Abstract
The main goals of the present study were to screen Iranian common bermudagrasses to find cold-tolerant accessions and evaluate their genetic and morphological variabilities. In this study, 49 accessions were collected from 18 provinces of Iran. One foreign cultivar of common bermudagrass was used as control. Morphological variation was evaluated based on 14 morphological traits to give information about taxonomic position of Iranian common bermudagrass. Data from morphological traits were evaluated to categorize all accessions as either cold sensitive or tolerant using hierarchical clustering with Ward’s method in SPSS software. Inter-Simple Sequence Repeat (ISSR) primers were employed to evaluate genetic variability of accessions. The results of our taxonomic investigation support the existence of two varieties of Cynodon dactylon in Iran: var. dactylon (hairless plant) and var. villosous (plant with hairs at leaf underside and/or upper side surfaces or exterior surfaces of sheath). All 15 primers amplified and gave clear and highly reproducible DNA fragments. In total, 152 fragments were produced, of which 144 (94.73%) being polymorphic. The polymorphic information content (PIC) values ranged from 0.700 to 0.928. The average PIC value obtained with 15 ISSR primers was 0.800, which shows that all primers were informative. Probability identity (PI) and discriminating power between all primers ranged from 0.029 to 0.185 and 0.815 to 0.971, respectively. Genetic data were converted into a binary data matrix. NTSYS software was used for data analysis. Clustering was done by the unweighted pair-group method with arithmetic averages and principle coordinate analysis, separated the accessions into six main clusters. According to both morphological and genetic diversity investigations of accessions, they can be clustered into three groups: cold sensitive, cold semi-tolerant, and cold tolerant. The most cold-tolerant accessions were: Taft, Malayear, Gorgan, Safashahr, Naein, Aligoudarz, and the foreign cultivar. This study may provide useful information for further breeding programs on common bermudagrass. Selected genotypes can be evaluated for other abiotic stresses such as drought and salinity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common bermudagrass (Cynodon dactylon [L.] Pers.) is the most extensive and widely planted warm season turfgrass in southern parts of Iran with subtropical climate. It is a prostrate and perennial grass belonging to the family Poaceae, using C4 carbon fixation pathway [9]. It is well adapted and grows across the tropical, subtropical, and transitional areas of the world distributed between 45°N and 45°S [13, 14]. Common bermudagrass originated from tropical Africa and Eurasia, and some countries in the Middle East such as Iran that is reported to be the center of diversity for this species [13]. Valuable genotypes with useful genes can be found in these countries. Because several biotypes of bermudagrasses are found in different regions of the world [16], potentially great genetic, morphological, and physiological variations exist. Common bermudagrass is tetraploid (2n = 4× = 36) [27], and its 2C nuclear DNA content is 1.95 ± 0.01 pg [5]; however, diploid cultivars (2n = 2× = 18) were reported as well [49].
Common bermudagrass is a major turf species in the transition zone, where they can be used for sports fields, pastures, forage, parks, lawns, roadsides, cemeteries, soil stabilization, remediation, and golf courses and residential and commercial landscapes. They show high tolerance to drought, salinity, wear, and most types of soil [7]. Less tolerance to low and freezing temperatures is one of the main roadblocks to widespread bermudagrass use throughout the world. Generally, common bermudagrass is sensitive to hard winters and has much potential for winterkill by freezing stress in the transitional regions [32]. There is great variability among common bermudagrass accessions in response to cold temperatures. Bermudagrass growth and development slow rapidly when mean daily temperature drops below 15 °C. Leaf senescence begins when temperature declines to 10 °C. The concept of LT50 has been used as a measure of cold hardiness and is defined as the predicted test temperature resulting in 50% or greater loss of total electrolytes [46]. According to Anderson et al. [3, 4], LT50 for many cultivars of bermudagrass ranges from − 5.9 to − 10.3 °C. Ibitayo and Butler [19] reported that the coldest killing temperature for bermudagrass ‘Brookings’ is − 17°C. Previous studies showed that in cold-acclimated plants, significant metabolic alterations take place against freezing stress [50]. Some of these alterations include changes in the levels of sugars, amino acids, proteins, compatible solutes and changes in the degree of fatty acid saturation level and antioxidant capacity that affect the freezing tolerance [24, 35]. Many reports in the literature indicate that freezing tolerance of common bermudagrass [35, 55] and other warm season grasses [8, 45] is attributed to these metabolic alterations. Some of these alterations are similar to drought tolerance, like the level of sugar [28]. There is much variation between natural accessions of common bermudagrass for tolerance to freezing and other environmental stresses. Genetic resources and wild species have important role in plant breeding programs [12]. Today, numerous and extensive wild bermudagrass germplasm collections exist in all over the world with the purpose of common bermudagrass improvement [26]. Traditionally, turfgrass variation researches have focused on phenotypic characteristics [25]. Some molecular methods have been established to significantly reveal natural germplasm diversity of crops [23]. In breeding programs of common bermudagrass, identifying the genetic variability among parental materials is a central task in development of highly stress-tolerant cultivars. One well-known method in studies on genetic variability is using molecular markers [51]. Molecular markers significantly discover polymorphism within the genome of an organism. Different molecular markers have been employed to determine genetic diversity of Cynodon accessions [17]. Some molecular markers have been successfully employed to analyze genetic diversity of bermudagrasses include DAF (DNA Amplification Fingerprinting) [2], RAPD (Random Amplified Polymorphic DNA) [10, 41], AFLP (Amplified Fragment Length Polymorphism) [22, 31, 52], ISSR [29, 33], SSR (Simple Sequence Repeats) [21], and CpSSRLP (Chloroplast-Specific Simple Sequence Repeat Length Polymorphism) [22]. The main advantage of RAPD markers is their technical simplicity, but they have poor consistency and discriminating power. Though the AFLP approach is a useful tool that has high multiplexing ratio, but usually its operation is fairly labor intensive, requires multiple steps, and the costs of reagents and equipments are relatively high. The EST (Expressed Sequence Tag) and SSR markers require knowledge of the flanking regions for designing species-specific primers. Compared with other DNA markers, ISSR is a technique that overcomes most of these restrictions. ISSR markers can detect polymorphism without prior DNA sequence information. Moreover, they can produce more reliable and reproducible bands due to their primer length and the higher annealing temperature. There are many reports proving that ISSR markers are useful tools for genome separation in many plants such as Miscanthus [15], date palms [54], mangrove [20], honeysuckle [48], pepper [11], and tall fescue [43]. Common bermudagrass is widely distributed in Iran. The extensive germplasm available for the species suggests the presence of great diversity in Iranian common bermudagrass accessions. Despite the widespread occurrence of wild bermudagrasses in Iran, little information is available on their diversity within Iranian bermudagrass germplasm pool. The present investigation was undertaken to check genetic diversity among 49 Iranian wild common bermudagrass accessions collected from 18 provinces along with one foreign commercial cultivar (as control) by ISSR markers, and evaluation of morphological characteristics to find accessions with good freezing tolerance.
Materials and Methods
Plant Materials
Forty-nine accessions of natural bermudagrass were collected from 18 provinces of Iran ranging from Shiraz, a city with subtropical climate to around Tabriz, with the temperate climatic condition (Table 1) along with one foreign cultivar (California origin) as control. Based on phenotypic traits described by Harlan and de Wet [13], Harlan [14], all collected samples were evaluated morphologically to be common bermudagrass. Originally, most of the accessions were collected from roadsides and transferred to the School of Agriculture, Shiraz University. They were kept in normal greenhouse condition. Turfgrasses were hand-clipped to a canopy height of 5 cm every 2 weeks.
Evaluation of Morphological Characteristics
Accessions were evaluated with 14 qualitative and quantitative morphological characters (Table 2). In qualitative morphological traits with multistate data, each state was converted into a binary (absent/present) character. Data were analyzed using NTSYS-pc 2.02 software [40]. Cluster analysis was done using the unweighted pair-group method with arithmetic mean (UPGMA).
For selecting cold-tolerant accessions, each accession was transplanted into 14 cm pots under normal greenhouse condition. The results of the evaluation of physiological characteristics are presented in another paper by Akbari and Salehi [1]. They transferred the pots to a growth chamber. Growth chamber was adjusted at 24/17, 7.5/0, − 7.5/− 12, and − 15/− 15 °C day/night cycles and a 10 h photoperiod under a light intensity of 300 µmol m−2 s−1. After 7 days for each temperature regime, fresh leaves and stolon sampling were done to assess their physiological traits including: SOD (Super Oxide Dismutase), CAT (Catalase), APX (Ascorbate Peroxidase) and POD (Peroxidase) activities, proline, electrolyte leakage (EL), and chlorophyll content. Turfgrass color and visual quality were measured after each treatment on a 1–9 scale, where 1 was the turf with very poor quality, 6 was minimally acceptable turf, and 9 was exceptional turf quality. In the present investigation, morphological data were evaluated for accession clustering as either cold tolerant or cold sensitive using Ward’s method of hierarchical cluster analysis in SPSS software.
ISSR Analysis
Two grams of fresh and healthy leaf tissue samples of each accession was detached for total DNA extraction. Harvested tissues got frozen in liquid nitrogen and ground to a fine powder with a sterile mortar and pestle. Genomic DNA isolation was performed using the EZ-10 spin column genomic DNA minipreps kit (Bio Basic, Inc) as per directions provided by the supplier. The DNA quantity testing was done using a nano-drop spectrophotometer (ND1000, Thermo Scientific). DNA was re-quantified by visualization under UV light after running on 1% agarose gel electrophoresis using 1× TBE buffer at 90 voltage in 15 min and staining with 0.5 µg/mL ethidium bromide. The final DNA concentration was diluted to 6 ng/ml for PCR amplification and kept in a freezer at – 20 °C. A total of 15 ISSR primers (Cinnagen Co, Iran) were used in lyophilized form, and optimum annealing temperature was determined for each primer. All reactions were performed in a BIO-RAD T100™ Thermal Cycler using the following protocol: one cycle of 5 min at 94 °C, 40 cycles of 45 s at 94 °C, 40 cycles of 45 s at annealing temperature, 40 cycles of 2 min at 72 °C, and final step one cycle of 10 min at 72 °C. The amplified PCR products from each accession were separated by gel electrophoresis using a 1.5% agarose gel, stained by soaking the gels in a 0.5 µg/mL ethidium bromide and revealed by a gel documentation unit (Syngene Bio Imaging). Observable polymorphic banding of ISSR markers was selected for analysis. In order to evaluate diversity among accessions, presence and absence of fragments were coded as 1 and 0 for each band, respectively. Data were constructed in a binary qualitative data matrix using MS Excel. NTSYS PC version 2.02 software was used for data analysis [40]. Cluster analysis was done by using two methods, namely principle coordinate analysis (PCoA) and the unweighted pair-group method with arithmetic averages (UPGMA). For each primer, polymorphism information content value (PIC; [37]), probability identity (PI; [38]) and discriminating power (D = 1 − PI) were calculated as follows:
where Pi and Pj are the frequencies of the ith and jth alleles within each locus, respectively.
Results and Discussion
The highest leaf width belonged to the Ilam Saymareh bridge (5.1 mm), and the lowest leaf width belonged to the Yazd (1.1 mm) accessions (Table 3). The highest leaf length belonged to the Malayear (10.7 cm), and the lowest leaf length belonged to the Doroud Siahvel (1.8 cm) accessions. The number of rachis was between 3 and 7. The spike length ranges between 1.0 cm (Gorgan accession) and 6.3 cm (Ilam Saymareh bridge). Observations indicated that accessions collected from Sarein and Talesh showed the highest visual quality (9), and accession from Semnan showed the lowest visual quality (4). The highest color belonged to the accessions collected from Boroujerd, Mahidasht, and Doroud Siahvel (9). The lowest color belonged to the accessions collected from Semnan (5).
In the dendrogram obtained based on 14 qualitative and quantitative morphological traits (Fig. 1), populations were divided into two major groups. Each group was divided into two subgroups. The first subgroup had 17 accessions comprised: Boroujerd, Doroud Daneshjo Park, Doroud Siahvel, Taft, Firouzan, Kamiaran, Holailan, Doroud Nahalestan, Homail, Malayear intersection, Mahidasht, Malayear, Ghidar, Damghan, Arak, Zanjan, and Tabriz. The second subgroup had 21 accession including Sarein, Talesh, Anzali, Nour, foreign cultivar, Maiami, Nahavand, Naien, Safashahr, Khorramabad, Ardakan 2, Yazd, Sari, Minoudasht, Chenaran, Gorgan, Mamulan, Mashhad, Shiraz, Tehran, and Poldokhtar. The third subgroup had three accessions including: Dehgolan, Ardakan 1, and Saimareh bridge. The fourth subgroup had nine accessions including: Abidar, Doroud Babahour, Azna, Aligoudarz, Daran, Abarkouh, Islamabad Gharb, Taghbostan, and Semnan. The accessions that isolated from north of Iran exhibited considerable separation after cold stress, but a weak association was found among accessions collected from west of Iran [1].
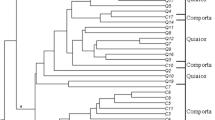
All 15 primers amplified and gave clear and highly reproducible DNA fragments. In total, 152 fragments were produced, of which 144 (94.73%) being polymorphic. Maximum bands (15) were produced with ISSR-5 and ISSR-7 while ISSR-1, ISSR-3, ISSR-6, ISSR-8, ISSR-11, ISSR-12, and ISSR-13 primers produced 100% polymorphic bands (Table 4). Polymorphic bands observed from 15 ISSR markers were ranged from 4 to 14, with an average of 9.60. PIC values ranged from 0.700 to 0.928, with a mean PIC value of 0.800. PI and discriminating power (D = 1 − PI) between the primers ranged from 0.029 to 0.185 and 0.815 to 0.971, respectively. Annealing temperature ranged from 42.5 °C (ISSR-10) to 56.1 °C (ISSR-2 and ISSR-8). The UPGMA clustering algorithm from ISSR analysis distributed the 50 accessions into six main clusters considered as A, B, C, D, E, and F with a similarity coefficient of 0.40 (Fig. 2). Cluster A consisted of Ardakan 1. Cluster B consisted of Khorramabad, Abidar, Tabriz, Homail, Chenaran, Boroujerd, Anzali, Dehgolan, Malayear, Nour, Abarkouh, Nahavand, Mamoulan, Mashhad, Islamabad Gharb, Minoudasht, Mahidasht, Semnan, Tehran, Arak, Kamiaran, Daran, Doroud Siahvel, Azna, Taghbostan Kermanshah, Doroud Babahour, Maiami, Holailan, Safashahr, Ardakan 2, Zanjan, Gorgan, Doroud Nahalestan, Damghan, Talesh, Doroud Daneshjo Park, Taft, Malayear intersection, Aligoudarz, Sarein, Sari, Saymareh bridge, and Poldokhtar. Cluster C consisted of foreign cultivar and Naein. Cluster D contained Shiraz and Yazd. Cluster E contained Ghidar. Lastly, Cluster F consisted of Firouzan. PCoA was carried out based on the genetic resemblance matrix to further understanding the relations between different accessions. Similar groupings with the UPGMA dendrogram were also revealed by the two-dimensional PCoA plot (Fig. 3). PCoA analysis also clearly showed great variation among accessions, supporting data from dendrogram. Similar grouping pattern of clustering was shown both in the PCoA biplot and in the dendrogram in other reports by Huang et al. [17], Li et al. [29], and Senthil Kumar et al. [44].
Dendrogram of 49 common bermudagrass accessions and one foreign cultivar derived from cluster analysis (UPGMA) based on genetic similarity estimates from the ISSR marker analysis. (1) Ardakan 1, (2) Khoram abad, (3) Homail, (4) Chenaran, (5) Abidar, (6) Anzali, (7) Shiraz, (8) Abarkouh, (9) Doroud babahour, (10) Mamoulan, (11) Tabriz, (12) Tehran, (13) Mashhad, (14) Semnan, (15) Foreign cultivar, (16) Naein, (17) Sari, (18) Ghidar, (19) Mahidasht, (20) Ardakan 2, (21) Islamabad gharb, (22) Minoudasht, (23) Daran, (24) Arak, (25) Kamiaran, (26) Doroud siahvel, (27) Boroujerd, (28) Nahavand, (29) Malayear, (30) Nour, (31) Miami, (32) Zanjan, (33) Damghan, (34) Yazd, (35) Holailan, (36) Dehgolan, (37) Doroud nahalestan, (38) Gorgan, (39) Saymareh bridge, (40) Firouzan, (41) Tagh bostan, (42) Doroud daneshjo park, (43) Talesh, (44) Taft, (45) Malayear intersection, (46) Aligoudarz, (47) Sarein, (48) Safashahr, (49) Azna, and (50) Poldokhtar
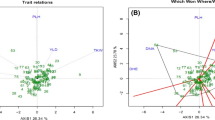
Two-dimensional projection of ISSR diversity calculated by PCA for 49 genotypes of common bermudagrass and one foreign cultivar. (1) Ardakan 1, (2) Khoram abad, (3) Homail, (4) Chenaran, (5) Abidar, (6) Anzali, (7) Shiraz, (8) Abarkouh, (9) Doroud babahour, (10) Mamoulan, (11) Tabriz, (12) Tehran, (13) Mashhad, (14) Semnan, (15) Foreign cultivar, (16) Naein, (17) Sari, (18) Ghidar, (19) Mahidasht, (20) Ardakan 2, (21) Islamabad gharb, (22) Minoudasht, (23) Daran, (24) Arak, (25) Kamiaran, (26) Doroud siahvel, (27) Boroujerd, (28) Nahavand, (29) Malayear, (30) Nour, (31) Miami, (32) Zanjan, (33) Damghan, (34) Yazd, (35) Holailan, (36) Dehgolan, (37) Doroud nahalestan, (38) Gorgan, (39) Saymareh bridge, (40) Firouzan, (41) Tagh bostan, (42) Doroud daneshjo park, (43) Talesh, (44) Taft, (45) Malayear intersection, (46) Aligoudarz, (47) Sarein, (48) Safashahr, (49) Azna, and (50) Poldokhtar
The results of morphological data showed a broad degree of variation among the accessions studied. Among all the observed phenotypic traits, presence of hair on leaf and sheath surfaces was a major factor for groupings in dendrogram. Hair presence on leaf and sheath surfaces was important characteristic that had been analyzed in some earlier reports [6, 36, 42]. Two forms glabrum Roshev. and villosus Regel. within C. dactylon were reported by Rozhevits and Shishkin [42] based on hair presence on leaves. Within C. dactylon, these traits were used by Davis [6] as diagnostic traits to discern var. dactylon and var. villosus Regel. Nasiri et al. [36] suggested that Iranian C. dactylon has two varieties: var. dactylon and var. villosus Regel. The results of our taxonomic investigation support the existence of two varieties of C. dactylon in Iran: var. dactylon (hairless plant) and var. villosous (plant with hairs at leaf underside and/or upper side surfaces or exterior surfaces of sheath).
Akbari and Salehi’s [1] results revealed that cold-tolerant common bermudagrass accessions had higher proline, protein, antioxidant enzymes, color, visual quality, and chlorophyll content under cold stress conditions. Selection of natural grasses with high stress tolerance is the most efficient and fastest method in plant improvement programs [30]. Great differences in freezing tolerance were observed among common bermudagrass accessions of Iran [1]. The major aim of many bermudagrass breeding programs is to select the genotypes with good freezing tolerance to increase winter survivability [3, 4].
Morphological traits in turfgrass may be affected by climatic conditions, and grasses grown in different climatic zones may be different morphologically. Thus, the use of morphological characters for grouping may result in inconsistency. Evaluation of genetic variability of accessions is a prerequisite to develop high stress-tolerant cultivars and to produce genetically diversified populations. A wide diversity has been reported for morpho-physiological and phylogenetic traits in common bermudagrass accessions [18, 33, 36]. Our results demonstrate that the ISSR markers are effective tools for identifying polymorphism between common bermudagrass accessions, as they can measure sufficient polymorphism and detect very low-level genetic variations and present high PIC, D, and low PI. A similar result was made by several other researches [17, 33, 47]. Our results clearly show that the polymorphism (94.73%) generated by ISSR markers was very high, and high level of variation exists in Cynodon accessions. Mohammadi Farsani et al. [33] in a survey on genetic variability of common bermudagrass genotypes from Iran by ISSR reported that of the 389 bands produced, 313 (80.5%) were polymorphic. High polymorphism can be due to high mutation chance in common bermudagrass DNA [34, 39] that can be related to the extensive geographical environment of Iran, which contained many climatic zones. This work showed also the good potential of random primers in assessing diversity and revealing high level of polymorphism among accessions. It has been proved that genetic variation in bermudagrass can be due to hybridization [47]). No correlation was found between the level of polymorphism and the number of amplified bands. This result is similar with results of Zhao et al. [56] using ISSR molecular markers.
The PIC was used to estimate the genetic variability levels in common bermudagrass genotypes. Three degrees of polymorphism include high (PIC > 0.5), medium (0.5 > PIC > 0.25), and low (PIC < 0.25) [53]. The high PIC values with a mean of 0.800 show that all primers are informative, and this can be related to high genetic variation among accessions used in this research. The abundance of genetic variability in natural Iranian common bermudagrass accessions can be the source of useful genes and genotypes, which is important in breeding programs. The genetic similarity coefficients of all accessions ranged from 0.43 to 0.78 (Fig. 2), with some accessions from the same area tending to form a subgroup. Lower genetic distance indicated lower relatedness among the accessions. No complete relatedness was observed between molecular clustering and geographical affinities. This might be due to different ploidy levels, cross-pollination, genetic overlap, germplasm exchange, and gene flow. Dendrogram generated from ISSR molecular markers may be used to discriminate between common bermudagrass accessions with different chromosome numbers, as reported by Anderson et al. [2] and Etemadi et al. [10].
According to morphological and genetic diversity of accessions, they can be clustered into three groups: cold sensitive, cold semi-tolerant, and cold tolerant. The most cold-tolerant accessions included: foreign cultivar, Naein, Safashahr, Gorgan, Malayear, Aligoudarz, and Taft. Accessions collected from Taft, Naein, and Malayear were the most cold-tolerant accessions compared to foreign cultivar. These accessions can be introduced as cold tolerant. Iran has a variable climate. Accessions from Taft and Naein were collected from center of Iran, with cold, dry winters and hot, dry summers. Accession from Malayear was collected from west of Iran. In the west, winters are cold with substantial snowfall and subfreezing temperatures, where summers are dry and mild. The UPGMA dendrogram grouped the accessions to some degree according to their geographical origin and some morphological traits, which is in agreement with earlier reports of Li et al. [29] and Mohammadi Farsani et al. [33]. In the present study, we integrate morphological and previously tested physiological traits with ISSR molecular markers, to introduce genotypes with high cold stress tolerance. This study may provide useful information for further breeding programs on common bermudagrass. Further studies are in progress to check other characteristics such as drought and salt tolerance of the selected genotypes.
References
Akbari, M., & Salehi, H. (2018). Biochemical and physiological evaluations revealed high diversity within common bermudagrass (Cynodon dactylon [L.] Pers.) Iranian accessions under cold stress. Advances in Horticultural Science, 32 (in press).
Anderson, M. P., Taliaferro, C. M., Martin, D. L., & Anderson, C. S. (2001). Comparative DNA profiling of U-3 turf bermudagrass strains. Crop Science, 41, 1184–1189.
Anderson, J. A., & Taliaferro, C. M. (2002). Freeze tolerance of seed-producing turf bermudagarsses. Crop Science, 42, 190–192.
Anderson, J. A., Taliaferro, C. M., & Martin, D. L. (2003). Longer exposure durations increase freeze damage to bermudagrasses. Crop Science, 43, 973–977.
Arumuganathan, K., Tallury, S. P., Fraser, M. L., Bruneau, A., & Qu, R. (1999). Nuclear DNA content of thirteen turfgrass species by flow cytometry. Crop Science, 39, 1518–1521.
Davis, P. H. (1985). Flora of Turkey and the east Aegean islands (p. 579). Edinburgh, UK: Edinburgh University Press.
Devitt, D. A., Bowman, D. C., & Schulte, P. J. (1993). Response of Cynodon dactylon to prolonged water deficits under saline conditions. Plant and Soil, 148, 239–251.
Dipaola, J. M., & Beard, J. B. (1992). Physiological effects of temperature stress. In D. V. Waddington, R. N. Carrow, & R. C. Shearman (Eds.), Turfgrass Agronomy Monograph (Vol. 32, pp. 231–262). Madison, WI: American Society of Agronomy.
Downton, W. J. (1975). The occurrence of C4 photosynthesis among plants. Photosynthetica, 9, 96–105.
Etemadi, N., Sayed-tabatabaei, B. E., Zamani, Z., Razmjoo, K. H., Khalighi, A., & Lessani, H. (2006). Evaluation of diversity among Cynodon dactylon (L.) Pers. using RAPD markers. Journal of Agriculture and Biology, 8, 198–202.
Finger, F. L., Lannes, S. E., Schuelter, A. R., Doege, G., Comerlato, A. P., Goncalves, L. S. A., et al. (2010). Genetic diversity of Capsicum chinensis (Solanaceae) accessions based on molecular markers and morphological and agronomic traits. Genetics and Molecular Research, 9, 1852–1864.
Hajjar, R., & Hodgkin, T. (2007). The use of wild relatives in crop improvement: A survey of developments over the last 20 years. Euphytica, 156, 1–13.
Harlan, J. R., & de Wet, J. M. J. (1969). Sources of variation in Cynodon dactylon (L.) Pers. Crop Science, 9, 774–778.
Harlan, J. R. (1970). Geographic distribution of the species of Cynodon L.C. Rich (Graminae). East African Agricultural and Forestry Journal, 36, 220–226.
Hodkinson, T. R., Chase, M. W., & Renvoize, S. A. (2002). Characterization of a genetic resource collection for Miscanthus (Saccharinae, Andropogoneae, Poaceae) using AFLP and ISSR PCR. Annals of Botany, 89, 627–636.
Holm, L. G., Plunknett, D. L., Pancho, J. V., & Herberger, J. P. (1991). The world’s worst weeds: Distribution and biology (p. 609). Malabar, Florida: Krieger Publishing Company.
Huang, C. Q., Liu, G. D., Bai, C. J., Wang, W. Q., Zhou, S., & Yu, D. Q. (2010). Estimation of genetic variation in Cynodon dactylon accessions using the ISSR technique. Biochemical Systematics and Ecology, 38, 993–999.
Huang, C. Q., Liu, G., Bai, C., & Wang, W. (2014). Genetic analysis of 430 Chinese Cynodon dactylon accessions using sequence-related amplified polymorphism markers. International Journal of Molecular Sciences, 15, 19134–19146.
Ibitayo, O. O., & Butler, J. D. (1981). Cold hardiness of bermudagrass and Paspalum vaginatum SW. HortScience, 16, 683–684.
Jian, S. G., Tang, T., Zhong, Y., & Shi, S. H. (2004). Variation in inter-simple sequencerepeat (ISSR) in mangrove and non-mangrove population of Heritieralittoralis (Sterculiaceae) from China and Australia. Aquatic Botany, 79, 75–86.
Kamps, T. L., Williams, N. R., Ortega, V. M., Chamusco, K. C., Harris-Shultz, K., Scully, B. T., et al. (2011). DNA polymorphisms at bermudagrass microsatellite loci and their use in genotype fingerprinting. Crop Science, 51, 1122–1131.
Karaca, M., Saha, S., Zipf, A., Jenkins, J. N., & Lang, D. J. (2002). Genetic diversity among bermudagrass (Cynodon spp.): Evidence from chloroplast and nuclear DNA fingerprinting. Crop Science, 42, 2118–2127.
Karp, A., Edwards, K. J., Bruford, M., Funk, S., Vosman, B., Morgante, M., et al. (1997). Molecular technologies for biodiversity evaluation: Opportunities and challenges. Nature Biotechnology, 15, 625–629.
Karpinski, S., Wingsle, G., Karpinska, B., & Hallgren, J. (2002). Low-temperature stress and antioxidant defense mechanisms in higher plants. In D. Inze & M. V. Montagu (Eds.), Oxidative stress in plants (pp. 69–104). London, UK: Taylor & Francis.
Kenworthy, K. E., Taliaferro, C. M., Carver, B. F., Martin, D. L., Anderson, J. A., & Bell, G. E. (2006). Genetic variation in Cynodon transvaalensis Burtt-Davy. Crop Science, 46, 2376–2381.
Kearns, R., Zhou, Y., Fukai, S., Ye, C., Loch, D., Godwin, I., et al. (2009). Eco-turf: Water use efficient turfgrasses from Australian biodiversity. Acta Horticulturae, 829, 113–118.
Kissmann, K. (1991). Plantas infestantes e nocivas. Basf Brasileira, 2, 317–321.
Koster, K. L., & Leopold, A. C. (1988). Sugars and desiccation tolerance. Plant Physiology, 88, 829–832.
Li, H., Liu, L., Lou, Y., Hu, T., & Fu, J. (2011). Genetic diversity of Chinese natural bermudagrass (Cynodon dactylon) germplasm using ISSR markers. Scientia Horticulturae, 127, 555–561.
Luo, N., Liu, J., Yu, X., & Jiang, Y. (2011). Natural variation of drought response in Brachypodium distachyon. Physiologia Plantarum, 141, 19–29.
Majidi, M. M., & Mirlohi, M. (2010). Genetic similarities among Iranian populations of Festuca, Lolium, Bromus and Agropyron using amplified fragments length polymorphism (AFLP) markers. Iranian Journal of Biotechnology, 8, 16–23.
McCarty, L. B. (2005). Best golf course management practices. Upper Saddle River, NJ: Prentice Hall.
Mohammadi Farsani, T., Etemadi, N., Sayed-Tabatabaei, B., & Talebi, M. (2012). Assessment of genetic diversity of bermudagrass (Cynodon dactylon) using ISSR markers. International Journal of Molecular Sciences, 13, 383–392.
Mohsen, H., & Ali, F. (2008). (. Study of genetic polymorphism of Artemisia herba-alba from Tunisia using ISSR markers. African Journal of Biotechnology, 7, 44–50.
Munshaw, G. C., Ervin, E. H., Shang, C., Askew, S. D., Zhang, X., & Lemus, R. W. (2006). Influence of late-season iron, nitrogen, and seaweed extract on fall color retention and cold tolerance of four bermudagrass cultivars. Crop Science, 46, 273–283.
Nasiri, A., Saiedi, H., & Rahiminejad, M. R. (2012). Morphological variation and taxonomic conclusion of Cynodon dactylon (L.) Pers. in Iran. Asian Journal of Plant Sciences, 11, 62–69.
Oliviera, E. J., Padua, J. G., Zucchi, M. I., & Venkovsky, R. (2006). Origin, evolution and genome distribution of microsatellites. Biology, 29, 294–307.
Pollefeys, P., & Bousquet, J. (2003). Molecular genetic diversity of the French-American grapevine hybrids cultivated in North America. Genome, 46, 1037–1048.
Raina, S. N., Rani, V., Kojima, T., Ogihara, Y., Singh, K. P., & Devarumath, R. M. (2001). RAPD and ISSR fingerprints for analysis of genetic diversity, varietal identification, and phylogenetic relationships in peanut (Arachis hypogaea) cultivars and wild species. Genome, 44, 763–772.
Rohlf, F. J. (1998). NTSYSpc: Numerical taxonomy and multivariate analysis system. Version 2.02.. New York: Exeter Publications.
Roodt, R., Spies, J. J., & Burger, T. H. (2002). Preliminary DNA fingerprinting of the turfgrass Cynodon dactylon (Poaceae: Chloridoideae). Bothalia, 32, 117–122.
Rozhevits, R. Y., & Shishkin, B. K. (1934). Gramineae. In Komarov, VL. (Ed.), Flora of the U.S.S.R. Vol. 2. Koeltz Scientific Books, USA, pp. 227–228.
Salehi, M., Salehi, H., Niazi, A., & Ghobadi, C. (2014). Convergence of goals: phylogenetical, morphological, and physiological characterization of tolerance to drought stress in tall fescue (Festuca arundinacea Schreb.). Molecular Biotechnology, 56, 248–257.
Senthil Kumar, R., Parthiban, K., & Govinda Rao, M. (2009). Molecular characterization of Jatropha genetic resources through inter-simple sequence repeat (ISSR) markers. Molecular Biology Reports, 36, 1951–1956.
Shahba, M. A., Qian, Y. L., Hughes, H. G., Koski, A. J., & Christensen, D. (2003). Relationships of soluble carbohydrates and freeze tolerance in saltgrass. Crop Science, 43, 2148–2153.
Shashikumar, K., & Nus, J. L. (1993). Cultivar and winter cover effects on bermudagrass cold acclimation and crown moisture content. Crop Science, 33, 813–817.
Shyan, Y. S., Juraimi, A. S., Rafiil, M. Y., Shabanimofrad, M., Alam, M. A., & Hakim, M. A. (2014). Genetic divergence of bermudagrass (Cynodon spp.). Population using ISSR markers. Life Science Journal, 11, 425–430.
Smolik, M., Ochmian, I., & Grajkowski, J. (2010). Genetic variability of Polish and Russian accessions of cultivated blue honeysuckle (Lonicera caerulea). Genetika, 46, 1079–1085.
Taliaferro, C. M. (2000). Bermudagrass has made great strides-and its diversity has barely been tapped. How many more superb cultivars may lurk in its germplasm? Diversity, 16, 23–24.
Thomashow, M. F. (1999). Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annual Review of Plant Physiology and Plant Molecular Biology, 50, 571–599.
Williams, K., Kubelik, J. G., Livak, A. R., Rafalski, K. J., & Tingey, S. V. (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research, 18, 6231–6235.
Wu, Y., Taliaferro, C. M., Bai, G. H., & Anderson, M. P. (2004). AFLP analysis of Cynodon dactylon (L.) Pers. Var. dactylon genetic variation. Genome, 47, 689–696.
Xie, W., Zhang, X., Cai, H., Liu, W., & Peng, Y. (2010). Genetic diversity analysis and transferability of cereal EST-SSR markers to orchardgrass (Dactylis glomerata L.). Biochemical Systematics and Ecology, 38, 740–749.
Zehdi, S., Trifi, M., Salem, A., Rhouma, A., & Marrakchi, M. (2002). Survey of inter simple sequence repeat polymorphisms in Tunisian date palms (Phoenix dactylifera L.). Journal of Genetics and Breeding, 56, 77–83.
Zhang, X., Ervin, E. H., & LaBranche, A. J. (2006). Metabolic defense responses of seeded bermudagrass during acclimation to freezing stress. Crop Science, 46, 2598–2605.
Zhao, W. G., Zhou, Z. H., Miao, X. X., Wang, S. B., Zhang, L., Yile, P., et al. (2006). Genetic relatedness among cultivated and wild mulberry (Moraceae: Morus) as revealed by inter-simple sequence repeat (ISSR) analysis in China. Canadian Journal of Plant Science, 86, 251–257.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Akbari, M., Salehi, H. & Niazi, A. Evaluation of Diversity Based on Morphological Variabilities and ISSR Molecular Markers in Iranian Cynodon dactylon (L.) Pers. Accessions to Select and Introduce Cold-Tolerant Genotypes. Mol Biotechnol 60, 259–270 (2018). https://doi.org/10.1007/s12033-018-0068-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-018-0068-5