Abstract
The major cause of cancer-related deaths in patients with lung adenocarcinoma (LAD) is due to distant metastasis. Many reports have indicated that miRNA plays a key role in tumour metastasis. The expression of miR-197 is correlated with LAD progression, however, the mechanism of miR-197 is still unknown in the processing of LAD. A Boyden chamber migration/invasion assay was used for the metastatic function study in vitro. Real-time PCR and Western blot assays were employed to analyse the EMT hallmark changes in both the mRNA and protein levels. \(3^{\prime }\)-UTR reporter luciferase assay was used to show that HIPK2 is a direct target of miR-197. miR-197 enhances LAD cell migration and invasion miR-197. The downregulation of miR-197 suppresses the EMT and migration ability. HIPK2 is a direct functional target of miR-197 in LAD metastasis. In summary, miR-197 controls EMT and metastasis by directly silencing HIPK2. The findings suggest that interfering with the miR-197-dependent regulation of HIPK2 could be a useful approach for the treatment of patients with late stage metastatic LAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the leading cause of cancer-related deaths in men worldwide, accounting for 1.8 million deaths annually. The most common type, lung adenocarcinoma (LAD), constitutes almost 40% of lung cancers. Approximately 90% of patients with LAD develop distant metastasis at the advanced stage (Sequist et al. 2013; Dey et al. 2015; Wu et al. 2015). Therefore, to design effective therapeutic strategies for patients with LAD, it is very important to understand the molecular mechanisms underlying distant metastasis. Epithelial–mesenchymal transition (EMT) is an evolutionarily conserved developmental process during which epithelial cells lose polarity and then acquire a mesenchymal phenotype, and this transition has been involved in the initiation of metastasis (Brabletz 2012; Tang et al. 2016).
MiR-197 enhances LAD cell migration and invasion in vitro. The migration and invasion abilities of A549 (a) and PC9 (b) cells when transfected with the miR-197 overexpression vector or an empty vector as assessed by Transwell migration and Matrigel invasion assay for 24 h. All experiments were performed at least thrice, and the data are expressed as means \(\pm \hbox { SD}\). The statistical significance of differences was measured by unpaired student’s t-test. *\(P<0.05\), **\(P<0.01\).
MicroRNAs (miRNAs) are small noncoding RNAs that upregulate or downregulate gene expression posttranscriptionally. It is well-known that miRNAs play a central role in tumour metastasis. For example, the miR-200 family members attenuate the EMT through targeting the repressors of E-cadherin that induce epithelial differentiation, or by targeting the EMT activators known as ZEB1/2 genes (Burk et al. 2008; Iliopoulos et al. 2010; Chang et al. 2011). Previously, miR-197 has reportedly been aberrantly expressed in different types of tumours. It plays a dual role by regulating tumour growth according to the tumour types, as oncogenes or tumour suppressors. On the one hand, miR-197 induces apoptosis, suppresses tumourigenicity, and inhibits cell proliferation and migration (Dai et al. 2014; Wang et al. 2015; Xin et al. 2015; Yang et al. 2015; Tian et al. 2016). Besides, miR-197 can suppress the expression of the tumour suppressor gene FUS1, and its expression is correlated with poor clinical outcomes in patients with LAD (Du et al. 2009; Mavridis et al. 2015). In these patients, miR-197 expression was significantly higher in tumours than in normal tissue and was associated with advanced tumour stage, overall poor survival and higher recurrence rates (Du et al. 2009; Mavridis et al. 2015), which suggest that miR-197 is important for LAD development. However, no studies have systematically determined the role of miR-197 in the development of metastatic disease in LAD.
In this study, we demonstrated the function of miR-197 in LAD and found that miR-197 strongly activates EMT and ultimately promotes LAD metastasis by targeting HIPK2. These observations suggest that miR-197 could be a therapeutic target for preventing LAD metastasis.
Materials and methods
Cell culture
The human LAD cell lines A549 and PC9 were obtained from the American Type Culture Collection (ATCC). The cell lines were maintained in Dulbecco’s modified Eagle’s medium (DMEM) with 10% foetal bovine serum (FBS), and they were incubated at 5% \(\hbox {CO}_{2}\) at \(37^{\circ }\hbox {C}\).
Real-time PCR
Total RNA was extracted from the LAD cells by using TRIzol (Invitrogen, Carlsbad, USA) following the manufacturer’s protocol. Quantification was determined by using NanoDrop 1000 spectrophotometer (Thermo Scientific, Pittsburgh, USA). Reverse transcription of RNA was performed using the ImProm-II reverse transcription system (Promega, Madison, USA) following the manufacturer’s instructions. Real-time PCR was performed in an ABI7500 Prism Sequence Detection System (Applied Biosystems, Foster City, USA) by using a SYBR Green kit (TaKaRa, Tokyo, Japan), and the relative changes were quantified. The \(2^{-\Delta \Delta \mathrm{CT}}\) method was used to measure gene expression. Each experiment was repeated at least thrice.
MiR-197 promoted LAD metastasis is mediated by the EMT. (a) The expression of EMT markers as analysed in miR-197 overexpressed LAD cell lines by real-time PCR. (b) EMT markers in two miR-197 overexpressed LAD cell lines as analysed by Western blots. All experiments were performed at least thrice and the data are expressed as means \(\pm \hbox { SD}\). The statistical significance of differences was measured by unpaired student’s t-test. *\(P<0.05\), **\(P<0.01\).
RNA interference
LAD cells were stably infected with the premicroRNA expression construct known as the lenti-miR expression plasmid, which contained the full-length miR-197 in the H1-MCS-CMV-EGFP vector (GeneChem, Shanghai, China). The sh-miR-197 was cloned into the H1-MCS-CMV-EGFP vector (GeneChem) to generate H1-MCS-CMV-EGFP-sh-miR-197 (GeneChem). A nontargeting sequence was used as a lentivirus negative control and was purchased from GeneChem.
Migration and invasion assays
A migration assay was performed using 24-well culture inserts with porous polycarbonate membranes (\(8.0~\mu \hbox {m}\), Millipore, USA). For the Matrigel invasion assay, the filters were precoated with \(30~\mu \hbox {L}\) of Matrigel (BD Biosciences, San Jose, USA) for 4 h. In brief, \(5\times 10^{4}\) cells in \(200~\mu \hbox {L}\) of serum-free medium were added to the upper chamber, and \(800~\mu \hbox {L}\) of medium with 10% serum was placed in the lower chamber. The plates were incubated for 24 h at \(37^{\circ }\hbox {C}\) in 5% \(\hbox {CO}_{2}\). Cells that did not migrate or invade through the pores were removed with the cotton swab. Cells on the lower surface of the membrane were examined and counted under a microscope. Each experiment was repeated at least thrice.
Western blot assay
LAD cells were washed with PBS, collected, and lysed with RIPA lysis buffer (Cell Signaling Technology, Danvers, USA). Protein concentrations were measured using Qubit Protein Assay Kit and Qubit 2.0 Fluorometer (Invitrogen). Approximately \(50~\mu \hbox {g}\) of total protein was loaded on a 10% or 12% SDS-PAGE gel, and transferred to polyvinylidene difluoride membranes for immunoblotting. Membranes were blocked in 5% milk in TBST and incubated with primary antibody at \(4^{\circ }\hbox {C}\) overnight, followed with horseradish peroxidase (HRP)-linked secondary antibody, and then detected with Pierce ECL western blotting substrate (Thermo Scientific). The following antibodies were used: E-cadherin (1:1000, Abcam, Cambridge), snail (1:1000, Abcam, Cambridge), vimentin (1:1000, Abcam, Cambridge), HIPK2 (1:1000, Cell Signaling Technology, USA), and \(\upbeta \)-actin (1:5000, Boster, China).
The downregulation of miR-197 suppresses the EMT and migration ability in LAD cells. The migration and invasion abilities of A549 (a) and PC9 cells (b) as transfected with the sh-miR-197 vector or an empty vector and assessed by Transwell migration and Matrigel invasion assay for 24 h. The expression of EMT markers analysed in sh-miR-197 transfected A549 cells (c) and PC9 cells (d) by real-time PCR. All experiments were performed at least thrice and the data are expressed as the means \(\pm \hbox { SD}\). The statistical significance of differences was measured by unpaired student’s t-test. *\(P<0.05\), **\(P<0.01\).
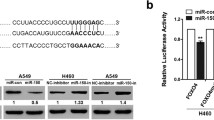
HIPK2 is a direct target of miR-197 in LAD cells. (a) A schematic representation of putative miR-197 binding in the \(3^{\prime }\)-UTR of HIPK2. (b) Dual-luciferase assays showing the repression of wild-type UTR (HIPK2-\(3^{\prime }\)-UTR) or mutant UTR (HIPK2-\(3^{\prime }\)-UTR mut) following the transfection of the vector or miR-197 overexpression vector. All experiments were performed at least thrice and the data are expressed as the means \(\pm \hbox { SD}\). The statistical significance of differences was measured by unpaired student’s t-test. **\(P<0.01\).
HIPK2 was a direct functional target of miR-197 in LAD metastasis. (a) The migration capacities of A549 cells after the indicated treatment as assessed by Transwell migration assay. *\(P<0.05\), **\(P<0.01\). (b) The invasive capacities of A549 cells after the indicated treatment as assessed by Matrigel invasion assay. All experiments were performed at least thrice and the data are expressed as the means \(\pm \hbox { SD}\). The statistical significance of differences was measured by unpaired student’s t-test. *\(P<0.05\), **\(P<0.01\).
3\(^{\prime }\)-UTR luciferase reporter assay
Lipofectamine-2000 (Invitrogen) was used to transfect A549 cells with luciferase vectors (an empty luciferase vector, a luciferase vector containing the \(3^{\prime }\)-UTR of the wild-type target gene, and a luciferase vector containing the \(3^{\prime }\)-UTR of the mutant-type target gene) for HIPK2 together with miR-197 or the negative control. After 48 h, transfected A549 cells in each well of 24-well plates were harvested with \(100~\mu \hbox {L}\) of lysis buffer. Ten \(\mu \hbox {L}\) of the cell extract was used to measure luciferase activity by using the Dual-Luciferase Reporter Assay System (Promega, Madison, USA). All assays were performed in triplicate and the data were presented as the ratios between firefly and Renilla fluorescence activities.
Statistical analyses
All data were presented as the means ± SD. The statistical analyses were performed using Student’s t-test. A difference was considered statistically significant when \(P<0.05\). All the statistical analyses were performed with SPSS 13.0 software.
Results
MiR-197 enhances LAD cell migration and invasion
First, we analysed the role of miR-197 in metastasis by migration and invasion studies. We generated LAD cell lines with stable miR-197 overexpression in A549 and PC9 cells using lentivirus transfection. Next, we performed a Boyden chamber migration/invasion assay to study metastatic function in vitro. The ectopic expression of miR-197 significantly increased the migratory and invasive abilities of both miR-197 overexpressing A549 and PC9 cells (figure 1, a–b). Collectively, our data suggested that miR-197 overexpression significantly increased the migratory and invasive abilities of LAD cells in vitro.
MiR-197 promotes LAD metastasis through EMT
EMT plays a critical role in the metastasis process, which provides the motility, invasion and migration properties of cancer cells (Brabletz 2012; Tang et al. 2016). Therefore, we further explored whether miR-197 promoted LAD metastasis is mediated through the EMT. To confirm this hypothesis, we first used miR-197 overexpressing A549 and PC9 cells to analyse the hallmark EMT changes at both the mRNA and protein levels. As shown in figure 2, a–b, the epithelial marker E-cadherin was dramatically decreased after miR-197 overexpression. However, the mesenchymal markers vimentin and snail were increased in miR-197 overexpressing cells compared to the vector cells. Taken together, these results demonstrate that miR-197 enhances the EMT process in LAD cells.
The downregulation of miR-197 suppresses the EMT and migration ability
Having shown that miR-197 overexpression could enhance the EMT process in LAD cells, we next employed a loss-of-function approach by using shRNA to investigate the role of miR-197 in the EMT process. As anticipated, the migratory and invasive capabilities of both A549 and PC9 cells were significantly reduced by miR-197 inhibition (figure 3, a–b). In addition, as shown in figure 3, c–d, the epithelial marker E-cadherin was increased, and the mesenchymal marker vimentin was decreased in sh-miR-197 transfected A549 and PC9 cells compared with the vector groups. Collectively, our findings suggest that miR-197-promoted LAD metastasis is mediated by EMT.
HIPK2 is a direct target of miR-197 in LAD cells
To investigate the roles for the target gene of miR-197 in the EMT process and metastasis in LAD, we first performed a miRNA target gene prediction with Targetscan database. We found that HIPK2 exhibited miR-197-binding sequences in its \(3^{\prime }\)-UTR regions (figure 4a). Next, the luciferase activity was decreased by miR-197 overexpression when the Luc-HIPK2-wt was present, compared with the luciferase activity in the Luc-HIPK2-mu, suggesting that miR-197 reduced the luciferase activity of Luc-HIPK2-wt but had no effect on Luc-HIPK2-mu (figure 4b). Taken together, our results suggest that miR-197 promotes LAD metastasis by directly targeting HIPK2.
HIPK2 is a direct functional target of miR-197 in LAD metastasis
We further tested whether HIPK2 is functionally regulated by miR-197 in LAD metastasis. Interestingly, the miR-197 overexpression-associated migration and invasion changes could be partially rescued by HIPK2 overexpression (figure 5, a–b). Collectively, these results demonstrated that miR-197 promotes LAD metastasis by targeting HIPK2.
Discussion
Many reports have demonstrated that the dysregulation of miRNAs contributes to cancer metastasis (Rabinowits et al. 2009; Martello et al. 2010; Song et al. 2013). However, it is not clear whether miRNAs play a key role in the regulation of LAD metastasis. The present study identified that miR-197 is involved in the promotion of LAD metastasis. For the first time, we found that miR-197 promoted the migratory and invasive abilities of LAD cells with two different cell lines and with overexpression and knockdown experiments. In this study, we provide strong evidence of the role of miR-197 through experiments performed in two different LAD cell lines. Moreover, we demonstrated that miR-197 enhanced EMT of LAD cells, an important process for metastasis, supporting the role of miR-197 as a factor that promotes LAD metastasis. This finding supports a previous report showing that miR-197 induces EMT by targeting p120 catenin in pancreatic cancer cells (Hamada et al. 2013).
The EMT process is a crucial step in initiating the metastatic spread of many tumour cells into distal organs in a variety of cancers (Tsai and Yang 2013; Puisieux et al. 2014). Importantly, miRNAs have been shown to regulate EMT (Song et al. 2014; Liang et al. 2015; Listing et al. 2015). Here, we found that miR-197 expression is involved in inducing the EMT process, as demonstrated by the gain of mesenchymal markers and loss of epithelial markers. Moreover, overexpression of miR-197 greatly promoted EMT in LAD cells, while knock down of miR-197 by shRNA inhibited EMT. Importantly, a high level of miR-197 expression was significantly associated with low E-cadherin expression and high vimentin and snail expression, which supports the loss-of-function and gain-of-function studies we performed in LAD tumour cell lines. To our knowledge, this study is the first to demonstrate that miR-197 enhances the EMT process in LAD.
It is well established that miRNA regulates a target gene expression to perform its function (Chen et al. 2015; Cekaite et al. 2016). Therefore, in this study, we investigated the functional target gene for miR-197 that was relative to LAD metastasis regulation. HIPK2 was predicted to be a functional target gene of miR-197 by a TargetScan database analysis. Moreover, we found that miR-197 directly bound to the \(3^{\prime }\)-UTR region of HIPK2 and suppressed HIPK2 expression. Previously, there was no direct evidence of a relationship between miR-197 and HIPK2 in cancer cells. This study provides the first evidence demonstrating that miR-197 directly regulates HIPK2 in LAD cells.
In conclusion, collectively our study has significant implications for understanding the underlying mechanisms of how miR-197 contributes to tumour progress in LAD. MiR-197 controls EMT and metastasis by directly silencing HIPK2. The findings from this study suggest that interfering with the miR-197 dependent regulation of HIPK2 could be a useful approach for the treatment of patients with late stage metastatic LAD.
References
Brabletz T. 2012 EMT and MET in metastasis: where are the cancer stem cells? Cancer Cell. 22, 699–701.
Burk U., Schubert J., Wellner U., Schmalhofer O., Vincan E., Spaderna S. et al. 2008 A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 9, 582–589.
Cekaite L., Eide P. W., Lind G. E., Skotheim R. I. and Lothe R. A. 2016 MicroRNAs as growth regulators, their function and biomarker status in colorectal cancer. Oncotarget 7, 6476–6505.
Chang C. J., Chao C. H., Xia W., Yang J. Y., Xiong Y., Li C. W. et al. 2011 P53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat. Cell Biol. 13, 317–323.
Chen W., Fan X. M., Mao L., Zhang J. Y., Li J., Wu J. Z. et al. 2015 MicroRNA-224: as a potential target for miR-based therapy of cancer. Tumour Biol. 36, 6645–6652.
Dai W., Wang C., Wang F., Wang Y., Shen M., Chen K. et al. 2014 Anti-miR-197 inhibits migration in HCC cells by targeting KAI 1/CD82. Biochem. Biophys. Res. Commun. 446, 541–548.
Dey S., Sayers C. M., Verginadis I. I., Lehman S. L., Cheng Y., Cerniglia G. J. et al. 2015 ATF4-dependent induction of heme oxygenase 1 prevents anoikis and promotes metastasis. J. Clin. Invest. 125, 2592–2608.
Du L., Schageman J. J., Subauste M. C., Saber B., Hammond S. M., Prudkin L. et al. 2009 miR-93, miR-98, and miR-197 regulate expression of tumor suppressor gene FUS1. Mol. Cancer Res. 7, 1234–1243.
Hamada S., Satoh K., Miura S., Hirota M., Kanno A., Masamune A. et al. 2013 miR-197 induces epithelial-mesenchymal transition in pancreatic cancer cells by targeting p120 catenin. J. Cell Physiol. 228, 1255–1263.
Iliopoulos D., Lindahl-Allen M., Polytarchou C., Hirsch H. A., Tsichlis P. N. and Struhl K. 2010 Loss of miR-200 inhibition of Suz12 leads to polycomb-mediated repression required for the formation and maintenance of cancer stem cells. Mol. Cell 39, 761–772.
Liang J., Li Y., Daniels G., Sfanos K., De Marzo A., Wei J. et al. 2015 LEF1 targeting EMT in prostate cancer invasion is regulated by miR-34a. Mol. Cancer Res. 13, 681–688.
Listing H., Mardin W. A., Wohlfromm S., Mees S. T. and Haier J. 2015 MiR-23a/-24-induced gene silencing results in mesothelial cell integration of pancreatic cancer. Br. J. Cancer 112, 131–139.
Martello G., Rosato A., Ferrari F., Manfrin A., Cordenonsi M., Dupont S. et al. 2010 A microRNA targeting dicer for metastasis control. Cell 141, 1195–1207.
Mavridis K., Gueugnon F., Petit-Courty A., Courty Y., Barascu A., Guyetant S. et al. 2015 The oncomiR miR-197 is a novel prognostic indicator for non-small cell lung cancer patients. Br. J. Cancer 112, 1527–1535.
Puisieux A., Brabletz T. and Caramel J. 2014 Oncogenic roles of EMT-inducing transcription factors. Nat. Cell Biol. 16, 488–494.
Rabinowits G., Gerçel-Taylor C., Day J. M., Taylor D. D. and Kloecker G. H. 2009 Exosomal microRNA: a diagnostic marker for lung cancer. Clin. Lung Cancer 10, 42–46.
Sequist L. V., Yang J. C., Yamamoto N., O’Byrne K., Hirsh V., Mok T. et al. 2013 Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J. Clin. Oncol. 31, 3327–3334.
Song S. J., Poliseno L., Song M. S., Ala U., Webster K., Ng C. et al. 2013 MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 154, 311–324.
Song Y., Li J., Zhu Y., Dai Y., Zeng T., Liu L. et al. 2014 MicroRNA-9 promotes tumor metastasis via repressing E-cadherin in esophageal squamous cell carcinoma. Oncotarget 5, 11669–11680.
Tang J., Li Y., Wang J., Wen Z., Lai M. and Zhang H. 2016 Molecular mechanisms of microRNAs in regulating epithelial-mesenchymal transitions in human cancers. Cancer Lett. 371, 301–313.
Tian L. Q., Liu E. Q., Zhu X. D., Wang X. G., Li J. and Xu G. M. 2016 MicroRNA-197 inhibits cell proliferation by targeting GAB2 in glioblastoma. Mol. Med. Rep. 13, 4279–4288.
Tsai J. H. and Yang J. 2013 Epithelial-mesenchymal plasticity in carcinoma metastasis. Genes Dev. 27, 2192–2206.
Wang H., Su X., Yang M., Chen T., Hou J., Li N. et al. 2015 Reciprocal control of miR-197 and IL-6/STAT3 pathway reveals miR-197 as potential therapeutic target for hepatocellular carcinoma. Oncoimmunology 4, e1031440.
Wu Y., Liu H., Shi X., Yao Y., Yang W. and Song Y. 2015 The long non-coding RNA HNF1A-AS1 regulates proliferation and metastasis in lung adenocarcinoma. Oncotarget 6, 9160–9172.
Xin J., Zhang X. K., Xin D. Y., Li X. F., Sun D. K., Ma Y. Y. et al. 2015 FUS1 acts as a tumor-suppressor gene by upregulating miR-197 in human glioblastoma. Oncol. Rep. 34, 868–876.
Yang Y., Li F., Saha M. N., Abdi J., Qiu L. and Chang H. 2015 miR-137 and miR-197 induce apoptosis and suppress tumorigenicity by targeting MCL-1 in multiple myeloma. Clin. Cancer Res. 21, 2399–2411.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Dhavendra Kumar
NZ carried out most of the experiments and drafted the manuscript. LT carried out migration and invasion assays. ZM designed the study and performed the statistical analysis. NG conceived the study, and designed and coordinated, and helped to draft the manuscript. All authors read and approved the final manuscript.
Rights and permissions
About this article
Cite this article
Zhang, N., Tian, L., Miao, Z. et al. MicroRNA-197 induces epithelial–mesenchymal transition and invasion through the downregulation of HIPK2 in lung adenocarcinoma. J Genet 97, 47–53 (2018). https://doi.org/10.1007/s12041-018-0881-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-0881-4