Abstract
Human bone morphogenetic protein 2 (BMP2) is a bone-growth regulatory factor involved in the formation of bone and cartilage, and has been recognized as an attractive therapeutic target for a variety of bone diseases and defects. Here, we report successful design of a head-to-tail cyclic peptide based on crystal structure to target BMP2. Computational alanine scanning identifies two hotspot regions at the crystal complex interface of BMP2 with its type-IA receptor; promising one is stripped from the interface to derive a linear self-inhibitory peptide RPS2[r78−94] that covers residues 78–94 of the receptor protein. Dynamics simulation and energetics analysis reveal that the peptide is highly flexible in isolated state and cannot spontaneously bind to BMP2. The RPS2[r78−94] peptide is further extended from its N- and C-termini until reaching two spatially vicinal residues 74 and 98 in the crystal structure of intact BMP2–receptor complex system, consequently resulting in a longer peptide RPS2[r74−98], which is then cyclized in a head-to-tail manner to obtain its cyclic counterpart cycRPS2[r74−98]. Computational analysis suggests that the cyclic peptide can well maintain in a conformation similar with its active conformation in complex crystal structure, exhibiting a smaller disorder and a larger potency than its linear counterpart. Further assays confirm that the two linear peptides RPS2[r78−94] and RPS2[r74−98] are nonbinders of BMP2, whereas, as designed, the cyclic peptide cycRPS2[r74−98] can bind to BMP2 with a moderate affinity. The cyclic peptide is expected as a lead molecular entity to develop new and potent peptide-based drugs for BMP2-targeted therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Human bone morphogenetic proteins (BMPs) are a group of growth factors that belong to the TGFβ superfamily of cytokines and metabologens (Chen et al. 2004). Similar to other TGFβ family proteins, BMPs are highly conserved across animal species, which has pivotal roles in the regulation of bone induction, maintenance and repair (Hogan 1996). BMPs act through an autocrine or paracrine mechanism by binding to cell surface receptors and initiating a cascading of downstream cell signaling events that have multiple effects on the formation of bone and cartilage (Sykaras and Opperman 2003). To date, thirteen more BMPs have been discovered, probably bringing the total to around twenty (Even et al. 2012), in which the BMP2 is one of the most documented members because of its functional importance and clinical significance (Agrawal and Sinha 2017). Over the past decades, accumulated evidences have shown that BMP2 is a central regulator of bone defects, non-union fractures, spinal fusion, osteoporosis and root canal surgery through induction of cartilage and bone cells (Geiger et al. 2003; Khan and Lane 2004), which also exhibits therapeutic benefits for a variety of bone diseases such as osteoporosis (Segredo-Morales et al. 2018), multiple myeloma (Seher et al. 2017), and osteonecrosis of femoral head (Vandermeer et al. 2011).
In recent years, BMP2-targeted therapy has been established as a potential and attractive strategy to modulate the bone-related TGFβ/BMP signaling at molecular level (Wu et al. 2011; Peng et al. 2018). Several forms of BMP2 exist, that is, a mature active 30 kDa homodimer, an N-terminal propeptide of 40–45 kDa, and a small amount of 60 kDa precursor protein (Israel et al. 1992). Proteolytic hydrolysis of the precursor protein produces variable-length propeptides, which can be further cleaved to the mature homodimer. The mature BMP2 dimer has a large hydrophobic surface exposed to solvent, contributing to its unusually low solubility in aqueous solution (Vallejo and Rinas 2013). The BMP2 monomer can also form a series of functionally active heterodimers with other members of this family, such as BMP2/6 and BMP2/7 (Valera et al. 2010; Morimoto et al. 2015). Like other growth factors such as EGF, FGF and HGF, the BMP2 is hard to be targeted by small-molecule chemical drugs due to its small size and smooth surface. Previously, biological agents such as monoclonal antibodies (Moshaverinia et al. 2013) and peptide ligands (Zhu et al. 2017) have been successfully applied to suppress the biological activity of several TGFβ/BMP family proteins.
Peptides possess many attractive features when compared to small molecule and protein therapeutics, such as high structural compatibility with target proteins and the ability to disrupt protein–protein interaction interface. This kind of biologics is also recognized for being highly selective and efficacious and, at the same time, relatively safe and well tolerated (Fosgerau and Hoffmann 2015). However, efficient development of high-affinity peptide ligands that can specifically target disease-related proteins has been a major obstacle to the development of this potential drug class (Vanhee et al. 2011; Ren et al. 2011). In recent years, computational peptidology has been recognized as a new and promising strategy to rationally design bioactive peptides (Zhou et al. 2013a, b; Li et al. 2019a). Previously, Song et al. have successfully grafted, striped and stapled of several helical peptides from the dimerization interface of BMP2 (Song et al. 2019). Here, we attempt to rationally design self-binding cyclic peptides (Yang et al. 2015a, 2016; Li et al. 2019b) based on the crystal complex structure of BMP2 with its type-IA receptor, which are expected to disrupt BMP2–receptor complex interaction by rebinding to their native sites at the complex interface. Here, a structure-based strategy that integrated computational analysis and experimental assay was described to dissect the high-resolution crystal structure of BMP2–receptor complex. A peptide segment was derived from the hotspot region of receptor protein, which was then extended, optimized and cyclized via a rational approach, aiming to improve its affinity and specificity for BMP2. The structural basis and energetic property underlying the intermolecular interaction between BMP2 and the designed peptides were also investigated systematically.
2 Materials and methods
2.1 Crystal structure of BMP2 in complex with its type-IA receptor
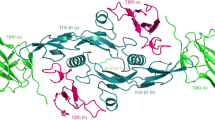
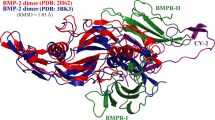
The 1.8 Å-resolution crystal structure of human BMP2 in complex with its cognate receptor was solved by Keller and co-workers using X-ray crystallography; the structure can be retrieved from the PDB database with accession code 1REW (Keller et al. 2004). The structure is a quaternate complex that contains a disulfide-bridged BMP2 homodimer bound with the extracellular domains of two type-IA receptors. As can be seen in figure 1, the two receptors are spatially separated from each other and can bind to two remote regions on BMP2 protein surface in an independent manner, suggesting no intermolecular interactions between them. Here, the cocrystallized water molecules, ions and cofactors were manually removed from the raw crystal structure (Luo et al. 2015).
2.2 Computational simulation and binding analysis
Molecular dynamics (MD) simulations of the investigated system of protein, peptide or their complex were carried out using AMBER ff03 force field (Duan et al. 2003). The system was immersed into an octahedral TIP3P water box (Jorgensen et al. 1983) with a 10 Å buffer extension from the solute. Counterions were added to make the system electroneutral. The system was heated from 0 to 300 K over 100 ps and then equilibrated for 500 ps (Yang et al. 2015b; Zhou et al. 2016). Next, 50–120-ns MD production simulations were performed with periodic boundary conditions (Bai et al. 2017; Zhou et al. 2018). A time step of 2 fs was set and the particle mesh Ewald (PME) method (Darden et al. 1993) was employed to calculate long-range electrostatic interactions for the simulations. A cut-off distance of 10 Å was used to calculate the short-range electrostatics and van der Waals interactions.
Structural snapshots were collected from the dynamics trajectory of production simulations and then employed to analyze the complex binding energetics by using molecular mechanics/Poisson-Boltzmann surface area (MM/PBSA) method (Tian et al. 2011, 2013). The method calculated the complex interaction energy ΔEint and the desolvation free energy ΔDdsv upon the complex binding using molecular mechanics (MM) approach and finite-difference solution of implicit solvent model (PBSA), respectively (Homeyer and Gohlke 2012). If the complex is formed by a BMP2–peptide interaction, conformational flexibility of the peptide ligand was dissected with normal mode analysis (NMA) to estimate entropy penalty –TΔS upon the peptide binding. Consequently, the total binding free energy ΔGttl can be expressed as follows (Wu et al. 2018):
or
where <···> represents average over the collected conformational snapshots and i corresponds to the ith snapshot of the complex.
2.3 Peptide affinity assay
Two linear peptides (RPS2[r78−94]: Ac-78MKYEGSDFQCKDSPKAQ94-NH2 and RPS2[r74−98]: Ac-74ASGCMKYEGSDFQCKDSPKAQLRRT98-NH2) and a head-to-tail cyclic peptide (cycRPS2[r74−98]: cyc[74ASGCMKYEGSDFQCKDSPKAQLRRT98]) were synthesized by Fmoc solid phase chemistry. The human BMP2 protein is natively in homodimer form stabilized by a disulfide bond (Cys78-Cys78) across two monomers. The fluorescence polarization (FP) assays were performed at 298 K using a protocol modified from previous reports (Tyler et al. 2010; Hu et al. 2017). Titrations were conducted by monitoring FP as a function of increasing amounts of BMP2 proteins added to 10 μM FITC-labeled peptides in a buffer containing 50 mM Tris-HCl, 100 mM NaCl and 5 mM EDTA. No DTT was added to avoid the reduction of BMP2 disulfide bond. Each assay was performed in triplicate.
3 Results and discussion
3.1 Derivation of linear peptide segments
The quaternate complex system of human BMP2 homodimer with the extracellular domains of its two type-IA receptors was subjected to 50-ns MD simulations. The dynamics trajectory indicated that the system can reach stable state after ~ 10 ns simulations. Subsequently, computational alanine scanning (Kortemme et al. 2004) was carried out to determine the residue importance of receptor binding to BMP2 based on analysis of the last 40-ns dynamics equilibrium trajectory. The scanning strategy separately mutated each residue of receptor protein to neutral alanine and then calculated change in total binding free energy ΔΔGttl upon the mutation. The resultant ΔΔGttl values can be used to measure the relative contribution of each residue in the receptor to its binding capability for BMP2; the favorable and unfavorable residues in the binding can be indicated by ΔΔGttl>0 and <0, respectively.
As can be seen in figure 2, most residues of receptor protein are favorable for BMP2 binding (ΔΔGttl>0). This is expected if considering that the sequence and structure of naturally evolved type-IA receptor have already been optimized to be well compatible with its cognate partner BMP2; the residue mutation would impair the compatibility and therefore cause unfavorable effect on the binding. Most mutations can only affect the BMP2–receptor interaction moderately or modestly, with ΔΔGttl<0.5 kcal/mol. Structural examination revealed that a wide contact interface can be observed in the interaction complex, at which a number of receptor residues are tightly packed against BMP2 active site, presenting two hotspot regions of residue ranges 43–64 and 78–94 as characterized by the alanine scanning. The two regions separately correspond to a β-strand/loop segment (peptide segment 1) and a α-helix/loop segment (peptide segment 2) in receptor protein, which can directly interact with BMP2 to confer strong binding affinity to the complex system. As shown in figure 2, the residues with effective favorable contribution in segment 1 are discontinuous; some residues in the region such as 49–52 and 59 contribute very modestly to the receptor binding (ΔΔGttl<0.3 kcal/mol). In addition, as compared to segment 2 the segment 1 is longer, more flexible and less residue contribution. Therefore, we herein only considered the segment 2 as a potential peptide candidate that is expected to competitively bind at its native site in the complex interface to target and disrupt the complex interaction.
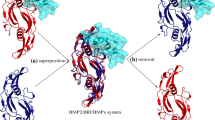
The Receptor Peptide Segment 2 (RPS2[r78−94]) is a 17-mer peptide (78MKYEGSDFQCKDSPKAQ94) that covers the hotspot region 2 of type-IA receptor and can be stripped from the crystal structure of the complex system (figure 3A). The RPS2[r78−94] peptide was then subjected to 120-ns MD simulations, during which the peptide conformation changed dramatically and, particularly, its local α-helical structure was totally disappeared, indicating that the peptide is highly flexible and possesses a large intrinsic disorder in isolated state (figure 3B). The total binding free energy of RPS2[r78−94] peptide to BMP2 homodimer was calculated as ΔGttl = 3.6 kcal/mol by using computational energetics analysis, indicating that the peptide cannot bind to BMP2 in a spontaneous manner. In order to elucidate the binding energetics, we have decomposed the total free energy ΔGttl into intermolecular interaction energy ΔEint, desolvation effect ΔDdsv and entropy penalty –TΔS (table 1). According to the decomposition the interaction energy is very favorable (ΔEint = –137.8 kcal/mol), which, however, is largely counteracted by the unfavorable desolvation effect (ΔDdsv = 98.2 kcal/mol) and entropy penalty (–TΔS = 43.2 kcal/mol). Consequently, the isolated RPS2[r78−94] peptide cannot bind to BMP2 effectively due to its high hydrophilicity and large flexibility without protein context support (Zhou et al. 2019). In order to substantiate this computational finding, the RPS2[r78−94] peptide (with N- and C-termini capped by –Ac and –NH2, respectively) was synthesized and purified commercially, and its binding affinity to the recombinant protein of human BMP2 homodimer was measured using FP assays. Consequently, no affinity can be observed (Kd = n.d.), suggesting that the linear peptide is a nonbinder of BMP2 as predicted by in silico calculations.
Crystal structure of BMP2–receptor complex (PDB: 1REW). (A) The segment is stripped from the complex system to derive a linear RPS2[r78−94] peptide. (B) The RPS2[r78−94] peptide is subjected to MD simulation, during which the peptide exhibits a large intrinsic disorder in isolated state. (C) The RPS2[r78−94] peptide is extended separately at its N- and C-termini to derive a longer linear peptide RPS2[r74−98]. (D) The RPS2[r74−98] peptide is then cyclized from its C- to N-termini to obtain a head-to-tail cyclic peptide cycRPS2[r74−98]. During MD simulations the cyclic peptide shows a larger rigidity and smaller disorder, which can well maintain in a conformation similar with its active conformation in complex crystal structure.
3.2 Extension and cyclization of RPS2[r78−94] peptide
Previously, Yu et al. found that entropy penalty has become the main content of the indirect readout energy in protein–peptide recognition instead of deformation energy as the major source of the indirect readout energy in classical biomolecular binding phenomena (Yu et al. 2014). Here, the entropy contribution can bring as much as 43.2 kcal/mol of unfavorable energetic effect to the total binding free energy of linear RPS2[r78−94] peptide. This is because, as MD simulations suggested, this linear peptide is highly flexible in isolated state and thus its conformational degrees of freedom would be reduced upon binding to BMP2, incurring considerable entropy penalty for the binding. By visually examining the native conformation of peptide segment 2 in the complex interface of intact BMP2–receptor crystal structure, it is revealed that the segment can form a partial cycle that is fixed by two β-strand arms on receptor protein surface. A further examination found that the N-terminal residue Ala74 and C-terminal residue Thr98 of the two arms are spatially vicinal to each other. Therefore, we considered to cyclize the peptide in a head-to-tail manner by covalently bonding the free amino and carboxyl groups of the two spatially vicinal residues.
As shown in figure 3C, the RPS2[r78−94] peptide is extended from its two termini until separately reaching the residues Ala74 and Thr98, resulting in a longer linear peptide RPS2[r74−98] (74ASGCMKYEGSDFQCKDSPKAQLRRT98). The amino and carboxyl groups of Ala74 and Thr98 is distanced by ~ 4.5 Å in crystal structure, and we performed force-directed MD simulations with an additional restraint (force constant = 10 kcal/mol/Å2) between them to gradually draw the two residues closer and link them together to model a head-to-tail cyclic counterpart cycRPS2[r74−98], (cyc[74ASGCMKYEGSDFQCKDSPKAQLRRT98]) of the linear RPS2[r78−94] peptide. After ~ 120-ns simulations the distance between two groups fluctuates around the standard length (~ 1.32 Å) of peptide bond in biological context (Crisma et al. 2015) and, during the simulations, the whole cyclic peptide system showed a larger rigidity and smaller disorder, which can well maintain in a conformation similar with its active conformation in complex crystal structure (figure 3D). As seen in Table 1, the total binding free energy of cycRPS2[r74−98] peptide was calculated as a negative value of ΔGttl = –7.2 kcal/mol, indicating that the cyclic peptide can spontaneously bind to BMP2 protein. In contrast, the RPS2[r74−98] peptide, the linear counterpart of cycRPS2[r74−98], has a positive value of ΔGttl = 6.8 kcal/mol, suggesting that the cyclization can restore the peptide potency from unbinding to binding. In fact, the linear RPS2[r74−98] peptide can interact tightly with BMP2 (ΔEint = –146.2 kcal/mol). However, strong entropy penalty (–TΔS = 52.7 kcal/mol) would considerably impair the favorable interaction. Although the cyclic peptide cycRPS2[r74−98] has only a lower interaction energy (ΔEint = –128.4 kcal/mol) as compared to the linear peptide (ΔEint = –146.2 kcal/mol), entropy penalty of the cyclic peptide is also reduced significantly (–TΔS = 25.8 kcal/mol) upon the cyclization. In addition, the desolvation effect seems not to be influenced substantially by the cyclization (ΔDdsv changes from 100.3 to 95.4 kcal/mol). Consequently, the cycRPS2[r74−98] peptide obtains a favorable binding energy (ΔGttl = –7.2 kcal/mol) — this can be confirmed by FP assays, that is, no binding affinity was detected for linear RPS2[r74−98] peptide (Kd = n.d.), while a moderate affinity was observed for cyclic cycRPS2[r74−98] peptide binding to BMP2 (Kd = 56 ± 8 μM).
References
Agrawal V and Sinha M 2017 A review on carrier systems for bone morphogenetic protein-2. J. Biomed. Mater Res. B Appl. Biomater. 105 904–925
Bai Z, Hou S, Zhang S, Li Z and Zhou P 2017 Targeting self-binding peptides as a novel strategy to regulate protein activity and function: a case study on the proto-oncogene tyrosine protein kinase c-Src. J. Chem. Inf. Model. 57 835–845
Chen D, Zhao M and Mundy GR 2004 Bone morphogenetic proteins. Growth Factors 22 233–241
Crisma M, Formaggio F and Toniolo C 2015 Insight into peptide bond formation from 3D-structural chemistry. Amino Acids, Peptides and Proteins 40 1–35
Darden T, York D and Pedersen L 1993 Particale mesh Ewald and N.log(N) method for Ewald sums in large systems. J. Chem. Phys. 98 10089–10092
Duan Y, Wu C, Chowdhury SS, Lee MC, Xiong GM, Zhang W, Yang R, Cieplak P, Luo R, Lee TS, Caldwell J, Wang JM and Kollman P 2003 A point-charge force field for molecular mechanics simulations of proteins. J. Comput. Chem. 24 1999–2012
Even J, Eskander M and Kang J 2012 Bone morphogenetic protein in spine surgery: current and future uses. J. Am. Acad. Orthop. Surg. 20 547–552
Fosgerau K and Hoffmann T 2015 Peptide therapeutics: current status and future directions. Drug Discov. Today 20 122–128
Geiger M, Li RH and Friess W 2003 Collagen sponges for bone regeneration with rhBMP-2. Adv. Drug Deliv. Rev. 55 1613–1629
Hogan BL 1996 Bone morphogenetic proteins: multifunctional regulators of vertebrate development. Genes Dev. 10 1580–1594
Homeyer N and Gohlke H 2012 Free energy calculations by the molecular mechanics Poisson-Boltzmann surface area method. Mol. Inf. 31 114–122
Hu H, Yang S, Zheng J and Mao G 2017 Structure-based derivation of peptide inhibitors to target TGF-β1 receptor for the suppression of hypertrophic scarring fibroblast activation. Chem. Biol. Drug Des. 90 345–351
Israel DI, Nove J, Kerns KM, Moutsatsos IK and Kaufman RJ 1992 Expression and characterization of bone morphogenetic protein-2 in Chinese hamster ovary cells. Growth Factors 7 139–150
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW and Klein ML 1983 Comparison of simple potential functions for simulating liquid water. J. Phys. Chem. 79 926–935
Kortemme T, Kim DE and Baker D 2004 Computational alanine scanning of protein-protein interfaces. Sci STKE 2004 pl2
Li Z, Miao Q, Yan F, Meng Y and Zhou P 2019a Machine learning in quantitative protein-peptide affinity prediction: implications for therapeutic peptide design. Curr. Drug Metab. 20 170–176
Li Z, Yan F, Miao Q, Meng Y, Wen L, Jiang Q and Zhou P 2019b Self-binding peptides: binding-upon-folding versus folding-upon-binding. J. Theor. Biol. 469 25–34
Luo H, Du T, Zhou P, Yang L, Mei H, Ng H, Zhang W, Shu M, Tong W, Shi L, Mendrick DL and Hong H 2015 Molecular docking to identify associations between drugs and class I human leukocyte antigens for predicting idiosyncratic drug reactions. Comb. Chem. High. Throughput Screen. 18 296–304
Keller S, Nickel J, Zhang JL, Sebald W and Mueller TD 2004 Molecular recognition of BMP-2 and BMP receptor IA. Nat. Struct. Mol. Biol. 11 481–488
Khan SN and Lane JM 2004 The use of recombinant human bone morphogenetic protein-2 (rhBMP-2) in orthopaedic applications. Expert. Opin. Biol. Ther. 4 741–748
Morimoto T, Kaito T, Matsuo Y, Sugiura T, Kashii M, Makino T, Iwasaki M and Yoshikawa H 2015 The bone morphogenetic protein-2/7 heterodimer is a stronger inducer of bone regeneration than the individual homodimers in a rat spinal fusion model. Spine J. 15 1379–1390
Moshaverinia A, Ansari S, Chen C, Xu X, Akiyama K, Snead ML, Zadeh HH and Shi S 2013 Co-encapsulation of anti-BMP2 monoclonal antibody and mesenchymal stem cells in alginate microspheres for bone tissue engineering. Biomaterials 34 6572–6579
Peng H, Lu SL, Bai Y, Fang X, Huang H and Zhuang XQ 2018 MiR-133a inhibits fracture healing via targeting RUNX2/BMP2. Eur. Rev. Med. Pharmacol. Sci. 22, 2519–2526
Ren Y, Chen X, Feng M, Wang Q and Zhou P 2011 Gaussian process: a promising approach for the modeling and prediction of peptide binding affinity to MHC proteins. Protein Pept Lett. 18 670–678
Segredo-Morales E, García-García P, Reyes R, Pérez-Herrero E, Delgado A and Évora C 2018 Bone regeneration in osteoporosis by delivery BMP-2 and PRGF from tetronic-alginate composite thermogel. Int. J. Pharm. 543 160–168
Seher A, Lagler C, Stühmer T, Müller-Richter UDA, Kübler AC, Sebald W, Müller TD, Nickel J 2017 Utilizing BMP-2 muteins for treatment of multiple myeloma. PLoS ONE 12 e0174884
Song W, Wang K, Wang W, Yang P and Dang X 2019 Grafting, stripping and stapling of helical peptides from the dimerization interface of ONFH-related bone morphogenetic protein-2. Protein J. 38 12–22
Sykaras N and Opperman LA 2003 Bone morphogenetic proteins (BMPs): how do they function and what can they offer the clinician? J. Oral. Sci. 45 57–73
Tian F, Lv Y, Zhou P and Yang L 2011 Characterization of PDZ domain-peptide interactions using an integrated protocol of QM/MM, PB/SA, and CFEA analyses. J. Comput. Aided Mol. Des. 25 947–958
Tian F, Tan R, Guo T, Zhou P and Yang L 2013 Fast and reliable prediction of domain–peptide binding affinity using coarse-grained structure models. Biosystems 113 40–49
Tyler RC, Peterson FC and Volkman BF 2010 Distal interactions within the par3-VE-cadherin complex. Biochemistry 49 951–957
Vallejo LF and Rinas U 2013 Folding and dimerization kinetics of bone morphogenetic protein-2, a member of the transforming growth factor-β family. FEBS J. 280 83–92
Valera E, Isaacs MJ, Kawakami Y, Izpisúa Belmonte JC and Choe S 2010 BMP-2/6 heterodimer is more effective than BMP-2 or BMP-6 homodimers as inductor of differentiation of human embryonic stem cells. PLoS ONE 5 e11167
Vandermeer JS, Kamiya N, Aya-ay J, Garces A, Browne R and Kim HK 2011 Local administration of ibandronate and bone morphogenetic protein-2 after ischemic osteonecrosis of the immature femoral head: a combined therapy that stimulates bone formation and decreases femoral head deformity. J. Bone Joint Surg. Am. 93 905–913
Vanhee P, van der Sloot AM, Verschueren E, Serrano L, Rousseau F and Schymkowitz J 2011 Computational design of peptide ligands. Trends Biotechnol. 29 231–239
Wu JB, Fu HQ, Huang LZ, Liu AW and Zhang JX 2011 Effects of siRNA-targeting BMP-2 on the abilities of migration and invasion of human liver cancer SMMC7721 cells and its mechanism. Cancer Gene Ther. 18 20–25
Wu T, He P, Wu W, Chen Y and Lv F 2018 Targeting oncogenic transcriptional corepressor Nac1 POZ domain with conformationally constrained peptides by cyclization and stapling. Bioorg. Chem. 80 1–10
Yang C, Zhang S, He P, Wang C, Huang J and Zhou P 2015a Self-binding peptides: folding or binding? J. Chem. Inf. Model. 55 329–342
Yang C, Wang C, Zhang S, Huang J and Zhou P 2015b Structural and energetic insights into the intermolecular interaction among human leukocyte antigens, clinical hypersensitive drugs and antigenic peptides. Mol. Simul. 41 741–751
Yang C, Zhang S, Bai Z, Hou S, Wu D, Huang J and Zhou P 2016 A two-step binding mechanism for the self-binding peptide recognition of target domains. Mol. Biosyst. 12 1201–1213
Yu H, Zhou P, Deng M and Shang Z 2014 Indirect readout in protein–peptide recognition: a different story from classical biomolecular recognition. J. Chem. Inf. Model. 54 2022–2032
Zhou P, Yang C, Ren Y, Wang C and Tian F 2013 What are the ideal properties for functional food peptides with antihypertensive effect? A computational peptidology approach. Food Chem. 141 2967–2973
Zhou P, Wang C, Tian F, Ren Y, Yang C and Huang J 2013 Biomacromolecular quantitative structure-activity relationship (BioQSAR): a proof-of-concept study on the modeling, prediction and interpretation of protein-protein binding affinity. J. Comput. Aided Mol. Des. 27 67–78
Zhou P, Zhang S, Wang Y, Yang C and Huang J 2016 Structural modeling of HLA-B*1502/peptide/carbamazepine/T-cell receptor complex architecture: implication for the molecular mechanism of carbamazepine-induced Stevens-Johnson syndrome/toxic epidermal necrolysis. J. Biomol. Struct. Dyn. 34 1806–1817
Zhou P, Hou S, Bai Z, Li Z, Wang H, Chen Z and Meng Y 2018 Disrupting the intramolecular interaction between proto-oncogene c-Src SH3 domain and its self-binding peptide PPII with rationally designed peptide ligands. Artif. Cells Nanomed. Biotechnol. 46 1122–1131
Zhou P, Miao Q, Yan F, Li Z, Jiang Q, Wen L and Meng Y 2019 Is protein context responsible for peptide-mediated interactions? Mol. Omics (https://doi.org/10.1039/c9mo00041k)
Zhu Z, Zhang C, Song W 2017 Rational derivation, extension, and cyclization of self-inhibitory peptides to target TGF-β/BMP signaling in ONFH. Amino Acids 49 283–290
Acknowledgements
This work was supported by the YCH Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Ravindra Venkatramani
Rights and permissions
About this article
Cite this article
Fan, X., Xia, H., Liu, X. et al. Rational design of type-IA receptor-derived cyclic peptides to target human bone morphogenic protein 2. J Biosci 44, 130 (2019). https://doi.org/10.1007/s12038-019-9945-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-019-9945-8