Abstract
Recent genome-wide association studies (GWAS) reported CR1 rs3818361 polymorphism to be an Alzheimer’s disease (AD) susceptibility variant in European ancestry. Three independent studies investigated this association in Chinese population. However, these studies reported weak or no significant association. Here, we reinvestigated the association using all the samples from three independent studies in Chinese population (N = 4047, 1244 AD cases and 2803 controls). We also selected three independent studies in European ancestry population (N = 11787, 3939 AD cases and 7848 controls) to evaluate the effect of rs3818361 polymorphism on AD risk in different ethnic backgrounds. In Chinese population, we did not identified significant heterogeneity using additive, recessive, and dominant genetic models. Meta-analysis showed significant association between rs3818361 and AD with P = 6.00E-03 and P = 5.00E-03. We further identified no heterogeneity of rs3818361 polymorphism between Chinese and European populations. We found that rs3818361 polymorphism contributed to AD with similar genetic risk in Chinese and European populations. In summary, this is the first study to show significant association between rs3818361 polymorphism and AD in Chinese population by a meta-analysis method. Our findings indicate that the effect of CR1 rs3818361 polymorphism on AD risk in Chinese cohorts is consistent with the increased risk observed in European AD cohorts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since 2009–2013, large-scale genome-wide association studies (GWAS) reported several Alzheimer’s disease (AD) susceptibility genes including CR1, BIN1, CLU, PICALM, MS4A4/MS4A6E, CD2AP, CD33, EPHA1, and ABCA7 [1–3], SORL1 [4], and TREM2 [5]. A recent large meta-analysis of AD GWAS in individuals of European ancestry identified 11 new AD genetic risk factors for AD, which include HLA-DRB5/DRB1, PTK2B, SLC24A4-0RING3, DSG2, INPP5D, MEF2C, NME8, ZCWPW1, CELF1, FERMT2, and CASS4 [3].
We previously analyzed the PICALM rs3851179 [6, 7], BIN1 rs744373 [8], CLU rs11136000 [1], rs2279590 [9], and rs9331888 [10], CD2AP rs9349407 [11], ABCA7 rs3764650 [12], and CD33 rs3865444 [13] polymorphisms, and identified significant association. In addition to these polymorphisms above, a single nucleotide polymorphism (SNP) rs3818361 in CR1 was reported to be significantly associated with AD in European ancestry with P = 8.5E-08, P = 9.2E-06, and P = 3.7E-14 [14–16]. The following studies confirmed the influence of CR1 on clinical and pathological measures of AD [17–20]. Three candidate gene studies investigated rs3818361 polymorphism in Chinese population [21–23]. However, one study reported weak association (P = 0.029 for allele test and P = 0.047 for genotype test [22]) and two reported no association (P = 0.07 for allele test and P = 0.18 for genotype test [21], P = 0.07 and P = 0.26 [23]) between rs3818361 and AD in Chinese population.
It is reported that meta-analysis method combines and analyzes quantitative evidence from related studies to produce results based on a whole body of research to aggregate information in order to achieve a higher statistical power [24]. Here, we reinvestigated the association using all the samples from three independent studies in Chinese population. We also selected three independent studies in European ancestry population to evaluate the effect of rs3818361 polymorphism on AD risk in different ethnic backgrounds.
Methods and Materials
Samples
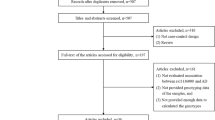
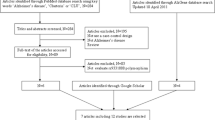
Our study included 4047 samples (1244 AD cases and 2803 controls) from previous three independent studies in Chinese population and 11787 samples (3939 AD cases and 7848 controls) from previous three independent studies in European ancestry population (Table 1).
Genetic Models
Both allele and genotype models were used to investigate the association between rs3818361 and AD. The CR1 rs3818361 polymorphism includes two alleles C and T. T is the minor allele. We assume that T is the high-risk allele and C is the lower-risk allele. We selected the additive (T allele versus C allele), recessive (TT genotype versus TC + CC genotypes), and dominant (TT + TC genotypes versus CC genotype) genetic models [25].
Quality Evaluation
Quality evaluation criteria were used to evaluate the quality of selected genetic association studies [26], which consist of ten components. Each component is scored as 1 if present or 0 if absent. The quality evaluation score was calculated by summing each component, resulting in a scoring range of 0–10 [26]. The selected studies were scored as “good” if the score was greater than or equal to 8, “mediocre” if the score was 5–7, and “poor” if the score was less than 4 [27]. Two authors performed the quality evaluation independently using the criteria proposed by Clark et al. A third author adjudicated any differences between the two authors as described in our previous study [12].
Heterogeneity Test
Genetic heterogeneity among the selected studies is evaluated using Cochran’s Q test and \( {I}^2=\raisebox{1ex}{$\left(Q-\left(k-1\right)\right)$}\!\left/ \!\raisebox{-1ex}{$Q$}\right.\times 100\% \) statistic. Cochran’s Q test approximately follows a χ 2 distribution with k − 1 degrees of freedom (k stands for the number of studies for analysis). I 2 ranges from 0 to 100 %, 0–24 % = no heterogeneity, 25–49 % = moderate heterogeneity, 50–74 % =large heterogeneity, and 75–100 % = extreme heterogeneity. P < 0.01 from Cochran’s Q test and I 2 > 50 % was considered to indicate statistically significant heterogeneity.
Meta-analysis
The pooled odds ratio (OR) is calculated by the fixed effect model (Mantel-Haenszel). Z test is used to determine the significance of OR. All statistical tests were computed using RevMan (v.5.1).
Sensitivity Analysis
We omit each study at a time to assess the influence of each individual study on the pooled OR.
Publication Bias Analysis
A funnel plot from Egger et al. is commonly used to check for the existence of publication bias [28, 29]. A regression-based approach proposed by Egger is used to test for publication bias to provide statistical evidence, with a P < 0.01 indicating that there was a significant publication bias [30]. All statistical tests were computed using R.
Results
Heterogeneity Test and Subgroup Meta-analysis
We first performed a subgroup analysis in Chinese and European populations, respectively. In Chinese population, we did not identify significant heterogeneity using additive (P = 0.41 and I 2 = 0 %), recessive (P = 0.84 and I 2 = 0 %), and dominant (P = 0.32 and I 2 = 13 %) genetic models. Meta-analysis showed significant association between rs3818361 and AD with P = 6.00E-03 and P = 5.00E-03 for additive and recessive genetic models but suggestive association for dominant genetic model with P = 5.00E-02.
In European population, we did not identify significant heterogeneity using additive (P = 0.87 and I 2 = 0 %), recessive (P = 0.81 and I 2 = 0 %), and dominant (P = 0.82 and I 2 = 13 %) genetic models. Meta-analysis showed significant association between rs3818361 and AD with P = 1.36E-05 and P = 1.24E-05 for additive and dominant genetic models but no association for recessive genetic model with P = 7.00E-02. More detailed results were described in Figs. 1, 2, and 3.
We further found that rs3818361 polymorphism contributed to AD with similar genetic risk in Chinese and European populations with OR = 1.16 and 1.16, OR = 1.36 and 1.19, and OR = 1.15 and 1.20 for additive, recessive, and dominant genetic models. More detailed results are described in Figs. 1, 2, and 3.
Heterogeneity Test and Meta-analysis in Pooled Population
We further evaluated the genetic heterogeneity of rs3818361 polymorphism between Chinese and European populations. We did not identify significant heterogeneity using additive (P = 0.84 and I 2 = 0 %), recessive (P = 0.91 and I 2 = 0 %), and dominant (P = 0.71 and I 2 = 0 %) genetic models. Meta-analysis showed significant association between rs3818361 and AD with P = 2.47E-07, P = 1.00E-03, and P = 2.03E-06 for additive, recessive, and dominant genetic models. More detailed results were described in Figs. 1, 2, and 3.
Sensitivity Analysis
We identified that the association between rs3818361 polymorphism and AD did not vary substantially, which suggested that the results from this meta-analysis were stable.
Publication Bias Analysis
The funnel plots of the selected studies investigating the association between rs3818361 and AD are symmetrical inverted funnels (Fig. 4). Egger’s test provides statistical evidence of symmetry with P = 0.637, P = 0.235, and P = 0.977 for additive, recessive, and dominant genetic models, respectively. These results indicated no evidence of publication bias.
Discussion
Original GWAS identified significant association between rs3818361 and AD in European ancestry [14–16]. Three independent studies investigated the rs3818361 polymorphism in Chinese population and reported a weak or negligible association between rs3818361 and AD. Growing evidence confirmed the influence of CR1 on clinical and pathological measures of AD [17–20]. Considering the important role of CR1 in AD, we reevaluated this association using large-scale samples from six independent studies in Chinese and European populations. Our results showed significant association between rs3818361 polymorphism and AD in Chinese population. We found no heterogeneity of rs3818361 polymorphism between Chinese and European populations.
Recent therapeutic strategies for treating AD mainly focused on reducing brain amyloid burden [18]. The influence of CR1 on clinical and pathological measures of AD cases and controls was also investigated. The results indicated that the higher expression of CR1 was associated with more advanced cognitive decline [19]. Chibnik et al. analyzed 1666 non-demented subjects to evaluate the associations between CR1 rs3818361 and rate of change in cognitive function [20]. The results showed that CR1 was associated with amyloid plaque burden and age-related cognitive decline [20]. Sweet et al. investigated 1831 non-demented subjects from Cardiovascular Health Study to determine the effects of CR1 on age and rate of decline. The results also indicated significant association between CR1 and more rapid cognitive decline [17].
In addition to rs3818361 polymorphism, CR1 rs6656401 polymorphism was also identified to be significantly associated with AD in European ancestry [14, 31]. Evidence showed that the OR was highest for the rs6656401 and rs3818361AA haplotype compared to the GG haplotype (OR = 1.22, P = 3.10E-10) [14]. Lambert et al. recently conducted a large, two-stage meta-analysis of AD GWAS in individuals of European ancestry [3]. They identified CR1 rs6656401 and rs3818361 polymorphisms to be significantly associated with AD reaching genome-wide significance with P = 5.70E-24 and 5.40E-14 [3]. We previously analyzed rs6656401 polymorphism in Asian and European populations. We did not identify significant heterogeneity [32]. We identified significant association between rs6656401 polymorphism and AD with P = 1.82E-26 [32]. We further performed a subgroup analysis in East Asian population. We did not identify significant heterogeneity and found significant association between rs6656401 polymorphism and AD in East Asian population [32].
In addition to the rs6656401 and rs3818361 polymorphisms, we previously analyzed PICALM rs3851179, BIN1 rs744373, and CLU rs11136000 and rs2279590 polymorphisms [1, 6–9]. We found that there was no significant genetic heterogeneity of these polymorphisms in Asian and European populations. We further analyzed relatively large-scale samples and reported significant association between these common variants and AD in Asian populations [1, 6–9].
Our study also has some limitations. In our research, we performed a publication bias analysis. However, it cannot adjust all the biases. Sampling bias and geographical bias analyses may be very helpful. Future studies are required to replicate our findings. In summary, our findings indicate that the effect of CR1 rs3818361 polymorphism on AD risk in Chinese cohorts is consistent with the increased risk observed in European AD cohorts. To our knowledge, this is the first study to show significant association between rs3818361 polymorphism and AD in Chinese population by a meta-analysis method.
References
Liu G, Wang H, Liu J, Li J, Li H, Ma G, Jiang Y, Chen Z et al (2014) The clu gene rs11136000 variant is significantly associated with alzheimer's disease in caucasian and asian populations. Neuromolecular Med 16:52–60
Reitz C, Jun G, Naj A, Rajbhandary R, Vardarajan BN, Wang LS, Valladares O, Lin CF et al (2013) Variants in the atp-binding cassette transporter (abca7), apolipoprotein e 4, and the risk of late-onset alzheimer disease in african americans. JAMA 309:1483–1492
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, DeStafano AL, Bis JC et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for alzheimer's disease. Nat Genet 45:1452–1458
Miyashita A, Koike A, Jun G, Wang LS, Takahashi S, Matsubara E, Kawarabayashi T, Shoji M et al (2013) Sorl1 is genetically associated with late-onset alzheimer's disease in japanese, koreans and caucasians. PLoS One 8:e58618
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E, Cruchaga C, Sassi C et al (2013) Trem2 variants in alzheimer's disease. N Engl J Med 368:117–127
Liu G, Zhang S, Cai Z, Ma G, Zhang L, Jiang Y, Feng R, Liao M et al (2013) Picalm gene rs3851179 polymorphism contributes to alzheimer's disease in an asian population. Neuromolecular Med 15:384–388
Liu G, Zhang L, Feng R, Liao M, Jiang Y, Chen Z, Zhao B, Li K (2013) Lack of association between picalm rs3851179 polymorphism and alzheimer's disease in chinese population and apoeepsilon4-negative subgroup. Neurobiol Aging 34(1310):e1319–1310
Liu G, Zhang S, Cai Z, Li Y, Cui L, Ma G, Jiang Y, Zhang L et al (2013) Bin1 gene rs744373 polymorphism contributes to alzheimer's disease in east asian population. Neurosci Lett 544:47–51
Zhang S, Zhang D, Jiang Y, Wu L, Shang H, Liu J, Feng R, Liao M et al (2015) Clu rs2279590 polymorphism contributes to alzheimer's disease susceptibility in caucasian and asian populations. J Neural Transm 122:433–439
Zhang S, Li X, Ma G, Jiang Y, Liao M, Feng R, Zhang L, Liu J, Wang G, Zhao B, Jiang Q, Li K, Liu G (2015) Clu rs9331888 polymorphism contributes to alzheimer's disease susceptibility in caucasian but not east asian populations. Mol Neurobiol
Chen H, Wu G, Jiang Y, Feng R, Liao M, Zhang L, Ma G, Chen Z, Zhao B, Li K, Yu C, Liu G (2014) Analyzing 54,936 samples supports the association between cd2ap rs9349407 polymorphism and alzheimer's disease susceptibility. Mol Neurobiol
Liu G, Li F, Zhang S, Jiang Y, Ma G, Shang H, Liu J, Feng R et al (2014) Analyzing large-scale samples confirms the association between the abca7 rs3764650 polymorphism and alzheimer's disease susceptibility. Mol Neurobiol 50:757–764
Li X, Shen N, Zhang S, Liu J, Jiang Q, Liao M, Feng R, Zhang L, Wang G, Ma G, Zhou H, Chen Z, Jiang Y, Zhao B, Li K, Liu G (2014) Cd33 rs3865444 polymorphism contributes to alzheimer's disease susceptibility in chinese, european, and north american populations. Mol Neurobiol
Lambert JC, Heath S, Even G, Campion D, Sleegers K, Hiltunen M, Combarros O, Zelenika D et al (2009) Genome-wide association study identifies variants at clu and cr1 associated with alzheimer's disease. Nat Genet 41:1094–1099
Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML, Pahwa JS, Moskvina V et al (2009) Genome-wide association study identifies variants at clu and picalm associated with alzheimer's disease. Nat Genet 41:1088–1093
Hollingworth P, Harold D, Sims R, Gerrish A, Lambert JC, Carrasquillo MM, Abraham R, Hamshere ML et al (2011) Common variants at abca7, ms4a6a/ms4a4e, epha1, cd33 and cd2ap are associated with alzheimer's disease. Nat Genet 43:429–435
Sweet RA, Seltman H, Emanuel JE, Lopez OL, Becker JT, Bis JC, Weamer EA, DeMichele-Sweet MA et al (2012) Effect of alzheimer's disease risk genes on trajectories of cognitive function in the cardiovascular health study. A J Psychiatry 169:954–962
Guan H, Liu Y, Daily A, Police S, Kim MH, Oddo S, LaFerla FM, Pauly JR et al (2009) Peripherally expressed neprilysin reduces brain amyloid burden: a novel approach for treating alzheimer's disease. J Neurosci Res 87:1462–1473
Karch CM, Jeng AT, Nowotny P, Cady J, Cruchaga C, Goate AM (2012) Expression of novel alzheimer's disease risk genes in control and alzheimer's disease brains. PLoS One 7, e50976
Chibnik LB, Shulman JM, Leurgans SE, Schneider JA, Wilson RS, Tran D, Aubin C, Buchman AS et al (2011) Cr1 is associated with amyloid plaque burden and age-related cognitive decline. Ann Neurol 69:560–569
Zhang Q, Yu JT, Zhu QX, Zhang W, Wu ZC, Miao D, Tan L (2010) Complement receptor 1 polymorphisms and risk of late-onset alzheimer's disease. Brain Res 1348:216–221
Chen LH, Kao PY, Fan YH, Ho DT, Chan CS, Yik PY, Ha JC, Chu LW et al (2012) Polymorphisms of cr1, clu and picalm confer susceptibility of alzheimer's disease in a southern chinese population. Neurobiol Aging 33(210):e211–e217
Liao YC, Lee WJ, Hwang JP, Wang YF, Tsai CF, Wang PN, Wang SJ, Fuh JL (2014) Abca7 gene and the risk of alzheimer's disease in han chinese in taiwan. Neurobiol Aging 35(2423):e2427–2423
Riley RD, Lambert PC, Abo-Zaid G (2010) Meta-analysis of individual participant data: rationale, conduct, and reporting. BMJ 340:c221
Lewis CM, Knight J (2012) Introduction to genetic association studies. Cold Spring Harb Protoc 2012:297–306
Clark MF, Baudouin SV (2006) A systematic review of the quality of genetic association studies in human sepsis. Intensive Care Med 32:1706–1712
Lv H, Jiang Y, Li J, Zhang M, Shang Z, Zheng J, Wu X, Liu P, Zhang R, Yu H (2014) Association between polymorphisms in the promoter region of interleukin-10 and susceptibility to inflammatory bowel disease. Mol Biol Rep
Egger M, Davey Smith G, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. BMJ 315:629–634
Sterne JA, Egger M (2001) Funnel plots for detecting bias in meta-analysis: guidelines on choice of axis. J Clin Epidemiol 54:1046–1055
Song F, Khan KS, Dinnes J, Sutton AJ (2002) Asymmetric funnel plots and publication bias in meta-analyses of diagnostic accuracy. Int J Epidemiol 31:88–95
Naj AC, Jun G, Beecham GW, Wang LS, Vardarajan BN, Buros J, Gallins PJ, Buxbaum JD et al (2011) Schellenberg GD: Common variants at ms4a4/ms4a6e, cd2ap, cd33 and epha1 are associated with late-onset alzheimer's disease. Nat Genet 43:436–441
Shen N, Chen B, Jiang Y, Feng R, Liao M, Zhang L, Li F, Ma G, Chen Z, Zhao B, Li K, Liu G (2014) An updated analysis with 85,939 samples confirms the association between cr1 rs6656401 polymorphism and alzheimer's disease. Mol Neurobiol
Acknowledgments
This work was supported by funding from the National Nature Science Foundation of China (Grant No. 81300945).
Conflict of Interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yongning Li and Dongjing Song contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, Y., Song, D., Jiang, Y. et al. CR1 rs3818361 Polymorphism Contributes to Alzheimer’s Disease Susceptibility in Chinese Population. Mol Neurobiol 53, 4054–4059 (2016). https://doi.org/10.1007/s12035-015-9343-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-015-9343-7