Abstract
Alzheimer's disease (AD), the most common type of dementia, is a serious neurodegenerative disease that has no cure yet, but whose symptoms can be alleviated with available medications. Therefore, early and accurate diagnosis of the disease and elucidation of the molecular mechanisms involved in the progression of pathogenesis are critically important. This study aimed to identify dysregulated miRNAs and their target mRNAs through the integrated analysis of miRNA and mRNA expression profiling in AD patients versus unaffected controls. Expression profiles in postmortem brain samples from AD patients and healthy individuals were extracted from the Gene Expression Omnibus database and were analyzed using bioinformatics approaches to identify gene ontologies, pathways, and networks. Finally, the module analysis of the PPI network and hub gene selection was carried out. A total of five differentially expressed miRNAs were extracted from the miRNA dataset, and 4312 differentially expressed mRNAs were obtained from the mRNA dataset. By comparing the DEGs and the putative targets of the altered miRNAs, 116 (3 upregulated and 113 downregulated) coordinated genes were determined. Also, six hub genes (SNAP25, GRIN2A, GRIN2B, DLG2, ATP2B2, and SCN2A) were identified by constructing a PPI network. The results of the present study provide insight into mechanisms such as synaptic machinery and neuronal communication underlying AD pathogenesis, specifically concerning miRNAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Complex biological processes are associated with a progressive and gradual decline in the biochemical and physiological activities of tissues and organs, leading to adult aging and age-related diseases such as cardiovascular disease, cancer, and dementia (Pugh and Wei 2001; Zhang et al. 2012; Gonul Baltaci et al. 2018; Kocpinar et al. 2020). Alzheimer's disease (AD) is a chronic neurodegenerative disease that is one of the most common forms of dementia among elderly people (Zhao et al. 2016; Gündoğdu et al. 2019; Istrefi et al. 2020). The pathogenesis of AD is very complex and is known to be caused by multiple factors (Ceylan et al. 2019; Turkes et al. 2019; Kalayci et al. 2021). The deposition of extracellular β-amyloid peptides and intracellular neurofibrillary tangles (NFTs) are considered major hallmarks of AD (Lichtenthaler et al. 2011; Wakasaya et al. 2011; Durgun et al. 2020; Sever et al. 2020). However, the full mechanisms and other principal molecular drivers that may affect AD pathogenesis remain unclear. Therefore, a further understanding and unraveling of key molecular pathways that lead to aging and age-associated diseases will provide insights into the development of new strategic approaches for healthy aging.
MicroRNAs (miRNAs), a class of non-coding RNAs, are small regulatory RNAs that can regulate gene expression post-transcriptionally (Chen 2010; Vazquez et al. 2010). miRNAs are critical for normal development and cellular homeostasis, and thus are involved in a variety of biological processes (Friedman and Jones 2009). Recent studies suggest that dysregulation and aberrant regulation and function of miRNAs may deeply impact cellular physiology, leading to the development of many diseases (Adams et al. 2014; Lai et al. 2016; Lee et al. 2016). Therefore, the identification and understanding of miRNA-target interactions are crucial for deciphering the regulatory mechanisms of miRNAs in various cellular processes and disease progression (Ceylan 2021a, 2021b). It has been shown that miRNAs participate in the regulation of gene expression in the aging process (Chen et al. 2010b; Lanceta et al. 2010). The high abundance of miRNAs within the nervous system, where they are key regulators of functions such as neuronal differentiation, neurogenesis, neurite growth, and synaptic plasticity, supports the hypothesis that miRNAs have a potential role in neurodegenerative diseases, especially in AD (Femminella et al. 2015; Moradifard et al. 2018).
The present study was carried out to demonstrate the association between miRNAs in AD brains and their target genes using bioinformatics analysis. Here we investigated and identified differentially expressed genes (DEGs), differentially expressed miRNAs (DEmiRs), and miRNA-mRNA interactions in the elderly AD brain, to provide novel insights into potential therapeutic targets that can be used to combat AD.
Methods
Microarray Data Profiles
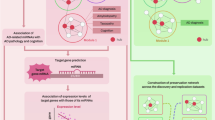
Datasets were retrieved from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/). The GSE26972 data set, which was extracted from the study of Berson et al. (Berson et al. 2012), has a total of six samples, containing three female non-demented controls (NDCs) and three female AD patients. The microarray platform was the GPL5188 [HuEx-1_0-st] Affymetrix Human Exon 1.0 ST Array [probe set (exon) version]. The GSE157239 data set was used for its miRNA expression profile. The miRNA data set samples were obtained from 16 postmortem cases, including eight control individuals without neuropathological lesions or neurological signs and eight AD patients (Braak stage III or above) (Henriques et al. 2020). The microarray platform was the GPL21572 [miRNA-4] Affymetrix Multispecies miRNA-4 Array [ProbeSet ID version]. A schematic representation of the methodology used in the present study is shown in Fig. 1.
Data Preprocessing, Screening of DEGs and DEmiRs
To identify DEGs and DEmiRs in AD compared to age-matched controls, the selected data sets were analyzed using the GEO web tool GEO2R (Barrett et al. 2013). |LogFC|> 1 and p < 0.05 were defined as the threshold for both data sets.
Acquiring the Intersection of DEG and DEmiR Targets
Firstly, the TargetScan online database (Agarwal et al. 2015) was used to predict the target genes of the up- and downregulated miRNAs with a high fold change (FC) (logFC > 1) value in the GSE157239 data set. The intersection of predicted targets by TargetScan was extracted using the Multiple List Comparator (http://www.molbiotools.com/listcompare.html). Next, to identify the interaction between DEmiRs and DEGs, the upregulated DEmiR targets were matched with downregulated DEGs, and downregulated DEmiR targets were matched with upregulated DEGs using the Multiple List Comparator. Consequently, the overlapping genes between the predicted target genes of DEmiRs and DEGs obtained from the GSE26972 data set were obtained.
Gene Ontology and Pathway Enrichment Analysis
Gene over-representation analyses were conducted using the ToppFun application in the ToppGene toolbox (Chen et al. 2009) to identify biological process (BP), cellular component (CC), and molecular function (MF) annotations of the identified common genes. Pathway enrichment analyses of DEGs was classified by the KEGG (Kyoto Encyclopedia of Genes and Genomes) using the DAVID pathway viewer. A p-value of < 0.05 was set as the cutoff value for all enrichment analyses.
Protein–Protein Interaction Network and Module Analyses
To evaluate the interrelationships among common DEGs, a protein–protein interaction network was constructed using the STRING (Search Tool for the Retrieval of Interacting Genes/Proteins) database (Jensen et al. 2009). Then, the PPI network was analyzed and visualized using Cytoscape software. Additionally, based on the above data, the Molecular Complex Detection (MCODE) app in Cytoscape was used for module selection. The CytoHubba plugin of Cytoscape was used to identify hub genes. The functions and pathway enrichment of candidate hub genes were analyzed using the DAVID (database for annotation, visualization, and integrated discovery) (da Huang et al. 2009) bioinformatics resource.
In Silico Validation of Hub Genes
To determine which cell type expressed the identified hub genes, we queried the genes in the Single-cell Atlas of the Entorhinal Cortex in Human Alzheimer’s Disease database (Grubman et al. 2019).
Results
Identification of DEmiRs and DEGs
In the present study, the differences in the miRNA expression profiles between six AD samples (62–94-year-old women) and five age-matched control samples (69–90-year-old women) of temporal cortex tissues were analyzed. A total of two (hsa-miR-1299 and hsa-miR-7150) upregulated and three (hsa-miR-4286, hsa-miR-4449, and hsa-miR-3195) downregulated miRNA were obtained after preprocessing of the GSE157239 data set. TargetScan software predicted 2182 common targets for hsa-miR-1299 and hsa-miR-7150 (Fig. 2a). By the same approach, 52 common targets were predicted by TargetScan for downregulated miRNAs (Fig. 2b). After data preprocessing, a total of 4312 DEGs were obtained from the GSE26972 data set, of which 3517 genes were upregulated and 795 were downregulated. Further analysis of these genes using a Venn diagram revealed 113 intersection genes between upregulated DEmiR targets and downregulated DEGs (Fig. 2c), and three intersection genes between downregulated DEmiR targets and upregulated DEGs (Fig. 2d).
Venn diagrams of DEGs, miRNAs, and the miRNA targets from GEO data sets. (a) Common upregulated miRNA targets, (b) common downregulated miRNA targets, (c) the intersection of downregulated DEGs and upregulated miRNA targets, (d) the intersection of upregulated DEGs and downregulated miRNA targets. DEGs, differentially expressed genes; GEO, Gene Expression Omnibus; miRNA, microRNA
Gene Ontology (GO) Function and KEGG Pathway Enrichment Analysis of Overlapped Genes
The functions and pathway enrichment of the DEGs were evaluated at the ToppGene website. Gene Ontology (GO) analysis further classified the DEGs into the three functional groups, i.e., biological process (BP), cellular component (CC), and molecular function (MF), as summarized in Table 1. The detailed GO analysis results for the DEGs are shown in Supplementary Table S1. According to the KEGG pathway enrichment analysis, upregulated genes were mainly enriched in adipocytokine signaling, PPAR signaling, glucagon signaling, insulin resistance, hepatitis C, non-alcoholic fatty liver disease (NAFLD), and cAMP signaling. The downregulated genes were mainly enriched in nicotine addiction and neuroactive ligand–receptor interaction (Table 2).
PPI Network Analyses and Module Analyses
Using the STRING online database and Cytoscape software, a total of 67 DEGs (2 upregulated and 65 downregulated) of the 116 altered DEGs from the entorhinal cortex were screened into the DEG PPI network, containing 67 nodes and 117 edges (Fig. 3). In total, one module (cluster 1) with a score > 5 was detected by the Cytoscape plug-in MCODE (Fig. 4a). Cluster 1 contained nine nodes including ANK3, CNTNAP2, NFASC, DLG2, SCN8A, GRIN2B, SNAP25, SCN2A, and ATP2B2 with 26 edges. The most connected nodes were identified from the PPI network using CytoHubba based on the betweenness algorithm (Fig. 4b). According to the betweenness scores, the ten highest-scored genes, including SNAP25, PACSIN1, GRIN2A, GRIN2B, SGIP1, GABRA4, DLG2, ATP2B2, GABRA3, and SCN2A, are listed in Table 3. Among the screened genes, SNAP25, GRIN2A, GRIN2B, DLG2, ATP2B2, and SCN2A were selected as hub (centrally located) genes. The MCODE score (> 3) and clustering status (found in cluster 1) were used as selection criteria.
The functional sub-network analysis of PPI network. (a) The sub-module from the PPI network of DEGs from samples of AD and healthy controls. The blue nodes represent the downregulated genes. (b) Construction of PPI network of hub genes. The grades of the colors represent the betweenness score. Lines indicate protein–protein interactions
GO Function and KEGG Pathway Enrichment Analysis of Hub Genes
To explore the biological functions and pathways of the genes, the enrichment analyses of the hub genes were evaluated using the DAVID online tool. GO annotation and pathway analysis results for six hub genes are summarized in Table 4.
Validation of Cell-Type-Specific Expression of Hub Genes
Major cell types of the human brain in which hub genes are expressed were identified using a single-cell/nuclei RNA-seq data set of AD samples of human brain tissues. Expression signatures detected from human postmortem AD brain tissues showed that all of the hub genes are predominantly expressed in neurons (Fig. 5).
Discussion
The pathogenesis of AD is extremely complex, involving multiple pathways and genes. As is the case with many other neurodegenerative diseases, the disease is predictable but remains incurable (Zarrin et al. 2021). Therefore, understanding the mechanisms underlying AD would aid in finding the best cure options as early as possible, which would improve the quality of life in patients with AD. Gene expression arrangements are altered in different stages of the disease. Therefore, the identification of differentially expressed genes in the brain tissues may help us understand the pathogenesis of AD. It is known that women are disproportionately affected by Alzheimer’s, and this disease forces them to make difficult choices about their lives. Additionally, World Health Organization reports and recent studies show that nearly two-thirds of deaths from the disease and other forms of dementia are in women (Snyder et al. 2016). Therefore, it is critical to identify risk factors that contribute to the risk of AD or that can be used as a target in treatment, especially in women. Hence, in this study, gene and miRNA expression profiles belonging to female patients were integrated to explore expression alterations in AD.
MicroRNAs are sophisticated regulatory non-coding RNAs that influence almost every aspect of cellular function (Ramakrishna and Muddashetty 2019). Due to their key roles in translational regulation and target mRNA decay, these molecules are implicated in the pathology of many disorders (Jo et al. 2015). Since miRNAs are involved in normal development and various biological processes, it is believed that their abnormal expression may be associated with many human diseases such as cardiovascular disorders (Wojciechowska et al. 2017), cancer (Peng and Croce 2016), inflammation (Contreras and Rao 2012), infectious diseases (Kim et al. 2017), and both central and peripheral neurological disorders (Zhang et al. 2018; Majidinia et al. 2020a). To date, many miRNAs have been identified as critical elements for the regulation of neuronal development, synapse formation, and cognitive functions lost in AD (Somel et al. 2010; Chen and Qin 2015). Moreover, recent studies have shown that an estimated 70% of all identified mature miRNAs in humans are expressed in the nervous system (Adlakha and Saini 2014). Therefore, elucidating the key mechanisms involving miRNAs and their targets might provide important insights into the pathogenesis of neurodegenerative diseases including AD.
In the present study, 116 DEmiR-targeted DEGs were identified in AD brain tissues compared to normal tissues based on gene expression profile data. Moreover, it was indicated that DEGs, including SNAP25, GRIN2A, GRIN2B, DLG2, ATP2B2, and SCN2A, were hub genes in the PPI network analysis (see Table 3). All six of these genes were downregulated in AD samples versus unaffected controls, suggesting that these genes may play critical roles in AD pathogenesis and/or progression.
Previous studies have shown that abnormalities in the pre-and/or postsynaptic machinery (neurotransmitters/neurotransmitter receptors and ion channels) were compromised in many age-related neurological disorders including AD (Mhatre et al. 2014; Ceylan and Erdogan 2017; Melland et al. 2020). According to GO and pathway enrichment analysis, hub genes are particularly enriched in synaptic function, synaptic regulation, learning, and memory (see Table 4). The majority of AD treatment approaches are focused on the beta-amyloid and tau protein pathologies (Klafki et al. 2006; Jackson et al. 2019). However, the synapse itself may also be an important endpoint for disease modification. It is known that the function, structure, and eventually the number of synapses change over time (Chen et al. 2019). This process, which is called synaptic plasticity, affects synaptic contact and optimizes neural connections, which are essential for cognitive functions (Fu and Ip 2017; Li et al. 2019). Therefore, determining the underlying mechanisms associated with the maintenance and recovery of synaptic function is important in the development of synaptic function-based therapeutic strategies in the treatment of dementia (Majidinia et al. 2020b).
Presynaptic terminals are spots of pathological alterations causing a synaptic loss in AD, and are also the site of many specific operations critical to standard neuronal functions (Overk and Masliah 2014). It was previously reported that multiple terminal proteins are affected in AD, which results in widespread synaptic pathology in multiple brain regions (Zhou and Liu 2015). Synaptosomal-associated protein 25 kDa (SNAP25), is a membrane-associated protein involved in synaptic transmission (Irfan et al. 2019). This protein is widely expressed but is highly variable in the mammalian nervous system. Furuya et al. (Furuya et al. 2012) reported that SNAP25 was expressed differentially across various brain regions in postmortem AD tissues. The fact that the protein is not uniformly expressed in all neuronal populations in the nervous system suggests that SNAP25 is involved in specific transmitter pathways (Minger et al. 2001). It has also been shown that SNAP25 levels are reduced in the temporal, occipital, frontal, and parietal cortex regions of the brain in patients with AD (Corradini et al. 2009). Other studies have similarly noted that SNAP25 was reduced in AD, and thus neurotransmission was impaired (Noor and Zahid 2017).
Neurotransmitter receptors are tuned by the signaling pathways of the brain in response to a pathological status to maintain homeostasis in the nervous network (Berg et al. 2013; Domercq et al. 2013). NMDA glutamatergic receptors (N-methyl-d-aspartate receptors; NMDARs) are one of the main receptors implicated in AD symptoms (Liu et al. 2019). Insufficient or excessive synaptic stimulation of NMDAR signaling compromises neuronal cell survival (Wang and Reddy 2017). Thus, NMDAR signaling must be tightly regulated because abnormal stimulation may disrupt the balance between neuronal survival and death. NMDARs are heterotetramers consisting of two GluN1 and two GluN2 subunits (Luo et al. 2011). GRIN2A and GRIN2B are the main subunits of functional NMDARs and they are the principal NMDARs found predominantly in brain regions affected by AD (Chen et al. 2010a). Numerous studies suggest that the GluN2B subunit, encoded by the GRIN2B gene, is of particular importance for the pathogenesis of AD, for several reasons: (1) GRIN2B is found to be abundant in areas where pathological signs of AD are concentrated and spread, including the neocortex (Hynd et al. 2004); (2) a GRIN2B-containing NMDA receptor antagonist prevents Aβ-induced synaptic plasticity disruption (Hu et al. 2009); (3) dysfunction of GRIN2B may contribute to memory impairment in AD patients (Clayton et al. 2002). Similarly, it was found that the GRIN2A subunit, encoded by the GRIN2A gene, plays essential roles not only in learning and memory but also in synaptic plasticity and is significantly reduced in sensitive regions in AD brains (Sun et al. 2017). Studies have also shown that the suppression of GRIN2A impairs the learning of complex motor skills (Lemay-Clermont et al. 2011). Therefore, these data suggest that the modulation of these subunits could be an effective strategy to treat brain diseases in the future. DLG2 encodes a membrane-associated protein, discs large MAGUK scaffold protein 2, which is a key scaffolding protein at postsynaptic sites and is required for NMDA receptor complex formation (Frank et al. 2016). It has been shown that DLG2 interacts with GRIN2A, and plays a role in synaptic dysfunction in AD (Irie et al. 1997). Also, it was previously reported that DLG2 expression is downregulated in the AD transcriptome and proteome network (Hallock and Thomas 2012). Recently, Yu et al. (Yu et al. 2017) revealed that DLG2 restoration in AD models ameliorated learning and memory impairment, improving Aβ-mediated cognitive dysfunction. Therefore, DLG2, which plays a critical role in the postsynaptic terminal, is predicted to contribute to cognitive flexibility, and ensuring proper modulation of DLG2 might be a potential therapy for AD.
It is known that calcium (Ca2+) is a ubiquitous second messenger, and is involved in neuronal life processes (Popugaeva et al. 2018). Numerous studies suggest that dysregulation of the intracellular Ca2+ levels and Ca2+-signaling pathways play major roles in the regulation of synaptogenesis and neuronal survival (Stafford et al. 2017), and thus they might act as a major progenitor of AD pathogenesis, such as Aβ accumulation and abnormal hyperphosphorylation of tau (Hermes et al. 2010). In the regulation of calcium homeostasis, many genes play a critical role, including ATP2A2 and ATP2B2. ATP2B2, which encodes plasma membrane calcium-transporting ATPase 2, is expressed at high levels in the brain, and plays a crucial role in removing divalent ions from cells to maintain intracellular calcium homeostasis (Kong et al. 2015). In Brendel et al.’s study (Brendel et al. 2014), the authors revealed that experimental downregulation of ATP2B2 increased the vulnerability of neuronal cells due to insufficient calcium efflux and significantly suppressed cell survival. Previous investigations also reported that ATP2B2 is primarily localized in specific parts of the CNS and is found predominantly in neurons, and the function of PMCA (plasma membrane calcium ATPases) in synaptic membranes of the brain decreases with aging (Zaidi et al. 1998; Garcia and Strehler 1999). All these findings raise the possibility that modulating this activity in vivo can help reduce the risk of AD development.
Electrical activity is a key component in the formation of synapses and brain development (Planells-Cases et al. 2000). Action potentials enable the release of neurotransmitters necessary for triggering neuronal signaling from the synaptic terminal. Voltage-gated sodium channels (NaChs) play major roles in the generation and propagation of action potentials in neurons (de Lera Ruiz and Kraus 2015). SCN2A (sodium voltage-gated channel alpha subunit 2) is the most abundant NaChα subunit postnatally, playing an important role in maintaining central nervous system function, and therefore its levels remain higher through adulthood (Israel et al. 2017). Moreover, studies have also shown that SCN2A deficiency may affect the interaction of other α subunits with the β subunits and thereby modulate the expression of functional channels (Lakhan et al. 2009). SCN2A has been previously reported to contribute to excitability for proper synaptic formation and development (Kruth et al. 2020). Additional studies using knockout (Scn2a−/−) and heterozygous haploinsufficient mouse models (Scn2a+/−) have revealed that Scn2a loss impairs neuronal excitability, synaptic function, and voltage-gated sodium channel-mediated currents, and leads to massive neuronal apoptosis (Planells-Cases et al. 2000; Spratt et al. 2019).
To summarize, dysregulated miRNAs and their target mRNAs involved in important biological processes such as postsynaptic machinery, neurotransmission, and neuronal life in AD are revealed together here for the first time. The results correlate well with many previous studies, proving that these genes have the potential to cause disease and/or trigger important disease-related events. Our integrative analysis highlights miRNA and mRNA candidates that may contribute to the disruption of synaptic plasticity observed in AD. Overall, we identified hub genes and crucial pathways, particularly associated with the synaptic mechanism that may affect the course of AD initiation and progression. These findings contribute to an understanding of the molecular mechanisms involved in AD pathogenesis in addition to other major hallmarks of AD, and may provide insights for new therapeutic strategies and targets.
Conclusion
Given the complexity observed in AD pathophysiology, the identification of differentially expressed miRNAs/mRNAs using integrative bioinformatics-based analyses can reveal specific mechanisms and molecular events for therapeutic intervention. In the present study, changes in the miRNA and mRNA expression patterns in the AD brain were investigated using miRNA and mRNA expression profiles. However, some limitations to this study should also be noted. Firstly, these differentially expressed genes and microRNAs need to be further examined to verify the analysis results. Secondly, these RNAs were screened from different cases and healthy samples, which could have affected the results.
Data Availability
All relevant data analyzed during the current study are available in the GEO repository.
References
Adams BD, Kasinski AL, Slack FJ (2014) Aberrant regulation and function of microRNAs in cancer. Curr Biol 24:R762–R776. https://doi.org/10.1016/j.cub.2014.06.043
Adlakha YK, Saini N (2014) Brain microRNAs and insights into biological functions and therapeutic potential of brain enriched miRNA-128. Mol Cancer 13:33. https://doi.org/10.1186/1476-4598-13-33
Agarwal V, Bell GW, Nam JW, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife 4. https://doi.org/10.7554/eLife.05005
Barrett T, Wilhite SE, Ledoux P et al (2013) NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res 41:D991–D995. https://doi.org/10.1093/nar/gks1193
Berg DA, Belnoue L, Song H, Simon A (2013) Neurotransmitter-mediated control of neurogenesis in the adult vertebrate brain. Development 140:2548–2561. https://doi.org/10.1242/dev.088005
Berson A, Barbash S, Shaltiel G et al (2012) Cholinergic-associated loss of hnRNP-A/B in Alzheimer’s disease impairs cortical splicing and cognitive function in mice. EMBO Mol Med 4:730–742. https://doi.org/10.1002/emmm.201100995
Brendel A, Renziehausen J, Behl C, Hajieva P (2014) Downregulation of PMCA2 increases the vulnerability of midbrain neurons to mitochondrial complex I inhibition. Neurotoxicology 40:43–51. https://doi.org/10.1016/j.neuro.2013.11.003
Ceylan H (2021a) A bioinformatics approach for identifying potential molecular mechanisms and key genes involved in COVID-19 associated cardiac remodeling. Gene Rep 24:101246. https://doi.org/10.1016/j.genrep.2021.101246
Ceylan H (2021b) Identification of hub genes associated with obesity-induced hepatocellular carcinoma risk based on integrated bioinformatics analysis. Med Oncol 38:63. https://doi.org/10.1007/s12032-021-01510-0
Ceylan H, Budak H, Kocpinar EF, Baltaci NG, Erdogan O (2019) Examining the link between dose-dependent dietary iron intake and Alzheimer’s disease through oxidative stress in the rat cortex. J Trace Elem Med Biol 56:198–206. https://doi.org/10.1016/j.jtemb.2019.09.002
Ceylan H, Erdogan O (2017) Cloning, expression, and characterization of human brain acetylcholinesterase in Escherichia coli using a SUMO fusion tag. Turk J Biol 41:77–87. https://doi.org/10.3906/biy-1602-83
Chen C, Li X, Wang T, Wang HH, Fu Y, Zhang L, Xiao SF (2010a) Association between NMDA receptor subunit 2b gene polymorphism and Alzheimer’s disease in Chinese Han population in Shanghai. Neurosci Bull 26:395–400. https://doi.org/10.1007/s12264-010-0729-2
Chen J, Bardes EE, Aronow BJ, Jegga AG (2009) ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res 37:W305–W311. https://doi.org/10.1093/nar/gkp427
Chen LH, Chiou GY, Chen YW, Li HY, Chiou SH (2010b) MicroRNA and aging: a novel modulator in regulating the aging network. Ageing Res Rev 9(Suppl 1):S59-66. https://doi.org/10.1016/j.arr.2010.08.002
Chen W, Qin C (2015) General hallmarks of microRNAs in brain evolution and development. RNA Biol 12:701–708. https://doi.org/10.1080/15476286.2015.1048954
Chen X (2010) Small RNAs - secrets and surprises of the genome. Plant J 61:941–958. https://doi.org/10.1111/j.1365-313X.2009.04089.x
Chen Y, Fu AKY, Ip NY (2019) Synaptic dysfunction in Alzheimer’s disease: Mechanisms and therapeutic strategies. Pharmacol Ther 195:186–198. https://doi.org/10.1016/j.pharmthera.2018.11.006
Clayton DA, Mesches MH, Alvarez E, Bickford PC, Browning MD (2002) A hippocampal NR2B deficit can mimic age-related changes in long-term potentiation and spatial learning in the Fischer 344 rat. J Neurosci 22:3628–37. https://doi.org/10.1523/jneurosci.22-09-03628.2002
Contreras J, Rao DS (2012) MicroRNAs in inflammation and immune responses. Leukemia 26:404–413. https://doi.org/10.1038/leu.2011.356
Corradini I, Verderio C, Sala M, Wilson MC, Matteoli M (2009) SNAP-25 in neuropsychiatric disorders. Ann N Y Acad Sci 1152:93–99. https://doi.org/10.1111/j.1749-6632.2008.03995.x
de Lera RM, Kraus RL (2015) Voltage-gated sodium channels: structure, function, pharmacology, and clinical indications. J Med Chem 58:7093–7118. https://doi.org/10.1021/jm501981g
Domercq M, Vazquez-Villoldo N, Matute C (2013) Neurotransmitter signaling in the pathophysiology of microglia. Front Cell Neurosci 7:49. https://doi.org/10.3389/fncel.2013.00049
Durgun M, Turkes C, Isik M et al (2020) Synthesis, characterisation, biological evaluation and in silico studies of sulphonamide Schiff bases. J Enzyme Inhib Med Chem 35:950–962. https://doi.org/10.1080/14756366.2020.1746784
Femminella GD, Ferrara N, Rengo G (2015) The emerging role of microRNAs in Alzheimer’s disease. Front Physiol 6:40. https://doi.org/10.3389/fphys.2015.00040
Frank RA, Komiyama NH, Ryan TJ, Zhu F, O’Dell TJ, Grant SG (2016) NMDA receptors are selectively partitioned into complexes and supercomplexes during synapse maturation. Nat Commun 7:11264. https://doi.org/10.1038/ncomms11264
Friedman JM, Jones PA (2009) MicroRNAs: critical mediators of differentiation, development and disease. Swiss Med Wkly 139:466–72. smw-12794
Fu AK, Ip NY (2017) Regulation of postsynaptic signaling in structural synaptic plasticity. Curr Opin Neurobiol 45:148–155. https://doi.org/10.1016/j.conb.2017.05.016
Furuya TK, Silva PN, Payao SL et al (2012) Analysis of SNAP25 mRNA expression and promoter DNA methylation in brain areas of Alzheimer’s Disease patients. Neuroscience 220:41–46. https://doi.org/10.1016/j.neuroscience.2012.06.035
Garcia ML, Strehler EE (1999) Plasma membrane calcium ATPases as critical regulators of calcium homeostasis during neuronal cell function. Front Biosci 4:D869–D882. https://doi.org/10.2741/garcia
Gonul Baltaci N, Guler C, Ceylan H, Kalin SN, Adem S, Kocpinar EF, Erdogan O, Budak H (2018) In vitro and in vivo effects of iron on the expression and activity of glucose 6-phosphate dehydrogenase, 6-phosphogluconate dehydrogenase, and glutathione reductase in rat spleen. J Biochem Mol Toxicol:e22229. https://doi.org/10.1002/jbt.22229
Grubman A, Chew G, Ouyang JF et al (2019) A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 22:2087–2097. https://doi.org/10.1038/s41593-019-0539-4
Gündoğdu S, Türkeş C, Arslan M, Demir Y, Beydemir Ş (2019) New isoindole-1,3dione substituted sulfonamides as potent inhibitors of carbonic anhydrase and acetylcholinesterase: Design, synthesis, and biological evaluation. ChemistrySelect 4:13347–13355. https://doi.org/10.1002/slct.201903458
Hallock P, Thomas MA (2012) Integrating the Alzheimer’s disease proteome and transcriptome: a comprehensive network model of a complex disease. OMICS 16:37–49. https://doi.org/10.1089/omi.2011.0054
Henriques AD, Machado-Silva W, Leite REP et al (2020) Genome-wide profiling and predicted significance of post-mortem brain microRNA in Alzheimer’s disease. Mech Ageing Dev 191:111352. https://doi.org/10.1016/j.mad.2020.111352
Hermes M, Eichhoff G, Garaschuk O (2010) Intracellular calcium signalling in Alzheimer’s disease. J Cell Mol Med 14:30–41. https://doi.org/10.1111/j.1582-4934.2009.00976.x
Hu NW, Klyubin I, Anwyl R, Rowan MJ (2009) GluN2B subunit-containing NMDA receptor antagonists prevent Abeta-mediated synaptic plasticity disruption in vivo. Proc Natl Acad Sci U S A 106:20504–20509. https://doi.org/10.1073/pnas.0908083106
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Hynd MR, Scott HL, Dodd PR (2004) Differential expression of N-methyl-D-aspartate receptor NR2 isoforms in Alzheimer’s disease. J Neurochem 90:913–919. https://doi.org/10.1111/j.1471-4159.2004.02548.x
Irfan M, Gopaul KR, Miry O, Hokfelt T, Stanton PK, Bark C (2019) SNAP-25 isoforms differentially regulate synaptic transmission and long-term synaptic plasticity at central synapses. Sci Rep 9:6403. https://doi.org/10.1038/s41598-019-42833-3
Irie M, Hata Y, Takeuchi M, Ichtchenko K, Toyoda A, Hirao K, Takai Y, Rosahl TW, Sudhof TC (1997) Binding of neuroligins to PSD-95. Science 277:1511–1515. https://doi.org/10.1126/science.277.5331.1511
Israel MR, Tay B, Deuis JR, Vetter I (2017) Sodium Channels and Venom Peptide Pharmacology. Adv Pharmacol 79:67–116. https://doi.org/10.1016/bs.apha.2017.01.004
Istrefi Q, Turkes C, Arslan M, Demir Y, Nixha AR, Beydemir S, Kufrevioglu OI (2020) Sulfonamides incorporating ketene N, S-acetal bioisosteres as potent carbonic anhydrase and acetylcholinesterase inhibitors. Arch Pharm (weinheim) 353:e1900383. https://doi.org/10.1002/ardp.201900383
Jackson J, Jambrina E, Li J, Marston H, Menzies F, Phillips K, Gilmour G (2019) Targeting the synapse in Alzheimer’s disease. Front Neurosci 13:735. https://doi.org/10.3389/fnins.2019.00735
Jensen LJ, Kuhn M, Stark M et al (2009) STRING 8–a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res 37:D412–D416. https://doi.org/10.1093/nar/gkn760
Jo MH, Shin S, Jung SR, Kim E, Song JJ, Hohng S (2015) Human argonaute 2 has diverse reaction pathways on target RNAs. Mol Cell 59:117–124. https://doi.org/10.1016/j.molcel.2015.04.027
Kalayci M, Turkes C, Arslan M, Demir Y, Beydemir S (2021) Novel benzoic acid derivatives: Synthesis and biological evaluation as multitarget acetylcholinesterase and carbonic anhydrase inhibitors. Arch Pharm (weinheim) 354:e2000282. https://doi.org/10.1002/ardp.202000282
Kim JK, Kim TS, Basu J, Jo EK (2017) MicroRNA in innate immunity and autophagy during mycobacterial infection. Cell Microbiol 19. https://doi.org/10.1111/cmi.12687
Klafki HW, Staufenbiel M, Kornhuber J, Wiltfang J (2006) Therapeutic approaches to Alzheimer’s disease. Brain 129:2840–2855. https://doi.org/10.1093/brain/awl280
Kocpinar EF, Gonul Baltaci N, Ceylan H, Kalin SN, Erdogan O, Budak H (2020) Effect of a prolonged dietary iron intake on the gene expression and activity of the testicular antioxidant defense system in rats. Biol Trace Elem Res 195:135–141. https://doi.org/10.1007/s12011-019-01817-0
Kong W, Mou X, Zhang N, Zeng W, Li S, Yang Y (2015) The construction of common and specific significance subnetworks of Alzheimer’s disease from multiple brain regions. Biomed Res Int 2015:394260. https://doi.org/10.1155/2015/394260
Kruth KA, Grisolano TM, Ahern CA, Williams AJ (2020) SCN2A channelopathies in the autism spectrum of neuropsychiatric disorders: a role for pluripotent stem cells? Mol Autism 11:23. https://doi.org/10.1186/s13229-020-00330-9
Lai CY, Lee SY, Scarr E et al (2016) Aberrant expression of microRNAs as biomarker for schizophrenia: from acute state to partial remission, and from peripheral blood to cortical tissue. Transl Psychiatry 6:e717. https://doi.org/10.1038/tp.2015.213
Lakhan R, Kumari R, Misra UK, Kalita J, Pradhan S, Mittal B (2009) Differential role of sodium channels SCN1A and SCN2A gene polymorphisms with epilepsy and multiple drug resistance in the north Indian population. Br J Clin Pharmacol 68:214–220. https://doi.org/10.1111/j.1365-2125.2009.03437.x
Lanceta J, Prough RA, Liang R, Wang E (2010) MicroRNA group disorganization in aging. Exp Gerontol 45:269–278. https://doi.org/10.1016/j.exger.2009.12.009
Lee JY, Ryu DS, Kim WJ, Kim SJ (2016) Aberrantly expressed microRNAs in the context of bladder tumorigenesis. Investig Clin Urol 57(Suppl 1):S52–S59. https://doi.org/10.4111/icu.2016.57.S1.S52
Lemay-Clermont J, Robitaille C, Auberson YP, Bureau G, Cyr M (2011) Blockade of NMDA receptors 2A subunit in the dorsal striatum impairs the learning of a complex motor skill. Behav Neurosci 125:714–723. https://doi.org/10.1037/a0025213
Li Q, Wu X, Na X et al (2019) Impaired cognitive function and altered hippocampal synaptic plasticity in mice lacking dermatan sulfotransferase Chst14/D4st1. Front Mol Neurosci 12:26. https://doi.org/10.3389/fnmol.2019.00026
Lichtenthaler SF, Haass C, Steiner H (2011) Regulated intramembrane proteolysis–lessons from amyloid precursor protein processing. J Neurochem 117:779–796. https://doi.org/10.1111/j.1471-4159.2011.07248.x
Liu J, Chang L, Song Y, Li H, Wu Y (2019) The role of NMDA receptors in Alzheimer’s disease. Front Neurosci 13:43. https://doi.org/10.3389/fnins.2019.00043
Luo T, Wu WH, Chen BS (2011) NMDA receptor signaling: death or survival? Front Biol (beijing) 6:468–476. https://doi.org/10.1007/s11515-011-1187-6
Majidinia M, Karimian A, Alemi F, Yousefi B, Safa A (2020a) Targeting miRNAs by polyphenols: Novel therapeutic strategy for aging. Biochem Pharmacol 173:113688. https://doi.org/10.1016/j.bcp.2019.113688
Majidinia M, Mir SM, Mirza-Aghazadeh-Attari M, Asghari R, Kafil HS, Safa A, Mahmoodpoor A, Yousefi B (2020b) MicroRNAs, DNA damage response and ageing. Biogerontology 21:275–291. https://doi.org/10.1007/s10522-020-09862-2
Melland H, Carr EM, Gordon SL (2020) Disorders of synaptic vesicle fusion machinery. J Neurochem. https://doi.org/10.1111/jnc.15181
Mhatre SD, Satyasi V, Killen M, Paddock BE, Moir RD, Saunders AJ, Marenda DR (2014) Synaptic abnormalities in a Drosophila model of Alzheimer’s disease. Dis Model Mech 7:373–385. https://doi.org/10.1242/dmm.012104
Minger SL, Honer WG, Esiri MM, McDonald B, Keene J, Nicoll JA, Carter J, Hope T, Francis PT (2001) Synaptic pathology in prefrontal cortex is present only with severe dementia in Alzheimer disease. J Neuropathol Exp Neurol 60:929–936. https://doi.org/10.1093/jnen/60.10.929
Moradifard S, Hoseinbeyki M, Ganji SM, Minuchehr Z (2018) Analysis of microRNA and Gene expression Profiles in Alzheimer’s Disease: A meta-analysis Approach. Sci Rep 8:4767. https://doi.org/10.1038/s41598-018-20959-0
Noor A, Zahid S (2017) A review of the role of synaptosomal-associated protein 25 (SNAP-25) in neurological disorders. Int J Neurosci 127:805–811. https://doi.org/10.1080/00207454.2016.1248240
Overk CR, Masliah E (2014) Pathogenesis of synaptic degeneration in Alzheimer’s disease and Lewy body disease. Biochem Pharmacol 88:508–516. https://doi.org/10.1016/j.bcp.2014.01.015
Peng Y, Croce CM (2016) The role of MicroRNAs in human cancer. Signal Transduct Target Ther 1:15004. https://doi.org/10.1038/sigtrans.2015.4
Planells-Cases R, Caprini M, Zhang J, Rockenstein EM, Rivera RR, Murre C, Masliah E, Montal M (2000) Neuronal death and perinatal lethality in voltage-gated sodium channel alpha(II)-deficient mice. Biophys J 78:2878–2891. https://doi.org/10.1016/S0006-3495(00)76829-9
Popugaeva E, Pchitskaya E, Bezprozvanny I (2018) Dysregulation of intracellular calcium signaling in Alzheimer’s disease. Antioxid Redox Signal 29:1176–1188. https://doi.org/10.1089/ars.2018.7506
Pugh KG, Wei JY (2001) Clinical implications of physiological changes in the aging heart. Drugs Aging 18:263–276. https://doi.org/10.2165/00002512-200118040-00004
Ramakrishna S, Muddashetty RS (2019) Emerging role of microRNAs in Dementia. J Mol Biol 431:1743–1762. https://doi.org/10.1016/j.jmb.2019.01.046
Sever B, Turkes C, Altintop MD, Demir Y, Beydemir S (2020) Thiazolyl-pyrazoline derivatives: In vitro and in silico evaluation as potential acetylcholinesterase and carbonic anhydrase inhibitors. Int J Biol Macromol 163:1970–1988. https://doi.org/10.1016/j.ijbiomac.2020.09.043
Snyder HM, Asthana S, Bain L et al (2016) Sex biology contributions to vulnerability to Alzheimer’s disease: A think tank convened by the Women’s Alzheimer’s Research Initiative. Alzheimers Dement 12:1186–1196. https://doi.org/10.1016/j.jalz.2016.08.004
Somel M, Guo S, Fu N et al (2010) MicroRNA, mRNA, and protein expression link development and aging in human and macaque brain. Genome Res 20:1207–1218. https://doi.org/10.1101/gr.106849.110
Spratt PWE, Ben-Shalom R, Keeshen CM, Burke KJ Jr, Clarkson RL, Sanders SJ, Bender KJ (2019) The Autism-associated gene Scn2a contributes to dendritic excitability and synaptic function in the prefrontal cortex. Neuron 103(673–685):e5. https://doi.org/10.1016/j.neuron.2019.05.037
Stafford N, Wilson C, Oceandy D, Neyses L, Cartwright EJ (2017) The plasma membrane calcium ATPases and their role as major new players in human disease. Physiol Rev 97:1089–1125. https://doi.org/10.1152/physrev.00028.2016
Sun Y, Cheng X, Zhang L, Hu J, Chen Y, Zhan L, Gao Z (2017) The functional and molecular properties, physiological functions, and pathophysiological roles of GluN2A in the central nervous system. Mol Neurobiol 54:1008–1021. https://doi.org/10.1007/s12035-016-9715-7
Turkes C, Arslan M, Demir Y, Cocaj L, Rifati Nixha A, Beydemir S (2019) Synthesis, biological evaluation and in silico studies of novel N-substituted phthalazine sulfonamide compounds as potent carbonic anhydrase and acetylcholinesterase inhibitors. Bioorg Chem 89:103004. https://doi.org/10.1016/j.bioorg.2019.103004
Vazquez F, Legrand S, Windels D (2010) The biosynthetic pathways and biological scopes of plant small RNAs. Trends Plant Sci 15:337–345. https://doi.org/10.1016/j.tplants.2010.04.001
Wakasaya Y, Kawarabayashi T, Watanabe M et al (2011) Factors responsible for neurofibrillary tangles and neuronal cell losses in tauopathy. J Neurosci Res 89:576–584. https://doi.org/10.1002/jnr.22572
Wang R, Reddy PH (2017) Role of glutamate and NMDA receptors in Alzheimer’s disease. J Alzheimers Dis 57:1041–1048. https://doi.org/10.3233/JAD-160763
Wojciechowska A, Braniewska A, Kozar-Kaminska K (2017) MicroRNA in cardiovascular biology and disease. Adv Clin Exp Med 26:865–874. https://doi.org/10.17219/acem/62915
Yu L, Liu Y, Yang H, Zhu X, Cao X, Gao J, Zhao H, Xu Y (2017) PSD-93 attenuates amyloid-beta-mediated cognitive dysfunction by promoting the catabolism of amyloid-beta. J Alzheimers Dis 59:913–927. https://doi.org/10.3233/JAD-170320
Zaidi A, Gao J, Squier TC, Michaelis ML (1998) Age-related decrease in brain synaptic membrane Ca2+-ATPase in F344/BNF1 rats. Neurobiol Aging 19:487–495. https://doi.org/10.1016/s0197-4580(98)00078-5
Zarrin P, Dehghani Ashkezari M, Seifati SM (2021) Liposomal form of L-Dopa and SH-Sy5y cell-derived exosomes modulate the tyrosine hydroxylase/dopamine receptor D2 signaling pathway in parkinson’s rat models. J Mol Neurosci. https://doi.org/10.1007/s12031-021-01853-3
Zhang J, Liu Y, Lu L (2018) Emerging role of MicroRNAs in peripheral nerve system. Life Sci 207:227–233. https://doi.org/10.1016/j.lfs.2018.06.011
Zhang X, Azhar G, Wei JY (2012) The expression of microRNA and microRNA clusters in the aging heart. PLoS One 7:e34688. https://doi.org/10.1371/journal.pone.0034688
Zhao Y, Tan W, Sheng W, Li X (2016) Identification of biomarkers associated with Alzheimer’s disease by bioinformatics analysis. Am J Alzheimers Dis Other Demen 31:163–168. https://doi.org/10.1177/1533317515588181
Zhou H, Liu G (2015) Regulation of density of functional presynaptic terminals by local energy supply. Mol Brain 8:42. https://doi.org/10.1186/s13041-015-0132-z
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
The concept, design, data collection and analysis, and writing processes of the article were performed by HC.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for Publication
The author consents to publication of this manuscript.
Competing Interests
The author declares that there is no conflict of interest with any financial organization or corporation or individual that can inappropriately influence this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ceylan, H. Integrated Bioinformatics Analysis to Identify Alternative Therapeutic Targets for Alzheimer’s Disease: Insights from a Synaptic Machinery Perspective. J Mol Neurosci 72, 273–286 (2022). https://doi.org/10.1007/s12031-021-01893-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-021-01893-9