Abstract
To investigate the roles of the active site residues in the catalysis of Bacillus thuringiensis WB7 chitinase, twelve mutants, F201L, F201Y, G203A, G203D, D205E, D205N, D207E, D207N, W208C, W208R, E209D and E209Q were constructed by site-directed mutagenesis. The results showed that the mutants F201L, G203D, D205N, D207E, D207N, W208C and E209D were devoid of activity, and the loss of the enzymatic activities for F201Y, G203A, D205E, W208R and E209Q were 72, 70, 48, 31 and 29%, respectively. The pH-activity profiles indicated that the optimum pH for the mutants as well as for the wildtype enzyme was 8.0. E209Q exhibited a broader active pH range while D205E, G203A and F201Y resulted in a narrower active pH range. The pH range of activity reduced 1 unit for D205E, and 2 units for G203A and F201Y. The temperature-activity profiles showed that the optimum temperature for other mutants as well as wildtype enzyme was 60°C, but 50°C for G203A, which suggested that G203A resulted in a reduction of thermostability. The study indicated that the six active site residues involving in mutagenesis played an important part in WB7 chitinase. In addition, the catalytic mechanisms of the six active site residues in WB7 chitinase were discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus thuringiensis (B. thuringiensis), a well-known entomopathogen, is a ubiquitous gram-positive spore-forming bacterium (Bravo et al. 2007; Grossi-de-Sa et al. 2007; Schnepf et al. 1998). It is considered as one of the most successful biopesticides that are widely used for biological control in agriculture and forestry. It produces insecticidal crystal proteins (ICPs) during the sporulation phase, which exhibit highly specific toxicity to a wide variety of insect orders as well as to other invertebrates (Bravo 1997; Chen et al. 2007; Fernández et al. 2008; Hofte and Whiteley 1989; Reyes-Ramírez and Ibarra 2008; Yu et al. 2007). However, with the extensive utilization of B. thuringiensis preparation, a variety of insect species have shown a high potential for the development of resistance to ICPs, which would greatly threaten the further popularization of B. thuringiensis formulations (Candas et al. 2003; Ferre and Van Rie 2002; Janmaat and Myers 2003; Janmaat et al. 2004; Yang et al. 2007). Therefore, alternative methods to improve the insecticidal activity of B. thuringiensis are urgently needed.
Chitinases (EC 3.2.1.14) are glycosyl hydrolases which catalyze the hydrolysis of chitin, the major constituent presenting in the cuticle and gut peritrophic membrane of insects (Hashimoto et al. 2000a, b; Lonhienne et al. 2001; Merzendorfer and Zimoch 2003; Orikoshi et al. 2003). They are widely distributed in microoganisms (e.g., fungi and bacteria), plants and animals (Bhattacharya et al. 2007; Souza et al. 2003). Generally, chitinases are classified into families 18 and 19 of glycosyl hydrolases according to the amino acid sequence similarity of their catalytic domains (Henrissat and Bairoch 1993). Family 18 includes chitinases from bacteria, fungi, animals, and certain plants, while family 19 consists of chitinases from plants. Family 18 chitinases usually contain multiple functional domains, such as signal peptide, catalytic domain, fibronectin typeIII-like domain and chitin-binding domain (ChBD) (Lonhienne et al. 2001). For decades, chitinases have received increasing attentions because of their potential applications in biocontrol of chitin-containing insects and other phytopathogenic fungi (Souza et al. 2003). Studies indicated that chitinases could improve the insecticidal activity of B. thuringiensis when added in B. thuringiensis preparation or when co-expressed with ICPs (Downing et al. 2000; Smirnoff 1971; Tantimavanich et al. 1997; Wiwat et al. 2000). To investigate the roles of the amino acid residues of the chitinase in the hydrolysis of chitin, site-directed mutagenesis was carried out to the putative active site residues of B. thuringiensis WB7 chitinase.
Family 18 chitinases have two highly conserved sequences, where putative active site residues are considered to be located (Kramer and Muthukrishnan 1997). One of the regions is particularly abundant in acidic amino acids and has the highly conserved sequence of FDGLD LDWEYP (Lu et al. 2002). Based on previous studies of chitinase active site residues from other members of this family, they might be involved in catalysis and/or substrate binding (Aronson et al. 2006; Lin et al. 1999; Lu et al. 2002; Thomas et al. 2000; Watanabe et al. 1993, 1994; Zhang et al. 2002).
In this study, six amino acid residues, namely phenylalanine 201 (F201), glycin 203 (G203), aspartic acid 205 (D205), aspartic acid 207 (D207), tryptophan 208 (W208) and glutamine 209 (E209), in the conserved region of chitinase from B. thuringiensis WB7, were substituted with other amino acids by site-directed mutagenesis. Twelve mutants, as well as the wildtype enzyme, were expressed in Escherichia coli (E. coli) BL21 (DE3) using pET-29a as the expression vector.
Materials and methods
Strains and plasmids
Bacillus thuringiensis WB7 used in this study is preserved in our laboratory. E. coli JM109 and pUC18 vector (Stratagene) were used for recombinant DNA cloning. E. coli XL1-Blue (Stratagene) was used as the host for the construction of mutants. E. coli BL21 (DE3) and pET-29a vector (Novagen) were used for the expression of chitinase from both the wildtype and mutant strains.
Construction of recombinant plasmid pUC-Chi
A pair of primers for amplification of the ORF of chi gene was designed based on the sequence deposited at GenBank (Accession Number: AY074882). Primer sequences were: Chi-F, 5′-ggAATTCAtggCTAtgAggTCTC-3′ (the bases underlined show EcoRI site) and Chi-R, 5′-CCCAAgCTTCTAgTTTTCgCTAATg-3′ (the bases underlined show HindIII site). Genomic DNA from B. thuringiensis WB7 was used as the template for PCR amplification with the primer pair Chi-F/Chi-R. The PCR product was then cloned into the pUC18 vector to obtain the recombinant plasmid pUC-Chi, and the selected clones were verified by DNA sequencing.
Site-directed mutagenesis
QuikChange Site-Directed Mutagenesis Kit (Stratagene) was used in this study. Twelve pairs of primers for mutagenesis were designed according to the protocol of QuikChange Site-Directed Mutagenesis Kit (Table 1). The recombinant plasmid pUC-Chi was used as the template for PCR amplification to obtain the mutated sequences. The PCR products were transformed into E. coli XL1-Blue after digested with DpnI. The mutants were then verified by DNA sequencing and were denominated as pUC-F201L, pUC-F201Y, pUC-G203A, pUC-G203D, pUC-D205E, pUC-D205N, pUC-D207E, pUC-D207N, pUC-E209D, pUC-E209Q, pUC-W208C and pUC-W208R, respectively.
Expression of chitinase and its mutant forms in E. coli BL21 (DE3)
From the plasmids that were confirmed by DNA sequencing to have the desired mutations, the chitinase gene coding fragments were cut out using EcoRI and HindIII and then inserted into pET-29a vector digested with the same restriction enzymes. The recombinant plasmids were then transformed into E. coli BL21 (DE3) and were denominated as pET-Chi, F201L, F201Y, G203A, G203D, D205E, D205N, D207E, D207N, E209D, E209Q, W208C and W208R, respectively. Protein expression was then carried out at 37°C in LB broth supplemented with kanamycin (50 μg/ml) and induced by isopropyl-β-d-thiogalacto-pyranoside (IPTG; 1 mM) as the culture reached an optical density at 600 nm of 0.6–0.8. After 24 h induction, proteins secreting into the medium were extracted by the trichloroacetic acid (TCA) precipitation method. Samples were boiled in the loading buffer for 5 min, and the mixtures were then analyzed by SDS–PAGE.
Enzyme assay
Enzymatic activity was determined using chitin colloid as the substrate. After induced by IPTG at 37°C for 24 h, culture solution was centrifuged at 10,000×g for 5 min at 4°C to remove cell debris. 1 ml culture supernatant was mixed with 1 ml phosphate buffer solution (PBS, pH 7.0) containing 1% chitin colloid. The mixture was then incubated at 37°C for 1 h. To stop the reaction, 2 ml 3,5-dinitrosalicylic acid (DNS: 20% potassium sodium tartrate, 1% NaOH, 0.2% phenol, 0.05% Na2SO3 and 1% DNS) was added and the mixture was boiled for 10 min. Then the reaction mixture was centrifuged at 12,000×g for 5 min and the absorbance of the supernatant was measured at 530 nm. The enzyme activity unit was defined as ΔNAG/μg/h.
The effect of pH on enzymatic activity was measured over the range of pH 4–10 at 37°C by using chitin colloid as the substrate. The effect of temperature on enzymatic activity was determined at pH 7.0 and following a procedure similar to that for measuring the pH-profile except for using temperature as the variable. The temperature ranged from 30 to 70°C.
Chitin colloidal degradation assay
Twelve mutants and the wildtype enzyme, as well as two controls of E. coli BL21(DE3) and E. coli BL21(DE3) harboring pET-29a, were inoculated on chitin colloidal plates covered with a layer of IPTG. The transparent circles of hydrolysis were observed after incubation of the plates at 37°C for 4–5 days.
Results
Sequence analysis of chi gene from B. thuringiensis WB7
The entire coding region of the chi gene was produced by PCR using Chi-F/Chi-R as the primers. The PCR product was then cloned into pUC18 vector and verified by DNA sequencing. It was 2,025 bp in length, encoding a protein of 674 amino acids with a calculated molecular weight of 74 kDa and a predicted pI value of 5.94. Blast analysis indicated that the DNA sequence of chi from B. thuringiensis WB7 had a high similarity of 98% to that of B. cereus ATCC 14579 (AE016877.1), B. thuringiensis serovar canadensis strain HD224 (AY455290.1), and B. thuringiensis BT010 (EU030 625.1).
Conserved Domains Search analysis showed that the chitinase from B. thuringiensis WB7 belonged to family 18 and consisted of three functional domains, including catalytic domain, fibronectin typeIII-like domain and chitin-binding domain. Signal peptide analysis by SignalP V2.0 and TMHMM2.0 Software indicated that a signal peptide present at the N-terminal of the protein, in which the most likely cleavage site was between Ala-32 and Asp-33. Analysis by ScanProsite revealed that the region from F201 to E209 (FDGVDLDWE) was the active center of the enzyme, which displayed high similarity to that of other members in family 18 chitinases (Table 2) (Lu et al. 2002).
Three-dimensional structure analysis
The three-dimensional structure of catalytic domain of WB7 chitinase was modelled by Homology Modelling method using Swiss Model Server. And the known tertiary structure of chitinase A1 (ChiA1) from B. circulans WL-12 (PDB: 1itx; Chain: A), which shared an approximately 57% amino acid identity with WB7 chitinase, was used as the template. The predicted tertiary structure of WB7 chitinase was then visualized using Accelrys Discovery Studio 2.1. The results showed that the catalytic domain of WB7 chitinase was composed of (α/β)8-TIM-barrel, β-domain 1 and β-domain 2. The two smallβ-domains were separately located on top of the TIM-barrel, which provided a deep cleft for substrate binding (Takuo et al. 1999).
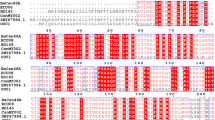
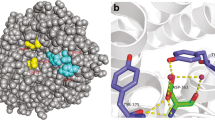
Structural comparison indicated that the catalytic domain of WB7 chitinase was well superimposed on that of B. circulans ChiA1 (Fig. 1a). The conserved region FDGVDLDWE (residues 201–209) of WB7 chitinase was also fully overlapped with that (residues 196–204) of B. circulans ChiA1 (Fig. 1b). Similar to other family 18 chitinases, the conserved region FDGVDLDWE of WB7 chitinase approximately corresponded to the amino acid residues forming the fourth β-strand of the TIM-barrel (Takuo et al. 1999; Terwisscha van Scheltinga et al. 1996; Perrakis et al. 1994). Due to the high conservation of the region FDGVDLDWE in amino acid sequence and spacial structure, they were believed to be important for stabilization of the barrel. To study the roles of the conserved amino acid residues performed in B. thuringiensis WB7 chitinase, six amino acid residues F201, G203, D205, D207, W208, E209 were substituted with other amino acids by site-directed mutagenesis.
Structural comparison of the chitinases between B. thuringiensis WB7 and B. circulans WL-12. a Superimposition of the overall structures of WB7 chitinase (white) and WL-12 chitinase (red). The white ball indicated the residue E209 in WB7 chitinase, while the red ball indicated the catalytic residue E204 in WL-12 chitinase. b Superimposition of the conserved regions FDGVDLDWE between WB7 chitinase and WL-12 chitinase (Color figure online)
Expression of chitinase in E. coli BL21 (DE3)
Twelve mutants, including F201L, F201Y, G203A, G203D, D205E, D205N, D207E, D207N, E209D, E209Q, W208C and W208R, were generated as described in “Materials and Methods” section. The mutants as well as the wildtype protein induced with IPTG were then identified by SDS–PAGE (Fig. 2). The results showed that the molecular weight of the expressed proteins were about 70 kDa, which were less than 74 kDa. The reason might be that the chitinase from B. thuringiensis WB7 was an extracellular enzyme, whose signal peptide was excised while the protein spanned the cell membrane, leading to the reduction of the molecular weight. The signal peptide was about 4 kDa, so the 70 kDa band probably was the supposed target protein. Meanwhile, the results indicated that the signal peptide from B. thuringiensis WB7 chitinase could be recognized by E. coli BL21 (DE3).
SDS–PAGE analysis of the chitinase produced in E. coli BL21(DE3). a Lane 1 ■pET-29a, lane 2 □pET-Chi, lane 3 ■pET-Chi, lane 4 □F201L, lane 5 ■F201L, lane 6 □F201Y, lane 7 ■F201Y, lane 8 TaKaRa Protein MW Marker (Broad). b Lane 1 ■pET-29a, lane 2 □G203A, lane 3 ■G203A, lane 4 □G203D, lane 5 ■G203D, lane 6 □D205E, lane 7 ■D205E, lane 8 □D205N, lane 9 ■D205N, lane 10 □D207E, lane 11 ■D207E, lane 12 TaKaRa Protein MW Marker (Broad). c Lane 1 ■pET-29a, lane 2 □D207N, lane 3 ■D207N, lane 4 □W208C, lane 5 ■W208C, lane 6 □W208R, lane 7 ■W208R, lane 8 □E209D, lane 9 ■E209D, lane 10 □E209Q, lane 11 ■E209Q, lane 12 TaKaRa Protein MW Marker (Broad). (“□”indicated the protein without IPTG induction, “■”indicated the protein induced with IPTG)
Enzyme assay
The detection of enzymatic activity was carried out under the condition of 37°C and pH 7.0 using chitin colloid as the substrate. The results demonstrated that the mutants F201L, G203D, D205N, D207E, D207N, W208C and E209D were devoid of activity, and the loss of the enzymatic activities for F201Y, G203A, D205E, W208R and E209Q were 72, 70, 48, 31 and 29%, respectively (Fig. 3).
The effect of pH on enzymatic activity was determined over the range of pH 4–10 at 37°C. The results indicated that the mutants, including F201L, G203D, D205N, D207E, D207N, W208C and E209D, were devoid of activity over the entire range of pH values tested. The pH-activity profiles of F201Y, G203A, D205E, W208R and E209Q were similar to that of the wildtype enzyme pET-Chi with highest activity at pH 8.0. However, D205E, F201Y and G203A exhibited a narrowing of active pH range, which reduced about 1 unit for D205E, and about 2 units for F201Y and G203A. E209Q showed a broader active pH range than that of the wildtype enzyme (Fig. 4).
The effect of temperature on enzymatic activity was investigated using chitin colloid as the substrate over the range of temperature 30–70°C at pH 7.0. It showed that the mutants, including F201L, G203D, D205N, D207E, D207N, W208C and E209D, had no detectable activity over the entire temperature range. The temperature-activity profiles of F201Y, D205E, W208R and E209Q were similar to that of the wildtype enzyme, all of which exhibited highest activity at 60°C. G203A showed highest activity at 50°C, but the activity sharply dropped as the temperature rose to 60oC, which indicated that the thermostability of G203A declined obviously (Fig. 5).
Chitin colloidal degradation assay
The mutants and the wildtype strain, as well as the controls of E. coli BL21 (DE3) and E. coli BL21 (DE3) harboring pET-29a were inoculated on chitin colloidal plates coated with IPTG. The plates were then placed in the incubator at 37°C. Five days later, transparent circles of hydrolysis were clearly observed around the mutants of F201Y, G203A, D205E, E209Q and W208R as well as the wildtype pET-Chi, but could not be found around other strains (Fig. 6). It revealed that the activity of the wildtype enzyme pET-Chi and the mutants F201Y, G203A, D205E, E209Q and W208R, could be detected on chitin colloidal plates. This result corresponded with that of enzymatic analysis.
Discussion
Based on the amino acid sequence identity (57%) and high similarity in tertiary structure, it was proposed that WB7 chitinases and B. circulans ChiA1 might share a similarity in their catalytic reaction mechanism. Studies indicated that B. circulans ChiA1 hydrolyzed scissile glycosidic bonds via “substrate-assisted catalysis” mechanism. In the mechanism, the glutamic acid residue (corresponding to E204 of B. circulans ChiA1) located at the end of the fourth β-strand of the barrel serves as a catalytic acid. The acetyl group of the substrate acts as the nucleophile and stabilizes the positive charge at C1 of the GlcNAc residue by forming an oxazolinium ion intermediate. Substrate-assisted catalysis has recently been generally accepted as the catalytic mechanism for family 18 chitinases (Aronson et al. 1997; Takuo et al. 1999; Hashimoto et al. 2000a, b).
In this paper, twelve mutants were constructed by site-directed mutagenesis to investigate the roles of the conserved amino acid residues in the catalysis of B. thuringiensis WB7 chitinase. The pH-activity profiles indicated that all the mutants as well as the wildtype enzyme showed higher activities under alkaline conditions, and exhibited very low activity or even inactivity under acid conditions. The chitinases from Manduca sexta, Bombyx mori and Bacillus sp. BG-11 were similar to that of this study, showing higher activity in the alkaline conditions (Zhang et al. 2002). However, most of the chitinases from plants, bacteria and animals had approximately optimum pH at 6.0, which would rapidly deactivate as the pH was beyond 10.0 (Zhang et al. 2002).
The mutant F201Y resulted in a loss of activity of 72% and a narrowing of active pH range. Both phenylalanine (F) and tyrosine (Y) are hydrophobic aromatic amino acid, whose structures and characteristics are close to each other. However, tyrosine could not completely replace phenylalanine at position 201 in B. thuringiensis WB7 chitinase, for the substitution of F201 for Y caused a significant decrease in enzymatic activity and a narrowing of active pH range. The side-chain and spatial structure of leucine (L) are quite different from those of phenylalanine. Maybe that was the reason for the inactivity of F201L. It was supposed that F201 might participate in substrate binding, but its exact role in B. thuringiensis WB7 chitinase needs to be investigated with further studies.
G203A caused a loss of 70% enzymatic activity and a reduction of thermostability while G203D was devoid of activity. Glycin(G) is the smallest and simplest amino acid of the 20 kinds of amino acids. It plays a unique role in maintaining the conformation of protein, because it can be filled in the gap which can not accommodate other amino acids for its simplest structure and smallest side-chain. The four amino acids, alanine (A), valine (V), leucine (L) and isoleucine (I), are devoid of active functional groups, but they play an important role in establishing and maintaining the three dimensional structure of the protein, because they trend to gather together to keep away from water due to their strongly hydrophobic side-chains. Maybe just for this point, alanine (A) is similar to glycin (G), thus G203A retained 30% of activity. Aspartic acid (D), a hydrophilic dicarboxylic amino acid with a negative charge, often appears on the surface of the protein. It is a very active amino acid whose structure and side-chain are much more complicated than that of glycin. Provided that G203 did take part in maintaining the three dimensional structure in B. thuringiensis WB7 chitinase, the substitution of glycin (G) for aspartic acid (D) probably destroy the structure of the protein, thus affecting the function of the protein. However, the actual role of G203 in WB7 chitinase needs to be further studied.
Mutant E209Q retained most of the enzymatic activity (71%) while E209D was devoid of activity, which were slightly different from the results in corresponding sites of other family 18 chitinases. Both mutants E204Q and E204D in B.circulans WL-12 caused inactivity, which were similar to the relevant mutants of E146Q and E146D in M. sexta chitinase (Lu et al. 2002; Watanabe et al. 1993). Glutamic acid (E) is an acid amino acid with a negative charge. Due to the high similarity in tertiary structure between WB7 chitinase and B. circulans ChiA1, it was supposed that E209 might play a role of catalytic acid in the “substrate-assisted catalysis” mechanism, which was similar to that of the corresponding residue E204 in B. circulans ChiA1 (Takuo et al. 1999; Hashimoto et al. 2000a, b).
It was suggested that D207 played an exclusive role in the catalysis of B. thuringiensis WB7 chitinase, because both mutants D207E and D207N were devoid of activity. The structure and characteristic of glutamic acid (E) are very similar to those of aspartic acid (D). However, the substitution of glutamic acid (E) for aspartic acid (D) still led to inactivity of D207E, which indicated that the carboxyl group of D207 is essential for catalysis. Similar results were obtained from B. circulans WL-12 chitinase with the mutants of D202E and D202N (Watanabe et al. 1994). Whereas, the results from M. sexta chitinase were slightly different from this paper. In that study, the relevant mutant D144E retained 70% enzymatic activity, while D144N lost most of the activity (Lu et al. 2002). D207 was supposed to be ionized at reaction pHs, and the negative charge it carried formed an ion-pair to help stabilize the positively charged oxazoline ion intermediate in the course of substrate-assisted catalysis. D207 then continued to promote the reaction to further stabilize the transition state (Aronson et al. 2006; Lu et al. 2002; Hashimoto et al. 2000a, b).
The studies showed that D205E retained 52% enzymatic activity while D205N was inactive under any conditions, which were similar to the results obtained from the B. circulans WL-12 chitinase. In that study, the mutant D200E retained most of the activity while D200N lost most of the enzymatic activity (Watanabe et al. 1993). Conversely, in the chitinase of Aeromonas caviae, the mutant D313E was inactive while D313N retained 50% activity (Lin et al. 1999). The mutant D142E in the M. sexta chitinase had nearly the same activity as the wildtype enzyme, and D142N led to a measurable increase in activity while using oligosaccharide as the substrate (Lu et al. 2002). It is supposed that there is a hydrogen bond between D205 and D207, which may have an effect on the ionization state of D207, and then influences the ionization state of active center (Lu et al. 2002). Therefore, the mutations of D205 probably destroyed the hydrogen bond, then subsequently destroyed the ionization state of the active center, thus affecting the enzymatic activity.
W208R retained 67% enzymatic activity while W208C was devoid of activity. Tryptophan (W), with a double-ring indole in the side-chain, is the largest residue in the 20 kinds of amino acids. Studies on lysozyme and some plant chitinases indicated that tryptophan might participate in substrate binding by hydrogen-bonding with substrate via the indole NH-group and by a stacking interaction with sugar rings of the substrate (Zhang et al. 2002). Given that the tryptophan did involve in interactions with sugars by stacking interactions and hydrogen bonding, the mutagenesis carried out in the residue of W208 may affect the recognition and binding between the chitinase and the substrate, and subsequently influenced the enzymatic activity.
In this paper, twelve mutants were successfully constructed to investigate the roles of the conserved amino acid residues in the catalysis of B. thuringiensis WB7 chitinase. The results indicated that the six conserved amino acid residues involving in mutagenesis played an important role in B. thuringiensis WB7 chitinase, for their mutants showed varying degrees of destruction in enzymatic activity. However, the actual roles of the corresponding amino acid residues in WB7 chitinase need to be confirmed with further work.
References
Aronson NN, Blanchard CJ, Madura JD (1997) Homology modeling of glycosyl hydrolase family 18 enzymes and proteins. J Chem Inf Comput Sci 37:999–1005
Aronson NN Jr, Halloran BA, Alexeyev MF, Zhou XE, Wang Y, Meehan EJ, Chen L (2006) Mutation of a conserved tryptophan in the chitin-binding cleft of Serratia marcescens chitinase A enhances transglycosylation. Biosci Biotechnol Biochem 70:243–251
Bhattacharya D, Nagpure A, Gupta RK (2007) Bacterial chitinases: properties and potential. Crit Rev Biotechnol 27:21–28
Bravo A (1997) Phylogenetic relationships of Bacillus thuringiensis delta-endotoxin family proteins and their functional domains. J Bacteriol 179:2793–2801
Bravo A, Gill SS, Soberón M (2007) Mode of action of Bacillus thuringiensis Cry and Cyt toxins and their potential for insect control. Toxicon 49:423–435
Candas M, Loseva O, Oppert B, Kosaraju P, Bulla LA Jr (2003) Insect resistance to Bacillus thuringiensis alterations in the indianmeal moth larval gut proteome. Mol Cell Proteomics 2:19–28
Chen J, Hua G, Jurat-Fuentes JL, Abdullah MA, Adang MJ (2007) Synergism of Bacillus thuringiensis toxins by a fragment of a toxin-binding cadherin. Proc Natl Acad Sci U S A 104:13901–13906
Downing KJ, Leslie G, Thomson JA (2000) Biocontrol of the sugarcane borer Eldana saccharina by expression of the Bacillus thuringiensis cry1Ac7 and Serratia marcescens chiA genes in sugarcane-associated bacteria. Appl Environ Microbiol 66:2804–2810
Fernández LE, Gómez I, Pacheco S, Arenas I, Gilla SS, Bravo A, Soberón M (2008) Employing phage display to study the mode of action of Bacillus thuringiensis Cry toxins. Peptides 29:324–329
Ferre J, Van Rie J (2002) Biochemistry and genetics of insect resistance to Bacillus thuringiensis. Annu Rev Entomol 47:501–533
Grossi-de-Sa MF, Quezado de Magalhaes M, Silva MS, Silva SM, Dias SC, Nakasu EY, Brunetta PS, Oliveira GR, Neto OB, Sampaio de Oliveira R, Soares LH, Ayub MA, Siqueira HA, Figueira EL (2007) Susceptibility of Anthonomus grandis (cotton boll weevil) and Spodoptera frugiperda (fall armyworm) to a cry1Ia-type toxin from a Brazilian Bacillus thuringiensis strain. J Biochem Mol Biol 40:773–782
Hashimoto M, Honda Y, Nikaidou N, Fukamizo T, Watanabe T (2000a) Site-directed mutagenesis of Asp280 suggests substrate-assisted catalysis of chitinase A1 from Bacillus circulans WL-12. J Biosci Bioeng 89:100–102
Hashimoto M, Ikegami T, Seino S, Ohuchi N, Fukada H, Sugiyama J, Shirakawa M, Watanabe T (2000b) Expression and characterization of the chitin-binding domain of chitinase A1 from Bacillus circulans WL-12. J Bacteriol 182:3045–3054
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Hofte H, Whiteley HR (1989) Insecticidal crystal proteins of Bacillus thuringiensis. Microbiol Rev 53:242–255
Janmaat AF, Myers J (2003) Rapid evolution and the cost of resistance to Bacillus thuringiensis in greenhouse populations of cabbage loopers, Trichoplusia ni. Proc Biol Sci 270:2263–2270
Janmaat AF, Wang P, Kain W, Zhao JZ, Myers J (2004) Inheritance of resistance to Bacillus thuringiensis subsp. kurstaki in Trichoplusia ni. Appl Environ Microbiol 70:5859–5867
Kramer KJ, Muthukrishnan S (1997) Insect chitinases: molecular biology and potential use as biopesticides. Insect Biochem Mol Biol 27:887–900
Lin FP, Chen HC, Lin CS (1999) Site-directed mutagenesis of Asp313, Glu315, and Asp391 residues in chitinase of Aeromonas caviae. IUBMB Life 48:199–204
Lonhienne T, Mavromatis K, Vorgias CE, Buchon L, Gerday C, Bouriotis V (2001) Cloning, sequences, and characterization of two chitinase genes from the Antarctic Arthrobacter sp. strain TAD20: isolation and partial characterization of the enzymes. J Bacteriol 183:1773–1779
Lu Y, Zen KC, Muthukrishnan S, Kramer KJ (2002) Site-directed mutagenesis and functional analysis of active site acidic amino acid residues D142, D144 and E146 in Manduca sexta (tobacco hornworm) chitinase. Insect Biochem Mol Biol 32:1369–1382
Merzendorfer H, Zimoch L (2003) Chitin metabolism in insects: structure, function and regulation of chitin synthases and chitinases. J Exp Biol 206:4393–4412
Orikoshi H, Baba N, Nakayama S, Kashu H, Miyamoto K, Yasuda M, Inamori Y, Tsujibo H (2003) Molecular analysis of the gene encoding a novel cold-adapted chitinase (ChiB) from a marine bacterium, Alteromonas sp. strain O-7. J Bacteriol 185:1153–1160
Perrakis A, Tews I, Dauter Z, Oppenheim AB, Chet I, Wilson KS, Vorgias CE (1994) Crystal structure of a bacterial chitinase at 2.3 Å resolution. Structure 2:1169–1180
Reyes-Ramírez A, Ibarra JE (2008) Plasmid patterns of Bacillus thuringiensis type strains. Appl Environ Microbiol 74:125–129
Schnepf E, Crickmore N, Van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62:775–806
Smirnoff WA (1971) Effect of chitinase on the action of Bacillus thuringiensis. Can Entomol 103:1829–1831
Souza RF, Gomes RC, Coelho RR, Alviano CS, Soares RM (2003) Purification and characterization of an endochitinase produced by Colletotrichum gloeosporioides. FEMS Microbiol Lett 222:45–50
Takuo M, Takamasa N, Masayuki H, Takeshi W, Yukio M (1999) Three-dimensional structure of the catalytic domain of chitinase Al from Bacillus circulars WL-12 at a very high resolution. Proc Jpn Acad 75:269–274
Tantimavanich S, Pantuwatana S, Bhumiratana A, Panbangred W (1997) Cloning of a chitinase gene into Bacillus thuringiensis subsp. aizawai for enhanced insecticidal activity. J Gen Appl Microbiol 43:341–347
Terwisscha van Scheltinga AC, Hennig M, Dijkstra BW (1996) The 1.8 Å resolution structure of hevamine, a plant chitinase/lysozyme, and analysis of the conserved sequence and structure motifs of glycosyl hydrolase family 18. J Mol Biol 262:243–257
Thomas CJ, Gooday GW, King LA, Possee RD (2000) Mutagenesis of the active site coding region of the Autographa californica nucleopolyhedrovirus chiA gene. J Gen Virol 81:1403–1411
Watanabe T, Kobori K, Miyashita K, Fujii T, Sakai H, Uchida M, Tanaka H (1993) Identification of glutamic acid 204 and aspartic acid 200 in chitinase A1 of Bacillus circulans WL-12 as essential residues for chitinase activity. J Biol Chem 268:18567–18572
Watanabe T, Uchida M, Kobori K, Tanaka H (1994) Site-directed mutagenesis of the Asp-197 and Asp-202 residues in chitinase A1 of Bacillus circulans WL-12. Biosci Biotechnol Biochem 58:2283–2285
Wiwat C, Thaithanun S, Pantuwatana S, Bhumiratana A (2000) Toxicity of chitinase- producing Bacillus thuringiensis ssp. kurstaki HD-1 (G) toward Plutella xylostella. J Invertebr Pathol 76:270–277
Yang Y, Chen H, Wu Y, Yang Y, Wu S (2007) Mutated cadherin alleles from a field population of Helicoverpa armigera confer resistance to Bacillus thuringiensis toxin Cry1Ac. Appl Environ Microbiol 73:6939–6944
Yu X, Huang T, Huang Z, Powell CA, Guan X (2007) Expression and characterization of inhA gene from Bacillus thuringiensis 8010. World J Microbiol Biotechnol 23:1621–1625
Zhang H, Huang X, Fukamizo T, Muthukrishnan S, Kramer KJ (2002) Site-directed mutagenesis and functional analysis of an active site tryptophan of insect chitinase. Insect Biochem Mol Biol 32:1477–1488
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (30571257) and the Natural Science Foundation of Fujian Province (2007J0324).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cai, W., Sha, L., Zhou, J. et al. Functional analysis of active site residues of Bacillus thuringiensis WB7 chitinase by site-directed mutagenesis. World J Microbiol Biotechnol 25, 2147–2155 (2009). https://doi.org/10.1007/s11274-009-0119-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-009-0119-y