Abstract
The skin’s epidermis is an essential barrier as the first guard against invading pathogens, and physical protector from external injury. The skin microbiome, which consists of numerous bacteria, fungi, viruses, and archaea on the epidermis, play a key role in skin homeostasis. Antibiotics are a fast-acting and effective treatment method, however, antibiotic use is a nuisance that can disrupt skin homeostasis by eradicating beneficial bacteria along with the intended pathogens and cause antibiotic-resistant bacteria spread. Increased numbers of antimicrobial peptides (AMPs) derived from humans and bacteria have been reported, and their roles have been well defined. Recently, modulation of the skin microbiome with AMPs rather than artificially synthesized antibiotics has attracted the attention of researchers as many antibiotic-resistant strains make treatment mediation difficult in the context of ecological problems. Herein, we discuss the overall insights into the skin microbiome, including its regulation by different AMPs, as well as their composition and role in health and disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
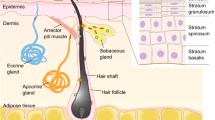
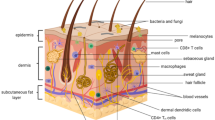
Skin microorganisms exert various effects on the skin through interactions between themselves or the host. The skin microbiome serves as a physical barrier that protects the skin from invading pathogens and plays an important role in the development of the host immune system (Belkaid & Segre, 2014). The importance of the skin microbiome is evidenced when the skin’s microbial barrier collapses and the resulting imbalance between commensal microorganisms and pathogens is closely related to skin health and diseases (Sanford & Gallo, 2013). Most skin commensal microorganisms are known to be harmless to humans, but certain strains like Staphylococcus aureus can cause infections triggered by host immunodeficiency. During the past 80 years, antibiotics have been widely used as a treatment for these microbial infections on the skin and other various body parts. Particularly, mupirocin, which can be hydrolyzed in vivo and is also used as a nasal ointment, is a representative antibiotic for the treatment of bacterial skin infections (van Rijen et al., 2008). It inhibits protein synthesis of various bacteria such as staphylococci, methicillin-resistant Staphylococcus aureus (MRSA), Streptococcus, Escherichia coli, and Haemophilus influenzae (Hughes & Mellows, 1978). In addition, fusidic acid, neomycin, and bacitracin have been used as topical treatments for skin infections in the form of ointments, creams, and gels (Werner & Russell, 1999). However, the widespread emergence of antibiotic resistance just over 60 years after antibiotics were introduced into clinical medicine continues to cause treatment failures and fatalities from various infectious diseases (Fair & Tor, 2014). Furthermore, overuse and misuse of antibiotics is a severe public health problem worldwide, especially in low and middle-income countries (Porter et al., 2020). Antibiotics reshape the skin microbiome ecology in profound ways during treatment (Langdon et al., 2016). Therefore, it is more important than ever to revisit how we use antibiotics, and treatment of the skin diseases must be complemented by efficient methods of restoring the microbiome into a balanced community after injury or infection (Sfriso et al., 2020). Antimicrobial peptides (AMPs) have been evaluated as novel antimicrobial drugs due to their extensively characterized mechanisms of action. Although concerns about the occurrence of bacteria that develop AMP resistance and their cross-resistance are still under debate, AMPs are worthy of study as it is used for therapeutic treatment to target microorganisms by preventing commensal microbiome collapse (Hancock & Sahl, 2006; Sang & Blecha, 2008) (Fig. 1).
Various bacteria cover the surface of the skin. Skin microbiome is shaped by interactions between microorganisms and the host. Antimicrobial peptides (AMPs) secreted by host and bacteria play an important role in infection and immunity, and bacteriocins involved in inter- and intra-species competition among Staphylococcus, Cutibacterium, Streptococcus have been widely reported. Taking it a step further than antibiotics and bacterial eluates, sustainable therapeutics should be developed in order to maintain a well-balanced microbiome
In this review, we discuss naturally occurring peptides resulting from bacterial interactions and their potential effects on the host skin microbiome. Furthermore, we introduce the types and mechanisms of natural molecules derived from the host and bacteria, and examples of their application in therapeutics and cosmetics.
Composition of the Skin Microbiome
Culture-based methods have been extensively used to explore skin microbial communities (Kong & Segre, 2012). However, some skin residential bacteria require demanding growth conditions and cannot be easily isolated in a laboratory environment (Kong, 2011). While skin microbiome research has focused on investigating a limited number of individual bacterial species associated with skin diseases, current human skin microbiome research uses culture-independent methods such as 16S ribosomal RNA (rRNA) genes and internal transcribed spacer (ITS) amplicon sequencing, metagenomics, and single cell-genomics to investigate the entire skin microbiome composition (De Filippis et al., 2017; Gao et al., 2010; Mathieu et al., 2014). Further, taxonomic and functional skin microbiome studies at the species or strain level are being conducted through metagenome-assumed genomes (MAGs), which recover the genomes of microorganisms from metagenomic data, and protocols for the discovery of skin MAGs have been published (Saheb Kashaf et al., 2022).
With the development of sequencing technology, various skin microbiome studies have been conducted, and it has been established that there are differences in microbial composition depending on the topological skin location. It was found that the lipophilic Cutibacterium was the most abundant bacteria on the sebaceous skin site, while Staphylococcus and Corynebacterium were preferentially abundant on the moist skin site (Grice & Segre, 2011). Many Gram-positive bacteria such as Staphylococcus spp., Cutibacterium spp., Corynebacterium spp., and Streptococcus spp. occupy most of the skin microbiome, and among them, Staphylococcus epidermidis, Staphylococcus aureus, and Cutibacterium acnes are attracting attention in skin microbiome research. (Chiller et al., 2001). Staphylococcus aureus, which most people carry asymptomatically, behaves as the main pathogen on the skin and is particularly associated with atopic dermatitis (Williams & Gallo, 2015). Cutibacterium acnes dominates significantly sebaceous skin like acne disease sites and produces many toxins and peptides related to immunomodulation (O’Neill & Gallo, 2018). Coagulase-negative staphylococci (CoNS), one of the most abundant colonizers of all skin regions, actively contributes to the maintenance of skin homeostasis by priming skin immunity, controlling other resident flora, and preventing opportunistic pathogen colonization (Parlet et al., 2019). Staphylococcus epidermidis, which can potentially cause nosocomial infections, is a mutually resident species that is essentially harmless to healthy humans. Staphylococcus epidermidis not only participates in a microbe-microbe interaction that secretes AMPs capable of killing S. aureus, but also engages host immunity by promoting AMPs secretion in human keratinocytes to protect the skin from infections (Chen et al., 2018).
Bacteria-Host Interaction via Host-Derived AMPs
Host defenses on the skin are composed of two complementary interacting systems: innate immunity, which protects against microorganisms in general, and adaptive immunity, which protects against a specific microorganism (Levinson et al., 2022). Among the various in vivo and in vitro host defense mechanisms, the type directly related to the skin microbiome is the physical barrier, such as the intact outer skin layer, which is classified as an innate defense system (Aristizábal & González, 2013). In addition to the physical barrier provided by the skin, the fatty acids secreted by the skin’s sebaceous glands have antibacterial activity (Drake et al., 2008). In turn, chemical barrier molecule-inducing proteases and cytokines that inhibit bacterial invasion are synthesized by epithelial cells of the host skin surface. These include free sphingosine, dihydrosphingosine, lauric acid, and sapienoic acid (Fischer et al., 2012).
There are several well-known human skin-derived antimicrobial peptides and proteins, ranging from 10–130 amino acids long, including human β-defensin 1,2,3 (hBD-1,2,3), cathelicidin LL-37, ribonuclease 7 (RNase 7), dermcidin and psoriasin S100A7 (Rademacher et al., 2021). Of these, only hBD-1,2,3 and LL-37 belong to the AMP classification with amino acids 10–50 in length (Cole and Nizet, 2016). They each exhibit their own antimicrobial killing efficacy, immunomodulatory functions, and chemotactic activity on the skin microbiome. For example, S. aureus growth can be regulated by blocking the antimicrobial activity of RNase 7, hBD-2, and hBD-3 (Kisich et al., 2007). However, bacteria have evolved strategies such as structural changes, protease secretion, and biofilm formation to overcome host immune response and defense mechanisms (Roy et al., 2018).
On the skin, Gram-positive pathogens such as staphylococci and enterococci adhere to the host surface by their microbial surface components recognizing adhesive matrix molecules (MSCRAMMs). These adhesins recognize fibronectin, collagen, and fibrinogen to form ligand-binding structures as virulence mediates microbial colonization (Foster, 2019).
Many virulence molecules including Intracellular adhesion (Ica), and Phenol-soluble modulins (PSMs), as well as MSCRAMMs, play critical roles. There are two steps in biofilm formation: the bacteria first adhere to the host surface and then accumulate into a complex biofilm architecture (Mack et al., 2006). In the accumulative phase of biofilm formation, most of the specific Ica from bacteria interact with extracellular matrix components on the surface. For example, S. epidermidis and S. aureus have an exclusive AMP sensor system (aps) and produce polysaccharide intercellular adhesin (PIA), which is a component of the extracellular matrix (Li et al., 2007a). Mutants deficient in both aps and PIA showed susceptibility toward cationic hBD-3 and LL-37. Homologs of the aps are also found in many Gram-positive bacteria, including S. haemolyticus, S. pneumonia, and Bacillus anthracis (Li et al., 2007b). In Gram-negative bacteria, the PhoPQ two-component system and its homologs are involved in AMPs resistance (Joo et al., 2016b).
Phenol-soluble modulins (PSMs) are a series of amphipathic peptides specifically found in Staphylococcus and, composed of 20–40 amino acids with a formyl-methionine group at the N-terminus (Wang et al., 2007). PSMs are named after a process in which a proinflammatory complex, which divides into three types: PSMα, PSMβ, PSMγ (= δ -toxin), was found in the supernatant of an S. epidermidis culture during high-temperature phenol extraction (Mehlin et al., 1999). PSMα is composed of 20–25 relatively short amino acids, contains an α-helix structure, and is cytotoxic. In contrast, PSMβ is composed of up to 44 amino acids and, unlike PSMα, only half of the C-terminal forms an α-helix structure and is not cytotoxic (Otto, 2014). It has been suggested that Staphylococcus, which cannot migrate on its own, increases its mobility by secreting PSMs on the surface of host cells. PSM has been used to emulsify cells to increase the uptake of extracellular nutrients for survival (Peschel & Otto, 2013). PSM’s representative function is to act as an immune modulator and toxin. When PSM concentration in the body is low, it binds to Formyl Peptide Receptor 2 (FPR2) of immune cells and induces various inflammation-promoting reactions in the body, such as chemotaxis and cytokine production. Otherwise, when PSM concentration in the body is high, PSM acts as a cytotoxin that dissolves host cells (Kretschmer et al., 2010). In addition, the amphiphilic nature of PSM, which has the properties of a unique surfactant, helps the structural maturation of the biofilm. When PSM forms a biofilm, it creates a microscopic pathway for the movement of nutrients and oxygen. In addition, PSMs sometimes detach the bacteria that are attached to the biofilm and transfer them to other host organs, causing infection (Periasamy et al., 2012). A recent study on the relationship between PSMs and biofilm stability found that S. aureus PSMs contribute to biofilm stabilization by forming amyloid, whereas S. epidermidis PSMs do not form amyloid and its infection phenotype is in accordance with the PSM biofilm structuring and detachment model in vivo (Le et al., 2019).
There are intimate associations between the host immune system and PSMs. T cells are the central lymphocytes of the adaptive immune system and are categorized by function, including helper T cells, cytotoxic T cells, regulatory T cells, and memory T cells. Among them, helper T cells, which include TH1, TH2, and TH17, assist with the maturation of B cells and activation of cytotoxic T cells and macrophages (Gutcher and Becher, 2007). The TH17 response has been shown to be clinically important in atopic dermatitis (AD) (Di Cesare et al., 2008). Proteases secreted from S. aureus play a critical role in their ability to penetrate deeper layers of the epidermis and dermis, thus triggering maximal TH2 and TH17 inflammatory responses to stimulate keratinocytes (Williams et al., 2019). According to previous reports that show PSMα could induce many proinflammatory cytokines, keratinocyte secretion may be important for subsequent TH17 responses in murine skin. This may explains an important component of how S. aureus accessory gene regulator (Agr)-modulating toxins, such as PSMα, stimulate skin inflammation (Damour et al., 2021). The positive contribution of bacteriocins to maintaining skin homeostasis in the host is not only their pathogen elimination activity but also their role as colonizing and signaling peptides.
Bacteria-Bacteria Communication on the Human Skin
The interactions between the human skin and the microorganisms that constitute the skin microbiome can be characterized by a variety of properties, ranging from mutual and symbiotic relationships to parasitic relationships. Also, there are noticeable interactions between different microbial species or strains of the skin microbiome where the skin may be in a state of homeostatic disturbance (Schommer & Gallo, 2013). Similarly, many microbial communities live in highly competitive surroundings, fight for survival, protect their genes, and pass on their genes to the next generation (Bauer et al., 2018).
There are two modes of bacterial competition: (1) exploitative and (2) interference through using specialized metabolites, multifunctional metabolites, secreted enzymes, and extracellular vesicles (Stubbendieck & Straight, 2016). Because microorganisms need protection from antibiotics or to compete with other microbes for energy sources, they engage in species-specific collective action called quorum sensing (QS) (Miller & Bassler, 2001). The QS system is the environmentally dependent chemical communication system between each bacterium and groups of bacteria. The basic role of QS seems to be the overall control of the physiology of bacterial populations in response to dynamically changing environments (Mukherjee & Bassler, 2019). This control is exerted both at different bacterial populations and the bacterial-host boundary as a synthesis of virulence factors, biofilm formation, and enzyme and siderophore production (Heilmann et al., 2015). It is known to control biofilm formation and the antibiotic resistance system by detecting the minimum threshold concentration. QS consists of signaling molecules, known as autoinducers (AIs), which are released as chemical messengers in proportion to cell density (Papenfort & Bassler, 2016). The Agr system is a representative QS system of the Gram-positive bacteria, Staphylococcus aureus (Yarwood et al., 2004). The Agr system produces and senses the auto-inducing peptide (AIP) produced by AgrD and AgrB (Ji et al., 1997). AgrD encodes the post-translationally modified AIP, and AgrB, the transmembrane protein, trims the AIP precursor to transport the AIP to the external cellular space (Vuong et al., 2000). When the AIP accumulates in the external cellular space, AgrC and AgrA, a two-component membrane regulator pair, become involved in the system (Geisinger et al., 2009). AIP binding to AgrC leads to the phosphorylation of AgrA which induces the expression of a regulatory RNA called RNAIII and the agrBDCA operon (Bronesky et al., 2016; Kavanaugh & Horswill, 2016). AIPs have been widely discovered in Gram-positive bacteria, and are specific to species and strains such as Staphylococcus, Clostridium, and Enterococcus (Monnet et al., 2016). In contrast, Gram-negative bacteria such as Pseudomonas, Acinetobacter, and Burkholderia use the acyl-homoserine lactones, which are composed of a lactone ring and an aliphatic acyl chain, as autoinducers (Schuster et al., 2013).
When different bacterial populations compete for common resources, the ability to disrupt QS can be advantageous to one bacterial species over another species that requires QS. Thus, QS interferences, called quorum quenching (QQ) mechanisms, have evolved to interfere with bacterial cell–cell communication (Fetzner, 2015). In the year 2000, the QQ phenomenon was first discovered in Erwinia carotovora by the identification of a QQ enzyme capable of cleaving autoinducer signaling (Dong et al., 2000). The outstanding difference between QS and QQ in wounded skin tissue is the sign of an infection. When a lesion occurs on the skin, bacteria colonize the wounded site to create a more hospitable environment by producing their own peptides. If the peptides are not degraded, they can initiate QS to secrete virulence factors and construct biofilms leading to infection. On the other hand, if the peptides are degraded, the bacteria remain in a harmless and defenseless state. Consequently, the wounded site remains colonized, but no infection occurs (Rémy et al., 2018).
PSM, the unique weapon of Staphylococcus, is useful not only for host cells but also for interacting with other bacteria. PSMs have been identified as major virulence factors in community-related MRSA and in the Staphylococcus species for which all genetic PSMs have been identified so far including S. epidermidis, S. aureus, and S. haemolyticus (Da et al., 2017; Peschel & Otto, 2013). PSM nomenclature varies by staphylococci species. In S. aureus, four types of PSMα (PSMα1-PSMα4) are encoded in the psmα operon, two types of PSMβ (PSMβ1 and PSMβ2) are encoded in the psmβ operon, and δ-toxin (also called PSMγ) is encoded in the hld (haemolysin delta) gene, which is located within the RNAIII coding sequence (Joo et al., 2016a). In S. epidermidis, the psmα and psmδ genes are located in an area of the genome corresponding to that of the S. aureus psmα operon. In addition, four types of PSMβ (PSMβ1a, PSMβ1b, PSMβ2, and PSMβ3) are encoded in the psmβ operon (Li et al., 2014). Although the role of individual PSMs in the biofilm formation process is not yet well understood, we demonstrate that PSMα and PSMβ have evolved to chemically cooperate to form stable amyloid structures while ensuring rapid and efficient biofilm formation (Zaman & Andreasen, 2020).
Bacterial Interspecies Competition Using AMPs
Bacteriocins, which are bacterial-derived antimicrobial peptides that kill other microorganisms, are a subset of antimicrobial peptides (Nishie et al., 2012).
Gram-positive bacteria-derived bacteriocins can be grouped into four classes: (I) lantibiotics, (II), non-lantibiotics, (III) large-sized bacteriocins, and (IV) uniquely structured bacteriocins (Simons et al., 2020). Lantibiotics, which are small peptides that contain lanthionine or 3-methyllanthionine and unusual post-translational modification residues, are further grouped into two categories: class Ia bacteriocins and class Ib bacteriocins (Lee & Kim, 2011). Lantibiotics Nisin A, subtilin, Pep5, epidermin, and gallidermin are representatives of class Ia lantibiotics (Bierbaum & Sahl, 2009). In contrast to class Ia lantibiotics, which are positively charged and are involved in bacterial membrane pore formation, class Ib lantibiotics are negatively charged and act in an enzyme-inhibiting manner of the target strain. Non-lantibiotics are heat-stable, naturally occurring small peptides, which include pediocin, sakacin, lactoccin, and their variants. This group can be subcategorized into four subclasses: class IIa, class IIb, class IIc, and class IId (Cotter et al., 2005). Staphylococcus is an affluent antibiotic source, most of which include Pep5, epidermin, and epicidin variants that are produced by S. epidermidis. Most bacteriocins from S. epidermidis strains belong to class II. In S. aureus, especially community-acquired methicillin-resistant S. aureus (MRSA), the bsa (bacteriocin of S. aureus) gene, which encodes lantibiotics with a broad antimicrobial spectrum, was found located on the vSaβ genomic island. It was found that this BSA conferred a competitive ecological advantage on the MRSA strain during infection (Daly et al., 2010).
Gram-negative bacteria-derived bacteriocins can be categorized into four classes: (I) colicins, (II) colicin-like, (III) microcins, and (IV) phage tail-like bacteriocins (Simons et al., 2020). Although most of the colicins from gram-negative bacteria are reportedly from E. coli, structurally similar colicins can be produced by other species, including Klebsiella spp. and Pseudomonas spp. (Riley & Chavan, 2007). Microcins and phage tail-like bacteriocins are mainly produced by bacteria in the genus belonging to the Enterobacteriaceae family and P. aeruginosa, respectively (Baquero et al., 2019; Michel-Briand & Baysse, 2002).
The skin microbiome and synthetic peptides from microbes offer many opportunities for skin therapeutics to support the treatment of infectious skin diseases. Although many virulence factors secreted from S. aureus have been suggested to contribute to AD pathogenesis, reduction of the S. aureus population through antibiotic therapy and bleach baths does not provide lasting symptom relief (Chopra et al., 2017). Recently, our understanding of environmental regulators and the distribution of the skin microbiome has been rapidly expanding through metagenomic approaches at the molecular level (Di Domenico et al., 2019). Specific bacteria can regulate the growth of S. aureus in diseased skin of AD patients. The therapeutic efficacy of QS inhibition has emerged as a promising paradigm for pharmacological interventions aimed at alleviating S. aureus-induced diseases (Tan et al., 2018). While relatively less studied than other small molecules, evidence suggests that staphylococcal interspecies competition exists. For example, S. epidermidis and S. caprae make an AIP that inhibits the S. aureus QS system. Cutaneous CoNS strains (S. epidermidis, S. haemolyticus, S. capitis, S. hominis, S. simulans, and S. warneri) can produce antimicrobial molecules that protect the skin from potential invaders including S. aureus (Nakatsuji et al., 2017). Among them, S. epidermidis and S. hominis produce bacteriocins targeting S. aureus. Cytoplasmic bacteriocin from live planktonic S. epidermidis exhibits antimicrobial activity against S. aureus and MRSA growth without activity against S. epidermidis and E. coli (Jang et al., 2020). Some staphylococci isolates showing antimicrobial activity against S. aureus have been collected from the nasal cavity. Among them, S. lugdunensis IVK28 produces lugdunin, a novel thiazolidine-containing cyclic peptide that has a particularly strong affinity for S. aureus eradication (Heilbronner & Foster, 2021). Gallidermin is a representative lantibiotic belonging to the large class of cationic antimicrobial peptides (CAMPs) derived from S. gallinarum, which not only inhibits the growth of staphylococci, especially MRSA, but also prevents biofilm formation (Saising et al., 2012).
In addition to the intraspecies interaction of Staphylococcus spp., we introduce some cases in which AMPs play a role as communicators from the main species present on the skin, Cutibacterium, Corynebacterium, Micrococcus, and Streptococcus. Cutibacterium acnes is related to acne, but how C. acnes mechanistically contributes to acne development is unclear. This bacterium produces coproporphyrin III, which induces the formation of S. aureus biofilms (Wollenberg et al., 2014). However, C. acnes may sometimes ferment glycerol into short-chain fatty acids, which suppress the growth of S. aureus (Nakamura et al., 2020). Corynebacterium accolens, a common skin resident, was recently shown to inhibit the growth of Streptococcus pneumoniae, a common respiratory tract pathogen (Horn et al., 2022). Corynebacterial lipase from C. accolens hydrolyses triolein to oleic acid, which inhibits pneumococcal growth. In addition, Corynebacterium pseudodiphtheriticum, a common member of the normal nasal microbiota, also mediates bactericidal activity against S. aureus, including MRSA. In response to these attacks, S. aureus gains resistance through insertional inactivation of agrC, which encodes the sensor kinase of the Agr QS system (Hardy et al., 2019). Furthermore, Corynebacterium striatum, suppresses pathogenic genes by shifting the global transcriptional program of co-cultured S. aureus and stimulating genes induced by the Agr QS system associated with commensalism (Ramsey et al., 2016). Antibiotic activity of the PSM derivative in which two N-terminal amino acids were removed from PSMα1 and PSMα2 (secreted by S. aureus against Micrococcus luteus and Streptococcus pyogenes) was several times higher than that of the complete PSM structure (Joo et al., 2011). For the immune-compromised host, the production of membrane-active PSMs by S. epidermidis can contribute to virulence leading to harmful cell lysis. However, it is important to note that, these PSMs have a unique ability to remove pathogens, such as S. aureus and Streptococcus pyogenes (Cheung et al., 2014). These immediate and selective mechanisms present an important strategy against colonization during sustentative microbiome preservation and could be used as next-generation anti-infective therapeutics.
Finally, to enable extracellular secretion, PSM self-regulates transcription factors of the ABC transporter operon (PSM transporter), which is a representative example of how toxins control their expression levels (Bojer et al., 2018). Almost all PSMs are regulated by the Agr quorum sensing system encoded within the agrBDCA operon. Preceding studies show that S. aureus toxins can exert antibacterial activity when chemically modified, but not as native peptides. In S. aureus, AgrA, not RNAIII, acts directly as a PSM operon regulator (Queck et al., 2008). Antimicrobial assays using peptides composed of four distinct PSMs, PSMβ1, 3, 4, and 6, showed that S. capitis E12 selectively inhibits the growth of C. acnes with greater potential than common antibiotics for acne treatment (O'Neill et al., 2020).
A common prokaryotic cell death mechanism used by the numerous bacteriocins mentioned above is the destruction of target cells by the inhibition of pore formation or cell wall synthesis (Bin Hafeez et al., 2021). As a defense mechanism against this process, bacteria also produce their own enzymes that resist AMPs. Bacterial proteases that hydrolyze the amide bonds of peptide units also contribute to polymicrobial dynamics, including how they are used in antimicrobial and anti-biofilm dynamics (Jiang et al., 2020). Lysostaphin, known as an example of the peptidoglycan hydrolases from Staphylococcus simulans, can degrade the cell wall of almost all Staphylococcus spp. It has been shown to target the unique structure of staphylococcal peptidoglycan to inhibit Staphylococcus spp., as well as kill MRSA by synergizing with bacteriophage lysins (Jayakumar et al., 2021). In addition, LasA protease and pyocyanin secreted from P. aeruginosa has similar anti-S. aureus activity, which has been used for the treatment of staphylococcal keratitis (Hotterbeekx et al., 2017). Finally, Bacillus tequilensis ZMS-2 was found to secrete an alkaline protease with antimicrobial activity against several clinically important human pathogens, including S. aureus, E. coli, and Klebsiella pneumoniae (Khan et al., 2019) (Table 1).
Current Application of the Skin Microbiome Regulation in Cosmetics
Deeper knowledge of the skin microbiome has widened perspectives on a revolution in dermo-cosmetic development, and these recent discoveries have changed our perception of the role of bacteria in skin health. New products that readjust the skin microbiome are ushering in a new trend in the dermo-cosmetics industry, and the development of skin microbiome analysis and diagnosis will enable the launch of microbiome-derived customized dermo-cosmetics optimized for each individual’s skin (Gueniche et al., 2022a; Kim et al., 2021a, b).
The International Scientific Association of Probiotics and Prebiotics (ISAPP) defined the scope of postbiotics as a preparation of inanimate microorganisms and/or their components that confers a health benefit on the host (Salminen et al., 2021). Ferments, extracts, lysates, and filtrates are used as postbiotic ingredients. Precedent research has suggested that the addition of bacterial lysates and eluates to cosmetics may provide benefits in skin infection treatment (Puebla-Barragan & Reid, 2021). Microbial lysates of Lactobacillus spp. are widely characterized and used in cosmetic products. Lysates of L. rhamnosus reconstruct the epidermis (Jung et al., 2019) and both L. salivarius and L. plantarum accelerate re-epithelialization to improve the skin barrier (Brandi et al., 2020). Lactobacillus plantarum CJLP55 was found to reduce erythema and reinforce the skin barrier in acne patients (Kim et al., 2021a, b). Lactobacillus paracasei NCC2461 (ST11) confers benefits to the skin to reinforce skin barrier function and decrease skin sensitivity and dandruff conditions (Reygagne et al., 2017). La1 from L. johnsonii has been shown to maintain the number and function of Langerhans cells after UV exposure to the skin and to regulate skin inflammation (Gueniche et al., 2022b).
Other bacterial extracts also have properties that may support wound healing of the skin. Lysates of Vitreoscilla filiformis significantly improved AD and seborrheic dermatitis by reinforcing the skin barrier and innate immunity via Toll-like receptor 2 activation (Guéniche et al., 2008). Extracts of S. thermophilus increase ceramide production to improve skin hydration and extracts of B. longum can decrease skin sensitivity by their tryptophan metabolite indole-3-carbaldehyde which activates the AHR-mediated immune signaling pathway based on the interactions of the gut-skin axis (Fang et al., 2022; Liu et al., 2022). A topical lotion containing an extract of Enterococcus faecalis SL-5 showed a significant C. acnes reduction in lesions (Kang et al., 2009). In a more recent study, LactoSporin, purified from Bacillus coagulans, was proven to be a postbiotic antibacterial agent for acne treatment that is more advanced in terms of safety and therapeutic effect (Majeed et al., 2020). Transplanting mixture of three C. acnes strains isolated from a healthy donor allows synergistic colonization in the diseased subject (Paetzold et al., 2019). Furthermore, there is abundant evidence that combining different strains tailored to the individual disease provides a greater therapeutic advantage as compared to applying only one specific strain (Kwoji et al., 2021).
Conclusion
While bacterial antibiotic resistance is the best adaptable mechanism to survive and evolve, it can be a potential hazard to alter the human microbiome, which can lead to the host becoming susceptible to infection. Therefore, when considering bacterial infections on the skin, it is important to differentiate whether the causative organism is an invading pathogen or a commensal opportunistic skin flora to choose the appropriate therapy. Although bacterial antibiotic resistance development has led to pharmacological advances, a crucial phase in which antibiotic resistance development is inevitable for all currently available antibiotics has been reached. Minimizing the development of antimicrobial resistance created by the existing methods of antibiotic therapy is a significant challenge.
It is now clear that diverse cell–cell communication peptides as well as human-derived AMPs mediate a prominent influence on the skin, as highlighted in this review. Since almost all antibiotics used to treat infections have side effects, the intercellular interaction of bacteria has recently attracted the attention of microbial ecologists. As stated above, some specific probiotic solutions based on the healthy skin microbiome may help to fluctuate the imbalanced microbiota back to a well-balanced state. However, there are no absolute guidelines on how to differentiate between “unconditionally good bacteria” and “unconditionally bad bacteria” or which peptides are beneficial for skin homeostasis at given skin areas and concentrations. Because the skin microbiome is a complicated environment where diverse microbes reside, there are bacteria that confer either beneficial or adverse effects on the skin relatively, and not unconditionally. Despite much research over the years, our knowledge about the role of bacterial-host/ bacterial-bacterial interaction to maintain skin homeostasis is still limited. Understanding the role of the skin microbiome in maintaining a normal skin condition and applying the knowledge gained by addressing the deleterious consequences of inflammation-causing microbial dysbiosis are promising directions for the development of novel therapies. The skin is a research-inexhaustible environment due to its complex composition; therefore, more functional and association studies are needed to gain more insight into AMPs-microbiota interactions.
References
Aristizábal, B., & González, Á. (2013). Innate immune system. In J. M. Anaya, Y. Shoenfeld, A. Rojas-Villarraga, R. A. Levy, & R. Cervera (Eds.), Autoimmunity: from bench to bedside. Bogota, Colombia: El Rosario University Press.
Baquero, F., Lanza, V. F., Baquero, M. R., del Campo, R., & Bravo-Vázquez, D. A. (2019). Microcins in Enterobacteriaceae: peptide antimicrobials in the eco-active intestinal chemosphere. Frontiers in Microbiology, 10, 2261.
Bauer, M. A., Kainz, K., Carmona-Gutierrez, D., & Madeo, F. (2018). Microbial wars: competition in ecological niches and within the microbiome. Microb. Cell, 5, 215–219.
Belkaid, Y., & Segre, J. A. (2014). Dialogue between skin microbiota and immunity. Science, 346, 954–959.
Bierbaum, G., & Sahl, H. G. (2009). Lantibiotics: Mode of action, biosynthesis and bioengineering. Current Pharmaceutical Biotechnology, 10, 2–18.
Bin Hafeez, A., Jiang, X., Bergen, P. J., & Zhu, Y. (2021). Antimicrobial peptides: an update on classifications and databases. International Journal of Molecular Sciences, 22, 11691.
Bojer, M. S., Lindemose, S., Vestergaard, M., & Ingmer, H. (2018). Quorum sensing-regulated phenol-soluble modulins limit persister cell populations in Staphylococcus aureus. Frontiers in Microbiology, 9, 255.
Brandi, J., Cheri, S., Manfredi, M., Di Carlo, C., Vita Vanella, V., Federici, F., Bombiero, E., Bazaj, A., Rizzi, E., Manna, L., et al. (2020). Exploring the wound healing, anti-inflammatory, anti-pathogenic and proteomic effects of lactic acid bacteria on keratinocytes. Science and Reports, 10, 11572.
Bronesky, D., Wu, Z., Marzi, S., Walter, P., Geissmann, T., Moreau, K., Vandenesch, F., Caldelari, I., & Romby, P. (2016). Staphylococcus aureus RNAIII and its regulon link quorum sensing, stress responses, metabolic adaptation, and regulation of virulence gene expression. Annual Review of Microbiology, 70, 299–316.
Chen, Y. E., Fischbach, M. A., & Belkaid, Y. (2018). Skin microbiota–host interactions. Nature, 553, 427–436.
Cheung, G. Y. C., Joo, H. S., Chatterjee, S. S., & Otto, M. (2014). Phenol-soluble modulins – critical determinants of staphylococcal virulence. FEMS Microbiology Reviews, 38, 698–719.
Chiller, K., Selkin, B. A., & Murakawa, G. J. (2001). Skin microflora and bacterial infections of the skin. The Journal of Investigative Dermatology. Symposium Proceedings, 6, 170–174.
Chopra, R., Vakharia, P. P., Sacotte, R., & Silverberg, J. I. (2017). Efficacy of bleach baths in reducing severity of atopic dermatitis: a systematic review and meta-analysis. Annals of Allergy, Asthma & Immunology, 119, 435–440.
Cole, J. N., & Nizet, V. (2016). Bacterial evasion of host antimicrobial peptide defenses. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.VMBF-0006-2015
Cotter, P. D., Hill, C., & Ross, R. P. (2005). Bacteriocins: developing innate immunity for food. Nature Reviews Microbiology, 3, 777–788.
Da, F., Joo, H. S., Cheung, G. Y. C., Villaruz, A. E., Rohde, H., Luo, X., & Otto, M. (2017). Phenol-soluble modulin toxins of Staphylococcus haemolyticus. Frontiers in Cellular and Infection Microbiology, 7, 206.
Daly, K. M., Upton, M., Sandiford, S. K., Draper, L. A., Wescombe, P. A., Jack, R. W., O’Connor, P. M., Rossney, A., Götz, F., Hill, C., et al. (2010). Production of the Bsa lantibiotic by community-acquired Staphylococcus aureus strains. Journal of Bacteriology, 192, 1131–1142.
Damour, A., Robin, B., Deroche, L., Broutin, L., Bellin, N., Verdon, J., Lina, G., Leclère, F. M., Garcia, M., Cremniter, J., et al. (2021). Phenol-soluble modulins α are major virulence factors of Staphylococcus aureus secretome promoting inflammatory response in human epidermis. Virulence, 12, 2474–2492.
De Filippis, F., Laiola, M., Blaiotta, G., & Ercolini, D. (2017). Different amplicon targets for sequencing-based studies of fungal diversity. Applied and Environment Microbiology, 83, e00905-e917.
Di Cesare, A., Di Meglio, P., & Nestle, F. O. (2008). A role for Th17 cells in the immunopathogenesis of atopic dermatitis? The Journal of Investigative Dermatology, 128, 2569–2571.
Di Domenico, E. G., Cavallo, I., Capitanio, B., Ascenzioni, F., Pimpinelli, F., Morrone, A., & Ensoli, F. (2019). Staphylococcus aureus and the cutaneous microbiota biofilms in the pathogenesis of atopic dermatitis. Microorganisms, 7, 301.
Dong, Y. H., Xu, J. L., Li, X. Z., & Zhang, L. H. (2000). AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proceedings of the National Academy of Sciences of the United States of America, 97, 3526–3531.
Drake, D. R., Brogden, K. A., Dawson, D. V., & Wertz, P. W. (2008). Thematic review series: skin lipids. Antimicrobial lipids at the skin surface. Journal of Lipid Research, 49, 4–11.
Fair, R. J., & Tor, Y. (2014). Antibiotics and bacterial resistance in the 21st century. Perspect. Medicin. Chem., 6, 25–64.
Fang, Z., Pan, T., Li, L., Wang, H., Zhu, J., Zhang, H., Zhao, J., Chen, W., & Lu, W. (2022). Bifidobacterium longum mediated tryptophan metabolism to improve atopic dermatitis via the gut-skin axis. Gut Microbes, 14, 2044723.
Fetzner, S. (2015). Quorum quenching enzymes. Journal of Biotechnology, 201, 2–14.
Fischer, C. L., Drake, D. R., Dawson, D. V., Blanchette, D. R., Brogden, K. A., & Wertz, P. W. (2012). Antibacterial activity of sphingoid bases and fatty acids against Gram-positive and Gram-negative bacteria. Antimicrobial Agents and Chemotherapy, 56, 1157–1161.
Foster, T. J. (2019). The MSCRAMM family of cell-wall-anchored surface proteins of Gram-positive cocci. Trends in Microbiology, 27, 927–941.
Gao, Z., Perez-Perez, G. I., Chen, Y., & Blaser, M. J. (2010). Quantitation of major human cutaneous bacterial and fungal populations. Journal of Clinical Microbiology, 48, 3575–3581.
Geisinger, E., Muir, T. W., & Novick, R. P. (2009). agr receptor mutants reveal distinct modes of inhibition by staphylococcal autoinducing peptides. Proceedings of the National Academy of Sciences of the United States of America, 106, 1216–1221.
Grice, E. A., & Segre, J. A. (2011). The skin microbiome. Nature Reviews Microbiology, 9, 244–253.
Guéniche, A., Cathelineau, A. C., Bastien, P., Esdaile, J., Martin, R., Queille Roussel, C., & Breton, L. (2008). Vitreoscilla filiformis biomass improves seborrheic dermatitis. Journal of the European Academy of Dermatology and Venereology, 22, 1014–1015.
Gueniche, A., Perin, O., Bouslimani, A., Landemaine, L., Misra, N., Cupferman, S., Aguilar, L., Clavaud, C., Chopra, T., & Khodr, A. (2022a). Advances in microbiome-derived solutions and methodologies are founding a new era in skin health and care. Pathogens, 11, 121.
Gueniche, A., Valois, A., Kerob, D., Rasmont, V., & Nielsen, M. (2022b). A combination of Vitreoscilla filiformis extract and Vichy volcanic mineralizing water strengthens the skin defenses and skin barrier. Journal of the European Academy of Dermatology and Venereology, 36, 16–25.
Gutcher, I., & Becher, B. (2007). APC-derived cytokines and T cell polarization in autoimmune inflammation. The Journal of Clinical Investigation, 117, 1119–1127.
Hancock, R. E. W., & Sahl, H. G. (2006). Antimicrobial and host-defense peptides as new anti-infective therapeutic strategies. Nature Biotechnology, 24, 1551–1557.
Hardy, B. L., Dickey, S. W., Plaut, R. D., Riggins, D. P., Stibitz, S., Otto, M., & Merrell, D. S. (2019). Corynebacterium pseudodiphtheriticum exploits Staphylococcus aureus virulence components in a novel polymicrobial defense strategy. mBio, 10, e02491-18.
Heilbronner, S., & Foster, T. J. (2021). Staphylococcus lugdunensis: a skin commensal with invasive pathogenic potential. Clinical Microbiology Reviews, 34, e00205-e220.
Heilmann, S., Krishna, S., & Kerr, B. (2015). Why do bacteria regulate public goods by quorum sensing?—How the shapes of cost and benefit functions determine the form of optimal regulation. Frontiers in Microbiology, 6, 767.
Horn, K. J., Jaberi Vivar, A. C., Arenas, V., Andani, S., Janoff, E. N., & Clark, S. E. (2022). Corynebacterium species inhibit Streptococcus pneumoniae colonization and infection of the mouse airway. Frontiers in Microbiology, 12, 804935.
Hotterbeekx, A., Kumar-Singh, S., Goossens, H., & Malhotra-Kumar, S. (2017). In vivo and in vitro interactions between Pseudomonas aeruginosa and Staphylococcus spp. Frontiers in Cellular and Infection Microbiology, 7, 106.
Hughes, J., & Mellows, G. (1978). Inhibition of isoleucyl-transfer ribonucleic acid synthetase in Echerichia coli by pseudomonic acid. The Biochemical Journal, 176, 305–318.
Jang, I. T., Yang, M., Kim, H. J., & Park, J. K. (2020). Novel cytoplasmic bacteriocin compounds derived from Staphylococcus epidermidis selectively kill Staphylococcus aureus, including methicillin-resistant Staphylococcus aureus (MRSA). Pathogens, 9, 87.
Jayakumar, J., Kumar, V. A., Biswas, L., & Biswas, R. (2021). Therapeutic applications of lysostaphin against Staphylococcus aureus. Journal of Applied Microbiology, 131, 1072–1082.
Ji, G., Beavis, R., & Novick, R. P. (1997). Bacterial interference caused by autoinducing peptide variants. Science, 276, 2027–2030.
Jiang, Y., Geng, M., & Bai, L. (2020). Targeting biofilms therapy: current research strategies and development hurdles. Microorganisms, 8, 1222.
Joo, H. S., Chatterjee, S. S., Villaruz, A. E., Dickey, S. W., Tan, V. Y., Chen, Y., Sturdevant, D. E., Ricklefs, S. M., & Otto, M. (2016a). Mechanism of gene regulation by a Staphylococcus aureus toxin. mBio, 7, e01579-16.
Joo, H.-S., Cheung, G. Y. C., & Otto, M. (2011). Antimicrobial activity of community-associated methicillin-resistant Staphylococcus aureus is caused by phenol-soluble modulin derivatives. Journal of Biological Chemistry, 286, 8933–8940.
Joo, H. S., Fu, C. I., & Otto, M. (2016b). Bacterial strategies of resistance to antimicrobial peptides. Philosophical Transactions of the Royal Society of London. Series B, 371, 20150292.
Jung, Y. O., Jeong, H., Cho, Y., Lee, E. O., Jang, H. W., Kim, J., Nam, K., & Lim, K. M. (2019). Lysates of a probiotic, Lactobacillus rhamnosus, can improve skin barrier function in a reconstructed human epidermis model. International Journal of Molecular Sciences, 20, 4289.
Kang, B. S., Seo, J. G., Lee, G. S., Kim, J. H., Kim, S. Y., Han, Y. W., Kang, H., Kim, H. O., Rhee, J. H., Chung, M. J., et al. (2009). Antimicrobial activity of enterocins from Enterococcus faecalis SL-5 against Propionibacterium acnes, the causative agent in acne vulgaris, and its therapeutic effect. Journal of Microbiology, 47, 101–109.
Kavanaugh, J. S., & Horswill, A. R. (2016). Impact of environmental cues on staphylococcal quorum sensing and biofilm development. Journal of Biological Chemistry, 291, 12556–12564.
Khan, Z., Shafique, M., Nawaz, H. R., Jabeen, N., & Naz, S. A. (2019). Bacillus tequilensis ZMS-2: A novel source of alkaline protease with antimicrobial, anti-coagulant, fibrinolytic and dehairing potentials. Pakistan Journal of Pharmaceutical Sciences, 32, 1913–1918.
Kim, G., Kim, M., Kim, M., Park, C., Yoon, Y., Lim, D. H., Yeo, H., Kang, S., Lee, Y. G., Beak, N. I., et al. (2021a). Spermidine-induced recovery of human dermal structure and barrier function by skin microbiome. Commun. Biol., 4, 231.
Kim, M. J., Kim, K. P., Choi, E., Yim, J. H., Choi, C., Yun, H. S., Ahn, H. Y., Oh, J. Y., & Cho, Y. (2021b). Effects of Lactobacillus plantarum CJLP55 on clinical improvement, skin condition and urine bacterial extracellular vesicles in patients with acne vulgaris: a randomized, double-blind, placebo-controlled study. Nutrients, 13, 1368.
Kisich, K. O., Howell, M. D., Boguniewicz, M., Heizer, H. R., Watson, N. U., & Leung, D. Y. M. (2007). The constitutive capacity of human keratinocytes to kill Staphylococcus aureus is dependent on β-defensin 3. The Journal of Investigative Dermatology, 127, 2368–2380.
Kong, H. H. (2011). Skin microbiome: genomics-based insights into the diversity and role of skin microbes. Trends in Molecular Medicine, 17, 320–328.
Kong, H. H., & Segre, J. A. (2012). Skin microbiome: looking back to move forward. The Journal of Investigative Dermatology, 132, 933–939.
Kretschmer, D., Gleske, A. K., Rautenberg, M., Wang, R., Köberle, M., Bohn, E., Schöneberg, T., Rabiet, M. J., Boulay, F., Klebanoff, S. J., et al. (2010). Human formyl peptide receptor 2 senses highly pathogenic Staphylococcus aureus. Cell Host & Microbe, 7, 463–473.
Kwoji, I. D., Aiyegoro, O. A., Okpeku, M., & Adeleke, M. A. (2021). Multi-strain probiotics: synergy among isolates enhances biological activities. Biology, 10, 322.
Langdon, A., Crook, N., & Dantas, G. (2016). The effects of antibiotics on the microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med., 8, 39.
Le, K. Y., Villaruz, A. E., Zheng, Y., He, L., Fisher, E. L., Nguyen, T. H., Ho, T. V., Yeh, A. J., Joo, H. S., Cheung, G. Y., et al. (2019). Role of phenol-soluble modulins in Staphylococcus epidermidis biofilm formation and infection of indwelling medical devices. Journal of Molecular Biology, 431, 3015–3027.
Lee, H., & Kim, H. Y. (2011). Lantibiotics, class I bacteriocins from the genus Bacillus. Journal of Microbiology and Biotechnology, 21, 229–235.
Levinson, W. E., Chin-Hong, P., Joyce, E. A., Nussbaum, J., & Schwartz, B. (2022). Review of medical microbiology and immunology (Vol. 17). New York, USA: McGraw Hill Professional.
Li, M., Cha, D. J., Lai, Y., Villaruz, A. E., Sturdevant, D. E., & Otto, M. (2007a). The antimicrobial peptide-sensing system aps of Staphylococcus aureus. Molecular Microbiology, 66, 1136–1147.
Li, M., Lai, Y., Villaruz, A. E., Cha, D. J., Sturdevant, D. E., & Otto, M. (2007b). Gram-positive three-component antimicrobial peptide-sensing system. Proceedings of the National Academy of Sciences of the United States of America, 104, 9469–9474.
Li, S., Huang, H., Rao, X., Chen, W., Wang, Z., & Hu, X. (2014). Phenol-soluble modulins: novel virulence-associated peptides of staphylococci. Future Microbiology, 9, 203–216.
Liu, C., Tseng, Y. P., Chan, L. P., & Liang, C. H. (2022). The potential of Streptococcus thermophiles (TCI633) in the anti-aging. Journal of Cosmetic Dermatology, 21, 2635–2647.
Mack, D., Rohde, H., Harris, L. G., Davies, A. P., Horstkotte, M. A., & Knobloch, J. K. M. (2006). Biofilm formation in medical device-related infection. International Journal of Artificial Organs, 29, 343–359.
Majeed, M., Majeed, S., Nagabhushanam, K., Mundkur, L., Rajalakshmi, H. R., Shah, K., & Beede, K. (2020). Novel topical application of a postbiotic, LactoSporin®, in mild to moderate acne: a randomized, comparative clinical study to evaluate its efficacy, tolerability and safety. Cosmetics, 7, 70.
Mathieu, A., Vogel, T. M., & Simonet, P. (2014). The future of skin metagenomics. Research in Microbiology, 165, 69–76.
Mehlin, C., Headley, C. M., & Klebanoff, S. J. (1999). An inflammatory polypeptide complex from Staphylococcus epidermidis: isolation and characterization. Journal of Experimental Medicine, 189, 907–918.
Michel-Briand, Y., & Baysse, C. (2002). The pyocins of Pseudomonas aeruginosa. Biochimie, 84, 499–510.
Miller, M. B., & Bassler, B. L. (2001). Quorum sensing in bacteria. Annual Review of Microbiology, 55, 165–199.
Monnet, V., Juillard, V., & Gardan, R. (2016). Peptide conversations in Gram-positive bacteria. Critical Reviews in Microbiology, 42, 339–351.
Mukherjee, S., & Bassler, B. L. (2019). Bacterial quorum sensing in complex and dynamically changing environments. Nature Reviews Microbiology, 17, 371–382.
Nakamura, K., O’Neill, A. M., Williams, M. R., Cau, L., Nakatsuji, T., Horswill, A. R., & Gallo, R. L. (2020). Short chain fatty acids produced by Cutibacterium acnes inhibit biofilm formation by Staphylococcus epidermidis. Science and Reports, 10, 21237.
Nakatsuji, T., Chen, T. H., Narala, S., Chun, K. A., Two, A. M., Yun, T., Shafiq, F., Kotol, P. F., Bouslimani, A., Melnik, A. V., et al. (2017). Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med., 9, eaah4680.
Nishie, M., Nagao, J. I., & Sonomoto, K. (2012). Antibacterial peptides “bacteriocins”: an overview of their diverse characteristics and applications. Biocontrol Science, 17, 1–16.
O’Neill, A. M., & Gallo, R. L. (2018). Host-microbiome interactions and recent progress into understanding the biology of acne vulgaris. Microbiome, 6, 177.
O’Neill, A. M., Nakatsuji, T., Hayachi, A., Williams, M. R., Mills, R. H., Gonzalez, D. J., & Gallo, R. L. (2020). Identification of a human skin commensal bacterium that selectively kills Cutibacterium acnes. The Journal of Investigative Dermatology, 140, 1619–1628.
Otto, M. (2001). Staphylococcus aureus and Staphylococcus epidermidis peptide pheromones produced by the accessory gene regulator agr system. Peptides, 22(10), 1603–1608.
Otto, M. (2014). Phenol-soluble modulins. International Journal of Medical Microbiology, 304, 164–169.
Paetzold, B., Willis, J. R., Pereira de Lima, J., Knödlseder, N., Brüggemann, H., Quist, S. R., Gabaldón, T., & Güell, M. (2019). Skin microbiome modulation induced by probiotic solutions. Microbiome, 7, 95.
Papenfort, K., & Bassler, B. L. (2016). Quorum sensing signal–response systems in Gram-negative bacteria. Nature Reviews Microbiology, 14, 576–588.
Parlet, C. P., Brown, M. M., & Horswill, A. R. (2019). Commensal staphylococci influence Staphylococcus aureus skin colonization and disease. Trends in Microbiology, 27, 497–507.
Periasamy, S., Chatterjee, S. S., Cheung, G. Y., & Otto, M. (2012). Phenol-soluble modulins in staphylococci: what are they originally for? Commun. Integr. Biol., 5, 275–277.
Peschel, A., & Otto, M. (2013). Phenol-soluble modulins and staphylococcal infection. Nature Reviews Microbiology, 11, 667–673.
Porter, G., Joshi, J., Bhullar, L., & Kotwani, A. (2020). Using ‘smart regulation’ to tackle antimicrobial resistance in low-income and middle-income countries. BMJ Global Health, 5, e001864.
Puebla-Barragan, S., & Reid, G. (2021). Probiotics in cosmetic and personal care products: trends and challenges. Molecules, 26, 1249.
Queck, S. Y., Jameson-Lee, M., Villaruz, A. E., Bach, T. H., Khan, B. A., Sturdevant, D. E., Ricklefs, S. M., Li, M., & Otto, M. (2008). RNAIII-independent target gene control by the agr quorum-sensing system: insight into the evolution of virulence regulation in Staphylococcus aureus. Molecular Cell, 32, 150–158.
Rademacher, F., Gläser, R., & Harder, J. (2021). Antimicrobial peptides and proteins: interaction with the skin microbiota. Experimental Dermatology, 30, 1496–1508.
Ramsey, M. M., Freire, M. O., Gabrilska, R. A., Rumbaugh, K. P., & Lemon, K. P. (2016). Staphylococcus aureus shifts toward commensalism in response to Corynebacterium Species. Frontiers in Microbiology, 7, 1230.
Rémy, B., Mion, S., Plener, L., Elias, M., Chabrière, E., & Daudé, D. (2018). Interference in bacterial quorum sensing: a biopharmaceutical perspective. Frontiers in Pharmacology, 9, 203.
Reygagne, P., Bastien, P., Couavoux, M. P., Philippe, D., Renouf, M., Castiel-Higounenc, I., & Gueniche, A. (2017). The positive benefit of Lactobacillus paracasei NCC2461 ST11 in healthy volunteers with moderate to severe dandruff. Benef. Microbes, 8, 671–680.
Riley, M. A., & Chavan, M. A. (2007). Bacteriocins. Berlin, Heidelbarg, Germany: Springer.
Roy, R., Tiwari, M., Donelli, G., & Tiwari, V. (2018). Strategies for combating bacterial biofilms: a focus on anti-biofilm agents and their mechanisms of action. Virulence, 9, 522–554.
Saheb Kashaf, S., Proctor, D. M., Deming, C., Saary, P., Hölzer, M., Comparative Sequencing Program, N. I. S. C., Taylor, M. E., Kong, H. H., Segre, J. A., Almeida, A., et al. (2022). Integrating cultivation and metagenomics for a multi-kingdom view of skin microbiome diversity and functions. Nature Microbiology, 7, 169–179.
Saising, J., Dube, L., Ziebandt, A. K., Voravuthikunchai, S. P., Nega, M., & Götz, F. (2012). Activity of gallidermin on Staphylococcus aureus and Staphylococcus epidermidis biofilms. Antimicrobial Agents and Chemotherapy, 56, 5804–5810.
Salminen, S., Collado, M. C., Endo, A., Hill, C., Lebeer, S., Quigley, E. M. M., Sanders, M. E., Shamir, R., Swann, J. R., Szajewska, H., & Vinderola, G. (2021). The international scientific association of probiotics and prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nature Reviews. Gastroenterology & Hepatology, 18, 649–667.
Sanford, J. A., & Gallo, R. L. (2013). Functions of the skin microbiota in health and disease. Seminars in Immunology, 25, 370–377.
Sang, Y., & Blecha, F. (2008). Antimicrobial peptides and bacteriocins: alternatives to traditional antibiotics. Animal Health Research Reviews, 9, 227–235.
Schommer, N. N., & Gallo, R. L. (2013). Structure and function of the human skin microbiome. Trends in Microbiology, 21, 660–668.
Schuster, M., Sexton, D. J., Diggle, S. P., & Greenberg, E. P. (2013). Acyl-homoserine lactone quorum sensing: From evolution to application. Annual Review of Microbiology, 67, 43–63.
Sfriso, R., Egert, M., Gempeler, M., Voegeli, R., & Campiche, R. (2020). Revealing the secret life of skin-with the microbiome you never walk alone. International Journal of Cosmetic Science, 42, 116–126.
Simons, A., Alhanout, K., & Duval, R. E. (2020). Bacteriocins, antimicrobial peptides from bacterial origin: overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms, 8, 639.
Stubbendieck, R. M., & Straight, P. D. (2016). Multifaceted interfaces of bacterial competition. Journal of Bacteriology, 198, 2145–2155.
Tan, L., Li, S. R., Jiang, B., Hu, X. M., & Li, S. (2018). Therapeutic targeting of the Staphylococcus aureus accessory gene regulator (agr) system. Frontiers in Microbiology, 9, 55.
van Rijen, M., Bonten, M., Wenzel, R., & Kluytmans, J. (2008). Mupirocin ointment for preventing Staphylococcus aureus infections in nasal carriers. The Cochrane Database of Systematic Reviews, 2008, CD006216.
Vuong, C., Saenz, H. L., Götz, F., & Otto, M. (2000). Impact of the agr quorum-sensing system on adherence to polystyrene in Staphylococcus aureus. Journal of Infectious Diseases, 182, 1688–1693.
Wang, R., Braughton, K. R., Kretschmer, D., Bach, T. H. L., Queck, S. Y., Li, M., Kennedy, A. D., Dorward, D. W., Klebanoff, S. J., Peschel, A., et al. (2007). Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nature Medicine, 13, 1510–1514.
Werner, A. H., & Russell, A. D. (1999). Mupirocin, fusidic acid and bacitracin: activity, action and clinical uses of three topical antibiotics. Veterinary Dermatology, 10, 225–240.
Williams, M. R., Costa, S. K., Zaramela, L. S., Khalil, S., Todd, D. A., Winter, H. L., Sanford, J. A., O’Neill, A. M., Liggins, M. C., Nakatsuji, T., et al. (2019). Quorum sensing between bacterial species on the skin protects against epidermal injury in atopic dermatitis. Science Translational Medicine, 11, eaat8329.
Williams, M. R., & Gallo, R. L. (2015). The role of the skin microbiome in atopic dermatitis. Current Allergy and Asthma Reports, 15, 65.
Wollenberg, M. S., Claesen, J., Escapa, I. F., Aldridge, K. L., Fischbach, M. A., Lemon, K. P., & Kolter, R. (2014). Propionibacterium-produced coproporphyrin III induces Staphylococcus aureus aggregation and biofilm formation. mBio, 5, e01286-14.
Yarwood, J. M., Bartels, D. J., Volper, E. M., & Greenberg, E. P. (2004). Quorum sensing in Staphylococcus aureus biofilms. Journal of Bacteriology, 186, 1838–1850.
Zaman, M., & Andreasen, M. (2020). Cross-talk between individual phenol-soluble modulins in Staphylococcus aureus biofilm enables rapid and efficient amyloid formation. eLife, 9, e59776.
Acknowledgements
This research was supported by the Chung-Ang University Graduate Research Scholarship in 2020 (to S.M.L).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors have no conflict of interest to report.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, S.M., Keum, H.L. & Sul, W.J. Bacterial Crosstalk via Antimicrobial Peptides on the Human Skin: Therapeutics from a Sustainable Perspective. J Microbiol. 61, 1–11 (2023). https://doi.org/10.1007/s12275-022-00002-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-022-00002-8