Abstract
Low-temperature-tolerant microorganisms and their cold-active enzymes could be an innovative and invaluable tool in various industrial applications. In the present study, bacterial isolates from the sediment samples of Kongsfjord, Norwegian Arctic, were screened for β-galactosidase production. Among the isolates, KS25, KS85, KS60, and KS92 have shown good potential in β-galactosidase production at 20 °C. 16SrRNA gene sequence analysis revealed the relatedness of the isolates to Enterobacter ludwigii. The optimum growth temperature of the isolate was 25 °C. The isolate exhibited good growth and enzyme production at a temperature range of 15–35 °C, pH 5–10. The isolate preferred yeast extract and lactose for the maximum growth and enzyme production at conditions of pH 7.0, temperature of 25 °C, and agitation speed of 100 rpm. The growth and enzyme production was stimulated by Mn2+ and Mg2+ and strongly inhibited by Zn2+, Ni2+, and Cu+. β-Galactosidases with high specific activity at low temperatures are very beneficial in food industry to compensate the nutritional problem associated with lactose intolerance. The isolate exhibited a remarkable capability to utilize clarified whey, an industrial pollutant, for good biomass and enzyme yield and hence could be well employed in whey bioremediation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Psychrophilic and psychrotrophic microorganisms are able to produce variety of technologically valuable cellular products such as cold-active enzymes that exhibit high catalytic activity at low temperature than the enzymes from their mesophilic counterparts [1]. Cold-active enzymes could have important industrial applications in various fields such as food processing, biomass conversion, molecular biology, environmental biosensors, and bioremediation [2]. Cold-active enzymes are thermolabile and are inactivated at moderate temperatures where the mesophilic enzymes are functional [3]. Practical utilization of cold-active enzymes will save energy consumptions, considerably. In food industry, the cold-active enzymes are in great demand because the use of these enzymes cuts down the process cost by reducing the investment in conventional cooling or heating treatments besides retaining the food quality with its own taste, flavor, and nutritional value [4]. The study of psychrotrophic microorganisms and their cold-active enzyme may lead to the discovery of the novel applications of these enzymes in various fields of biotechnology and simpler route for the synthetic processes of economically important metabolites.
Among cold-active enzymes, cold-active β-galactosidases (β-galactoside galactohydrolase, EC3.2.1.23), commonly known as lactases, is one of the most important food industrial enzyme. It catalyzes two specific reactions such as (i) the hydrolysis of the β (1 → 4) linkage of lactose to glucose and galactose, and (ii) transglycosylation reaction where a galactose molecule, by-product of lactose hydrolysis, is transferred to the galactose moiety of lactose to form galacto-oligosaccharides [5]. In dairy industry, cold-active β-galactosidases are primarily used for the production of lactose-free milk for lactose intolerant people. Besides eliminating the nutritional problem, the low-temperature optimum cold-active β-galactosidases offer some other important advantages such as (i) the lactose hydrolysis that can run during shipping and storage of milk that shorten the entire production process (save energy), (ii) eliminating the chance of mesophilic microflora contamination, and (iii) preventing the formation of nonenzymatic browning products formed at higher temperatures. Use of beta-galactosidase during milk processing is technologically important because lactose hydrolysis decreases the hydroscopicity of milk and suppresses lactose crystallization in ice cream and condensed milk. It can also be well employed for the conversion of lactose to easily fermentable sugars, glucose and galactose, in whey, and primary by-product of cheese industry and reduces the water pollution related to the dairy industry [6, 7].
In addition, the transglycosylation activity of β-galactosidase yields water soluble, nondigestible, and mildly sweet galacto-oligosaccharides which are widely used as food additives in infant milk powders, beverages, yogurts, and other dairy products for the enhancement of desirable flavors during food processing and as prebiotics to promote the growth of beneficial bifidobacteria in the large intestine [8, 9].
An ideal cold-active β-galactosidase for treating milk should work well at approximately 10 °C, be highly active at pH 6.7–6.8, not be inhibited by Na+, Ca2+ ions, galactose, and glucose, and be specific for lactose. Most of the currently applied β-galactosidases in food industry such as Maxilact, Lactozym, and Yeast neutral lactase (DYL) are mesophilic and are harvested from microorganisms such as Kluyveromyces lactis and Kluyveromyces fragilis, Validase fungal lactase concentrate (VFL) from Aspergillus oryzae which has an optimum temperature of approximately 35 to 50 °C and displays poor activity below 20 °C [10, 11]. Recently, a number of cold-active β-galactosidases from different microorganisms viz. Arthrobacter sp. [12, 13], Bacillus subtilis KL88, Carnobacterium piscicola BA, Pseudoalteromonas haloplanktis [14, 15], and from Planococcus [16] have been studied. In the present study, psychrotrophic bacteria from the sediment samples of Kongsfjord, Arctic, were screened for cold-active β-galactosidase production and identified using molecular methods. Effect of various physical and chemical parameters on the growth and cold-active β-galactosidase production by Enterobacter ludwigii KS85, which was to be a prospective strain, was also analyzed.
Materials and methods
Bacterial isolates, screening, and culture medium
Bacterial isolates (100 No.s) from the sediment samples of Kongsfgord, Arctic (KS1-KS100), were enriched in ZoBell’s Marine Broth at 20 °C. The primary screening of cold-active β-galactosidase producers among isolates was carried on modified LB agar plates containing 10 g peptone, 10 g yeast extract, 5 g lactose, 15 g agar, 20 g NaCl, and 40 mg of X-gal (5-bromo-4-chloro-3-indolyl-b-d-galactopyranoside) per liter, pH 7.0. The enriched cultures were spread plated on the x-gal plates and incubated at 20 °C for 48 h. At the end of incubation, the colonies with blue color indicating β-galactosidase production were selected and used for the study. Among the isolates that are capable of producing β-galactosidase, potent ones were selected by quantifying the enzyme activity using a chromogenic substrate o-nitrophenyl-β-d-galactopyranoside (ONPG). Modified LB broth (10-g peptone, 10-g yeast extract, 10-g lactose, and 20 g NaCl per liter, pH 7.0) was used as culture medium for the preparation of starter culture to evaluate the effect of various physical and chemical parameters on the growth and enzyme production.
Characterization of selected isolates
Isolation of genomic DNA
DNA from selected isolates was extracted by the boiling method [17]. Isolated pure colonies were inoculated to 5-mL sterile modified LB medium (10-g peptone, 10-g yeast extract, 20 g NaCl (w/v) per liter, pH 7.0) and incubated at 20 °C for 48 h with an agitation speed of 100 rpm. The cells were harvested by centrifuging 2 mL of the cultures at 10,000 rpm for 2 min at 4 °C. The pellets were then washed with normal saline (0.85 % NaCl, w/v) and re-suspended in 0.5-mL DNA-free sterile distilled water. Cell lysis, to release the DNA, was performed by heating the tubes that were kept at 98 ± 2 °C for 10–15 min. The lysate was centrifuged at 10,000 rpm, 4 °C for 5 min, and the supernatant containing the genomic DNA was collected and stored at −20 °C until further use.
16S rRNA amplification and phylogenetic analysis
The genus and species level identification of the selected isolates KS25, KS85, KS90, and KS92 was carried out by 16S rRNA gene amplification using the universal primer set (27F: 5′-AGA GTT TGA TCC TGG CTC AG-3′, 149 R: 5′ GGT TAC CTT GTT ACG ACT T-3′). The amplification was performed according to previously described PCR conditions by Reshma et al. (2015) [18]. Briefly, an initial denaturation at 95 °C for 2 min, followed by 30 cycles of denaturation at 95 °C for 2 min, annealing at 58 °C for 1 min, extension at 72 °C for 2 min, and a final extension at 72 °C for 10 min were included. The PCR products were sequenced at Scigenom (Kochi, India) and the sequences were submitted to BLAST (http://www.ncbi.nim.noh.gov/BLAST) to check their homology with already available 16S rRNA gene sequences in the GenBank. The sequences were submitted in the GenBank and were allotted with accession numbers. Multiple alignments of the sequences were also carried out using ClustalW software, and the phylogenetic tree was constructed by neighbor-joining method using Mega 6 software.
Detection of cellular location of the enzyme and cell permeabilization
Cellular location of cold-active β-galactosidase in E. ludwigii KS85 was detected by measuring the enzyme activity in culture broth, clear broth (supernatant), and in buffered cell pellet. To measure the intracellular β-galactosidase activity, 10-μL buffered cells were permeabilized by vortexing thoroughly with 200 μL of a modified Miller’s permeabilization solution containing 100 mM Na2HPO4, 20 mM KCl, 2 mM MgSO4, 0.8 mg/mL CTAB (Cetyl trimethyl ammonium bromide), 0.4 mg/mL sodium cholate, and 5.4 μL/mL beta-mercaptoethanol for 2 min. The permeabilized cells were examined under scanning electron microscope to analyze the role of permeabilizing agents on the cell membrane structure integrity.
Assay of β-galactosidase activity
β-Galactosidase activity was measured using the protocol given by Miller [19] with slight modification. The production medium containing 1 % (w/v) lactose was inoculated with 2 % inoculum and incubated at 20 °C for 48 h. After recording OD600 of the cultures, the cells were harvested by centrifuging at 6000 rpm at 4 °C for 10 min. The pellets were collected and re-suspended in 100-mM phosphate buffer out of which 10 μL is transferred to a 2 mL sterile Eppendorf tube containing 200-μL modified permeabilization solution. The contents were vortexed for 2 min to maximize the permeabilization of the cell membrane. The reaction was carried out by adding 0.25 mL of phosphate-buffered o-nitrophenyl-β-d-galactopyranoside (ONPG, 4 mg/mL) containing beta-mercaptoethanol (2.7 μL/mL). After 15 min, the reaction was stopped by adding 1 mL of 1 M Na2CO3. The absorbance of yellow-colored product, ONP, was determined at 420 and 550 nm after centrifuging at 10,000 rpm at 4 °C for 10 min. One unit of β-galactosidase is expressed as the change in absorbance of the product o-nitrophenol per minute per cell under assay conditions [20] and is calculated as β-galactosidase units = 1000 × OD420 − (1.73 × OD550) / (t × V × OD600) where t is the incubation time in minutes and v is the volume of culture in milliliters. Values expressed are mean ± standard error of three independent experiments.
Effect of various physicochemical parameters on β-galactosidase production
Influence of following physical and chemical parameters on bacterial growth and cold-active β-galactosidase production was analyzed by varying one factor at a time.
Incubation period and inoculum size
The kinetics of growth and β-galactosidase production was followed at different time intervals. The Arctic bacterial isolate KS85 cultivated in modified LB medium containing 1 % (w/v) lactose at 20 °C for 48 h was used as the starter culture during study. Erlenmeyer flasks containing 100 mL sterile modified LB medium (10-g peptone, 10-g yeast extract, 10-g lactose, 20 g NaCl per liter, pH 7.0) were inoculated with 2 % starter culture, and the flasks were incubated for the periods ranging from 0 to 60 h in static condition at 20 °C. The cultures were examined for the growth and enzyme production at regular intervals of 6 h. The effect of inoculum size was evaluated by inoculating 25 mL of the modified LB medium with different volumes of inoculum (0.5–4 %, v/v).
Temperature, pH, and Agitation
The role of various physical factors viz., temperature, pH, and agitation on the growth and enzyme productivity was studied in modified LB medium. To study the effect of temperature, the isolate was grown at 15, 20, 25, 30, 37, and 43 °C for optimized period of time (48 h) under standard conditions. The effect of the initial pH value on the β-galactosidase production was investigated by varying the initial pH of the culture medium from 4 to 10 at optimized conditions of temperature (25 °C) and incubation period. The growth and β-galactosidase activity were determined in fermentation flasks. The influence of agitation was studied by incubating the bacterial culture at a range of 0–150 rpm with 50.0 U variation under standard conditions.
Nitrogen source, carbon source, and metal ions
Influence of various organic nitrogen sources such as yeast extract, beef extract, malt extract, peptone, soya peptone, tryptone, and inorganic nitrogen sources like ammonium sulfate, ammonium nitrate, potassium nitrate, and urea on the growth and enzyme production was evaluated at 1 % (w/v) concentration in basal medium supplemented with 1 % lactose and 2 % sodium chloride (pH 7.0). The ability of the isolate to assimilate various carbon sources during growth and enzyme production was also assessed. The carbon sources used were glucose, galactose, xylose (monosaccharides), mannitol, sorbitol (sugar alcohols), sucrose, lactose, maltose (disaccharides), starch, carboxy methyl cellulose (polysaccharides), and glycerol. All the carbon sources were employed at 1 % concentration in a basal medium containing 1 % yeast extract (w/v), 1 % (w/v) tryptone, and 2 % (w/v) NaCl.
The potential of the isolate to utilize residual nutrients in clarified whey as the sources for nitrogen and carbon was also tested. Whey was prepared from the cow’s whole milk. The pH of the milk (250 mL) was adjusted to 3.5 with 1 N HCl and boiled for 20 min. It was then filtered through muslin cloth after cooling. Further clarification of whey was done by passing through Whatman filter paper No. 1. During the study, the clarified whey (pH 7.0) containing 2 % sodium chloride was used directly without supplementing any listed nitrogen and carbon sources. The pH of the whey was adjusted to 7.0 using 1 N NaOH and sterilized by autoclaving.
Effect of various metal ions such as Mg2+, Mn2+, Zn2+, Ni2+, Ca2+, Na+, K+, Cu2+, and Li+ at 5 mM concentration on the growth and enzyme productivity was also assessed in 25 mL of sterile modified LB medium. The fermentation medium containing metal ions was inoculated with 1 % (v/v) of 48-h inoculum and incubated at 20 °C. The growth and enzyme production in the flasks were evaluated after 48 h of incubation.
Results
Screening of cold-active β-galactosidase producers
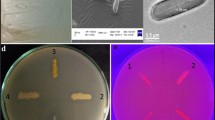
Among the 100 bacterial isolates (KS1–KS100) from Arctic fjord sediment screened, KS1, KS10, KS22, KS25, KS30, KS48, KS52, KS85, KS90, and KS92 produced blue colonies on X-gal containing LB plate indicating the production of cold-active beta galactosidase (data not shown). All the positive isolates were subcultured in lactose broth and were grown at standard conditions for 48 h. Out of the 10 bacterial isolates selected on x-gal plate, four isolates (KS25, KS85, KS60, and KS90) which shown good cold-active β-galactosidase enzyme production potential (4431.77, 4598.32, 4478.34, and 4670.07 U/min/cell, respectively) were chosen for further characterization.
Characterization of isolates and phylogenetic analysis
Preliminary examination of KS25, KS85, KS90, and KS92 indicated that these bacteria were gram negative short rods. All the isolates were indole-negative, methyl red-negative, Voges-Proskauer-positive, and citrate-positive. The morphological and biochemical characterization revealed that these isolates belonged to Enterobacter genus. Further characterization was carried out based on 16S rRNA gene sequencing. The homology analysis using 16S rRNA sequence revealed that all the bacterial isolates exhibited 99 % similarity to Enterobacter ludwigii. The phylogenetic analysis (Fig. 1) of the sequences with the available sequences in the GenBank database of NCBI revealed the closest relation of KS25, KS 85, KS90, and KS92 to E. ludwigii EN119 (99 % similarity), and the sequences were allotted with the accession numbers KT887886, KT199710, KT887887, and KT887885, respectively.
In recent years, Enterobacter sp. has been extensively studied by scientists all over the world because of their immense potential in various biotechnological applications. E. cloacae had been previously explored for their potential in the production of thermostable intracellular and extracellular β-galactosidase [21]. It was reported in a recent publication that E. cloacae is a preferred candidate for the production of diacetyl, a high value product that can be extensively used as a food ingredient [22]. Various species of Enterobacter was also reported to have many other applications such as in the production of polygalacturonase [23], galacto-oligosaccharides (GOS) [24], cellulase [25], antimicrobial lipopeptides [26], chitosan hydrolysate [27], and bioremediation of heavy metals [28]. There are reports on E. ludwigii regarding its potential for promoting plant growth [29], expression and protein modeling of CspE gene [30], and potential production of calcite nanocrystal [31]. The present investigation is focused on the cold-active β-galactosidase production potential of a psychrotrophic bacterium, E. ludwigii KS85.
Cellular location of β-galactosidase and cell permeabilization
The cellular location of cold-active β-galactosidase in E. ludwigii KS85 was determined (data not shown). It was found that all the enzyme activity was associated with the cells, and hence we can conclude that E. ludwigii KS85 produces an intracellular β-galactosidase. To overcome the cell membrane barrier of biocatalyst and also to increase the enzyme-substrate accessibility, the cells were permeabilized with a permeabilization solution. SEM images (Fig. 2) clearly showed that control cells had an intact smooth outer surface, whereas the membranes of the permeabilized cells were rough and crumpled. The morphological changes (shrinkage, formation of gaps on membrane, and reduced cell size) on the membrane structure clearly explained the role of different agents in the permeabilization solution in disorganizing the membrane by removing cell membrane lipids and generating some membrane pores through which an intracellular enzyme can easily access its substrate available in the reaction mixture. Our results were similar to the reports of researchers who described the role of certain chemical agents and organic solvents in permeabilizing the outer cell membrane of Kluyeveromyces marxianus NCIM 3465 and Ochrobactrum anthropi SY509 effectively and decreasing the membrane barrier of whole cell biocatalysts [32, 33].
Effect of various physicochemical factors for cold-active β-galactosidase production
Incubation period and inoculum size
The data obtained (data not shown) clearly shows that the growth and enzyme production were proportionate with the time of incubation up to 24 h. The maximum enzyme production occurred during the 48th hour after inoculation. Out of the different inoculums levels (0.5–4 %, v/v) tested, the growth and enzyme production was found maximum in fermentation flasks inoculated with 2 % inoculums (data not shown), whereas a slight decrease in growth and enzyme activity was observed with further increase in inoculums size beyond 3 %. Sabu et al. (2006) [34] proved that any increase in inoculum size beyond optimum level disturbed the equilibrium between the proliferating bacterial population and substrate availability in the medium which is highly inevitable for maximum enzyme yield.
Effect of temperature, pH, and agitation
KS85 was able to grow and produce cold-active β-galactosidase enzyme at temperatures 15–37 °C with an optimum production at 25 °C. At temperatures of 15 and 20 °C, the isolate has shown good enzyme activity (90 and 94 %, respectively). The growth and enzyme production by KS85 was found to be restricted at and beyond 43 °C (Fig. 3a). After 72 h of incubation at 8 °C (lowest temperature selected for this study), E. ludwigii KS85 showed cell growth indicating the psychrotrophic nature of the organism [35, 36].
The effect of pH on the growth and enzyme activity was determined at 20 °C (pH 4–10). E. ludwigii was able to grow and produce enzyme at pH ranging from 5 to 10. The enzyme displayed maximum activity at pH 7.0 and 8.0 (Fig. 3b). However, there was good growth and activity at pH 5, 6, 9, and 10 (72, 81, 93, and 84 %, respectively) claiming the broad pH range activity of β-galactosidase [24, 37].
Agitation of the culture flasks was effective over static cultivation condition for better bacterial growth and enzyme production. Over the different agitation speed studied, the maximum growth as well as β-galactosidase production was detected at 90 rotations per minute (data not shown). Further increase in agitation speed led to the decrease in growth and enzyme activity. The studies on Kluyveromyces marxianus CCT 7082 proved that the volumetric β-galactosidase production was strongly influenced by mixing conditions [38]. According to Zambare et al. (2014) [39], agitation of fermentation flask produces highly homogenized environment for better fermentation with maximum dispersion of both oxygen and nutrients and they reported that the cell biomass yield and protease production were poor at lower agitation rate which may be due to decreased oxygen mass energy transfer.
Nitrogen source, carbon source, and metal ions
Several nitrogen sources, one at a time, were tested in the minimal medium to check their influence on growth and enzyme production (Fig. 4). The bacterial growth and enzyme productivity were found to be strongly influenced by the presence of various organic nitrogen sources compared to inorganic representatives. The fermentation medium containing nitrogen sources such as yeast extract and beef extract favored maximum enzyme production with relative activities of 99 and 96 %, respectively, followed by tryptone (91 %) and peptone (88 %). It has been reported that Pantoea agglomerance, a close relative of E. ludwigii, preferred yeast extract as best nitrogen source for the growth [40, 41]. Bifidobacterium animalis Bb12 and Lactobacillus delbrueckii ssp. bulgaricus ATCC 11842 preferred whey medium supplemented with yeast extract, peptone, and casein hydrolysate for maximum β-galactosidase production [42]. Lactobacillus acidophilus preferred yeast extract for the production of β-glucosidase [43]. The preference of organic nitrogen sources over inorganic nitrogen sources may be due to the complex nature of these sources with vitamins, minerals, and amino acids which can add to the nutritional value of the growth medium and contribute much to the growth as well as enzyme production. All the inorganic nitrogen sources contributed much less to the growth and enzyme production by E. ludwigii KS85.
Though E. ludwigii KS85 was able to assimilate various carbon sources supplemented in the basal medium during growth, β-galactosidase production was found to be induced by lactose, glycerol, and whey. All the other carbon sources supplemented in the medium failed to stimulate enzyme production (Fig. 5). Results revealed that lactose was the most effective carbon supplement for cold-active β-galactosidase production by E. ludwigii KS85 (4502 U/min/cell). Different researchers observed that β-galactosidase induction strictly depends on the carbon source provided in the medium and the organism used. Lactose was found to promote β-galactosidase production in Enterobacter aerogenes [44] and Geobacillus stearothermophilus [45], whereas Bacillus licheniformis ATCC 12759 β-galactosidase induction was done by xylose and galactose in submerged fermentation [46]. In the medium with 1 % level, glycerol relative activity of β-galactosidase was found to be 48 %. It had already been proved that a good yield of β-galactosidase occurred when Escherichia coli strains were grown in M9 medium containing 0.2 % glycerol as carbon source [47].
The study also aimed to check the ability of organism to utilize soluble proteins and lactose present in the clarified whey as nitrogen and carbon sources, respectively. The relative enzyme production in whey fermentation flask was 93 %. High activity in whey medium suggests that whey could be a cost-effective raw medium for the production of cold-active β-galactosidase. Recycling of whey, an industrial pollutant can reduce the problems associated with its disposal. During the last decades, whey has been employed as good medium for the microbial production of industrially valuable products [48–50].
The data (Fig. 6) clearly shows that at 5-mM concentrations, Mn2+ and Mg2+ slightly enhanced the bacterial growth and enzyme production. At the same concentrations, β-galactosidase production was found partially inhibited by K+, Na+, Li2+, and Ca2+ with relative activities of 96, 94, 94, and 85 %, respectively. The presence of Zn2+, Ni2+, and Cu+ in the medium strongly inhibited both bacterial growth and β-galactosidase production.
Conclusion
Though the cold adapted bacteria and their enzymes have immense potentials in industrial application, they were found underutilized in biotechnology. In the present study, we have described a psychrotrophic bacterium E. ludwigii KS85 from the sediments of Kongsfjord, Arctic with very good β-galactosidase production potential. The study revealed that the medium components and fermentation parameters such as carbon and organic nitrogen sources, pH, temperature, inoculum size, and incubation period are significant in the β-galactosidase production by E. ludwigii KS85. The ability of the organism to assimilate the soluble nutrients in whey, an industrial pollutant, could greatly reduce the overall cost of production medium as well as the environmental problems associated with whey disposal. To the best of our knowledge, there is no report on psychrotrophic E. ludwigii in this regard and hence the studies on purified cold-active β-galactosidase from E. ludwigii would be helpful in its commercial exploitation in the future.
References
Cavicchioli, R., Charlton, T., Ertan, H., Omar, S. M., Siddiqui, K. S., & Williams, T. J. (2011). Biotechnological uses of enzymes from psychrophiles. Microbial Biotechnology, 4, 449–460.
Ghosh, M., Pulicherla, K. K., Rekha, V. P. B., Raja, P. K., & Sambasiva Rao, K. R. S. (2012). Cold active β-galactosidase from Thalassospira sp. 3SC-21 to use in milk lactose hydrolysis: a novel source from deep waters of Bay-of-Bengal. World Journal of Microbiology and Biotechnology, 28, 2859–2869.
Vester, J. K., Glaring, M. A., & Stougaard, P. (2014). Discovery of novel enzymes with industrial potential from a cold and alkaline environment by a combination of functional metagenomics and culturing. Microbial Cell Factories, 13, 72.
Nakagawa, T., Ikehata, R., Uchino, M., Miyaji, T., Takano, K., & Tomizuka, N. (2006). Cold-active acid β-galactosidase activity of isolated psychrophilic-basidiomycetous yeast Guehomyces pullulans. Microbiological Research, 161, 75–79.
Panesar, P. S., Kumari, S., & Panesar, R. (2010). Potential applications of immobilized β-galactosidase in food processing industries. Enzyme Research, 2010, 473137.
Białkowska, A. M., Cieśliński, H., Nowakowska, K. M., Kur, J., & Turkiewicz, M. (2009). A new β-galactosidase with a low temperature optimum isolated from the Antarctic Arthrobacter sp. 20B: gene cloning, purification and characterization. Archives of Microbiology, 191, 825–835.
Husain, Q. (2010). Beta galactosidases and their potential applications: a review. Critical Reviews in Biotechnology, 30(August 2009), 41–62.
Karasová-Lipovová, P., Strnad, H., Spiwok, V., Malá, Š., Králová, B., & Russell, N. J. (2003). The cloning, purification and characterisation of a cold-active β-galactosidase from the psychrotolerant Antarctic bacterium Arthrobacter sp. C2-2. Enzyme and Microbial Technology, 33, 836–844.
Sako, T., Matsumoto, K., & Tanaka, R. (1999). Recent progress on research and applications of non-digestible galactooligosaccharides. International Diary Journal, 9, 69–80.
Wierzbicka-Woś, A., Cieśliński, H., Wanarska, M., Kozłowska-Tylingo, K., Hildebrandt, P., & Kur, J. (2011). A novel cold-active β-D-galactosidase from the Paracoccus sp. 32d - gene cloning, purification and characterization. Microbial Cell Factories, 10, 108.
Horner, T. W., Dunn, M. L., Eggett, D. L., & Ogden, L. V. (2011). β-Galactosidase activity of commercial lactase samples in raw and pasteurized milk at refrigerated temperatures. Journal of Dairy Science, 94(2007), 3242–3249.
Nakagawa, T., Ikehata, R., Myoda, T., Miyaji, T., & Tomizuka, N. (2007). Overexpression and functional analysis of cold-active β-galactosidase from Arthrobacter psychrolactophilus strain F2. Protein Expression and Purification, 54, 295–299.
Coker, J., Sheridan, P. P., Loveland-Curtze, J., Gutshall, K. R., Auman, A. J., & Brenchley, J. E. (2003). Biochemical characterization of a β-galactosidase with a low temperature optimum obtained from an Antarctic Arthrobacter isolate. Journal of Bacteriology, 185(18), 5473–5482.
Hoyoux, A., Jennes, I., Dubois, P., Genicot, S., Dubail, F., François, J. M., Baise, E., Feller, G., & Gerday, C. (2001). Cold-adapted β-galactosidase from the antarctic psychrophile Pseudoalteromonas haloplanktis. Applied and Environmental Microbiology, 67(4), 1529–1535.
Fernandes, S., Geueke, B., Delgado, O., Coleman, J., & Hatti-Kaul, R. (2002). β-Galactosidase from a cold-adapted bacterium: purification, characterization and application for lactose hydrolysis. Applied Microbiology and Biotechnology, 58, 313–321.
Hu, J. M., Li, H., Cao, L. X., Wu, P. C., Zhang, C. T., Sang, S. L., Zhang, X. Y., Chen, M. J., Lu, J. Q., & Liu, Y. H. (2007). Molecular cloning and characterization of the gene encoding cold-active beta-galactosidase from a psychrotrophic and halotolerant Planococcus sp. L4. Journal of Agricultural and Food Chemistry, 55, 2217–2224.
Devi, R., Surendran, P. K., & Chakraborty, K. (2009). Antibiotic resistance and plasmid profiling of Vibrio parahaemolyticus isolated from shrimp farms along the Southwest coast of India. World Journal of Microbiology and Biotechnology, 25(1603), 2005–2012.
Silvester, R., Alexander, D., & Ammanamveetil, M. H. A. (2015). Prevalence, antibiotic resistance, virulence and plasmid profiles of Vibrio parahaemolyticus from a tropical estuary and adjoining traditional prawn farm along the southwest coast of India. Annals of Microbiology. doi:10.1007/s13213-015-1053-x.
Miller, J. H. (1972). Experiments in molecular genetics (pp. 352–355). Cold Spring Harbor: Cold Spring Harbor Laboratory.
Miller, J. H. (1992). A short course in bacterial genetics. Cold Spring Harbor: CSH Laboratory Press.
Ghatak, A., Guha, A. K., & Ray, L. (2010). β-D-Galactosidase from Enterobacter cloacae: production and some physicochemical properties. Applied Biochemistry and Biotechnology, 162, 1678–1688.
Zhang, L., Zhang, Y., Liu, Q., Meng, L., Hu, M., Lv, M., Li, K., Gao, C., Xu, P., & Ma, C. (2015). Production of diacetyl by metabolically engineered Enterobacter cloacae. Scientific Reports, 5, 9033.
Darah, I., Nisha, M., & Lim, S. H. (2013). Enhancement of polygalacturonase production from enterobacter aerogenes NBO2 by submerged fermentation. Advanced Studies in Biology, 5(5), 173–189.
Lu, L. L., Xiao, M., Li, Z. Y., Li, Y. M., & Wang, F. S. (2009). A novel transglycosylating β-galactosidase from Enterobacter cloacae B5. Process Biochemistry, 44, 232–236.
Lokapirnasari, W. P., Nazar, D. S., Nurhajati, T., Supranianondo, K., & Yulianto, A. B. (2015). Production and assay of cellulolytic enzyme activity of Enterobacter cloacae WPL 214 isolated from bovine rumen fluid waste of Surabaya abbatoir. Indonesia, 8, 367–371.
Mandal, S. M., Sharma, S., Pinnaka, A. K., Kumari, A., & Korpole, S. (2013). Isolation and characterization of diverse antimicrobial lipopeptides produced by Citrobacter and Enterobacter. BMC Microbiology, 13(1), 152.
Yamasaki, Y., Fukumoto, I., Kumagai, N., Ohta, Y., Nakagawa, T., Kawamukai, M., & Matsuda, H. (1992). Continuous chitosan hydrolyzate production by immobilized chitosanolytic enzyme from Enterobacter sp. G-1. Bioscience, Biotechnology, and Biochemistry, 56(June 2015), 1546–1551.
Banerjee, G., Pandey, S., Ray, A. K., & Kumar, R. (2015). Bioremediation of heavy metals by a novel bacterial strain Enterobacter cloacae and Its antioxidant enzyme activity, Flocculant production, and protein expression in presence of lead, cadmium, and nickel. Water, Air, & Soil Pollution, 226.
Shoebitz, M., Ribaudo, C. M., Pardo, M., Cantore, M. L., Ciampi, L., & Curá, J. (2009). Plant growth promoting properties of a strain of Enterobacter ludwigii isolated from Lolium perenne rhizosphere. Soil Biology and Biochemistry, 41, 1768–1774.
Kandasamy, P., Chaturvedi, N., Sisodia, B. S., Shasany, A. K., Gahoi, S., Marla, S. S., & Goel, R. (2013). Expression of CspE by a psychrotrophic bacterium Enterobacter ludwigii PAS1, isolated from Indian Himalayan soil and in silico protein modelling, prediction of conserved residues and active sites. Current Microbiology, 66, 507–514.
Ghashghaei, S., & Emtiazi, G. (2013). Production of calcite nanocrystal by a urease-positive strain of Enterobacter ludwigii and study of its structure by SEM. Current Microbiology, 67, 406–413.
Kumari, S., Panesar, P. S., Bera, M. B., & Singh, B. (2011). Permeabilization of Yeast Cells for beta galactosidase activity using mixture of organic solvants: a response surface methodology approach. Asian Journal of Biotechnology, 3(4), 406–414.
Choi, K. O., Song, S. H., & Yoo, Y. J. (2004). Permeabilization of Orchrobactrum anthropi SY509 cells with organic solvents for whole cell biocatalyst. Biotechnology and Bioprocess Engineering, 9, 147–150.
Sabu, A., Augur, C., Swati, C., & Pandey, A. (2006). Tannase production by Lactobacillus sp. ASR-S1 under solid-state fermentation. Process Biochemistry, 41, 575–580.
Baghel, V. S., Tripathi, R. D., Ramteke, P. W., Gopal, K., Dwivedi, S., Jain, R. K., Rai, U. N., & Singh, S. N. (2005). Psychrotrophic proteolytic bacteria from cold environment of Gangotri glacier, Western Himalaya, India. Enzyme and Microbial Technology, 36, 654–659.
Jadhav, V. V., Pote, S. S., Yadav, A., Shouche, Y. S., & Bhadekar, R. K. (2013). Extracellular cold active lipase from the psychrotrophic Halomonas sp. BRI 8 isolated from the Antarctic sea water. Songklanakarin Journal of Science and Technology, 35(6), 623–630.
Beniwal, V., Chhokar, V., Singh, N., & Sharma, J. (2010). Optimization of process parameters for the production of tannase and gallic acid by Enterobacter cloacae MTCC 9125. Journal of American Science, 6(July 2015), 389–397.
Alves, F. G., Filho, F. M., De Medeiros Burkert, J. F., & Kalil, S. J. (2010). Maximization of β-Galactosidase production: a simultaneous investigation of agitation and aeration effects. Applied Biochemistry and Biotechnology, 160, 1528–1539.
Zambare, V. P., Nilegaonkar, S. S., Kshirsagar, P. R., & Kanekar, P. P. (2014). Scale up production of Protease using Pseudomonas aeruginosa MCM B-327 and its detergent compatibility. Journal of Biochemical Technology, 5, 698–707.
Costa, E., Teixidó, N., Usall, J., Atarés, E., & Viñas, I. (2002). The effect of nitrogen and carbon sources on growth of the biocontrol agent Pantoea agglomerans strain CPA-2. Letters in Applied Microbiology, 35, 117–120.
Konsoula, Z., & Liakopoulou-Kyriakides, M. (2007). Co-production of α-amylase and β-galactosidase by Bacillus subtilis in complex organic substrates. Bioresource Technology, 98, 150–157.
Laxmi, N. P., Mutamed, M., & Nagendra, P. S. (2011). Effect of nitrogen sources on production of β-galactosidase from Bifidobacterium animalis Bb12 and Lactobacillus delbrueckii ssp. bulgaricus ATCC 11842 grown in whey under different culture conditions. International Food Research Journal, 18, 445–450.
Mahajan, P. M., Desai, K. M., & Lele, S. S. (2012). Production of cell membrane-bound α-and β-glucosidase by lactobacillus acidophilus. Food and Bioprocess Technology, 5, 706–718.
Khleifat, K. M., Abboud, M. M., Al-Mustafa, A. H., & Al-Sharafa, K. Y. (2006). Effects of carbon source and Vitreoscilla hemoglobin (VHb) on the production of β-galactosidase in Enterobacter aerogenes. Current Microbiology, 53, 277–281.
Soliman, N. A. (2008). Coproduction of thermostable amylase and β-galactosidase enzymes by Geobacillus stearothermophilus SAB-40: application of Plackett-Burman design to evaluate culture requirements affecting enzyme production. Journal of Microbiology and Biotechnology, 18, 695–703.
Nurullah, A. K. C. A. N. (2011). High level production of extracellular β-galactosidase from Bacillus licheniformis ATCC 12759 in submerged fermentation. African Journal of Microbiology Research, 5(26), 4615–4621.
Chan, V., Dreolini, L. F., Flintoff, K. A., Lloyd, S. J., & Mattenley, A. A. (2002). The effect of glycerol, glucose, glactose, lactose with galactose on the induction of -galactosidase in Escherichia coli. Journal of Experimental Microbiology and Immunology, 2(April), 130–137.
Panesar, P. S., Kennedy, J. F., Gandhi, D. N., & Bunko, K. (2007). Food chemistry Bioutilisation of whey for lactic acid production. Food Chemistry, 105, 1–14.
Foda, M. I., Dong, H., & Li, Y. (2010). Study the suitability of cheese whey for bio-butanol production by Clostridia. Journal of American Science, 6(July), 39–46.
Zohri, A. A., Gomah, N. H., & Ali, M. A. (2014). Utilization of cheese whey for bio-ethanol production. Universal Journal of Microbiology Research, 2(4), 57–69.
Acknowledgments
We are thankful to the Department of Marine Biology, Microbiology and Biochemistry and Sophisticated Testing and Instrumentation Centre at Cochin University of Science and Technology and National Centre for Antarctic and Ocean Research for providing the facilities and financial assistance to carry out the research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alikkunju, A.P., Sainjan, N., Silvester, R. et al. Screening and Characterization of Cold-Active β-Galactosidase Producing Psychrotrophic Enterobacter ludwigii from the Sediments of Arctic Fjord. Appl Biochem Biotechnol 180, 477–490 (2016). https://doi.org/10.1007/s12010-016-2111-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2111-y