Abstract
Powdery mildew (Blumeria graminis f. sp. Tritici, (Bgt)) is an important worldwide fungal foliar disease of wheat (Triticum aestivum) responsible for severe yield losses. The development of resistance genes and dissection of the resistance mechanism will therefore be beneficial in wheat breeding. The Bgt resistance gene PmAS846 was transferred to the hexaploid wheat lines N9134 from Triticum dicoccoides, and it is still one of the most effective resistance genes. Here, by RNA sequencing, we identified three co-expressed gene modules using pairwise comparisons and weighted gene co-expression network analysis during wheat–Bgt interactions compared with mock-infected plants. Hub genes of stress-specific modules were significantly enriched in spliceosomes, phagosomes, the mRNA surveillance pathway, protein processing in the endoplasmic reticulum, and endocytosis. Induced module genes located on chromosome 5BL were selected to construct a protein–protein interaction network. Several proteins were predicted as the key hub node, including Hsp70, DEAD/DEAH box RNA helicase PRH75, elongation factor EF-2, cell division cycle 5, ARF guanine-nucleotide exchange factor GNOM-like, and protein phosphatase 2C 70 protein, which interacted with several disease resistance proteins such as RLP37, RPP13 and RPS2 analogues. Gene ontology enrichment results showed that wheat could activate binding functional genes via an mRNA transcription mechanism in response to Bgt stress. Of these node genes, GNOM-like, PP2C isoform X1 and transmembrane 9 superfamily member 9 were mapped onto the genetic fragment of PmAS846 with a distance of 4.8 Mb. This work provides the foundations for understanding the resistance mechanism and cloning the resistance gene PmAS846.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Powdery mildew, caused by Blumeria graminis f. sp. tritici (Bgt), is an important fungal foliar disease of wheat (Triticum aestivum) that is prevalent in many wheat-growing regions of the world (Dean et al. 2012). Rising trends in yield losses caused by the Bgt virulent races are becoming increasingly serious because of the climate change and crop planting structure adjustments. Breeding for resistance is the most profitable and environmentally acceptable strategy to control damage caused by powdery mildew disease. However, the development of specific resistant varieties lags far behind the frequency of genetic resistance loss. Therefore, there is an urgent need to identify novel powdery mildew resistance genes and to develop durable resistant varieties though converging multispecific resistance genes.
The development of effective genetic resistance for breeding commercial varieties is an increasingly time-consuming practice due to the declining genetic diversity of germplasms in the current agricultural systems. Thus, dissecting disease resistance mechanisms has become a hotspot for research in determining R genes in wheat pathogen defense, although many breeding efforts are still based on nucleotide-binding leucine-rich repeat (NLR)-based disease resistance. During plant–pathogen interactions, plants wield resistance (R) gene-mediated defense mechanisms and mount a mass defensive response to pathogen attack to delay or arrest potential pathogenic microorganism growth (Schwessinger and Ronald 2012; Bouktila et al. 2015). Compared with model plants, the use of classical genetics to isolate R genes has been limited because the hexaploid wheat has a large and complex genome, although the powdery mildew resistance gene (Pm3) has been identified (Hurni et al. 2013). In wheat–Bgt interactions, a number of studies using RNA sequencing (RNA-seq) (Zhang et al. 2014), GeneChip, microRNA analysis (Xin et al. 2010) and proteomics have identified thousands of functional genes and noncoding genes (Xin et al. 2011; Zhang et al. 2016). However, the resistant trait is well known as a monogenic control phenotype in classical genetics, raising the troublesome question of how to select a key R gene from the big data of gene transcription expression profiles. Additionally, the expression of determinative proteins is highly complex because of posttranscriptional, translational, and/or posttranslational regulatory mechanisms (Janke and Bulinski 2011), alternative splicing (Reddy et al. 2013) and protein degradation; as a result, the model of wheat resistance activation in responding to powdery mildew is still unclear. Moreover, in contrast to its importance in breeding, research into the genetic and molecular basis of R is still in its infancy.
Isobaric tags for relative and absolute quantification proteomics technology is a key tool that contributes to the explanation of complex biological processes at the protein level. Recently, we analysed changes in the protein profiles of resistant wheat in response to powdery mildew (Bgt) infection, and identified some stress-related and defense-related protein species (Fu et al. 2016). However, this was achieved by covering a wide range of potential defense-related proteins which was not sufficiently narrow to recognize critical genes involved in defense mechanism of wheat undergoing Bgt infection. Weighted gene correlation network analysis (WGCNA) implements methods that are conducive in generating testable hypotheses for validation in independent datasets, such as putative pathways associated with developing receptacles (Hollender et al. 2014) and disease or defense outcomes (Rasmussen et al. 2013). Previously, we used WGCNA and transcriptome–proteome-associated analysis to construct a model of gene activation in the wheat defense response to stripe rust (Zhang et al. 2019a), and narrowed the number of key resistance genes to three dozen. Unfortunately, large-scale transcriptome comparison results revealed the wheat response to stripe rust and powdery mildew stress through distinct gene activations (Zhang et al. 2014). The wheat line N9134 had maintained a high level of resistance to powdery mildew because of the resistance genes in the long arm of chromosome 5B (Xue et al. 2012), with the exception of the stripe rust resistance gene in chromosome 1BS.
In the present study, we focussed on the networks involved in the wheat–Bgt interaction by merging data from RNA-seq and WGCNA, and then used the genetic fragment analysis to further narrow the key resistance genes. The main objective of this study was to identify a spectrum model of wheat resistance activation in response to powdery mildew by identifying changes in expression patterns after inoculation with Bgt. WGCNA elucidated the higher-order relationships between genes based on their co-expression relationships, delineated modules of biologically related genes, and permitted a robust view of transcriptome organization in the response of the resistant wheat line N9134 to Bgt E09. The most highly connected or central genes, referred to as ‘hubs’ were further employed to construct protein–protein interaction (PPI) networks. Finally, taking the classical genetics resistance gene locus into consideration, we predicted the possible involvement of the candidate gene PmAS846.
Materials and methods
Plant materials and pathogen stress treatment
The winter wheat line N9134, developed by Northwest A&F University (Yangling, China) shows high immune resistance to all Bgt races in China. This resistance was conferred by one all-stage resistance gene located on chromosome 5BL bin 0.75–0.76, named PmAs846 (Xue et al. 2012). A pair of contrasting BC7F2 homozygous lines were developed form backcross SY225/7 × PmAS846, which differ only regarding PmAS846 on 5BL. The Bgt isolate E09 was maintained by the College of Agronomy.
Weighted gene correlation network analysis
To carry out the co-expression network analysis on the response of resistant wheat line N9134 to powdery mildew, we separated RNA-seq data of nine Bgt-infected samples from PRJNA243835 (Zhang et al. 2014), which included 21 different leaf samples at 0, 24, 48 and 72 h postinfection (hpi) after fungal infection. Briefly, after the total RNA was extracted and modified with DNase digestion, Oligo(dT)-magnetic beads were used to enrich the mRNA, which was then broken into fragments. Following cDNA preparation with random hexamers, adaptor sequences were ligated to the ends of the repaired double-stranded cDNA after purification. Finally, EST libraries were constructed by PCR amplification and sequenced with an Illumina HiSeq 2000 platform. Here, all reliable readings were assembled using TopHat2 and Cufflinks software (Trapnell et al. 2012; Kim et al. 2013) to reconstruct the gene libraries from reference wheat (Chinese spring) genome sequences from Unité de Recherche Génomique Info (v. v2.2) (Brenchley et al. 2012) after cleaning low-quality reads and screening ribosomal RNA with bowtie. Fragments per kilobase of exon model per million mapped reads (FPKM) values were used to examine the gene expression levels in each sample. With a fold change of ≥2 and a false discovery rate (FDR) ≤0.05, differentially expressed genes (DEGs) were selected with DESeq software among the three treatment groups compared with mock-inoculated leaves. Principal component analysis and Pearson’s correlation of FPKM were used to test the repeatability of samples. Co-expression networks were constructed using the WGCNA package (v1.47) in R (Langfelder and Horvath 2008). Among all 75,906 assembled unigenes, 30,828 genes with FPKM >5 were used for the WGCNA unsigned co-expression network analysis. The modules were obtained using the automatic network construction function on large expression datasets, blockwiseModules with default settings, except that power was 10, the similarity degree was 0.75, minModuleSize was 30, and mergeCutHeight was 0.3. The eigengene value was calculated for each module and used to test the association with each time point sample. The total connectivity and intramodular connectivity, kME (for modular membership, i.e. eigengene-based connectivity), and kME-P value were calculated as previously described (Zhang et al. 2019a). The expression trends of DEGs were classified using short time-series expression miner (STEM; http://www.cs.cmu.edu/~jernst/stem) with log-transformed normalized data (Ernst and Bar-Joseph 2006).
PPI network construction
PPI networks were constructed and visualized using STRING v10.5 to analyse DEG-encoding proteins identified in the main modules (Szklarczyk et al. 2015).
Informatics enrichment and k-means clustering analysis
To organize the genes into hierarchical categories, the DEGs were mapped to gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways using the MAS molecular function annotation system (http://mas.capitalbiotech.com/mas3/). GO terms and KEGG pathways with FDR-corrected P values <0.05 and Q-values ≤0.05 were considered statistically significant. Gene expression pattern analysis was used to cluster genes showing similar expression trends at 0, 24, 48 and 72 hpi. To examine the DEG expression patterns in each sample, the expression data were normalized to 0, log2 (v1/v0), log2 (v2/v0), and log2 (v3/v0), respectively. Clusters were generated using STEM with previously described standard parameters (Zhang et al. 2019a). P ≤0.05 was also set as the threshold for statistical significance for the profiles generated.
Quantitative real-time PCR analysis
SYBR Green-based quantitative PCR system was used for qRT-PCR analysis. The template cDNAs were prepared from the infected contrasting wheat NILs samples, which were collected at 6, 12, 24, 36, 48, 72 and 96 hpi with Bgt E09. The uninoculated plant samples at the same time points were set as the controls. Three independent biological replicates were carried out for each time point. Standard protocol was used to quantify relative gene expression levels with specific primers (table 1 in electronic supplementary material at http://www.ias.ac.in/jgenet) and TaActin (endogenous reference). The qRT-PCR was completed with the QuantStudio 7 Flex real-time PCR system.
Results
Co-expression network analysis of wheat responding to powdery mildew
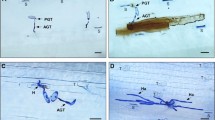
For Bgt-infected samples, the average number of high quality clean reads per library was around 37.42 million 101 bp paired-end reads, compared to 35.99 million clean reads per library constructed from Pst-infected samples. Based on the bread wheat reference transcripts, 66727, 67280, 67764 and 67799 genes were assembled at 0, 24, 48 and 72 hpi, respectively. After the co-expression networks were constructed on the basis of pairwise correlations between genes in their common expression trends across all sampled leaves, 18 main modules with mergeCutHeight 0.3 were further classified from 79 distinct modules, as shown previously (Zhang et al. 2019a). Taking replication into consideration, three of the 18 co-expression modules, including black, darkolivegreen4, and plum1 were shown to be comprised of genes that are highly specifically expressed in Bgt test at different time points (r > 0.8, P < 10−3) (figure 1, a–d). Here, we identified 2684 Bgt-induced specific genes at 24 hpi classifying into the black module, 4285 specific genes in the darkolivegreen4 module at 24, 48 and 72 hpi, and 11601 genes in the plum1 module at 48 and 72 hpi. Among the three modules, plum1 comprised the most sample-specific expressed genes, which were mainly enriched in spliceosomes, peroxisomes, the mRNA surveillance pathway, RNA transport and degradation, and valine, leucine, and isoleucine degradation (table 1; figure 1 in electronic supplementary material). Genes in the black module were enriched in phagosomes, biosynthesis/metabolism of amino acids, protein processing in the endoplasmic reticulum, and protein export, while the darkolivegreen4 module consisted of genes enriched in proteasomes, ribosomes, phagosomes, endocytosis, N-glycan biosynthesis, valine, leucine, and isoleucine degradation, sphingolipid metabolism, SNARE interactions in vesicular transport, arginine biosynthesis, ascorbate and aldarate metabolism, and the citrate cycle.
Developing the Bgt-induced specific gene module. (a) Correlation heat map between modules. Heat maps and eigengene expression profiles for specific modules: (b) black, (c) darkolivegreen4, and (d) plum1 in inoculated leaves of N9134. Sample names are given at the top of each panel. Heat maps show the relative normalized RPKM of each gene. The y-axis indicates the value of the module eigengene; the x-axis indicates the sample type.
Trends analysis was carried out using the earlier K-means clustering method (figure 2 in electronic supplementary material). In the significant profile 16, the gene expression pattern was upregulated at 48 and 72 hpi but not at 24 hpi, and was enriched in spliceosomes, RNA transport, and the mRNA surveillance pathway (table 2). Figure 2 in electronic supplementary material shows that the gene expression pattern was steeply upregulated at 24 hpi, then returned to normal levels as a control in the significant profile 18. These genes were enriched in the biosynthesis of amino acids, protein processing in the endoplasmic reticulum, carbon metabolism, and the cysteine and methionine metabolism pathway as shown in table 2. This indicates that the wheat expresses genes to modulate transcripts underlying the stress signal and dysregulation of amino acid metabolism.
PPI network construction for DEGs induced by Bgt stress
Although WGCNA analysis narrowed the range of genes responding to Bgt infection, the priming or core control genes/locus that trigger other defense-related and downstream genes was still difficult to determine. The main reason is that a typical plant genome contains hundreds of NLR-encoding genes distributed in each chromosome, of which many reside in complex clusters. Adversely, in classical genetics, most resistance genes are well known as a monogenic control phenotype. Considering that the powdery mildew resistance gene, a dominant genetic gene locus in N9134, is located on the short arm of chromosome 5BL (Xue et al. 2012), we further focussed on candidate genes on 5BL. A total of 780 hub genes located on chromosome 5BL were upregulated at the transcript and/or protein level during Bgt stress. Table 2 in electronic supplementary material shows that the first quarter of these genes function in pathways involved in genetic information processing and regulation, plant–pathogen interaction, and oxidative phosphorylation. They include heat stress transcription factors, ribosomal protein S2, probable ubiquitin receptor RAD23, and disease resistance proteins RPS2, RPM1 and RPP13. Some genes involved in the peroxisome pathway were also predicted as candidate genes, such as peroxisomal-2-hydroxy-acid oxidase GLO1, pathogenesis-related protein and subtilisin-like protease. Disease resistance protein RPS2 and DnaJ protein-like protein gene were detected with higher connectivity than other hub genes, reaching 2881.7 and 2676.0, respectively.
To further investigate interactions among Bgt stress-induced DEGs, the top 50% of high connectivity DEGs predicatively clustered into plum1, darkolivegreen4 and black modules were integrated with information from the STRING database to construct a PPI network. Five interaction networks were predicted from 114 nodes proteins with enrichment P values < 1.03E-16 (figure 2; table 3 in electronic supplementary material provided the supporting information). Hsp70 was accompanied by DEAD/DEAH box RNA helicase PRH75, elongation factor EF-2 (LOS), cell division cycle 5, ARF guanine-nucleotide exchange factor GNOM-like, cleavage/polyadenylation specificity factor CPCF, AT4G25550-like protein, PCF11P-similar protein, and protein phosphatase 2C 70 protein (KAPP) in the powerful PPI network, which contained and interacted with several receptor kinases such as cell wall-associated kinase (WAK), disease resistance protein RPM1, RLP37, RPP13, RPS2 (At4g26090), and AT5G46520 homologue. Figure 3 shows the interaction between zinc-finger CCCH domain-containing protein 16 with calmodulin-binding transcription activator (CAMTA)2 (AT5G64220), CAMTA3 and cyclic nucleotide-gated ion channel 2 (DND1). Moreover, WPP domain protein 2 interacted with Hsp70, the cysteine-rich receptor-like protein kinases (CRKs) interacted with each other, and we predicted an interaction between both of two CRKs with PP2C.
Interaction networks of related DEGs in plum1, darkolivegreen4, and black modules identified by WGCNA analysis. Protein–protein interaction networks were constituted by String software in the Arabidopsis experiment or text mining databases. Purple lines represent experimental evidence, green lines represent the gene neighbourhood, blue lines represent gene co-occurrence database evidence, yellow lines represent text mining evidence, black lines represent co-expression evidence. Hsp70, heat shock protein 70; PRH75, DEAD/DEAH box RNA helicase, LOS1; elongation factor EF-2; CDC5, cell division cycle 5; GN, ARF guanine-nucleotide exchange factor GNOM-like; KAPP, protein phosphatase 2C 70 protein; RABA2c, RAB GTPase homolog A2C; CPSF100, cleavage and polyadenylation specificity factor 100; At4g25550, cleavage/polyadenylation specificity factor CPSF6; PCFS, PCF11P-similar protein 4; GR-RBP2, glycine-rich RNA-binding protein 2; At1g75340, zinc finger CCCH domain-containing protein 16; RLP37, receptor like protein 37; RPL3B, R-protein L3 B; Y14, RNA-binding protein 8A. A description of the other nodes is given in table 1 in electronic supplementary material.
GO enrichment results showed that the PPI network was significantly enriched in binding and catalytic activity for the molecular function domain as shown in figure 3; table 4 in electronic supplementary material, including ion binding, protein binding, ATP and ADP binding, and nucleotide binding. For the cellular component domain, pathways involving the intracellular membrane, plasma membrane and organelle membrane were remarkably enriched in GO analysis, while the organic substance metabolic process, biological regulation, and response to stimulus were significantly enriched in the biological process domain. KEGG pathways were enriched in mRNA surveillance and RNA transport, suggesting that wheat activates the binding of functional genes via mRNA transcription mechanisms to respond to Bgt pathogen stress.
Prediction of the powdery mildew resistance gene PmAS846 as a candidate gene
Previous work showed that PmAs846 was cosegregated with AL819406, CJ694617 and CJ540214, which were flanked by expressed sequence tag (EST) markers BJ261635 and XFCP620 with a genetic distance of 2.3 cM (Xue et al. 2012). However, after we mapped all predicted gene and EST marker sequences onto the reference genome of Chinese Spring, the candidate gene PmAs846 was predicted to be located in the physical region from 541,341,767 to 546,569,512 bp in chromosome 5B of Chinese Spring. We further mapped all node genes onto the Chinese Spring reference genome. Most of the aforementioned hub genes are annotated outside the region, although many similar proteins can be found here, such as cyclic nucleotide-gated ion channel (CNGC)17, STRUBBELIG-receptor SRF8, and wall-associated receptor kinase. However, ARF guanine-nucleotide exchange factor GNOM-like gene, protein phosphatase 2C 70 isoform X1 (PP2C), and transmembrane 9 superfamily member 9 gene (TM9SF9) were located in this region, which were named GN, KAPP and At5G25100 in the network, respectively. Considering that the limitation of the protein in PPI database, we mapped the hub genes with higher connectivity of WGCNA module as well. Several hub genes of WGCNA were similarly mapped into this region, such as mediator of RNA polymerase II transcription subunit 8 (MED8) isoform X2, NADPH-cytochrome P450 reductase (CPR)-like and synaptic vesicle 2 related. Thus, these genes are possible candidate genes of PmAS846.
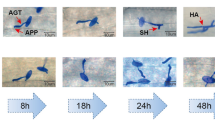
To further evaluate the possibility of candidate genes, we analysed three gene expression pattern comparing BC7F2 resistance with susceptible line, which differ only regarding PmAS846 on chromosome 5BL. The expression profiles of GNEF-5B, MED8X2-5B and CPRL-5B in the inoculated resistant and susceptible plants are presented in figure 4. Following the inoculation with Bgt, the accumulation of three tested transcripts were 3.2, 5.1 and 2.6 times upregulated in resistant plants at 36 hpi, respectively. The expression levels of GNEF-5B was 2.4 times upregulated in susceptible plants at 24 dpi, while MED8X2 was 2.6 times dysregulated at 36 and 72 hpi. Generally, the transcription levels of GNEF, MED8X2 and CPRL were distinctly lower in susceptible line than that in resistant line. This demonstrated that the expression of GNEF-5B, MED8X2-5B, and CPRL-5B homolog were induced and implicated in resistance to powdery mildew.
Gene expression patterns of GNEF, MED8X2 and CPRL in resistance and susceptible lines infected by Bgt pathogen. Gene expression levels were assessed by qRT-PCR at 6, 12, 24, 36, 48, 72 and 96 hpi. Data were normalized to the Actin expression level. The mean expression value was calculated from three independent replicates, and the standard deviation was given at each time points. The dotted line indicated the uninfected controls at the same time points. The corresponding name of genes are listed on top of each panel. R and S represent gene expression in resistance and susceptible line respectively.
Discussion
Plants have the capacity to initiate innate immune systems that recognize the presence of potential pathogens and trigger effective defense responses in the PAMP-triggered immunity (PTI) and effector-triggered immunity (ETI) pathways (Dodds and Rathjen 2010; Miller et al. 2017). PTI was thought to be the first line of activating plant defense and to trigger immediate defense responses leading to basal or nonhost resistance (Miller et al. 2017; Nejat and Mantri 2017). However, PTI does not always fully restrict pathogen proliferation, and can lead to a weak defense response (Schwessinger and Ronald 2012). ETI is activated upon recognition of highly variable microbial molecules (effectors) and is often associated with a hypersensitive response (HR). R genes encode resistance proteins that directly or indirectly detect isolate-specific pathogen effectors encoded by avirulence genes in phytopathology. Unfortunately, there is not always a conceptually clear distinction between PAMPs and effectors, and receptors blur the borderline between PTI and ETI. Thus, knowledge of the plant immune system remains incomplete despite considerable ongoing scientific progress into pathogen sensing and plant immune response mechanisms. Here, we identified several dozens of related resistance genes including RLP, RPM1, RPS2, and so on, which were reported as R genes in the ETI pathway (Bouktila et al. 2015). However, the pathogen could not seem break through the immune protective screen of the leaves of N9134 because there have not any hypersensitive cell death symptom after inoculation with Bgt. Additionally, many models are valid in Arabidopsis but signalling may occur differently in other plant species (Staal and Dixelius 2009). This hinted that both PTI and ETI may also be involved together in pathogen defense because of differences in the diversity of microbial species and fungal race virulence in plants grown in the field.
Wheat (Triticum aestivum L.) is one of the four major cereal grains, but its growth and production are severely affected by pathogens in wheat growing areas worldwide. In practice, the same wheat variety may preform differential resistance levels to different races, including perfect immune, robust HR, middle HR, and susceptible due to gene-for-gene. This makes the ‘resistance’ question more complex for important crop plants. In the present study, we used the triplicate RNA-seq database in leaves of the wheat immune resistance line N9134 inoculated with Bgt to attempt to understand the resistance mechanism to Bgt using WGCNA methods. We found that cochaperones, transcription regulator genes and disease resistance genes were together involved in the wheat defense network, although the interaction net should be further tested in future studies. These findings indicate that R genes were not functioning alone in the resistance phenotype, but instead aided the regulatory mechanism. In wheat, three chromosome sets function together, but the most specific resistance phenotypes are thought to involve a single gene loci controller in classical genetics. Adversely, many genes, especially R gene analogues, were shown to have undergone tandem duplication or multicopy characterization (Baggs et al. 2017; IWGSC 2018). Additionally, around 12% of genes undergo alternative splicing transcription (Zhang et al. 2019b). Taken these results together, we can inferred that the resistance phenotype is a systemic syndrome, which could balance the confusion on the perception of R genes between genetics and phytopathology.
Some key node genes identified in the present study are very similar to those that function in the response of wheat to stripe rust pathogen (Zhang et al. 2019a), although the activated genes are different. For example, the serine/threonine-protein phosphatase PP2A, CNGC2, and calmodulin-binding transcription activator 2 (CAMTA) were implicated in protecting wheat seedlings from infection by Bgt. Serine/threonine protein phosphatases PP1 and PP2A were shown to play key roles in apoptosis, which is a genetically programmed form of cell death (Garcia et al. 2003). The ‘defense, no death’ (dnd1) phenotype is caused by mutation of the gene encoding AtCNGC2, which directly participates in the calcium influx pathogen response signalling cascade and mediates the initiation of cell death programmes during plant defense responses to pathogens (Jurkowski et al. 2004). CAMTA is a small transcription factor family with a broad range of functions in response to environmental stress that regulates the expression of downstream genes in plants (Liu et al. 2015; Rahman et al. 2016). In the present study, the CAMTA2 homologue TraesCS5B02G521200 showed high connectivity with a value of 2362.43 in the WGCNA net and was partially similar to protein phosphatase 2C.
WAKs are receptor-like protein kinases in plant cell walls which have the ability to transmit signals using their cytoplasmic serine/threonine kinase domains (Anderson et al. 2001). They are primarily involved in regulating cell expansion and pathogen responses, and also protect plants from detrimental effects depending upon the state of pectin (Kohorn 2016). Recently, the disease resistance function of ZmWAK that confers quantitative resistance to head smut was reported (Zuo et al. 2015), while two WAK like genes, XA4 and Stb6, conferred race-specific resistance to Xanthomonas oryzae pv. oryzae (Xoo) and Zymoseptoria tritici, respectively (Ning et al. 2017; Saintenac et al. 2018). Here, the WAK protein identified in the plum1 module with a high connectivity value of 1583.83 was predicted to be a subcentre node protein interacting with RLP37, RPM1 and RPS2. However, the aforementioned hub genes and acknowledged R genes are annotated outside the genetic region of PmAS846 flanked by BJ261635 and XFCP620 (Xue et al. 2012) with a distance of 4.8 Mb (IWGSC 2018). Adversely, GNOM-like gene, PP2C isoform X1, and the TM9SF9 gene were located in this region, so are more likely candidate genes of PmAS846 than RPM1, RPP13 and RPS2. These findings stress the fact that resistance genes should be diversified beyond leucine-rich repeat-type genes (Maekawa et al. 2011) such as NLR helper (including but not limited to NLRs) ADR (Bonardi et al. 2011), caffeoyl-CoA O-methyltransferase ZmCCoAOMT2 (Yang et al. 2017), C2H2-type transcription factor bsr-d1 (Li et al. 2017), and actin-depolymerizing factor TaADF7 (Fu et al. 2014). Taken together, our findings suggest another possible function of R genes in the specific activation or modulation of defense pathways including quality and/or quantitative trait loci/genes; this may occur through several genes or gene clusters in resistant plants.
In conclusion, in this study, we reveal the immense complexity of the mechanisms underlying the responses of wheat to powdery mildew. We performed a transcriptome WGCNA analysis of RNA-seq data and identified three co-expressed gene modules. The sample-specific expressed genes in the biggest module were mainly enriched in spliceosomes, peroxisomes, the mRNA surveillance pathway, RNA transport and degradation. Genes in the black module were enriched in phagosomes, biosynthesis/metabolism of amino acids, protein processing in the endoplasmic reticulum, and protein export, while the darkolivegreen4 module consisted of genes enriched in proteasomes, ribosomes, phagosomes, endocytosis, N-glycan biosynthesis, sphingolipid metabolism, and SNARE interactions in vesicular transport. Further, predicted PPI networks from the STRING database substantiated Hsp70, PRH75, LOS, CDC5, GNOM-like, CPCF and KAPP as key hub nodes involved in mRNA surveillance, RNA transport, genetic information processing and regulation, plant–pathogen interaction, and oxidative phosphorylation pathways. Three protein-coding genes induced in chromosome 5BL were identified as potential candidates for Bgt resistance. This study provides important insights into the molecular networks underlying the mechanism of wheat defense against Bgt, while the methodology here can also be used as a reference to narrow down the field of potential key resistance genes using omics and multi-discipline-associated analyses.
References
Anderson C. M., Wagner T. A., Perret M., He Z. H., He D. and Kohorn B. D. 2001 WAKs: cell wall-associated kinases linking the cytoplasm to the extracellular matrix. Plant Mol. Biol. 47, 197–206.
Baggs E., Dagdas G. and Krasileva K. V. 2017 NLR diversity, helpers and integrated domains: making sense of the NLR IDentity. Curr. Opin. Plant Biol. 38, 59−67.
Bonardi V., Tang S., Stallmann A., Roberts M., Cherkis K. and Dangl J. L. 2011 Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proc. Natl. Acad. Sci. USA 108, 16463–16468.
Bouktila D., Khalfallah Y., Habachi-Houimli Y., Mezghani-Khemakhem M., Makni M. and Makni H. 2015 Full-genome identification and characterization of NBS-encoding disease resistance genes in wheat. Mol. Genet. Genomics 290, 257−271.
Brenchley R., Spannagl M., Pfeifer M., Barker G. L., D’Amore R., Allen A. M. et al. 2012 Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491, 705–710.
Dean R., Van Kan J. A., Pretorius Z. A., Hammond-Kosack K. E., Di Pietro A., Spanu P. D. et al. 2012 The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414−430.
Dodds P. N. and Rathjen J. P. 2010 Plant immunity: towards an integrated view of plant-pathogen interactions. Nat. Rev. Genet. 11, 539–548.
Ernst J. and Bar-Joseph Z. 2006 STEM: a tool for the analysis of short time series gene expression data. BMC Bioinformatics 7, 191.
Fu Y., Duan X., Tang C., Li X., Voegele R. T., Wang X. et al. 2014 TaADF7, an actin-depolymerizing factor, contributes to wheat resistance against Puccinia striiformis f. sp. tritici. Plant J. 78, 16−30.
Fu Y., Zhang H., Mandal S. N., Wang C., Chen C. and Ji W. 2016 Quantitative proteomics reveals the central changes of wheat in response to powdery mildew. J. Proteomics 130, 108–119.
Garcia A., Cayla X., Guergnon J., Dessauge F., Hospital V., Rebollo M. P. et al. 2003 Serine/threonine protein phosphatases PP1 and PP2A are key players in apoptosis. Biochimie 85, 721−726.
Hollender C. A., Kang C., Darwish O., Geretz A., Matthews B. F., Slovin J. et al. 2014 Floral transcriptomes in woodland strawberry uncover developing receptacle and anther gene networks. Plant Physiol. 165, 1062–1075.
Hurni S., Brunner S., Buchmann G., Herren G., Jordan T., Krukowski P. et al. 2013 Rye Pm8 and wheat Pm3 are orthologous genes and show evolutionary conservation of resistance function against powdery mildew. Plant J. 76, 957−969.
IWGSC, Appels R., Eversole K., Stein N., Feuillet C., Keller B. et al. 2018 Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361, 661–674.
Janke C. and Bulinski J. C. 2011 Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 12, 773−786.
Jurkowski G. I., Smith R. K., Yu I. C., Ham J. H., Sharma S. B., Klessig D. F. et al. 2004 Arabidopsis DND2, a second cyclic nucleotide-gated ion channel gene for which mutation causes the “defense, no death” phenotype. Mol. Plant-Microbe Interact. 17, 511–520.
Kim D., Pertea G., Trapnell C., Pimentel H., Kelley R. and Salzberg S. L. 2013 TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36.
Kohorn B. D. 2016 Cell wall-associated kinases and pectin perception. J. Exp. Bot. 67, 489−494.
Langfelder P. and Horvath S. 2008 WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559.
Li W., Zhu Z., Chern M., Yin J., Yang C., Ran L. et al. 2017 A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126.
Liu J., Whalley H. J. and Knight M. R. 2015 Combining modelling and experimental approaches to explain how calcium signatures are decoded by calmodulin-binding transcription activators (CAMTAs) to produce specific gene expression responses. New Phytol. 208, 174−187.
Maekawa T., Kufer T. A. and Schulze-Lefert P. 2011 NLR functions in plant and animal immune systems: so far and yet so close. Nat. Immunol. 12, 817–826.
Miller R. N., Costa Alves G. S. and Van Sluys M. A. 2017 Plant immunity: unravelling the complexity of plant responses to biotic stresses. Ann. Bot. 119, 681−687.
Nejat N. and Mantri N. 2017 Plant immune system: crosstalk between responses to biotic and abiotic stresses the missing link in understanding plant defence. Curr. Issues Mol. Biol. 23, 1–16.
Ning Y., Liu W. and Wang G. L. 2017 Balancing immunity and yield in crop plants. Trends Plant Sci. 22,1069−1079.
Rahman H., Xu Y. P., Zhang X. R. and Cai X. Z. 2016 Brassica napus genome possesses extraordinary high number of CAMTA genes and CAMTA3 contributes to PAMP triggered Immunity and Resistance to Sclerotinia sclerotiorum. Front Plant Sci. 7, 581.
Rasmussen S., Barah P., Suarez-Rodriguez M. C., Bressendorff S., Friis P., Costantino P. et al. 2013 Transcriptome responses to combinations of stresses in Arabidopsis. Plant Physiol. 161, 1783–1794.
Reddy A. S., Marquez Y., Kalyna M. and Barta A. 2013 Complexity of the alternative splicing landscape in plants. Plant Cell 25, 3657−3683.
Saintenac C., Lee W. S., Cambon F., Rudd J. J., King R. C., Marande W. et al. 2018 Wheat receptor-kinase-like protein Stb6 controls gene-for-gene resistance to fungal pathogen Zymoseptoria tritici. Nat. Genet. 50, 368–374.
Schwessinger B. and Ronald P. C. 2012 Plant innate immunity: perception of conserved microbial signatures. Annu. Rev. Plant Biol. 63, 451−482.
Staal J. and Dixelius C. 2009 Plant Innate Immunity. In Encyclopedia of life sciences. John Wiley, Chichester.
Szklarczyk D., Franceschini A., Wyder S., Forslund K., Heller D., Huerta-Cepas J. et al. 2015 STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–452.
Trapnell C., Roberts A., Goff L., Pertea G., Kim D., Kelley D. R. et al. 2012 Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562−578.
Xin M., Wang Y., Yao Y., Xie C., Peng H., Ni Z. et al. 2010 Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol. 10, 123.
Xin M., Wang Y., Yao Y., Song N., Hu Z., Qin D. et al. 2011 Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biol. 11, 61.
Xue F., Ji W., Wang C., Zhang H. and Yang B. 2012 High-density mapping and marker development for the powdery mildew resistance gene PmAS846 derived from wild emmer wheat (Triticum turgidum var. dicoccoides). Theor. Appl. Genet. 124, 1549–1560.
Yang Q., He Y., Kabahuma M., Chaya T., Kelly A., Borrego E. et al. 2017 A gene encoding maize caffeoyl-CoA O-methyltransferase confers quantitative resistance to multiple pathogens. Nat. Genet. 49, 1364−1372.
Zhang H., Yang Y., Wang C., Liu M., Li H., Fu Y. et al. 2014 Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genomics 15, 898.
Zhang H., Hu W., Hao J., Lv S., Wang C., Tong W. et al. 2016 Genome-wide identification and functional prediction of novel and fungi-responsive lincRNAs in Triticum aestivum. BMC Genomics 17, 238.
Zhang H., Fu Y., Guo H., Zhang L., Wang C. Y., Song W. N. et al. 2019a Transcriptome and proteome-based network analysis reveals a model of gene activation in wheat resistance to stripe rust. Inter. J. Mol. Sci. 20, 1106.
Zhang H., Mao R., Wang Y. Z., Zhang L., Wang C. Y., Lv S. K., et al. 2019b Transcriptome-wide alternative splicing modulation during plant-pathogen interactions in wheat. Plant Sci. 288, 110160.
Zuo W., Chao Q., Zhang N., Ye J., Tan G., Li B. et al. 2015 A maize wall-associated kinase confers quantitative resistance to head smut. Nat. Genet. 47, 151–157.
Acknowledgments
This work was financially supported by the National Key Technologies Research and Development Programme of China (Grant No. 2017YFD0100701), and by the National Natural Science Foundation of China (31971941). We thank Sarah Williams for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Corresponding editor: Manoj Prasad
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, W., Wang, Q., Wang, S. et al. Gene co-expression network analysis provides a novel insight into the dynamic response of wheat to powdery mildew stress. J Genet 99, 44 (2020). https://doi.org/10.1007/s12041-020-01206-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12041-020-01206-w