Abstract
Diabetes mellitus (DM), a metabolic and endocrine condition, poses a serious threat to human health and longevity. The emerging role of gut microbiome associated with bioactive compounds has recently created a new hope for DM treatment. UHPLC-HRMS methods were used to identify these compounds in a poly herbal ethanolic extract (PHE). The effects of PHE on body weight (BW), fasting blood glucose (FBG) level, gut microbiota, fecal short-chain fatty acids (SCFAs) production, and the correlation between DM-related indices and gut microbes, in rats were investigated. Chebulic acid (0.368%), gallic acid (0.469%), andrographolide (1.304%), berberine (6.442%), and numerous polysaccharides were the most representative constituents in PHE. A more significant BW gain and a reduction in FBG level towards normal of PHE 600 mg/kg treated rats group were resulted at the end of 28th days of the study. Moreover, the composition of the gut microbiota corroborated the study’s hypothesis, as evidenced by an increased ratio of Bacteroidetes to Firmicutes and some beneficial microbial species, including Prevotella copri and Lactobacillus hamster. The relative abundance of Bifidobacterium pseudolongum, Ruminococcus bromii, and Blautia producta was found to decline in PHE treatment groups as compared to diabetic group. The abundance of beneficial bacteria in PHE 600 mg/kg treatment group was concurrently associated with increased SCFAs concentrations of acetate and propionate (7.26 nmol/g and 4.13 nmol/g). The findings of this study suggest a promising approach to prevent DM by demonstrating that these naturally occurring compounds decreased FBG levels by increasing SCFAs content and SCFAs producing gut microbiota.
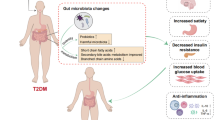
Graphical Abstract
Flow chart summarizing research on the dual therapeutic approach to diabetes mellitus via bioactive chemicals found in a poly herbal extract and the management of gut microbiota in relation to DM. (SCFAs, short chain fatty acids; SMB53, a genus of bacterial microbiota of small intestine; LPS, lipopolysaccharide)

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diabetes mellitus (DM) is a serious and complicated metabolic condition that lasts for a long time [1]. There are currently over 536.6 million people with diabetes globally, despite the availability of numerous therapeutic approaches, according to a recent report released from the International Diabetes Federation for 2022 [2]. A recent study predicts that without quick and efficient interventions, the occurrence of diabetes will grow sharply, reaching 11.3% (642.7 million individuals) by 2030 and 12.2% (783.2 million people) by 2045 [2]. The prevalence of diabetes in India is rising rapidly, and it now affects approximately 8.7% of people aged 20–70. A longer life expectancy, unhealthy lifestyles, poor diets, smoking, and rapid urbanization are all factors in the rise in diabetes cases [3]. It costs at least $966 million a year, which is 9% of all global health spending [4]. Drugs like pioglitazone, acarbose, metformin, and others [5], which are sold on the market, can help prevent diabetes. The potential adverse side effects of these anti-diabetic medicines, however, must not be neglected [6]. Innovative, effective, and secure anti-diabetic medications are desperately needed to combat the rising prevalence of diabetes. Therefore, finding alternative active substances to treat this condition that are non-toxic and have fewer side effects is an urgent priority.
Ayurveda is recognized as one of the oldest of the traditional systems of medicine (TSMs) accepted worldwide because it has drawn considerable attention in recent years due to its holistic approach to disease management [7,8,9]. Ancient Indian medicines, which are defined through systematic and fundamental regulation by the utilization of multiple medicinal plants and their polyherbal formulations, contain multiple active components, act through multiple pathways and targets, and thus have an increasing impact on the development of anti-diabetic drugs [10,11,12,13]. Here we texted some of the medicinal plants with reported anti-diabetic activity potential (Aegle marmelos, Andrographis paniculata, Citrullus colocynthis, Gymnema sylvestre, Momordica charantia, Ocimum sanctum, Swertia chirayita, Trigonella foenum graceum, etc.) [9]. The Indian traditional medical system has developed several effective anti-diabetic medicines for the treatment of diabetes in accordance with advances in medicinal technology [12], for example, Madhumeh churna, Triphala churna, Vilva kudineer, Diabet, Diasol, Dianex, Diakyur, Diasulin, Diabecure, and others [3, 9]. Here, we formulated the PHE from our earlier study that summarizes the pharmacological actions like antimicrobial, anti-infalmmatory, and antidiabetic, as well as the antioxidant potential that would be due to the bioactive molecules of the ingredient plants [3]. The crude source, short name, and family of six medicinal plants in PHE are illustrated herewith, with their voucher specimens granted: fruits of Terminalia chebula Retz. (TC), Combretaceae (No. Combreta. 2020/02); fruits of Terminalia bellerica (Gaertn.) Roxb. (TB), Combretaceae (No. Combreta. 2020/02); whole herb of Andrographis paniculata (Buem.f.) Nees. (AP), Acanthaceae (No. Acantha. 2020/01); stem of Berberis aristata DC. (BA), Berberidaceae (No. Berberida. 2020/02); leaves of Nyctanthes arbor-tristis L. (NA), Oleaceae (No. Olea. 2020/01); and leaves of Premna integrifolia L. (PI), Lamiaceae (No. Lamia. 2020/01) [3].
Modern metabolic profiling techniques are highly sensitive and selective, allowing for the successful acquisition of novel insights into the medicinal compounds of plants [14]. Metabolomics, which is defined as both the quantitative and qualitative examination of all small metabolites in a biological specimen whose presence is influenced at a specific time by specific conditions [15], grows our understanding of daily life by highlighting how the variety of chemicals in herbal remedies is still far beyond the researchers’ thoughts, even on plant matrices, which are thought to have already undergone extensive research [16]. Therefore, in this paper, ultra-high-performance liquid chromatography (UHPLC) techniques were applied to PHE in conjunction with a high-resolution accurate mass spectrometry system (HRMS), as the examination of the chemical constituents of plant extracts stands as an important component for understanding their bioactivity, both in terms of efficacy and safety. The method has the potential to be much more sensitive than the current standard for disclosing secondary metabolites because of its ability to simultaneously measure mass with high precision and identify characteristic fragments. The phytoconstituents may influence our body physiology through various mechanisms, even through modulation of the gut microbiota and the synthesis of micronutrients [17]. The gut microbiome is made up of eukaryotes, archaea, and bacteria. Bacterial phyla are the most common type [18]. Maintaining the diversity and abundance of the human gut microbiota has been found to be important for maintaining intestinal homeostasis [19, 20]. This is because the microbiota protects the host from pathogens by competing for niches and nutrients, boosting immune functions, and regulating metabolism. The gut microbiota is a dynamic ecosystem that is constantly changing and interacting with the host in a mutualistic manner. It may, in fact, have an impact on a number of physiological processes linked to human health, including immune system function, defense against infections, synthesis of micronutrients, and control of glucose and fat metabolism [17]. Therefore, modulating the gut microbiota may be an effective means of warding off metabolic and inflammatory disorders. Obesity, beta-cell exhaustion, insulin resistance, and T2DM are all conditions that have been linked to changes in the gut microbiota, according to animal studies [21]. Furthermore, observational investigations have demonstrated that there is evidence of altered microbial communities in the gut in prediabetic or DM patients in comparison to healthy subjects, a condition referred to as “gut microbiota dysbiosis” [22, 23]. Furthermore, “gut microbiota dysbiosis” may disrupt the equilibrium between commensal species and different types of pathogens, and it may also reduce the release of metabolic SCFAs and antimicrobial molecules like bacteriocins [24]. However, there is now more evidence on the activities of microbial species associated with the gut microbiota’s protective effect against DM as a metabolic disorder [25].
The management of DM through gut microbiota may be based on the following mechanisms: (1) keeping the intestinal barrier strong; (2) stopping the movement of bacteria and, as a result, reducing systemic inflammation (endotoxemia); and (3) generation of SCFAs (acetate, propionate, and butyrate), which can change metabolic pathways [26]. In addition to SCFAs and bacterial toxins, the microbiota may also affect the gut barrier through the endocannabinoid system [27, 28] and the metabolism of bile acids (BAs) [27]. There is growing evidence that DM is strongly linked to gut microbiota dysbiosis, damages the gut wall, and encourages the movement of endotoxemia from the gut into the bloodstream, which leads to inflammation, autoimmune responses, and oxidative stress [29, 30]. Numerous studies showed that people with different types of DM had an increased abundance of Firmicutes and a decreased abundance of Bacteroidetes, which had a negative impact on their metabolic regulation of nutrients, their glucose tolerance, and their inflammatory responses [31]. Anti-inflammatory SCFA levels, specifically butyrate, were lower in people with diabetes because they had a decreased abundance of SCFA-producing microorganisms such as Roseburia intestinalis and Faecalibacterium prausnitzii [30, 32, 33]. The mechanism of action of SCFAs mainly depends on the intestinal bacteria, as illustrated in the flow chart figure, which may be more likely to enter the systemic circulation if SCFA synthesis is impaired, which may trigger or exacerbate integrated inflammatory responses [34]. Positive effects of SCFAs for diabetics include lowering serum glucose levels, improving insulin sensitivity, decreasing inflammation, and increasing the production of the protective hormones glucagon-like peptide 1 (GLP-1) [35] and peptide YY (PYY) that increase satiety feelings through the gut-brain axis [36]. In the liver, SCFAs have been shown to decrease glycolysis and gluconeogenesis and increase glycogen synthesis and fatty acid oxidation. Additionally, SCFAs have been shown to improve glucose uptake in skeletal muscle and adipose tissue by elevating the expression of GLUT4 through AMP kinase (AMPK) activity. Moreover, in the skeletal muscle, SCFAs reduce glycolysis with a consequent accumulation of glucose-6-phosphate and an increase in glycogen synthesis [36].
It makes sense to think that these results will help us understand how these bioactive compounds work together to prevent DM. They will likely provide strong evidence for at least some of the metabolic diseases that this PHE could potentially prevent. In this study, we used high-throughput 16S rRNA gene sequencing to look at how PHE affected the gut microbiota of DM rats and find key genera that were closely linked to PHE treatment. This helped us figure out if the gut microbiota was an appropriate target for PHE during the improvement of DM symptoms. Additionally, we correlated diabetic parameters with gut microbiota enrichment linked to increased SCFA content in order to ameliorate DM.
Materials and Methods
Chemicals and Drugs
Details of all the chemicals utilized for the preparation of polyherbal extract (PHE) and further required chemicals and drugs for evaluations were described in our previous study [3]. The analytical-grade solvents used in the study were bought from Sigma Aldrich through local vendors.
Preparation and Evaluation of PHE
All the crude materials for the preparation of PHE were scientifically collected from the medicinal garden at BHU, Varanasi. The crude materials were identified and authenticated taxonomically. The preparation steps, preliminary qualitative phytochemical tests, GC–MS fingerprint analysis of the prepared PHE, and their in-vitro anti-inflammatory activity with potential antioxidant values were reported in our previous study [3]. The already-prepared PHE from the previous study was used for the present study as well.
UHPLC-HRMS Analysis
Analyses were performed using a modified version of the high-resolution accurate mass spectrometry system (HRMS) published by Abu-Reidah [37], and data were collected with the help of a Thermo Scientific Orbitrap Eclipse Tribrid mass spectrometer (MS) running on the small molecule application H-ESI mode. To achieve improved MS detection of ethanol extract derived from PHE, a suitable full MS/ddMS2 (OT HCD) technique was created, which included the use of a UHPLC system (Dionex Ultimate 3000 RS Series). These were the source parameters utilized here, such as formic acid at 0.1 percent in either water (A), acetonitrile (B), or methanol (C) as the mobile phase and a spray voltage of + ve 3.5 kV and − ve 2.5 kV. Another factor, like the ion transfer tube temperature of 325 °C, the sheath gas flow rate of 50 arb, the vaporizer temperature of 350 °C, the aux gas flow rate of 10 arb, as well as the sweep gas flow rate of 1 arb, were also the corresponding determinants of the present study. The initial charge state was determined to be 1. MS2 spectra of PHE glycosylated with various sugars were obtained by using the stepped collision energy mode (SCEM) at collision energies of 30, 45, and 60% of the maximum achievable in high-energy collision-induced dissociation (HCD). The data was tracked, processed with Xcalibur 4.1 software from Thermo Fisher Scientific SII, and collected.
Ethical Approval
The laboratory animal care and use manual was followed for all experimental rats utilized in this investigation. All experimental protocols were approved by the Central Animal Ethical Committee of Banaras Hindu University (Reg. No. 542/GO/Rebi//S/02/CPCSEA dated 26.5.2017), which provided an approval number (Approval number: Dean/2021/IAEC/2560) for this research.
Induction of Diabetes Mellitus
Rats were induced with DM using the established animal model outlined by Hilary [38]. Male Wistar rats (n = 36) weighing between 180.32 and 230.54 g were acclimated and then starved on water alone for 12 h. Prior to receiving streptozotocin, the rats’ fasting blood sugar levels were measured and recorded. After that, we induced insulin resistance in all rats except normal group rats by administering for 24 h orally a 5% glucose solution to prevent mortality in rats after an I.P. injection of streptozotocin single at pH 4.5 dissolved in citrate buffer at a dose of 60 mg/kg. After 72 h of inducing DM in rats, we tested their FBG levels to confirm the diagnosis. Diabetic rats had a FBG level higher than 250 mg/dl.
Animal Groups and Treatment Protocol with PHE
Thirty-six male Wistar rats chosen in this study were divided into six groups of six. These classifications were created below:
-
GP-1: Normal rats + drinking water
-
GP-2: Diabetic control untreated rats
-
GP-3: Diabetic rats + 250 mg/kg BWt of metformin (oral gavage)
-
GP-4: Diabetic rats + 300 mg/kg BWt of PHE (oral gavage)
-
GP-5: Diabetic rats + 600 mg/kg BWt of PHE (oral gavage)
-
GP-6: Diabetic rats + 1700 mg/kg BWt of DMR (oral gavage)
Each rat group (N = 6)’s feces was collected at the completion of the entire study and frozen at − 80 °C for later assessment. Image (Fig. 1) shows the treatment plan of the study.
Body Weight and Glucose Estimation
Every seventh day, the body weight was measured. For blood glucose determination, blood was withdrawn using the tail snipping technique, and the rats’ fasting blood glucose (FBG) concentrations were measured weekly using a glucometer (One Touch Select Plus, Lifescan, Europe) based on the glucose oxidase method [39].
Fecal DNA Extraction and Sequencing
By using a DNA extraction kit (Clevergene Biocorp Private Limited, Banaswadi, Bangalore, India), the complete genomic DNA of microorganisms in the fecal samples of several rat groups was retrieved. The concentrations of the DNA samples were tested for purity using a NanoDrop™ 2000 Spectrophotometer (ND2000, Thermo Scientific). The bacterial 16S rRNA gene was amplified using a forward and a reverse primer like 5′-CCTACGGGNGGCWGCAG-3′ and 5′-GACTACHVGGGTATCTAATCC-3′, respectively, on the highly variable region V3–V4 using 25 ng of DNA. Designed 341F and 785R primers are present in the process at 100 nM final concentrations, together with KAPA HiFi HotStart Ready Mix [40]. In order to prepare the genome sequencing libraries, an additional 8 cycles of PCR utilizing Illumina barcoded adapters were carried out. The sequence data was produced by Illumina MiSeq, and FastQC and MultiQC software were used to assess its quality.
Composition and Diversity of the Gut Microbiota
Alpha and beta diversity parameters were applied to characterize variations in species diversity within as well as between habitats, respectively, in order to conduct an all-encompassing evaluation of species’ overall diversity. Alpha diversity refers to diversity on a local scale, describing the species diversity (richness) within a functional community. Beta diversity, on the other hand, describes the amount of differentiation between species communities. Different indices are used to characterize the alpha diversity of the gut microbiota. In this study, species richness was characterized by the Chao1 estimator, the ACE estimator, and observed species indices; species diversity was characterized by the Shannon, Simpson, InvSimpson, and Fisher indices. The fecal gut flora was identified by sequencing the 5′ and 3′ untranslated regions of the 16S rRNA gene of bacteria on the Illumina MiSeq technology. GreenGenes v.13.8–99 [41] was used to filter, analyze, merge, and classify the acquired sequences into taxonomic outlines. Following this, 97% sequence similarity was used to group the contigs into OTUs.
Determination of SCFAs by HPLC in the Fecal Sample
The fecal sample SCFA concentration was determined using a modified version of the procedure described by Sassaki [42]. Each of the six groups of rats was sampled for its feces. After centrifuging each feces pellet at 4 °C over 5 min at 13,000 rpm, we mixed in 1 ml of sterile 18Ω de-ionized water and mixed at 3200 rpm to homogenize the mixture. Whatman™ UNIFLO™ 13 mm sterile syringe filters with 0.2 µm PES Filter Media were used to collect and filter the supernatant. The filtrate underwent HPLC-based SCFA analysis. Aminex HPX-87H carbohydrate column (4.6 × 250 mm cartridge), particle size 4 µm, flow rate 0.6 ml/min, run time 60 min, HPLC system (Waters 2000). The eluent employed was 5 mM H2SO4, and the column temperature was adjusted to 50 °C. Injection standards were generated by serially diluting a 10 mM stock solution of acetate and propionate and then filtering the resulting solution using Whatman™ UNIFLO™ 13 mm sterile syringe filters with 0.2 µm PES Filter Media. The concentration of each sample was determined by plotting and analyzing the customary plots. GraphPad Prism 5 was used to generate the graphs representing the results of the various test groups.
Statistical Analysis
GraphPad Prism 5 was used for statistical analysis. With the help of Tukey’s test and one-way analysis of variance (ANOVA) for various comparisons, the significance of differences between groups was examined. Additionally, the Fishers exact test significance was run to evaluate the impact of each differentially abundant taxon, or OUT. A significant difference was defined as one with a p value of < 0.05. To find out whether there are any connections between metabolic parameters related to diabetes and gut microbiota, CCA analysis was performed using XLSTAT 2023.03.23.
Result
Bioactive Components and UHPLC-HRMS Metabolite Profiling of PHE
Through early qualitative tests and GC–MS analyses, it was determined that PHE exhibited synergistic antioxidant capacity as well as strong in vitro anti-inflammatory activity because of the presence of several bioactive components in our previous study [3]. Therefore, PHE can be utilized as a complementary therapy for the treatment of inflammatory conditions and metabolic diseases like diabetes. Through UHPLC-HRMS analysis of bioactive chemicals and regulation of gut bacteria and SCFAs in diabetic rats, we were able to extend the results of prior work on PHE.
Metabolic profiling led to the identification of numerous phytochemicals in PHE, including phenolic acids, organic acids, flavonoids, triterpenes, fatty acids, sesquiterpenes, coumarins, lignans, and sugars (Supplementary Table S1). Annotation sources included predicted composition, mzCloud search, chemSpider search, and massList search (Arita Lab 6549 Flavonoid Structure Database). The UHPLC-HRMS metabolite fingerprint revealed numerous bioactive compounds provided in the supplementary file along with the article. Out of 1759 screening checks, flavonoids were identified in 361 checks, with 9.9% calculated on the basis of the total peak percent area (Supplementary Table S1). Biomarker compounds like chebulic acid (0.368%), gallic acid (0.759%), ellagic acid (0.469%), andrographolide (1.304%), berberin (6.442%), kaempferol (0.069%), quercetin (0.020%), apigenin (0.040%), and other bioactive compounds like luteolin (0.034%), oxyberberine (0.141%), rutin (0.017%), genistein (0.022%), ferulic acid (0.148%), hesperidin (0.336%), and isorhamnetin (0.015%) were identified in good proportion from PHE shown in supplementary Table S1. The total metabolite profile of PHE was expressed in a total ion chromatogram at the various retention times (RT) for bioactive compounds obtained in both the negative and positive modes of analysis (Fig. 2). In this study, MS chromatograms with MS1 and MS2 peaks of well-known biomarker compounds like chebulic acid (Fig. 3), gallic acid (Fig. 4), andrographolide (Fig. 5), and berberine (Fig. 6) were of particular interest.
Body Weight of Diabetic Rats Administered PHE
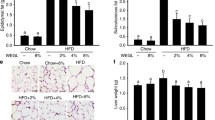
The animals’ body weights dropped significantly after streptozotocin treatment (Fig. 7A), in contrast to the normal control group, which gained weight. Diabetic rats treated with PHE at doses of 300 mg/kg and 600 mg/kg gained weight. The body weight of GP-4 and GP-5 rats increased more significantly than that of GP-3 and GP-6 rats (Supplementary Table S2). Diabetic rats exhibited a therapeutic change, as evidenced by a dose-dependent alteration in body weight across treatment groups.
Fasting Blood Glucose Level
Male rats in the untreated group had FBG levels above 250 mg/dl for the entirety of the experiment, even after 72 h of streptozotocin induction (Fig. 7B). PHE at 300 mg/kg, 600 mg/kg, metformin at 250 mg/kg, and DMR at 1700 mg/kg body weight all reduced FBG levels, as seen in Supplemental Table S3. FBG levels were reduced to levels similar to those in the non-diabetic group in the groups of rats given PHE at 300 mg/kg and 600 mg/kg, as well as metformin and DMR (Fig. 7B). PHE was found to be more effective than metformin and DMR in lowering FBG levels in the present animal models. At 600 mg/kg of body weight, we saw the greatest change in FBG levels.
Effect of PHE on Gut Microbiota
Sequence score cards (Supplementary Figs. S1A and S1B) were used to verify that an entire amplification at around 450 bp was present in each of the six samples (Supplementary Table S4). The recorded gel picture (Fig. 7C) demonstrates the QC of samples from each group of experiments in our study. High coverage was attained across all rat groups, as shown by the distribution plot of sequence (contig) length (Fig. 7D) and the trend for each of the rarefaction curves (Fig. 7E). The diversity estimators and summary statistics showed that the gut microorganisms in the normal group (GP-1), diabetes group (GP-2), and treatment groups like metformin (GP-3), PHE (GP-4 and GP-5), and DMR (GP-6) were significantly different from one another. The diversity of the treated rats was significantly higher than that of the normal and diabetic rat groups. Based on the results of the microbial diversity analysis, we can see in Fig. 8 that there are 10 main phyla covered in the present study of gut microbiota.
Alpha Diversity
Table 1 demonstrates the non-significant prevalent alpha diversity indices in the different groups of experiments (GP-1 to GP-6). GP-5 rats’ feces demonstrated similar ACE and Chao1, as well as observed species indices towards GP-1 but not GP-2. GP-4 and GP-6 got similar outcomes as GP-2 in these tests (supplementary Fig. S2). Thus, GP-5, as well as other investigational treatment groups, had rich microbial communities. GP-2 diabetic rats had higher Fisher indices than GP-1 control rats. Fisher indices for the treatment animals (GP-5) were equivalent to those of metformin-treated rats (GP-3) and restored the diversity of bacteria to normal (GP-1). Table 1 shows that the treatment groups GP-4 and GP-6 have opposite Fisher indices. Thus, sequencing was sufficient to assess gut microbiota diversity. Table 1 shows that the Simpson, InvSimpson, and Shannon indices for both GP-5 and GP-1 did not differ significantly. Community richness and rarely-seen amplicon sequence variations (ASVs) affect the Shannon index more. A total of 28 days of high-dose PHE (GP-5) treatment may improve rat feces species diversity.
Beta Diversity
This study has many microbial species. PCoA cluster analysis was used to investigate gut microbial populations in the feces of experimental rats. Based on taxonomical distribution, the principal coordinate analysis (PCoA) was employed to compare microbiota architectures. Gut microbial structure differed among rat groups. The two-axis principal component assessments explained 50.7% and 21.5% of the total variances (Fig. 8A). Axis 1 shows that GP-1 and GP-5 have a significant difference in PCoA1 compared to GP-2. GP-1 and GP-5 exhibit similar gut microbiota diversity at PCoA1, while GP-3, GP-4, followed by GP-6 remain close to GP-2. In addition, GP-3 and GP-4 were nearer to GP-2 on PCoA2, but GP-1, GP-5, followed by GP-6 were significantly different on Axis 2. Thus, the GP-3 and GP-4 in the present study had fewer microbial differences on PCoA2 and PCoA1 (Fig. 8A). The results of GP-1 and GP-5 were more similar, suggesting that rats’ intestinal microbial communities might be altered by gavage with 600 mg/kg body weight of PHE for 28 days.
Comparison of the Gut Microflora Community Composition
According to the results of the metagenomic study, the 10 phyla of the microbiome are the representatives of the present study like Actinobacteria, Bacteroidetes, Chlamydiae, Euryarchaeota, Firmicutes, Planctomycetes, Proteobacteria, Tenericutes, TM7, as well as Verrucomicrobia (Fig. 8B). The phylum Firmicutes was present in the highest percentage among samples. The results showed that a bloom of Actinobacteria was associated with normal (GP-1), diabetic control (GP-2), metformin (GP-3), PHE (GP-4 to GP-5), and DMR (GP-6)-treated groups samples. An increase of Firmicutes and a reduction of Bacteroidetes along with Euryarchaeota, Planctomycetes, Proteobacteria, Tenericutes, TM7, and Verrucomicrobia in GP-2 rats were detected compared with GP-1 rats. In contrast, overall treatment groups reduced the prevalence of Firmicutes relative to Bacteroidetes as compared to diabetic rats of GP-2. However, treatment groups GP-3, GP-4, and GP-6 as compared to GP-5 remarkably enriched the amount of Firmicutes. The treatment GP-5 showed a reduction of Firmicutes, Chlamydiae, and Euryarchaeota associated with an elevation of the amount of Bacteroidetes, Planctomycetes, Proteobacteria, Tenericutes, and TM7 in comparison to GP-2. The microbial diversity in GP-3, GP-4, and GP-6 was endorsed by microbiota phyla lateral to GP-5 and restored the microbial phyla abundance compared to GP-1, as seen in Fig. 8B.
In our investigation, we found that there were substantial differences between the groups of rats in terms of 10 different genera of bacterial microbiome. Figure 8C showed the gut microbiota: Aerococcus, Allobaculum, Atopostipes, SMB53, and Staphylococcus genera, which obviously increased, while Clostridium, Lactobacillus, Oscillospira, Prevotella, and Ruminococcus groups decreased in GP-2 rat samples in comparison to GP-1 rat samples. The gut microbiota: Allobaculum, Atopostipes, Lactobacillus, SMB53, and Staphylococcus declined, while Oscillospira, Prevotella, and Ruminococcus increased in the GP-5 rats in comparison to the GP-2 and restored microbiota abundance as in the GP-1. The microbial diversity at the genus level in GP-3, GP-4, and GP-6 was also regulated, as seen in Fig. 8C. The gut microbiota diversity of [Prevotella], Bacteroides, Desulfovibrio, Adlercreutzia, and Bifidobacterium (Supplementary Table S5) was also regulated for treatment groups including PHE to rehabilitate good health.
Furthermore, as shown in Fig. 8D, there were notable differences in the abundance of species-level gut microbiota among the various rat groups. Expression in various conditions is controlled by the species composition and abundance of gut bacteria. The result revealed that the abundance of Atopostipes suicloacalis, Bifidobacterium pseudolongum, Lactobacillus hamsteri, Jeotgalicoccus psychrophilus, Staphylococcus sciuri, and Blautia producta was increased, while the abundance of Clostridium celatum, Prevotella copri, and Ruminococcus bromii was decreased in GP-1 as compared to GP-2. Clostridium celatum, Atopostipes suicloacalis, Jeotgalicoccus psychrophilus, Staphylococcus sciuri, and Ruminococcus albus showed regulated abundance in different treatment groups as compared to group of diabetic rats. The relative abundance of Bifidobacterium pseudolongum was found to decline in PHE treatment groups as compared to diabetic, metformin, and DMR groups. The relative abundance of Lactobacillus hamsteri was elevated in PHE treatment groups as compared to diabetic while it was lower to metformin, and DMR groups. Prevotella copri abundance was also found to increase in PHE treatment groups as well as metformin and DMR groups as compared to the diabetic group. Ruminococcus bromii and Blautia producta showed almost similar declines in abundance in different treatment groups as compared to the diabetic group. At the species level, 10 species shown in Fig. 8D had markedly changed abundance levels with the treatment doses (supplementary Table S5). The taxonomic composition with relative abundance heat map of gut microbial in rat fecal sample of different experimental groups was also shown a significant diversity between groups (supplementary Fig. S2 and S3). The results indicated that the administration of 600 mg/kg body weight of PHE (GP-5) for 28 days was sufficient to alter the gut microbiota composition of the rat. Therefore, the PHE groups were selected for the subsequent study of SCFAs in this study.
Correlation of Gut Microflora with Metabolic Parameters
DM-related indices (body weight and FBG) are shown in Fig. 9i along with their correlations with the microbiota of the gut (markedly modified ten genera of microorganisms by PHE) using canonical correspondence analysis (CCA) on multiple samples. The resultant ordination plots showed that the first axis for both body weight and FBG against gut microbiota explained the majority of the variance (73.82 and 52.42 percent, respectively), while the corresponding second axis accounted for only 26.18 and 29.03 percent, respectively. Based on CCA, we found that Lactobacillus, Oscillospira, Prevotella, Clostridium, Ruminococcus, Desulfovibrio, [Prevotell], Bacteroides, and Aerococcus were positively correlated with body weight (final weight) on the first axis. In contrast, the other bacteria (Adlercreutzia, Bifidobacterium, Allobaculum, Atopostipes, Staphylococcus, and SMB53 group) were negatively correlated with the body weight on the same axis (Fig. 9i (A)). Furthermore, the gut microbiota Lactobacillus, Oscillospira, Prevotella, Ruminococcus, Desulfovibrio, [Prevotell], and Bacteroides were associated with a negative correlation with the level of fasting blood glucose (FBG) on the first axis, while the other bacteria (Adlercreutzia, Bifidobacterium, Allobaculum, Clostridium, Atopostipes, Aerococcus, Staphylococcus, and SMB53 group) were positively correlated with FBG on the same axis (Fig. 9i (B)). Both T2DM-related indicators were found to be more significantly correlated with the gut microbiota species Lactobacillus, Oscillospira, Prevotella, Ruminococcus, Desulfovibrio, [Prevotell], and Bacteroides.
i Correlation analysis between DM-related parameters (body weight and FBG) and gut microbiota at genus level. Canonical correspondence analysis. (A) Body weight Vs gut microbiota and (B) FBG vs gut microbiota. ii SCFAs analysis in the fecal sample of different rat groups. (A) The concentration of acetic acid and (B) the concentration of propionic acid
SCFAs Content of Fecal Sample
Standard graphs from 5 dilutions (50–5000 nM) were plotted for peak area versus concentration. A linear relationship was established with regression coefficients of 0.998 and 0.996 for acetate and propionate, respectively. Details of molecular weight (g/mol), density (g/ml), standard equation, retention time, molecular formula, and R2 values of standards run on HPLC are shown in Table 2, with the standard graph in supplementary Figure S4. Subsequent calculations of SCFAs concentration in the sample groups were done using the standard linear equation derived and summarized in Table 2. Fecal SCFAs can indirectly reflect colon SCFAs, which are linked to intestinal health. The content two dominant main SCFAs in rat feces were affected through PHE as shown in Fig. 9ii (A, B). After 28 days of treatment with different doses of GP-3, GP-4, GP-5, and GP-6, all groups had higher SCFAs concentrations than GP-1 at the end of the experiment. GP-2 had lower SCFAs concentrations among other groups. GP-4 and GP-5 had significantly higher acetate content than GP-1 (p < 0.05 and 0.01, respectively). GP-4 and GP-5 increase feces acetate levels (Fig. 9ii (A)). GP-3 and GP-6 doses also increased acetic acid concentrations in feces (p < 0.05), but GP-2 decreased acetate (Fig. 9ii (A)). Both PHE dosages increased fecal acetic acid concentrations after 28 days of continuous administration in rats from various groups. A consistent finding for propionate concentration for both doses of PHE within GP-4 as well as GP-5 was also seen, with large and highly significant increases in propionic acid concentration in feces compared to other groups (Fig. 9ii (B)). This indicated that both dosages of PHE administered to rats for 28 days could also influence the intestinal environment. These studies demonstrated that PHE’s effect on the gut is dose-dependent; particularly that 600 mg/kg body weight of PHE-treated rats for 28 days increased the primary SCFA content of their feces.
Discussion
With the recent advancement of metabolic and other analytical technologies in the field of scientific research, phytoconstituents introduce a new era of medicinals that may influence our body physiology through various mechanisms utilizing several microneutrients. Thus, the botanical products enriched with bioactive compounds known from earlier investigations can prevent diabetes by changing gut microbiota dysbiosis and inducing microbiome-associated SCFA production. The UHPLC-HRMS analysis of PHE examined numerous biomarkers and bioactive substances. The bioactive substances restored the gut microbiota in rats with DM. Previous investigations (GC–MS and LC–MS) on the bioactive compounds of PHE were consistent and supported our conclusion in the present study. Chebulic acid, chebulagic acid, and ellagic acid identified in PHE may be used as hepatoprotective agents due to the enrichment of metabolites in the gut through gut-microbial interaction [43, 44]. Gallic acid and andrographolide therapy in animals were also supported to restore the metabolic profile and eliminate the risk of developing diabetes [45, 46]. An anti-diabetic effect of berberine, the most concentrated bioactive compound in the current study, has been postulated to occur via regulation of the gut microbiota in an earlier study [47]. A list of bioactive compounds such as berberine [48], oxyberberine [49], kaempferol [50], quercetin [51], apigenin [52], luteolin [53, 54], rutin [55], genistein [56,57,58], hesperidin [59], isorhamnetin [60], ferulic acid [61], and several others have been shown to have anti-diabetic activity by altering gut microbial structures in the studies. These bioactive compounds, their derivatives, and several others were identified (tentively) in good proportion in PHE through UHPLC-HRMS analysis, especially chebulic acid, gallic acid, andrographolide, and berberine. We found that the above-described bioactive compounds improved diabetes-related insulin resistance and the ability to tolerate glucose by strengthening the integrity of the intestinal membrane barrier and restructuring the microbiota of the gut [62], which would be responsible and consistent for the effective anti-diabetic activity of PHE in the current study.
As stated earlier, diabetes has been linked to gut dysbiosis, increased intestinal permeability, and alterations in the overall structure of the gut microbiota [63]. Therefore, altering the composition of the microbiota of the gut is a potential method of reducing hyperglycaemic index in the DM [64]. In the current study, we carried out sequencing of the 16S rRNA using Illumina by the V3 and V4 regions genes and determined the fecal microbiota composition at different taxonomic levels from the studied rat groups. DM rats had increased gut microbial diversity, as measured by Chao1, ACE, and the observed species estimator. These findings corroborated earlier studies showing that a host disease might increase the variety of microorganisms in the digestive tract [65]. The results indicated that the relative abundance of groups like Lactobacillus, Clostridium, Oscillospira, Prevotella, and Ruminococcus was restored in DM rats via PHE treatment, while the relative abundance of groups like Aerococcus, Staphylococcus, Allobaculum, Atopostipes, and SMB53 was reduced. The study of gut microbiota dysbiosis found that Firmicutes and Bacteroidetes were two of the most common bacterial phyla in both rats and humans. A growing amount of literature shows that the Bacteroidetes are enriched for numerous pathways involved in glucose uptake, while the Firmicutes contribute to glucose transport systems [66]. The ratio of Bacteroidetes to Firmicutes was expected to rise after andrographolide treatment, as shown in the study [67], and was consistent in this study. Overall, our findings are consistent with the hypothesis that PHE promotes metabolic health by increasing the abundance of beneficial Bacteroidetes microorganisms in the body. Increasing data revealed that higher levels of Bacteroidetes and Firmicutes were linked to higher BMIs, lower glucose levels, and less insulin resistance, as also stated by Clemente [68]. Diabetic rats had an increased abundance of certain microbiota, including Adlercreutzia, Bifidobacterium, Allobaculum, Clostridium, Atopostipes, Aerococcus, Staphylococcus, and the SMB53 group, all of which linked positively with FBG level and negatively with final body weight in the current study. Concurrently, the other bacteria, including Lactobacillus, Oscillospira, Prevotell, and Bacteroides, were associated positively with the final body weight of the rats and inversely with their FBG levels. As their inverse correlation with FBG and positive correlation with body weight attest, these microbiotal communities have a therapeutic impact [69]. All of these results converged on the conclusion that the PHE treatment would be beneficial for the reshaping of gut microbiota dysbiosis.
In addition, PHE was able to significantly alter the diversity of the microbiota in the intestines of DM rats, specifically influencing certain bacteria associated with SCFAs production as well as anti-inflammation. Bacterial fermentation produces the majority of SCFAs in the major gastrointestinal tract [70] as acetate, propionate, and butyrate. Several positive impacts on host metabolism were mediated by SCFAs. As we know, strong cellular junctions and permeability are both negatively affected by a lack of SCFAs [71]. Total SCFAs, especially butyric, acetic, propionic, valeric acids, and other SCFAs, are reported to increase after treatment with bioactive compounds such as chebulic acid [43], gallic acid [45], andrographolide [67], and berberine [48]. The elevation of SCFAs content with the treatment was also approved in the studies on other bioactive compounds of PHE described and referenced earlier. In addition to the above, the SCFAs appear to promote epithelial regeneration and prevent the inflammation that is partly produced by LPS [72]. SCFAs can also enhance glucose homeostasis as well as insulin sensitivity by altering the function of skeletal muscle, the liver, and adipose tissue [73,74,75]. Diabetic rats had an increased generation of butyric, propionic, and acetic acids due to the enrichment of SCFAs-producing bacteria [63]. Therefore, the hypoglycemic effects of PHE treatment in this study could possibly be attributable to the elevated levels of SCFAs and the growing number of bacteria capable of generating SCFAs. Another study [76] showed that a high concentration of SCFAs was inversely related to the prevalence of overweight, diabetes, cardiovascular disease, and metabolic syndrome. The symbiotic bacteria genus bacterium known as Prevotella copri can generate SCFAs [77]. In another study, it was stated that the commensal obligate anaerobic gram-negative bacterium (Prevotella copri) in the gut has been shown to reduce blood glucose levels in rodents and healthy humans [78], making it a promising candidate to prevent or treat obesity and DM. At the mechanistic level, Prevotella copri enhanced the bile acid (BA) metabolism signal [79]. Despite the fact that Prevotella copri was shown to be the primary species driving the relationship between branch-chain amino acid (BCAA) production and insulin resistance, it was also found that BCAAs themselves can cause insulin resistance and exacerbate glucose intolerance in DM [62]. As stated, the presence of Prevotella copri is an indicator of food, lifestyle, and health status [80]. Thus, collectively, in a mechanistic way, it increased hepatic glycogen through modulating BA metabolism [79], which was consistent with our result. From our data, it was identified that Blautia, a gut microbe, belonged to the Lachnospiraceae family within the Firmicutes phylum and that it exhibited a counterintuitive variation trend among distinct groups, many of which were known to be powerful SCFAs producers [81]. Blautia producta, an organism involved in the formation of acetate or propionate, was discovered in significantly higher amounts in this study, with similar findings in HFD-fed rats compared to control animals [82]. The findings are also consistent with those of Zhang [83], who found that rats given an HFD had considerably higher amounts of acetic acid as well as propionic acid. Our findings suggested that this family could have a major beneficial effect on PHE’s ability to treat DM. Blautia may convert undigested carbs and proteins into acetate, which could subsequently be used to provide energy for the organism [84].
Bacteroidetes, Proteobacteria, Firmicutes, and Verrucomicrobia have all been found to increase in abundance in studies showing that berberine can alter the composition of gut microbiota in normal animals [85]. There is a wide variety of metabolites that can be produced and released by the gut microbiota, including but not limited to SCFAs, BCAAs, BAs, and TMA [86]. Berberine-deprived species that mainly create single sugars, or SCFAs from metabolizing polysaccharides or oligosaccharides, such as Ruminococcus bromii and Bifidobacterium spp. [87], were often observed to cross-feed with other saccharide degraders [88]. Metformin treatment generated the same species enrichment seen with berberine [89], including two Bacteroides spp. and many taxa of γ-Proteobacteria. Herbal extracts have been shown to be more effective at treating diabetes than the commonly used drugs acarbose and metformin, according to the literature [89, 90]. Researchers also found that BA metabolism was controlled by Ruminococcus strains [91]. This suggests that Ruminococcus bromii in the gut microbiome could be the objective of berberine in order to decrease the microbial synthesis of secondary BA, which is linked to the successful glucose management produced with berberine [87]. Multiple Bifidobacterium spp. were inhibited by berberine in both humans and animals [92]. Therefore, Bifidobacterium strains may have an especially positive effect on elderly people with DM who are taking Berberine. Bifidobacterium pseudolongum was also shown to control the number of bacteria in the gut, implying that its presence was essential for the proliferation of the genus [93]. Overgrowth organisms like Staphylococci coliforms were prevented by berberine, but native lactobacilli and bifidobacteria were unaffected [47]. The quantity of gut microbes such as Actinobacteria, Lactobacillus, and Firmicutes was decreased in an animal study with DM [48], and berberine was able to effectively manage glucose metabolism and re-establish glucose homeostasis. Previous research revealed that a novel gut microbiota-mediated metabolite of berberine called oxyberberine, which is also an important bioactive constituent of PHE, was able to keep the intestinal microbiota in equilibrium by altering the structure of the gut microbiota [49]. Andrographolide enhanced Prevotella and Adlercreutzia at the genus level [67]. Kaempferol, quercetin, luteolin, and rutin administration in the experimental animals with a specific protocol individually was found to be effective in reversing the gut dysbiosis associated with metabolic diseases, as determined in the different studies by sequencing the fecal microbiota [51, 53, 54]. Thus, PHE containing bioactive compounds like andrographolide, kaempferol, quercetin, luteolin, and rutin may change the composition of the gut microbiota by raising the number of good bacteria. Together with each other, our results showed that genistein [56,57,58], hesperidin [59], isorhamnetin [60], and ferulic acid [61] are the important contents of the PHE in the present study, which may have a direct interaction on the gut microbiota and can ameliorate diabetic symptoms in DM rats by up-regulating SCFA concentrations. Our results were consistent with those of previous studies.
Our analysis of SCFAs in fecal samples from all rats groups revealed that PHE treatment compared with metformin and DMR could significantly alter the composition and function of the beneficial gut microbiota. This finding of the study would be due to the presence of numerous biomarkers and bioactive compounds in PHE [69]. In the current study, more than ten bacterial phyla controlled by PHE-contents exhibited a significant link with being healthy (body weight) or with beneficial glycemic effects (FBG levels). Thus, the collective presence of numerous beneficial bacteria like Lactobacillus, Oscillospira, Prevotella, Desulfovibrio, [Prevotella], and Bacteroides in the PHE has the potential to produce SCFAs such as acetate and propionate in good proportion, which could activate gut gluconeogenesis via complementary mechanisms [69] and reduce blood glucose levels as well as increase body mass index (good health). These findings were supported according to a correlation analysis between the abundance of gut bacteria with PHE treatment and a couple of DM-related indices. We generalized our prior study’s [3] findings that ingesting PHE in DM rats had a hypoglycemic effect by targeting particular gut microbiota, which is the study’s supportive approach for the treatment of DM at multiple pharmacological sites. Our results demonstrated that PHE, an anti-diabetic medicine, was able to reduce and mitigate inflammation as well as hyperglycemia, insulin resistance, and lipid metabolism abnormalities brought on by DM in the present experimental rat groups through modulation of the gut microbiome.
Conclusion
In conclusion, the presence of several bioactive substances such as chebulic acid, ellagic acid, gallic acid, andrographolide, berberine, kaempferol, quercetin, luteolin, rutin, apigenin, and others in PHE suggested that it could alleviate the hyperglycemia in DM rats. Furthermore, PHE was able to change the gut microbiome and enhance SCFAs, both of which were significant in the pathophysiology of DM. Finally, we hypothesized that SCFAs-producing as well as anti-inflammatory bacterial genera like Lactobacillus, Oscillospira, Prevotella, Desulfovibrio, [Prevotella], and Bacteroides were involved in PHE’s beneficial effects on DM rats. Specifically, the enrichment of bacterial strains like Prevotella copri and Lactobacillus hamster and the decline in relative abundance of Bifidobacterium pseudolongum, Ruminococcus bromii, and Blautia producta through PHE treatment produce health benefits because the generated metabolites increase epithelial barrier function, improve gut permeability, inhibit inflammation, reduce insulin resistance, and reduce the severity of DM by modulating the multiple metabolic pathways with multiple targets involved. We found unanticipated impacts of PHE’s systemically hypoglycemic effects using dual therapy. PHE phytochemicals, regenerated gut microbiota, and SCFAs treated DM in this study. In the future, using a mechanistic approach to study the gut microbiota’s anti-diabetic properties in vivo and apply them to clinical practice may help to treat DM.
Data Availability
Not applicable.
References
Bunsroem, K., Prinyawiwatkul, W., & Thaiudom, S. (2022). The influence of whey protein heating parameters on their susceptibility to digestive enzymes and the antidiabetic activity of hydrolysates. Foods, 11(6), 829. https://doi.org/10.3390/foods11060829
Inayati, A., Lee, B. O., Wang, R. H., Chen, S. Y., Hsu, H. C., Lu, C. H., & Lee, Y. J. (2022). Determinants of fear of falling in older adults with diabetes. Geriatric Nursing, 46, 7–12. https://doi.org/10.1016/j.gerinurse.2022.04.017
Singh, A. K., Kumar, P., Rajput, V. D., Mishra, S. K., Tiwari, K. N., Singh, A. K., Minkina, T., & Pandey, A. K. (2023). Phytochemicals, antioxidant, anti-inflammatory studies, and identification of bioactive compounds using GC-MS of ethanolic novel polyherbal extract. Applied Biochemistry and Biotechnology. https://doi.org/10.1007/s12010-023-04363-7
Sun, H., Saeedi, P., Karuranga, S., Pinkepank, M., Ogurtsova, K., Duncan, B. B., Stein, C., Basit, A., Chan, J. C. N., Mbanya, J. C., Pavkov, M. E., Ramachandaran, A., Wild, S. H., James, S., Herman, W. H., Zhang, P., Bommer, C., Kuo, S., Boyko, E. J., & Magliano, D. J. (2022). IDF diabetes atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Research and Clinical Practice, 183, 109119. https://doi.org/10.1016/j.diabres.2021.109119
Lorenzati, B., Zucco, C., Miglietta, S., Lamberti, F., & Bruno, G. (2010). Oral hypoglycemic drugs: Pathophysiological basis of their mechanism of actionoral hypoglycemic drugs: Pathophysiological basis of their mechanism of action. Pharmaceuticals, 3, 3005–3020. https://doi.org/10.3390/ph3093005
Calcutt, N. A., Cooper, M. E., Kern, T. S., & Schmidt, A. M. (2009). Therapies for hyperglycaemia-induced diabetic complications: From animal models to clinical trials. Nature Reviews. Drug Discovery, 8(5), 417–429. https://doi.org/10.1038/nrd2476
Jaiswal, Y. S., & Williams, L. L. (2017). A glimpse of Ayurveda-The forgotten history and principles of Indian traditional medicine. Journal of Traditional and Complementary Medicine, 7(1), 50–53.
Rathor, L., Pant, A., Awasthi, H., Mani, D., & Pandey, R. (2017). An antidiabetic polyherbal phytomedicine confers stress resistance and extends lifespan in Caenorhabditis elegans. Biogerontology, 18, 131–147.
Srivastava, S., Lal, V. K., & Pant, K. K. (2012). Polyherbal formulations based on Indian medicinal plants as antidiabetic phytotherapeutics. Phytopharmacology, 2(1), 1–15.
Girish, S., Kuber, S., & Nataraj, H. R. (2015). Review on Kataka (Strychnous potatorum Linn). International Journal of Research in Ayurveda and Pharmacy, 6(1), 86–89.
Duraiswamy, A., Shanmugasundaram, D., Sasikumar, C. S., Cherian, S. M., & Cherian, K. M. (2015). Development of an antidiabetic formulation (ADJ6) and its inhibitory activity against α-amylase and α-glucosidase. Journal of Traditional and Complementry Medicine, 6(3), 204–208. https://doi.org/10.1016/j.jtcme.2014.12.006
Kumudhaveni, B., & Radha, R. (2017). Anti-diabetic potential of a traditional Polyherbal formulation-A review. Research Journal of Pharmacy and Technology, 10(6), 1865–1869.
Wediasari, F., Nugroho, G. A., Fadhilah, Z., Elya, B., Setiawan, H., & Mozef, T. (2020). Hypoglycemic effect of a combined Andrographis paniculata and Caesalpinia sappan extract in streptozocin-induced diabetic rats. Advances in Pharmacological and Pharmaceutical Sciences, 2020, 8856129. https://doi.org/10.1155/2020/8856129
Piccolella, S., Crescente, G., Candela, L., & Pacifico, S. (2019). Nutraceutical polyphenols: New analytical challenges and opportunities. Journal of Pharmaceutical and Biomedical Analysis, 175, 112774. https://doi.org/10.1016/j.jpba.2019.07.022
Turi, C. E., Finley, J., Shipley, P. R., Murch, S. J., & Brown, P. N. (2015). Metabolomics for phytochemical discovery: Development of statistical approaches using a cranberry model system. Journal of Natural Products, 78(4), 953–966. https://doi.org/10.1021/np500667z
Brahmi-Chendouh, N., Piccolella, S., Crescente, G., Pacifico, F., Boulekbache, L., Hamri-Zeghichi, S., Akkal, S., Madani, K., & Pacifico, S. (2019). A nutraceutical extract from Inula viscosa leaves: UHPLC-HR-MS/MS based polyphenol profile, and antioxidant and cytotoxic activities. Journal of Food and Drug Analysis, 27(3), 692–702. https://doi.org/10.1016/j.jfda.2018.11.006
Makki, K., Deehan, E. C., Walter, J., & Bäckhed, F. (2018). The impact of dietary fiber on gut microbiota in host health and disease. Cell Host and Microbe, 23(6), 705–715. https://doi.org/10.1016/j.chom.2018.05.012
Sharma, S., & Tripathi, P. (2019). Gut microbiome and type 2 diabetes: Where we are and where to go? The Journal of Nutritional Biochemistry, 63, 101–108. https://doi.org/10.1016/j.jnutbio.2018.10.003
Xu, W. T., Nie, Y. Z., Yang, Z., & Lu, N. H. (2016). The crosstalk between gut microbiota and obesity and related metabolic disorders. Future Microbiology, 11, 825–836. https://doi.org/10.2217/fmb-2015-0024
Singer-Englar, T., Barlow, G., & Mathur, R. (2019). Obesity, diabetes, and the gut microbiome: An updated review. Expert Review of Gastroenterology and Hepatology, 13(1), 3–15. https://doi.org/10.1080/17474124.2019.1543023
Puddu, A., Sanguineti, R., Montecucco, F., & Viviani, G. L. (2014). Evidence for the gut microbiota short-chain fatty acids as key pathophysiological molecules improving diabetes. Mediators of Inflammation, 2014, 162021. https://doi.org/10.1155/2014/162021
Ley, R. E., Bäckhed, F., Turnbaugh, P., Lozupone, C. A., Knight, R. D., & Gordon, J. I. (2005). Obesity alters gut microbial ecology. Proceedings of the National Academy of Sciences of the United States of America, 102(31), 11070–11075. https://doi.org/10.1073/pnas.0504978102
Kreznar, J. H., Keller, M. P., Traeger, L. L., Rabaglia, M. E., Schueler, K. L., Stapleton, D. S., Zhao, W., Vivas, E. I., Yandell, B. S., Broman, A. T., Hagenbuch, B., Attie, A. D., & Rey, F. E. (2017). Host genotype and gut microbiome modulate insulin secretion and diet-induced metabolic phenotypes. Cell reports, 18(7), 1739–1750. https://doi.org/10.1016/j.celrep.2017.01.062
Drissi, F., Buffet, S., Raoult, D., & Merhej, V. (2015). Common occurrence of antibacterial agents in human intestinal microbiota. Frontiers in Microbiology, 6, 441. https://doi.org/10.3389/fmicb.2015.00441
Federici, M. (2019). Gut microbiome and microbial metabolites: A new system affecting metabolic disorders. Journal of Endocrinological Investigation, 42(9), 1011–1018. https://doi.org/10.1007/s40618-019-01022-9
Salgaço, M. K., Oliveira, L. G. S., Costa, G. N., Bianchi, F., & Sivieri, K. (2019). Relationship between gut microbiota, probiotics, and type 2 diabetes mellitus. Applied Microbiology and Biotechnology, 103(23–24), 9229–9238. https://doi.org/10.1007/s00253-019-10156-y
Fiorucci, S., & Distrutti, E. (2015). Bile acid-activated receptors, intestinal microbiota, and the treatment of metabolic disorders. Trends in Molecular Medicine, 21(11), 702–714. https://doi.org/10.1016/j.molmed.2015.09.001
Cani, P. D., Plovier, H., Van Hul, M., Geurts, L., Delzenne, N. M., Druart, C., & Everard, A. (2016). Endocannabinoids—At the crossroads between the gut microbiota and host metabolism. Nature Reviews. Endocrinology, 12(3), 133–143. https://doi.org/10.1038/nrendo.2015.211
Sohail, M. U., Althani, A., Anwar, H., Rizzi, R., & Marei, H. E. (2017). Role of the gastrointestinal tract microbiome in the pathophysiology of diabetes mellitus. Journal of Diabetes Research, 2017, 9631435. https://doi.org/10.1155/2017/9631435
Larsen, N., Vogensen, F. K., van den Berg, F. W., Nielsen, D. S., Andreasen, A. S., Pedersen, B. K., Al-Soud, W. A., Sørensen, S. J., Hansen, L. H., & Jakobsen, M. (2010). Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE, 5(2), e9085. https://doi.org/10.1371/journal.pone.0009085
Wei, X., Tao, J., Xiao, S., Jiang, S., Shang, E., Zhu, Z., Qian, D., & Duan, J. (2018). Xiexin Tang improves the symptom of type 2 diabetic rats by modulation of the gut microbiota. Scientific Reports, 8(1), 3685. https://doi.org/10.1038/s41598-018-22094-2
Forslund, K., Hildebrand, F., Nielsen, T., Falony, G., Le Chatelier, E., Sunagawa, S., Prifti, E., Vieira-Silva, S., Gudmundsdottir, V., Pedersen, H. K., Arumugam, M., Kristiansen, K., Voigt, A. Y., Vestergaard, H., Hercog, R., Costea, P. I., Kultima, J. R., Li, J., Jørgensen, T., Levenez, F., … Pedersen, O. (2015). Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature, 528(7581), 262–266. https://doi.org/10.1038/nature15766
Harbison, J. E., Roth-Schulze, A. J., Giles, L. C., Tran, C. D., Ngui, K. M., Penno, M. A., Thomson, R. L., Wentworth, J. M., Colman, P. G., Craig, M. E., Morahan, G., Papenfuss, A. T., Barry, S. C., Harrison, L. C., & Couper, J. J. (2019). Gut microbiome dysbiosis and increased intestinal permeability in children with islet autoimmunity and type 1 diabetes: A prospective cohort study. Pediatric diabetes, 20(5), 574–583. https://doi.org/10.1111/pedi.12865
van Passel, M. W., Kant, R., Zoetendal, E. G., Plugge, C. M., Derrien, M., Malfatti, S. A., Chain, P. S., Woyke, T., Palva, A., de Vos, W. M., & Smidt, H. (2011). The genome of Akkermansia muciniphila, a dedicated intestinal mucin degrader, and its use in exploring intestinal metagenomes. PLoS ONE, 6(3), e16876. https://doi.org/10.1371/journal.pone.0016876
Yan, Z., Wu, H., Zhou, H., Chen, S., He, Y., Zhang, W., Chen, T., Yao, H., & Su, W. (2020). Integrated metabolomics and gut microbiome to the effects and mechanisms of naoxintong capsule on type 2 diabetes in rats. Scientific reports, 10(1), 10829. https://doi.org/10.1038/s41598-020-67362-2
Salamone, D., Rivellese, A. A., & Vetrani, C. (2021). The relationship between gut microbiota, short-chain fatty acids and type 2 diabetes mellitus: The possible role of dietary fibre. Acta Diabetologica, 58(9), 1131–1138.
Abu-Reidah, I. M., Critch, A. L., Manful, C. F., Rajakaruna, A., Vidal, N. P., Pham, T. H., Cheema, M., & Thomas, R. (2021). Effects of pH and temperature on water under pressurized conditions in the extraction of nutraceuticals from Chaga (Inonotus obliquus) Mushroom. Antioxidants, 10(8), 1322. https://doi.org/10.3390/antiox10081322
Hilary, S., Mohamed, O., Platat, C., Qureshi, M. A., Kizhakkayil, J., Al-Meqbaali, F., & Howarth, F. C. (2023). Supplemental ferulic acid does not affect metabolic markers and improves some oxidative damage parameters in diabetic rats. Heliyon, 9(6), e17313. https://doi.org/10.1016/j.heliyon.2023.e17313
Ojo, O. A., Amanze, J. C., Oni, A. I., Grant, S., Iyobhebhe, M., Elebiyo, T. C., Rotimi, D., Asogwa, N. T., Oyinloye, B. E., Ajiboye, B. O., & Ojo, A. B. (2022). Antidiabetic activity of avocado seeds (Persea americana Mill.) in diabetic rats via activation of PI3K/AKT signaling pathway. Scientific Reports, 12(1), 2919. https://doi.org/10.1038/s41598-022-07015-8
Klindworth, A., Pruesse, E., Schweer, T., Peplies, J., Quast, C., Horn, M., & Glöckner, F. O. (2013). Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Research, 41(1), e1. https://doi.org/10.1093/nar/gks808
DeSantis, T. Z., Hugenholtz, P., Larsen, N., Rojas, M., Brodie, E. L., Keller, K., Huber, T., Dalevi, D., Hu, P., & Andersen, G. L. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Applied and Environmental Microbiology, 72(7), 5069–5072. https://doi.org/10.1128/AEM.03006-05
Sasaki, D., Sasaki, K., Ikuta, N., Yasuda, T., Fukuda, I., Kondo, A., & Osawa, R. (2018). Low amounts of dietary fibre increase in vitro production of short-chain fatty acids without changing human colonic microbiota structure. Scientific Reports, 8(1), 435. https://doi.org/10.1038/s41598-017-18877-8
Seçkin, H. (2022). Microflora in rats metagenomic analysis of intestinal microflora in rats treated with ellagic acid and sinapic acid using 16 S rDNA gene region. Indian Journal of Pharmaceutical Education and Research, 56(3).
Kim, D. H., Sim, Y., Hwang, J. H., Kwun, I. S., Lim, J. H., Kim, J., Kim, J. I., Baek, M. C., Akbar, M., Seo, W., Kim, D. K., Song, B. J., & Cho, Y. E. (2021). Ellagic acid prevents binge alcohol-induced leaky gut and liver injury through inhibiting gut dysbiosis and oxidative stress. Antioxidants (Basel, Switzerland), 10(9), 1386. https://doi.org/10.3390/antiox10091386
Chao, J., Huo, T. I., Cheng, H. Y., Tsai, J. C., Liao, J. W., Lee, M. S., Qin, X. M., Hsieh, M. T., Pao, L. H., & Peng, W. H. (2014). Gallic acid ameliorated impaired glucose and lipid homeostasis in high fat diet-induced NAFLD mice. PLoS ONE, 9(2), e96969. https://doi.org/10.1371/journal.pone.0096969
Brahmachari, G. (Ed.). (2017). Discovery and development of neuroprotective agents from natural products. Elsevier.
Han, J., Lin, H., & Huang, W. (2011). Modulating gut microbiota as an anti-diabetic mechanism of berberine. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research, 17(7), RA164–RA167.
Cheng, H., Liu, J., Tan, Y., Feng, W., & Peng, C. (2022). Interactions between gut microbiota and berberine, a necessary procedure to understand the mechanisms of berberine. Journal of pharmaceutical analysis, 12(4), 541–555. https://doi.org/10.1016/j.jpha.2021.10.003
Li, C., Ai, G., Wang, Y., Lu, Q., Luo, C., Tan, L., Lin, G., Liu, Y., Li, Y., Zeng, H., Chen, J., Lin, Z., Xian, Y., Huang, X., Xie, J., & Su, Z. (2020). Oxyberberine, a novel gut microbiota-mediated metabolite of berberine, possesses superior anti-colitis effect: Impact on intestinal epithelial barrier, gut microbiota profile and TLR4-MyD88-NF-κB pathway. Pharmacological Research, 152, 104603. https://doi.org/10.1016/j.phrs.2019.104603
Bian, Y., Lei, J., Zhong, J., Wang, B., Wan, Y., Li, J., Liao, C., He, Y., Liu, Z., Ito, K., & Zhang, B. (2022). Kaempferol reduces obesity, prevents intestinal inflammation, and modulates gut microbiota in high-fat diet mice. The Journal of Nutritional Biochemistry, 99, 108840. https://doi.org/10.1016/j.jnutbio.2021.108840
Etxeberria, U., Arias, N., Boqué, N., Macarulla, M. T., Portillo, M. P., Martínez, J. A., & Milagro, F. I. (2015). Reshaping faecal gut microbiota composition by the intake of trans-resveratrol and quercetin in high-fat sucrose diet-fed rats. The Journal of Nutritional Biochemistry, 26(6), 651–660. https://doi.org/10.1016/j.jnutbio.2015.01.002
Qiao, Y., Zhang, Z., Zhai, Y., Yan, X., Zhou, W., Liu, H., Guan, L., & Peng, L. (2022). Apigenin alleviates obesity-associated metabolic syndrome by regulating the composition of the gut microbiome. Frontiers in Microbiology, 12, 805827. https://doi.org/10.3389/fmicb.2021.805827
Ge, X., He, X., Liu, J., Zeng, F., Chen, L., Xu, W., Shao, R., Huang, Y., Farag, M. A., Capanoglu, E., El-Seedi, H. R., Zhao, C., & Liu, B. (2023). Amelioration of type 2 diabetes by the novel 6, 8-guanidyl luteolin quinone-chromium coordination via biochemical mechanisms and gut microbiota interaction. Journal of Advanced Research, 46, 173–188. https://doi.org/10.1016/j.jare.2022.06.003
Ge, X., Wang, C., Chen, H., Liu, T., Chen, L., Huang, Y., Zeng, F., & Liu, B. (2020). Luteolin cooperated with metformin hydrochloride alleviates lipid metabolism disorders and optimizes intestinal flora compositions of high-fat diet mice. Food and Function, 11(11), 10033–10046. https://doi.org/10.1039/d0fo01840f
Cai, C., Cheng, W., Shi, T., Liao, Y., Zhou, M., & Liao, Z. (2023). Rutin alleviates colon lesions and regulates gut microbiota in diabetic mice. Scientific Reports, 13(1), 4897. https://doi.org/10.1038/s41598-023-31647-z
Huang, G., Xu, J., Lefever, D. E., Glenn, T. C., Nagy, T., & Guo, T. L. (2017). Genistein prevention of hyperglycemia and improvement of glucose tolerance in adult non-obese diabetic mice are associated with alterations of gut microbiome and immune homeostasis. Toxicology and Applied Pharmacology, 332, 138–148. https://doi.org/10.1016/j.taap.2017.04.009
Li, S., Zhou, L., Zhang, Q., Yu, M., & Xiao, X. (2022). Genistein improves glucose metabolism and promotes adipose tissue browning through modulating gut microbiota in mice. Food and Function, 13(22), 11715–11732. https://doi.org/10.1039/d2fo01973f
Yang, R., Jia, Q., Mehmood, S., Ma, S., & Liu, X. (2021). Genistein ameliorates inflammation and insulin resistance through mediation of gut microbiota composition in type 2 diabetic mice. European Journal of Nutrition, 60(4), 2155–2168. https://doi.org/10.1007/s00394-020-02403-0
Estruel-Amades, S., Massot-Cladera, M., Pérez-Cano, F. J., Franch, À., Castell, M., & Camps-Bossacoma, M. (2019). Hesperidin effects on gut microbiota and gut-associated lymphoid tissue in healthy rats. Nutrients, 11(2), 324. https://doi.org/10.3390/nu11020324
Li, J., Yuan, H., Zhao, Z., Li, L., Li, X., Zhu, L., Wang, X., Sun, P., & Xiao, Y. (2022). The mitigative effect of isorhamnetin against type 2 diabetes via gut microbiota regulation in mice. Frontiers in Nutrition, 9, 1070908. https://doi.org/10.3389/fnut.2022.1070908
Tian, B., Geng, Y., Wang, P., Cai, M., Neng, J., Hu, J., Xia, D., Cao, W., Yang, K., & Sun, P. (2022). Ferulic acid improves intestinal barrier function through altering gut microbiota composition in high-fat diet-induced mice. European Journal of Nutrition, 61(7), 3767–3783. https://doi.org/10.1007/s00394-022-02927-7
O’Keefe, S. J. (2008). Nutrition and colonic health: The critical role of the microbiota. Current Opinion in Gastroenterology, 24(1), 51–58. https://doi.org/10.1097/MOG.0b013e3282f323f3
Nie, Q., Hu, J., Gao, H., Li, M., Sun, Y., Chen, H., Zuo, S., Fang, Q., Huang, X., Yin, J., & Nie, S. (2021). Bioactive dietary fibers selectively promote gut microbiota to exert antidiabetic effects. Journal of Agricultural and Food Chemistry, 69(25), 7000–7015. https://doi.org/10.1021/acs.jafc.1c01465
Su, H., Xie, L., Xu, Y., Ke, H., Bao, T., Li, Y., & Chen, W. (2020). Pelargonidin-3-O-glucoside derived from wild raspberry exerts antihyperglycemic effect by inducing autophagy and modulating gut microbiota. Journal of Agricultural and Food Chemistry, 68(46), 13025–13037. https://doi.org/10.1021/acs.jafc.9b03338
Ling, Z., Liu, X., Jia, X., Cheng, Y., Luo, Y., Yuan, L., Wang, Y., Zhao, C., Guo, S., Li, L., Xu, X., & Xiang, C. (2014). Impacts of infection with different toxigenic Clostridium difficile strains on faecal microbiota in children. Scientific Reports, 4, 7485. https://doi.org/10.1038/srep07485
Turnbaugh, P. J., Hamady, M., Yatsunenko, T., Cantarel, B. L., Duncan, A., Ley, R. E., Sogin, M. L., Jones, W. J., Roe, B. A., Affourtit, J. P., Egholm, M., Henrissat, B., Heath, A. C., Knight, R., & Gordon, J. I. (2009). A core gut microbiome in obese and lean twins. Nature, 457(7228), 480–484. https://doi.org/10.1038/nature07540
Su, H., Mo, J., Ni, J., Ke, H., Bao, T., Xie, J., Xu, Y., Xie, L., & Chen, W. (2020). Andrographolide exerts antihyperglycemic effect through strengthening intestinal barrier function and increasing microbial composition of Akkermansia muciniphila. Oxidative Medicine and Cellular Longevity, 2020, 6538930. https://doi.org/10.1155/2020/6538930
Clemente, J. C., Ursell, L. K., Parfrey, L. W., & Knight, R. (2012). The impact of the gut microbiota on human health: An integrative view. Cell, 148(6), 1258–1270. https://doi.org/10.1016/j.cell.2012.01.035
Holmes, D. (2016). Gut microbiota: Antidiabetic drug treatment confounds gut dysbiosis associated with type 2 diabetes mellitus. Nature Reviews. Endocrinology, 12(2), 61. https://doi.org/10.1038/nrendo.2015.222
Chen, G., Ran, X., Li, B., Li, Y., He, D., Huang, B., Fu, S., Liu, J., & Wang, W. (2018). Sodium butyrate inhibits inflammation and maintains epithelium barrier integrity in a TNBS-induced inflammatory bowel disease mice model. eBioMedicine, 30, 317–325. https://doi.org/10.1016/j.ebiom.2018.03.030
Shi, X., Wei, X., Yin, X., Wang, Y., Zhang, M., Zhao, C., Zhao, H., McClain, C. J., Feng, W., & Zhang, X. (2015). Hepatic and fecal metabolomic analysis of the effects of Lactobacillus rhamnosus GG on alcoholic fatty liver disease in mice. Journal of Proteome Research, 14(2), 1174–1182. https://doi.org/10.1021/pr501121c
Sturm, A., & Dignass, A. U. (2008). Epithelial restitution and wound healing in inflammatory bowel disease. World Journal of Gastroenterology, 14(3), 348–353. https://doi.org/10.3748/wjg.14.348
Canfora, E. E., Jocken, J. W., & Blaak, E. E. (2015). Short-chain fatty acids in control of body weight and insulin sensitivity. Nature Reviews. Endocrinology, 11(10), 577–591. https://doi.org/10.1038/nrendo.2015.128
Sanna, S., van Zuydam, N. R., Mahajan, A., Kurilshikov, A., Vich Vila, A., Võsa, U., Mujagic, Z., Masclee, A. A. M., Jonkers, D. M. A. E., Oosting, M., Joosten, L. A. B., Netea, M. G., Franke, L., Zhernakova, A., Fu, J., Wijmenga, C., & McCarthy, M. I. (2019). Causal relationships among the gut microbiome, short-chain fatty acids and metabolic diseases. Nature Genetics, 51(4), 600–605. https://doi.org/10.1038/s41588-019-0350-x
Tremaroli, V., & Bäckhed, F. (2012). Functional interactions between the gut microbiota and host metabolism. Nature, 489(7415), 242–249. https://doi.org/10.1038/nature11552
Jung, M. J., Lee, J., Shin, N. R., Kim, M. S., Hyun, D. W., Yun, J. H., Kim, P. S., Whon, T. W., & Bae, J. W. (2016). Chronic repression of mTOR complex 2 induces changes in the gut microbiota of diet-induced obese mice. Scientific Reports, 6, 30887. https://doi.org/10.1038/srep30887
Vuillermin, P. J., O’Hely, M., Collier, F., Allen, K. J., Tang, M. L. K., Harrison, L. C., Carlin, J. B., Saffery, R., Ranganathan, S., Sly, P. D., Gray, L., Molloy, J., Pezic, A., Conlon, M., Topping, D., Nelson, K., Mackay, C. R., Macia, L., Koplin, J., Dawson, S. L., … BIS Investigator Group (2020). Maternal carriage of Prevotella during pregnancy associates with protection against food allergy in the offspring. Nature communications, 11(1), 1452https://doi.org/10.1038/s41467-020-14552-1
Verbrugghe, P., Brynjólfsson, J., Jing, X., Björck, I., Hållenius, F., & Nilsson, A. (2021). Evaluation of hypoglycemic effect, safety and immunomodulation of Prevotella copri in mice. Scientific Reports, 11(1), 21279. https://doi.org/10.1038/s41598-021-96161-6
Péan, N., Le Lay, A., Brial, F., Wasserscheid, J., Rouch, C., Vincent, M., Myridakis, A., Hedjazi, L., Dumas, M. E., Grundberg, E., Lathrop, M., Magnan, C., Dewar, K., & Gauguier, D. (2020). Dominant gut Prevotella copri in gastrectomised non-obese diabetic Goto-Kakizaki rats improves glucose homeostasis through enhanced FXR signalling. Diabetologia, 63(6), 1223–1235. https://doi.org/10.1007/s00125-020-05122-7
Gorvitovskaia, A., Holmes, S. P., & Huse, S. M. (2016). Interpreting Prevotella and Bacteroides as biomarkers of diet and lifestyle. Microbiome, 4, 15. https://doi.org/10.1186/s40168-016-0160-7
Daniel, H., Gholami, A. M., Berry, D., Desmarchelier, C., Hahne, H., Loh, G., Mondot, S., Lepage, P., Rothballer, M., Walker, A., Böhm, C., Wenning, M., Wagner, M., Blaut, M., Schmitt-Kopplin, P., Kuster, B., Haller, D., & Clavel, T. (2014). High-fat diet alters gut microbiota physiology in mice. The ISME Journal, 8(2), 295–308. https://doi.org/10.1038/ismej.2013.155
Liu, S., Qin, P., & Wang, J. (2019). High-fat diet alters the intestinal microbiota in streptozotocin-induced type 2 diabetic mice. Microorganisms, 7(6), 176. https://doi.org/10.3390/microorganisms7060176
Zhang, C., Li, S., Yang, L., Huang, P., Li, W., Wang, S., Zhao, G., Zhang, M., Pang, X., Yan, Z., Liu, Y., & Zhao, L. (2013). Structural modulation of gut microbiota in life-long calorie-restricted mice. Nature Communications, 4, 2163. https://doi.org/10.1038/ncomms3163
Park, S. K., Kim, M. S., & Bae, J. W. (2013). Blautia faecis sp. nov., isolated from human faeces. International Journal of Systematic and Evolutionary Microbiology, 63(pt 2), 599–603. https://doi.org/10.1099/ijs.0.036541-0
Ji, Y., Yin, Y., Li, Z., & Zhang, W. (2019). Gut microbiota-derived components and metabolites in the progression of non-alcoholic fatty liver disease (NAFLD). Nutrients, 11(8), 1712. https://doi.org/10.3390/nu11081712
Tian, Y., Cai, J., Gui, W., Nichols, R. G., Koo, I., Zhang, J., Anitha, M., & Patterson, A. D. (2019). Berberine directly affects the gut microbiota to promote intestinal Farnesoid X receptor activation. Drug Metabolism and Disposition: The Biological Fate of Chemicals, 47(2), 86–93. https://doi.org/10.1124/dmd.118.083691
Zhang, Y., Gu, Y., Ren, H., Wang, S., Zhong, H., Zhao, X., Ma, J., Gu, X., Xue, Y., Huang, S., Yang, J., Chen, L., Chen, G., Qu, S., Liang, J., Qin, L., Huang, Q., Peng, Y., Li, Q., Wang, X., … Wang, W. (2020). Gut microbiome-related effects of berberine and probiotics on type 2 diabetes (the PREMOTE study). Nature communications, 11(1), 5015. https://doi.org/10.1038/s41467-020-18414-8
Rios-Covian, D., Gueimonde, M., Duncan, S. H., Flint, H. J., & de Reyes-Gavilanlos, C. G. (2015). Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiology Letters, 362(21), fnv176. https://doi.org/10.1093/femsle/fnv176
Sun, L., Xie, C., Wang, G., Wu, Y., Wu, Q., Wang, X., Liu, J., Deng, Y., Xia, J., Chen, B., Zhang, S., Yun, C., Lian, G., Zhang, X., Zhang, H., Bisson, W. H., Shi, J., Gao, X., Ge, P., Liu, C., … Jiang, C. (2018). Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nature medicine, 24(12), 1919–1929. https://doi.org/10.1038/s41591-018-0222-4
Gu, Y., Wang, X., Li, J., Zhang, Y., Zhong, H., Liu, R., Zhang, D., Feng, Q., Xie, X., Hong, J., Ren, H., Liu, W., Ma, J., Su, Q., Zhang, H., Yang, J., Wang, X., Zhao, X., Gu, W., Bi, Y., … Wang, W. (2017). Analyses of gut microbiota and plasma bile acids enable stratification of patients for antidiabetic treatment. Nature communications, 8(1), 1785. https://doi.org/10.1038/s41467-017-01682-2
Staley, C., Weingarden, A. R., Khoruts, A., & Sadowsky, M. J. (2017). Interaction of gut microbiota with bile acid metabolism and its influence on disease states. Applied Microbiology and Biotechnology, 101(1), 47–64. https://doi.org/10.1007/s00253-016-8006-6
Sun, R., Yang, N., Kong, B., Cao, B., Feng, D., Yu, X., Ge, C., Huang, J., Shen, J., Wang, P., Feng, S., Fei, F., Guo, J., He, J., Aa, N., Chen, Q., Pan, Y., Schumacher, J. D., Yang, C. S., Guo, G. L., … Wang, G. (2017). Orally administered berberine modulates hepatic lipid metabolism by altering microbial bile acid metabolism and the intestinal FXR signaling pathway. Molecular pharmacology, 91(2), 110–122. https://doi.org/10.1124/mol.116.106617
Centanni, M., Lawley, B., Butts, C. A., Roy, N. C., Lee, J., Kelly, W. J., & Tannock, G. W. (2018). Bifidobacterium pseudolongum in the Ceca of rats fed hi-maize starch has characteristics of a keystone species in bifidobacterial blooms. Applied and Environmental Microbiology, 84(15), e00547-e618. https://doi.org/10.1128/AEM.00547-18
Acknowledgements
The author is thankful to the Indian Institute of Technology (BHU) and MHRD, India, for fellowship. The authors thankfully acknowledge to Prof. Sushant Kumar Shrivastava formal Head of Department, Pharmaceutical Engineering and Technology, IIT (BHU) Varanasi-221 005, Uttar Pradesh, India, for guidance during the work. The author is also thankful to SATHI BHU for the High-Resolution Accurate Mass Spectrometry (HRMS) study for plant metabolomics.
Author information
Authors and Affiliations
Contributions
Amit Kumar Singh: conceptualization, methodology, writing-original draft preparation. Pradeep Kumar: methodology, writing—original draft preparation. Vishnu D Rajput: conceptualization, formal analysis, validation. Sunil Kumar Mishra: conceptualization, methodology, formal analysis, supervision. Kavindra Nath Tiwari: conceptualization, formal analysis, validation. Anand Kumar Singh: formal analysis, investigation, writing—review and editing. Tatiana Minkina: formal analysis, writing—review and editing, validation. Ajay Kumar Pandey: formal analysis, validation. Prabhat Upadhyay: formal analysis, validation. All authors have read and agreed to the final version of the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
All of the tests were carried out in compliance with Institutional animal experimentation rules. The Central Animal Ethics Committee, as per CPCSEA guidelines at IMS BHU Varanasi 221005 (Reg. No. 542/GO/Rebi//S/02/CPCSEA dated 26.5.2017), approved this study (Approval number: Dean/2021/IAEC/2560). The CPCSEA guidelines were fully followed in the experiment.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, A.K., Kumar, P., Mishra, S.K. et al. A Dual Therapeutic Approach to Diabetes Mellitus via Bioactive Phytochemicals Found in a Poly Herbal Extract by Restoration of Favorable Gut Flora and Related Short-Chain Fatty Acids. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-024-04879-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-024-04879-6