Abstract
Genetic information of polymerase chain reaction (PCR)-based markers, one of the main tools of genetics and genomics research in wheat, have been well documented in wheat. However, the physical position in relation to these markers has not yet been systematically characterized. Aim of this study was to characterize the physical information of thousands of widely used molecular markers. We first assigned 2705 molecular markers to wheat physical map, of which 86.1% and 84.7% were the best hits to chromosome survey sequencing (CSS) project (CSS-contigs) and International Wheat Genome Sequencing Consortium Reference Sequence v1.0 (IWGSC RefSeq v1.0), respectively. Physical position of 96.2% markers were predicated based on BLAST analysis, were in accordance with that of the previous nullisomic/aneuploidy/linkage analysis. A suggestive high-density physical map with 4643 loci was constructed, spanning 14.01 Gb (82.4%) of the wheat genome, with 3.02 Mb between adjacent markers. Both forward and reverse primer sequences of 1166 markers had consistent best hits to IWGSC RefSeq v1.0 based on BLAST analysis, and the corresponding allele sizes were characterized. A detailed physical map with 1532 loci was released, spanning 13.93 Gb (81.9%) of the wheat genome, with 9.09 Mb between adjacent markers. Characteristic of recombination rates in different chromosomal regions was discussed. In addition, markers with multiple sites were aligned to homoeologous sites with a consistent order, confirming that a collinearity existed among A, B and D subgenomes. This study facilitates the integration of physical and genetical information of molecular markers, which could be of value for use in genetics and genomics research such as gene/QTL map-based cloning and marker-assisted selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Molecular markers play important roles in wheat genetic and genomic study, such as gene tagging/cloning (Gupta et al. 2008; Zhang et al. 2017), marker-assisted selection (MAS) (Varshney et al. 2007; Gupta et al. 2010), genome structure and evolution study (Gupta et al. 2008; Feldman and Levy 2012). PCR-based markers are timesaving, cost-effective and easy to use, to achieve much higher throughput analysis in the selection of DNA markers. Consequently, PCR-based markers have become one of the main tools for wheat genetic and genomic analyses (Harushima et al. 1998; Ishikawa et al. 2007; Cui et al. 2014, 2017). Various PCR-based markers, such as amplified fragment length polymorphism (AFLP), randam amplified polymorphism DNA (RAPD), genomic-simple sequence repeat (g-SSR), expressed sequence tag (EST)-SSR and sequence-tagged site (STS)-PCR have been used to construct genetic and cytological chromosome maps of wheat (Röder et al. 1995, 1998; Messmer et al. 1999; Paillard et al. 2003; Sourdille et al. 2003, 2004; Xue et al. 2008). Of these, g-SSR and EST-SSR are the most stable and usually inherited in a codominant Mendelian manner. As compared to any other molecular markers, SSR markers are most useful for marker-assisted selection (MAS) in breeding programmes due to their hypervariability and ease of handling (Röder et al. 1995). ESTs represent partial complementary DNA (cDNA) sequences from expressed genes. In addition to EST-SSR markers, EST-derived STS (STS-PCR) have also been extensively explored in cereal crops including wheat (Qi et al. 2004; Xue et al. 2008).
To date, numerous studies regarding quantitative trait locus (QTL) mapping of biologically or agronomically important traits based on g-SSR, EST-SSR and STS-PCR markers have been documented in wheat (Anderson et al. 2001; Cui et al. 2014, 2017). Most of these studies were conducted based on different mapping population with distinct population sizes. In addition, common molecular markers among different studies are limited, which makes it difficult to integrate different maps into a consensus integrative map (Somers et al. 2004; Francki et al. 2009). All these limitations hampered the comparison of common QTL across different mapping populations. Moreover, density, coverage and resolution of current genetic maps are less than satisfactory, especially around the centromeric regions where the recombination events tend to be suppressed (Sourdille et al. 2004; Cui et al. 2017). Compared with genetic mapping information, physical mapping information of molecular markers is the authentic rather than the relative position, which is of great value in genetic and genomic study especially in gene tagging. Previously, most of the wheat molecular markers were assigned to a physical position by nullisomic–tetrasomic (NT) or deletion lines due to the lack of reference genome sequence (Kota et al. 1993; Mickelson-Young et al. 1995; Sorrells et al. 2003; Sourdille et al. 2004). Such physical maps cannot meet the requirements of some further genetic and genomic research such as gene map-based cloning. Wheat genomics research is lagging behind with most of the other important crops such as rice and maize etc. due to the large genome size (17 Gb), hexaploid nature (AABBDD) and high percentage of repetitive regions. However, a rapid progress has been made in the recent five years in wheat genomic research (Jia et al. 2013; Ling et al. 2013, 2018; Mayer et al. 2014; Choulet et al. 2014; Luo et al. 2017; Zhao et al. 2017). The genome assembly of Triticum aestivum cv. Chinese Spring (CS) (https://urgi.versailles.inra.fr/download/iwgsc/IWGSC_RefSeq_Assemblies/v1.0/) and the IWGSC RefSeq v1.0 (the first version of the reference sequence of the bread wheat variety CS) (https://urgi.versailles.inra.fr/download/iwgsc/IWGSC_RefSeq_Assemblies/v1.0/) assembly have been released. These genomic sequence resources facilitate the mapping of molecular markers to physical map in detail. The aim of this study was to characterize the information on physical position of 2705 widely used PCR-based markers and evaluate their potential use in genetic and genomic research. This study is expected to facilitate the integration of physical and genetical information of molecular markers, which is of great value in further genetics and genomics research.
Materials and methods
Primer sequence information for 2705 PCR-based molecular markers
A total of 2705 PCR-based molecular markers were used in this study. These molecular markers included g-SSR, EST-SSR and STS-PCR markers. Of these, the relevant primer sequence information for g-SSR markers, including USDA-ARS Beltsville Agricultural Research Center (BARC), Clermont Ferrand A-genome (CFA), Clermont Ferrand D-genome (CFD), Clermont Ferrand 3B-chromosome (CFT), Gatersleben Wheat Microsatellite (GWM), Gatersleben D-genome Microsatellite (GDM), SSR markers developed and mapped by Génoplante (GPW), Wheat Microsatellite Consortium (WMC) and Markers from P. Stephenson (PSP) codes, as well as PCR-based STS markers of the markers of applied genomics (MAG) code were taken from the Grain Genes website (http://wheat.pw.usda.gov). Relevant primer sequence information about EST-SSR markers prefixed Clermont-Ferrand EST-SSR (CFE), Kansas State University Microsatellite (KSUM) and Microsatellite developed in Conell University Laboratory (CNL) are publicly available (http://wheat.pw.usda.gov/ITMI/EST-SSR/). Primer sequence information for EST-SSR markers of SDAU Wheat EST-SSR (SWES) and randomly named (ww) codes were kindly provided by Prof. Sishen Li, College of Agronomy, Shandong Agricultural University, China. Primer sequences of EST-SSR markers with the prefixes Colorado wheat EST-derived microsatellite (CWEM), EST-derived microsatellite (EDM) and wheat microsatellites derived from cDNAs (CWM) were published in reference articles by Peng and Lapitan (2005), Mullan et al. (2005) and Gao et al. (2004), respectively. Genetic position information for these markers was taken from the corresponding web sites and/or references.
Basic local alignment search tool (BLAST) analysis against wheat genome
We used the BLAST (ftp://ftp.ncbi.nlm.nih.gov/) to align the PCR-based marker probes to the contigs assembled in the chromosome survey sequencing (CSS) project (CSS-contigs), with an aim to determine the chromosomal arms of the markers. All CSS-contigs were downloaded from http://www.wheatgenome.org/. The databases of CSS-contigs of the 21 wheat chromosomes were divided into three groups which describe A, B and D genomes, respectively. Both forward and reverse primer sequences of each PCR-based marker were used for BLAST analysis against these three databases, respectively. Further BLAST analysis against the genome assembly of T. aestivum cv. CS (IWGSC RefSeq v1.0; https://urgi.versailles.inra.fr/download/iwgsc/IWGSC_RefSeq_Assemblies/v1.0/) was performed to obtain their physical locations. As mentioned above, databases of the IWGSC RefSeq v1.0 were divided into three groups which describe A, B and D genomes, respectively. Both forward and reverse primer sequences of each PCR-based marker were used for BLAST analysis against these three databases, respectively. An expectation value (E) of 1E-3 was used as the significance threshold. A suggestive high-density physical map was released based on BLAST analysis against IWGSC RefSeq v1.0 using the forward and/or reverse primer sequences as queries. A detailed physical map was released using markers with consistency physical position in BLAST analysis based on both forward and reverse primer sequences as queries. MapChart 2.2 (http://www.biometris.nl/uk/Software/MapChart/) was used to draw the physical and genetic maps.
Results
General genetic information of the 2705 PCR-based markers
The 2705 PCR-based markers fall into three categories, 1140 were g-SSR markers, 1220 were EST-SSR markers, and 345 were STS-PCR-based markers. All primer sequence information has been listed in table 1 in electronic supplementary material at http://www.ias.ac.in/jgenet. Genetic information of all the 1140 g-SSR markers has been previously reported (http://wheat.pw.usda.gov; Somers et al. 2004) of these, 1039 were reported to be locus-specific that were mapped to a certain unique chromosome; 101 were reported to be assigned on more than one chromosomes, 72.3% of which were assigned to homoeologous sites. To our knowledge, the genetic information for 455 EST-SSR of the 1220 markers are available (http://wheat.pw.usda.gov; http://wheat.pw.usda.gov/ITMI/EST-SSR/; Gao et al. 2004; Mullan et al. 2005; Peng and Lapitan 2005) of which, 281 were locus-specific that were mapped to a unique chromosome; 174 were reported to be assigned on more than one chromosomes, and of which 66.7% were assigned to homoeologous chromosomes. All the 345 STS-PCR markers have been mapped to wheat chromosomes based on linkage/nullisomic analysis (http://wheat.pw.usda.gov) of which, 308 were assigned to a unique wheat chromosome as locus-specific markers; 37 were reported to be assigned on more than one chromosomes, of which 45.9% were assigned to homoeologous sites.
Assignment of 2705 PCR-based marker probes to the CSS-contigs
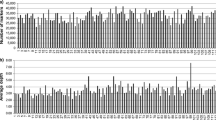
Using 2705 forward primer sequences as queries, BLAST analysis against the three databases of CSS-contigs was performed (table 1 in electronic supplementary material). The results showed that 1316, 1371 and 1386 markers were assigned to chromosomes of wheat A, B and D genomes, respectively (figure 1). A total of 683 markers were assigned to A, B and D genomes simultaneously, 310, 61, 95 and 87 of which were assigned to homoeologous chromosomes of A–B–D (e.g., 2AL_6366997, 2BL_7950323 and 2DL_9909165 for BE404098-2A), A–B (e.g., 3AS_3440701, 3B_10730855 and 4DL_14163974 for Xbarc147-3BS), B–D (e.g., 5AL_2792085, 4BL_6881842 and 4DL_14470349 for Xksum245) and A–D (e.g., 3AL_4291626, 5BL_10845438 and 3DL_6926940 for Xbarc1060-3A) genomes, respectively (figure 2). In addition, 130 markers were assigned to chromosomes of A, B and D genomes in disorder (e.g., 4AL_7061052, 5BS_2278815 and 6DL_1043317 for BE517017-4A). A total of 171, 231 and 200 markers, respectively, were assigned to pairwise chromosomes of A–B, A–D and B–D genomes simultaneously; 53.2%, 68.4% and 61.0% of these markers were assigned to homoeologous chromosomes of A–B (e.g., 3AL_4338880 and 3B_10424015 for Xbarc1044-3A/3B), A–D (e.g., 5AL_1602962 and 5DL_4307567 for Xcfe16-5D) and B–D (e.g., 2BL_8039964 and 2DL_9794606 for Xcinau180-2B/2D) genomes; the remaining markers were assigned to pairwise chromosomes of A–B (e.g., 7AL_4557926 and 6BS_779935 for Xcwm42), A–D (e.g., 7AL_4557920 and 2DS_5318891 for Xgwm261-2D) and B–D (e.g., 2BL_5745690 and 7DL_3387755 for Xmag892-7D) genomes, randomly. Moreover, 231, 317 and 272 markers were assigned to unique chromosome of A (e.g., 7AL_2454499 for Xbarc029-7AL), B (e.g., 7BS_3068985 for Xbarc072-7BS), or D (e.g., 5DL_4529348 for Xcfe27-5A/5B/5D) genomes.
Using 2705 reverse primer sequences as queries, BLAST analysis against the three databases of CSS-contigs was performed again (table 1 in electronic supplementary material). A total of 1328, 1368 and 966 markers were assigned to chromosomes of wheat A, B and D genomes, respectively (figure 1). Of these, 500 were assigned to A, B and D genomes simultaneously, 277, 30, 56 and 45 were assigned to homoeologous sites of A–B–D (e.g., 5AL_2795946, 5BL_10823191 and 5DL_4499420 for BE794904-5A/5B), A–B (e.g., 3AS_3435153, 3B_10656454 and 4DL_14330503 for Xcnl125), B–D (e.g., 4AL_7162314, 1BS_3483987 and 1DS_1913693 for Xcwm70) and A–D (e.g., 7AL_4553763, 2BL_8041131 and 7DL_3313587 for Xedm142-2AS/2DS/4AL/4BS/4DS/7BL/7DS) genomes, respectively (figure 2). In addition, 92 markers were assigned to chromosomes of A, B and D genomes in disorder (e.g., 3AL_4286564, 1BS_3450042 and 2DL_9848697 for Xgdm87-2D). A total of 351, 143 and 141 markers, respectively, were assigned to pairwise chromosomes of A–B, A–D and B–D genomes simultaneously; 55.0%, 67.8% and 67.4% of these markers were assigned to homoeologous chromosomes of A–B (e.g., 3AL_4341827 and 3B_10763162 for Xgwm340-3B), A–D (e.g., 5AL_1164146 and 5DS_2722709 for Xgwm358-5D) and B–D (e.g., 7BL_6697023 and 7DL_3370690 for Xksum198) genomes; the remaining markers were assigned to pairwise chromosomes of A–B (e.g., 4AS_5945307 and 3B_10756713 for Xcau9-1-4B), A–D (e.g., 5AS_854549 and 6DL_2258032 for Xcfd189-5D) and B–D (e.g., 5BL_10793809 and 6DL_3230782 for Xcfd287-6D) genomes randomly. Moreover, 334, 376 and 182 markers were assigned to unique chromosome of A (e.g., 5AL_1954377 for Xcfe069), B (e.g., 4BL_7041683 for Xcfe046), or D (e.g., 6DS_1580594 for Xcfd13-6B/6D) genomes.
In total, 2330 (86.1%) of the 2705 markers have been assigned to CSS-contigs, 1002 of which were verified by blast analysis based on both forward and reverse primer sequences as queries (table 2 in electronic supplementary material). Of these, 39 were assigned to ternate homoeologous chromosomes of A–B–D, simultaneously (e. g., 7AS_4244756, 7BS_3162040 and 7DS_3956966 for Xedm34-7BS); 154 were assigned to diploid homoeologous chromosomes of A–B/A–D/B–D genomes, simultaneously (e.g., 2AS_5307484 and 2BS_5246378 for Xmag4094-2B); 809 were assigned to an unique chromosome of A/B/D genomes (e.g., 7AL_4552970 for Xmag4134-7A; 6BL_4335220 for Xswes1-6B; 1DS_1519366 for Xcfd61-1D).
Alignment of the PCR-based marker probes to IWGSC RefSeq v1.0
Using 2705 forward primer sequences as queries, we conducted BLAST analysis against the three databases of IWGSC RefSeq v1.0 (table 1 in electronic supplementary material). The results showed that 1161, 1251 and 1177 markers were aligned to the wheat chromosomes of A, B and D genomes, respectively (figure 3). A total of 552 markers were assigned to A, B and D genomes simultaneously, 390, 86, 41 and 29 of which were assigned to homoeologous chromosomes of A–B–D (e.g., chr2A656241766, chr2B602764294 and chr2D512687689 for BE404098-2A), A–B (e.g., chr1A115006037, chr1B199487268 and chr2D129818690 for Xcfd17-2D), B–D (e.g., chr5A661148272, chr4B610563857 and chr4D481548817 for Xcfd39-5A) and A–D (e.g., chr2A656241939, chr1B332585855 and chr2D512687872 for Xksum146) genomes, respectively (figure 4). In addition, 92 markers were aligned to chromosomes of A, B and D genomes in disorder (e.g., chr4A313375423, chr2B373755829 and chr5D5597846 for Xcfd183-5D). A total of 186, 150 and 154 markers, respectively, were assigned to pairwised chromosomes of A–B, A–D and B–D genomes simultaneously; 67.2%, 75.3% and 73.4% of these markers were assigned to homoeologous chromosomes of A-B (e.g., chr6A20248916 and chr6B34217980 for Xcfd13-6B/6D), A-D (e.g., chr5A562895693 and chr5D445321992 for Xcfd12-5D) and B–D (e.g., chr6B646268602 and chr6D429073436 for Xbarc204-6DL/6AL) genomes; the remaining markers were assigned to pairwise chromosomes of A–B (e.g., chr7A438170786 and chr6B17721589 for Xcwm42), A–D (e.g., chr7A714684018 and chr2D19623173 for Xgwm261-2D) and B–D (e.g., chr2B798058352 and chr4D499405457 for Xgwm265-2A) genomes randomly. Additionally, 258, 315 and 362 markers were assigned to an unique chromosome of A (e.g., chr5A205213643 for Xgwm129-2B/5A), B (e.g., chr1B684861788 for Xgwm140-1B), or D (e.g., chr3D7094923 for Xgwm161-3D) genome.
The 2705 reverse primer sequences were used as queries to perform BLAST analysis against the three databases of IWGSC RefSeq v1.0 (table 1 in electronic supplementary material). A total of 1209, 1247 and 1341 markers were aligned to the wheat chromosomes of A, B and D genomes, respectively (figure 3). Of these, 652 were assigned to A, B and D genomes simultaneously; 450, 43, 54 and 35 of which were assigned to ternate homoeologous chromosomes of A–B–D (e.g., chr4A51139015, chr4B509036280 and chr4D412716679 for Xgwm165-4D), diploid homoeologous chromosomes of A–B (e.g., chr5A637056686, chr5B640401877 and chr2D155414883 for Xksum242), B–D (e.g., chr4A605665638, chr5B709423235 and chr5D557181863 for Xksum177) and A–D (e.g., chr1A229288898, chr5B191820848 and chr1D310466472 for Xwmc278-1A), respectively (figure 4). Concerning markers aligned to pairwise chromosomes, 75 of the 126, 116 of the 173 and 116 of the 173, respectively, were assigned to homoeologous chromosomes of A–B (e.g., chr2A197790820 and chr2B239195435 for Xww104), A–D (e.g., chr3A739613471 and chr3D607167914 for Xmag896-3A) and B–D (e.g., chr5B6654357 and chr5D5170389 for Xmag705-5B) genomes simultaneously. Moreover, 258, 315 and 362 markers were assigned to unique chromosome of A (e.g., chr5A417896057 for Xmag3794-5A), B (e.g., chr1B660879874 for Xmag322-1B), or D (e.g., chr5D528215209 for Xksum153) genomes.
In total, 2290 (84.7%) of the 2705 markers have been assigned to the IWGSC RefSeq v1.0, 1166 of which were verified by BLAST analysis based on both forward and reverse primer sequences (table 3 in electronic supplementary material). Of these, 83 were assigned to ternate homoeologous chromosomes of A–B–D, simultaneously (e.g., chr4A453604987, chr4B184233671 and chr4D121432813 for Xcwem35-4BS/4AL/4DS); 209 were assigned to diploid homoeologous chromosomes of A–B/A–D/B–D, simultaneously (e.g., chr2B659532169 and chr2D553728845 for Xcfd73-2B/2D); 874 were assigned to an unique chromosome of A/B/D genomes (e.g., chr7A434548295 for Xpsp3050-7A; chr3B818390587 for Xpsp3001-3B; chr2D130832852 for Xwmc18-2D).
The 1166 PCR-based markers that were verified by BLAST analysis based on both forward and reverse primer sequences were aligned to the wheat physical map (figure 5; tables 1 & 4, and figure 1 in electronic supplementary material). Of these, 534 were g-SSR markers, 450 were EST-SSR markers and 182 were STS-PCR-based markers. A total of 292 markers were aligned to multiple chromosomes, 95.2% of which were aligned to homoeologous sites (figure 6; figure 2 in electronic supplementary material). The integrative physical map covered 13927.4 Mb with 4893.7 Mb (35.1%) for the A genome, 5106.9 Mb (36.7%) for the B genome, and 3926.8 Mb (28.2%) for the D genome. Most markers were mapped to the D (37.1%) and B genomes (32.6%), with an average of 81.1 and 71.4 markers per chromosome, respectively. The remaining markers (30.3%) were mapped to the A genome, with an average of 66.3 markers per chromosome. Chromosome 6D had the highest level of marker average density with one marker per 6.21 Mb, whereas chromosome 6B had the lowest with an average of 17.1 Mb between adjacent loci. Moreover, orders of common markers in the physical map were generally in accordance with that of the genetic map reported by Somers et al. (2004) (figure 5; figure 1 in electronic supplementary material). In addition, markers with multiple sites were aligned to homoeologous sites with a consistent order, indicating that collinearity existed among A, B and D subgenomes of wheat (figure 6; figure 2 in electronic supplementary material).
Detailed physical map of the markers that were verified by blast analysis based on both forward and reverse primer sequences and the syntenic relationship between physical and genetic (Somers et al. 2004) positions (group 1 of chromosomes 1A, 1B and 1D). The left map numbered ‘-1’ is the physical map, and the right map numbered ‘-2’ is the high-density consensus integrative genetic map reported by Somers et al. (2004). Green dashed lines connect common loci mapped in both physical and genetic map.
Discussion
Genetic and physical location of PCR-based marker
To date, many PCR-based molecular markers have been physically and genetically mapped based on nullisomic/aneuploidy/linkage analysis (Röder et al. 1998; Sourdille et al. 2003, 2004; Somers et al. 2004; Song et al. 2005; Torada et al. 2006; Li et al. 2007; Francki et al. 2009; Gadaleta et al. 2009; Cui et al. 2014, 2017). In total, 71.7% of the 2705 PCR-based markers reported here have previously been genetically mapped to their corresponding chromosomes, which facilitate their use in genetic analysis and molecular breeding programmes. However, comparison of genetic positions between studies have identified ambiguities because of the use of distinct mapping population sizes, various molecular marker types, different mapping functions, etc. Combining single population genetic maps into a consensus map can resolve disagreements in marker order and distances to some extent. This approach was taken by Somers et al. (2004) who joined four genetic maps into a single consensus map, which resulted in a comprehensive analysis of marker order and distance of DNA markers. As we all known that genetic map construction is conducted based on linkage analysis. Recombination rates between loci varied greatly in different chromosomal regions, which complicate the unit conversions between genetic and physical distance (Sourdille et al. 2003, 2004; Torada et al. 2006; Cui et al. 2014).
Physical mapping information of molecular markers is of great value in genetic study especially in map-based cloning study. Previously, most markers were assigned to a physical position by NT and deletion lines (Kota et al. 1993; Mickelson-Young et al. 1995; Qi et al. 2004; Sourdille et al. 2004; Gupta et al. 2008). Such physical maps remain too rough to meet with the demand of some further genetic analysis such as map-based cloning. Recently, the genome assembly of CS and the IWGSC RefSeq v1.0 assembly are now available. We first assigned 2330 PCR-based molecular markers to CSS-contigs, 1002 of which were verified by blast analysis based on both forward and reverse primer sequences (table 2 in electronic supplementary material). Moreover, 2290 PCR-based molecular markers have been assigned to the IWGSC RefSeq v1.0, 1166 of which were verified by BLAST analysis based on both forward and reverse primer sequences as queries (table 3 in electronic supplementary material). The corresponding physical position information is of great value for their use in targeting additional markers in specific chromosomal regions for fine mapping and map-based clone of a certain QTL. Moreover, physical position information facilitate the integration of some QTL results from separate studies via meta-analysis, thus to determine reliable markers that can be implemented in the context of MAS programmes.
A total of 86.1% and 84.7% of the 2705 markers could be best hits to CSS-contigs and IWGSC RefSeq v1.0 based on primer sequences, respectively, only 37.0% and 43.1 of which could be verified by BLAST analysis based on both forward and reverse primer sequences. These results might be attributed to a fact that many gaps exist in the present reference sequence. In addition, the short queried sequences, chromosomal rearrangement, and the nonspecificity of primer design (e.g., the addition of an untemplated G–C clamp to the 5\(^{\prime }\)-end of primers for Xbarc series markers, Song et al. 2005) might also result in the failure of aligning markers to CSS-contigs and IWGSC RefSeq v1.0. It is worth mentioning that physical position of the 96.2% markers predicated based on BLAST analysis were in accordance with that of the previous nullisomic/aneuploidy/linkage analysis, no matter if it was based on one or two primer sequences as a query (figure 7; figure 3 and table 1 in electronic supplementary material). This finding indicated that most, if not all, of the physical position information listed in table 5 in electronic supplementary material are referable for further genetic and genomic research. A suggestive high-density physical map with 4643 loci was constructed, spanning 14.01 Gb (82.4%) of the wheat genome with 3.02 Mb between adjacent markers (figure 3; table 2 in electronic supplementary material).
Comparison of the present physical map with previous genetic map
A high-density microsatellite consensus map with 1235 microsatellite loci was constructed by Somers et al. (2004). Five hundred and nine (43.7%) of the 1166 physical-mapped markers in the present study had previously been documented by Somers et al. (2004). The marker order in the present physical map was in good agreement with that of the genetic map, with the exception of chromosome 4B, in which an inversion relationship between physical and genetical map was identified (figures 5&8; figure 1 in electronic supplementary material). This consistency not only confirmed the authenticity and credibility of the physical position released here but also facilitate the integration of physical and genetic information.
The comparison of genetic distance with the corresponding physical distances between adjacent markers allowed estimation of information about variations in recombination frequencies and cryptic structural changes (if any) in different regions of individual chromosomes (Gupta et al. 2008). In wheat, a majority of the recombination events occurred on the most distal portions of the chromosomal arms, whereas the recombination events tend to be suppressed around the centromere (Sourdille et al. 2003, 2004; Torada et al. 2006; Cui et al. 2014). These characteristics result in a low resolution of the genetic map in the centromeric region, which was evident in the small genetic distance in the genetic map corresponding to a large physical region around the centromere compared with the most distal portions of the chromosomal arms (figure 5; figure 1 in electronic supplementary material). A comparison of the present physical map with previously published genetical map confirmed this conclusion (Somers et al. 2004). For example, a 2-cM genetic distance of Xcfd36-Xgwm512 on most distal portions of chromosome 2AS corresponds to 4.2 Mb in physical distance, with 1 cM corresponding to 2.1 Mb; a 2-cM genetic distance of Xgwm372-Xwmc261 on chromosome 2A nearby centromere corresponds to 45.3 Mb in physical distance, with 1 cM corresponding to 22.7 Mb. The present study indicates that the unit conversions between genetical and physical distance varied greatly in different wheat chromosomal region. Moreover, much less available markers were distributed in the centromeric region than that of the most distal portions of the chromosomal arms (table 4 in electronic supplementary material). These findings indicate the difficulty of high-resolution mapping and map-based cloning of a QTL around the centromere.
The suggestive physical map of markers determined by blast analysis based on forward and/or reverse primer sequences and the synteny between genetic and physical positions based on common markers reported by Somers et al. (2004) (group 3 of chromosomes 3A, 3B and 3D). The left map numbered ‘-1’ is the physical map, and the right map numbered ‘-2’ is the high-density consensus integrative genetic map reported by Somers et al. (2004). Green dashed lines connect common loci mapped in both physical and genetic map.
Prediction of amplified products sizes of the 1166 PCR-based markers
Both forward and reverse primer sequences of 1166 markers could be best hits for IWGSC RefSeq v1.0 based on BLAST analysis. The corresponding information enables us to determine, or at least predict amplified product sequence and size in CS, thus enabling users to predict product sequence and allele sizes in newly breeding populations and develop molecular breeding and genomics strategies (table 3 in electronic supplementary material). In total, 87.5% of the predicted amplified product sizes were not more than 500 bp, 74.2% of which were within 100–300 bp, consistent with the common PCR-based marker’s amplification fragment size (Röder et al. 1998; Somers et al. 2004; Xue et al. 2008).
Schematic representation of the syntenic relationships between any common marker in wheat physical and genetic maps. Phy-1A to Phy-7D represent the 21 wheat chromosomal physical maps released in this paper; Gen-1A to Gen-7D represent the 21 wheat chromosomal genetic maps reported by Somers et al. (2004).
Allele sizes of most g-SSR and STS-PCR based markers are available (Röder et al. 1998; Somers et al. 2004; http://wheat.pw.usda.gov; http://wheat.pw.usda.gov/ITMI/EST-SSR/). We randomly selected 347 common g-SSR markers reported by Somers et al. (2004) to evaluate the reliability of the predicted allele size in the present study. The results showed that 81.5% of predicted sizes were identical or at least similar (no more than 50 bp in amplified fragment size differences) with those of previous report (figure 9). For example, the predicted alleles of Xgwm95 on chromosome 2A was 122 bp in CS; the alleles reported by Somers et al. (2004) in different varieties were 128 bp, 116 bp, 122 bp, 111 bp, 124 bp, and 126 bp, respectively. The predicted allele of Xwmc216 on chromosome 1D was 126 bp in CS; the alleles reported by Somers et al. (2004) in different varieties were 119 bp, 127 bp, 128 bp and 130 bp. These coincidences further confirmed the authenticity and credibility of the physical position information released here. Sequence of the amplified fragment also could be predicted, which is of great value to some genetic analysis such as fine-mapping and map-based cloning of a target QTL.
Conclusion
We first reported the physical position information of 2705 PCR-based molecular markers, 86.1% and 84.7% of which could be best hits for CSS-contigs and IWGSC RefSeq v1.0, respectively. Physical position of 96.2% markers predicated based on BLAST analysis were in accordance with that of the previous nullisomic/aneuploidy/linkage analysis. A suggestive high-density physical map with 4643 loci was constructed, spanning 14.01 Gb (82.4%) of the wheat genome with 3.02 Mb between adjacent markers. Both forward and reverse primer sequences of 1166 markers could be best hits for a similar position in IWGSC RefSeq v1.0 based on BLAST analysis, and the corresponding allele sizes were characterized. A detailed physical map with 1532 loci was released, spanning 13.93 Gb (81.9%) of the wheat genome with 9.09 Mb between adjacent markers. Characteristic of recombination rates in different chromosomal regions were discussed. In addition, markers with multiple sites were aligned to homoeologous sites with a consistent order, confirming that collinearity existed among A, B, and D subgenomes in wheat. This study facilitates the integration of physical and genetic information of molecular markers, which is of great value for their use in genetic and genomic study such as gene/QTL map-based cloning, MAS, etc.
References
Anderson J. A., Stack R. W., Liu S., Waldron B. L., Fjeld A. D., Coyne C. et al. 2001 DNA markers for Fusarium head blight tolerance QTL in two wheat populations. Theor. Appl. Genet. 102, 1164–1168.
Choulet F., Alberti A., Theil S., Glover N., Barbe V., Daron J. et al. 2014 Structural and functional partitioning of bread wheat chromosome 3B. Science 345, 1249721.
Cui F., Zhao C., Ding A., Li J., Wang L., Li X. et al. 2014 Construction of an integrative linkage map and QTL mapping of grain yield-related traits using three related wheat RIL populations. Theor. Appl. Genet. 127, 659–675.
Cui F., Zhang, N., Fan X., Zhang W., Zhao C., Yang L. et al. 2017 Utilization of a Wheat660K SNP array-derived high-density genetic map for high-resolution mapping of a major QTL for kernel number. Sci. Rep. 7, 3788.
Feldman M. and Levy A. A. 2012 Genome evolution due to allopolyploidization in wheat. Genetics 192, 3763–3774.
Francki M. G., Walker E., Crawford A. C., Broughton S., Ohm H. W., Barclay I. et al. 2009 Comparison of genetic and cytogenetic maps of hexaploid wheat (Triticum aestivum L.) using SSR and DArT markers. Mol. Genet. Genomics 281, 181–191.
Gadaleta A., Giancaspro A., Giove S. L., Zacheo S., Mangini G., Simeone R. et al. 2009 Genetic and physical mapping of new EST-derived SSRs on the A and B genome chromosomes of wheat. Theor. Appl. Genet. 118, 1015–1025.
Gao L. F., Jing R. L., Huo N. X., Li Y., Li X. P., Zhou R. H. et al. 2004 One hundred and one new microsatellite loci derived from ESTs (EST-SSR) in bread wheat. Theor. Appl. Genet. 108, 1392–1400.
Gupta P. K., Mir R. R., Mohan A. and Kumar J. 2008 Wheat genomics: present status and future prospects. Int. J. Plant Genome 2008, 1–36.
Gupta P. K., Langridge P. and Mir R. R. 2010 Marker-assisted wheat breeding: present status and future possibilities. Mol. Breed. 26, 145–161.
Harushima Y., Yano M., Shomura A., Sato M., Shimano T., Kuboki Y. et al. 1998 A high-density rice genetic linkage map with 2275 markers using a single \(\text{ F }_{{2}}\) population. Genetics 148, 479–494.
Ishikawa G., Yonemaru J., Saito M. and Nakamura T. 2007 PCR-based landmark unique gene (PLUG) markers effectively assign homoeologous wheat genes to A, B and D genomes. BMC Genomics 8, 135.
Jia J., Zhao S., Kong X., Li Y., Zhao G., He W. et al. 2013 Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496, 91–95.
Kota R. S., Gill K. S., Gill B. S. and Endo T. R. 1993 A cytogenetically based physical map of chromosome 1B in common wheat. Genome 36, 548–554.
Li S. S., Jia J. Z., Wei X. Y., Zhang X. C., Li L. Z., Chen H. M. et al. 2007 A intervarietal genetic map and QTL analysis for yield traits in wheat. Mol. Breed. 20, 167–178.
Ling H. Q., Zhao S., Liu D., Wang J., Sun H., Zhang C. et al. 2013 Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 496, 87–90.
Ling H. Q., Ma B., Shi X. L., Liu H., Dong L. L., Sun H. et al. 2018 Genome sequence of the progenitor of wheat A subgenome Triticum urartu. Nature 557, 424–428.
Luo M. C., Gu Y. Q., Puiu D., Wang H., Twardziok S. O., Deal K. R. et al. 2017 Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature 551, 498–502.
Mayer K. F. X., Rogers J., Doležel J., Pozniak C., Eversole K., Feuillet C. et al. 2014 A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345, 1251788.
Messmer M. M., Keller M., Zanetti S. and Keller B. 1999 Genetic linkage map of wheat \(\times \) spelt cross. Theor. Appl. Genet. 98, 1163–1170.
Mickelson-Young L., Endo T. R. and Gill B. S. 1995 A cytogenetic laddermap of the wheat homoeologous group-4 chromosomes. Theor. Appl. Genet. 90, 1007–1011.
Mullan D. J., Platteter A., Teakle N. L., Appels R., Colmer T. D., Anderson J. M. and Francki M. G. 2005 EST-derived SSR markers from defined regions of the wheat genome to identify Lophopyrum elongatum specific loci. Genome 48, 811–822.
Paillard S., Schnurbusch T., Winzeler M., Messmer M., Sourdille P., Abderhalden O. et al. 2003 An integrative genetic linkage map of winter wheat (Triticum aestivum L.). Theor. Appl. Genet. 107, 1235–1242.
Peng J. H. and Lapitan N. L. V. 2005 Characterization of EST-derived microsatellites in the wheat genome and development of eSSR markers. Funct. Integr. Genomics 5, 80–96.
Qi L. L., Echalier B., Chao S., Lazo G. R., Butler G. E., Anderson O. D. et al. 2004 A chromosome bin map of 16,000 EST loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168, 701–712.
Röder M. S., Plaschke J., König S. U., Börner A., Sorrells M. E., Tanksley S. D. et al. 1995 Abundance, variability and chromosomal location of microsatellites in wheat. Mol. Gen. Genet 246, 327–333.
Röder M. S, Korzun V., Wendehake K., Tixier M. H., Leroy P. and Ganal M. W. 1998 A microsatellite map of wheat. Genetics 149, 2007–2023.
Somers D. J., Isaac P. and Edwards K. 2004 A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 109, 1105–1114.
Song Q. J., Shi J. R., Singh S., Fickus E. W., Costa J. M., Lewis J. et al. 2005 Development and mapping of microsatellite (SSR) markers in wheat. Theor. Appl. Genet. 110, 550–560.
Sorrells M. E., Rota M. L., Bermudez-Kandianis C. E., Greene R. A., Kantety R., Munkvold J. D. et al. 2003 Comparative DNA sequence analysis of wheat and rice genomes. Genome Res. 13, 1818–1827.
Sourdille P., Cadalen T., Guyomarc’h H., Snape J. W., Perretant M. R., Charmet G. et al. 2003 An update of the Courtot\(\times \)Chinese Spring intervarietal molecular marker linkage map for the QTL detection of agronomic traits in wheat. Theor. Appl. Genet. 106, 530–538.
Sourdille P., Singh S., Cadalen T., Gina L., Brown-Guedira G. L., Gay G. et al. 2004 Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct. Integr. Genomics 4, 12–25.
Torada A., Koike M., Mochida K. and Ogihara Y. 2006 SSR-based linkage map with new markers using an intraspecific population of com-mon wheat. Theor. Appl. Genet. 112, 1042–1051.
Varshney R. K., Langridge P. and Graner A. 2007 Application of genomics to molecular breeding of wheat and barley. Adv. Genet. 58, 121–155.
Xue S., Zhang Z., Lin F., Kong Z., Cao Y., Li C. et al. 2008 A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor. Appl. Genet. 117, 181–189.
Zhang W. Z., Chen S. S., Abate Z. D., Nirmala J. N., Rouse M., Dubcovsky J. et al. 2017 Identification and characterization of Sr13, a tetraploid wheat gene that confers resistance to the Ug99 stem rust race group. Proc. Natl. Acad. Sci. USA 114, E9483–E9492.
Zhao G. Y., Zou C., Li K., Wang K., Li T. B., Gao L. F. et al. 2017 The Aegilops tauschii genome reveals multiple impacts of transposons. Nat. Plants 946, 946–955.
Acknowledgements
This research was supported by the Shandong Provincial Science Foundation for Outstanding Youth (ZR2017JL017), National Natural Science Foundation of China (31701505, 31671673, 31871612), Yantai Key Research and Development Plan (2017ZH052), Shandong Provincial Science Foundation (ZR2018PC016), the Open Project Programme (CSBAAKF2018003) of State Key Laboratory of Crop Stress Biology for Arid Areas, NWAFU, the Key Research Programme of the Chinese Academy of Sciences (KFZD-SW-110) and Shandong key research and development plan (2017NC210012).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Arun Joshi
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, C., Sun, H., Guan, C. et al. Physical information of 2705 PCR-based molecular markers and the evaluation of their potential use in wheat. J Genet 98, 69 (2019). https://doi.org/10.1007/s12041-019-1114-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12041-019-1114-1