Abstract
Recent studies have revealed the physiological significance of post-translational lysine acylations such as acetylation in the regulation of various cellular processes. Here, we characterized lysine propionylation, a recently discovered post-translational acylation, in five representative bacteria: Geobacillus kaustophilus, Thermus thermophilus, Escherichia coli, Bacillus subtilis, and Rhodothermus marinus. Using antibody-based propionyl peptide enrichment followed by identification with nano-liquid chromatography tandem mass spectrometry, we showed that proteins were subject to lysine propionylation in all five bacterial species analyzed. Notably, many propionylations were identified in the Bacillus-related, thermophilic eubacterium G. kaustophilus, but fewer in the mesophilic eubacterium B. subtilis, suggesting that propionylation event abundance is independent of phylogenetic relationship. We further found propionylation sites in the thermophilic eubacterium T. thermophilus, but the thermophilic eubacterium R. marinus showed the fewest number of sites, indicating that growth temperature is not a determinant of propionylation state. In silico analyses demonstrated that lysine propionylation is related to metabolic pathways, particularly those controlled by acyl-CoA synthetases, similar to lysine acetylation. We also detected dozens of propionylation sites at positions important for protein functions across bacteria, demonstrating the regulatory mechanisms affected by lysine propionylations. Our proteome-wide analyses across bacteria thus provide insights into the general functions of lysine propionylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Post-translational modification (PTM) is an important cellular mechanism for regulating protein function (Walsh 2006). The recent development of mass spectrometric methods has enabled us to detect and map covalent modifications of proteins on a proteome-wide scale. Previous studies have reported over 300 types of PTMs such as Ser, Thr, and Tyr phosphorylation as well as Lys ubiquitination, sumoylation, and acylation, which have expanded our understanding of PTMs (Witze et al. 2007; Doll and Burlingame 2015). In both eukaryotes and prokaryotes, PTMs control numerous proteins involved in a wide range of cellular functions such as metabolism, signaling pathways, and transcriptional regulation (Cousin et al. 2013; Kyriakis 2014; Tessarz and Kouzarides 2014; Shi and Tu 2015). Furthermore, the cross-talk between PTMs has also been shown to be important for modulating protein functions in both eukaryotes (Snider and Omary 2014; Tessarz and Kouzarides 2014) and bacteria (Soufi et al. 2012). For example, the prevalence and pattern of several types of PTMs dynamically changed in microbial communities, suggesting the physiological significance of PTMs in cell growth, microbial evolution, ecological adaptation against stresses, and environmental change (Li et al. 2014), and indicating the complex regulatory mechanisms of cellular processes by various PTMs. To further expand our knowledge of PTMs, we need to examine which types of PTMs are present in vivo and which kinds of proteins are subject to the various PTMs.
Development of mass spectrometric methods and antibody-based enrichment techniques of acylated peptides have enabled the proteome-wide identification of lysine acylation both in eukaryotes and bacteria (Kim et al. 2006). Lysine acetylation, one of the well-studied PTMs, regulates various cellular processes, particularly metabolism (Guan and Xiong 2011; Shi and Tu 2015). For example, lysine acetylation controls acetate metabolism through modification of the conserved lysine residue in the active site of acetyl-CoA synthetase in mammals (Hallows et al. 2006; Schwer et al. 2006) and bacteria (Starai et al. 2002). Other acyl-CoA synthetases are also regulated by acetylation of the corresponding lysine residues, suggesting that acetylation can regulate various acyl-CoA metabolisms (Crosby et al. 2010, 2012; Crosby and Escalante-Semerena 2014). As another example, lysine acetylation can control the metabolic flux of glycolytic/gluconeogenesis and the TCA cycle (Wang et al. 2010). Acetyl-CoA, a key metabolic intermediate, is an acetyl donor for protein acetylation by lysine acetyltransferases. It is therefore reasonable to consider that acetylation mechanisms can sense the metabolic state through the concentration of acetyl-CoA, and regulate metabolic flux accordingly (Guan and Xiong 2011). Recent proteome-wide studies on lysine acetylation have identified hundreds to thousands of acetylation sites in organisms ranging from humans to bacteria (Kim et al. 2006; Yu et al. 2008; Choudhary et al. 2009; Zhang et al. 2009a, 2013; Wang et al. 2010; Okanishi et al. 2013; Weinert et al. 2013a, b).
In addition, other types of post-translational lysine acylation have been discovered as well (Chen et al. 2007; Peng et al. 2011; Tan et al. 2011; Zhang et al. 2011; Tan et al. 2014). Similar to lysine acetylation, succinylation (Zhang et al. 2011; Park et al. 2013; Weinert et al. 2013b) in organisms from humans to bacteria, malonylation (Peng et al. 2011) and glutarylation (Tan et al. 2014) in eukaryotes, and propionylation in bacteria (Okanishi et al. 2014) were recently revealed as prevalent PTMs. These findings suggested that the regulation of metabolic flux by different types of lysine acylation is dependent on the levels of each acyl donor (Choudhary et al. 2014).
Lysine propionylation is one of the newly discovered lysine acylations, and was first reported in histones (Chen et al. 2007) and subsequently in several non-histone proteins (Cheng et al. 2009; Liu et al. 2009; Zhang et al. 2009b; Tweedie-Cullen et al. 2012; Fritz et al. 2013). In myeloid precursor leukemia cells, the level of propionylation on Lys-23 of histone H3 was shown to decrease during monocytic differentiation, which suggests that it is epigenetically regulated in a similar manner as is lysine acetylation (Liu et al. 2009). In bacteria, propionylation can regulate propionyl-CoA synthetase similar to the acetylation of acetyl-CoA synthetase as described above (Garrity et al. 2007). Most recently, we identified 361 propionylation sites in 183 mid-exponential and late stationary phase proteins in Thermus thermophilus, an extremely thermophilic eubacterium (Okanishi et al. 2014), which was the first report showing the prevalence of post-translational lysine propionylation on diverse proteins. The identified propionylation sites (Okanishi et al. 2014) were approximately as much as that of acetylation in T. thermophilus (Okanishi et al. 2013), raising the possibility that lysine propionylation rivals lysine acetylation as a major PTM. Therefore, we hypothesized that lysine propionylation regulates various cellular processes similarly to lysine acetylation. However, it remains unknown whether lysine propionylation is observed in bacteria other than T. thermophilus, whether the observed propionylations are conserved at the corresponding residues of the protein homologs, and which functions are regulated thereby.
Another point to consider is that lysine propionylation as a prevalent PTM may be peculiar to thermophiles. At present, lysine propionylation has not been emerged as a major PTM in mesophiles including Escherichia coli and Bacillus subtilis and even in other thermophiles. Lysine acetylation, another protein acylation, has already been reported for an extreme thermophile T. thermophilus (Okanishi et al. 2013), a moderate thermophile Geobacillus kaustophilus (Lee et al. 2013) and a hyperthermophile Sulfolobus solfataricus (Bell et al. 2002). Lysine methylation, another type of lysine PTM, is found to be prevalent in Sulfolobus proteins (Botting et al. 2010) and considered to have a role in their thermal stabilization (Febbraio et al. 2004). There is a possibility that other PTMs in thermophilic proteins also contribute to maintaining their thermal activity and stability. Therefore, it is important to investigate the prevalence of lysine propionylation in a wide range of organisms.
Here, we characterized lysine propionylation in five bacteria during stationary phase in complex media: E. coli W3110, B. subtilis 168, G. kaustophilus HTA426, T. thermophilus HB8, and Rhodothermus marinus. Only a few propionylation sites were identified in E. coli (a γ-proteobacterium) and B. subtilis (a Firmicutes). In contrast, in G. kaustophilus, a moderately thermophilic Bacillus-related Firmicutes (Nazina et al. 2001; Takami et al. 2004), as well as in T. thermophilus, an extremely thermophilic eubacterium, many lysine propionylation sites were detected. However, only two lysine propionylation sites were identified in R. marinus, a moderately halophilic, extremely thermophilic Bacteroidete (Sako et al. 1996; Silva et al. 2000). We also performed functional classification of propionyl proteins, comparison of propionylation sites among bacteria, and mapping of propionylation site on protein structure. This represents the first global analysis describing lysine propionylation in E. coli, B. subtilis, G. kaustophilus, and R. marinus. These results provide insights into the physiological significance of lysine propionylation in enzyme regulation.
Materials and methods
Materials and cell cultivation
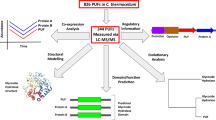
Yeast extract was purchased from Nihon Seiyaku (Tokyo, Japan), and tryptone and marine broth from Difco Laboratories (Detroit, MI, USA). Gellan gum and agar powder for solid cultivation were obtained from Wako (Tokyo, Japan) and Nacalai Tesque (Kyoto, Japan), respectively. Formic acid and acetonitrile for liquid chromatography and other chemical compounds were purchased from Wako. R. marinus (JCM9785) was obtained from the Japan Collection of Microorganisms (RIKEN BioResource Center, Saitama, Japan). The temperature of cultivation was as follows: 37 °C for B. subtilis and E. coli; 60 °C for G. kaustophilus; and 70 °C for T. thermophilus and R. marinus. Cells of G. kaustophilus, B. subtilis, and E. coli were cultivated on LB plates (1% tryptone, 0.5% yeast extract, 0.5% NaCl, and 1.5% agar powder); T. thermophilus on TT plates (0.4% tryptone, 0.2% yeast extract, 0.1% NaCl, 1.5 mM MgCl2, 1.5 mM CaCl2 and 1.5% gellan gum); and R. marinus on marine plates (37.4 g/L marine broth and 1.5% gellan gum). The types of culture media used were as follows: LB broth (1% tryptone, 0.5% yeast extract, and 0.5% NaCl) for G. kaustophilus, B. subtilis, and E. coli; TT broth (0.4% tryptone, 0.2% yeast extract, 0.1% NaCl, 1.5 mM MgCl2, and 1.5 mM CaCl2) for T. thermophilus; and marine broth (37.4 g/L marine broth) for R. marinus. Marine broth was dissolved by heating at 70 °C for 30 min and sterilized by filtration (0.2 μm, Millipore, Billerica, MA, USA). The other broths were sterilized by autoclave. Each single colony on a plate was inoculated into 5 mL of each broth with vigorous shaking. After overnight cultivation, 50 μL of culture was inoculated into 5 mL of each respective broth. When cells reached the mid-exponential growth phase (OD600 = 0.8), a 1.5 mL culture aliquot was inoculated into 150 mL of each culture media. After the cells reached the stationary phase (OD600 = 8.5 for G. kaustophilus, 8 h cultivation; OD600 = 5.5 for B. subtilis, 10 h cultivation; OD600 = 6.0 for E. coli, 12 h cultivation; OD600 = 3.5 for T. thermophilus, 8 h cultivation; and OD600 = 2.0 for R. marinus, 11 h cultivation), the cells were harvested by centrifugation at 7000 g for 5 min. The cell pellets were then washed twice by phosphate-buffered saline (PBS) (pH 7.4). To provide reproducibility, cell samples from three independent biological replicates were prepared (Fig. 1).
Workflow of proteome-wide identification of lysine propionylation in five bacteria; G. kaustophilus, T. thermophilus, B. subtilis, E. coli, and R. marinus. Tryptic peptide samples from three biological replicates of each bacterium were prepared (a total of 15 peptide samples). From each tryptic peptide sample, two propionyl peptide samples were enriched by immunoprecipitation using an anti-propionyl-lysine antibody (a total of 30 immunoprecipitates). Then, each propionyl peptide sample was analyzed twice by nano-LC–MS/MS (a total of 60 runs). Data of the 60 MS/MS runs were processed and propionylation sites were identified by Mascot database search
Preparation of tryptic peptides
Harvested cells were resuspended in cold PBS (pH 7.4) containing 5 mM EDTA and 1 mM phenylmethylsulfonyl fluoride. The cells were then disrupted using a UD-201 ultrasonic disruptor (TOMY, Tokyo, Japan) at 4 °C. The extracts were immediately precipitated with 80% acetone to minimize non-enzymatic modification. After overnight incubation of the extract at −20 °C, the pellet was collected by centrifugation at 7000 g for 20 min. Before acetone precipitation, small aliquots of the lysates were retained and used for Western blotting and determination of the protein concentration in the extracts by the Bradford assay. The acetone pellets were completely dried and then resuspended in 50 mM ammonium bicarbonate (pH 8.0) containing 6 M guanidine hydrochloride. After protein denaturation by heating at 90 °C for 10 min, the protein mixtures were diluted with 50 mM ammonium bicarbonate to a final concentration of 1.5 M guanidine hydrochloride. Because protein pellets were not completely dissolved due to the usage of the large amount of proteins, proteins were digested with trypsin prior to reduction and alkylation as previously described (Kim et al. 2006; Zhang et al. 2009a). From this, 60 mg protein was then digested with TPCK trypsin (Thermo Fisher Scientific, Waltham, MA, USA) at 1:100 (w/w) enzyme/substrate ratio at 37 °C overnight. Then the tryptic peptides were further digested with 1:100 (w/w) of TPCK trypsin at 37 °C overnight. The trypsin was subsequently inactivated by heating at 95 °C for 10 min. The tryptic peptides were alkylated with 10 mM iodoacetamide at 37 °C for 30 min after reduction with 5 mM dithiothreitol at 50 °C for 30 min. The alkylation reaction was quenched by adding dithiothreitol to a 10 mM final concentration. The peptide samples were desalted using a Sep-Pak Plus C18 cartridge (Waters, MA, USA). The eluted peptides were lyophilized using a Freezone 4.5 (Labconco, MO, USA).
Enrichment of propionyl peptides
An anti-propionyl-lysine antibody was generated as previously described (Guan et al. 2010). Briefly, bovine serum albumin (BSA) was chemically propionylated with propionic anhydride. The efficiency of propionylation of the ε-amino group of the lysine residues in the proteins was evaluated using a ninhydrin reaction, demonstrating that over 90% of the lysine residues were propionylated. Using this propionylated BSA as an antigen, the anti-propionyl-lysine antiserum was generated by Kitayama Labes Co. (Nagano, Japan).
A 50 μL aliquot of the anti-propionyl-lysine antibody was mixed with 100 μL of protein A agarose beads (Santa Cruz Biotechnology, Dallas, TX, USA) and 350 μL of PBS (pH 7.4). The antibody was conjugated to the beads by gentle rotation at 4°C for 4 h using an RT-30 mini culture rotator (TAITEC, Tokyo, Japan). The antibody-conjugated beads were washed three times with 1 mL of NETN buffer (50 mM Tris–HCl (pH 8.0) containing 100 mM NaCl, 1.0 mM EDTA, and 0.5% Nonidet P40). Concurrently, peptides digested from 5 mg proteins were dissolved in 500 μL of NETN buffer. The peptide solution was mixed with the antibody-conjugated beads and gently rotated at 4 °C for 6 h. The beads were washed with 1 mL of NETN buffer and then washed twice with 1 mL of 50 mM Tris–HCl (pH 8.0) containing 100 mM NaCl and 1.0 mM EDTA. Next, the propionyl peptides were eluted twice with 300 μL of 0.1% trifluoroacetic acid. After desalting of the eluate with a Sep-Pak Plus C18 cartridge, the immunoprecipitated propionyl peptides were lyophilized in a Freezone 4.5 for 3 h. The lyophilized samples were resuspended in 20 μL of 0.1% formic acid. Two immunoprecipitation samples were prepared for each biological replicate. A total of 6 samples for each bacterium were analyzed by mass spectrometry (MS).
Mass spectrometric analysis of propionyl peptides
Half of each propionyl peptide sample (10 μL) was injected via the autosampler of a Proxeon EASY-nLC (Bruker Daltonics, MA, USA) into an NS-MP-10 BioSphere trap column (C18, 5 μm, 120 Å, 100 μm i.d. and 20 mm length; Nanoseparations, Nieuwkoop, The Netherlands). The peptides on the column were washed with solvent A (0.1% formic acid in water) at a flow rate of 8 μL/min for 3 min. Then the peptides were separated on an NS-AC-11-C18 BioSphere analytical column (C18, 5 μm, 120 Å, 75 μm i.d., and 150 mm length; Nanoseparations) at a flow rate of 200 nL/min with the following gradient: 5% solvent B (0.1% formic acid in acetonitrile) for 5 min; 5–40% solvent B for 60 min; 40–90% solvent B for 5 min; and 90% solvent B for 40 min. The eluted peptides were ionized by electrospray ionization, typically at a capillary voltage of −1.6 kV and a drying gas temperature of 150 °C, and then introduced into a micrOTOF-Q II mass spectrometer (Bruker Daltonics). The three most abundant ions within the m/z range from 300 to 3000 were data-dependently selected and subjected to tandem mass spectrometry (MS/MS) analysis by micrOTOF control software (version 3.0, Bruker Daltonics). Mass window for precursor ion selection was set in AutoMS(n) mode as follows: 3.0 for m/z 300, 5.0 for m/z 500, 7.0 for m/z 1000, and 10.0 for m/z 2000. Charge state screening was enabled and +2 and +3 charged ions were preferentially subject to MS/MS analysis. The collision energy of the fragmentation of precursor ions by collision-induced dissociation with Ar gas was changed at 20–40 eV depending on the charge states and m/z values of the precursor ions. Each compound was analyzed by MS/MS three times, and then ions with the m/z value of the analyzed compound were excluded for MS/MS analysis for 1 min. Each propionyl peptide sample was analyzed twice by nano-liquid chromatography/quadrupole time-of-flight (nano-LC/Q-TOF) MS.
In summary, peptide samples from three biological replicates of each bacterium (G. kaustophilus, T. thermophilus, E. coli, B. subtilis, and R. marinus) were prepared (a total of 15 peptide samples). Two propionyl peptide samples were enriched from each biological replicate by immunoprecipitation using an anti-propionyl-lysine antibody (a total of 30 immunoprecipitates). Each propionyl peptide sample was analyzed twice by nano-LC–MS/MS, leading to a total of 60 runs.
Data processing and peptide identification
The MS/MS data from a total of 60 runs (2 runs for each immunoprecipitated sample, 2 immunoprecipitations for each biological sample, and 3 biological replicates for the five respective bacteria) were analyzed by DataAnalysis 4.0 SP5 software (Bruker Daltonics). The software generated peak list files including the m/z and intensity values of the precursor ions and of their product ions using the Compound-Auto MS(n) option. Only charge-assigned peaks after deconvolution and the 50 most abundant non-charge-assigned peaks above the intensity threshold of 150 from each MS/MS spectrum were exported to the peak list files. The created peaks were searched against a database of each bacterium: G. kaustophilus HTA426 database containing 3540 protein sequences (NC_006510 and NC_006509), E. coli W3110 database containing 4227 protein sequences (NC_007779), B. subtilis 168 database containing 4175 protein sequences (NC_000964), R. marinus DSM4252 database containing 2863 protein sequences (NC_013501 and NC_013502), and T. thermophilus HB8 database containing 2238 protein sequences (NC_006461, NC_006462, and NC_006463) using the Mascot search engine version 2.4 (Matrix Science, London, U.K.). The search parameters included a mass tolerance of ± 50 ppm for the precursor ions and ± 0.05 Da for the product ions, and a maximum of 2 miscleavages were allowed for trypsin. Carbamidomethylation of cysteine residues was selected as a fixed modification. Propionylation of lysine residues and oxidation of methionine residues were selected as variable modifications. Peptides with Mascot ion scores in the > 99% confidence range (p value <0.01) were considered to be identified, and false discovery rates (FDRs) were calculated (supplemental Table S3). The possibility of propionylation on C-terminal lysine residues was filtered out because these propionylated lysine residues are not cleaved by trypsin. All MS/MS spectral data and the identification results files (dataset PXD002832) were deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) and the PRIDE database (http://www.ebi.ac.uk/pride/) (Vizcaino et al. 2013, 2014). The FDR was determined by a Mascot decoy database search (supplemental Table S3). To increase confidence in the results, unreliable identified peptides were excluded by manual inspection, such as low-score peptides derived from misassignments of charge states and those with abnormal errors of product ions derived from mismatch by chance in database search. The Mascot results are presented in Supplemental Fig. S1.
In silico analysis
The identified propionyl proteins were functionally annotated based on Clusters of Orthologous Groups of Proteins (COGs; http://www.ncbi.nlm.nih.gov/COG/) (Tatusov et al. 2003) and KEGG Orthology (Kanehisa et al. 2004). The functional annotation was compensated using BLAST2GO (Conesa et al. 2005). The definitions of the proteins were updated based on the annotated functions.
The sequence logos of the normalized amino acid sequence frequencies around the propionylated lysine residues were generated using iceLogo software (Colaert et al. 2009). Ten amino acids on the N-terminal and C-terminal sides of each propionylated lysine were normalized by amino acids on both sides of all lysine residues from whole proteins in each bacterial protein sequence database described above.
Results
Identification of lysine propionylation in five bacteria
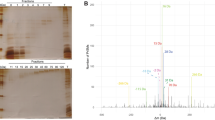
Lysine propionylation was recently shown to be abundant in T. thermophilus, an extremely thermophilic Gram-negative bacterium that belongs to the Deinococcus–Thermus phylum (Okanishi et al. 2014). To reveal how widely distributed lysine propionylation is across different bacterial taxa, we first investigated this PTM in another thermophilic bacterium. We selected G. kaustophilus, a moderately thermophilic Gram-positive Firmicutes of the Bacillaceae family, and analyzed the affinity-enriched propionyl peptides generated from stationary phase cells of G. kaustophilus and T. thermophilus by nano-LC–MS/MS. We designed the experiment as follows: three biological replicates were run for each bacterial cell which reached the mid-stationary phase in rich medium; two immunoprecipitates using an anti-propionyl-lysine antibody were prepared for each biological replicate; and two nano-LC–MS/MS runs were performed for each affinity-enriched peptide (Fig. 1). Figure 2a shows a representative MS/MS spectrum assigned to propionyl peptides from G. kaustophilus proteins. We identified propionylation sites at the 99% confidence range (p value <0.01). We used the mid-stationary-phase cells for exploring lysine propionylation in five kinds of bacteria, because our previous study showed more propionylation events in stationary phase than in exponential phase and because it was relatively easy to recognize when the bacteria entered mid-stationary phase compared to late stationary phase, and thus to harvest cells of different kinds of bacteria under a similar growth condition.
From this, we identified 83 propionylation sites from 55 G. kaustophilus proteins and 129 propionylation sites from 87 T. thermophilus proteins. Approximately 60% of the propionylation sites were found in biological triplicates (51/83 sites in G. kaustophilus and 101/129 in T. thermophilus), and ~80% of the propionylation sites (65/83 sites in G. kaustophilus and 80/129 in T. thermophilus) were identified at least twice (Fig. 2b and supplemental Table S1 and S2). In addition, we validated the specificity of the anti-propionyl-lysine antibody with a competition assay using propionylated BSA in Western blotting (supplemental Fig. S2). These results show that propionylation is abundant not only in T. thermophilus but also in G. kaustophilus cells (supplemental Table S1 and Fig. 2).
To reveal whether propionylation occurs in a broader range of bacteria, we examined lysine propionylation in two other bacteria: E. coli, a mesophilic Proteobacteria and B. subtilis, a mesophilic Firmicutes. We identified 10 propionylation sites from 9 E. coli proteins and 7 propionylation sites from 4 B. subtilis proteins (Fig. 2b and supplemental Table S1). We identified 6 of 7 propionylation sites in B. subtilis proteins and 8 of 10 propionylation sites in E. coli proteins in biological triplicates (Fig. 2b). Notably, although G. kaustophilus is a Bacillus-related eubacterium (Nazina et al. 2001; Takami et al. 2004), propionylation was detected approximately 12-fold more often in G. kaustophilus than in B. subtilis. It should be noted that more propionylation sites were found in the thermophilic bacteria (T. thermophilus and G. kaustophilus) than in the mesophilic bacteria (E. coli and B. subtilis). We therefore hypothesized that thermophilic bacteria exhibit many propionylation events. It is reasonable that high growth temperature for thermophile increases the rate of chemical reaction and thus leads to many propionylation events in thermophilic bacteria.
Next, we further investigated propionylation in R. marinus, an extremely thermophilic Bacteroidetes. As a result, we identified two propionylation sites from two proteins in R. marinus (Fig. 2b and supplemental Table S1). We identified one propionylation site twice and one site once (Fig. 2b). This is the smallest number of propionylation events identified among the five examined organisms. These results show that propionylation can occur in a wide range of bacteria. These suggest that the phylogenetic distance is not a determining factor of propionylation state and that the high growth temperature for thermophile is not mainly responsible for many propionylation events. In summary, the percentage of propionylated proteins in the respective proteomes varied among the five bacteria tested. It was higher in G. kaustophilus (55 of 3540 proteins, 1.6%) and T. thermophilus (87 of 2173 proteins, 4.0%) compared to E. coli (9 of 4226 proteins, 0.2%), B. subtilis (4 of 4175 proteins, 0.1%), and R. marinus (2 of 2863 proteins, 0.07%).
We investigated the dependence of the propionylation state on growth condition by comparing the profiles of the propionylation sites in T. thermophilus identified at different growth conditions. Propionylations were found at 129 sites on the mid-stationary phase proteins in this study, in comparison to the 121 and 323 sites identified on mid-exponential and late stationary phase proteins, respectively, in our previous study (Okanishi et al. 2014). Among these, 26 of 121 mid-exponential (21.5%), 26 of 129 mid-stationary (20.1%), and 200 of 323 late stationary (61.9%) phase-specific propionylation sites were observed, whereas 51 propionylation sites were detected in all three growth phases (supplemental Fig. S3 and Table S6). Furthermore, 14 of 26 mid-exponential (53.8%), 18 of 26 mid-stationary (69.2%), and 129 of 200 late stationary (64.5%) phase-specific sites were reproducibly identified (supplemental Table S6), demonstrating the confidence of the identifications. These results indicated that lysine propionylation can be regulated depending on the growth conditions.
Comparison of lysine propionylation sites among bacteria
To evaluate the general functions of propionylation, we compared the propionylation sites among the five bacteria tested. We found that several propionyl proteins were commonly found throughout these bacteria. In G. kaustophilus, its propionyl proteins contained 3 B. subtilis, 6 E. coli, 2 R. marinus, and 18 T. thermophilus homologs that were subject to propionylation in each organism (supplemental Table S4). We also found that nine propionylation sites identified in G. kaustophilus were propionylated at the corresponding sites in the homologs of three other bacteria; five sites in conjunction with B. subtilis, one with E. coli, and four with T. thermophilus (supplemental Table S4), suggesting that propionylation at these sites might function in a similar manner.
Among these nine propionylation sites, propionylation was detected at the lysine residue in the active site of acyl-CoA synthetases. Previous studies have reported that acetylation at this lysine residue inhibits the enzymatic activity of several types of acyl-CoA synthetases (Starai et al. 2002; Hallows et al. 2006; Schwer et al. 2006; Crosby et al. 2010, 2012; Crosby and Escalante-Semerena 2014). Notably, the critical lysine residues of four acyl-CoA synthetases in G. kaustophilus (Lys-526 of GK1598, Lys-542 of GK2136, Lys-535 of GK2759, and Lys-548 of GK2806), and one acyl-CoA synthetase in B. subtilis (Lys-549 of BSU29680) were shown to be propionylated in this study (supplemental Table S1 and S4). Our previous report showed that the corresponding lysine residue of T. thermophilus acyl-CoA synthetase was also propionylated (Okanishi et al. 2014). These results suggested that lysine propionylation regulates acyl-CoA synthetase in bacteria in a similar manner as does lysine acetylation.
Comparison of propionyl protein functions among the tested bacteria
Previous reports have shown that the majority of identified acyl-proteins represented metabolic enzymes and translation-related proteins (Kim et al. 2006; Yu et al. 2008; Zhang et al. 2009a, 2011, 2013; Wang et al. 2010; Okanishi et al. 2013; Okanishi et al. 2014; Kosono et al. 2015). To elucidate which functions are affected by lysine propionylation in bacteria, we preformed functional classification of the propionyl proteins identified in this study (Fig. 3a and supplemental Fig. S4A). In G. kaustophilus and T. thermophilus, 69.1% (38/55 proteins) and 67% (58/87 proteins) of the propionyl proteins were metabolic enzymes, respectively (Fig. 3a and supplemental Fig. S4A). The further classification of metabolic enzymes revealed a slight difference therein: the majority of metabolic proteins in G. kaustophilus were related to energy production (39.5%, 15/38) and the metabolism of lipids (15.8%, 6/38), carbohydrates (13.1%, 5/38), and amino acids (10.5%, 4/38), whereas those in T. thermophilus were related to energy production (34.5%, 20/58) and the metabolism of amino acids (15.5%, 9/58), coenzymes (12.1%, 7/58), and lipids (10.3%, 6/58) (Fig. 3b and Supplemental Fig. S4B). In the other three bacteria, 2 of 4 B. subtilis, 7 of 9 E. coli, and all two R. marinus proteins were related to metabolism (supplemental Table S5). In addition, 14.5% (8/55) of the proteins identified among G. kaustophilus and 12.6% in T. thermophilus propionyl proteins were related to translation (Fig. 3a and supplemental Fig. S4A). These results indicated that proteins subject to lysine propionylation show a similar trend with respect to their functional categories as do other types of lysine acylation.
Functional classification of the identified propionyl proteins and the amino acid sequence context around propionylation sites in G. kaustophilus. a Distributions of the functional classes of 55 propionyl proteins, based on COGs (Tatusov et al. 2003) and KEGG Orthology (Kanehisa et al. 2004). b Distributions of the functional subclasses of 38 propionyl proteins in the “metabolism” class. c The normalized frequencies of amino acid residues around the propionylated lysines, which were prepared by iceLogo software (Colaert et al. 2009) (p value = 0.05)
General characterization of the amino acid sequences around identified propionylation sites
In both eukaryotes and bacteria, lysine acylation including propionylation has been reported to be reversibly controlled by two types of enzymes: lysine acyltransferases (KATs) and lysine deacylases (KDACs) (Lin et al. 2012). The sequence specificity of these enzymes might be reflected in the frequencies of amino acid occurrences around the modification sites. We therefore generated the plot of the amino acid sequence frequencies from the −10 to +10 positions around the propionylation sites identified in the two propionylation-rich bacteria: G. kaustophilus and T. thermophilus (Fig. 3c and Supplemental Fig. S4C). Notably, Asp and Glu appeared in the +1 position in the two bacteria. Acidic residues were also abundant in the +3 and +7 positions, and basic residues such as Lys were broadly preferred in the −10 to −1 positions in both bacteria. However, marked exclusion of hydrophobic residues, primarily Leu at the +1 position and a preference for Glu at the −1 position was observed only in T. thermophilus. These characteristics of the amino acid sequence around propionylated lysines in T. thermophilus at mid-stationary phase are similar to those observed at both mid-log and late stationary phases in this organism (Okanishi et al. 2014). G. kaustophilus and T. thermophilus might therefore have both common and distinct types of enzymes catalyzing propionylation.
Discussion
Here, we identified propionylation in five representative bacteria with high confidence (Fig. 2), showing that all five bacteria contain propionylated proteins. Notably, a large number of propionylation sites were found not only in T. thermophilus, as previously reported (Okanishi et al. 2014), but also in another thermophilic bacterium, G. kaustophilus (Fig. 2 and Supplemental Table S1). Among mesophilic bacteria, E. coli and B. subtilis, propionylations were also observed but the numbers were less than seen in G. kaustophilus and T. thermophilus. G. kaustophilus is a thermophilic Bacillus-related bacterium. Although G. kaustophilus is closely related to B. subtilis phylogenetically, their propionylation states were distinct. This difference suggests that the degree of phylogenetic relatedness is not critical for lysine propionylation. We therefore deduced that the propionylation state depends on factors within the intra- and/or extracellular environments, such as growth temperature, the concentration of metabolites including propionyl-CoA, and expression levels of mRNA and protein related in propionylation (Garrity et al. 2007; Seidel et al. 2016). Although propionylation-rich bacteria, G. kaustophilus and T. thermophilus, are thermophiles, we found the lowest number of propionylation sites in another thermophilic bacterium, R. marinus (Fig. 2b and Supplemental Table S1), suggesting that high growth temperature optimal for thermophile itself is not mainly responsible for many propionylation events. In addition, a previous study showed that the profiles of lysine acetylation, another type of post-translational acylation, dramatically changed depending on cultivation conditions (Wang et al. 2010; Weinert et al. 2013a; Kosono et al. 2015; Schilling et al. 2015; Mizuno et al. 2016); therefore, lysine propionylation might also be dramatically altered by cultivation conditions. Indeed, the number of lysine propionylation were detected ~2.5-fold more often at late stationary phase (323 sites) (Okanishi et al. 2014) than mid-stationary phase (129 sites; this study). Acetylation levels are regulated by acetyl-CoA levels (Choudhary et al. 2014). Interestingly, a previous study showed that concentration of propionyl-CoA is lower than that of acetyl-CoA in E. coli grown in LB broth (Febbraio et al. 2004). In E. coli cultivated in LB broth, the number of propionylation events in this study was smaller than that of acetylation events previously reported (Yu et al. 2008; Zhang et al. 2009a). This suggests that propionylation levels are regulated by propionyl-CoA concentration similarly to acetylation. Also, the intracellular supply of propionyl-CoA from metabolic pathways such as β-oxidation of fatty acids might be related to the finding that lysine propionylation occurs more frequently in G. kaustophilus and T. thermophilus than the other three bacteria.
Additionally, we found that certain propionylation sites identified in one organism were propionylated at the corresponding sites in the homologs of other examined bacteria, which implied the existence of general functions of lysine propionylation throughout bacterial taxa (supplemental Table S4). We further investigated the propionylation sites identified in our previous report, which presented the prevalent lysine propionylation sites in T. thermophilus (Okanishi et al. 2014). From this analysis, we found that an additional eight G. kaustophilus propionylation sites were also propionylated in T. thermophilus (supplemental Table S4). Overall, a total of 13 propionylation sites in G. kaustophilus were shown to be propionylated in the other bacteria. Notably, 10 of these 13 overlapping sites were represented lysine residues of metabolic enzymes. Four acyl-CoA synthetases in G. kaustophilus were propionylated at highly conserved lysine residues, and each homolog was also propionylated at the corresponding residues in B. subtilis and T. thermophilus. Acyl-CoA synthetases control various metabolic pathways by the conversion of acetate to acetyl-CoA in all three kingdoms. Recent studies have revealed that various types of acyl-CoA synthetases were inactivated by acetylation at a conserved lysine residue through disruption of the hydrogen bond between the lysine residue and the substrate (Starai et al. 2002; Hallows et al. 2006; Schwer et al. 2006; Crosby et al. 2010, 2012; Crosby and Escalante-Semerena 2014). Propionylation at Lys-592 of propionyl-CoA synthetase in Salmonella enterica is also known to inactivate its enzymatic activity (Garrity et al. 2007). In this study, we identified five propionylation sites at conserved lysine residues of acyl-CoA synthetases (supplemental Table S1 and S4). We also reported that the corresponding lysine residue of T. thermophilus acyl-CoA synthetase was propionylated (Okanishi et al. 2014). These suggest that lysine propionylation also regulates different types of acyl-CoA synthetases in other bacteria and controls various metabolic pathways in the same manner as does lysine acetylation.
Furthermore, we found propionylation at a conserved lysine residue of l-alanine dehydrogenase in G. kaustophilus and T. thermophilus (Lys-75 of GK3448 and Lys-73 of TTHA0216). This conserved residue forms a hydrogen bond with its substrate, pyruvate, which can polarize and activate the substrate (Fig. 4) (Baker et al. 1998). Propionylation can disrupt the hydrogen bonding network, and therefore block the function of this enzyme in these bacteria. Together, these findings reveal some of the general functions of lysine propionylation that are maintained throughout bacteria.
Representative structure of propionylation site found at functionally important lysine residue. a The overall tertiary structure of Phormidium lapideum alanine dehydrogenase (PDB ID, 1SAY), which is the homolog of the identified propionyl proteins, GK3448 of G. kaustophilus and TTHA0216 of T. thermophilus (Blast E value is 3 × 10−118 and 4 × 10−133, respectively). In this study, propionylation was identified at a conserved lysine residue, Lys-75 of GK3448 and Lys-73 of TTHA0216. The conserved lysine residue forms a hydrogen bond with its substrate which can polarize and activate the substrate (Baker et al. 1998). b An extended view of the red square of (a). The conserved lysine residue and its binding substrate are shown as sticks. Blue dashed line indicates hydrogen bonding
To further understand the functions of lysine propionylation, we investigated the sites identified in proteins other than in the above-mentioned proteins. One of the propionylation sites identified in this study, Lys-242 of isocitrate dehydrogenase (NADP+) (Y75_p1106) of E. coli, has been reported to be succinylated, and it was shown that mutation of this residue (K242E) led to a decrease of enzymatic activity (Zhang et al. 2011). This suggests that competition between propionylation and succinylation at the same lysine residue might serve as a novel mechanism to regulate enzyme activity.
Furthermore, we investigated whether the identified propionylation sites are located at or near positions important for protein function (Table 1 and supplemental Fig. S5). We obtained structural data of authentic and homologous proteins from the Research Collaboratory for Structural Bioinformatics Protein Data Bank (PDB, http://www.rcsb.org/pdb) and mapped all the identified propionylation sites onto the protein structures. This analysis is cumbersome, but can provide invaluable information with regard to the role of each propionylation event. We used the structural data of homologs only for lysine residues conserved at the position corresponding to the propionylation site in the homolog. Supplemental figure S5 shows protein tertiary structures with 29 propionylation sites, which might play important roles in protein function; 14 sites in G. kaustophilus, 1 site in B. subtilis, 1 site in E. coli, 1 site in R. marinus, and 12 sites in T. thermophilus. Among these, 20 sites are at or near ligand-binding sites. The representative sites in G. kaustophilus are as follows: Lys-87 of ferrochelatase (GK0662) at 3.7 Å from porphyrin (substrate); Lys-374 of glycerol-3-phosphate dehydrogenase (GK2153) at 3.9 Å from dihydroxyacetone phosphate (product); Lys-132 of inorganic pyrophosphatase (GK2246) at 2.8 Å from pyrophosphate (substrate); Lys 289 of electron transfer flavoprotein α subunit (GK2686) at 3.8 Å from FAD (cofactor); Lys215 of triose-phosphate isomerase (GK3056) at 5.0 Å from 2-phosphoglycolate (competitive inhibitor); and Lys-86 of acetyl-CoA acetyltransferase (GK3397) at 4.2 Å from the chloride ion stabilizing enzymatic active loop. In T. thermophilus, the following sites are representative: Lys55 of nucleoside diphosphate kinase (TTHA0188) at 4.2 Å from ATP (substrate); Lys-50 of GroEL chaperone (TTHA0271) at 2.4 Å from ADP; Lys-58 of 3-hydroxybutyryl-CoA dehydrogenase (TTHA1262) at 3.2 Å from acetoacetyl-CoA (substrate); Lys-81 of 3-hydroxybutyryl-CoA dehydratase (TTHA1434) at 2.7 Å from acetoacetyl-CoA (competitive inhibitor); and Lys-190 of isocitrate lyase (TTHA1836) at 4.6 Å from succinic acid (product). Interestingly, we found propionylations at or near ligand-binding sites even in the bacteria with less propionylation: Lys-549 of B. subtilis acetyl-CoA synthetase (BSU29680) at 3.1 Å from ATP (substrate); Lys-143 of E. coli elongation factor G (Y75_p3836) at 3.2 Å from GTP (substrate); and Lys-273 of R. marinus succinate-semialdehyde dehydrogenase (Rmar_2008) at 4.8 Å from NADP (substrate). In addition, we found four propionylation sites near nucleic acids bound to proteins. In G. kaustophilus, Lys-9 of the transition state regulator abh (GK0030) at 2.3 Å from the phosphate group of double-stranded DNA and Lys-74 of the 50S ribosomal protein L24 (GK0117) at 3.2 Å from the phosphate group of 23S rRNA were propionylated. In T. thermophilus, Lys-190 of the single-stranded DNA-binding protein (TTHA0244) at 2.2 Å from the phosphate group of single-stranded DNA and Lys-42 of elongation factor P (TTHA1125) at 2.5 Å from the phosphate group of 16S rRNA were propionylated. These findings suggest the significance of lysine propionylation in the regulation of protein functions in bacteria.
Our propionyl-proteome analysis on five different bacteria showed 29 propionylation sites located at tertiary positions important for protein function (Table 1 and supplemental Fig. S5). Among them, 19 propionylation sites (65.5% of 29 sites) were located near ligands (substrates, products, cofactors, and their analogs) in the active sites (Table 1 and supplemental Fig. S5). This suggests that lysine propionylation regulates enzymatic activities in the broad range of physiological processes. However, it remains unknown whether such functional role is specific to lysine propionylation or common among lysine acylations. In our recent study, we investigated the tertiary positions of succinylation, which were identified in the same five bacterial species (unpublished data). A total of 44 succinylation sites were located at positions important for protein functions. Among them, only 8 succinylation sites (18.2% of 44 sites) were located near ligands in the active sites. Lysine propionylation and succinylation have distinct chemical properties because propionylation neutralizes positively charged lysine residue and succinylation changes charge states of lysine from +1 to −1. Our findings suggest that the two types of acylation with distinct chemical characteristics have different functions in the cells, and especially propionylation plays roles in enzyme regulation.
We also analyzed the amino acid frequencies around the propionylated lysine residues of G. kaustophilus and T. thermophilus (Fig. 3c and supplemental Fig. S4C). As a result, we identified several common characteristics as well as some differences. A previous study revealed that propionyl-CoA synthetase of S. enterica is propionylated at Lys-592 by two types of KATs, S. enterica Pat and B. subtilis AcuA, and depropionylated by a KDAC, S. enterica CobB (Garrity et al. 2007). Bacteria have several types of KATs such as Pat and AcuA as well as KDACs such as AcuC and SrtN. The G. kaustophilus genome has only a single KAT, AcuA (GK2807), and three kinds of KDACs, AcuC (GK2809) and two SrtNs (GK1653 and GK1437), whereas the T. thermophilus genome also has only a single KAT, Pat (TTHA0276) but two kinds of KDACs, AcuC (TTHA0475) and SrtN (TTHA1392). The sets of KDACs in the two bacteria are similar, but the KATs are not. Thus, there is the possibility that the similarities and differences in amino acid frequencies reflect the respective sets of encoded KATs and KDACs between bacteria.
Conclusions
In summary, we investigated lysine propionylation among five representative bacteria on a proteome-wide scale. Lysine propionylation was found to occur broadly and not only in certain bacteria, and the propionylation state was not dependent on phylogenetic lineage but was dependent on intra- and/or extracellular environmental conditions except for growth temperature. In silico analysis suggested a link between lysine propionylation and metabolism similar to that observed for lysine acetylation and provided new clues to the possible regulatory mechanisms exerted by the dozens of identified propionylation sites. This study established a foundation for further understanding of the molecular and cellular biology of lysine propionylation. It remains unknown how KATs and KDACs regulate propionylation state. It is also important to reveal crosstalks between lysine propionylation and other PTMs. Our investigation may help future studies to address these issues and reveal the roles of lysine propionylation.
Abbreviations
- PTM:
-
Post-translational modification
- PBS:
-
Phosphate-buffered saline
- BSA:
-
Bovine serum albumin
- MS:
-
Mass spectrometry
- MS/MS:
-
Tandem mass spectrometry
- nano-LC:
-
Nano-liquid chromatography
- FDR:
-
False discovery rate
- Q-TOF:
-
Quadrupole time-of-flight
- COGs:
-
Clusters of Orthologous Groups
- KAT:
-
Lysine acyltransferase
- KDAC:
-
Lysine deacylase
- PDB:
-
Protein data bank
References
Baker PJ, Sawa Y, Shibata H, Sedelnikova SE, Rice DW (1998) Analysis of the structure and substrate binding of Phormidium lapideum alanine dehydrogenase. Nat Struct Biol 5:561–567
Bell SD, Botting CH, Wardleworth BN, Jackson SP, White MF (2002) The interaction of Alba, a conserved archaeal chromatin protein, with Sir2 and its regulation by acetylation. Science 296:148–151
Botting CH, Talbot P, Paytubi S, White MF (2010) Extensive lysine methylation in hyperthermophilic crenarchaea: potential implications for protein stability and recombinant enzymes. Archaea 2010:106341
Chen Y, Sprung R, Tang Y, Ball H, Sangras B, Kim SC, Falck JR, Peng J, Gu W, Zhao Y (2007) Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol Cell Proteomics 6:812–819
Cheng Z, Tang Y, Chen Y, Kim S, Liu H, Li SS, Gu W, Zhao Y (2009) Molecular characterization of propionyllysines in non-histone proteins. Mol Cell Proteomics 8:45–52
Choudhary C, Kumar C, Gnad F, Nielsen ML, Rehman M, Walther TC, Olsen JV, Mann M (2009) Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325:834–840
Choudhary C, Weinert BT, Nishida Y, Verdin E, Mann M (2014) The growing landscape of lysine acetylation links metabolism and cell signalling. Nat Rev Mol Cell Biol 15:536–550
Colaert N, Helsens K, Martens L, Vandekerckhove J, Gevaert K (2009) Improved visualization of protein consensus sequences by iceLogo. Nat Methods 6:786–787
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Cousin C, Derouiche A, Shi L, Pagot Y, Poncet S, Mijakovic I (2013) Protein-serine/threonine/tyrosine kinases in bacterial signaling and regulation. FEMS Microbiol Lett 346:11–19
Crosby HA, Escalante-Semerena JC (2014) The acetylation motif in AMP-forming Acyl coenzyme A synthetases contains residues critical for acetylation and recognition by the protein acetyltransferase pat of Rhodopseudomonas palustris. J Bacteriol 196:1496–1504
Crosby HA, Heiniger EK, Harwood CS, Escalante-Semerena JC (2010) Reversible Nε-lysine acetylation regulates the activity of acyl-CoA synthetases involved in anaerobic benzoate catabolism in Rhodopseudomonas palustris. Mol Microbiol 76:874–888
Crosby HA, Pelletier DA, Hurst GB, Escalante-Semerena JC (2012) System-wide studies of N-lysine acetylation in Rhodopseudomonas palustris reveal substrate specificity of protein acetyltransferases. J Biol Chem 287:15590–15601
Doll S, Burlingame AL (2015) Mass spectrometry-based detection and assignment of protein posttranslational modifications. ACS Chem Biol 10:63–71
Febbraio F, Andolfo A, Tanfani F, Briante R, Gentile F, Formisano S, Vaccaro C, Scire A, Bertoli E, Pucci P, Nucci R (2004) Thermal stability and aggregation of Sulfolobus solfataricus β-glycosidase are dependent upon the N-ε-methylation of specific lysyl residues: critical role of in vivo post-translational modifications. J Biol Chem 279:10185–10194
Fritz KS, Green MF, Petersen DR, Hirschey MD (2013) Ethanol metabolism modifies hepatic protein acylation in mice. PLoS One 8:e75868
Garrity J, Gardner JG, Hawse W, Wolberger C, Escalante-Semerena JC (2007) N-lysine propionylation controls the activity of propionyl-CoA synthetase. J Biol Chem 282:30239–30245
Guan KL, Xiong Y (2011) Regulation of intermediary metabolism by protein acetylation. Trends Biochem Sci 36:108–116
Guan KL, Yu W, Lin Y, Xiong Y, Zhao S (2010) Generation of acetyllysine antibodies and affinity enrichment of acetylated peptides. Nat Protoc 5:1583–1595
Hallows WC, Lee S, Denu JM (2006) Sirtuins deacetylate and activate mammalian acetyl-CoA synthetases. Proc Natl Acad Sci USA 103:10230–10235
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Kim SC, Sprung R, Chen Y, Xu Y, Ball H, Pei J, Cheng T, Kho Y, Xiao H, Xiao L, Grishin NV, White M, Yang XJ, Zhao Y (2006) Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell 23:607–618
Kosono S, Tamura M, Suzuki S, Kawamura Y, Yoshida A, Nishiyama M, Yoshida M (2015) Changes in the acetylome and succinylome of Bacillus subtilis in response to carbon source. PLoS One 10:e0131169
Kyriakis JM (2014) In the beginning, there was protein phosphorylation. J Biol Chem 289:9460–9462
Lee DW, Kim D, Lee YJ, Kim JA, Choi JY, Kang S, Pan JG (2013) Proteomic analysis of acetylation in thermophilic Geobacillus kaustophilus. Proteomics 13:2278–2282
Li Z, Wang Y, Yao Q, Justice NB, Ahn TH, Xu D, Hettich RL, Banfield JF, Pan C (2014) Diverse and divergent protein post-translational modifications in two growth stages of a natural microbial community. Nat Commun 5:4405
Lin H, Su X, He B (2012) Protein lysine acylation and cysteine succination by intermediates of energy metabolism. ACS Chem Biol 7:947–960
Liu B, Lin Y, Darwanto A, Song X, Xu G, Zhang K (2009) Identification and characterization of propionylation at histone H3 lysine 23 in mammalian cells. J Biol Chem 284:32288–32295
Mizuno Y, Nagano-Shoji M, Kubo S, Kawamura Y, Yoshida A, Kawasaki H, Nishiyama M, Yoshida M, Kosono S (2016) Altered acetylation and succinylation profiles in Corynebacterium glutamicum in response to conditions inducing glutamate overproduction. Microbiologyopen 5:152–173
Nazina TN, Tourova TP, Poltaraus AB, Novikova EV, Grigoryan AA, Ivanova AE, Lysenko AM, Petrunyaka VV, Osipov GA, Belyaev SS, Ivanov MV (2001) Taxonomic study of aerobic thermophilic bacilli: descriptions of Geobacillus subterraneus gen. nov., sp. nov. and Geobacillus uzenensis sp. nov. from petroleum reservoirs and transfer of Bacillus stearothermophilus, Bacillus thermocatenulatus, Bacillus thermoleovorans, Bacillus kaustophilus, Bacillus thermodenitrificans to Geobacillus as the new combinations G. stearothermophilus, G. thermoglucosidasius and G. thermodenitrificans. Int J Syst Evol Microbiol 51:433–446
Okanishi H, Kim K, Masui R, Kuramitsu S (2013) Acetylome with structural mapping reveals the significance of lysine acetylation in Thermus thermophilus. J Proteome Res 12:3952–3968
Okanishi H, Kim K, Masui R, Kuramitsu S (2014) Lysine propionylation is a prevalent post-translational modification in Thermus thermophilus. Mol Cell Proteomics 13:2382–2398
Park J, Chen Y, Tishkoff DX, Peng C, Tan M, Dai L, Xie Z, Zhang Y, Zwaans BM, Skinner ME, Lombard DB, Zhao Y (2013) SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol Cell 50:919–930
Peng C, Lu Z, Xie Z, Cheng Z, Chen Y, Tan M, Luo H, Zhang Y, He W, Yang K, Zwaans BM, Tishkoff D, Ho L, Lombard D, He TC, Dai J, Verdin E, Ye Y, Zhao Y (2011) The first identification of lysine malonylation substrates and its regulatory enzyme. Mol Cell Proteomics 10(M111):012658
Sako Y, Takai K, Ishida Y, Uchida A, Katayama Y (1996) Rhodothermus obamensis sp. nov., a modern lineage of extremely thermophilic marine bacteria. Int J Syst Bacteriol 46:1099–1104
Schilling B, Christensen D, Davis R, Sahu AK, Hu LI, Walker-Peddakotla A, Sorensen DJ, Zemaitaitis B, Gibson BW, Wolfe AJ (2015) Protein acetylation dynamics in response to carbon overflow in Escherichia coli. Mol Microbiol 98:847–863
Schwer B, Bunkenborg J, Verdin RO, Andersen JS, Verdin E (2006) Reversible lysine acetylation controls the activity of the mitochondrial enzyme acetyl-CoA synthetase 2. Proc Natl Acad Sci USA 103:10224–10229
Seidel J, Klockenbusch C, Schwarzer D (2016) Investigating feformylase and deacylase activity of mammalian and bacterial sirtuins. ChemBioChem 17:398–402
Shi L, Tu BP (2015) Acetyl-CoA and the regulation of metabolism: mechanisms and consequences. Curr Opin Cell Biol 33:125–131
Silva Z, Horta C, da Costa MS, Chung AP, Rainey FA et al (2000) Polyphasic evidence for the reclassification of Rhodothermus obamensis Sako et al. 1996 as a member of the species Rhodothermus marinus Alfredsson et al. 1988. Int J Syst Evol Microbiol 50:1457–1461
Snider NT, Omary MB (2014) Post-translational modifications of intermediate filament proteins: mechanisms and functions. Nat Rev Mol Cell Biol 15:163–177
Soufi B, Soares NC, Ravikumar V, Macek B (2012) Proteomics reveals evidence of cross-talk between protein modifications in bacteria: focus on acetylation and phosphorylation. Curr Opin Microbiol 15:357–363
Starai VJ, Celic I, Cole RN, Boeke JD, Escalante-Semerena JC (2002) Sir2-dependent activation of acetyl-CoA synthetase by deacetylation of active lysine. Science 298:2390–2392
Takami H, Takaki Y, Chee GJ, Nishi S, Shimamura S, Suzuki H, Matsui S, Uchiyama I (2004) Thermoadaptation trait revealed by the genome sequence of thermophilic Geobacillus kaustophilus. Nucleic Acids Res 32:6292–6303
Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E, Buchou T, Cheng Z, Rousseaux S, Rajagopal N, Lu Z, Ye Z, Zhu Q, Wysocka J, Ye Y, Khochbin S, Ren B, Zhao Y (2011) Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146:1016–1028
Tan M, Peng C, Anderson KA, Chhoy P, Xie Z, Dai L, Park J, Chen Y, Huang H, Zhang Y, Ro J, Wagner GR, Green MF, Madsen AS, Schmiesing J, Peterson BS, Xu G, Ilkayeva OR, Muehlbauer MJ, Braulke T, Muhlhausen C, Backos DS, Olsen CA, McGuire PJ, Pletcher SD, Lombard DB, Hirschey MD, Zhao Y (2014) Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab 19:605–617
Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV, Krylov DM, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, Smirnov S, Sverdlov AV, Vasudevan S, Wolf YI, Yin JJ, Natale DA (2003) The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4:41
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708
Tweedie-Cullen RY, Brunner AM, Grossmann J, Mohanna S, Sichau D, Nanni P, Panse C, Mansuy IM (2012) Identification of combinatorial patterns of post-translational modifications on individual histones in the mouse brain. PLoS One 7:e36980
Vizcaino JA, Cote RG, Csordas A, Dianes JA, Fabregat A, Foster JM, Griss J, Alpi E, Birim M, Contell J, O’Kelly G, Schoenegger A, Ovelleiro D, Perez-Riverol Y, Reisinger F, Rios D, Wang R, Hermjakob H (2013) The Proteomics Identifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res 41:D1063–D1069
Vizcaino JA, Deutsch EW, Wang R, Csordas A, Reisinger F, Rios D, Dianes JA, Sun Z, Farrah T, Bandeira N, Binz PA, Xenarios I, Eisenacher M, Mayer G, Gatto L, Campos A, Chalkley RJ, Kraus HJ, Albar JP, Martinez-Bartolome S, Apweiler R, Omenn GS, Martens L, Jones AR, Hermjakob H (2014) ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat Biotechnol 32:223–226
Walsh CT (2006) Posttranslational modification of proteins: Expanding nature’s inventory. Roberts and Co. Publishers, Greenwood Village
Wang Q, Zhang Y, Yang C, Xiong H, Lin Y, Yao J, Li H, Xie L, Zhao W, Yao Y, Ning ZB, Zeng R, Xiong Y, Guan KL, Zhao S, Zhao GP (2010) Acetylation of metabolic enzymes coordinates carbon source utilization and metabolic flux. Science 327:1004–1007
Weinert BT, Iesmantavicius V, Wagner SA, Scholz C, Gummesson B, Beli P, Nystrom T, Choudhary C (2013a) Acetyl-phosphate is a critical determinant of lysine acetylation in E. coli. Mol Cell 51:265–272
Weinert BT, Scholz C, Wagner SA, Iesmantavicius V, Su D, Daniel JA, Choudhary C (2013b) Lysine succinylation is a frequently occurring modification in prokaryotes and eukaryotes and extensively overlaps with acetylation. Cell Rep 4:842–851
Witze ES, Old WM, Resing KA, Ahn NG (2007) Mapping protein post-translational modifications with mass spectrometry. Nat Methods 4:798–806
Yu BJ, Kim JA, Moon JH, Ryu SE, Pan JG (2008) The diversity of lysine-acetylated proteins in Escherichia coli. J Microbiol Biotechnol 18:1529–1536
Zhang J, Sprung R, Pei J, Tan X, Kim S, Zhu H, Liu CF, Grishin NV, Zhao Y (2009a) Lysine acetylation is a highly abundant and evolutionarily conserved modification in Escherichia coli. Mol Cell Proteomics 8:215–225
Zhang K, Chen Y, Zhang Z, Zhao Y (2009b) Identification and verification of lysine propionylation and butyrylation in yeast core histones using PTMap software. J Proteome Res 8:900–906
Zhang Z, Tan M, Xie Z, Dai L, Chen Y, Zhao Y (2011) Identification of lysine succinylation as a new post-translational modification. Nat Chem Biol 7:58–63
Zhang K, Zheng S, Yang JS, Chen Y, Cheng Z (2013) Comprehensive profiling of protein lysine acetylation in Escherichia coli. J Proteome Res 12:844–851
Acknowledgements
We thank Dr. Yuichi Otsuka (Osaka University) for giving us E. coli W3110 strain. The mass spectrometry proteomics data have been deposited with the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) (Vizcaino et al. 2014) via the PRIDE partner repository (Vizcaino et al. 2013) with the dataset identifier PXD002832. This research was supported in part by the Ministry of Education, Science, Sports and Culture, Grant-in-Aid for Challenging Exploratory Research 25650008 (to RM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no conflict of interest.
Additional information
Communicated by A. Oren.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Okanishi, H., Kim, K., Masui, R. et al. Proteome-wide identification of lysine propionylation in thermophilic and mesophilic bacteria: Geobacillus kaustophilus, Thermus thermophilus, Escherichia coli, Bacillus subtilis, and Rhodothermus marinus . Extremophiles 21, 283–296 (2017). https://doi.org/10.1007/s00792-016-0901-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-016-0901-3