Abstract
A thermotolerant bacterial strain 1D isolated from refinery oil-contaminated soil was identified as Gordonia sp. based on the analysis of 16S rRNA and gyrB gene sequences. The strain was found to utilize crude oil, diesel fuel, and a wide spectrum of alkanes at temperatures up to 50 °C. Strain 1D is the first representative of Gordonia amicalis capable of utilizing alkanes of chain length up to С36 at a temperature of 45–50 °C. The degree of crude oil degradation by Gordonia sp. 1D at 45 °C was 38% in liquid medium and 40% in soil (with regard to abiotic loss). There are no examples of so effective hydrocarbon-oxidizing thermotolerant Gordonia in the world literature. The 1D genome analysis revealed the presence of two alkane hydroxylase gene clusters, genes of dibenzothiophene cleavage, and the cleavage of salicylate and gentisate – naphthalene metabolism intermediates. The highly efficient thermotolerant strain Gordonia sp. 1D can be used in remediation of oil-contaminated soils in hot climates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oil spills and refinery accidents are the main causes of hydrocarbon emission into the environment. Hydrocarbon pollution poses a serious man-made threat for the modern world. Microbial degradation is considered as one of the most effective and environmentally friendly approaches to removing hydrocarbons from contaminated ecosystems.
Gordoniae encompass a rich reservoir of metabolic diversity, which makes them attractive candidates for crude oil bioremediation (Arenskötter et al. 2004). The presence of the dibenzothiophene (DBT) desulfurization pathway in Gordonia amicalis DSM44461T is consistent with their emerging status as a source of metabolic diversity rivaling that of rhodococci (Kim et al. 2000). Gordonia strains utilize aliphatic (Shen et al. 2010; Lo Piccolo et al. 2011) and aromatic (Lin et al. 2012) hydrocarbons and heterocyclic compounds (Santos et al. 2005).
However, the data about the ability of Gordoniae to degrade hydrocarbons at elevated temperatures are very limited (Young et al. 2005; Lin et al. 2008). Thus, it is of great importance to isolate and characterize novel effective Gordonia strains capable of oil degradation at high temperatures. Such strains can be effective agents of remediation of oil-contaminated soils at hot climates with temperatures up to 50 °C. In addition, of importance is further research into the mechanism of alkane degradation by Gordonia strains based on genomics because they are some of the most frequently reported bacterial strains for alkane degradation.
In this study, we describe the isolation and characterization of Gordonia sp. strain 1D capable of vigorous utilization of crude oil and a wide range of alkanes. The peculiar properties of the degradation of main crude oil fractions during the growth of strain 1D at different temperatures were also investigated. In addition, we made the whole genome sequencing and assembly to analyze clusters of genes responsible for alkane degradation.
Materials and methods
Chemicals and media
Hydrocarbons (phenanthrene, anthracene, fluorene, octane, nonane, decane, hexadecane, heptamethyl nonane) of high-grade purity (> 98%) (Sigma–Aldrich, USA; Merck and Fluka, Germany) were used. Crude oil used as a carbon source was characterized by concentration of 0.868 g/mL and the following contents: water, 0.06%; salts, 45 mg/mL; mechanical impurities, 0.008%; and sulfur, 1.42%.
Lysogeny broth (LB) liquid medium consisted of (per liter of distilled water) 10 g of tryptone, 5.0 g of yeast extract, and 5.0 g of sodium chloride. Evans mineral medium contained the following components (per L): 8.71 g K2HPO4, 1 mL 5 mol solution of NH4Cl, 1 mL 0.1 mol solution of Na2SO4, 1 mL 62 mmol solution of MgCl2, 1 mL 1 mmol solution of CaCl2, 1 mL 0.005 mmol solution of (NH4)6Mo7O24·4H2O; trace elements, 1 mL; pH 7.0. The trace element solution consisted of the following components (in 1% HCl, g/L): ZnO, 0.41; FeCl2·6H2O, 5.4; MnCl2·4H2O, 2.00; CuCl2·2H2O, 0.17; CoCl2·6H2O, 0.48; H3BO3, 0.06.
Isolation and identification of crude oil-degrading microorganism
A crude oil-contaminated soil sample was obtained from the territory of a petroleum refinery plant in the Moscow Region (Russia). To enrich petroleum-degrading bacteria, the sample was cultivated in Erlenmeyer flasks with 100 mL of Evans medium with crude oil (2% v/v) for 3 weeks. The cultivation temperature was 30 °C. After that, the sample of enrichment culture was transferred to a flask with the same media and substrate and cultivated for 2 weeks at a temperature of 45 °C. That technique allowed us to obtain thermotolerant strains able to grow at medium as well as elevated temperatures.
To obtain single bacterial colonies, a series of standard tenfold dilutions of enrichment culture was carried out. Then, 100 μL from the last dilution was plated on a petri dish with agarized Evans medium supplemented with 2% of crude oil as a sole carbon and energy source. Single colonies were thrice re-plated by dashing, using inoculation loop, on the same medium to obtain pure cultures (Miller 1972).
Pure cultures were phylogenetically characterized using 16S rRNA and gyrB gene sequencing. Universal primers 27f/1492r (Eden et al. 1991) were used for PCR amplification of the 16S rRNA genes. The PCR thermal cycling consisted of an initial denaturation at 95 °C for 10 min, followed by 35 cycles at 94 °C for 45 s, 56 °C for 45 s, and 72 °C for 90 s, plus the final step at 72 °C for 10 min. Amplification was performed with a GeneAmp PCR System 2400. For PCR amplification of a gyrB gene fragment, we used the primer pair UP1/UP2r (Yamamoto and Harayama 1995). The conditions of PCR cycling were as follows: a total of 30 cycles of amplification was performed with template DNA denaturation at 94 °C for 1 min, primer annealing at 60 °C for 1 min, and primer extension at 72 °C for 2 min.
All amplification products were checked by electrophoresis on 1% agarose gels stained with ethidium bromide. The products were sequenced with an Applied Biosystems 3130 × 1 sequencer (USA) and a BigDye v.3.1 sequencing kit. The obtained sequences were subjected to BLAST homology search (http://www.ncbi.nlm.nih.gov/BLAST). We used the Clustal algorithm for sequence alignment and MEGA7 software for phylogenetic tree building (Tamura et al. 2013). All the sequences of type strains to compare with the sequences of our strains were taken from the Genbank NCBI database.
Hydrocarbon utilization experiments
To examine the ability of strain Gordonia sp. 1D to degrade various hydrocarbons, it was cultivated in 10 mL of test tubes with 5 mL of liquid Evans medium and the substrates at an initial concentration of 2% (w/v or v/v) (100 μL of liquid hydrocarbons or 100 mg solid hydrocarbons per tube). As it is impossible to assess the microbial growth on hydrocarbons by measuring the increase of biomass by absorbance (A 600), we assessed the growth visually and classified the ability of the strain to grow on different substrates in three levels: vigorous, normal growth, and no growth.
Evaluation of the efficiency of crude oil utilization by strain 1D under various conditions
Thermotolerant strain Gordonia sp. 1D was cultivated in flasks with 50 mL of Evans medium and crude oil (2%) for 14 days at 24 and 45 °C. The total content of oil hydrocarbons was measured by IR spectrometry. After the cultivation, the residual oil was extracted with carbon tetrachloride (1:1). The hydrocarbon content in the extract was determined with an AN-2 oil product analyzer (Russia). Sterile Evans medium with 2% oil was used as a control.
The crude oil concentration in liquid samples was calculated by the formula:
where С is the crude oil concentration in the eluate found by instrument’s readings or the calibration dependence, mg/L; VI is eluate volume, L; V is the water sample volume, L; and η is the degree of eluate dilution. In the case of no dilution, η = 1.
To study the process of crude oil destruction in soil (sand), we constructed a model system with the following contents: sterile sand with sea salt (final concentration, 3%), crude oil (2%), sterile liquid mineral medium (10%), and strain Gordonia sp. 1D to a final concentration of 1 × 104 CFU/g. The sand was loosened every 3–4 days. The experiment was carried out both at 24 and at 45 °C. After the experiment, the residual concentration of hydrocarbons was measured by IR spectroscopy on an AN-2 oil analyzer (Russia).
The crude oil concentration in soil samples was calculated by the formula:
where D is the proportionality coefficient obtained by processing the calibration dependence based on actual and measured oil concentrations in the soil (established for this type of soil); K is the coefficient of eluate dilution by CCl4, mL/mL; \( {C}_{H_k} \) is the crude oil concentration in the diluted eluate determined based on the calibration dependence, mg/L; \( {C}_{H_0} \) is the concentration of non-specific humus components and residual oil products that can occur in the soil taken as a control, mg/L; V0 is the initial volume of CCl4 taken for crude oil extraction from a soil sample, L; and P is the soil sample, kg.
Genome sequencing, assembly, and annotation
High molecular mass genomic DNA was isolated from Gordonia sp. 1D using a DNA isolation kit (QIAGEN DNeasy Blood and Tissue Kit, Cat No/ID: 69506).
An Illumina Nextera XT reagent kit (FC-131-1024) was used to prepare a DNA library. Sequencing was carried out using the Illumina MiSeq platform with an MiSeq Reagent Kit v3 (600-cycle), MS-102-3003.
We obtained 1,143,191 paired-end reads of < 300 bp in size. Adapter and low-quality sequences (Q < 20) were deleted using Trimmomatic-0.36 software (Bolger et al. 2014). The remaining sequences were assembled to contigs using SPAdes-3.10.1 software (Bankevich et al. 2012). The number of contigs obtained was 191 with average k-mer coverage (k = 127) of 26.5659. The contigs were filtered using BLASTN and only 176 contigs were retained for further assembly.
Initial genome annotation of Gordonia sp. strain 1D was carried out using Prokka software (Seemann 2014). Additional gene prediction and functional annotation analyses were performed using RAST (Aziz et al. 2008) with k-mer size of 12 and Tigr Glimmer with minimum gene length of 300 (Delcher et al. 2007). Functions of some genes were checked manually by BLAST.
The circular map of the chromosome was made with DNAPlotter (Carver et al. 2009). Alkane hydroxylase cluster maps were performed using Artemis software (Carver et al. 2012).
Results and discussion
Isolation and identification of thermotolerant alkane-degrading strains
Several thermotolerant bacterial strains were isolated from refinery oil-contaminated soil (Moscow, Russia) after enrichment cultivation in Evans medium with crude oil for 3 weeks (Delegan et al. 2016). Gram-positive strain named 1D and capable of highly efficient degradation of crude oil at a temperature up to 50 °C was selected for further study. Colonies of the strain were orange-colored; cells were aerobic, non-motile, and featured short rods. The phylogenetic analysis of 16S rRNA and gyrB gene fragments showed the strain to be the closest to G. amicalis (99% similarity of the 16S rRNA gene, 94% similarity of the gyrB gene), but the similarity percentage of the marker genes, especially gyrase-encoding, was too low to unambiguously attribute the strain to G. amicalis. Based on the results of the morphological and genetic analysis, strain 1D was identified as Gordonia sp. (Fig. 1). The strain was deposited at the All-Russian Collection of Microorganisms as VKM Ac-2120 D.
Phylogenetic tree derived from a 16S rRNA and bgyrB gene sequences and the sequences of the closest neighbor type strains. The distances were calculated using the neighbor-joining method. Numbers at branch points are bootstrap values based on 1000 samplings. Dietzia kunjamensis type strain (FJ438561.1) was used as the outgroup
Gordoniae are known for their ability to utilize crude oil and its individual components (alkanes, polycyclic aromatic compounds), including sulfur-containing components (DBT). However, there are little data about the metabolic peculiarities of Gordonia at elevated temperatures. Young et al. (2005) isolated strain G. alkanivorans CC-JG39 from oil-contaminated dryer sludge of a gas station in Taiwan. The strain grew on media with diesel fuel at temperatures up to 45 °C.
Analysis of the growth kinetics and temperature dependence of the specific growth rate of Gordonia sp. strain 1D within the range of 20–56 °C showed a maximum at 35–37 °C; at the same time, the maximum cell number was observed at the beginning of the stationary phase of culture growth. This temperature was optimal for strain growth. The maximal specific growth rate of strain 1D at 45 °C was 0.33 1/h (0.39 1/h at 35 °C) (Fig. 2).
The results imply that the thermotolerant representatives of the genus Gordonia may have a great scientific significance due to physiological and possible genetic differences from other Gordonia species that are mainly mesophilic.
Substrate spectrum of Gordonia sp. strain 1D
The degradation of crude oil, diesel fuel, alkanes of different chain length, and polycyclic aromatic hydrocarbons (PAHs) with two or three rings was measured. We analyzed the growth of bacteria at temperatures of 24 and 45 °C and found no differences between their hydrocarbon-oxidizing capacities at different temperatures in a good agreement with our preliminary results (Delegan et al. 2016) (Table 1).
We established that Gordonia sp. 1D could not degrade any of the tested PAHs. Amplification with specific primers also showed that there were no genes of PAH catabolism in the strain. Meanwhile, the strain was a vigorous alkane degrader; it utilized alkanes with a wide range of chain length.
The results of the fractional analysis showed that strain 1D was capable of utilizing a wide spectrum of odd and even alkanes from C8 to C36, including branched ones (pristane, phytane). There are no examples of such effective hydrocarbon-oxidizing thermotolerant Gordonia species in the world literature. The strain utilized petroleum alkanes at temperatures up to 50 °C.
Efficiency of crude oil utilization by Gordonia sp. 1D under different conditions
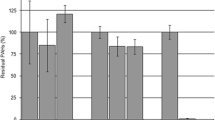
The results of our investigations have shown that Gordonia representatives can actively degrade crude oil and a wide spectrum of hydrocarbons not only under moderate conditions. To evaluate the efficiency of crude oil degradation by strain 1D, it was grown both in Evans medium with crude oil and introduced into oil-contaminated soil at a temperature 45–50 °C. After the cultivation, the residual hydrocarbon content was measured and the degree of crude oil degradation was evaluated (Fig. 3).
This paper is the first report of the peculiarities of crude oil degradation by a strain close to G. amicalis at different temperatures. The full spectrum of alkanes utilized by the thermotolerant Gordonia strain was also investigated for the first time.
Overview of Gordonia sp. 1D genome
The draft genome of Gordonia sp. 1D has a single chromosome and no plasmids (Fig. 4). The size of the genome is 5,151,623 bp with a G + C content of 67.3%. Average gene size is 965 bp, giving a coding density of 89.4%. The genome contains 4772 predicted protein coding sequences (CDS), 47 tRNA genes, and 3 rRNA operons. Average nucleotide identity (ANI) parameter analysis revealed that the closest phylogenetic relatives of strain 1D are Gordonia sp. KTR9 (NC_018581.1) and Gordonia terrae 3612 (CP016594.1) both having the OrthoANIu value of 80.9%. There are no fully assembled genomes of G. amicalis strains in the Genbank NCBI database, so we could compare the genome of 1D only with suitable contigs of G. amicalis strains CCMA-559 and NBRC 100051.
Of the 4772 CDS found, 4091 could be assigned to 23 different categories of clusters of orthologous groups (COGs). These results suggest that the thermotolerant strain 1D possesses efficient lipid, carbohydrate, amino acid transport, and metabolism (Fig. 5).
COG function classification of thermotolerant strain Gordonia sp. 1D. (A) RNA processing and modification. (B) Chromatin structure and dynamics. (C) Energy production and conversion. (D) Cell cycle control, cell division, chromosome partitioning. (E) Amino acid transport and metabolism. (F) Nucleotide transport and metabolism. (G) Carbohydrate transport and metabolism. (H) Coenzyme transport and metabolism. (I) Lipid transport and metabolism. (J) Translation, ribosomal structure, and biogenesis. (K) Transcription. (L) Replication, recombination, and repair, (M) Cell wall/membrane/envelope biogenesis. (N) Cell motility. (O) Posttranslational modification, protein turnover, chaperones. (P) Inorganic ion transport and metabolism. (Q) Secondary metabolites biosynthesis, transport, and catabolism. (R) General function prediction only. (S) Function unknown. (T) Signal transduction mechanisms. (U) Intracellular trafficking, secretion, and vesicular transport. (V) Defense mechanisms. (Z) Cytoskeleton
In the present paper, the thermotolerant strain Gordonia sp. 1D was found to be able to utilize alkanes of different chain length. The analysis of the assembled sequence of the strain confirmed that Gordonia sp. 1D possesses different alkane hydroxylase systems. Moreover, the strain bears the enzymes of haloalkanes and alkanal degradation.
The alkane hydroxylase systems in Gordonia sp. strain 1D
The first group of alkane catabolic genes in strain 1D was represented by a cluster of four genes consisting of an alkane 1-monooxygenase alkB, two rubredoxin units and rubredoxin reductase (Fig. 6). It differed from the same cluster in Gordonia sp. KTR9 by the absence of transposase gene directly after the gene alkB, as shown in strain KTR9.
Organization of the genes coding for the enzymes of AlkB hydroxylase cluster: alkB, alkane 1-monooxygenase, EC 1.14.15.3; rubA, rubA2, rubredoxins; rubR, rubredoxin reductase. The closest neighbors of the cluster are (left) hypothetical protein of 199 aa and (right) transcriptional regulator, TetR family, of 200 aa
The RubR amino acid sequence has a 67% similarity with that of Gordonia sp. KTR9, and the sequence of the gene alkB has an 89% similarity.
The search for genes coding for the enzyme system CYP153 in the genome of strain 1D revealed the presence of ten copies of cytochrome P450 monooxygenase, only one of which was included in a full cluster with 2Fe-2S ferredoxin and ferredoxin reductase. The enzymes encoded by the genes of this cluster probably gave strain 1D the ability to oxidize short (from C8) alkanes. This process was also active at high temperatures. Strain Gordonia sp. KTR9, the closest phylogenetic relative of strain 1D (ANI value 80.9%), does not possess cytochrome P450 monooxygenases. There are also no data about the presence of these genes in other close relatives of strain 1D—G. amicalis CCMA-559 and NBRC 100051. Cloning and expression of these putative alkane hydroxylases would be helpful for the better understanding the mechanism of alkane degradation by Gordonia strains, including that process at high temperatures.
Besides the genes of alkane degradation, the genome of Gordonia sp. 1D contained genes of DBT cleavage, as well as the cleavage of salicylate and gentisate–naphthalene metabolism intermediates. The naphthalene cleavage pathway via salicylate and gentisate is present in most actinobacteria especially in Rhodococci (Di Gennaro et al. 2001). However, naphthalene degradation genes were not found in the genome of strain 1D. In this study, we observed no naphthalene degradation, and it was not surprising that no naphthalene catabolic gene cluster was identified by the genomic analysis of strain 1D. Meanwhile, the metabolic capabilities of Gordonia sp. 1D as a promising hydrocarbon degrader could be enhanced by introducing genetic constructions in the strain. Construction of genetically engineered strains based on 1D can overcome the incomplete degradation of naphthalene during the recultivation process. Further research is planned to test this possibility.
Conclusions
Thermotolerant strain 1D capable of vigorously degrading crude oil, diesel fuel, and alkanes of different chain lengths was isolated from refinery oil-contaminated soil (Moscow, Russia). 16S rRNA and gyrB gene sequencing and analysis have revealed that the strain belongs to the genus Gordonia. The results showed that hydrocarbon degradation occurred actively both at moderate and high (up to 50 °C) temperatures in liquid media and soil. Strain 1D is the first representative of G. amicalis capable of utilizing alkanes of chain length up to С36 at a temperature of 45–50 °C. All the strains of G. amicalis known to date are mesophilic. There are no examples of such effective hydrocarbon-oxidizing thermotolerant Gordonia in the world literature. Information about the genome sequence of Gordonia sp. 1D offered an opportunity to expand the data about the metabolic potential of Gordoniae. Further experiments will be helpful to verify the functions of alkane degradation annotated genes.
References
Arenskötter M, Bröker D, Steinbuchel A (2004) Biology of the metabolically diverse genus Gordonia. Appl Environ Microbiol 70:3195–3204. https://doi.org/10.1128/AEM.70.6.3195-3204.2004
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 8(9):75. https://doi.org/10.1186/1471-2164-9-75
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19(5):455–477. https://doi.org/10.1089/cmb.2012.0021
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Carver T, Thomson N, Bleasby A, Berriman M, Parkhill J (2009) DNAPlotter: circular and linear interactive genome visualization. Bioinformatics 25(1):119–120. https://doi.org/10.1093/bioinformatics/btn578
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA (2012) Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28:464–469. https://doi.org/10.1093/bioinformatics/btr703
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with glimmer. Bioinformatics 23(6):673–679. https://doi.org/10.1093/bioinformatics/btm009
Delegan Y, Vetrova A, Akimov V, Titok M, Filonov A, Boronin M (2016) Thermotolerant oil-degrading bacteria isolated from soil and water of geographically distant regions. Appl Biochem Microbiol 52(4):389–396. https://doi.org/10.1134/S0003683816040025
Di Gennaro P, Rescalli E, Galli E, Sello G, Bestetti G (2001) Characterization of Rhodococcus opacus R7, a strain able to degrade naphthalene and o-xylene isolated from a polycyclic aromatic hydrocarbon-contaminated soil. Res Microbiol 152(7):641–651. https://doi.org/10.1016/S0923-2508(01)01243-8
Eden PA, Schmidt TM, Blakemore RP, Pace NR (1991) Phylogenetic analysis of Aquaspirillum magnetotacticum using polymerase chain reaction-amplified 16S rRNA-specific DNA. Int J Syst Bacterifol 41(2):324–325. https://doi.org/10.1099/00207713-41-2-324
Kim SB, Brown R, Oldfield C, Gilbert SC, Iliarionov S, Goodfellow M (2000) Gordonia amicalis sp. nov., a novel dibenzothiophene-desulphurizing actinomycete. Int J Syst Evol Microbiol 50:2031–2036. https://doi.org/10.1099/00207713-50-6-2031
Lin CL, Shen FT, Tan CC, Huang CC, Chen BY, Arun AB, Young CC (2012) Characterization of Gordonia sp. strain CC-NAPH129-6 capable of naphthalene degradation. Microbiol Res 167:395–404. https://doi.org/10.1016/j.micres.2011.12.002
Lin TC, Chang JS, Young CC (2008) Exopolysaccharides produced by Gordonia alkanivorans enhance bacterial degradation activity for diesel. Biotechnol Lett 30:1201–1206. https://doi.org/10.1007/s10529-008-9667-8
Lo Piccolo L, De Pasquale C, Fodale R, Puglia AM, Quatrini P (2011) Involvement of an alkane hydroxylase system of Gordonia sp. strain SoCg in degradation of solid n-alkanes. Appl Environ Microbiol 77:1204–1213. https://doi.org/10.1128/AEM.02180-10
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory. 466 pages
Santos SCC, Alviano DS, Alviano CS, Pádula M, Leitão AC et al (2005) Characterization of Gordonia sp. strain F.5.25.8 capable of dibenzothiophene desulfurization and carbazole utilization. Appl Microbiol Biotechnol 71:355–362. https://doi.org/10.1007/s00253-005-0154-z
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Shen FT, Young LS, Hsieh MF, Lin SY, Young CC (2010) Molecular detection and phylogenetic analysis of the alkane 1-monooxygenase gene from Gordonia spp. Syst Appl Microbiol 33:53–55. https://doi.org/10.1016/j.syapm.2009.11.003
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Yamamoto S, Harayama S (1995) PCR amplification and direct sequencing of gyrB genes with universal primers and their application to the detection and taxonomic analysis of Pseudomonas putida strains. Appl Environ Microbiol 61:1104–1109
Young CC, Lin TC, Yeh MS, Shen FT, Chang JS (2005) Identification and kinetic characteristics of an indigenous diesel-degrading Gordonia alkanivorans strain. World J Microbiol Biotechnol 21:1409–1414. https://doi.org/10.1007/s11274-005-5742-7
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. The work was financially supported by RFBR (project №18-34-00329_mol_а).
Rights and permissions
About this article
Cite this article
Delegan, Y.A., Valentovich, L.N., Shafieva, S.M. et al. Characterization and genomic analysis of highly efficient thermotolerant oil-degrading bacterium Gordonia sp. 1D. Folia Microbiol 64, 41–48 (2019). https://doi.org/10.1007/s12223-018-0623-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-018-0623-2