Abstract
Non-histone protein acetylation is involved in key cellular processes both in eukaryotes and prokaryotes. Acetylation in bacteria is used to modify proteins involved in metabolism and allow the bacteria to adapt to their environment. TTE (Thermoanaerobacter tengcongensis) is an anaerobic, thermophilic saccharolytic bacterium that grows at extreme temperature range between 50 and 80 ℃. The annotated TTE proteome contains less than 3000 proteins. We analyzed the proteome and acetylome of TTE using 2DLC-MS/MS (2-dimensional liquid chromatography mass spectrum). We evaluated the ability of mass spectrometry technology to cover a relatively small proteome as much as possible. And we also observed wide spread of acetylation in TTE, which changed under different temperatures. A total of 2082 proteins were identified, which accounts for about 82% of the database. A total of 2050 (~ 98%) proteins were quantified in at least one culture condition and 1818 proteins were quantified in all 4 conditions. The result also consisted 3457 acetylation sites corresponding to 827 distinct proteins, which covered 40% of the proteins identified. Bioinformatics analysis reported that proteins related to replication, recombination, repair, and extracellular structure cell wall biogenesis had more than half members acetylated, while energy production, carbohydrate transport, and metabolism related proteins were least acetylated. Our result suggested that acetylation affects the ATP-related energy metabolism and energy-dependent biosynthesis process. Comparing the enzymes related with lysine acetylation and acetyl-CoA (acetyl-coenzyme A) metabolism, we suggested that the acetylation of TTE took a non-enzymatic mechanism and affected by abundance of acetyl-CoA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-histone protein acetylation is involved in key cellular processes both in eukaryotes and prokaryotes. It is generated by transfer of an acetyl group from acetyl-CoA (acetyl-coenzyme A) to the ε-amino side chain of lysine, which is catalyzed by KATs (lysine acetyltransferases) and is reversable by KDACs (lysine deacetylases) [1]. There is also non-enzymatic acetylation affected by pH, proximal amino acid residues, and the availability of acetyl-CoA [2]. It can affect the activity of enzymes, the stability of proteins, and the interaction of proteins with other molecules. Acetylation can also play a role in the transport of metabolites and in the assembly of multi-protein complexes [3, 4]. In bacteria, this process is used to modify proteins involved in metabolism. The acetylation of these proteins can change their function and allow the bacteria to adapt to their environment [5]. Research of Salmonella enterica samples has found that acetylation modification occurs in cellular cytosol, metabolic enzymes are highly acetylated, and acetylation modification regulates the activity of metabolic and metabolic enzymes [6].
TTE (Thermoanaerobacter tengcongensis) is an anaerobic, thermophilic, rod-shaped, gram-negative saccharolytic bacterium. In 2004, TTE was systematically classified and renamed as Caldanaerobacter subterraneus subsp. tengcongensis [7]. It was first found in a hot spring in Tengchong, Yunnan Province, China, in 1998 and reported in 2001 [8]. It can grow at temperature range between 50 and 80 ℃, with an optimum of 75 ℃. In 2002, the first genome sequencing and annotation were done by Bao and colleagues [9]. The organism can utilize glucose, galactose, and other carbohydrates. Acetate was the main end product from glucose fermentation. The PTM (post-translational modification) research of TTE was rarely involved, with preliminary investigations of dozens of sites of scale in phosphorylation modifications and large-scale study absent [10, 11]. The ability of TTE to grow at in extreme environments makes it a potential candidate for use in thermal-related biotechnology applications and the temperature-dependent biological investigations.
The human proteome project strives to profile proteome or modifications in the sample. However, it is difficult to define whether the unidentified protein is biologically absent or not detectable by current instrument. One approach was to start from the genome as simple as possible. The TTE proteome contains 2545 proteins (UNIPROT) and we used 2DLC-MSMS technology to identify proteins and acetylation sites. On the one hand, we investigated the coverage of mass spectrometry technology on the research object; and on the other hand, we observed the identification process of acetylation in the bacterial genome and the changes under different temperatures. We used various bioinformatic annotations and methods to orient the extraction of trends of acetylation alteration for further discussion.
Methods
Materials
Urea, thiourea, Tris–HCl, CHAPS, and guanine-HCl were purchased from Sigma-Aldrich (St. Louis, MO, USA). LC–MS grade water, acetonitrile, and formic acid were purchased from Merck Millipore (Billerica, MA, USA). Sequencing-grade trypsin was purchased from Promega (Madison, WI, USA).
Sample Preparation and Digestion
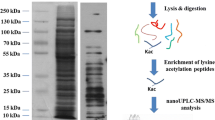
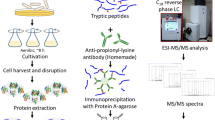
The bacteria were first inoculated at 75 ℃ to the log phase, then distributed to the media (at a ratio of 1:50) prewarmed at 55, 65, 75, and 80, respectively. The bacterial cells were harvested at log phase by centrifugation at 4000 g at 4 ℃, and the pellets were washed with the buffer of 50 mM Tris–HCl, pH 7.8. The washed bacteria were stored at – 80 ℃. Extraction buffer containing 8 M urea, 2 M thiourea, 4% CHAPS, 10 mM DTT (dithiotreitol), and 50 mM Tris–HCl, pH 8.0, was added to extract the proteome with MiniBeadbeater-16 bead grinding tissue grinder. After protein quantitation, 2 SDS-PAGE (sodium dodecyl sulfate–polyacrylamide gel electrophoresis) experiments were performed simultaneously and 0.2 mg protein was loaded. Only 1 piece of gel was dyed using CBB (Coomassie brilliant blue) and named sample CBB in further analysis, while the other transparent piece named noCBB. Another aliquot of 0.2 mg protein was cleansed by acetone precipitation and the leftover was stored at – 80 ℃.
The SDS-PAGE gel pieces were cut evenly to 20 fragments, then in-gel digestion was performed as previously described [12]. For solution sample, sequencing-grade trypsin was added to the guanine-HCl re-lysed proteins by a ratio of 1:50. Then the mixture was incubated at 37 ℃ overnight with constant shaking. Formic acid was added to 5% concentration (vol/vol) to terminate the digestion. After desalting, the lyophiled samples were stored at – 20 ℃ before further analysis.
High pH LC
High-pH LC separation was performed on an HPLC system composed of an LC-20AB LC pumps and an SIL-20 AC auto-sampler (all Shimadzu, Tokyo, Japan). Mobile phase A was 2% acetonitrile and 20 mM NH4FA (pH adjusted to 10 with ammonium); mobile phase B was 90% ACN and 20 mM NH4FA (pH also adjusted to 10). Elution profile included 2–8% of solution B in the first 10 min, then solution B from 8 to 35% in 55 min, then solution B from 35 to 60% in 15 min. The flow rate was 0.2 mL/min and the peptides were monitored at 214 nm. The fractioned peptides were collected at one tube/min during the linear elution period then mixed to 33 samples for LC–MS/MS injection.
Nano-LC–MS/MS
Nano-LC MS/MS experiment was performed on an HPLC system composed of two LC-20AD nano-flow LC pumps, an SIL-20 AC auto-sampler, and an LC-20AB micro-flow LC pump (all Shimadzu, Tokyo, Japan) connected to a Q exactive mass spectrometer (ThermoFisher, San Jose, CA). Sample was loaded directly onto a C18 reverse-phase column (0.075 × 500 mm, Monolith column, Omicsolution Co. Ltd. Shanghai, China) at a flow rate of 300 nL/min. The mobile phases were 2% acetonitrile with 0.1% formic acid (phase A and the loading phase) and 90% acenitrile with 0.1% formic acid (phase B). To achieve proper separation, a 35-min linear gradient from 2 to 45% phase B was employed. The separated sample was introduced into the mass spectrometer via the nano-spray interface supplied by the mass spectrometer vendor. The spray voltage was set at 2.2 kV and the heated capillary at 180 ℃. The mass spectrometer was operated in data-dependent mode and each cycle of duty consisted one full-MS survey scan at the mass range 300–1400 Da with resolution power of 70,000, followed by MS/MS experiments for 15 strongest peaks. Peptides were fragmented using HCD (high-energy collision-induced dissociation) and the normalized collision energy value set at 30%. 2 + , 3 + , and 4 + peaks were selected for MS/MS run and previously fragmented peptides were excluded for 60 s. Each sample was analyzed twice.
Database Search
The TTE proteome database was downloaded from the UNIPROT website (https://www.uniprot.org/proteomes/UP000000555). This database contains 2545 entries. Protein identification and quantitation were performed with the FRAGPIPE software suite. Module MSFragger [13, 14] did the database search using the database containing the TTE proteome and common contaminant proteins and their reversed counterparts. Cysteine carbamidomethylation was set as fixed modification and protein N-term acetylation, lysine acetylation, and methione oxidation were set as variable modifications. We used module Philosopher [15] to accomplish the validation step. Quantitation was done with module IonQuant [16] running MS1 quant mode and LFQ option. All other parameters of modules mentioned above were set default. After the tabular result files were created, downstream analysis was carried out using Microsoft Excel or in-house composed R script.
Bioinformatics Analysis
CGVIEW was used to generate circular visualization for TTE genome [17]. The proteome profiles were clustered based on Euclidean distances with average linkage by R pheatmap package. We only clustered the expression profiles and the temperatures were not clustered. The functional annotations and sequence features on each protein of TTE were derived from the UNIPORT database. DAVID (https://david.ncifcrf.gov/) was used to annotate the result then categorized into molecular function, biological process, and cellular component. We used GPS-PAIL [18] and modified the parameters to predict the acetylation probability of the lysines in TTE proteins. All 7 types of acetyltransferases supported by GPS were included in the prediction, which were CREBBP, EP300, HAT1, KAT2A, KAT2B, KAT5, and KAT8, respectively. Simple prediction mode, which took no account of annotations of secondary structure and surface accessibility, was used. Its result cutoff was set that prediction scores of high, medium, and low for each site were picked out.
Results
Overview of Proteome and Acetylated Proteome of TTE
We used 2DLC-MS/MS to quantitatively profile the proteome and acetylome of TTE at 4 temperatures. According to literature [8], the growth rate is approximately 0.04/h, 0.12/h, 0.27/h, and 0.07/h at each temperature. All identified proteins were aligned to their locus in the TTE genome and plotted as bars. The acetylation analysis result was also compared with predicted acetylation sites (Fig. 1). All the acetylation sites were color coded and aligned as the 4 outmost rings of the Circos diagram. The colors of the bars were temperature dependent and the selected gene tags were given different colors based on the clustering results described below. The acetylation sites distributed quite evenly on the genome rather than several gene clusters. The mid ring was the predicted acetylation site distribution. The alignment of the outer and mid rings showed that most gaps in outer rings located at the gaps of prediction. The inner rings were results aligned to the genome research and predictions. The gaps in the acetylation distribution were also aligned to the peaks and valleys of the GC content ring. The prediction showed that there were more acetylation incidents in the minus skew of genome and our result fits it well. As counted in Table 1, 2082 proteins and 64,493 peptides were identified, which accounts for about 82% of total 2545 entries of the database. On the quantitative side, 2050 proteins (98% of all proteins) were quantified in at least one culture condition and 1818 proteins were quantified in all 4 conditions (Fig. 2A). The number of proteins quantified for each temperature was 1950, 1974, 1950, and 1981, respectively. The protein abundance span 5 orders. Average quantified peptide number of each protein was 21 for the whole experiment and ranged from 17 to 19 for samples of different temperatures. A total of 44,160 peptides (68% of all peptides) were quantified and the peptide abundance also span 5 orders. Comparing our result with previous study [19] for TTE cultured at the same temperature series using iTRAQ technology, 1515 proteins (95% of the proteins in [19] and 83% of our result) were quantified throughout all 4 conditions in both studies.
Overview of acetylation sites and prediction sites identified by TTE. Circular genome plot contained the genome information and annotation as the most inner circles, acetylation sites predicted as the middle circle, and the acetylation sites from each sample stacked as the outer circles. The locus and amino acid sites were aligned
Comparison of proteome data between each temperature for TTE. A Sample-wise protein distribution. The lower-left section: horizontal bar plot of protein identification. Protein distribution was displayed as dot and connected line beneath the bar plot on the right side. B Sample-wise acetylated protein distribution. C Sample-wise acetylation site distribution
The result also consisted 3457 acetylation sites corresponding to 827 distinct proteins, which covered 40% of the proteins identified. The temperature-wise protein counts were 748, 679, 753, and 671, while acetylation site distribution was 1823, 1849, 2621, and 2354 for each temperature. Although acetylated protein number varied slightly with different temperatures, acetylation sites differed significantly at higher temperatures, suggesting more acetylation activity. A total of 388 (~ 47%) acetylated proteins were identified from all 4 conditions, while 62 (~ 7%) acetylated proteins were identified from only 1 condition (Fig. 2B). A total of 632 (~ 18%) acetylation sites were identified from all 4 conditions, while 472 (~ 14%) sites were identified from only 1 condition (Fig. 2C). The site distribution indicated that there were most lysine acetylation events when TTE growth flourished. Moreover, the heterogeneity of acetylation pattern suggested that culture temperature has complicated effect on acetylation and deacetylation process at each condition.
The UNIPROT TTE proteome database only included 13 proteins with PTM annotation and 70 proteins with modified residue information; none of them was about lysine acetylation. To assess the acetylation sites reported, we used the GPS [18] software suite to provide the possibility of each protein recorded in the database of TTE proteome being acetylated. The prediction was present as high, median, and low, suggesting the possibility of specific lysine site being acetylated. Among all 66,951 lysine sites in the proteome database, number of high, median, and low were 4893, 10,143, and 16,966, respectively, forming a roughly 1:2:3.5 distribution and the site distribution alongside the genome was roughly even. The corresponding numbers in our data were 202, 461, and 836. We identified relatively more in-silico medium and low confident acetylation sites, which indicated the benefit of wet experiment as supplement to software prediction. From sample cultured in 80 ℃, PKN1_CALS4 K23 was one of the acetylation sites that were not scored by GPS and identified in the LC–MS data. METK_CALS4 K276 was one of the acetylation sites that were scored “high” by GPS and identified in the LC–MS data. We made no discrimination to acetylation sites with or without GPS prediction. Both peptides received expectation score < 0.0001 and treated equally in further analysis.
Proteome of TTE Derived at Different Temperatures
Temperature-dependent proteome expression profiles were clustered into 7 clusters using hierarchy clustering method (Supple. Table 1). The only 2 clusters (clusters 2 and 5) with consistent tendency had most cluster members, where 566 (counts for 27.6% of total proteome) proteins showed consistent escalating trend, and DAVID analysis for GO-annotation enrichment found that BP of translation, pentose-phosphate shunt, transcription, DNA-templated and MF of DNA binding, structural constituent of ribosome, rRNA binding, GTP binding, GTPase activity, DNA-directed 5′–3′ RNA polymerase activity, NADP binding enriched in cluster 2 (Figs. 3 and 4). The 374 (counts for 18.2%) proteins’ abundance kept declining as temperature rose and BP of DNA recombination, “de novo” UMP biosynthetic process and MF of catalytic activity, peptidase activity, electron carrier activity, sequence-specific DNA binding enriched in cluster 5. Considering the optimum culture temperature 75 ℃, cluster 1 (definite minimum at 75 ℃) enriched in biosynthesis including branched-chain amino acid, leucine, fatty acid, and valine biosynthetic process and metal ion including 4 iron–4 sulfur cluster and iron-sulfur cluster binding. The profile suggested that corresponding proteins were metal-binding enzymes taking part in amino acids and fatty acid biosynthesis processes. While those enzyme abundance were at their bottom and TTE growth rate was largest at 75 ℃, such deficit supported an opinion that there was other mechanism such as chaperones assured the growth.
Acetylation Proteome of TTE Derived at Different Temperatures
Temperature-dependent acetylated proteome expression profiles were also clustered (Supple. Table 2). Acetylated proteins identified at four different temperatures were with smaller difference in quantity than proteome. Hierarchy clustering resulted into 9 clusters. Similar to the clusters of protein abundance, the 2 clusters with consistent change tendency also had most cluster member counts, where cluster 4 with 166 (counts for 20% of total acetylated proteome) proteins showed consistent escalating trend with different temperatures and cluster 6 with 135 (counts for 16%) proteins declining. DAVID GO-annotation enrichment analysis found that MF term ATP binding enriched in cluster 4. BP term “de novo” UMP biosynthetic process, MF term oxidoreductase activity, and GTPase activity enriched in cluster 6. Both clusters enriched in energy metabolism.
Analysis of Cluster of Orthologous Group
According to the analysis of COG (cluster of orthologous group), we could categorize the temperature-dependent proteins into 25 groups (Fig. 5A). Full annotation for each symbol of COG was derived at NCBI COG database (https://www.ncbi.nlm.nih.gov/research/cog/). Because 2 groups were poorly characterized and other 2 groups were ambiguous, the functional categories for the temperature-dependent proteins were simplified to 23 groups. We analyzed the function distributions of the 2050 identified proteins derived from the TTE genome based on these 23 function groups. We also analyzed the acetylation data. We estimated the proportional distributions for the two data sets of the COG analysis. After excluding general function prediction only or COG entries with less than 10 members, 2 groups, namely “replication, recombination, and repair” and extracellular structures cell wall/membrane/envelope biogenesis had more than half members acetylated. Defense mechanisms, amino acid transport and metabolism, energy production and conversion, and carbohydrate transport and metabolism were least acetylated and the portion was less than 20%.
Correlation analysis between protein expression and acetylation sites. All intensity data were log10 transformed before plotting. A COG analysis resulted in categorizing the proteome into 25 groups; the expression pattern of each COG group was drawn as broken line connecting the median abundance of each temperature beneath the heatmap and the zenith of each pattern was highlighted in red. B Spearman correlation coefficient of each protein expression cluster and cluster of acetylated proteins; the overlay of protein cluster 2 and acetylation cluster 7 (the only pair that its coefficient = 1) labeled as star symbol. C Bar plot: summed raw intensity of each protein (orange) and each acetylated peptide (blue); dotted line: ratio of the 2 sums above. D Spearman correlation coefficient of temperature-wise acetylation abundance and protein abundance of the 827 acetylated proteins plotted at descending style; GO terms enriched for proteins that its coefficient > 0.8 or < − 0.8 labeled at corners of the plot
Correlation of TTE Proteome with Acetylation Proteome at Different Temperatures
Spearman correlation (Fig. 5B) was calculated across the abundance pattern of all clusters. DAVID analysis was performed for the overlay protein members of cluster pairs whose Spearman correlation coefficient > 0.95 (protein cluster 2 and acetylation cluster 7, protein 6 and acetylation 5, protein 3 and acetylation 4) or < − 0.95 (protein 5 and acetylation 4). The molecular function category was dominated by enzymes, and the biological process category was composed of ribosomal proteins and proteins involved in cell wall biogenesis. The cellular component category was represented by proteins related to the cell membrane and proteins related with DNA, RNA, and proteins. The result was limited due to member counts of the scarce overlay protein counts. Only the overlay proteins of protein cluster 2 and acetylation cluster 7 (coefficient = 1) showed enrichment of “ATP binding” on the MF GO term and “nucleotide binding” on the keyword ligand term. This result suggests acetylation impacts on the energy-dependent biosynthesis process.
Level of acetylation was calculated as ratio of summed acetylated peptide signal intensity to summed intensity of all peptides at each temperature (Fig. 5C). Due to the low abundance nature of acetylation and the lack of modification enrichment method, only about 5% of acetylated peptides were successfully quantified. However, we obtained a non-ambiguous result. When plotted against temperature, the data showed a “smile” shape curve. It halved when comparing samples from 6 and 55 ℃, while escalated at temperatures of 75 ℃ and 80 ℃. Despite having most acetylation sites, the optimum temperature of 75 ℃ did not bring to highest level of acetylation. Level of acetylation was higher as it reached close to the lower and high limit of TTE survival temperature [8], suggesting that increasing level of acetylation may function as counter measurement of TTE against extreme environment.
The overlap of proteome and acetylome of TTE contained 827 proteins. Spearman correlation coefficient was used to analyze the correlation between the expression profile of acetylated protein at different temperatures (Fig. 5D). Spearman correlation coefficient is arranged in descending order. Proteins at either end of 285 proteins’ coefficient were larger than 0.8 and 112 proteins less than − 0.8. After functional analysis, proteins enriched in those their abundance and acetylation profile correlated well were aminopeptidases and magnesium ion binding enzymes, indicating that acetylation regulates translation at the same orientation as these proteins. While ATP binding was the sole enriched function proteins in the lower coefficient section, indicating that the correlation between acetylation and energy metabolism for these proteins were at the opposite orientation.
Discussion
Acetylation Sites on Protein Fragments with Biological Functions
The results of bioinformatic analysis showed that the change of acetylome was connected to metabolism. The sequence feature analysis of sequences around the acetylation sites, ATP binding emerged as majority in the “binding site” profile (Fig. 6A). In general, the neutral acetyl group neutralizes the positive charge of lysine side chain, therefore, affecting the binding events [20]. Although according to function annotation, the proteins binding ATP were all ATPase including subunits or have the ability to hydrolyze ATP. Most acetylation level differs slightly at all 4 temperatures, except Q8R6W0_CALS4 (iron-regulated ABC transporter ATPase subunit SufC, product of gene TTE2673). There is limited function annotation of TTE2673. Another protein that significantly changed acetylation level was GPMA_CALS4 (2,3-bisphosphoglycerate dependent phosphoglycerate mutase, product of gene gpmA), a vital member of glycolysis that catalyzes the interconversion of 2-phosphoglycerate and 3-phosphoglycerate [21]. The acetylation site was K98, which is annotated to the substrate binding site. Acetylation can affect the binding efficiency of substrate and enzyme, thus, the glycolysis pathway. In Escherichia coli, acetylation at or close to binding sites limits or inhibits the protein’s ability of binding [22, 23]. Our data indicated that TTE glycolysis pathway participates in growth and alters at different temperatures.
Analysis of acetylation sites appearing in functional segments of protein sequences. The data were log10 transformed before plotting. A Intensity of each acetylated peptide at all temperatures, which has annotation of “binding site” of energy metabolism-related chemicals. B Intensity distribution of all AccA2 acetylated peptides. C Intensity distribution of all glmU acetylated peptides. D Abundance distribution of PaaJ, plotted as intensity IonQuant reported. E Abundance distribution of cobB1 and cobB2. G Intensity distribution of cobB1 and cobB2
We found that SYI_CALS4 (product of gene ileS) was acetylated at the site K600. The modification occurred only at 5 and 65 ℃ with a declining trend (Fig. 6F). SYI is isoleucine-tRNA ligase and catalyzes the attachment of isoleucine to tRNA (Ile). The acetylation site was at the beginning of annotated motif of “KMSKS” motif, motif that catalyzes the amino acid activation with ATP [24]. As false chemical such as structurally similar amino acid valine can inadvertently cause false accommodation and processing, SYI has two additional distinct tRNA (Ile)-dependent editing activities. One is function through the hydrolysis of activated Val-AMP and the other activity involves deacylation of mischarged Val-tRNA (Ile) [25]. The structure of SYI_CALS4 predicted by AlphaFold2 (https://alphafold.ebi.ac.uk/entry/Q8R9L3) suggests there was a hydrogen bond between K600 and N607. The neutral acetyl group neutralizes the positive charge of lysine side chain, therefore, affecting the hydrogen bond and structure. In Escherichia coli, acetylation regulates protein translation [26] and acetylation close to the “KMSKS” motif in leucyl-tRNA synthetase (LeuRS) downregulated its enzymatic activity [27]. We hereby suggest reduction of SYI_CALS4 enzymatic activity at unfavorable culture temperature that changes the protein structure and leads to acetylation at the ATP recognition motif, therefore, downgraded the protein translation related.
Plausible Acetylation Mechanism
The enzymes responsible for lysine acetylation are the lysine N-acetyltransferase (KAT) that catalyze the transfer of acetyl groups from acetyl-CoA to lysine residues, whereas lysine deacetylase (KDAC) removes acetyl groups from histones. Limited number of proteins (510 entries out of 2545) have enzyme class (EC, https://enzyme.expasy.org/enzyme-byclass.html) annotation. No protein has annotation of EC 2.3.1.32: lysine N-acetyltransferase and 2 proteins (NPD1_CALS4 and NPD2_CALS4, namely, NAD-dependent protein deacetylase 1 and 2, products of gene cobB1 and cobB2) were annotated as EC 2.3.1.286: protein acetyl-lysine N-acetyltransferase and noted as deacetylases. Both proteins’ abundance was much higher at higher culture temperature (Fig. 6E). We observed that the level of acetylation and number of acetylated lysine sites were both higher at 75 ℃ and 80 ℃. With abundance of deacetylases increasing, this phenomenon indicated that the acetylation pattern of TTE took a non-enzymatic mechanism and affected by abundance of acetyl-CoA. Acetyl-CoA carboxylase, a primary acetyl-CoA consumer which transfers the CO2 group to acetyl-CoA to form malonyl-CoA, was reported as a regulator to histone acetylation and can affect tumor recurrence upon primary tumor resection in mice [28, 29]. We found that acetyl-CoA carboxylase alpha subunit Q8R7L8_CALS4 (product of gene AccA2) changed less than 20% throughout the temperature span (Fig. 6G) while going through multiple acetylation. With insufficient feature description of the sequence, we roughly categorized the sequence from 1 to 200 as miscellaneous and 201 to end as sequence similar to ACCA_ECOLI. At higher temperature, all acetylations were at the second moiety (Fig. 6B) that could affect activity. Bifunctional protein GLMU_CALS4 (product of gene glmU, the C-terminal domain catalyzes the transfer of acetyl group from acetyl-CoA to glucosamine-1-phosphate to produce N-acetylglucosamine-1-phosphate) [30] also changed insignificantly but acetylated at multiple sites (Fig. 6C). K349 is annotated as UDP-N-acetyl-alpha-D-glucosamine binding site. Acetylation of K349 can hinder the binding ability and reached apex at 75 ℃, leading to least activity. We also observed Q8RC88_CALS4 (acetyl-CoA acetyltransferases, which namely removes acetyl group from acetyl-CoA to generate CoA [31], product of gene PaaJ) at their lowest abundance. Summing up the acetyl-CoA-consuming enzymes above, the acetyl-CoA reservoir was at peak level at 75 ℃ and resulted in a maximum number of acetylation sites and relatively high acetylation level following the non-enzymatic mechanism.
CBB and noCBB
As we suggest promoting TTE for biological and technology research, we checked the impact of CBB staining, the most commonly used technique in these areas which can cause loss of information with experimental destain operations to remove excessive CBB before MS analysis. We compared the results of using and not using CBB in the analysis of TTE proteome and acetylome of sample cultured at 65 °C (Fig. 7). Briefly, 2 parallel SDSPAGE were run and only one gel was stained with CBB. The results showed that the identified number of TTE proteins changed slightly (about 3%), and the identification rate of acetylation sites was greatly improved (about 70% more sites identified with more than 60% sites overlapped) when omitting the usage of CBB. We reasoned that the gain of acetylation sites was consequence of less experimental operation, which had potential to guide future experiment designs. We took a closer look at the protein level, the 58 proteins only identified from noCBB sample; about half of them were located to membrane (31 proteins) or had transmembrane (28 proteins) feature and the function annotation with most members were transport (9 proteins, Fig. 7C). We attributed the tendency of loss of membrane-related proteins to the hydrophobicity-related binding mechanism that CCB binding can be applied as an indicator of protein surface hydrophobicity [32]. For example, protein Q8R8R3_CALS4 (product of gene MelB), a well-studied Melibiose permease in prokaryotes [33, 34], was missing in the CBB sample. We suggest that future studies on hydrophobic protein with modern instruments should use CBB carefully.
Comparison of identification of protein and acetylation sites between CBB and noCBB. A Venn plot of identified proteins from CBB and noCBB samples. B Venn plot of identified acetylated sites from CBB and noCBB samples. C Feature distribution of the 58 proteins only identified from noCBB sample. The intensity of each protein was plotted at descending order. The title of each slice indicated the enriched features of the 58 proteins. Proteins fit the corresponding feature were highlighted with red color
Conclusion
TTE has the possibility of wide acetylation. By using 2DLC-MSMS, we investigated the profile and change of TTE proteome and acetylome at 4 different temperatures. Our result covered 82% of the TTE genome and quantified 98% of them. A total of 3457 acetylation sites corresponding to 827 distinct proteins were found, which covered 40% of the proteins identified. We found that acetylome changed more extensively with temperature than proteome. We connected the change of acetylation of ileS with temperature to its enzymatic activity. After reviewing the abundance and acetylation alteration of related enzymes, we suggested that the acetylome of TTE takes a non-enzymatic mechanism and affected by abundance of acetyl-CoA. The TTE is a model organism with small genome, which can be used to investigate the ability of mass spectrometry technology. We suggest promoting such TTE for further biological and technology research, while CBB should be used with caution.
Data Availability
The raw data of proteome and acetylome were deposited at iProx (www.iprox.org) with the accession number of IPX0005333000 or ProteomeXchange (http://proteomecentral.proteomexchange.org) with the accession number of PXD038221.
Abbreviations
- 2DLC-MS/MS:
-
2-Dimensional liquid chromatography mass spectrum
- CBB:
-
Coomassie brilliant blue
- COG:
-
Cluster of orthologous group
- DAVID:
-
Database for Annotation, Visualization and Integrated Discovery
- DEPs:
-
Differential expressed proteins
- GO:
-
Gene Ontology
- GPS:
-
Group-based prediction system
- HCD:
-
High-energy collision-induced dissociation
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- noCBB:
-
Sample that was not stained by Coomassie brilliant blue
- SDSPAGE:
-
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- PSMs:
-
Peptide spectrum matches
- PTMs:
-
Post-translational modifications
- TTE:
-
Thermoanaerobacter tengcongensis
References
Narita, T., Weinert, B. T., & Choudhary, C. (2019). Functions and mechanisms of non-histone protein acetylation. Nature Reviews Molecular Cell Biology, 20, 156–174.
James, A. M., Hoogewijs, K., Logan, A., Hall, A. R., Ding, S., Fearnley, I. M., & Murphy, M. P. (2017). Non-enzymatic N-acetylation of lysine residues by acetylCoA often occurs via a proximal S-acetylated thiol intermediate sensitive to glyoxalase II. Cell Reports, 18, 2105–2112.
Drazic, A., Myklebust, L. M., Ree, R., & Arnesen, T. (2016). The world of protein acetylation. Biochimica Et Biophysica Acta-Proteins and Proteomics, 1864, 1372–1401.
Shvedunova, M., & Akhtar, A. (2022). Modulation of cellular processes by histone and non-histone protein acetylation. Nature Reviews Molecular Cell Biology, 23, 329–349.
Carabetta, V. J., Cristea, I. M. (2017). Regulation, function, and detection of protein acetylation in bacteria. Journal of Bacteriology 199.
Wang, Q., Zhang, Y., Yang, C., Xiong, H., Lin, Y., Yao, J., Li, H., Xie, L., Zhao, W., Yao, Y., Ning, Z.-B., Zeng, R., Xiong, Y., Guan, K.-L., Zhao, S., & Zhao, G.-P. (2010). Acetylation of metabolic enzymes coordinates carbon source utilization and metabolic flux. Science, 327, 1004.
Fardeau, M.-L., Salinas, M. B., Haridon, S., Jeanthon, C., Verhé, F., Cayol, J.-L., Patel, B. K. C., Garcia, J.-L., & Ollivier, B. (2004). Isolation from oil reservoirs of novel thermophilic anaerobes phylogenetically related to Thermoanaerobacter subterraneus: Reassignment of T. subterraneus, Thermoanaerobacter yonseiensis, Thermoanaerobacter tengcongensis and Carboxydibrachium pacificum to Caldanaerobacter subterraneus gen. nov., sp. nov., comb. nov. as four novel subspecies. International Journal of Systematic and Evolutionary Microbiology, 54, 467–474.
Xue, Y. F., Xu, Y., Liu, Y., Ma, Y. H., & Zhou, P. J. (2001). Thermoanaerobacter tengcongensis sp nov., a novel anaerobic, saccharolytic, thermophilic bacterium isolated from a hot spring in Tengcong, China. International Journal of Systematic and Evolutionary Microbiology, 51, 1335–1341.
Bao, Q. Y., Tian, Y. Q., Li, W., Xu, Z. Y., Xuan, Z. Y., Hu, S. N., Dong, W., Yang, J., Chen, Y. J., Xue, Y. F., Xu, Y., Lai, X. Q., Huang, L., Dong, X. Z., Ma, Y. H., Ling, L. J., Tan, H. R., Chen, R. S., Wang, J., … Yang, H. M. (2002). A complete sequence of the T tengcongensis genome. Genome Research, 12, 689–700.
Lin, W., Wang, J., Ying, W., & Qian, X. (2012). Optimization of titanium dioxide enrichment of phosphopeptides and application in the Thermoanaerobacter tengcongensis phosphoproteome analysis. Se pu = Chinese journal of chromatography, 30, 763–769.
Lee, K. O., & Suh, J.-Y. (2017). Backbone assignment of phosphorylated cytoplasmic domain B of mannitol transporter IIMtl in Thermoanaerobacter tengcongensis. Journal of the Korean Magnetic Resonance Society, 21, 20–25.
Hu, Y., Yao, J., Wang, Z., Liang, H., Li, C., Zhou, X., Yang, F., Zhang, Y., & Jin, H. (2022). Comparative proteomic analysis of drug trichosanthin addition to BeWo cell line. Molecules, 27, 1603.
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D., & Nesvizhskii, A. I. (2017). MSFragger: Ultrafast and comprehensive peptide identification in mass spectrometry–based proteomics. Nature Methods, 14, 513–520.
Teo, G. C., Polasky, D. A., Yu, F., & Nesvizhskii, A. I. (2021). Fast Deisotoping Algorithm and Its Implementation in the MSFragger Search Engine. Journal of proteome research, 20, 498–505.
da Veiga Leprevost, F., Haynes, S. E., Avtonomov, D. M., Chang, H.-Y., Shanmugam, A. K., Mellacheruvu, D., Kong, A. T., & Nesvizhskii, A. I. (2020). Philosopher: A versatile toolkit for shotgun proteomics data analysis. Nature Methods, 17, 869–870.
Yu, F., Haynes, S. E., & Nesvizhskii, A. I. (2021). IonQuant enables accurate and sensitive label-free quantification with FDR-controlled match-between-runs. Molecular & Cellular Proteomics, 20, 100077.
Stothard, P., & Wishart, D. S. (2005). Circular genome visualization and exploration using CGView. Bioinformatics, 21, 537–539.
Wang, C. W., Xu, H. D., Lin, S. F., Deng, W. K., Zhou, J. Q., Zhang, Y., Shi, Y., Peng, D., & Xue, Y. (2020). GPS 5.0: An update on the prediction of kinase-specific phosphorylation sites in proteins. Genomics Proteomics & Bioinformatics, 18, 72–80.
Chen, Z., Wen, B., Wang, Q., Tong, W., Guo, J., Bai, X., Zhao, J., Sun, Y., Tang, Q., Lin, Z., Lin, L., & Liu, S. (2013). Quantitative proteomics reveals the temperature-dependent proteins encoded by a series of cluster genes in Thermoanaerobacter tengcongensis. Molecular & Cellular Proteomics, 12, 2266–2277.
Ali, I., Conrad, R. J., Verdin, E., & Ott, M. (2018). Lysine acetylation goes global: From epigenetics to metabolism and therapeutics. Chemical Reviews, 118, 1216–1252.
Bond, C. S., White, M. F., & Hunter, W. N. (2002). Mechanistic implications for Escherichia coli cofactor-dependent phosphoglycerate mutase based on the high-resolution crystal structure of a vanadate complex1 1Edited by R. Huber. Journal of Molecular Biology, 316, 1071–1081.
Molloy, D., Mapp, K. L., Webster, R., Gallimore, P. H., & Grand, R. J. A. (2006). Acetylation at a lysine residue adjacent to the CtBP binding motif within adenovirus 12 E1A causes structural disruption and limited reduction of CtBP binding. Virology, 355, 115–126.
Li, S., Zhang, Q., Xu, Z., Yao, Y.-F. (2017). Acetylation of lysine 243 inhibits the oriC binding ability of DnaA in Escherichia coli. Frontiers in Microbiology 8.
Kobayashi, T., Takimura, T., Sekine, R., Vincent, K., Kamata, K., Sakamoto, K., Nishimura, S., & Yokoyama, S. (2005). Structural snapshots of the KMSKS loop rearrangement for amino acid activation by bacterial tyrosyl-tRNA synthetase. Journal of Molecular Biology, 346, 105–117.
Bishop, A. C., Beebe, K., & Schimmel, P. R. (2003). Interstice mutations that block site-to-site translocation of a misactivated amino acid bound to a class I tRNA synthetase. Proceedings of the National Academy of Sciences, 100, 490–494.
Feid Sarah, C., Walukiewicz Hanna, E., Wang, X., Nakayasu Ernesto, S., Rao Christopher, V., & Wolfe Alan, J. (2022). Regulation of translation by lysine acetylation in Escherichia coli. mBio, 13, e01224-01222.
Moras, D. (1992). Structural and functional relationships between aminoacyl-tRNA synthetases. Trends in Biochemical Sciences, 17, 159–164.
Galdieri, L., & Vancura, A. (2012). Acetyl-CoA carboxylase regulates global histone acetylation. Journal of Biological Chemistry, 287, 23865–23876.
Rios Garcia, M., Steinbauer, B., Srivastava, K., Singhal, M., Mattijssen, F., Maida, A., Christian, S., Hess-Stumpp, H., Augustin, H. G., Müller-Decker, K., Nawroth, P. P., Herzig, S., & Berriel Diaz, M. (2017). Acetyl-CoA carboxylase 1-dependent protein acetylation controls breast cancer metastasis and recurrence. Cell Metabolism, 26, 842-855.e845.
Green, O. M., McKenzie, A. R., Shapiro, A. B., Otterbein, L., Ni, H., Patten, A., Stokes, S., Albert, R., Kawatkar, S., & Breed, J. (2012). Inhibitors of acetyltransferase domain of N-acetylglucosamine-1-phosphate-uridyltransferase/glucosamine-1-phosphate-acetyltransferase (GlmU). Part 1: Hit to lead evaluation of a novel arylsulfonamide series. Bioorganic & Medicinal Chemistry Letters, 22, 1510–1519.
Teufel, R., Mascaraque, V., Ismail, W., Voss, M., Perera, J., Eisenreich, W., Haehnel, W., & Fuchs, G. (2010). Bacterial phenylalanine and phenylacetate catabolic pathway revealed. Proceedings of the National Academy of Sciences, 107, 14390–14395.
Cao, Y., Zhao, J., & Xiong, Y. L. (2016). Coomassie brilliant blue-binding: A simple and effective method for the determination of water-insoluble protein surface hydrophobicity. Analytical Methods, 8, 790–795.
Pourcher, T., Leclercq, S., Brandolin, G., & Leblanc, G. (1995). Melibiose permease of Escherichia coli: Large scale purification and evidence that H+, Na+, and Li+ sugar symport is catalyzed by a single polypeptide. Biochemistry, 34, 4412–4420.
Guan, L., Nurva, S., & Ankeshwarapu, S. P. (2011). Mechanism of melibiose/cation symport of the melibiose permease of Salmonella typhimurium*. Journal of Biological Chemistry, 286, 6367–6374.
Acknowledgements
We are so grateful to the selfless help from the Shanghai Huisen Science & Technology Company for data analysis.
Funding
This research was supported by the National Key R&D Program of China (2021YFF0703702) and the National Nature Science Foundation of China (32070605).
Author information
Authors and Affiliations
Contributions
Conception and design: Jun Yao and Yang Zhang. Administrative support: Hong Jin and Yang Zhang. Collection and assembly of data: Jun Yao. Data analysis and interpretation: Yang Zhang and Zening Wang. Manuscript writing: all authors. Final approval of manuscript: all authors.
Corresponding authors
Ethics declarations
Ethics Approval
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yao, J., Wang, Zn., Liu, H. et al. Survey of Acetylation for Thermoanaerobacter tengcongensis. Appl Biochem Biotechnol 195, 6081–6097 (2023). https://doi.org/10.1007/s12010-023-04361-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-023-04361-9