Abstract
Chitin is the second most widely found natural polymer next to cellulose. Chitinases degrade the insoluble chitin to bioactive chitooligomers and monomers for various industrial applications. Based on their function, these enzymes act as biocontrol agents against pathogenic fungi and invasive pests compared with conventional chemical fungicides and insecticides. They have other functional roles in shellfish waste management, fungal protoplast generation, and Single-Cell Protein production. Among the chitinases, thermophilic and thermostable chitinases are gaining popularity in recent years, as they can withstand high temperatures and maintain the enzyme stability for longer periods. Not all chitinases are thermostable; hence, tailor-made thermophilic chitinases are designed to enhance their thermostability by direct evolution, genetic engineering involving mutagenesis, and proteomics approach. Although research has been done extensively on cloning and expression of thermophilic chitinase genes, there are only few papers discussing on the mechanism of chitin degradation using thermophiles. The current review discusses the sources of thermophilic chitinases, improvement of protein stability by gene manipulation, metagenomics approaches, chitin degradation mechanism in thermophiles, and their prospective applications for industrial, agricultural, and pharmaceutical purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thermophilic extremozymes are gaining wide interest globally over chemical catalysts due to their high tolerance and catalytic action at elevated temperatures. Predominantly, industries use chemical solvents that decrease the viscosity and elevate the diffusion content and reaction rate at high temperatures [1, 2]. The intrinsic properties of thermophilic enzymes are ideal in catalyzing chemical reactions at high temperatures. For industrial purposes, high temperatures can enhance the solubility of hydrophobic compounds, prevent contamination of microbes, and aid in the biodegradation process by environmentally friendly approaches. The microbial physiology of the thermophilic microbes, the nature of their enzyme catalytic center, and evolutionary diversity are helpful in protein engineering for industrial applications. Thermophiles are adapted to thrive at high-temperature sites like volcanic and geothermal springs [3]. Scientists are interested in unveiling the mysteries involved in the protein stability of these thermophiles and looking for possibilities in the industrial and health care sector.
Considering the availability of natural organic resources, chitin is the second most abundant organic polymer, next to cellulose. The exoskeletons of crustaceous sea animals like prawn, lobster, sea krill, crab, fungal cell wall, and the wings and skeletal remains of some insects contained chitin. The chitinases degrade the β-1, 4 linkages of chitin to yield N-acetyl glucosamine units. Bioconversion of chitin using chitinases can produce chitooligosaccharides and chitosan, which are widely used in the medical, industrial, agricultural, and environmental sectors. For agricultural applications, chitinases eliminate fungal diseases affecting crops by acting as a lytic enzyme that binds to the fungal cell wall containing chitin, inhibiting the growth and development of pathogenic fungi, and thus, they function as biocontrol agents [4]. Chitinases are used in the removal of shellfish wastes generated by the seafood industry and dumped into the water bodies. For the shellfish waste management, a higher temperature is required for the demineralization and deproteination of chitin wastes. The application of chitinases can convert these chitin raw materials to chitooligosaccharides compared with thermochemical processes generating low-quality heterogeneous mixtures [5]. Byproducts of chitin hydrolysis generate chitooligosaccharides (CHOs) that have antitumor activity [6]. Thus, the usage of stable and eco-friendly thermophilic enzymes can reduce the cost and increase reliability in various applications. In the present scenario, not much information is available to enhance the stability of thermophilic chitinases. Therefore, this review focuses on the various sources of thermophilic chitinases, engineering strategies, their applications, and future perspectives to improve their thermostability by biotechnological approaches.
Physiochemical Properties of Chitin and Chitosan
Chitin is a crystalline natural polymer that is widely found in the exoskeleton of crustaceans like shrimp, shellfish, lobster, and Antarctic krill in insects and cell walls of fungi. It is insoluble, inert, and inelastic resembling the cellulose molecule except that the hydroxyl groups of the C2 position are replaced by acetamide functional groups (−NH−CO−CH3). They are insoluble in organic solvents and water, and based on the structural nature, they are classified as alpha, beta, and gamma chitin. Alpha chitins are found mainly in crustacean species containing alternating anti-parallel polysaccharide chains, tightly packed with inter- and intramolecular hydrogen bonding. Beta chitin is found predominantly in squid with parallel polysaccharide chains that are loosely packed due to the lack of intermolecular hydrogen bonds. Gamma chitin is detected in fungi, and contains two parallel polysaccharide strands alternating with an anti-parallel polysaccharide strand [7].
Chitin is extracted from shellfish wastes by the removal of proteins and calcium carbonate from their exoskeleton. It involves the process of demineralization (DM) to remove the minerals and deproteination (DP) to remove the proteins. The process of demineralization and deproteination can be achieved either chemically or enzymatically [8]. Some methods use chemicals at high temperature for deproteination followed by demineralization using lactic acid bacteria or proteolytic enzymes followed by acid treatment at high temperature for demineralization [9]. One of the major derivatives of chitin is chitosan formed by the application of deacetylases [10]. Chitin is converted to chitosan by deacetylation using a concentrated base like NaOH at high temperatures or by fungal deacetylases derived from Mucor rouxii [11].
The major significance of using thermophilic chitinases is that their stability is maintained at higher temperature during industrial process. These thermophilic chitinases can reduce the cooling temperature, making the shellfish waste utilization and valorization process economically feasible. For example, Krolicka et al. [12], studied the application of thermostable chitinase Chi1 from Myceliophthora thermophila C1 that had thermostability at 55 °C with a half-life of 48 h. The chitinase Chi1 had broad host specificity utilizing colloidal chitin, chitosan, and chitooligosaccharides, and their activity was affected by molecular weight, degree of deacetylation (DD), and derivatization of chitosan.
Compared with chitin, chitosan is soluble in dilute acids like formic acid and acetic acid and are widely applied in the medical and pharmaceutical field. They are used as a nanoparticle carrier for delivery of therapeutic drugs to the targeted site, wound dressings to prevent scar-forming, and in the development of films that aid in wastewater management and water purification with antibacterial and antifungal functions [13]. Both chitin and chitosan are having versatile applications in various fields, as they are non-toxic, biocompatible, adsorbent, biodegradable, and found abundantly in nature and are deacetylated to form oligomers and monomers respectively.
Mechanism and Classification of Chitinases
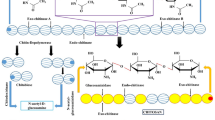
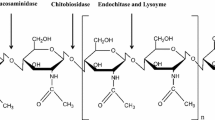
Chitin is hydrolyzed by the chitinases. Based on the cleavage action, they are classified into endochitinases and exochitinases. Endo-chitinase (E.C. 3.2.1.14) cleaves the internal β 1, 4 linkages to form oligomers (Fig. 1). These oligomers are further degraded by exochitinase (E. C. 3.2.1.52) to form N-acetyl glucosamine monomers. N-Acetyl glucosamine is deacetylated to glucosamine with the help of N-acetyl glucosamine deacetylases (E. C. 3.5.1.33) [14]. Chitinases belong to the group of CAZY databases and are grouped as 18, 19, and 20 glycosyl hydrolases based on their amino acid sequence similarity [15]. Group 18 chitinases are evolutionarily diverse having different molecular mechanisms and are found originally in bacteria, fungi, some animals, and plants. In bacteria, GH 18 chitinases are classified as class A and class B. Group 18 chitinases carry out transglycosylation reaction and are found mainly in Trichoderma harzianum (Chit 33 and Chit 42), Aspergillus fumigatus (Chi B1), etc. Group 18 chitinases have an enzymatic core with 8 parallel beta sheets. Group 19 chitinases are mainly found in the plant (I, II, and IV classes) and some Streptomyces. Groups 18 and 19 chitinases are likely to have evolved from different ancestral lines due to their evolutionary diversity and molecular mechanisms. GH 19 chitinases have an active site with deep cleft, and the catalytic domain has α-helix-rich lysozyme region. They are composed of various loops and helices in their catalytic subunits, which are differentiated as loopful or loopless. Extra loops are found on either side of the substrate-binding clefts in the case of loopful chitinases, which provides more sites for binding longer chitin. The loopless members do not have these extra loops. Group 20 chitinases are found in some Streptomyces, humans, and bacteria. They have a TIM barrel catalytic domain with deep and open cleft active site, e.g., Sm chitobiase [16]. Based on origin, mode of action, and amino acid sequences, chitinase classification is enlisted in Table 1.
Sources of Thermophilic Chitinases
Chitinases are glycosyl hydrolases having a molecular size ranging from 20 to 90 kDa and are found in bacteria, fungi, yeasts, actinomycetes, plants, and animals [37, 38]. They are prevalent in other sources like protozoans, arthropods, molluscs, nematodes, and coelenterates [39]. These enzymes have attracted researchers globally due to their diverse applications in various fields like agriculture, medicine, and environment management. Based on their stability at higher temperatures, thermophilic chitinases are isolated preferably from high-temperature zones like geothermal springs, hot springs, and volcanoes [3]. Thermophilic chitinases can be genetically engineered and protein engineered, apart from isolated from natural resources. Thermophilic chitinases are functional between temperatures ranging from 50 °C and above and are used in industrial and medical research for assays controlled at higher temperatures [37]. Some thermophilic chitinases reported in archaea, bacteria, and fungi and in plants are depicted in Table 2.
Thermophilic Chitinases from Bacteria
From Archaea
Thermostable chitinases are gaining attention for the past few years, due to their rise in industrial applications that require synthesis of products at high temperatures. They are isolated from extremely high-temperature environments, and these diverse cultures are maintained for future industrial applications. Thermostable chitinases are genetically engineered in host organisms and are also applied for various purposes [4]. The hyperthermophilic archaea species have higher resistance at elevated temperatures in comparison with the thermophilic bacterial chitinase [71]. Thermococcus chitonophagus was the first novel anaerobic hyperthermophilic archaeal strain isolated from the deep-sea hypothermal vents and found to break down chitin for nutritional functions [40, 72]. Another hyperthermophilic archaea Thermococcus kodakarensis KOD1 (earlier named as Pyrococcus kodakarensis KOD1) is the first chitinase, characterized by biochemical and structural evaluations [41]. A hyperthermophilic archaeal bacteria, Pyrococcus furiosus, contained 2 chitinase-producing genes Chi A and Chi B genes belonging to family 18 glycosyl hydrolases [42].
General Bacteria
Based on their different catalytic centers and active sites containing putative residues, the bacterial chitinases are different from the plant chitinases [31]. Most of the bacterial species belong to the GH18 family. Bacterial chitinases are divided into two subfamilies A and B based on their catalytic subunit with or without the chitin insertion domain (CID). The synergistic action of subfamilies A and B are well-explained in Serratia marcescens [50]. Serratia marcescens consists of SmChiA, SmChiB, and SmChiC that belong to GH18 families [73]. According to Brurberg et al. 1996 [74], the synergistic action of SmChi A and SmChi B enhanced the degradation rate of colloidal chitin in Serratia marcescens. Several bacterial chitinases were purified and characterized. A moderately tolerant thermophilic bacterium, from the genus Ralstonia, was isolated from the compost system containing chitin waste [75]. Similarly, thermostable endochitinases from Bacillus sp. MH-1 and thermostable exochitinases from Bacillus stearothermophilus CH-4 were isolated from compost pits respectively [38, 48]. Toharosman et al., 2005 [49] isolated Bacillus licheniformis MB-2 from the geothermal springs, situated in Indonesia with a molecular weight of 67 kDa and optimal pH 6.0 and optimal temperature of 70 °C. Also, other thermophilic chitinases from Paenibacillus species [76], Bacillus sp. [77], were isolated from hot springs located in Indonesia. Aeromonas sp. DYU-too 7 produces extracellular thermophilic chitinases. GlcN acted as an inducer in the production of chitinase [47]. Thermostable chitinases were obtained from Staphylococcus species that had an optimum temperature of 60 °C and pH 7.0 [78]. Thermophilic bacterium was isolated from Clostridial species; Clostridium thermocellum, isolated from hot springs, produced cellulosomes that contained catalytic and non-catalytic modules held together by protein-protein interaction. The chitinase gene chiA belonged to glycosyl hydrolase family 18 was found to be located downstream of the endoglucanase gene cel A in the cellulosome [43]. The extracellular chitinase produced by Cohnella sp. A01 had an optimum temperature of 70 °C and optimum pH 5.0 and could utilize colloidal chitin [51].
Actinomycetes
Currently, only a few thermophilic actinomycetes have been explored, although they have a wide range of bioactives and novel enzymes. Streptomyces sp. AC4 and AC7 contained thermophilic chitinases that were isolated from the compost containing predominantly horse litter [56]. Streptomyces species belong to a group of gram-positive bacteria that are having chitinolytic enzymes capable of degrading chitin in the soil, thus obtaining their carbon and nitrogen source [53, 79]. Streptomyces thermoviolaceus OPC-520 consists of thermostable chitinase genes having an optimum of 80 °C and having high pH ranging from 8.0 to 10.0 [53]. Another actinomycete, Microbispora sp. V2, contains mesophilic and thermophilic enzymes and is isolated from the hot springs. They had thermophilic chitinases with a molecular weight of 35 kDa and an optimum temperature of 60 °C [54]. Recently, thermostable chitinases from Streptomyces sp. F-3 showed higher chitin degradation activities [57, 58].
Thermophilic Chitinases from Fungi
Fungal chitinases belong to the group of class 18 glycohydrolases and show resemblance to class III plant chitinases [80]. GH family 18 chitinases have 5 domains comprising of (a) catalytic domain, (b) N- terminal signal peptide region, (c) chitin-binding domain, (d) serine-/threonine-rich region, and (e) C-terminal extension region [81]. The amino acid sequences of GH 18 chitinases revealed the presence of two highly conserved motifs: DXXDXDXE and SXGG. Fungal chitinases are classified as 3 subgroups A, B, and C based on their available phylogenetic sequences. Subgroup A fungal chitinases are extracellular in nature with molecular mass ranging from 40 to 50 kDa containing catalytic domain, but lacking chitin-binding domain. Subgroup B had a molecular mass ranging from 30 to 90 kDa and variable catalytic domain with chitin-binding sites. Subgroup C forms a novel category with molecular masses ranging from 140 to 170 kDa. They have a unique architecture with the presence of LysM motifs and CBM18 [82].
In the case of fungal chitinases, the source of the medium plays a primary role in chitinase production. For example, addition of colloidal chitin in the medium helped in the production of chitinase in Thermomyces lanuginosus SSBP. An inducer repressor system is followed in the chitinase production of T. lanuginosus. It was observed that as the byproducts increased after 96 h, it inhibited the chitinase production by feedback inhibition. Another thermophilic chitinases, produced by Humicola grisea ITCC 10,360.16 by submerged fermentation was optimized by response surface methodology using various parameters like colloidal chitin, chitin, yeast extract, and KCl [66]. Some fungal species represented both chitinase and amylolytic activities, e.g., Rhizopus oryzae [63]. Most thermophilic fungal chitinases were found to be acidophilic. For example, a commercial preparation from Trichoderma viride, designated as Usukizyme, showed multiple chitinolytic activities. One of these enzymes was found to be highly acidic (ranging from 3.5 to 6.0) and thermophilic with temperature ranging from 50 to 55 °C [62]. Other fungi like Talaromyces emersonii [61] and Gliocladium catenulatum [65] utilized chitin to form extracellular chitinases and chitobiase. The fungal chitinases had a wide range of properties having thermophilic, acidophilic, or amylolytic functions.
Plant Chitinases
The plant chitinases are purified and display antifungal activity in vitro. They are found in various parts of monocotyledonous and dicotyledonous plants; located in embryos, seeds, cotyledons, stem, leaves, flowers, leaf abscission zones, suspension cultures, and tissue-cultured callus [83]. In crop plants, they are found in peas, carrot, tomato, potato, rice, chickpea, rye, cabbage, cucumber, oat, tobacco, sugar beet, yam, turnip, wheat, etc. and non-crop plants like Arabidopsis, spruce, poplar, rubber, petunia, bent grass, and chestnut [83]. Plant chitinases belong to a group of pathogenesis-related (PR) and PR-3 proteins, which are induced in the host cells because of pathogenesis [84, 85]. PR proteins are also activated in the presence of various stress-related factors like salt, cold, UV light, drought, heavy metal, and addition of plant growth regulators [86]. Although there are few reports of thermostable plant chitinases, they have exochitinases with lysozymal activity (E. C. 3.2.1.17) that break down the β-1,4 linkages of N-acetyl glucosamine and N-acetyl muramic acid present in the peptidoglycan layer of bacteria [87]. Based on their amino acid residues, the plant chitinases are classified mainly as five different classes. Classes I, II, and IV contain globular domains and belong to GH 19 family [88]. Class I contains the presence of chitin-binding domain at the N terminus, whereas class II does not contain chitin-binding domain (CBD) at the N terminus. The CBD contains approximately 40 amino acids that have highly conserved cysteine residues. The GH 18 family comprises classes III and V plant chitinases which have 8 α and β strands and displays lysozymal activity [37]. Class III chitinases bear similarity with bacterial and fungal chitinases that lacks CBD. Class V chitinases contain 2 CBD and with C-terminal extension aiding in vacuolar targeting and have similarity to class III chitinases and bacterial chitinases. Class VI chitinases are found in sugar beet having ½ Chitin-binding domain [89]. Class VII chitinases shared 30% similarity to class I and class II chitinases. The catalytic domain of the plant chitinases comprises about 220 to 230 amino acid residues. The N terminal region separates the catalytic domain by a hinge region and is followed by a C-terminal region in many cases. Plant chitinases are involved in carrying symbiotic associations with mycorhizza and rhizobial bacteria for plant development, in contrast to their properties as a biocontrol agent against pathogenic fungi [86]. Some of the chitinases are involved in embryogenesis and ethylene synthesis. A thermostable legume chitinase was isolated, purified, and characterized from Phaseolus vulgaricus (Canadian Cranberry Beans). The molecular mass was determined as 30.6 kDa and was a monomeric protein with an optimum pH of 5.4 and optimum temperature of 40–55 °C and showed antifungal activity against Physalospora piricola, Pythium aphanidermatum, Botrytis cinerea, and Fusarium oxysporum [68]. Similarly, thermostable chitinase was isolated and purified from seeds of Adenanthera pavonina with optimum temperature 60 °C, but could be stable at 70 °C with optimum pH 4.0 [69]. The plant endochitinases had relatively less molecular mass ranging from 25 to 45 kDa in comparison with insect chitinases [37].
Insect Chitinases
Insect chitinases belong to GH 18 family and have high amino acid sequence similarity. They have three domains (a) catalytic domain; (b) PEST-like region containing proline, serine, threonine, and glutamate amino acids in majority; and (c) a cysteine rich region. These chitinases are reported to have molecular mass ranging from 40 to 85 kDa with optimal pH of 4.0–8.0 [81]. They are detected in Manduca sexta and Bombyx mori, which functions in the breakdown of CHOs during the process of ecdysis [90]. These insect chitinases are used for new cuticle generation and in the defense mechanism against other parasitic insects. These enzymes are inhibited by allosamindin and regulated by hormones during metamorphosis [90]. So far, there are no reports of thermophilic insect chitinases.
Metagenome-Sourced Thermostable Chitinases
Some of the thermophilic microorganisms are difficult to culture under harsh conditions and are not cultivable from natural sources. In the recent decade, microbial chitinases have been isolated from various habitats using degenerate primers specific for conserved catalytic regions of GH18 and GH19 by uncultivable methods. Cretoiu et al. 2012 [91] screened the presence of chitin-degrading bacterial communities from untapped terrestrial and aquatic habitats by culture-independent methods like DGGE using ChiA gene and ChiA gene pyrosequencing. The mining of such novel chitinase-producing communities could be beneficial for future biotechnological applications. Recently, uncultivable chitinases like Chi18H8 discovered from phytopathogen suppressive soil and 53D1 from chitin-supplemented agricultural soil was tested in vitro and in vivo against Bombyx mori. This is the first study to use metagenome-sourced chitinases as a bioinsecticide agent [92]. Similarly, various uncultivable novel thermostable chitinases from thermophilic natural habitats like volcanoes and hot springs can be discovered by metagenomics approaches and be used for various industrial applications.
Genome Analysis and Proteomics as a Tool for Elucidating the Mechanism of Chitin Degradation by Thermophilic Chitinase
Genome analysis can be a potential tool to prospect thermophilic chitinases from microorganism. Understanding the various mechanisms adopted by different bacteria to decompose chitin substrates possess an inevitable tool for biotechnological applications. Bacteria and fungi are the main mediators of polysaccharide decomposition in nature; however, the highly complex machinery used to sense and decompose these is not established [93]. Recently, a thermophilic T. chitonophagus has been reported to grow in chitin-containing media, and the bacteria are expected to possess an efficient system for chitin degradation and assimilation. To elucidate candidate genes involved in chitin degradation, Horiuchi et al (2016) [94] performed a genomic analysis of T. chitonophagus. Genome analysis revealed the presence of structurally novel chitinase, Tc-ChiD. The gene encoding Tc-ChiD contains regions corresponding to a signal sequence, two chitin-binding domains, and a putative catalytic domain. This catalytic domain shows no similarity with previously characterized chitinases. Furthermore, they found that the presence of the two chitin-binding domains significantly enhanced the thermostability of the catalytic domain [94].
Proteome analysis is another important technique to elucidate the mechanism behind the action of thermophilic chitinases. Sun et al. [57] identified a highly thermophilic bacterium Streptomyces sp. from the soil which can grow very well at 50 °C. They found that the bacterium produces a high amount of chitinases when chitin was supplied as the carbon source. To find out the mechanism and novel proteins involved in chitin degradation, Sun et al. [58] profiled the extracellular proteome and identified six chitin degradation–related enzymes like three chitinases of GH18 family, one GH19 chitinase, one GH20 β-N-acetylhexosaminidase, and polysaccharide monooxygenase from the AA10 family. They overexpressed all these chitin-degrading enzymes and characterized biochemically for industrial applications. From their study, the substrate binding, synergistic structures of the chitinases, and catalytic patterns were analyzed, and results indicated that three different GH18 subfamilies namely SsChi18A, SsChi18B, and SsChi1C had different specific substrate binding modes and different biological structures. To elucidate the structure of catalytic domains of SsChi18A, SsChi18B, and SsChi1C, PyMOL molecular visualization system was used (http://pymol.org/). Although all the three enzymes had a TIM barrel structure corresponding to the GH18 family, they had functional variations. SsChi18A acted as a processive chitinase, which is an essential property in chitin degradation. SsChi18A catalytic domain had an open deep active-binding cleft and a large α + β domain with 81 amino acids, a characteristic trait of processive enzymes. FACE analysis confirmed that SsChi18A enzymes could hydrolyze colloidal chitin and chitooligosaccharides (CHOs) to chitobiose. SsChi18A had a conserved motif SXGGW with Trp residue to be necessary for processivity. SsChi18B and SsChi1C appeared to be non-processive. SsChi18B has a shallow active-binding cleft with the accumulation of aromatic amino acids near the catalytic center. SsChi18B had a special β hairpin subdomain at loop 6 and resembled the predicted model of SmChiC from Serratia marcescens. The SsChi18C had 7 loops (a comparatively larger 7 loop) unlike 8 loops found in SsChi18A and SsChi18B without the presence of α + β domain in loop 7. Trp-257 and Arg-261 in loop 7 increased their substrate-binding ability from subsites − 1 and − 2. SsChi18C is reported to be an active enzyme with the catalytic motif DXXDXDXE with the first conserved aspartate replaced by threonine. One lytic polysaccharide monooxygenase SsLPMO10A helped in the high-efficiency degradation of chitin. Although the three chitinases in Streptomyces F-3; SsChi18A, SsChi18B, and SsChi1C, were having different structural and functional differences, they acted synergistically in the degradation of chitin. Similar kinds of the synergistic mechanism were observed in mesophilic bacteria like Serratia marcescens [73, 74] and non-thermophilic chitinase like Cellvibrio japonicus [91]. From the in vitro and in vivo approaches, it was identified that 4 enzymes, CjChi18A, CjChi18B, CjChi18C, and CjLPMO10A, from GH 18 family helped in chitin degradation.
Bioinformatic approaches using molecular stimulation by semi-rational design and structure analysis was used to enhance the thermostability of chitinases by improving their half-life period. The thermostability of mesophilic microorganisms could be improved by the introduction of disulfide bonds. An example is PpChi1 chitinase of mesophilic Paenibacillus pasadenensis [95]. The chitin-binding domains (CBDs) play an important role in degradation of chitin. Chitin-binding domains (CBDs) of chitinase-producing microorganisms can be incorporated into non-CBD producers to create thermostable and chemically stable chitinases by genetic engineering. Matroodi et al. [96] designed hybrid chitinases using the CBD of S. marcescens chitinase B with fungal chitinase of Trichoderma atroviridae Chit 42 that lacks CBD, to enhance their biocontrol functions.
Heterologous Gene Expression of Thermophilic Chitinase
One of the major challenges concerning application of thermophilic enzymes is hindrances in obtaining higher expression levels for industrial applications. Difficulties were encountered in cloning the thermophilic genes into the E. coli expression system as archaeal systems are closely related to eukaryotes than bacteria requiring strong promoters (T7 RNA polymerase, plac, ptac) for gene expression [97]. Low expression levels were observed in the E. coli expression system due to different codon applications in the expressed gene [98]. Among the E. coli expressed thermophilic genes, only 10% of the hyperthermophilic enzymes were found to display thermostability and catalytic and structural properties differing from the native enzymes [99].
Another limiting factor for cloning the genes is obtaining positive transformants with antibiotic selection. The commonly used antibiotics targeting mesophiles are ineffective against thermophiles that function at higher temperatures, as they degrade. In recent years, the antibiotic simvastatin was found to be effective in targeting the archaeal membrane and was tolerant at 100 °C, devoid of oxygen and successfully used in genetic engineering of T. kodakarensis [100] and P. furiosus [101]. Considering the limitations in antibiotic selection, nutritional selection with uracil prototrophic selection from an auxotrophic parental strain is preferred [102].
For cloning and expression of thermophilic chitinases, various host expression systems like E. coli and Pichia pastoris was used. There are several reports of thermophilic chitinase genes cloned and expressed in mesophilic organisms [103, 104]. Duffner et al. 1996 [103] cloned chiA gene which is labeled as Chia_Entag from Enterobacter agglomerans and expressed in E. coli JM109. This protein (Chia_Entag) contained 61 kDa molecular mass and showed antifungal activities against phytopathogens like Fusarium oxysporum and Rhizoctonia solani.
Large-scale overexpression of thermostable chitinase gene Chi 40 from Streptomyces thermoviolaceus OPC-520 was cloned and expressed in several expression vectors of E. coli [53]. Another thermostable chitinase from Chromobacterium violaceum expressed in Escherichia coli had an optimum pH 5.0 and optimum temperature 60 °C and maintained their thermostability at 60 °C for 30 min, retaining 32 % of the chitinase activity [105, 106]. A highly thermostable chitinase gene chi A of family 18 glycosyl hydrolase from Rhodothermus marinus was cloned and expressed in E. coli. The Chi A enzyme was modified due to poor expression by designing a signal peptide lacking mutant Chi AΔsp. The optimal pH for the chitinase activity was 70 °C and pH 4.5–5.0 for both Chi A and Chi AΔsp [45]. Due to their high growth rate and ability to grow in a simplified medium, Pichia pastoris is used as a host expression system. Sequencing of Thermomyces lanuginosus SSBP genome revealed the presence of 4 putative family 18 chitinases, chi 1, chit 2, chit 3, and chit4. chit 1 and chit2 were cloned and expressed in Pichia pastoris. The recombinant chit1 was optimally active at 50 °C, and the recombinant chit 2 had an optimum temperature of 40 °C [107]. Efficient expressions of thermostable chitinases in different hosts are enlisted in Table 3.
Recently, the Crispr cas 9 technology for gene editing has transformed the biotechnological research. Mougiakos et al. [113] developed ThermoCas9, from the thermophilic bacteria Geobacillus thermodenitrificans T12, that was used for gene deletion and transcriptional silencing of mesophilic microorganisms. Thermocas9 could serve as gene-editing tool for bacteria covering a broad temperature range, applicable for the production of valuable chemicals and fuels.
Protein Engineering
The thermostability of an enzyme is characterized by the primary structure of the protein and the catalytic activity of the enzyme can be increased by protein engineering. Various external factors like substrates, polyols, modulators proteins, and coenzymes increase thermostability. Thermodynamic studies are initiated recently to evaluate the thermostability of enzymes [114]. Various important parameters like entropy, enthalpy, and Gibbs free energy are used to determine the thermodynamics of the thermostable enzymes [115]. Other methods to improve the thermostability of enzymes are shortening or deletion of loops, smaller and lesser number of cavities, more proline accumulation, increased polar area and helical content, substitution of amino acids outside and within the secondary structures, and increase in hydrogen bonds and salt bridges with a decrease in the occurrence of thermolabile compounds [116]. The factors that are enhancing the thermostability of thermophilic enzymes can be analyzed by calculating the minimum folding energy of their secondary mRNA structures and the amino acid substitution. It is crucial to study the location of the amino acids and their energy differences responsible for causing the thermostability of enzymes as they were dependent on amino acid substitutions. For example, in the case of T. thermophilus HB27, arginine was substituted by glutamine and lysine followed by alanine substituted with serine and threonine [117]. It was observed that the stabilizing mutations were found on the loops and surfaces of the protein.
The main focus in chitinase research is to improve its catalytic activity which can be achieved through protein engineering. Two major approaches for engineering protein are directed evolution and mutagenesis (random and site-directed).
Directed Evolution
Directed evolution is the natural process of evolving the genes after several rounds of mutation, recombination, and selection [118]. In the directed evolution approach, by error-prone PCR, chitinase was randomly generated to produce a mutant library that is screened using fluorescent-activated cell sorter (FACS). Mutants having a higher catalytic activity compared with the wild-type clones were selected. DNA shuffling is also applied to shuffle the genes to generate a large number of thermostable organisms. After DNA shuffling, the thermostable enzymes are screened based on their temperature tolerance and percentage of enzyme retained at higher temperature [119]. DNA shuffling and screening enhanced the catalytic ability of the chitinase Beauveria bassiana BbChit1. After three rounds of error-prone PCR, and 1, 50, and 000 variants of the Bbchit1 library were expressed in E. coli and signal peptide Pel B for extracellular chitinase production, only two mutants SHU-1 and SHU-2 were tested positive for chitinase activity [120]. A direct evolution method for the improvement of chitinase activity in Bacillus licheniformis was established [121]. Out of 517 colonies, one mutant showed improved chitinase activity that was 2.7-fold higher than the wild-type at pH 3.0. This method is ideal to generate mutant chitinases for various industrial applications.
Random Mutagenesis and Site-Directed Mutagenesis
Mutation by chemical and physical methods, followed by selection and screening, is used to determine the enhanced thermostable activity of the enzymes for various applications [122]. Physical mutations using UV radiation are one of the most common methods used for strain improvement. Vahed et al. [52] improved the halotolerant nature of chitinase ChiL produced from Bacillus pumilus SG2 by UV radiation and nitrous acid treatment. A single-base mutation (point mutation) in the catalytic domain of Chi L enhanced the chitinase activity. There was a single-base change from GGA coding for Gly residue to GAA coding for Glu at position 432. The mutant ChiL designated as ChiLm showed more thermostability than the wild-type at 70 °C, and their activity continued to be stable after sequential sub-culturing for 6 months. According to Chen et al. [123], a thermophilic chitinase Pachi, from P. aeruginosa, increased their nematocidal activity against C. elegans when associated with Cry 21Aa of B. thuringiensis. Pachi chitinases produced PachiN35D, which increased the substrate specificity and catalytic efficiency by random mutagenesis and screening using bacteriophage T7. One base substitution occurred in N35 that is situated near the catalytic domain and closer to the TIM barrel, which occurs in GH 18 family. The wild-type Pachi chitinase had an optimal temperature of 60 °C, whereas the mutant PachiN35D had a temperature optimum of 50 °C [124].
In Site-Directed Mutagenesis (SDM), thermostable chitinases are designed by rational engineering. Site-Directed Mutagenesis of bacterial chitinases is shown in Table 4. Some methods of rational engineering include (a) introduction of the disulfide linkages, (b) α-helix dipole stabilization, (c) point mutation to reduce the entropy of the unfolded state, and (d) hydrophobic core improvement [14]. Disulfide bridges have covalent bonds in reducing the entropy of the unfolded polypeptide chain and improve the protein rigidity. The dissociation energy of disulfide bridges is approximately 60 Kcal/mol, which helps in imparting thermostability to the enzymes [130]. Recently, computation approaches were designed to introduce disulfide bridges to increase the thermostability of DFPase enzyme that helped in organophosphorus degradation [131]. The hydrophobic core containing aromatic amino acids like tyrosine, tryptophan, and phenylalanine engage in energy-favoring reactions. They are found mainly in globular proteins and can maintain compact arrangements even in an aqueous medium, thus maintaining their stability [14]. Another area to increase the protein thermostability is by domain swapping. It is used to study the domain compatibility for increasing thermostability and aid the binding ability of chitinases to insoluble chitin. Neeraja et al. [132] cloned and characterized the chitinase from Bacillus thuringiensis and B. licheniformis consisting of a catalytic domain of GH 18 family at the N terminal and chitin-binding domain (CBD) at the C-terminal. The recombinant chitinases (BtGH-BliChBD and BliGH-BtChBD) were designed by domain swapping as a means to exchange the chitin-binding domain (CBD) of BliGH-ChBD with BtGH-ChBD and vice versa. Both the chimeric chitinases improved the conformational stability, functional stability, and thermostability of the chitinases. Refolding experiments followed by CD analysis indicated that different chimeric chitinases showed 60–65 °C (BtGH-ChBD), 40 °C (BtGH-BliChBD), 60–65 °C (BliGH-ChBD), and 60–65 °C (BliGH-BtChBD), proving that the domain-swapped chitinases were thermostable.
Immobilization of Thermophilic Chitinases
Chitinase immobilization improves pH and thermal stability, catalytic activity, and enzyme reusability and improves chitin hydrolysis. The selection of a proper low-cost carrier to reuse the enzyme efficiently is highly preferred [133]. Prasad and Palanivelu [134] immobilized the recombinant thermostable fungal chitinase of T. lanuginosus on glutaraldehyde cross-linked chitosan beads. The immobilized enzyme was stable at alkaline pH and optimally active at 60 °C and stable at 50 °C for 3 h. Mansour et al. [67] studied the production of thermophilic chitinase from Penicillium chyrsogenum. The enzyme was partially purified and immobilized by modified bentonite and had a 100% relative activity at 55 °C. The purified enzyme from Bacillus sp. BG-11 was immobilized using chitosan and calcium alginate and had an optimal pH, and the temperatures of the purified immobilized enzymes were 8.5 and 50 °C, respectively [135].
Functional Application of Thermophilic Chitinases
Thermostable chitinases can tolerate temperature ranging from 65 to 75 °C and help in the conversion of chitin to chitobiose, chitin to N-acetyl glucosamine by chitobiase and chitin to chitosan with deacetylases [10]. The chitosan generated by chitinases using deacetylases has applications in wastewater removal, textile treatment, and development of photographic products, construction materials, and heavy metal–chelating agents [10]. In the chemical process, deacetylation from chitin to chitosan is carried out using 40% NaOH, which is harmful to the water bodies. This chemical process can be replaced by environmentally friendly methods using thermostable bacterial or fungal chitin deacetylases. Since chitin is insoluble in water and crystalline in nature, it can be degraded efficiently using thermostable chitinase. Recycling solid organic wastes using microorganisms at higher temperatures is considered an effective way for the degradation of wastes which results in the formation of fertilizers that can nourish the soil. Chitooligosaccharides are generated by the action of thermophilic bacterial and fungal chitinases on chitin. Chitinases from Rhodothermus marinus hydrolyzed 73% of the deacetylated chitosan [45]. Similarly, many thermophilic bacterial chitinases of GRAS status like Bacillus licheniformis and Cohnella sp. with low pH and high-temperature tolerance can be considered for the generation of CHOs on a larger scale. Thermophilic chitinases can be used in the development of medicinal drugs. Bacillus altitudinus displayed antifungal activity against vaginal infection causing Candida species and thus can be considered a potential drug [136]. Thermophilic chitinases from gene Enterobacter agglomerans expressed in E. coli [104] and Streptomyces AC4 and AC7 can be used as a biocontrol agent against pathogenic fungi affecting beneficial plants, thus can be used for horticultural and agricultural purposes [56]. Chitinases from Microbispora sp. V2 and thermophilic fungi T. aurantiacus var. levisporus and C. thermophilum cloned and expressed in P. pastoris GS115 can be ideally used in the recycling of chitinous wastes generated from the fish industry, due to their acidophilic and thermophilic properties [54, 59].
Fungal chitinases aid in the production of single-cell protein, fungal protoplast generation, and for the production of various valuable chitooligosaccharides, which finds applications in medical and industry, e.g., Myceliophthora thermophila C1 [12]. Chitinases obtained from thermophilic fungi Thermomyces lanuginosus SSBP (Chit1) and Bacillus licheniformis (Chit lic) could be used as a potential biopesticide against the second instar larvae of Eldana saccharina and showed antifungal activity against pathogenic fungi [60, 137]. Usukizyme, from T. viridae under cellulose-induced conditions, is used in the protoplast fusion studies [138]. A highly thermostable chitinase was isolated from pineapple (Annanas comosus), which is found to be the most stable plant chitinase reported so far and can be used for various industrial applications [70]. Chitinase production can be improved by optimizing the various process parameters by solid-state fermentation, which is studied in Penicillium chrysogenum [139] and Trichoderma harzianum [140]. Also, crude chitinases were generated by submerged fermentation using colloidal chitin in Lecanicillium fungicola for the production of oligosaccharides by the hydrolysis of chitin [141]. A thermophilic chitinase Pachi, from Pseudomonas aeruginosa, showed nematocidal effects against C. elegans. On random mutagenesis, the mutant of chitinase Pachi (PachiN35D) showed increased nematocidal activity than their wild-types by mortality assays, quantitative growth measurements, and brood tests [124]. Thermophilic immobilized chitinases from Penicillium chrysogenum were reported to be having an insecticidal activity against larvae of Cx. pipiens L, a pest responsible for transmitting lymphatic filariasis, Rift Valley fever virus, and West Nile virus [67]. Thus, thermophilic chitinases are beneficial and applied as in mesophilic chitinases [142]. Some of the recent potential applications of thermophilic chitinases are mentioned in Table 5.
Conclusion and Future Perspectives
Thermophilic chitinases are gaining significance among researchers considering their wide range of applications in the field of medicine, agriculture, and the environment. The usage of these enzymes can minimize the cost in industrial applications, as they are less prone to microbial contamination and could be functional at higher temperatures. Various gene-cloning methods have been promoted to introduce the thermophilic chitinase gene into mesophilic microorganisms. Random mutagenesis and site-directed mutagenesis, followed by selection and screening, can generate thermophilic chitinases of greater activity than their wild-types. The reusability of the thermophilic enzymes is achieved by immobilization for repeated usage in the industry, thus making it economically feasible.
Chitinases are generally known biocontrol agents against phytopathogenic fungi and pests under normal environmental conditions. Thermophilic chitinases can be used against fungi that can withstand arid and high-temperature conditions. Leafcutter ants are agricultural pests found in the tropical areas of Central and South America, and Mexico, and can affect the citrus vegetation by consuming the leaves voraciously and spread opportunistic diseases due to microorganisms as a result of foraging. The addition of thermophilic chitinases can remove the presence of such kinds of pests. Chitinases can also be applied to stagnant water bodies to eliminate mosquito larvae at various instars to prevent the incidence of malaria, dengue, etc., as chitinases can degrade the insect cuticle containing chitin. In the health care sector, thermophilic chitinases can be used as a microbicide in ophthalmic formulations and as an antifungal agent in lotions and creams that are applied topically. Since the thermophilic chitinases can be thermostable, they have a longer shelf life and preserve the integrity of the drug for an extended period. Thermophilic chitinases expressed in yeast like S. pompe cause the yeast cells to be swollen and elongated. This chitinase expressed fission yeast can be used in tea brewing or possibly alcohol fermentation. To conclude, various proteomics and bioinformatics tools can be used to analyze the structure of the chitinases and their conserved sequences to improve their stability and high temperatures. Genetic engineering has also paved a way to create hybrid chitinases that can be secreted by mesophilic organisms. Thus, novel applications can initiate using engineered thermophilic chitinases in the future.
References
Krahe, M., Antranikian, G., & Märkl, H. (1996). Fermentation of extremophilic microorganisms. FEMS Microbiology Reviews, 18, 271–285.
Becker, P., Abu-Reesh, I., Markossian, S., Antranikian, G., & Märkl, H. (1997). Determination of the kinetic parameters during continuous cultivation of the lipase-producing thermophile Bacillus sp. IHI-91 on olive oil. Applied Microbiology and Biotechnology, 48, 184–190.
Niehaus, F., Bertoldo, C., Kahler, M., & Antranikian, G. (1999). Extremophiles as a source of novel enzymes for industrial applications. Applied Microbiology and Biotechnology, 51(6), 711–729.
Neeraja, C., Anil, K., Purushotham, P., Suma, K., Sarma, P. V. S. R. N., Moerschbacher, B. M., & Podile, A. R. (2010a). Biotechnological approaches to develop bacterial chitinases as a bio shield against fungal diseases of plants. Critical Reviews in Biotechnology, 30(3), 231–241.
Liaqat, F., & Eltem, R. (2018). Chitooligosaccharides and their biological activities: A comprehensive review. Carbohydrate Polymers, 184, 243–259.
Liang, T.-W., Chen, Y.-J., Yen, Y.-H., & Wang, S.-L. (2007). The antitumor activity of the hydrolysates of chitinous materials hydrolyzed by crude enzyme from Bacillus amyloliquefaciens V656. Process Biochemistry, 42, 527–534.
Kaya, M., Mujtaba, M., Ehrlich, H., Salaberria, A. M., Baran, T., Amemiya, C. T., Galli, R., Akyuz, L., Sargin, I., & Labidi, J. (2017). On chemistry of γ-chitin. Carbohydrate Polymers, 176, 177–186.
Younes, I., & Rinaudo, M. (2015). Chitin and chitosan preparation from marine sources. Structure, properties and applications. Marine Drugs, 13(3), 1133–1174.
Healy, M., Green, M., & Healy, A. (2003). Bioprocessing of marine crustacean shell waste. Acta Biotechnologica, 23(23), 151–160.
Haki, G. D., & Rakshit, S. K. (2003). Developments in industrially important thermostable enzymes: a review. Bioresource Technology, 89, 17–34.
Kafetzopoulos, D., Martinou, A., & Bouriotis, V. (1993). Bioconversion ofchitin to chitosan: purification and characterization of chitin deacetylase from Mucor rouxii. Proceedings of the National Academy of Sciences of the United States of America, 90(7), 2564–2568.
Krolicka, M., Hinz, S. W. A., Koetsier, M. J., Joosten, R., Eggink, G., van den Broek, L. A. M., & Boeriu, C. G. (2018). Chitinase Chi1 from Myceliophthora thermophila C1, a thermostable enzyme for chitin and chitosan depolymerization. Journal of Agricultural and Food Chemistry, 66(7), 1658–1669.
Aam, B. B., Heggset, E. B., Norberg, A. L., Sørlie, M., Vårum, K. M., & Eijsink, V. G. (2010). Production of chitooligosaccharides and their potential applications in medicine. Marine Drugs, 8, 1482–1517.
Sarma, P. V. S. R. N., Prakash, J. M., Das, S. N., et al. (2013). Microbial chitinases: Natural sources, mutagenesis, and directed evolution to obtain thermophilic counterparts. In T. Satyanarayana, Littlechild, & J. Y. Kawarabayasi (Eds.), Thermophilic microbes in environmental and industrial biotechnology: Biotechnology of thermophiles, Vol. 2. Dordrecht: Springer.
Henrissat, B., & Bairoch, A. (1993). New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. The Biochemical Journal, 293, 781–788.
Oyeleye, A., & Normi, Y. M. (2018). Chitinase: diversity, limitations, and trends in engineering for suitable applications. Bioscience Reports, 38(4), BSR2018032300.
Patil, S. R., Ghormade, V., & Deshpande, M. V. (2000). Chitinolytic enzymes: An exploration. Enzyme and Microbial Technology, 26(7), 473–483.
Veliz, E. A., Martínez-Hidalgo, P., & Hirsch, A. M. (2017). Chitinase-producing bacteria and their role in biocontrol. AIMS Microbiology, 3, 689–705.
Park, J. K., Morita, K., Fukumoto, I., Yamasaki, Y., Nakagawa, T., Kawamukai, et al. (1997). Purification and characterisation of the chitinase (ChiA) from Enterobacter sp. G-1. Bioscience, Biotechnology, and Biochemistry, 61(4), 684–689.
Wang, S. L., & Chang, W. T. (1997). Purification and characterization of two bifunctional chitinases/lysozymes extracellularly produced by Monascus purpureus CCRC31499 in a shrimp and crab shell powder medium. Journal of Agricultural and Food Chemistry, 50, 2249–4085.
Rush, C. L., Schüttelkopf, A. W., Hurtado-Guerrero, R., Blair, D. E., Ibrahim, A. F. M., Desvergnes, S., Eggleston, I. M., & van Aalten, D. M. F. (2010). Natural product-guided discovery of a fungal chitinase inhibitor. Chemistry & Biology, 17(12), 1275–1281.
Yang, J., Gan, Z., Lou, Z., Tao, N., Mi, Q., Liang, L., Sun, Y., Guo, Y., Huang, X., Zou, C., Rao, Z., Meng, Z., & Zhang, K.-Q. (2010). Crystal structure and mutagenesis analysis of chitinase CrChi1 from the nematophagous fungus Clonostachys rosea in complex with the inhibitor caffeine. Microbiology., 156(12), 3566–3574.
Chen, L., Liu, T., Zhou, Y., Chen, Q., Shen, X., & Yang, Q. (2014). Structural characteristics of an insect group I chitinase, an enzyme indispensable to moulting. Acta Crystallographica Section D, 70(4), 932–942.
Cavada, B. S., Moreno, F. B., da Rocha, B. A., et al. (2006). cDNA cloning and 1.75 A crystal structure determination of PPL2, an endochitinase and N-acetylglucosamine-binding hemagglutinin from Parkia platycephala seeds. The FEBS Journal, 273(17), 3962–3974.
Ohnuma, T., Numata, T., Osawa, T., Mizuhara, M., Vårum, K. M., & Fukamizo, T. (2011). Crystal structure and mode of action of a class V chitinase from Nicotiana tabacum. Plant Molecular Biology, 75(3), 291–304.
Sutherland, T. E., Andersen, O. A., Betou, M., Eggleston, I. M., Maizels, R. M., van Aalten, D., & Allen, J. E. (2011). Analyzing airway inflammation with chemical biology: dissection of acidic mammalian chitinase function with a selective drug-like inhibitor. Chemistry & Biology, 18(5), 569–579.
Tran, T. N., Doan, C. H., Nguyen, M. H., Nguyen, V. B., Vo, T. P. K., Nguyen, A. D., & Wang, S.-L. (2019). An exochitinase with N-acetyl-β-glucosaminidase-like activity from shrimp head conversion by Streptomyces speibonae and its application in hydrolyzing β-chitin powder to produce N-acetyl-d-glucosamine. Polymers, 11(10), 1600.
Ruiz-Roldan, C., & Roncero, M. I. (2016). Quantification of chitinase activity in Fusarium oxysporum. Bio-Protocol, 6, e1882.
Graham, L. S., & Sticklen, M. B. (1994). Plant chitinases. Canadian Journal of Botany, 72, 1057–1083.
Terwisscha van Scheltinga, A. C., Kalk, K. H., Beintema, J. J., & Dijkstra, B. W. (1994). Crystal structures of hevamine, a plant defence protein with chitinase and lysozyme activity, and its complex with an inhibitor. Structure., 2(12), 1181–1189.
Hoell, I. A., Dalhus, B., Heggset, E. B., Aspmo, S., & Eijsink, V. G. H. (2006). and I. Crystal structure and enzymatic properties of a bacterial family 19 chitinase reveal differences from plant enzymes. The FEBS Journal, 273, 4889–4900.
Hahn, M., Hennig, M., Schlesier, B., & Höhne, W. (2000). Structure of jack bean chitinase. Acta Crystallographica. Section D, Biological Crystallography, 56(9), 1096–1099.
Ubhayasekera, W., Tang, C. M., Ho, S. W. T., Berglund, G., Bergfors, T., Chye, M. L., & Mowbray, S. L. (2007). Crystal structures of a family 19 chitinase from Brassica juncea show flexibility of binding cleft loops. The FEBS Journal, 274(14), 3695–3703.
Ubhayasekera, W., Rawat, R., Ho, S. W., Wiweger, M., Von Arnold, S., Chye, M. L., & Mowbray, S. L. (2009). The first crystal structures of a family 19 class IV chitinase: The enzyme from Norway spruce. Plant Molecular Biology, 71(3), 277–289.
Tews, I., Perrakis, A., Oppenheim, A., Dauter, Z., Wilson, K. S., & Vorgias, C. E. (1996). Bacterial chitobiase structure provides insight into catalytic mechanism and the basis of Tay-Sachs disease. Nature Structural Biology, 3(7), 638–648.
Swiontek Brzezinska, M., Jankiewicz, U., Burkowska, A., & Walczak, M. (2014). Chitinolytic microorganisms and their possible application in environmental protection. Current Microbiology, 68(1), 71–81.
Hamid, R., Khan, M. A., Ahmad, M., Ahmad, M. M., Abdin, M. Z., Musarrat, J., & Javed, S. (2013). Chitinases: an update. Journal of Pharmacy & Bioallied Sciences, 5, 21–29.
Sakai, K., Yokota, A., Kurokawa, H., Wakayama, M., & Moriguchi, M. (1998). Purification and characterization of three thermostable endochitinases of a noble Bacillus strain, MH-1, isolated from chitin-containing compost. Applied and Environmental Microbiology, 64(9), 3397–3402.
Muzzarelli, R. A. A. (1996). Chitosan-based dietary foods. Carbohydrate Polymers, 29(4), 309–316.
Huber, R., Stöhr, J., Hohenhaus, S., Rachel, R., Burggraf, S., Jannasch, H. W., & Stetter, K. O. (1995). Thermococcus chitonophagus sp. nov., a novel, chitin-degrading, hyperthermophilic archaeum from a deep-sea hydrothermal vent environment. Archives of Microbiology, 164(4), 255–264.
Tanaka, T., Fujiwara, S., Nishikori, S., Fukui, T., Takagi, M., & Imanaka, T. (1999). A unique chitinase with dual active sites and triple substrate binding sites from the hyperthermophilic archaeon Pyrococcus kodakaraensis KOD1. Applied and Environmental Microbiology, 65(12), 5338–5344.
Gao, J., Bauer, M. W., Shockley, K. R., Pysz, M. A., & Kelly, R. M. (2003). Growth of hyperthermophilic archaeon Pyrococcus furiosus on chitin involves two family 18 chitinases. Applied and Environmental Microbiology, 69, 3119–3128.
Zverlov, V. V., Fuchs, K.-P., & Schwarz, W.-H. (2002). Chi18A, the endochitinase in the cellulosome of the thermophilic, cellulolytic bacterium Clostridium thermocellum. Applied and Environmental Microbiology, 68(6), 3176–3179.
Takayanagi, T., Ajisaka, K., Takiguchi, Y., & Shimahara, K. (1991). Isolation and characterization of thermostable chitinases from Bacillus licheniformis X_7u. Biochimica et Biophysica Acta, 1078(3), 404–410.
Hobel, C. F. V., Hreggvidsson, G. O., Marteinsson, V. T., Bahrani-Mougeot, F., Einarsson, J. M., & Kristjánsson, J. K. (2005). Cloning, expression, and characterization of a highly thermostable family18 chitinase from Rhodothermus marinus. Extremophiles, 9, 53–64.
Gohel, V., & Naseby, D. C. (2007). Thermal stabilization of chitinolytic enzymes of Pantoea dispersa. Biochemical Engineering Journal, 35(2), 150–157.
Lien, T. S., Yu, S. T., Wu, S. T., & Too, J. R. (2007). Induction and purification of a thermophilic chitinase produced by Aeromonas sp. DYU-too7 using glucosamine. Biotechnology and Bioprocess Engineering, 12, 610–617.
Sakai, K., Narihara, M., Kasama, Y., Wakayama, M., & Moriguchi, M. (1994). Purification and characterization of thermostable β-N-acetylhexosaminidase of Bacillus stearothermophilus CH-4 isolated from chitin-containing compost. Applied and Environmental Microbiology, 60(8), 2911–2915.
Toharisman, A., Suhartono, M. T., Spindler-Barth, M., Hwang, J.-K., & Pyun, Y.-R. (2005). Purification and characterization of a thermostable chitinase from Bacillus licheniformis MB-2. World Journal of Microbiology and Biotechnology, 21, 733–738.
Vaaje-Kolstad, G., Horn, S. J., Sørlie, M., & Eijsink, V. G. H. (2013). The chitinolytic machinery of Serratia marcescens – a model system for enzymatic degradation of recalcitrant polysaccharides. The FEBS Journal, 280(13), 3028–3049.
Aliabadi, N., Aminzadeh, S., Karkhane, A. A., & Haghbeen, K. (2016). Thermostable chitinase from Cohnella sp. A01: isolation and product optimization. Brazilian Journal of Microbiology, 47(4), 931–940.
Vahed, M., Motalebi, E., Rigi, G., Noghabi, K. A., Soudi, M. R., Sadeghi, M., & Ahmadian, G. (2013). Improving the chitinolytic activity of Bacillus pumilus SG2 by random mutagenesis. Journal of Microbiology and Biotechnology, 23(11), 1519–1528.
Tsujibo, H., Endo, H., Miyamoto, K., & Inamori, Y. (1995). Expression in Escherichia coli of a gene encoding a thermostable chitinase from Streptomyces thermoviolaceus OPC-520. Bioscience, Biotechnology, and Biochemistry, 59(1), 145–146.
Nawani, N. N., Kapadnis, B. P., Das, A. D., Rao, A. S., & Mahajan, S. K. (2002). Purification and characterization of a thermophilic and acidophilic chitinase from Microbispora sp V2. Journal of Applied Microbiology, 93, 965–975.
Gomes, R. C., Semedo, L. T. A. S., Soares, R. M. A., Linhares, L. F., Ulhoa, C. J., Alviano, C. S., & Coelho, R. R. R. (2001). Purification of a thermostable endochitinase from Streptomyces RC1071 isolated from a cerrado soil and its antagonism against phytopathogenic fungi. Journal of Applied Microbiology, 90(4), 653–661.
González-Franco, A. C., Robles-Hernández, L., & Strap, J. L. (2017). Chitinase, chitosanase, and antifungal activities from thermophilic Streptomycetes isolated from compost. ϕYTON International Journal of Experimental Botany, 86, 14–27.
Sun, X., Meng, J., Liu, S., Zhang, H., & Wang, L. (2016). Draft genome sequence of Streptomyces sp. F-3. Genome Announcements, 4, e00780–e00816.
Sun, X., Li, Y., Tian, Z., Qian, Y., Zhang, H., & Wang, L. (2019). A novel thermostable chitinolytic machinery of Streptomyces sp. F-3 consisting of chitinases with different action modes. Biotechnology for Biofuels, 12, 136.
Li, A. N., Yu, K., Liu, H. Q., Zhang, J., Li, H., & Li, D. C. (2010). Two novel thermostable chitinase genes from thermophilic fungi: Cloning, expression and characterization. Bioresource Technology, 101(14), 5546–5551.
Khan, F. I., Bisetty, K., Singh, S., Permaul, K., & Hassan, M. I. (2015). Chitinase from Thermomyces lanuginosus SSBP and its biotechnological applications. Extremophiles., 19(6), 1055–1066.
McCormack, J., Hackett, T. J., Tuohy, M. G. and Coughlan, M.P. (1991). Chitinase production by Talaromyces emersonii. Biotechnol. Lett.13, 677–682.
Omumasaba, C. A., Yoshida, N., & Ogawa, K. (2001). Purification and characterization of a Chitinase from Trichoderma viride. The Journal of General and Applied Microbiology, 47(2), 53–61.
Chen, W.-M., Chen, C.-S., & Jiang, S.-T. (2013). Purification and characterisation of extracellular chitinase from Rhizopus oryzae. Journal of Marine Science and Technology, 21, 361–366.
Xia, G., Jin, C., Zhou, J., Yang, S., Zhang, S., & Jin, C. A. (2001). A novel chitinase having a unique mode of action from Aspergillus fumigatus YJ-407. European Journal of Biochemistry, 268(14), 4079–4085.
Ma, G.-Z., Gao, H.-L., Zhang, Y.-H., Joosten, R., Eggink, G., van den Broek, L. A. M., & Boeriu, C. G. (2012). Purification and characterization of Chitinase from Gliocladium catenulatum strain HL-1. African Journal of Microbiology Research, 6, 4377–4383.
Kumar, M., Brar, A., Vivekanand, V., & Pareek, N. (2017). Production of chitinase from thermophilic Humicola grisea and its application in production of bioactive chitooligosaccharides. International Journal of Biological Macromolecules, 104(Pt B), 1641–1647.
Mansour, A., Abdel-Fattah, M., Esawy, M. A., et al. (2019). Immobilization, thermodynamic studies and application of chitinase enzyme from Penicillium chrysogenum. Egyptian Journal of Aquatic Biology and Fisheries, 23(3), 527–544.
Wang, S., Shao, B., Fu, H., & Rao, P. (2009). Isolation of a thermostable legume chitinase and study on the antifungal activity. Applied Microbiology and Biotechnology, 85, 313–321.
Santos, I. S., Da Cunha, M., Machado, O. L. T., & Gomes, V. M. (2004). Chitinase from Adenanthera pavonina L. seeds: purification, characterisation and immunolocalisation. Plant Science, 167, 1203–1210.
Onaga, S., Chinen, K., Ito, S., & Taira, T. (2011). Highly thermostable chitinase from pineapple: Cloning, expression, and enzymatic properties. Process Biochemistry, 46(3), 695–700.
Eichler, J. (2001). Biotechnological uses of archaeal extremozymes. Biotechnology Advances, 19(4), 261–278.
Tanaka, T., Fukui, T., Atomi, H., & Imanaka, T. (2003). Characterization of an exo-β-D-glucosaminidase involved in a novel chitinolytic pathway from the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. Journal of Bacteriology, 185(17), 5175–5181.
Horn, S. J., Sørbotten, A., Synstad, B., Sikorski, P., Sørlie, M., Varum, K. M., & Eijsink, V. G. H. (2006). Endo/exo mechanism and processivity of family 18 chitinases produced by Serratia marcescens. The FEBS Journal, 273(3), 491–503.
Brurberg, M. B., Nes, I. F., & Eijsink, V. G. H. (1996). Comparative studies of chitinases A and B from Serratia marcescens. Microbiology, 142, 1581–1589.
Sutrisno, A., Ueda, M., Abe, Y., Nakazawa, M., & Miyatake, K. (2004). A chitinase with high activity toward partially N-acetylated chitosan from a new, moderately thermophilic, chitin-degrading bacterium, Ralstonia sp. A-471. Applied Microbiology and Biotechnology, 63(4), 398–406.
Chrisnasari, R., Yasaputera, S., Christianto, P., Santoso, V. I., & Pantjajani, T. (2016). Production and characterization of chitinases from thermophilic bacteria isolated from Prataan hot spring. East Java. J. Math .Fund. Sci, 48, 149–163.
Yuli, P. E., Suhartono, M. T., Rukayadi, Y., Hwang, J. K., & Pyun, Y. R. (2004). Characteristic of thermostable chitinase enzymes from the Indonesian Bacillus sp.13.26. Enzyme and Microbial Technology, 35, 147–153.
Basu, D., & Chaudhuri, A. N. (2014). Purification and characterization of chitinase from thermophilic Staphylococcus sp. Journal of Environmental Sciences, 4, 458–467.
Watanabe, T., Kanai, R., Kawase, T., Tanabe, T., Mitsutomi, M., Sakuda, S., & Miyashita, K. (1999). Family 19 chitinases from Streptomyces species: Characterization and distribution. Microbiology, 145, 3353–3363.
Hayes, C. K., Klemsdal, S., Lorito, M., Di Petro, A., Peterbauer, C., Nakas, J. P., Tronsmo, A., & Harman, G. E. (1994). Isolation and sequence of an endochitinase-encoding gene from a cDNA library of Trichoderma harzianum. Gene., 138(1-2), 143–148.
Merzendorfer, H., & Zimoch, L. (2003). Chitin metabolism in insects: structure, function and regulation of chitin synthases and chitinases. The Journal of Experimental Biology, 206(24), 4393–4412.
Hartl, L., Zach, S., & Seidl-Seiboth, V. (2012). Fungal chitinases: diversity, mechanistic properties and biotechnological potential. Applied Microbiology and Biotechnology, 93(2), 533–543.
Punja, Z. K., & Zhang, Y. Y. (1993). Plant chitinases and their roles in resistance to fungal disease. Journal of Nematology, 25, 526–540.
Van Loon, L. C., Rep, M., & Pieterse, C. M. J. (2006). Significance of inducible defense related proteins in infected plants. Annual Review of Phytopathology, 44(1), 135–162.
Legrand, M., Kaufmann, S., Geoffroy, P., & Fritig, B. (1987). Biological functions of ‘pathogenesis-related’ proteins: four tobacco PR proteins are chitinases. Proceedings of the National Academy of Sciences, 84(19), 6750–6754.
Grover, A. (2012). Plant chitinases: genetic diversity and physiological roles. Critical Reviews in Plant Sciences, 31(1), 57–73.
Collinge, D. B., Kragh, K. M., Mikkelsen, J. D., Nielsen, K. K., Rasmussen, U., & Vad, K. (1993). Plant chitinases. The Plant Journal, 3(1), 31–40.
Santos, P., Fortunato, A., Ribeiro, A., & Pawalowski, K. (2008). Chitinases in root nodules. Plant Biotechnology Journal, 25(3), 299–307.
Berglund, L., Brunstedt, J., Nielsen, K. K., Chen, Z., Mikkelsen, J. D., & Marcker, K. A. (1995). A proline-rich chitinase from Beta vulgaris. Plant Molecular Biology, 27, 211–216.
Koga, D., Sasaki, Y., Uchiumi, Y., Hirai, N., Arakane, T., & Nagamatsu, Y. (1997). Purification and characterization of Bombyx mori chitinases. Insect Biochemistry and Molecular Biology, 27(8-9), 757–767.
Cretoiu, M. S., Kielak, A. M., Al-Soud, W. A., Sørensen, S. J., & van Elsas, J. D. (2012). Mining of unexplored habitats for novel chitinases - chiA as a helper gene proxy in metagenomics. Applied Microbiology and Biotechnology, 94(5), 1347–1358.
Berini, F., Morena, C., Aurora, M., Reguzzoni, M., Tettamanti, G., & Marinelli, F. (2019). Metagenome-sourced microbial chitinases as potential insecticide proteins. Frontiers in Microbiology, 10, 1358.
Monge, E. C., Tuveng, T. R., Vaaje-Kolstad, G., Eijsink, V. G. H., & Gardner, J. G. (2018). Systems analysis of the family glycoside hydrolase family 18 enzymes from Cellvibrio japonicas characterizes essential chitin degradation functions. The Journal of Biological Chemistry, 293(10), 3849–3859.
Horiuchi, A., Aslam, M., Kanai, T., & Atomi, H. A. (2016). Structurally novel chitinase from the chitin-degrading hyperthermophilic archaeon Thermococcus chitonophagus. Applied and Environmental Microbiology, 82, 3554–3562.
Xu, P., Ni, Z.-F., Zong, M.-H., Ou, X.-Y., Yang, J.-G., & Lou, W.-Y. (2020). The thermostability and activity of Paenibacillus pasadenensis chitinase through semi-rational design. International Journal of Biological Macromolecules, 150, 9–15.
Matroodi, S., Zamani, M., Haghbeen, K., Motallebi, M., & Aminzadeh, S. (2013). Physicochemical study of a novel chimeric chitinase with enhanced binding ability. Acta Biochimica et Biophysica Sinica, 45, 845–856.
Vieille, C., & Zeikus, G. J. (2001). Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiology and Molecular Biology Reviews, 65(1), 1–43.
Uriarte, M., Marina, A., Ramon-Maiques, S., Fita, I., & Rubio, V. (1999). The carbamoyl-phosphate synthetase of Pyrococcus furiosus is enzymologically and structurally a carbamate kinase. The Journal of Biological Chemistry, 274(23), 16295–16303.
Cacciapuoti, G., Fusco, S., Caiazzo, N., Zappia, V., & Porcelli, M. (1999). Heterologous expression of 5′-methylthioadenosine phosphorylase from the archaeon Sulfolobus solfataricus: Characterization of the recombinant protein and involvement of disulfide bonds in thermophilicity and thermostability. Protein Expression and Purification, 16, 125–135.
Matsumi, R., Manabe, K., Fukui, T., Atomi, H., & Imanaka, T. (2007). Disruption of a sugar transporter gene cluster in a hyperthermophilic archaeon using a host-marker system based on antibiotic resistance. Journal of Bacteriology, 189(7), 2683–2269.
Waege, I., Schmid, G., Thumann, S., Thomm, M., & Hausner, W. (2010). Shuttle vector-based transformation system for Pyrococcus furiosus. Applied and Environmental Microbiology, 76(10), 3308–3313.
Zeldes, B. M., Keller, M. W., Loder, A. J., Straub, C. T., Adams Michael, W. W., & Kelly, R. M. (2015). Extremely thermophilic microorganisms as metabolic engineering platforms for production of fuels and industrial chemicals. Frontiers in Microbiology, 6, 1209.
Duffner, F., Christodoulou, E. and Vorgias, C. E. (1996) Expression and correct folding of a thermostable chitinase in E. coli. In: First International Congress on Extremophiles, Estoril, Portugal, 1996.
Chernin, L. S., de la Fuente, L., Sobolev, V., Haran, S., Vorgias, C. E., Oppenheim, A. B., & Chet, I. (1997). Molecular cloning, structural analysis and expression in E. coli of a chitinase gene from Enterobacter agglomerans. Applied and Environmental Microbiology, 63(3), 834–839.
Lobo, M. D., Silva, F. D., Landim, P. G., et al. (2013). Expression and efficient secretion of a functional chitinase from Chromobacterium violaceum in Escherichia coli. BMC Biotechnology, 13, 46.
Sousa, A. J. S., Silva, C. F. B., Sousa, J. S., Monteiro Jr., J. E., Freire, J. E. C., Sousa, B. L., Lobo, M. D. P., Monteiro-Moreira, A. C. O., & Grangeiro, T. B. (2019). A thermostable chitinase from the antagonistic Chromobacterium violaceum that inhibits the development of phytopathogenic fungi. Enzyme and Microbial Technology, 126, 50–61.
Zhang, M., Puri, A. K., Govender, A., Wang, Z., Singh, S., & Permaul, K. (2015). The multi-chitinolytic enzyme system of the compost-dwelling thermophilic fungus Thermomyces lanuginosus. Process Biochemistry, 50, 237e244.
Yang, S., Fu, X., Yan, Q., Guo, Y., Liu, Z., & Jiang, Z. (2016). Cloning, expression, purification and application of a novel chitinase from a thermophilic marine bacterium Paenibacillus barengoltzii. Food Chemistry, 1, 1041–1048.
Fu, X., Yan, Q., Yang, S., Yang, X., Guo, Y., & Jiang, Z. (2014). An acidic, thermostable exochitinase with β-N-acetylglucosaminidase activity from Paenibacillus barengoltzii converting chitin to N-acetyl glucosamine. Biotechnology for Biofuels, 10, 174.
Yan, Q., & Fong, S. S. (2018). Cloning and characterization of a chitinase from Thermobifida fusca reveals Tfu_0580 as a thermostable and acidic endochitinase4. Biotechnology Reports (Amsterdam, Netherlands), 19, e0027.
Dua, A., Joshi, S., & Satyanarayana, T. (2017). Recombinant exochitinase of the thermophilic mould Myceliopthora thermophila BJA: Characteristics and utility in generating N-acetyl glucosamine and in biocontrol of phytopathogenic fungi. Biotechnology Progress, 33(1), 70–80.
Prasad, M., & Palanivelu, P. (2012). Overexpression of a chitinase gene from the thermophilic fungus, Thermomyces lanuginosus in Saccharomyces cerevisiae and characterization of the recombinant chitinase. Microbial Technology and Biochemical Technology Journals, 4, 3.
Mougiakos, I., Mohanraju, P., Bosma, E. F., Vrouwe, V., Finger Bou, M., Naduthodi, M. I. S., Gussak, A., Brinkman, R. B. L., van Kranenburg, R., & van der Oost, J. (2017). Characterizing a thermostable Cas9 for bacterial genome editing and silencing. Nature Communications, 8(1), 1647.
Karam, E. A., Wahab, W. A. A., Saleh, S. A. A., Hassan, M. E., Kansoh, A. L., & Esawy, M. A. (2017). Production, immobilization and thermodynamic studies of free and immobilized Aspergillus awamori amylase. International Journal of Biological Macromolecules, 102, 694–703.
Mostafa, F. A., Abdel Wahab, W. A., Salah, H. A., Nawwar, G. A. M., & Esawy, M. A. (2018). Kinetic and thermodynamic characteristic of Aspergillus awamori EM66 levansucrase. International Journal of Biological Macromolecules, 119, 232–239.
Kumar, S., Tsai, C-J.and Nussinov, R. (2000). Factor’s enhancing protein stability. Protein. Engin.13, 179-191.
Sadeghi, M., Naderi-Manesh, H., Zarrabi, M., & Ranjbar, B. (2006). Effective factors in thermostability of thermophilic proteins. Biophysical Chemistry, 119(3), 256–270.
Farinas, E. T., Schwaneberg, U., Glieder, A., & Arnold, F. H. (2001). Directed evolution of a cytochrome P450 monooxygenase for alkane oxidation. Advanced Synthesis and Catalysis, 343(6-7), 601–606.
Stephens, D. E., Khan, F. I., Singh, P., Bisetty, K., Singh, S., & Permaul, K. (2014). Creation of thermostable and alkaline stable xylanase variants by DNA shuffling. Journal of Biotechnology, 187, 139–146.
Fan, Y., Fang, W., Xiao, Y., Yang, X., Zhang, Y., Bidochka, M. J., & Pei, Y. (2007). Directed evolution for increased chitinase activity. Applied Microbiology and Biotechnology, 76, 135–139.
Songsiriritthigul, C., Pesatcha, P., Eijsink, V. G., & Yamabhai, M. (2009). Directed evolution of a Bacillus chitinase. Biotechnology Journal, 4(4), 501–509.
Burg, B. V. D., & Eijsink, V. G. (2002). Selection of mutations for increased protein stability. Current Opinion in Biotechnology, 13(4), 333–337.
Chen, L., Jiang, H., Cheng, Q., Chen, J., Wu, G., Kumar, A., Sun, M., & Liu, Z. (2015). Enhanced nematicidal potential of the chitinase pachi from Pseudomonas aeruginosa in association with Cry21Aa. Scientific Reports, 5(1), 14395.
Chen, J., An, Y., Kumar, A., & Liu, Z. (2017). Improvement of chitinase Pachi with nematicidal activities by random mutagenesis. International Journal of Biological Macromolecules, 96, 171–176.
Hardt, M., & Laine, R. A. (2004). Mutation of active site residues in the chitin-binding domain ChBDChiA1 from chitinase A1 of Bacillus circulans alters substrate specificity: Use of a green fluorescent protein binding assay. Archives of Biochemistry and Biophysics, 426(2), 286–297.
Gåseidnes, S., Synstad, B., Jia, X., Kjellesvik, H., Vriend, G., & Eijsink, V. G. H. (2003). Stabilization of a chitinase from Serratia marcescens by Gly Ala and Xxx Pro mutations. Protein Engineering, 16, 841–846.
Lin, F. P., Chen, H. C., & Lin, C. S. (1999). Site-directed mutagenesis of Asp313, Glu315, and Asp391 residues in chitinase of Aeromonas caviae. IUBMB Life, 48(2), 199–204.
Suginta, W., Songsiriritthigul, C., Kobdaj, A., Opassiri, R., & Svasti, J. (2007). Mutations of Trp275 and Trp397 altered the binding selectivity of Vibrio carchariae chitinase A. Biochimica et Biophysica Acta, 1770(8), 1151–1160.
Cai, W., Sha, L., Zhou, J., Huang, Z., & Guan, X. (2009). Functional analysis of active site residues of Bacillus thuringiensis WB7 chitinase by site-directed mutagenesis. World Journal of Microbiology and Biotechnology, 25(12), 2147–2155.
Alam, A., Takaguchi, Y., & Tsuboi, S. (2005). Simple, Extremely fast, and high-yielding oxidation of thiols to disulfides. Synthetic Communications, 35(10), 1329–1333.
Mohammadi, M., Sakhteman, A., Ahrari, S., Hassanpour, K., Hashemi, S. E., & Farnoosh, G. (2018). Disulfide bridge formation to increase thermostability of DFPase enzyme: A computational study. Computational Biology and Chemistry, 77, 272–278.
Neeraja, C., Subramanyam, R., Moerschbacher, B. M., & Podile, A. R. (2010b). Swapping the chitin-binding domain in Bacillus chitinases improves the substrate binding affinity and conformational stability. Molecular BioSystems, 6, 1492–1502.
Longo, M. A., Novella, I. S., Gracia, L. A., & Díaz, M. (1992). Diffusion of proteases in calcium alginate beads. Enzyme and Microbial Technology, 14(7), 586–590.
Prasad, M., & Palanivelu, P. (2015). Immobilization of a thermostable, fungal recombinant chitinase on biocompatible chitosan beads and the properties of the immobilized enzyme. Biotechnology and Applied Biochemistry, 62(4), 523–529.
Bhushan, B. (2000). Production and characterization of a thermostable chitinase from a new alkalophilic Bacillus sp. BG-11. Journal of Applied Microbiology, 88(5), 800–808.
Abassi, S., Emtiazi, G., Hosseini-Abari, A., & Kim, B. G. (2020). Chitooligosaccharides and thermostable chitinase against vulvovaginal candidiasis and saprophyte fungi: LC mass studies of shrimp shell fermentation by Bacillus altitudinis. Current Microbiology, 77(1), 40–48.
Okongo, R. N., Puri, A. K., Wang, Z., Singh, S., & Permaul, K. (2019). Comparative biocontrol ability of chitinases from bacteria and recombinant chitinases from the thermophilic fungus Thermomyces lanuginosus. Journal of Bioscience and Bioengineering, 127(6), 663–671.
Ogawa, K. (1994). Approaches to hybridization and applied genetics by protoplast fusion: Koji-molds and mushrooms. In Y. Murooka & T. Imanaka (Eds.), Recombinant microbes for industrial and agricultural applications (pp. 581–603). New York: Marcel Dekker, Inc..
Patidar, P., Agrawal, D., Banerjee, T., & Patil, S. (2005). Optimisation of process parameters for chitinase production by soil isolates of Penicillium chrysogenum under solid substrate fermentation. Process Biochemistry, 40(9), 2962–2967.
Nampoothiri, K. M., Baiju, T. V., Sandhya, C., Sabu, A., Szakacs, G., & Pandey, A. (2004). Process optimization for antifungal chitinase production by Trichoderma harzianum. Process Biochemistry, 39(11), 1583–1590.
Ramírez-Coutiño, L., Marín-Cervantes, M. C., Revah, S., & Shirai, K. (2006). Enzymatic hydrolysis of chitin in the production of oligosaccharides using Lecanicillium fungicola chitinases. Process Biochemistry, 41(5), 1106–1110.
Dahiya, N., Tewari, R., & Hoondal, G. S. (2006). Biotechnological aspects of chitinolytic enzymes: A review. Applied Microbiology and Biotechnology, 71, 773–782.
Acknowledgments
Aravind Madhavan and KB Arun acknowledge the Department of Health Research, Ministry of Health and Family Welfare for sanctioning a project under Young Scientist Scheme. Raveendran Sindhu acknowledges the Department of Science and Technology for sanctioning projects under DST WOS-B scheme.
Funding
Gincy Marina Mathew received financial support from the Kerala State Council for Science, Technology and Environment under the “Back-to-Lab” Post-Doctoral Fellowship Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mathew, G.M., Madhavan, A., Arun, K.B. et al. Thermophilic Chitinases: Structural, Functional and Engineering Attributes for Industrial Applications. Appl Biochem Biotechnol 193, 142–164 (2021). https://doi.org/10.1007/s12010-020-03416-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-020-03416-5