Abstract
To accurately reveal the scenario and mecahnism of gastrointestinal diseases, the establishment of in vitro models of intestinal diseases and drug screening platforms have become the focus of attention. Over the past few decades, animal models and immortalized cell lines have provided valuable but limited insights into gastrointestinal research. In recent years, the development of intestinal organoid culture system has revolutionized in vitro studies of intestinal diseases. Intestinal organoids are derived from self-renewal and self-organization intestinal stem cells (ISCs), which can replicate the genetic characteristics, functions, and structures of the original tissues. Consequently, they provide new stragety for studying various intestinal diseases in vitro. In the review, we will discuss the culture techniques of intestinal organoids and describe the use of intestinal organoids as research tools for intestinal diseases. The role of intestinal epithelial cells (IECs) played in the pathogenesis of inflammatory bowel diseases (IBD) and the treatment of intestinal epithelial dysfunction will be highlighted. Besides, we review the current knowledge on using intestinal organoids as models to study the pathogenesis of IBD caused by epithelial dysfunction and to develop new therapeutic approaches. Finally, we shed light on the current challenges of using intestinal organoids as in vitro models.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the human body, the gut is one of the most complicated organs. Different research models have been developed to understand intestinal development and disease. Digestion and absorption of nutrients are the main functions of the gut. The gut lumen is surrounded by a single layer of epithelium, which forms a selective barrier to prevent entering of intramural pathogens into the lumen. When IECs are damaged and intestinal function is disrupted, it may lead to diseases [1]. However, intestinal disease research relies primarily on animal studies, which tend to be low-throughput and could not fully refelect physiologic setting of desease. Therefore, in vitro culture of human intestinal tissue offers a new hope for modeling human intestinal diseases.
In recent years, the development of 3D culture systems using stem cells (also called "organoids") has generated widespread utilization of organoids as models for intestinal disease research. Organoids are stem cell-originated and self-organized 3D clusters that have the ability to self-renew and spatially organize. Organoids can mimic the structure and functionality of the originating organ and display the same key features as those that occur in vivo, including tissue architecture, cellular function, and genetic signature [2,3,4,5]. Organoids in 3D culture mode can reach the level of microorgans, and can be combined with cutting-edge technologies such as single-cell sequencing (scRNA-seq) [6, 7], gene editing [8, 9] and microfluidic organ-chips [10], demonstrating high potency in various medical research fields. These properties make 3D organoid culture widely employed in the research on disease model construction [3, 11, 12], therapeutic development [13,14,15], drug screening and toxicity testing [16, 17] and biomolecule delivery [18, 19]. Multiple organoid culture platforms have now been developed for different organs, including the intestine [20], brain [21], liver [22], kidney [23], etc.
Intestinal organoids have the ability to replicate the characteristics and cell types of the native intestinal epithelium [24, 25]. Since the first culture system of mouse small intestinal organoids was generated in 2009 [24], intestinal organoids have been widely applied to the research of a series of diseases, including cystic fibrosis [26], Crohn's disease [27], intestinal cancer [28], etc. In this review, Firstly, we outline pluripotent stem cells (PSCs) and adult stem cells (ASCs)-derived intestinal organoid culture systems and their application in intestinal diseases. A focus is placed on the role of intestinal epithelial dysfunction in the pathogenesis of IBD and current associated treatment method. Secondly, we elucidated the use of intestinal organoids to construct IBD models and develop new therapeutic methods. Finally, the challenges in using intestinal organoids as models are described.
Culture Systems of Intestinal Organoids
The type of intestinal organoids can be divided into two groups: (1) PSCs, including embryonic stem cells (ESCs) or induced pluripotent stem cells (iPSCs) [29], and (2) organ-specific ASCs [30]. Intestinal organoids show all the features of the original intestine in terms of genetic signature and cell type composition [24, 31]. Intestinal organoids are cultured in a unique serum-free and mesenchymal-free growth medium, which requires extracellular supporting substrates and large amounts of additional growth factors. The extracellular matrix (ECM) provides stem cells with tissue integrity and elastic support while transmitting biochemical cues required for regulating cell adhesion, migration, proliferation and differentiation [32]. The key ingredients of the WENR medium commonly used in intestinal organoid culture include: the WNT signaling pathway agonist R-spondin1, epidermal growth factor (EGF) as facilitating the mitosis of stem cells, and Noggin to hold up bone morphogenetic protein (BMP) activity [33]. The growth and development of intestinal organoids are critically dependent on the WENR medium that replicate the signaling pathways in vivo stem cell niche [34, 35]. Furthermore, mouse colon organoid growth requires Wnt-3A, while human intestinal organoid growth requires extra gastrin, nicotinamide, N-acetyl-L-cysteine, the transforming growth factor receptor type I (TGF-βR1) inhibitor A 83–01 and p38 mitogen-activated kinase (MAPK) inhibitor SB202190 [5, 36, 37]. After SB202190 was found to prevent the differentiation of secretory cell lineages in human intestinal organoid culture, insulin-like growth factor 1 (IGF1) and fibroblast growth factor 2 (FGF2) were added to replace this inhibitor [38]. This modification promotes simultaneous multi-lineage differentiation and self-renewal of human intestinal organoids. The percentages of various IEC types in the intestinal organoids are almost comparable with the types in vivo. For a better understanding the function of each cell type, individual culture condition has been established that favor the generation of each one [39]. Although many studies have reported the composition of intestinal organoid culture medium [37, 40, 41], it is constantly optimized according to different schemes to reach the desired result.
ASC-derived Intestinal Organoids
ASC-derived intestinal organoids are acquired by collecting crypts or single Leu-rich repeat-containing G protein-coupled receptor 5 (Lgr5)-positive ISCs (Fig. 1a) and their 3D cultures are grown in the presence of growth factors (R-spondin-1, EGF, Noggin) and laminin-rich Matrigel [42]. Within days, the crypt forms a cysts-like spheroids with a central lumen. Subsequently, it continues to grow into a budding structures and develops into a crypt-villus structure within 1–2 weeks. ISCs and Paneth cells exist at the bottom of the crypt. The villus domain contains secretory goblet cells, enteroendocrine cells (EECs), and absorptive enterocytes [24]. The resulting 3D organoids exhibit normal polarization, with microvilli in the intestinal epithelium facing the central lumen and the basolateral side facing outwards [43]. These organoids are so stable that they can be passed every 5–7 days, allowing long-term culture while maintaining genetic stability.. Interestingly, organoids can arise from any part of the gut and retain the regional identity to their tissues of origin [44]. These organoids can be produced directly human intestinal isolated biopsy specimens and are easy to obtain, which may provide a promising research model for patient-specific therapy and the development of personalized medicine.
PSC-derived Intestinal Organoids
Another exciting method of organoid technology is to generate 3D intestinal organoids from iPSCs and ESCs (Fig. 1b). Spence et al. were the first research group to set up human intestinal organoids (HIOs) from iPSC using continuous manipulations of growth factors that mimic embryonic gut development [45]. There are two major differences between PSC-derived organoids and ASC-organoids: (i) Undifferentiated PSCs can generalize the process of intestinal development, and (ii) differentiation of both the epithelial and submucosal tissues can be theoretically possible. PSC-derived HIOs are produced by guiding differentiation protocols that mimic intestinal developmental processes in vivo [45, 46]. Initially, 2D cultures of FOXA2+SOX17+ endoderm differentiated from PSCs needs addition of activin A, a multifunctional cytokine of the TGF-β family. After endoderm formation, PSCs are induced to differentiate into CDX2+ midgut and hindgut tissues by adding FGF4 and Wnt3a. After four days, gut tube spheroids are formed by polarized columnar epithelium with villus-like structures and crypt-like areas surrounded by mesenchymal cells. During this patterning step, these 3D spheroids are further cultured in Matrigel along with tissue-specific growth factors. They can expand and differentiate into HIOs, which have mesenchymal cells and all major types of epithelial cells [43, 47]. HIOs has a phenotype similar to fetal gut, but lacks regional specificity in ASC-derived organoids [48]. The combination of IECs and mesenchymal cells in HIOs is closer to the original intestinal, which plays an crucial role in epithelial function [49]. These intestinal organoids have already been applied to research areas such as disease modeling and drug development.
Applications of Intestinal Organoids in the Study of Intestinal Diseases
At present, human normal and cancer-derived intestinal epithelial cell lines are only one dimention of cell type, which cannot fully refect the diversity of intestinal epithelial cell types. Although animal models provide a wide range of systemic responses, the gaps between animal models and human diseases may exist and could be one of the key factors leading to clinical trials failures [50]. In contrast, differentiated organoids possesse crypt-villus structures and major IECs types (ISCs, Paneth cells, goblet cells, EECs, enterocytes, transit-amplifying cells, tuft cells, and microfold cells) [24]. Besides, polarized intestinal organoids retain functional characteristics such as absorption of nutrients and working as epithelial barriers and can remain stable in vitro for long periods without genetic or physiological changes [51, 52]. Therefore, intestinal organoids can overcome the limitations of animal models and immortalized cell lines and become reliable in vitro tools to reflect various intestinal physiological states and diseases.
Application of Intestinal Organoids in Regenerative Medicine
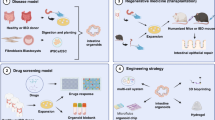
Organ transplantation and repair are promising new treatments for human bowel diseases in recent years (Fig. 2a). Yui et al. first described the ability of orgaoids to regenerate the colonic epithelium when mouse colonic epithelium organoids were transplanted into host animals [39]. They transferred cultured postive GFP colon organoids into ostensibly impaired mouse colon. The grafted donor cells attach to the damaged tissue, after 4 weeks, they rebuild well epithelial tissue and formed a self-renewal crypts of normal function and histology. Watson et al. [53] first constructed an in vivo HIOs engraftment model to assess the role of in vivo environment in the development and maturation of HIOs. HIOs derived from hPSCs were transplanted into the the kidney capsule of immunocompromised mice. After transplantation, intestinal morphology with mucus-filled lumen and well-developed sheets of villi were observed, and each villi had its own central capillary network, demonstrating tissue regeneration. Furthermore, the transplanted HIOs epithelium showed peptide uptake and permeability, confirming the existence of digestive functions [53, 54]. This indicated that although HIOs exhibited fetal intestinal phenotypes early in vitro, HIOs could be transformed into more mature IEC after transplantation in vivo. In addition, the following studies demonstrate that HIOs developed into almost the same tissue as adult intestinal tissue and promote colonic wound repair when they were inoculated onto synthetic polyglycolic/poly-L-lactic acid scaffolds and then implanted into the the abdomen of mice [55] or when directly delivered to mechanically induced colonic mucosal wounds using hydrogel as an injection vehicle [56]. To further diversify the HIOs culture system, Workman et al. [57] utilized a tissue engineering approach to add hPSC-derived neural crest cells (NCCs) components to HIOs to generate human intestinal tissue containing functional enteric nervous system (ENS). This integration offers significant promise for the generation of tissue-engineered intestines containing ENS that controls intestinal motility. Tsai et al. [58] used human ESC-derived intestinal organoids to reveal the machine-processed that regulate regional specificity. They controlled gene expression patterns in specific regions by manipulating the time of exposure to active FGF and WNT signals. Short-term exposure produce duodenal-like organoids, while longer exposure produces ileal organoids. The resulting organoids retain their regional characteristics when cultured in vitro or transplanted in vivo. Short bowel syndrome (SBS) is a clinical syndrome caused by the decrease of small intestine absorption area due to different reasons. Sugimoto et al. [59] produced functional small intestinalized colon (SIC) by taking the place of colon epithelium with small intestinal epithelium through organotransplantation. Transplanting it into SBS rat models ameliorated intestinal failure. It was also confirmed that the graft was a functional small intestine with with absorption. This organ remodeling method provides a feasible strategy for SBS treatment. Overall, this area of research makes it possible to use intestinal organoids as a source of regenerative medicine. With further translational research, HIOs may have the potential to be used for repairing damaged intestines or compensating for the loss of human intestinal function.
Application of Intestinal Organoids in Drug Screening
During the development of drugs for intestinal diseases, intestinal organoids retain the physiological and pathological features of human intestinal segments, which can be applied to a high-throughput drug screening platform to more accurately predict human drug reactions (Fig. 2b). A recent study described robust and repeatable monolayer growth conditions for enteroid monolayers with complete secretion and absorption. A high-throughput compound screening platform was established by miniaturizing it into 96 well plates, and phenotypic screening of approximately 2000 drug candidates was successfully performed [60]. Furthermore, some researchers have developed a series of high-throughput screening platforms using intestinal organoids. Du et al. [61] established a high-density plate organoid screening system using genetically engineered human colon organoids and demonstrated that the system has stable properties for high throughput primary compound screening. Norkin et al. [62] developed a high-throughput, high-content drug discovery system, targeted organoid sequencing (TORNADO-seq). The platform can be used to identify small molecule drugs that induce differentiation between normal and cancer cells in the gut. This approach could not only discover drugs, but also measure gene expression profiles to further elucidate the underlying biological mechanisms. Hirokawa et al. established a low viscosity matrix (LVM) suspension culture platform to produce organoids and tumoroids, which are compatible with engineered bioreactors and can be used to establish biobanks and perform high throughput tumoroid drug sensitivity testing [63]. Researchers have developed a colorectal cancer patient-derived organoids culture system that faithfully recapitulate the genetic changes in cancer samples [64]. In addition, the tumor organoids can better mimic the tumor response to drug stimuli and can detect correlations between cancer genomics and anti-cancer drugs through high-throughput drug screening [65]. Therefore, organoids can be used as a system for high-throughput searching for new drugs and personalizing precision therapy.
Application of Intestinal Organoids in Genetic Engineering
Using intestinal organoid models, the relationship between the genotype of interest and the disease phenotype can be explored (Fig. 2c). Based on gene delivery methods such as lipofection, DNA electroporation and recombinant lentivirus transduction, gene knock down, knockout, knockin or overexpression can be performed in intestinal organoids [66]. Studies have shown that CRISPR-Cas9 system was used to modify the intestinal organoid genome to correct the cystic fibrosis transmembrane conductance regulator (CFTR) locus and repair its function [67]. An organoid culture combined with CRISPR-Cas9 genome editing offers a new way to study to study normal and cancer stem cells. Organoids derived from normal tissue can also model cancer. In one study, the most commonly mutated genes in colorectal cancer (i.e., APC, TP53, SMAD4, KRAS and PIK3CA) were introduced by CRISPR/Cas9 into normal intestinal epithelium to obtain genetically mutated organoids. Transplantation of them into the subperitoneum of the mouse kidney resulted in tumor formation. Compared with other colon cancer models, this model can be better mimic the situation of tumor formation in patients [68]. Therefore, the model may provide new hope for studying colon cancer progression and discovering anti-cancer drugs.
Application of Intestinal Organoids in Host-microbial Interactions
The human gut is ceaselessly challenged by microorganisms and toxins. It is very complex to explore the interaction between microorganisms and the gut epithelium or the destruction of gut epithelium by toxins in vivo. Organoids, by contrast, can be much more easily manipulated. Therefore, intestinal organoids are widely used to explore the mechanisms of epithelial cell destruction caused by microorganisms or toxins (Fig. 2d). By co-culturing organoids with microorganisms or injecting microorganisms into organoids, a model system has been established to study how microorganisms adhere to and invade the intestinal epithelium. Human norovirus (Hunov) and rotavirus are the most common pathogens causing acute gastroenteritis worldwide. Currently, intestinal organoids have been successfully appiled to construct disease models of norovirus and rotavirus infection in vitro [69, 70]. These systems can promote the development of new diagnostics and, vaccines for specific viral pathogens. Other bacterial pathogens can also cause gastrointestinal illness and diarrhea, such as Salmonella enterica and Clostridium difficile. By microinjecting bacterial pathogens into iPSCs or ASCs-derived intestinal organoids to establish the infection, these models can be used to study the interactions between host IECs and microorganisms [71, 72]. These researches may contribute to explain the pathophysiology of infectious diseases and provide new targets for the development of other therapeutic pharmaceutical in the future. Although injecting microorganisms into 3D organoids can study physiological responses in epithelial cells, the technique remains challenging. In recent years, 2D organoid culture system has made it more convenient to study the interaction between IECs and microorganisms and to observe the physiological changes of epithelial cells [73].
The Pressing Need of Intestinal Organoid Models in the IBD Research
IBD is the result of a combination of genetic variations, immune, microbial alterations, and environmental factors [74]. In healthy individuals, intestinal homeostasis is guaranteed by a complicated system called the "intestinal barrier". The intestinal epithelium forms a mechanical barrier, with its apex exposed to the lumen microorganisms and environmental factors, and its base acting on immune cells in the lamina propria to regulate antigen transport (Fig. 3a). However, in IBDs, interference from the intestinal microbiome and breakdown of the intestinal epithelial barrier allow pathogens to invade IECs (Fig. 3b). Further, anomalous immune responses to antigens of the intestinal contents result in persistent inflammatory states [75]. Thus, the intestinal epithelium is located at the crossroads of four factors in the pathogenesis of IBD. In addition, several recent GWASs studies on the pathogenesis of IBD have identified some IBD susceptibility genes related to epithelial barrier functions [76]. Thus, these studies indicate that barrier dysfunction of the intestinal epithelium is one of the critical factors promoting the pathogenesis of IBD.
The Role of Intestinal Epithelial Dysfunction in the Pathogenesis of IBD and the Current Treatment
As a mechanical barrier, IEC is tightly arranged through cell junctions, which consist of tight junctions(TJs), gap junctions(GJs), adhesion junctions(AJs) and desmosomes(DMs). Polymorphism of some IBD susceptible genes adjust the expression and assembly of TJs and AJs, and increase the permeability of IECs. This would promote connections between antigens, luminal microbes, and the host immune system [77]. In order to ensure the continuity of the epithelial barrier after tissue damage, epithelial barrier is rapidly resealed by neighboring cells, and then the wound healing process is delayed. ISCs continue to differentiate and proliferated to form newly developed IECs [78]. At present, several drug candidates have been reported that target epithelial cell proliferation or TJ assembly to correct physical barrier dysfunction. For example, several growth factors, such as EGF, GLP-2 analogue Teduglutide and recombinant HGF, can promote ISC proliferation and have potential effects on mucosal healing. Preliminary studies have shown that these growth factors are moderately effective in treating patients with IBD [79,80,81]. Additionally, one recent study has shown that activating the IL-22-STAT3 pathway enables cell proliferation and increases the expression of secretion factors such as RegIIIβ, RegIIIγ or S100A9 to promote epithelial regeneration. Anti-TNFα therapy can play a therapeutic role in mucosal healing by inhibiting the expression of the endogenous IL-22 inhibitor IL-22BP [82, 83]. In addition, vitamin D deficiency may impair the mucosal barrier and increase the risk of IBD [84]; Estradiol regulated permeability and TJ function, enhancing intestinal epithelial barrier function [85]; Metformin was shown to activate AMPK to promote TJ expression and assembly, which significantly controlled the progression of colitis in mouse models of IBD [86].
In addition to forming a physical barrier between the lumen and the mucosa, IECs also act as a biochemical barrier. Goblet cells secrete and produce abundant highly glycosylated MUC2, which forms mucus matrix on the surface of intestinal epithelium to prevent the invasion of pathogens [87]. Goblet cell reduction is a common pathological characteristic of IBD. It has been shown that the mucous layer becomes thinner and the expression level of MUC2 is decreased due to goblet cell depletion in active UC patients [88]. In addition, Muc2-deficient mice developed spontaneous colitis, which emphasizes the significance of mucin production by goblet cells. The loss of normal MUC2 secretion may directly contribute to chronic inflammation [89]. Goblet cells also secrete and produce another small antiprotease peptide trefoil factors (TFFs). Mashimo et al. [90] suggested that TFF-deficient mice take orally dextran sodium sulfate (DSS) resulted in impaired mucosal healing and death. TFFs are considered as initiators of mucosal healing and can promote the repair of IECs, which plays an significant role in the sustenance of intestinal mucosa [91]. Recent studies have shown that a newly discovered bicarbonate transporter (bestrophin 2, Best2) is specifically expressed in colonic goblet cells. In UC patients, Best2 was found to be down-regulated in their inflamed colon mucosa, which was related to mucous dysfunction and pathogenesis of UC [92, 93]. Enterocytes and Panth cells secrete AMPs to further enhance intestinal barrier function. Enterocytes produce the C-type lectin regenerating islet derived protein IIIγ (REGIIIγ), which can keep bacteria out the surface of the intestinal epithelium, and its production depends on the IECs innate recognition of symbiotic microbial signals [94]. Paneth cells based on intestinal crypts can maintain homeostasis of intestinal stem cell niche [95]. At the same time, these cells maintain host immunity by secreting a variety of antibacterial molecules (defensins, cathelicidins, lysozymes) to form micropores in bacterial membranes, destroying the integrity of the bacteria [96]. Paneth cells are also involved in the pathogenesis of IBD. Several genetic susceptibility factors of CD such as the intracellular pattern recognition receptor NOD2, the key molecule in autophagy ATG16L1, and endoplasmic reticulum stress-induced transcription factor (XBP1) are related to Paneth cell function [97,98,99]. In conclusion, Paneth cells are of great significance in the study of CD pathophysiology. NOD2 and ATG16L1 risk variants are associated with impaired autophagy and AMP secretion in Paneth cells. Loss of Paneth cells function cause CD-like ileitis in mice deficient in both ATG16L1 and XBP1 [99]. Finally, IECs transport secretory immunoglobulin produced by lamina propria cells across the epithelial barrier into the intestinal lumen to maintain the homeostasis of IEC and immune cells. Therefore, understanding the regulatory processes of mucus, AMPs and secretory immunoglobulin will help develop new therapeutic interventions to maintain IEC barrier function. At present, there are several potential therapies to treat the dysfunction of IECs. For example, the prostaglandin E2 receptor 4 (EP4) agonist ONO-4819CD induced mucus production by IECs and improved histological scores in UC patients. Phosphatidylcholine (PC), one component of mucus, is thought to be arranged as a continuous lamellar layers in the apical mucus. Clinical studies have shown that oral PC preparation (rPC) can effectively resolve rectal mucous PC deficiency [100]. Interestingly, the change of gut microbiota are also associated with early gut barrier dysfunction. López-Cauce et al. [101] observed significant differences in bacterial composition in the stool and colonic mucosa from week 5 in wild-type and IL-10−/− mice, and Akkermansia muciniphila deficiency was remarkably observed in IL-10−/−mice. Subsequently, intestinal permeability increased, the number of goblet cells, and the expression of MUC-2, IL-18, WAP Four-Disulfide Core Domain 2(WFDC2) and X-box-binding protein(Xbp-1) decreased significantly. This suggests that Akkermansia muciniphila deficiency and mucin depletion affect intestinal barrier function. Also, studies have shown that the fecal bacterial composition of UC patients in long-term remission is more similar to that of healthy control subjects, and the abundance of Akkermansia muciniphila is positively correlated with the duration of remission [102]. Therefore, Akkermansia muciniphila might play an important role in maintaining the integrity of the intestinal barrier in IBD, and the composition of beneficial gut microbiota should be further investigated in the future.
As one of the regulatory factors of immune homeostasis, IECs can be used as a non-specific antigen presenting cell to process antigens and present them to intestinal immune cells. Under normal physiological conditions, IECs, as a sentinel of innate immunity, recognize commensal bacteria through PRRs including Toll-like receptors (TLRs) and NOD-like receptor (NLRs) [103]. IECs-derived thymic stromal lymphopoietin (TSLP), TGF-β and retinoic acid (RA) react to the symbiotic bacteria signals and transform dendritic cells and macrophages into tolerant phenotypes [87]. These dendritic cells promote the differentiation of naive CD4+ T cells into regulatory T (Treg) cells and maturation B cells into IgA-secreting plasma cells. The IECs can then transport secreted IgA into the intestinal lumen to maintain immune homeostasis [87]. Treg cells and tolerant macrophages secrete IL-10 and other anti-inflammatory cytokines to help maintain immune homeostasis [104]. Under the condition of IBD, IECs barrier function is destroyed, and activated macrophages in lamina propria munch bacterial invaders and produce TNFα, IL-6, IL-12 and IL-23, facilitating inflammatory response [105]. Recent studies have shown that normal IECs preferentially activate CD8+ regulatory T cells through non-classical (MHC) Class I molecules CD1d and costimulatory glycoprotein gp-180 as non-specialized antigen presenting cells, resulting in suppression of immune responses. In contrast, the expression of gp-180 in IECs was abnormal in IBD patients, especially in the CD group. IECs from IBD patients preferentially stimulated proliferation of CD4+ T cell and secretion of IFN-γ through MHC class II [106]. Finally, etrolizumab, an anti-β7 integrin monoclonal antibody blocks the action of αE (CD103) on intestinal T cells to E-cadherin on IECs, thereby inhibiting the retention of CD4+ and CD8+ T cells in the inflammatory sites of IBD [107]. These studies highlighted the need to develop new drugs that specifically target the innate immune dysfunction in IECs for IBD treatment.
Construct IBD Models and Develop New Therapeutic Methods using Intestinal Organoids
Current models for studying the pathogenesis of epithelial barrier dysfunction in IBD have certain limitations and there are only a few therapeutic agents targeting epithelial barrier dysfunction. Recently, Kollmann et al. [108] proposed an optimized in vitro model and demonstrated that human organoids are superior to cell culture models in intestinal barrier studies. In this study, they cultured differentiated intestinal organoids and added cytokine mixture (TNF-α, IFN-γ, IL-1β) to simulate the occurrence of intestinal inflammation. Inflammatory stimulation induces loss of intercellular adhesion, redistribution and loss of junctional proteins resulting in loss of barrier function with consecutive epithelial death. This is consistent with what has been observed in patients with intestinal inflammation. Therefore, the use of differentiated organoids can more reliably simulate inflammation in vivo, which is very helpful for further analysis of the molecular mechanism of inflammatory barrier dysfunction. Organoids provides a powerful tool for powerful tools for modeling IBD and developing new drugs to treat IBD.
Genetic changes affecting IECs function play a crucial role in IBD. Since organoids hold on to the genetic makeup of the primary tissue, the effects of genetic and epigenetic background investigated using patient-derived intestinal organoids on IBD epithelium. Mutations in several genes related to IECs function have been indicated to be associated with epithelial phenotype abnormalities. Two recent studies using intestinal organoids have shown that two proteins known to be concerned with autophagy, ATG16L1 and ATG5, have a protective effect against TNFα-mediated cell death in Paneth cells [109, 110]. This study has provided new insights into the function of Paneth cells and its molecular basis, contributing to the development of new therapeutics for IBD. Moreover, host genetics in some patients with very early-onset IBD (VEO-IBD) that occurs in children under the age of five, plays an important role in disease development. Genetic deficiencies influencing epithelial function have also been found in some patients with these Mendelian IBD (e.g. ADAM17, TRIM22, TTC7A) [111,112,113]. Intestinal organoids can provide sufficient and genetically stable experimental materials for exploring the genetic and phenotypic relationships in IBD. In a recent study, Howell et al. [114] found that changes in DNA methylation in IECs of IBD patients were stable over time compared to controls. This change was partially preserved in patient derived organoid cultures. These organoids can be passaged and frozen and they maintain the stability of their genetic materials. Furthermore, CRISPR/Cas9 technology can import mutations into normal organoids to study the role of gene mutations on IECs function, providing a useful tool for exploring the mechanism of intestinal epithelial function in IBD [115]. Therefore, organoids can be used in the analysis of genetic and epigenetic changes in IBD. Recently, Boye et al. [116] discussed for the first time clinically the possibility of achieving phenotype, genotype, and epigenetic recovery in IBD through molecular manipulation and stem cell treatment. This was achieved by targeting a genetically susceptible IBD risk gene in patient-derived intestinal organoids with CRISPR/Cas9 and then reintroducing the in vitro CRISPR/Cas9-manipulated intestinal organoids into damaged tissues. Stem cells manipulated by CRISPR/Cas9 recolonize the gut and alter the molecular balance, extending remission. This patient-based ISC organoid technology combines stem cell therapy with (epi)drugs and/or CRISPR-Cas9 genome editing technology, opening up new opportunities for molecular therapies. Overall, healthy organoids constructed with IBD associated mutations or organoids from IBD individuality can be used to study the influence of genetic and proinflammatory environments on epithelial barrier function. The organoid models have the potential for diagnosing IBD, finding therapeutic targets, and predicting the risk of cancer recurrence and inflammation.An organoid model of based on simulated IBD conditions has been reported. Intestinal microorganisms and their metabolites, cytokines, immune cells and other intracavitary and lamina propria factors lead to epithelial dysfunction, which leads to the occurrence and development of IBD. The complex pathogenesis of IBD was replicated in vitro to model IBD epithelial barrier dysfunction by introducing these elements into healthy intestinal organoids. For example, IFN-γ and TNF-α induce apoptosis and loss of epithelial cells in intestinal organoids [117, 118]. Therefore, intestinal organoids can regarded as as an effective model to study epithelial cell dysfunction in IBD patients, and may provide a new strategy for correcting epithelial cell dysfunction and promoting survival in IBD patients. Similarly, organoid culture systems can serve as assessing the effects of microorganisms and their metabolites on intestinal epithelium. To explore the mutual effect of Enterohemorrhagic Escherichia coli (EHEC) with epithelium, colonic organoids derived monolayers were differentiated to mimic IBD host–pathogen interactions. Studies have shown that colonizing EHEC reduced the production of mucus and destroys the mucous layer in the colon [119]. Escherichia coli O157:H7 is well known to infect host epithelial cells and hold up the IFN-γ/STAT1 signaling, a mechanism very be alike to the pathogenic features of adherent invasive Escherichia coli (AIEC) separated from patients with CD. Therefore, these intestinal organoids monolayer cultures may also be used as related pathophysiological models to investigate the role of AIEC in IBD biochemical barrier dysfunction [120, 121]. Giri et al. [122] have shown that organoid-based in vitro methods can be used to screen bacteria and their metabolites that inhibit NF-kB activation in the context of IBD and identify strains that alleviate colitis. This makes the study of complex interactions between epithelium and gut microbes or their metabolites another promising application of organoid systems.
IBD is a chronic disease of repeated injury and repair of intestinal epithelium. Epithelial regeneration is an important process to achieve mucosal healing and represent the golden standard for UC and CD therapy. Using intestinal organoids, it has been shown that IL-22 produced by innate lymphoid cells (ILCs) after intestinal damage promoted organoid growth and ISCs proliferation through the IL-22-induced STAT3 phosphorylation pathway. Therefore, this signal transduction pathway may be a prospective therapeutic target for IBD mucosal healing [123]. During intestinal epithelial repair, epithelial cells were transiently reprogrammed to their original state by remodeling the ECM and activating transcription factors YAP/TAZ. The intestinal epithelium transiently acquired fetal phenotype [124]. In addition, the wound-associated epithelium (WAE) loses polarity as the epithelium heals and begins to migrate to the lesion to restore its integrity [125]. A previous study showed that PGE2-PTGER4 signaling promoted post-injury epithelial regeneration by driving ISC differentiation into WAE cells [126]. These findings may help identify new curative targets for IBD to promote IECs regeneration.
During the pathogenesis of IBD, IECs interact with multiple cell types (neuronal cells, mesenchymal cells, andimmune cells). Recent studies suggest that mesenteric fat in CD has a certain relationship with intestinal inflammation, suggesting that mesenteric fat is obviously related to changes in the connective tissue of the intestinal wall (e.g. muscle hypertrophy, fibrosis, and stenosis formation). So mesenteric fat has been described as a characteristic of CD [127]. Recently, co-culture of intestinal organoids with differentiated adipocytes in a transwell has revealed mutual inflammatory activation between these cell types, which provides important new information for future research. Fat cells could provide a potential drug target for the prevention or therapy of CD and obesity-related enterocolitis [128]. Doublecortin-like kinase 1 protein (DCLK1) is a marker of gastrointestinal epithelial cluster cells. In the gastrointestinal crypt 3D culture, DLCK+ tuft cells were observed to completely lose over time. A recently established coculture model of intestinal organoids and primary neurons has shown that intestinal nervous system signaling prolonged the survival of DLCK1+ tuft cells in vitro. DLCK+ tuft cells promote the progression of DSS-induced colitis to poorly differentiated colon cancer in adenomatous polyposis coli (APC)-deficient mice. This suggests that DLCK+ tuft cells may be participated in the initiation of IBD colitis-associated cancer (CAC) [129]. Therefore, this system should provide a new tool for the further development of the CAC model of IBD.
Challenges of Intestinal Organoid Modeling
Although organoids have many advantages over traditional experimental models and their clinical applications and future prospects are encouraging, many issues still need to be resolved. In vitro culture of intestinal organoids requires appropriate ECM and media rich in growth factors. However, the Matrixgel from mouse sarcoma cells is an unambiguouse mixture of proteins, and the growth factors in conditioned culture-medium also contain different number of heterogeneous and undefined components. This increases the introduction of multiple unknown variables into organoids, potentially decreasing their ability to accurately restate physiology and hindering their potential application for therapeutic purposes [130, 131]. Therefore, optimization of organoid culture conditions is still an urgent problem to be solved.
Currently, intestinal organoids transplantation may have the potential to repair injured intestines or intestinal function in humans. This area of research makes it possible to use gut organoids as a source of regenerative medicine. Much needed intestinal organoid scaffolds materials can not only support growth, proliferation, differentiation and inward growth but also induce no acute inflammation or chronic immune response when implanted in vivo. Therefore, the development of new intestinal organoid scaffolds will accelerate the great potential of advances in regenerative medicine.
The transfection efficiency for gene editing in intestinal 3D organoid culture is not the same as that achieved in cell lines. Although detailed protocols have been reported for successfully acquiring gene edited organoids, there is an urgent need for future efforts to overcome the inefficiencies of gene-editing. This could provide a more efficient intestinal disease model to accelerate drug development and clinical treatment of intestinal diseases. In addition, some late-stage tumor organoids are difficult to grow in vitro. It is also unclear whether tumor organoids can capture the full heterogeneous tumor cell population from the original tumor. Therefore, this will also pose a certain challenge for some cancer drug screening.
In recent years, human intestinal organoids have been used as effective models for studying host–pathogen interactions. How to co-culture the dominant anaerobic intestinal microbiota with aerobic cells under aseptic culture conditions is an important challenge. Presently, a promising microfluidic cavity system that combines peristaltic stretching with apical and basolateral fluid flow (intestine-on-a-chip) may be a method to surmount the shortcomings of intestinal organoids [132]. In addition, the intestinal organoid system lacks a natural microenvironment composed of nerve cells, blood vessels, and immune cells. These factors are the key factors in the pathophysiology of intestinal infection, tumor, genetic process. Therefore, further research is needed to incorporate these factors into the model. However, so far, the coculture conditions of intestinal organoids and other cell types are still limited, and further exploration and optimization of co-culture conditions are needed.
In the mucosa of active IBD, the quantity and quality of intestinal crypts were low, which resulted in low production of intestinal organoids and did not guarantee follow-up tests. This suggests that the existing methods for the separation and growth of intestinal organoids are not sufficient for the preparation of the organoids sample bank of active inflammatory IBD. Currently used organoids lack certain elements of complete organs found in the body such as mesenchymal tissue, immune cells and nerve cells. The intestinal epithelial structures of organoids differentiated from iPSCs are more representative of the gut, but it takes longer time to produce and still lacks the immune cells necessary to tackle IBD.
Conclusions
Intestinal organoids are self-organized multicellular culture systems that reproduce the cellular composition, structure, and function of the gut. Intestinal organoids can be cultivated in vitro for a long time and retain host genetic characteristics, which provides hope for the modeling of intestinal diseases. With advances in organoid culture techniques, intestinal organoid modeling techniques have expanded to multiple fields, including intestinal tissue regeneration, genetic manipulation, drug screening analysis, and co-culture systems with various microorganisms. In addition, current intestinal organoids can also simulate multifactorial complex intestinal diseases such as IBD. Currently, intestinal organoids can be used as pathophysiological models to study genetic abnormalities and mucosal injury of IBD. Despite the limitation of organoid technology, the rapid advances in the field of technology and the efforts of scientists in many disciplines will make human organoids more reliable and create extraordinary value in the field of medicine.
Data Availability
None declared.
References
Turner, J. R. (2009). Intestinal mucosal barrier function in health and disease. Nature Reviews. Immunology, 9, 799–809.
Foley, K. E. (2017). Organoids: A better in vitro model. Nature Methods, 14, 559–562.
Dutta, D., Heo, I., & Clevers, H. (2017). Disease modeling in stem cell-derived 3D organoid systems. Trends in Molecular Medicine, 23, 393–410.
Huch, M., Knoblich, J. A., Lutolf, M. P., et al. (2017). The hope and the hype of organoid research. Development (Cambridge, England), 144, 938–941.
Date, S., & Sato, T. (2015). Mini-gut organoids: Reconstitution of the stem cell niche. Annual Review of Cell and Developmental Biology, 31, 269–289.
Stanifer, M. L., & Boulant, S. (2021). Adapting gastrointestinal organoids for pathogen infection and single cell sequencing under biosafety level 3 (BSL-3) conditions. Journal of visualized experiments: JoVE, e62857.
Yin, Y., Liu, P. Y., Shi, Y., et al. (2021). Single-cell sequencing and organoids: A powerful combination for modelling organ development and diseases. Reviews of Physiology, Biochemistry and Pharmacology, 179, 189–210.
Ramakrishna, G., Babu, P. E., Singh, R., et al. (2021). Application of CRISPR-Cas9 based gene editing to study the pathogenesis of colon and liver cancer using organoids. Hepatology International, 15, 1309–1317.
Zhou, H., Wang, Y., Liu, L. P., et al. (2021). Gene editing in pluripotent stem cells and their derived organoids. Stem Cells International, 2021, 8130828.
Osaki, T., Sivathanu, V., & Kamm, R. D. (2018). Vascularized microfluidic organ-chips for drug screening, disease models and tissue engineering. Current Opinion in Biotechnology, 52, 116–123.
Dekkers, J. F., Wiegerinck, C. L., de Jonge, H. R., et al. (2013). A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nature Medicine, 19, 939–945.
Garcez, P. P., Loiola, E. C., Madeiro da Costa, R., et al. (2016). Zika virus impairs growth in human neurospheres and brain organoids. Science (New York, N.Y.), 352, 816–818.
Bouchi, R., Foo, K. S., Hua, H., et al. (2014). FOXO1 inhibition yields functional insulin-producing cells in human gut organoid cultures. Nature Communications, 5, 4242.
Chen, K. G., Mallon, B. S., Park, K., et al. (2018). Pluripotent stem cell platforms for drug discovery. Trends in Molecular Medicine, 24, 805–820.
Liu, F., Huang, J., Ning, B., et al. (2016). Drug discovery via human-derived stem cell organoids. Frontiers in Pharmacology, 7, 334.
Leslie, J. L., Huang, S., Opp, J. S., et al. (2015). Persistence and toxin production by Clostridium difficile within human intestinal organoids result in disruption of epithelial paracellular barrier function. Infection and Immunity, 83, 138–145.
Mittal, R., Woo, F. W., Castro, C. S., et al. (2019). Organ-on-chip models: Implications in drug discovery and clinical applications. Journal of Cellular Physiology, 234, 8352–8380.
Davoudi, Z., Peroutka-Bigus, N., Bellaire, B., et al. (2018). Intestinal organoids containing poly(lactic-co-glycolic acid) nanoparticles for the treatment of inflammatory bowel diseases. Journal of Biomedical Materials Research. Part A, 106, 876–886.
Peng, H., Wang, C., Xu, X., et al. (2015). An intestinal Trojan horse for gene delivery. Nanoscale, 7, 4354–4360.
Pleguezuelos-Manzano, C., Puschhof, J., van den Brink, S., et al. (2020). Establishment and culture of human intestinal organoids derived from adult stem cells. Current Protocols in Immunology, 130, e106.
Lee, C. T., Bendriem, R. M., Wu, W. W., et al. (2017). 3D brain Organoids derived from pluripotent stem cells: Promising experimental models for brain development and neurodegenerative disorders. Journal of Biomedical Science, 24, 59.
Broutier, L., Andersson-Rolf, A., Hindley, C. J., et al. (2016). Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nature Protocols, 11, 1724–1743.
Takasato, M., Er, P. X., Becroft, M., et al. (2014). Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nature Cell Biology, 16, 118–126.
Sato, T., Vries, R. G., Snippert, H. J., et al. (2009). Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature, 459, 262–265.
Barker, N., van Es, J. H., Kuipers, J., et al. (2007). Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature, 449, 1003–1007.
de Poel, E., Lefferts, J. W., & Beekman, J. M. (2020). Intestinal organoids for cystic fibrosis research. Journal of Cystic Fibrosis: Official Journal of the European Cystic Fibrosis Society, 19(Suppl 1), S60-s64.
Lee, C., Hong, S. N., Kim, E. R., et al. (2021). Epithelial regeneration ability of Crohn's disease assessed using patient-derived intestinal organoids. International Journal of Molecular Sciences, 22, 6013.
Szvicsek, Z., Oszvald, Á., Szabó, L., et al. (2019). Extracellular vesicle release from intestinal organoids is modulated by Apc mutation and other colorectal cancer progression factors. Cellular and Molecular Life Sciences: CMLS, 76, 2463–2476.
Tsuruta, S., Uchida, H., & Akutsu, H. (2020). Intestinal organoids generated from human pluripotent stem cells. JMA Journal, 3, 9–19.
Middendorp, S., Schneeberger, K., Wiegerinck, C. L., et al. (2014). Adult stem cells in the small intestine are intrinsically programmed with their location-specific function. Stem Cells (Dayton, Ohio), 32, 1083–1091.
Lancaster, M. A., & Knoblich, J. A. (2014). Organogenesis in a dish: modeling development and disease using organoid technologies. Science (New York, N.Y.), 345, 1247125.
Rezakhani, S., Gjorevski, N., & Lutolf, M. P. (2021). Extracellular matrix requirements for gastrointestinal organoid cultures. Biomaterials, 276, 121020.
Almeqdadi, M., Mana, M. D., Roper, J., et al. (2019). Gut organoids: Mini-tissues in culture to study intestinal physiology and disease. American Journal of Physiology. Cell Physiology, 317, C405-c419.
Artegiani, B., & Clevers, H. (2018). Use and application of 3D-organoid technology. Human Molecular Genetics, 27, R99-r107.
Kriz, V, & Korinek, V. (2018). Wnt, RSPO and Hippo signalling in the intestine and intestinal stem cells. Genes (Basel), 9, 20.
Fujii, M., Matano, M., Nanki, K., et al. (2015). Efficient genetic engineering of human intestinal organoids using electroporation. Nature Protocols, 10, 1474–1485.
Sato, T., Stange, D. E., Ferrante, M., et al. (2011). Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology, 141, 1762–1772.
Fujii, M., Matano, M., Toshimitsu, K., et al. (2018). Human intestinal organoids maintain self-renewal capacity and cellular diversity in niche-inspired culture condition. Cell Stem Cell, 23, 787-793.e786.
Yui, S., Nakamura, T., Sato, T., et al. (2012). Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nature Medicine, 18, 618–623.
Leushacke, M., & Barker, N. (2014). Ex vivo culture of the intestinal epithelium: Strategies and applications. Gut, 63, 1345–1354.
Merker, S. R., Weitz, J., & Stange, D. E. (2016). Gastrointestinal organoids: How they gut it out. Developmental Biology, 420, 239–250.
Mahe, M. M., Sundaram, N., Watson, C. L., et al. (2015). Establishment of human epithelial enteroids and colonoids from whole tissue and biopsy. Journal of Visualized Experiments: JoVE, e52483.
Tsakmaki, A., Fonseca Pedro, P., & Bewick, G. A. (2017). 3D intestinal organoids in metabolic research: Virtual reality in a dish. Current Opinion in Pharmacology, 37, 51–58.
Zachos, N. C., Kovbasnjuk, O., Foulke-Abel, J., et al. (2016). Human enteroids/colonoids and intestinal organoids functionally recapitulate normal intestinal physiology and pathophysiology. The Journal of Biological Chemistry, 291, 3759–3766.
Spence, J. R., Mayhew, C. N., Rankin, S. A., et al. (2011). Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature, 470, 105–109.
McCracken, K. W., Howell, J. C., Wells, J. M., et al. (2011). Generating human intestinal tissue from pluripotent stem cells in vitro. Nature Protocols, 6, 1920–1928.
Wells, J. M., & Spence, J. R. (2014). How to make an intestine. Development (Cambridge, England), 141, 752–760.
Hynds, R. E., Bonfanti, P., & Janes, S. M. (2018). Regenerating human epithelia with cultured stem cells: Feeder cells, organoids and beyond. EMBO Molecular Medicine, 10, 139–150.
Dedhia, P. H., Bertaux-Skeirik, N., Zavros, Y., et al. (2016). Organoid models of human gastrointestinal development and disease. Gastroenterology, 150, 1098–1112.
Foulke-Abel, J., In, J., Kovbasnjuk, O., et al. (2014). Human enteroids as an ex-vivo model of host-pathogen interactions in the gastrointestinal tract. Experimental Biology and Medicine (Maywood, N.J.), 239, 1124–1134.
Zietek, T., Rath, E., Haller, D., et al. (2015). Intestinal organoids for assessing nutrient transport, sensing and incretin secretion. Scientific Reports, 5, 16831.
Jung, P., Sato, T., Merlos-Suárez, A., et al. (2011). Isolation and in vitro expansion of human colonic stem cells. Nature Medicine, 17, 1225–1227.
Watson, C. L., Mahe, M. M., Munera, J., et al. (2014). An in vivo model of human small intestine using pluripotent stem cells. Nature Medicine, 20, 1310–1314.
Finkbeiner, S. R., Hill, D. R., Altheim, C. H., et al. (2015). Transcriptome-wide analysis reveals hallmarks of human intestine development and maturation in vitro and in vivo. Stem Cell Reports, 4, 1140–1155.
Finkbeiner, S. R., Freeman, J. J., Wieck, M. M., et al. (2015). Generation of tissue-engineered small intestine using embryonic stem cell-derived human intestinal organoids. Biology Open, 4, 1462–1472.
Cruz-Acuña, R., Quirós, M., Farkas, A. E., et al. (2017). Synthetic hydrogels for human intestinal organoid generation and colonic wound repair. Nature Cell Biology, 19, 1326–1335.
Workman, M. J., Mahe, M. M., Trisno, S., et al. (2017). Engineered human pluripotent-stem-cell-derived intestinal tissues with a functional enteric nervous system. Nature Medicine, 23, 49–59.
Tsai, Y. H., Nattiv, R., Dedhia, P. H., et al. (2017). In vitro patterning of pluripotent stem cell-derived intestine recapitulates in vivo human development. Development (Cambridge, England), 144, 1045–1055.
Sugimoto, S., Kobayashi, E., Fujii, M., et al. (2021). An organoid-based organ-repurposing approach to treat short bowel syndrome. Nature, 592, 99–104.
Kozuka, K., He, Y., Koo-McCoy, S., et al. (2017). Development and characterization of a human and mouse intestinal epithelial cell monolayer platform. Stem Cell Reports, 9, 1976–1990.
Du, Y., Li, X., Niu, Q., et al. (2020). Development of a miniaturized 3D organoid culture platform for ultra-high-throughput screening. Journal of Molecular Cell Biology, 12, 630–643.
Norkin, M., Ordóñez-Morán, P., & Huelsken, J. (2021). High-content, targeted RNA-seq screening in organoids for drug discovery in colorectal cancer. Cell Reports, 35, 109026.
Hirokawa, Y., Clarke, J., Palmieri, M., et al. (2021). Low-viscosity matrix suspension culture enables scalable analysis of patient-derived organoids and tumoroids from the large intestine. Communications Biology, 4, 1067.
Weeber, F., van de Wetering, M., Hoogstraat, M., et al. (2015). Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proceedings of the National Academy of Sciences of the United States of America, 112, 13308–13311.
van de Wetering, M., Francies, H. E., Francis, J. M., et al. (2015). Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell, 161, 933–945.
Zhao, Q., Guan, J., & Wang, X. (2020). Intestinal stem cells and intestinal organoids. Journal of Genetics and Genomics = Yi Chuan Xue Bao, 47, 289–299.
Schwank, G., Koo, B. K., Sasselli, V., et al. (2013). Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell, 13, 653–658.
Matano, M., Date, S., Shimokawa, M., et al. (2015). Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nature Medicine, 21, 256–262.
Ettayebi, K., Crawford, S. E., Murakami, K., et al. (2016). Replication of human noroviruses in stem cell-derived human enteroids. Science (New York, N.Y.), 353, 1387–1393.
Finkbeiner, S. R., Zeng, X. L., Utama, B., et al. (2012). Stem cell-derived human intestinal organoids as an infection model for rotaviruses. mBio, 3, e00159-00112.
Engevik, M. A., Yacyshyn, M. B., Engevik, K. A., et al. (2015). Human Clostridium difficile infection: altered mucus production and composition. American Journal of Physiology. Gastrointestinal and Liver Physiology, 308, G510-524.
Wilson, S. S., Tocchi, A., Holly, M. K., et al. (2015). A small intestinal organoid model of non-invasive enteric pathogen-epithelial cell interactions. Mucosal Immunology, 8, 352–361.
Fujii, M., & Sato, T. (2021). Somatic cell-derived organoids as prototypes of human epithelial tissues and diseases. Nature Materials, 20, 156–169.
Schirbel, A., & Fiocchi, C. (2010). Inflammatory bowel disease: Established and evolving considerations on its etiopathogenesis and therapy. Journal of Digestive Diseases, 11, 266–276.
Scaldaferri, F., & Fiocchi, C. (2007). Inflammatory bowel disease: Progress and current concepts of etiopathogenesis. Journal of Digestive Diseases, 8, 171–178.
Graham, D. B., & Xavier, R. J. (2020). Pathway paradigms revealed from the genetics of inflammatory bowel disease. Nature, 578, 527–539.
Pastorelli, L., De Salvo, C., Mercado, J. R., et al. (2013). Central role of the gut epithelial barrier in the pathogenesis of chronic intestinal inflammation: Lessons learned from animal models and human genetics. Frontiers in Immunology, 4, 280.
Sturm, A., & Dignass, A. U. (2008). Epithelial restitution and wound healing in inflammatory bowel disease. World Journal of Gastroenterology, 14, 348–353.
Sinha, A., Nightingale, J., West, K. P., et al. (2003). Epidermal growth factor enemas with oral mesalamine for mild-to-moderate left-sided ulcerative colitis or proctitis. The New England Journal of Medicine, 349, 350–357.
Numata, M., Ido, A., Moriuchi, A., et al. (2005). Hepatocyte growth factor facilitates the repair of large colonic ulcers in 2,4,6-trinitrobenzene sulfonic acid-induced colitis in rats. Inflammatory Bowel Diseases, 11, 551–558.
Buchman, A. L., Katz, S., Fang, J. C., et al. (2010). Teduglutide, a novel mucosally active analog of glucagon-like peptide-2 (GLP-2) for the treatment of moderate to severe Crohn’s disease. Inflammatory Bowel Diseases, 16, 962–973.
Pickert, G., Neufert, C., Leppkes, M., et al. (2009). STAT3 links IL-22 signaling in intestinal epithelial cells to mucosal wound healing. The Journal of Experimental Medicine, 206, 1465–1472.
Pelczar, P., Witkowski, M., Perez, L. G., et al. (2016). A pathogenic role for T cell-derived IL-22BP in inflammatory bowel disease. Science (New York, N.Y.), 354, 358–362.
Kong, J., Zhang, Z., Musch, M. W., et al. (2008). Novel role of the vitamin D receptor in maintaining the integrity of the intestinal mucosal barrier. American Journal of Physiology. Gastrointestinal and Liver Physiology, 294, G208-216.
Braniste, V., Leveque, M., Buisson-Brenac, C., et al. (2009). Oestradiol decreases colonic permeability through oestrogen receptor beta-mediated up-regulation of occludin and junctional adhesion molecule-A in epithelial cells. The Journal of Physiology, 587, 3317–3328.
Chen, L., Wang, J., You, Q., et al. (2018). Activating AMPK to restore tight junction assembly in intestinal epithelium and to attenuate experimental colitis by metformin. Frontiers in Pharmacology, 9, 761.
Peterson, L. W., & Artis, D. (2014). Intestinal epithelial cells: Regulators of barrier function and immune homeostasis. Nature Reviews. Immunology, 14, 141–153.
Tytgat, K. M., van der Wal, J. W., Einerhand, A. W., et al. (1996). Quantitative analysis of MUC2 synthesis in ulcerative colitis. Biochemical and Biophysical Research Communications, 224, 397–405.
Van der Sluis, M., De Koning, B. A., De Bruijn, A. C., et al. (2006). Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology, 131, 117–129.
Mashimo, H., Wu, D. C., Podolsky, D. K., et al. (1996). Impaired defense of intestinal mucosa in mice lacking intestinal trefoil factor. Science (New York, N.Y.), 274, 262–265.
Taupin, D., & Podolsky, D. K. (2003). Trefoil factors: Initiators of mucosal healing. Nature Reviews. Molecular Cell Biology, 4, 721–732.
Yu, K., Lujan, R., Marmorstein, A., et al. (2010). Bestrophin-2 mediates bicarbonate transport by goblet cells in mouse colon. The Journal of Clinical Investigation, 120, 1722–1735.
Ito, G., Okamoto, R., Murano, T., et al. (2013). Lineage-specific expression of bestrophin-2 and bestrophin-4 in human intestinal epithelial cells. PLoS ONE, 8, e79693.
Vaishnava, S., Yamamoto, M., Severson, K. M., et al. (2011). The antibacterial lectin RegIIIgamma promotes the spatial segregation of microbiota and host in the intestine. Science (New York, N.Y.), 334, 255–258.
Salzman, N. H., Underwood, M. A., & Bevins, C. L. (2007). Paneth cells, defensins, and the commensal microbiota: A hypothesis on intimate interplay at the intestinal mucosa. Seminars in Immunology, 19, 70–83.
Clevers, H. C., & Bevins, C. L. (2013). Paneth cells: Maestros of the small intestinal crypts. Annual Review of Physiology, 75, 289–311.
Cadwell, K., Liu, J. Y., Brown, S. L., et al. (2008). A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature, 456, 259–263.
Wehkamp, J., Harder, J., Weichenthal, M., et al. (2004). NOD2 (CARD15) mutations in Crohn’s disease are associated with diminished mucosal alpha-defensin expression. Gut, 53, 1658–1664.
Adolph, T. E., Tomczak, M. F., Niederreiter, L., et al. (2013). Paneth cells as a site of origin for intestinal inflammation. Nature, 503, 272–276.
Stremmel, W., Hanemann, A., Ehehalt, R., et al. (2010). Phosphatidylcholine (lecithin) and the mucus layer: Evidence of therapeutic efficacy in ulcerative colitis? Digestive Diseases (Basel, Switzerland), 28, 490–496.
López-Cauce, B., Puerto, M., García, J. J., et al. (2023). Akkermansia deficiency and mucin depletion are implicated in intestinal barrier dysfunction as earlier event in the development of inflammation in interleukin-10-deficient mice. Frontiers in Microbiology, 13, 1083884.
Herrera-deGuise, C., Varela, E., Sarrabayrouse, G., et al. (2023). Gut microbiota composition in long-remission ulcerative colitis is close to a healthy gut microbiota. Inflammatory Bowel Diseases, 29, 1362–1369.
Otte, J. M., Cario, E., & Podolsky, D. K. (2004). Mechanisms of cross hyporesponsiveness to Toll-like receptor bacterial ligands in intestinal epithelial cells. Gastroenterology, 126, 1054–1070.
Yoshimatsu, Y., Mikami, Y., & Kanai, T. (2021). Bacteriotherapy for inflammatory bowel disease. Inflammation and Regeneration, 41, 3.
Chang, J. T. (2020). Pathophysiology of inflammatory bowel diseases. The New England Journal of Medicine, 383, 2652–2664.
Roda, G., Sartini, A., Zambon, E., et al. (2010). Intestinal epithelial cells in inflammatory bowel diseases. World Journal of Gastroenterology, 16, 4264–4271.
Dotan, I., Allez, M., Nakazawa, A., et al. (2007). Intestinal epithelial cells from inflammatory bowel disease patients preferentially stimulate CD4+ T cells to proliferate and secrete interferon-gamma. American Journal of Physiology. Gastrointestinal and Liver Physiology, 292, G1630-1640.
Kollmann, C., Buerkert, H., Meir, M., et al. (2023). Human organoids are superior to cell culture models for intestinal barrier research. Frontiers in Cell and Developmental Biology, 11, 1223032.
Matsuzawa-Ishimoto, Y., Shono, Y., Gomez, L. E., et al. (2017). Autophagy protein ATG16L1 prevents necroptosis in the intestinal epithelium. The Journal of Experimental Medicine, 214, 3687–3705.
Burger, E., Araujo, A., López-Yglesias, A., et al. (2018). Loss of paneth cell autophagy causes acute susceptibility to toxoplasma gondii-mediated inflammation. Cell Host & Microbe, 23, 177-190.e174.
Avitzur, Y., Guo, C., Mastropaolo, L. A., et al. (2014). Mutations in tetratricopeptide repeat domain 7A result in a severe form of very early onset inflammatory bowel disease. Gastroenterology, 146, 1028–1039.
Li, Q., Lee, C. H., Peters, L. A., et al. (2016). Variants in TRIM22 that affect NOD2 signaling are associated with very-early-onset inflammatory bowel disease. Gastroenterology, 150, 1196–1207.
Blaydon, D. C., Biancheri, P., Di, W. L., et al. (2011). Inflammatory skin and bowel disease linked to ADAM17 deletion. The New England Journal of Medicine, 365, 1502–1508.
Howell, K. J., Kraiczy, J., Nayak, K. M., et al. (2018). DNA methylation and transcription patterns in intestinal epithelial cells from pediatric patients with inflammatory bowel diseases differentiate disease subtypes and associate with outcome. Gastroenterology, 154, 585–598.
Fujii, M., Clevers, H., & Sato, T. (2019). Modeling human digestive diseases with CRISPR-Cas9-modified organoids. Gastroenterology, 156, 562–576.
Boye, T. L., Steenholdt, C., Jensen, K. B., et al. (2022). Molecular manipulations and intestinal stem cell-derived organoids in inflammatory bowel disease. Stem Cells (Dayton, Ohio), 40, 447–457.
Farin, H. F., Karthaus, W. R., Kujala, P., et al. (2014). Paneth cell extrusion and release of antimicrobial products is directly controlled by immune cell-derived IFN-γ. The Journal of Experimental Medicine, 211, 1393–1405.
Grabinger, T., Luks, L., Kostadinova, F., et al. (2014). Ex vivo culture of intestinal crypt organoids as a model system for assessing cell death induction in intestinal epithelial cells and enteropathy. Cell Death & Disease, 5, e1228.
In, J., Foulke-Abel, J., Zachos, N. C., et al. (2016). Enterohemorrhagic Escherichia coli reduce mucus and intermicrovillar bridges in human stem cell-derived colonoids. Cellular and Molecular Gastroenterology and Hepatology, 2, 48-62.e43.
Martinez-Medina, M., & Garcia-Gil, L. J. (2014). Escherichia coli in chronic inflammatory bowel diseases: An update on adherent invasive Escherichia coli pathogenicity. World Journal of Gastrointestinal Pathophysiology, 5, 213–227.
Ho, N. K., Crandall, I., & Sherman, P. M. (2012). Identifying mechanisms by which Escherichia coli O157:H7 subverts interferon-γ mediated signal transducer and activator of transcription-1 activation. PLoS ONE, 7, e30145.
Giri, R., Hoedt, E. C., Khushi, S., et al. (2022). Secreted NF-κB suppressive microbial metabolites modulate gut inflammation. Cell Reports, 39, 110646.
Lindemans, C. A., Calafiore, M., Mertelsmann, A. M., et al. (2015). Interleukin-22 promotes intestinal-stem-cell-mediated epithelial regeneration. Nature, 528, 560–564.
Yui, S., Azzolin, L., Maimets, M., et al. (2018). YAP/TAZ-dependent reprogramming of colonic epithelium links ECM remodeling to tissue regeneration. Cell Stem Cell, 22, 35-49.e37.
Seno, H., Miyoshi, H., Brown, S. L., et al. (2009). Efficient colonic mucosal wound repair requires Trem2 signaling. Proceedings of the National Academy of Sciences of the United States of America, 106, 256–261.
Miyoshi, H., VanDussen, K. L., Malvin, N. P., et al. (2017). Prostaglandin E2 promotes intestinal repair through an adaptive cellular response of the epithelium. The EMBO Journal, 36, 5–24.
Mao, R., Kurada, S., Gordon, I. O., et al. (2019). The mesenteric fat and intestinal muscle interface: Creeping fat influencing stricture formation in Crohn’s disease. Inflammatory Bowel Diseases, 25, 421–426.
Takahashi, Y., Sato, S., Kurashima, Y., et al. (2017). Reciprocal inflammatory signaling between intestinal epithelial cells and adipocytes in the absence of immune cells. eBioMedicine, 23, 34–45.
Westphalen, C. B., Asfaha, S., Hayakawa, Y., et al. (2014). Long-lived intestinal tuft cells serve as colon cancer-initiating cells. The Journal of Clinical Investigation, 124, 1283–1295.
Hughes, C. S., Postovit, L. M., & Lajoie, G. A. (2010). Matrigel: A complex protein mixture required for optimal growth of cell culture. Proteomics, 10, 1886–1890.
Noben, M., Vanhove, W., Arnauts, K., et al. (2017). Human intestinal epithelium in a dish: Current models for research into gastrointestinal pathophysiology. United European Gastroenterology Journal, 5, 1073–1081.
Jalili-Firoozinezhad, S., Gazzaniga, F. S., Calamari, E. L., et al. (2019). A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nature Biomedical Engineering, 3, 520–531.
Funding
This work was supported by National Natural Sciences Foundation of China (32070138; 82060105), Talent Introduction Plan of the Lanzhou University Second Hospital (yjrckyqdj-2022–01; yjrckyqdj-2021–03), the Central University Excellent Youth Team Project (537000–561223001), and Cuiying Scientific and Technological Innovation Program of the Lanzhou University Second Hospital (CY2021-MS-A05).
Author information
Authors and Affiliations
Contributions
Wenxiu Liu: writing original draft; Qian Wang, Yanrui Bai, Han Xiao, Zhunduo Li, Yan Wang: participate in the discussion; Qi Wang, Jing Yang, Hui Sun provided advice and revised the manuscript.
Corresponding authors
Ethics declarations
Ethical Approval
This article does not contain any studies with human or animal participants performed by any of the authors.
Consent to Participate
None declared.
Consent to Publish
None declared.
Conflicts of Interest
The research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, W., Wang, Q., Bai, Y. et al. Potential Application of Intestinal Organoids in Intestinal Diseases. Stem Cell Rev and Rep 20, 124–137 (2024). https://doi.org/10.1007/s12015-023-10651-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-023-10651-w