (P = 4.4E−5). Importantly, these same processes were similarly enriched when using the more stringent statistical cutoff for the DCIS versus IBC comparison (Supplementary Fig. 2). These findings suggest that alterations in these key biological processes play a fundamental role in the transition from DCIS to IBC.
Highly significant differences in gene expression between DCIS and IBC. a Heatmaps depict genes with an absolute fold change difference of >1.5 and P < 0.01 in each tissue compartment. b Heatmaps of genes with that demonstrated a fold change difference of >1.5 and FDR < 40% in each of the tissue compartment are provided
Validation of tissue-compartment-specific gene expression in independent data sets
Stroma-specific and tumor-specific gene expression profiles defined from microdissected samples were applied to three independent gene expression datasets comprising DCIS and IBC cases to verify that these expression profiles were distinctly associated with invasive disease [13, 29, 36]. These analyses show that the classifiers of DCIS versus IBC in the epithelial compartment are highly effective at segregating these two forms of disease (Fig. 4a). Genes differentially expressed in the stromal compartment were also capable of effectively separating DCIS and IBC when applied to an independent set of stromal samples (Fig. 4b). Combined, these analyses indicate that alterations in gene expression within the epithelial compartment are particularly relevant for the transition between DCIS and IBC and are highly reproducible through the independent data sets evaluated.
DCIS versus IBC classifiers validated on independent data sets. a The genes identified as differentially expressed in DCIS versus IBC effectively segregated disease state in three independent data sets. b The genes differentially expressed in stromal tissue associated with DCIS versus IBC effectively segregated disease in one independent data set where stromal compartment samples were available
Distinction between pure DCIS and DCIS associated with IBC
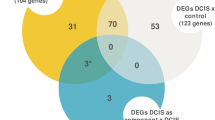
Several prior studies have analyzed DCIS co-existing in one tumor with IBC [12–14, 29, 36]. Here, we interrogated the gene expression profile as observed in our pure DCIS cases (TJU) versus those DCIS samples that were obtained in conjunction with IBC (GSE14548). These analyses show that the DCIS cases present in concert with IBC harbor gene expression profiles more similar to IBC (Fig. 5a). Similarly, powered comparisons show differential expression in IBC and pure DCIS (P = 7.51 × 10−7), but not in IBC versus mixed cases (P = 0.7). In particular, those genes that we defined as being upregulated in IBC are already elevated in the DCIS associated with IBC. This finding may provide an explanation for why relatively few changes in the epithelial compartment were seen in studies analyzing DCIS associated with IBC, and suggest that upregulation of these genes in the epithelial compartment of DCIS is associated with disease progression. Consistent with that supposition, mixed DCIS/IBC have an expression profile magnitude that is indistinguishable from IBC (Fig. 5b).
DCIS associated with invasive disease already harbor gene expression profile similar to IBC. a Relative expression levels of epithelial (top panels) and stromal classifiers (bottom panels) as identified in pure DCIS (TJU) were compared with data from DCIS associated with invasive disease (GSE1458). Left panels show genes that are upregulated from DCIS to IBC, right panels show genes that are downregulated from DCIS to IBC. b Relative expression levels of epithelial classifiers were compared with data from pure DCIS and mixed DCIS/IBC from an independent cohort (GSE26304). Left panels show genes that are upregulated from DCIS to IBC, right panels show genes that are downregulated from DCIS to IBC
Association of differentially expressed DCIS/IBC genes with breast cancer outcome
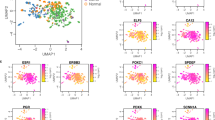
Presumably, the altered gene expression associated with progression to IBC could yield more aggressive forms of disease that would have significant effect on disease outcome. Consistent with this concept, box/whisker plots demonstrate that the genes differentially expressed between DCIS and IBC are significantly deregulated in a consistent manner between healthy/normal breast and breast tumor tissue in a large (N > 1,000) integrated breast cancer data set (Fig. 6a). This finding indicates a relatively general role for dysregulation of the genes identified in our study across all forms of breast cancer and suggests that such genes could be utilized as markers. Interestingly, while these genes were generally dysregulated across all IBC, their prognostic significance was only observed in ER-negative breast cancer—where elevated DCIS/IBC signature expression (top 25% as determined by quartile survival analyses) was associated with poor outcome (Fig. 6b).
Genes dysregulated between DCIS and IBC are deregulated in IBC relative to healthy controls. a The gene expression classifiers were applied to a large microarray database of IBC cases. These discriminatory gene profiles were significantly deregulated in ER-positive and ER-negative breast cancer. b Investigating the prognostic significance of this gene expression program revealed that it is associated with poor outcome in ER-negative breast cancer (P = 0.0184)
DCIS to IBC is associated with EMT and myoepithelial gene expression
To determine how the genes defined in our analyses relate to prior gene expression studies on breast cancer disease progression, gene set enrichment analyses (GSEA) was performed. These data revealed there was an over-representation of genes involved in EMT (EMT-Quiagen/Taube et al.), myoepithelial cell specific genes (Grigoriadis et al.) and disease progression (Schuetz et al./Kim et al.) [26–28] (Fig. 7a, b). These findings suggest that transcriptional reprogramming within the epithelia toward a more invasive state is a critical and common feature of breast cancer disease progression.
EMT and myoepithelial expression programs are enriched in IBC. a GSEA analyses was performed on a collection of gene expression signatures relevant to breast cancer disease progression. These data revealed a highly consistent enrichment for gene sets involved in EMT and myoepithelial cell-specific genes. b Representative enrichment plots are shown. c Box and whisker plots demonstrate the elevation of EMT and myoepithelial cell-specific genes in IBC versus DCIS
To investigate specific biological facets of the gene expression differences defined between DCIS and IBC epithelia, we specifically focused on significant differentially expressed genes that have previously been identified as EMT- and myoepithelial-associated genes. As shown in the box and whisker plots, EMT associated genes were consistently upregulated in invasive disease relative to pure DCIS (Fig. 7c). This observation was specific to the epithelial compartment, and was not observed in the stroma (not shown). Analysis of the EMT signature across breast cancer subtypes demonstrated an enrichment in all breast cancer subtypes. Importantly, in the context of ER-negative breast cancer this signature was associated with poor disease outcome as determined by quartile survival analyses (Fig. 8a). Similarly, application of myoepithelial-specific genes across breast cancer subtypes demonstrated an overall induction in breast cancer, but association with poor disease outcome specifically in ER-negative breast cancer (Fig. 8b). Thus, the deregulated programs of gene expression related to EMT and myoepithelial differentiation as observed at the transition to IBC are relevant to the outcome of ER-negative breast cancer.
EMT and myoepithelial cell specific signatures are associated with poor disease outcome in ER-negative breast cancer. EMT (a) and myoepithelial cell-specific (b) gene expression signatures were evaluated for relative expression across ER-positive and ER-negative breast cancer showing consistent deregulation in disease. However, both signatures were only associated with poor disease outcome in ER-negative breast cancer (P < 0.05)
Discussion
The management of breast cancer has been dramatically influenced by the definition of specific subtypes of disease that have known prognosis and treatment susceptibilities. In contrast, our understanding of DCIS biology and the mechanisms of progression to IBC is less mature. There is significant interest in identifying the nature of molecular events involved in disease progression to determine those precursor lesions likely to progress to malignancy and therefore to provide individualized treatment to patients with DCIS. To date, molecular analyses of DCIS have been limited by scant availability of frozen tissue for molecular studies, shortage of DCIS tissue biorepositories with long-term follow up, and varying treatment modalities. In addition, the small size of DCIS lesions in many cases precludes analysis of whole tissue sections and requires microdissection to analyze lesional cells and avoid contamination with normal tissue.
Several studies evaluating gene expression profiles in DCIS demonstrated that there was remarkable similarity of the neoplastic epithelial cells of DCIS and IBC [12, 13, 14]. However, other investigators comparing transcriptome of DCIS and IBC have identified stage-specific markers and a gene expression classifier that differentiate DCIS and IBC [29, 30]. Another recent study demonstrated that genes conferring invasive growth are present only in a subset of DCIS cases [36]. Our findings indicate that there are intrinsic differences in the gene expression program of DCIS and IBC. Importantly, these alterations are distinct from tumor grade, ER/PR status, and Her2 status, indicating that there is a general basis for invasive behavior. It has been postulated that alterations in the tumor microenvironment (i.e., stroma) are crucial for progression to invasive disease. In fact, prior studies reported most transcriptional changes are evident in the stroma surrounding DCIS versus IBC, as opposed to the lesions themselves [12–14]. This study demonstrated that the largest degree of alterations in gene expression between pure DCIS and IBC is present in the epithelial/tumor compartment. Importantly, the classification potential of these alterations was subsequently validated in multiple independent data sets. While there are clearly differences in the stroma as well, the overall number of significant genes and relevant pathways in that compartment involved in progression to invasive behavior remain elusive. While we did initially observe statistically significant differences between DCIS and IBC stroma, after adjusting for multiple testing, very few genes passed this more rigorous statistical filtering and relevant pathways in that compartment involved in progression to invasive behavior are unclear.
To examine the concept that invasive potential is predetermined at the early stages of breast carcinogenesis, we compared data sets of DCIS associated with the IBC against our pure DCIS that did not progress to IBC on at least 8 years follow up. These analyses indicate that in DCIS that is associated with IBC there is already a significant induction of gene expression, and particularly genes associated with invasion are largely deregulated in DCIS associated with IBC. This finding likely explains why there is little difference in gene expression in the epithelial compartment of DCIS that are microdissected from invasive disease [12–14]. Correspondingly, cases of mixed DCIS/IBC behave essentially as IBC, suggesting that such lesions that have the capacity to progress largely express the gene expression program of invasive disease [36]. These findings illustrate that there exists a clear molecular distinction between pure DCIS and DCIS which is associated with invasive disease and that may already have undergone changes in gene expression profiles to become more similar to that of IBC.
Gene expression programs associated with invasion are associated with the progression from DCIS to IBC. Upregulation of genes associated with EMT and myoepithelial cell specific genes was seen in IBC when compared to pure DCIS, and these changes were highly reproducible between independent data sets. The development of the ability to invade the surrounding tissue is perhaps the most critical event in cancer progression. Tumor cell invasion is a multistep process, of which the key events include increased migration, increased protease secretion, and altered adhesion to allow dissemination from primary tumor sites [37, 38]. While EMT has been implicated in tumor invasion partly by reducing cell–cell adhesion, the myoepithelial phenotype is specifically implicated in increased cell motility [39]. Thus, such alterations in gene expression specifically associated with disease state (in situ versus invasive carcinoma), are clearly consistent with multi-step processes driving the invasive behavior. This finding is unique as many processes remain common between DCIS and IBC. For example, both DCIS and IBC demonstrate similar proliferation rates, and similar levels of cell cycle-related genes were observed between these stages of disease (not shown). GSEA showed marked overlap between genes upregulated in IBC transcriptome in our study and myoepithelial genes described by Jones and Grigoriadis [27, 40]. Consistent with these findings, other studies have previously reported expression of several other myoepithelial markers identified in our study (e.g., POSTN, FN, SPARC, LUM, COL1A2, COL1A11, and CAV-1) in IBC. Furthermore and consistent with our findings expression of several myoepithelial markers such as S100A2, SPARC, and maspin was associated with a poor prognosis in IBC, especially in ER-negative breast cancers. Correspondingly, EMT has been implicated in mediating both invasion and metastases. During EMT epithelial cells convert to a mesenchymal cell phenotype after losing cell polarity, disassembling cell–cell adhesion machinery, and subsequently acquiring increased cell mobility [39]. An EMT core signature has been recently described using gene expression changes induced in human mammary epithelial cells induced to undergo EMT [26]. GSEA showed enrichment of EMT core signature genes among genes differentially expressed between IBC and DCIS. EMT genes were clearly upregulated in IBC and were associated similarly with poor outcome in ER-negative breast cancer. These findings agree well with the prior association of EMT markers with poor outcome in basal like IBC [26]. At present, how EMT influences prognosis versus therapeutic response in ER-negative breast cancer remains unknown. In our cohorts, there are insufficient patient samples annotated for therapy to rigorously support conclusions based on treatment (n = 26). However, in patients treated with adjuvant chemotherapy a high expression of EMT signature was associated with poor overall survival (P < 0.01). Together, these findings indicate that myoepithelial and EMT phenotypes are associated with the conversion to invasive disease in all subtypes of breast cancer, but associate specifically with poor outcome in ER-negative disease.
Recently, it has been suggested that pathologic features and markers driving DCIS recurrence and progression to IBC may vary. From the clinical history of DCIS it is clear that while some patients will experience multiple recurrences in the form of DCIS over extended periods of time, others will rapidly progress to IBC. In a case–control study of 619 DCIS cases treated with a breast conserving surgery without radiation or hormonal therapy (225 DCIS with the recurrence and 395 without recurrence), larger DCIS lesions and close/positive excision margins were associated with a DCIS recurrence but did not predict increased risk of invasive recurrence. In fact, none of the evaluated pathologic factors was predictive of the invasive recurrence. Additional studies with larger cohorts of DCIS will be needed to determine if the relative expression of the DCIS/IBC classifiers defined here are relevant prognosticators for disease progression. However, one can envision that the gene expression program of the DCIS at the time of the diagnosis could determine the rate of the progression to IBC. Such knowledge in association with clinical features (age, co-morbidities) could become a base for the development of the clinical tests allowing for the accurate prognostication of the DCIS patients. This would have a dramatic impact on DCIS treatment and allow identification of a low risk group of the patients who require follow up only, and a high risk group needing additional therapy.
Ethical standards
The experiments detailed in this article comply with the current laws of the country in which they were performed.
References
Leonard GD, Swain SM (2004) Ductal carcinoma in situ, complexities and challenges. J Natl Cancer Inst 96(12):906–920
Collins LC, Tamimi RM, Baer HJ, Connolly JL, Colditz GA, Schnitt SJ (2005) Outcome of patients with ductal carcinoma in situ untreated after diagnostic biopsy: results from the Nurses’ Health Study. Cancer 103(9):1778–1784
Wang SY, Shamliyan T, Virnig BA, Kane R (2011) Tumor characteristics as predictors of local recurrence after treatment of ductal carcinoma in situ: a meta-analysis. Breast Cancer Res Treat 127(1):1–14
Schnitt SJ (2010) Local outcomes in ductal carcinoma in situ based on patient and tumor characteristics. J Natl Cancer Inst Monogr 2010(41):158–161
Tsikitis VL, Chung MA (2006) Biology of ductal carcinoma in situ classification based on biologic potential. Am J Clin Oncol 29(3):305–310
Sakorafas GH, Farley DR, Peros G (2008) Recent advances and current controversies in the management of DCIS of the breast. Cancer Treat Rev 34(6):483–497
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Eystein Lonning P, Borresen-Dale AL (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
Buyse M, Loi S, van’t Veer L, Viale G, Delorenzi M, Glas AM, d’Assignies MS, Bergh J, Lidereau R, Ellis P, Harris A, Bogaerts J, Therasse P, Floore A, Amakrane M, Piette F, Rutgers E, Sotiriou C, Cardoso F, Piccart MJ (2006) Validation and clinical utility of a 70-gene prognostic signature for women with node-negative breast cancer. J Natl Cancer Inst 98(17):1183–1192
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, Bryant J, Wolmark N (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351(27):2817–2826
Ma XJ, Hilsenbeck SG, Wang W, Ding L, Sgroi DC, Bender RA, Osborne CK, Allred DC, Erlander MG (2006) The HOXB13:IL17BR expression index is a prognostic factor in early-stage breast cancer. J Clin Oncol 24(28):4611–4619
Ma XJ, Salunga R, Tuggle JT, Gaudet J, Enright E, McQuary P, Payette T, Pistone M, Stecker K, Zhang BM, Zhou YX, Varnholt H, Smith B, Gadd M, Chatfield E, Kessler J, Baer TM, Erlander MG, Sgroi DC (2003) Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci USA 100(10):5974–5979. doi:10.1073/pnas.0931261100
Ma XJ, Dahiya S, Richardson E, Erlander M, Sgroi DC (2009) Gene expression profiling of the tumor microenvironment during breast cancer progression. Breast Cancer Res 11(1):R7
Allinen M, Beroukhim R, Cai L, Brennan C, Lahti-Domenici J, Huang H, Porter D, Hu M, Chin L, Richardson A, Schnitt S, Sellers WR, Polyak K (2004) Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 6(1):17–32
Holland R, Peterse JL, Millis RR, Eusebi V, Faverly D, van de Vijver MJ, Zafrani B (1994) Ductal carcinoma in situ: a proposal for a new classification. Semin Diagn Pathol 11(3):167–180
Elston EW, Ellis IO (1993) Method for grading breast cancer. J Clin Pathol 46(2):189–190
Moeder CB, Giltnane JM, Harigopal M, Molinaro A, Robinson A, Gelmon K, Huntsman D, Camp RL, Rimm DL (2007) Quantitative justification of the change from 10% to 30% for human epidermal growth factor receptor 2 scoring in the American Society of Clinical Oncology/College of American Pathologists guidelines: tumor heterogeneity in breast cancer and its implications for tissue microarray based assessment of outcome. J Clin Oncol 25(34):5418–5425
Fuller AP, Palmer-Toy D, Erlander MG, Sgroi DC (2003) Laser capture microdissection and advanced molecular analysis of human breast cancer. J Mammary Gland Biol Neoplasia 8(3):335–345
Storey JD (2002) A direct approach to false discovery rates. J R Stat Soc Ser B 64(3):479–498. doi:10.1111/1467-9868.00346
Gruvberger-Saal SK, Bendahl PO, Saal LH, Laakso M, Hegardt C, Eden P, Peterson C, Malmstrom P, Isola J, Borg A, Ferno M (2007) Estrogen receptor beta expression is associated with tamoxifen response in ERalpha-negative breast carcinoma. Clin Cancer Res 13(7):1987–1994. doi:10.1158/1078-0432.CCR-06-1823
Zaravinos A, Lambrou GI, Boulalas I, Delakas D, Spandidos DA (2010) Identification of common differentially expressed genes in urinary bladder cancer. PLoS One 6(4):e18135. doi:10.1371/journal.pone.0018135
Hammoud SS, Nix DA, Zhang H, Purwar J, Carrell DT, Cairns BR (2009) Distinctive chromatin in human sperm packages genes for embryo development. Nature 460(7254):473–478. doi:10.1038/nature08162
Reiner A, Yekutieli D, Benjamini Y (2003) Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics 19(3):368–375
Hackstadt AJ, Hess AM (2009) Filtering for increased power for microarray data analysis. BMC Bioinform 10:11. doi:10.1186/1471-2105-10-11
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci 98(9):5116–5121. doi:10.1073/pnas.091062498
Taube JH, Herschkowitz JI, Komurov K, Zhou AY, Gupta S, Yang J, Hartwell K, Onder TT, Gupta PB, Evans KW, Hollier BG, Ram PT, Lander ES, Rosen JM, Weinberg RA, Mani SA (2010) Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc Natl Acad Sci USA 107(35):15449–15454. doi:10.1073/pnas.1004900107
Grigoriadis A, Mackay A, Reis-Filho JS, Steele D, Iseli C, Stevenson BJ, Jongeneel CV, Valgeirsson H, Fenwick K, Iravani M, Leao M, Simpson AJ, Strausberg RL, Jat PS, Ashworth A, Neville AM, O’Hare MJ (2006) Establishment of the epithelial-specific transcriptome of normal and malignant human breast cells based on MPSS and array expression data. Breast Cancer Res 8(5):R56. doi:10.1186/bcr1604
Kim H, Watkinson J, Varadan V, Anastassiou D (2010) Multi-cancer computational analysis reveals invasion-associated variant of desmoplastic reaction involving INHBA, THBS2 and COL11A1. BMC Med Genomics 3:51. doi:10.1186/1755-8794-3-51
Schuetz CS, Bonin M, Clare SE, Nieselt K, Sotlar K, Walter M, Fehm T, Solomayer E, Riess O, Wallwiener D, Kurek R, Neubauer HJ (2006) Progression-specific genes identified by expression profiling of matched ductal carcinomas in situ and invasive breast tumors, combining laser capture microdissection and oligonucleotide microarray analysis. Cancer Res 66(10):5278–5286
Hannemann J, Velds A, Halfwerk JB, Kreike B, Peterse JL, van de Vijver MJ (2006) Classification of ductal carcinoma in situ by gene expression profiling. Breast Cancer Res 8(5):R61. doi:10.1186/bcr1613
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102(43):15545–15550. doi:10.1073/pnas.0506580102
Huang da W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. doi:10.1038/nprot.2008.211
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles–database and tools update. Nucleic Acids Res 35(suppl_1):D760–D765. doi:10.1093/nar/gkl887
Brazma A, Parkinson H, Sarkans U, Shojatalab M, Vilo J, Abeygunawardena N, Holloway E, Kapushesky M, Kemmeren P, Lara GG, Oezcimen A, Rocca-Serra P, Sansone SA (2003) ArrayExpress—a public repository for microarray gene expression data at the EBI. Nucleic Acids Res 31(1):68–71
Ertel A, Dean JL, Rui H, Liu C, Witkiewicz AK, Knudsen KE, Knudsen ES (2010) RB-pathway disruption in breast cancer: differential association with disease subtypes, disease-specific prognosis and therapeutic response. Cell Cycle 9(20):4153–4163
Muggerud AA, Hallett M, Johnsen H, Kleivi K, Zhou W, Tahmasebpoor S, Amini RM, Botling J, Borresen-Dale AL, Sorlie T, Warnberg F (2010) Molecular diversity in ductal carcinoma in situ (DCIS) and early invasive breast cancer. Mol Oncol 4(4):357–368
Liotta LA, Stetler-Stevenson WG (1991) Tumor invasion and metastasis: an imbalance of positive and negative regulation. Cancer Res 51(18 Suppl):5054s–5059s
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70
Guarino M (2007) Epithelial-mesenchymal transition and tumour invasion. Int J Biochem Cell Biol 39(12):2153–2160
Jones C, Mackay A, Grigoriadis A, Cossu A, Reis-Filho JS, Fulford L, Dexter T, Davies S, Bulmer K, Ford E, Parry S, Budroni M, Palmieri G, Neville AM, O’Hare MJ, Lakhani SR (2004) Expression profiling of purified normal human luminal and myoepithelial breast cells: identification of novel prognostic markers for breast cancer. Cancer Res 64(9):3037–3045
Conflict of interest
Authors Knudsen, Ertel, Kline, Schwartz, and Witkiewicz declare that they have no conflict of interest. Author Dr. Elai Davicioni declares remuneration and stock ownership in GenomeDx Biosciences, Inc.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Knudsen, E.S., Ertel, A., Davicioni, E. et al. Progression of ductal carcinoma in situ to invasive breast cancer is associated with gene expression programs of EMT and myoepithelia. Breast Cancer Res Treat 133, 1009–1024 (2012). https://doi.org/10.1007/s10549-011-1894-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1894-3