Abstract
Pig farming is one of the major sources of greenhouse gas (GHG) emissions in the agricultural sector; nevertheless, few studies have been undertaken to directly measure or estimate GHGs, particularly carbon dioxide (CO2) from pig barns. Therefore, the main objective of the present research was to estimate and predict CO2 emission rate as a function of the mass of pigs and feed consumption. Two identical experiments were carried out in experimental pig barns in 2020 and 2021 to develop and evaluate the performance of CO2 emission model. The CO2 emission data (ppm) were collected utilizing Livestock Environment Management Systems (LEMS) and weather sensors, respectively within the pig barns and the outside environment. The models were built using seven statistical and machine learning–based regression algorithms, i.e., linear, multiple linear, polynomial, exponential, ridge, lasso, and elastic net. The findings of the study revealed that among the seven models, the exponential-based regression model performed the best, with a coefficient of determination (R2) greater than 0.78 in the training stage and 0.75 in the testing stage being suitable to describe the relationship between the feed intake and the rate of CO2 emission. However, when compared to the other models in the testing stage, the lasso model had the worst performance (R2 < 0.65 and RMSE > 20.00 ppm). In conclusion, this study recommends employing an exponential-based regression model by taking feed intake as an input variable in predicting CO2 for a small number of the experimental dataset.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increasing carbon dioxide (CO2) emissions from livestock farming, as well as their impact on global climate, poses a significant challenge to reduce greenhouse gas (GHG) emissions. Enteric fermentation and manure management, feed production, and energy consumption all contributed significantly to CO2 emission in the livestock sector, either directly or indirectly (Rojas-Downing et al. 2017; Ambade et al. 2021a; Maharjan et al. 2021; Ambade et al. 2022a; Kurwadkar et al. 2022). Several authors have recently quantified the amount of CO2 emitted by livestock and its contribution to global agricultural-induced GHGs emissions (Philippe and Nicks 2014; González et al. 2014). It was discovered that livestock production alone emitted approximately 8.1 gigatons (Gt) of CO2 (FAO 2010; Petrovic et al. 2015), accounting for 14.5% of global anthropogenic GHGs emissions in 2013 (Gerber et al. 2013). According to Prasad et al. (2015), the livestock production process generates nearly 9% of CO2, out of which 75% emits only from ruminants.
Pig production is the second largest source of GHGs emissions in the livestock industry (FAO 2015), and this contribution is on the rise, due to global dietary shifts. Pig meat accounts for 45% of the world’s total meat consumption (Pork Chekoff 2018). According to a recent report by Gerber et al. (2013), the global demand for pig meat will rise by 32% between 2005 and 2030, resulting in increased emissions of greenhouse gases. For instance, CO2 emissions from pig production accounted for approximately 13% of the total emissions of the livestock sector in 2010, which is higher than in previous decades, and this proportion is expected to grow significantly in the upcoming years (FAO 2013). Therefore, determining the CO2 emission from a pig farm is a crucial issue for mitigating the adverse effects of GHGs on the global climate. Within this context, this paper aims to quantify the amount of CO2 emission from an experimental pig barn.
Given that a variety of environmental, management, and housing factors have been shown to affect CO2 emission, it is critical to identify precise input parameters that are more informative for CO2 emission modeling. Additionally, it is critical to develop a rigorous model with the optimal architecture in order to maximize model performance. Recently, a variety of models have been used to estimate the rate of CO2 emission, including regression-based equations (Keat et al. 2015; Zhou et al. 2021), decomposition models (Cansino et al. 2015; Xiao-wen et al. 2021), and empirical models (Lashkenari and KhazaiePoul 2015). According to the decomposition analysis, structural adjustment in agriculture, growing affluence, and population growth contributed to an increase in the GHG emissions of pork production by 23, 41, and 13 Mt CO2 eq, respectively (Xiao-wen et al. 2021). According to a life cycle analysis (LCA) that takes into account the entire process of producing animal products, the livestock sector contributes 9% of CO2 to total GHGs emissions (Oonincx et al., 2010). Regrettably, the application of these methods for CO2 measurement is further limited by the complexities associated with conducting experiments in pig barns. On the other hand, machine learning techniques appear to be promising in terms of addressing and mitigating the limitations of existing methods (Rybarczyk and Zalakeviciute 2018).
Since monitoring CO2 in the air is important for pigs’ health and their living environment, it is essential to regulate effective controlling systems. The CO2-based model can act as an initial step in controlling mechanisms. Evaluating the performance of statistical and machine learning models in the field of air quality modeling and forecasting has been undertaken in several studies (Austin 2007; Kisi et al. 2017; Kang et al. 2018; Shahriar et al. 2020; Ambade et al. 2021b; Ambade et al. 2021c; Ambade et al. 2021d; Ambade et al. 2022b). However, based on relevant literature, the study of regression-based CO2 prediction model was limited in the experimental pig barn. In this study, regression-based models were developed using statistical and machine learning algorithms to measure the CO2 emission as a function of feed intake and mass of pigs.
These models are effective in predicting CO2 concentrations in experimental pig barns in two ways. To begin, the models can be used to understand the emission pattern of CO2 in different stages of pig growth, which may lead to improved knowledge. Second, they are useful for monitoring the inside environment to describe the relationships between feed intake, pig mass, and CO2 emission. The primary goal of these methods is to investigate the relationship between observed and predicted parameters (Boldina and Beninger 2016). The most common type of regression-based model is linear form, which is used in a wide variety of fields. Regression analysis was used in some studies, such as the characterization of spatial patterning (Beninger and Boldina 2014), and demonstration of the relationship between phytoplankton cell size and abiotic parameters in dynamic energy budget (DEB) models (Duarte et al. 2012; Agüera et al. 2017). Like linear models, polynomial regression models are also used in a variety of research fields (Legendre and Legendre 2012). Furthermore, the polynomial model is preferable because it can easily fit the data. However, higher-order polynomials can be difficult to obtain any meaningful meaning, which is actually unacceptable in some cases (Peng et al. 2017; Muthusamy et al. 2021). On the other hand, for nonlinear data, a linear or polynomial-based regression model is difficult to outperform. In this case, exponential, ridge, lasso, or elastic net-based machine learning performed well (Glazier 2013; Packard 2013; Joharestani et al. 2019; Shahriar et al. 2020, Basak et al. 2022). Such regression-based statistical and machine learning models for estimating CO2 emission rate from feed intake and pig mass can reduce sampling effort and cost while potentially increasing precision in cases where samples are difficult to handle.
Measuring CO2 concentrations in an experimental pig barn entails a lengthy process that includes setting up the experimental setup and maintaining internal environmental conditions, dietary composition, and feed intake, among other things. Additionally, the CO2 emission rate varies significantly within the same experimental pig barn year after year, making measurement even more difficult. In this regard, regression-based statistical and machine learning models may be advantageous for forecasting CO2 emissions. As such, the research objectives are to quantify the daily CO2 emission rate per pig using livestock environment management systems (LEMS, AgriRoboTech Co., Ltd, Republic of Korea) and then to use regression methods to model CO2 emission rate as a function of feed intake and mass of pigs.
Materials and methods
Animal Resources, experimental design, and data collection
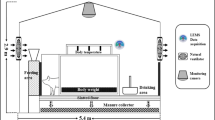
The experimental procedures were approved by Gyeongsang National University’s ethical and animal experimentation committee (certification #GNU-150508-R0029). Two independent tests were carried out in the Smart Farm Systems Laboratory at Gyeongsang National University with six, 2-month-old crossbreed pigs (American Yorkshire × Duroc) in experimental pig barns from September 1, to December 15, 2020 and 2021. During the both experimental periods, only one pig barn was used for rearing pigs which was mentioned as Livestock 2 (LS-2), and another one was mentioned as Livestock 1 (LS-1). The average dimension of each experiment pig barn was 3.3 m (width) × 5.4 m (length) × 2.9 m (height) and 0.05 m thickness of roof and wall (Fig. 1). The sidewalls and roof were made of galvanized steel and expanded polystyrene, respectively. These materials were chosen for their ability to maintain a comfortable environment within the pig barn (Moon et al. 2016). A constantly ventilated fan and an air inlet damper (Auto-Damper 250, Sanison Co., Ltd., Korea) were installed at 1.44 m and 1.72 m above ground level, respectively, for air circulation with an average flow rate of 0.16 m3s−1. We recorded CO2 data from both experimental pig barns (LS-1 and LS-2) at the same flow rate of the ventilated fan throughout the entire experimental period.
In both experimental periods, similar sizes of the pigs (ages and weights) were studied by providing Growing Pigs Late Feed 10 (Nonghyupfeed Co., Ltd., Seoul, Republic of Korea) concentrated diet (Table 1). Pigs of similar sizes were retained, implying that CO2 generation rates would be comparable. Each pig received an identical amount of feed twice daily at 10 a.m. and 5 p.m., and the amount of feed consumed was estimated using daily records of feed offered and leftovers from each pig. Additionally, the mass of the pigs was determined by averaging weights taken twice daily with a load cell. Moreover, the LS-2 barn was equipped with a drinker and feeder so that pigs could be restrained with halters while eating and drinking (Fig. 1). The LS-2 was cleaned of pig manure, including urine and fecal matter every 15-day intervals. The experimental setup view of the LS-1 and LS-2 was represented in the Supplementary Section (Fig. S1).
Environmental data inside the two pig barns such as temperature, humidity, and CO2 concentration were measured by using LEMS every 5-min intervals. The collected CO2 data from the two pig barns were later analyzed to obtain the emission rate of CO2 per pig (ppm pig−1). To obtain the CO2 emission rate, the emitted CO2 was subtracted by LS-1 from LS-2, after it was scaled by the number of pigs in the pig barn to obtain CO2 produced by a pig. Weather sensors were used to collect data on the outside temperature, humidity, and carbon dioxide levels (MetPRO, Producer: Campbell Scientific, United States of America) (Fig. S2).
Data pre-processing and model development
Data collected from the two experimental pig barns in 2020 and 2021 were used to build models for predicting CO2 emission rate. Throughout the model preparation stage, pig mass (MP), age, and feed intake (FI) were all considered as input variables. However, a high-multicollinearity value was discovered between body weight and age (correlation coefficient (r) > 0.94); therefore, MP and FI were chosen as model input variables. The Z-score data normalization technique (Eq. 1) was used to keep values within a scale that was applied across all numeric columns in the model (Jain et al. 2018):
where Z is the standard score; X is the value in the input dataset; μ is the mean of the variable of X, and σ is the standard deviation.
Due to the limited number of experimental data on input variables (the number of input data is 182), seven regression-based statistical and machine learning models were evaluated based on how well these algorithms predicted CO2 emission rate from the three types of datasets presented in Table 2. The mathematical approaches used to predict CO2 emission rate were briefly explained in the following sections of the methodology part.
Linear regression model
One of the primary objectives of this study was to establish relationships between pig body mass, feed intake, and CO2 emission rate. These relationships are frequently difficult to detect. The linear regression model is one of the most frequently used approaches for fitting a study variable (dependent variable) to a set of covariables (independent variables). The linear regression equation is denoted by the following notation (Permai and Tanty 2018):
where dependent variable Y indicates approximately a linear function of independent variable X, while ε refers to the degree of discrepancy of this approximation, \({\beta }_{0}\) is the model intercept, and \({\beta }_{1}\) is the slope of the straight line. The term ε denotes the error of approximating the observed value Y by means of the linear estimation obtained from the model.
In cases where we have more than one independent variable, the multiple linear regression (MLR) model is based on the same assumptions as simple linear regression. The purpose of this MLR model was to investigate and quantify the relationship between explanatory (MP and FI) and response variables in the current study (CO2 emission rate). The MLR model is structured as follows (Darlington and Hayes 2016):
where Yi is the CO2 emission rate, \({\beta }_{0}\) is the model intercept, β1–βn are the coefficients of regression, X1–Xn are the input variables, and ε is the error associated with the ith observation.
PR
Polynomial regression describes a nonlinear relationship in which the independent variable X and the dependent variable Y are modeled as a degree polynomial (Ostertagová 2012). Polynomial regression (PR) models comprise squared and higher order terms of the dependent variables creating the response line curvilinear form (Gendy et al. 2015). A polynomial mathematical model based on experimental data was investigated in different orders to study this phenomenon and to find the best correlation. The general form of a complete second degree PR model with two independent variables X1 and X2 is shown in Eq. (4) (Gavrilova 2021):
where \({\mu }_{y\mathrm{\rm I}\left(X1, X2\right)}\) refers to the true mean response for the independent variables, \({\beta }_{0}\) is the model intercept, β (1, 2, 3, 4, 5) are the model parameters, and X1 and X2 (MP and FI) are the independent variables. After examining different degrees of polynomials (order = 2, 3, 4, 5), in this study, we decided to perform 4-degree due to its better performance.
ER
Exponential regression model (ER) model is normally used to explain the situation in which growth starts gradually and then accelerates quickly without bound; otherwise, decline starts quickly and gradually slows down, getting closer and closer to zero (Dettea and Neugebauer 1997; Silva and Almeida 2018). The ER model is defined as
where the input variable X follows as an exponent. The exponential curve is determined by the exponential function, and it depends on the value of the X.
RR
A penalty-based regression approach (i.e., ridge regression) was also used to model CO2 emission in this study. The ridge regression model (RR) has been frequently used to test a variety of features in a single sample at the same time (Ransom et al. 2019; Wieringen 2020). It is a continuous method of shrinkage in which the residual sum of squares is diminished, as each coefficient is shrunk towards zero, thus decreasing the significance or impact of any specific factor (Nagesha et al. 2019; Ransom et al. 2019). Although the RR equation is quite similar to the least-squares equation, the minimization equation is slightly different, as seen in Eq. (6):
where \({\Arrowvert{y-X}_\beta^\wedge\Arrowvert}^2\) is called the sum of the squares of all coefficients (RSS), and it is also denoted as a loss function, and the λ parameter is the regularization penalty.
LaR
Likewise ridge regression, Lasso regression model (LaR) also minimizes the residual sum of squares, subject to a constraint on the sum of the absolute values of the regression coefficients (Rajaratnam et al. 2019). The estimation of regression coefficients by the lasso regression model can be defined as:
where \(\left({}_\beta^\wedge\right)\) is a p-dimensional vector containing the lasso estimates of the slope coefficients, and λ is a regularization parameter (Rajaratnam et al. 2019).
Elastic net model (ENR)
Elastic net was developed through critiques of lasso and ridge regression algorithms, whose variable selection is highly dependent on data and thus unstable. Like ridge and lasso, elastic net is also used to improve the performance of linear regression model. To get the best performance, elastic net combined the penalties of both ridge and lasso regression (Mol et al. 2009). In order to have a good understanding of ridge and lasso, and elastic net regression model, it was more described in an earlier study noted in Aldabal (2020). Elastic net aims at minimizing the following loss function:
where α is the mixing parameter between ridge (α = 0) and lasso (α = 1).
Experimental setup and performance metrics
In this study, all models were developed using open source libraries in a Python (Python 3.7.0) environment. Python is a powerful, interpreted programming language with a wide range of applications, including scientific research (Tran et al. 2020). Different libraries including NumPy (Van Der Walt et al. 2011), Pandas (McKinney 2010), and Matplotlib (Hunter 2007) in the Python platform were employed for processing, manipulating, and displaying data. In this study, 75% of the experimental data were used as the training while 25% was used as the testing dataset. Two model evaluation metrics such as root mean square error (RMSE) and coefficient of determination (R2) were used to assess the models’ performance. The Statistical Package for the Social Sciences (IBM SPSS Statistics 22.0.0.0, New York, USA) and Origin Pro 9.5.5 were used to perform all statistical calculations including analysis of variance (ANOVA), correlation of input variables in this work (OriginLab, Northampton, Massachusetts, USA).
Results and discussions
Data overview
The descriptive statistics for CO2 concentration, air temperature, and relative humidity data in LS-1, LS-2, and outside the livestock are presented in Table 3. Among the three locations, the highest mean CO2 emission (797.55 ppm in 2020 and 752.84 ppm in 2021) and relative humidity (67.85% in 2020 and 60.20% in 2021) were observed in LS-2 and the lowest CO2 (121.92 ppm in 2020 and 119.02 ppm in 2021) and relative humidity (45.68% in 2020 and 46.98% in 2021) were found in LS-1 and outside environment, respectively. While the outside environment of livestock was found to have the highest air mean temperature of 21.5 °C (21.42°C in 2020 and 21.54 °C in 2021), LS-1 was found to have the lowest (17.23 °C in 2020 and 19.80 °C in 2021) (Table 3).
Figure 2 depicts the pattern of CO2 emission rate changes prior to 15 days of pigs entering LS-2 and throughout the experimental period. It was demonstrated that when the pigs began rearing in LS-2, the CO2 emission rate increased dramatically. However, a similar pattern was observed in LS-1 and the surrounding environment during the whole experimental time. Additionally, the current study examined that the differences in CO2 and relative humidity among the three locations (LS-1, LS-2, and outside) were statistically significant (p < 0.05), whereas the variation in air temperature between LS-1 and LS-2 was statistically insignificant (p = 0.168 in 2020 and p = 1.00 in 2021). To gain a better understanding of the relationship between ambient environmental parameters inside and outside of pig barns, the results of our previous study were described (Basak et al. 2020). Figure 3 shows the relationships between the mass of pigs, age, feed intake, and CO2 emission in 2020 and 2021.
Model execution
The microclimatic parameters of swine buildings, such as temperature, relative humidity, and CO2 concentration are widely quantified using statistical models (Besteiro et al. 2017; Tuomisto et al. 2017). The current study investigated widely used regression-based statistical and machine learning models for predicting CO2 levels in a swine building. The evaluation results from the model execution were classified into four categories: input datasets, model performance, model comparison, and proposed model. The “Input datasets” section discusses the results obtained using datasets D1, D2, and D3. The model performance section discusses the results obtained during the training and testing stages of those models. Finally, the “Model comparison and proposed model” section discuss the percentage differences between the results of all models.
Input datasets
The current study considered the use of three datasets, i.e., D1, D2 and D3, in predicting CO2 emission. The performances of all models with three input datasets were evaluated in the training and testing stages (Table 4). For example, the ER model obtained the best performance with D2 (R2 = 0.757 and RMSE = 16.45 ppm pig−1) in the testing stage. The results also indicated that the selected ER model with D2 dataset could predict CO2 emission rate for testing stage with a 1.67% increase in R2 and a reduction of 18.36% in RMSE compared to D1 dataset. Like the ER model, the LaR model also had a better performance using D2 (13.44% and 7.65% increase in R2 and a reduction of 15.62% and 10.34% in RMSE, respectively) compared to D1 and D3 datasets. Similar results were also obtained for the other regression-based models; however, the performances of those models in training stage were nearly identical (Table 4; Figs. 4, 5, and 6).
According to the analysis of input datasets, it was found that the amount of feed intake was the most influential factor in generating CO2. Numerous studies have been conducted to determine the relationship between CO2 emission rate and feed intake and body weight of pigs (Gerber et al. 2013; Philippe and Nicks 2014; Van Mierlo et al. 2021). According to Philippe and Nicks (2015), the feed intake, body weight, production level, and physiological stage of the pigs have an effect on the amount of CO2 exhaled (E-CO2, pig). Aubry et al. (2004) proposed a mathematical equation to predict CO2 exhalation (E-CO2, pig, in kg CO2 day−1) in pigs weighing 20–120 kg body weight, and they estimated that a pig weighing 70 kg produces approximately 1.55 kg CO2 per day via the respiratory process.
Model performance
In the present study, seven regression-based statistical and machine learning models, including linear, multiple linear, polynomial, exponential, ridge, lasso, and elastic net models, were used. Therefore, it is important to figure out which one performs the best. The best model selection for the datasets D1, D2, and D3 is depicted in Fig. 7. The results indicate that overfitting was perfectly controlled in this study. In CO2 prediction, most of the models obtained almost the same performance (R2 and RMSE) during the training stage. The study found that the R2 and RMSE values ranged from 0.692 to 0.785 and 17.44 ppm pig−1 to 20.85 ppm pig−1 for the D1 dataset, 0.673 to 0.788 and 17.10 ppm pig−1 to 21.48 ppm pig−1 for the D2 dataset, and 0.752 to 0.817 and 16.08 ppm pig−1 to 18.72 ppm pig−1 for the D3 dataset, respectively, during the training period. Among the models, ER showed the best performance in predicting CO2 emission particularly for D1 (R2 = 0.660 and RMSE = 19.47 ppm pig−1) and D2 (R2 = 0.757 and RMSE = 16.45 ppm pig−1) in testing period. Due to its simplicity, this model has been used widely to predict greenhouse gas emission (Petersen et al. 2016; Ngwabie et al. 2018; Hempel et al. 2020). The exponential equation was used for CH4 emission modeling as a function of pigs’ mass, where the models explained about 88% of the variations in the measured and predicted data (Ngwabie et al. 2018).
The worst performance of ENR (R2 = 0.692 and RMSE = 20.85 ppm pig−1 for D1; R2 = 0.673 and RMSE = 21.48 ppm pig−1 for D2 and R2 = 0.752 and RMSE = 18.72 ppm pig−1 for D3) was observed in the training period; nevertheless, it performed better compared to RR and LaR models during the testing period. According to a previous study, the ENR model is particularly useful because it solves the limitations of both RR and LaR methods (Zou and Hastie 2005). Additionally, though the PR model with D3 dataset had a high training accuracy (R2 = 0.817 and RMSE = 16.08 ppm pig−1), it performed less in the testing stage (R2 = 0.606 and RMSE = 20.95 ppm pig−1). This finding indicated that there was a strong correlation between the predicted and measured data in the training stage; however, it was not consistent during the testing phase of the PR model, which might have happened due to the data splitting process (Kim and Oh 2021). The study also found that in terms of the percentage differences between ER’s training and testing, the R2 value in the training stage was 3.93% higher, and the RMSE value was 3.80% lower compared to the testing stage. In comparison to the testing stage, the training stage’s R2 value was 25.73% higher, and the RMSE value was 30.27% lower for the PR model (Table 4).
Model comparison and proposed model
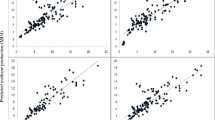
In this study, ER model performed better in the testing period compared to all the statistical and machine learning–based regression model (Table 4). In addition, the PR model had the highest performance during the training period for D3 dataset among all models; however, the testing performance was lower compared to the ER model. Since the training was supervised, the testing results were considered for model evaluation substantially (Krishan et al. 2019; Arulmozhi et al. 2021). Other than those two models, RR, LaR, MLR, linear regression (LR), and ENR were performed in ascending order (Fig. 7). When compared with ER results, RMSE values were 21.64%, 19.03%, 21.64%, 21.64%, and 18.30% higher and R2 values were 15.46%, 13.47%, 15.45%, 15.45%, and 20.61% lower for the models LR, PR, RR, LaR, and ENR, respectively, in the testing stage for the D2 dataset. The overall comparison between actual and predicted CO2 data is illustrated along with the coefficient of determination (Figs. 4, 5, and 6).
Additionally, a graphical representation of the actual and predicted CO2 values on a graph (Fig. 8) can aid in comprehending the capability of ER models. The cumulative distribution function calculated from measured and predicted CO2 data using the ER model indicated that the predicted CO2 values using the D2 dataset were more accurate compared to the D1 dataset. As illustrated in Fig. 8, 73.91% of the data in D2 dataset had a residual value between − 20 and 20, whereas it was 65.22% in D1 dataset for the same range. Additionally, the S-curve revealed that a substantial portion of the relationships between independent variables and CO2 emission is nonlinear, which may affect the performance of this model. As a result, the exponential model was chosen as the optimal model for predicting CO2 emission rate when feed intake data was used as an input dataset (D2). The exponential method has a superior ability to describe the CO2 scenarios as the emission begins slowly and then accelerates rapidly, resembling the nature of exponential function. Similar findings have been reported in a variety of modeling studies (Basak et al. 2019; Borrani et al. 2021; Rahimpour et al. 2021).
Conclusion
The experiment was designed to assess the model relationships between CO2 emission rate, pig mass, and feed intake. Seven regression-based statistical and machine learning models were developed in this study to predict CO2 emission rate using three selected input variables, i.e., feed intake, pig mass, and the combination of feed intake and pig mass. The study discovered that among the seven models, an exponential model with a coefficient of determination (R2) greater than 78% in the training stage and 75% in the testing stage was the most suitable for explaining the relationships between feed intake and CO2 production rate. Apart from the exponential model, the polynomial, ridge, lasso, multiple linear, linear, and elastic net models placed first, second, third, fourth, fifth, and sixth, respectively. The exponential method has a superior ability to describe the CO2 scenarios as the emission begins slowly and then accelerates rapidly, resembling the nature of exponential function. Sensitivity analysis revealed that quantity of feed intake was the most influential factor in predicting CO2, followed by the combination of body mass and feed intake and the pig’s body mass. Due to the ease of computing, simplicity, and interpretability of the parameters as well as the better performance of exponential models, the study may be useful for CO2 emission modeling. However, feed intake and body mass may not always be the same when associated with CO2 emissions from pig barn. Additionally, attempting to achieve high prediction efficiency of CO2 emission modeling by applying the same parameters may lead to changes in the performance of the models. Therefore, additional research may be conducted to optimize this model’s performance efficiency by providing a diverse range of diets and management conditions.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Agüera A, Ahn I-Y, Guillaumot C, Danis B (2017) A dynamic energy budget (DEB) model to describe Laternula elliptica (King, 1832) seasonal feeding and metabolism. PLoS ONE 12(8):1–20. https://doi.org/10.1371/journal.pone.0183848
Aldabal M (2020) A comparative study of ridge, LASSO and elastic net estimators. Dissertation, Carleton University, Ottawa, Ontario
Ambade B, Sethi SS, Kurwadkar S, Kumar A, Sankar TK (2021) Toxicity and health risk assessment of polycyclic aromatic hydrocarbons in surface water, sediments and groundwater vulnerability in Damodar River Basin. Groundw Sustain Dev 13:1–12. https://doi.org/10.1016/j.gsd.2021.100553
Ambade B, Sankar TK, Panicker AS, Gautam AS, Gautam S (2021) Characterization, seasonal variation, source apportionment and health risk assessment of black carbon over an urban region of East India. Urban Clim 38:1–12. https://doi.org/10.1016/j.uclim.2021.100896
Ambade B, Sankar TK, Gautam Kumar A, S, (2021) COVID-19 lockdowns reduce the Black carbon and polycyclic aromatic hydrocarbons of the Asian atmosphere: source apportionment and health hazard evaluation. Environ Dev Sustain 23:12252–12271. https://doi.org/10.1007/s10668-020-01167-1
Ambade B, Sethi SS, Chintalacheruvu MR (2022) Distribution, risk assessment, and source apportionment of polycyclic aromatic hydrocarbons (PAHs) using positive matrix factorization (PMF) in urban soils of East India. Environ Geochem Health. https://doi.org/10.1007/s10653-022-01223-x
Ambade B, Kumar A, Kumar A, Sahu LK (2022) Temporal variability of atmospheric particulate-bound polycyclic aromatic hydrocarbons (PAHs) over central east India: sources and carcinogenic risk assessment. Air Qual Atmos Health 15:115–130. https://doi.org/10.1007/s11869-021-01089-5
Ambade B, Sethi SS, Kumar A, Sankar TK (2021a) Solvent extraction coupled with gas chromatography for the analysis of polycyclic aromatic hydrocarbons in riverine sediment and surface water of Subarnarekha River and its tributary, India. In: Kailasa SK, Hussain CM (ed) Miniaturized Analytical Devices: Materials and Technology. Wiley Online Library. https://doi.org/10.1002/9783527827213.ch4
Arulmozhi E, Basak JK, Sihalath T, Park J, Kim HT, Moon BE (2021) Machine learning-based microclimate model for indoor air temperature and relative humidity prediction in a swine building. Animals 11(1):1–24. https://doi.org/10.3390/ani11010222
Aubry A, Quiniou N, Cozler YL, Querne M (2004) New standardized criteria for GTE performances. J Rech Porcine 36:409-422. https://www.researchgate.net/publication/287174879
Austin M (2007) Species distribution models and ecological theory: a critical assessment and some possible new approaches. Ecol Modell 200(1–2):1–19. https://doi.org/10.1016/j.ecolmodel.2006.07.005
Basak JK, Qasim W, Okyere FG, Khan F, Lee YJ, Park J, Kim HT (2019) Regression analysis to estimate morphology parameters of pepper plant in a controlled greenhouse system. J Biosyst Eng 44:57–68. https://doi.org/10.1007/s42853-019-00014-0
Basak JK, Okyere FG, Arulmozhi E, Park J, Khan F, Kim HT (2020) Artificial neural networks and multiple linear regression as potential methods for modelling body surface temperature of pig. J Appl Anim Res 48(1):207–219. https://doi.org/10.1080/09712119.2020.1761818
Basak JK, Arulmozhi E, Moon BE, Bhujel A, Kim HT (2022) Modelling methane emissions from pig manure using statistical and machine learning methods. Air Qual Atmos Health. https://doi.org/10.1007/s11869-022-01169-0
Beninger PG, Boldina I (2014) Fine-scale spatial distribution of the temperate in faunal bivalve Tapes (=Ruditapes) philippinarum (Adams and Reeve) on fished and unfished intertidal mudflats. J Exp Mar Biol Ecol 457:128–134. https://doi.org/10.1016/j.jembe.2014.04.001
Besteiro R, Ortega JA, Arango T, Rodriguez MR, Fernandez MD, Ortega JA (2017) ARIMA modeling of animal zone temperature in weaned piglet buildings: design of the model. Trans ASABE 60:2175–2183. https://doi.org/10.13031/trans.12372
Borrani F, Solsona R, Candau R, Méline T, Sanchez AM (2021) Modelling performance with exponential functions in elite short-track speed skaters. J Sports Sci 39(20):2378–2385. https://doi.org/10.1080/02640414.2021.1933351
Cansino JM, Sánchez-Braza A, Rodríguez-Arévalo ML (2015) Driving forces of Spain’s CO2 emissions: A LMDI decomposition approach. Renew Sust Energ Rev 48:749–759. https://doi.org/10.1016/j.rser.2015.04.011
Darlington RB, Hayes, AF (2016) Regression analysis and linear models: concepts, applications, and implementation. Guilford Publications
Dettea H, Neugebauer HM (1997) Bayesian D-optimal designs for exponential regression models. J Stat Plan Inference 60(2):331–349. https://doi.org/10.1016/S0378-3758(96)00131-0
Duarte P, Fernández-Reiriz MJ, Labarta U (2012) Modelling mussel growth in ecosystems with low suspended matter loads using a Dynamic Energy Budget approach. J Sea Res 67(1):44–57. https://doi.org/10.1016/j.seares.2011.09.002
FAO (2013) GLEAM 2.0 Assessment of greenhouse gas emissions and mitigation potential. Food and Agriculture Organization (FAO), Rome, Italy. https://www.fao.org/gleam/results/en/. Accessed 21 October 2021
FAO (2015) World Livestock 2011-Livestock in Food Security. Food and Agricultural Organization (FAO), Rome, Italy. https://www.fao.org/policy-support/tools-and-publications/resources-details/en/c/1262785/. Accessed 30 October 2021
Gavrilova Y (2021) Introduction to polynomial regression analysis. Serokell Developers. https://serokell.io/blog/polynomial-regression-analysis. Accessed 21 January 2022
Gendy TS, El-Shiekh TM, Zakhary AS (2015) A polynomial regression model for stabilized turbulent confined jet diffusion flames using bluff body burners. Egypt J Pet 24(4):445–453. https://doi.org/10.1016/j.ejpe.2015.06.001
Gerber PJ, Steinfeld H, Henderson B, Mottet A, Opio C, Dijkman J, Falcucci A, Tempio G (2013) Tackling climate change through livestock -a global assessment of emissions and mitigation opportunities. Food and Agriculture Organization of the United Nations (FAO), Rome, Italy. https://www.fao.org/3/i3437e/i3437e.pdf. Accessed 10 July 2021
Glazier DS (2013) Log-transformation is useful for examining proportional relationships in allometric scaling. J Theor Biol 334:200–203. https://doi.org/10.1016/j.jtbi.2013.06.017
González PF, Landajo M, Presno MJ (2014) Tracking European Union CO2 emissions through LMDI (logarithmic-mean Divisia index) decomposition The activity revaluation approach. Energy 73:741–750. https://doi.org/10.1016/j.energy.2014.06.078
Hempel S, Adolphs J, Landwehr N, Willink D, Janke D, Amon T (2020) Supervised machine learning to assess methane emissions of a dairy building with natural ventilation. Appl Sci 10(19):1–21. https://doi.org/10.3390/app10196938
Hunter JD (2007) Matplotlib: A 2D graphics environment. Comput Sci Eng 9(3):90–95. https://doi.org/10.1109/MCSE.2007.55
Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S (2018) High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun 9:1–8. https://doi.org/10.1038/s41467-018-07641-9
Joharestani MZ, Cao C, Ni X, Bashir B, Talebiesfandarani S (2019) PM2.5 prediction based on random forest, XGBoost, and deep learning using multisource remote sensing data. Atmosphere 10(7):373. https://doi.org/10.3390/atmos10070373
Kang GK, Gao JZ, Chiao S, Lu S (2018) Air quality prediction: big data and machine learning approaches. Int J Environ Sci Develop 9(1):8-16. https://doi.org/10.18178/ijesd.2018.9.1.1066
Keat SC, Chun BB, San LH, Jafri MZM (2015) Multiple regression analysis in modelling of carbon dioxide emissions by energy consumption use in Malaysia. AIP Conference Proceedings 1657(1). https://doi.org/10.1063/1.4915185
Kim Y, Oh H (2021) Comparison between multiple regression analysis, polynomial regression analysis, and an artificial neural network for tensile strength prediction of BFRP and GFRP. Materials 14(17):1–13. https://doi.org/10.3390/ma14174861
Kisi O, Parmar KS, Soni K, Demir V (2017) Modeling of air pollutants using least square support vector regression, multivariate adaptive regression spline, and M5 model tree models. Air Qual Atmos Health 10:873–883. https://doi.org/10.1007/s11869-017-0477-9
Krishan M, Jha S, Das J, Singh A, Goyal MK, Sekar C (2019) Air quality modelling using long short-term memory (LSTM) over NCT-Delhi, India. Air Qual Atmos Health 12:899–908. https://doi.org/10.1007/s11869-019-00696-7
Kurwadkar S, Dane J, Kanel SR, Nadagouda MN, Cawdrey RW, Ambade B, Struckhoff GC, Wilkin R (2022) Per- and polyfluoroalkyl substances in water and wastewater: a critical review of their global occurrence and distribution. Sci Total Environ 809:1–19. https://doi.org/10.1016/j.scitotenv.2021.151003
Lashkenari MS, KhazaiePoul A (2015) Application of KNN and semi-empirical models for prediction of polycyclic aromatic hydrocarbons solubility in supercritical carbon dioxide. Polycycl Aromat Compd 37(5):415–425. https://doi.org/10.1080/10406638.2015.1129976
Legendre P, Legendre L (2012) Numerical ecology (3rd ed.). Amsterdam, Boston: Elsevier. https://www.elsevier.com/books/numerical-ecology/legendre/978-0-444-53868-0. Accessed 2 March 2021
Maharjan L, Tripathee L, Kang S, Ambade B, Chen P, Zheng H, Li Q, Shrestha KL, Sharma CM (2021) Characteristics of atmospheric particle-bound polycyclic aromatic compounds over the Himalayan middle hills: implications for sources and health risk assessment. Asian J Atmos Environ 15(4):1–19. https://doi.org/10.5572/ajae.2021.101
McKinney W (2010) Data structures for statistical computing in Python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June-3 July 2010. https://doi.org/10.25080/Majora-92bf1922-012
Mol CD, Vito ED, Rosasco L (2009) Elastic-net regularization in learning theory. J Complex 25(2):201–230. https://doi.org/10.1016/j.jco.2009.01.002
Moon BE, Kim HT, Kim JG, Ryou YS, Kim HT (2016) A fundamental study for development of unglazed transpired collector control system in window less pig house. J Agric Life Sci 50(2):175-185. https://doi.org/10.14397/jals.2016.50.2.175
Muthusamy B, Ramalingam S, Chandran SK, Kannaiyan SK (2021) Multivariate polynomial fit: decay heat removal system and pectin degrading Fe3O4-SiO2 nanobiocatalyst activity. IET Nanobiotechnol 15(2):173–196. https://doi.org/10.1049/nbt2.12034
Nagesha KV, Kumar H, Muralidhar Singh M (2019) Development of statistical models to predict emission rate and concentration of particulate matters (PM) for drilling operation in opencast mines. Air Qual Atmos Health 12:1073–1079. https://doi.org/10.1007/s11869-019-00723-7
Ngwabie NM, Chungong BN, Yengong FL (2018) Characterisation of pig manure for methane emission modelling in sub-Saharan Africa. Biosyst Eng 170:31–38. https://doi.org/10.1016/j.biosystemseng.2018.03.009
Oonincx DGAB, van Itterbeeck J, Heetkamp MJW, van den Brand H, van Loon JJA, van Huis A (2010) An exploration on greenhouse gas and ammonia production by insect species suitable for animal or human consumption. PLoS ONE 5(12):1–7. https://doi.org/10.1371/journal.pone.0014445
Ostertagová E (2012) Modelling using polynomial regression. Procedia Eng 48:500–506. https://doi.org/10.1016/j.proeng.2012.09.545
Packard GC (2013) Fitting statistical models in bivariate allometry: scaling metabolic rate to body mass in mustelid carnivores. Comp Biochem Physiol Mol Amp Integr Physiol 166(1):70–73. https://doi.org/10.1016/j.cbpa.2013.05.013
Peng H, Lima AR, Teakles A, Jin J, Cannon AJ, Hsieh WW (2017) Evaluating hourly air quality forecasting in Canada with nonlinear updatable machine learning methods. Air Qual Atmos Health 10:195–211. https://doi.org/10.1007/s11869-016-0414-3
Permai SD, Tanty H (2018) Linear regression model using Bayesian approach for energy performance of residential building. 3rd International Conference on Computer Science and Computational Intelligence 2018. Procedia Computer Science 135:671-677. https://doi.org/10.1016/j.procs.2018.08.219
Petersen SO, Olsen AB, Elsgaard L, Triolo JM, Sommer SG (2016) Estimation of methane emissions from slurry pits below pig and cattle confinements. PLoS ONE. 11(8):1–16. https://doi.org/10.1371/journal.pone.0160968
Petrovic Z, Djordjevic V, Milicevic D, Nastasijevic I, Parunovid N (2015) Meat production and consumption: environmental consequences. Procedia Food Sci 5:235–238. https://doi.org/10.1016/j.profoo.2015.09.041
Philippe F-X, Nicks B (2015) Review on greenhouse gas emissions from pig houses: production of carbon dioxide, methane and nitrous oxide by animals and manure. Agric Ecosyst Environ 199:10–25. https://doi.org/10.1016/j.agee.2014.08.015
Pork Chekoff (2018) World per capita pork consumption. https://porkcheckoff.org/. Accessed 12 May 2021
Prasad RJ, Sourie SJ, Cherukuri VR, Fita L, Merera CE (2015) Global warming: genesis, facts and impacts on livestock farming and mitigation strategies. Int J Agric Innov Res 3(5):1494–1503
Rahimpour A, Amanollahi J, Tzanis CG (2021) Air quality data series estimation based on machine learning approaches for urban environments. Air Qual Atmos Health 14:191–201. https://doi.org/10.1007/s11869-020-00925-4
Rajaratnam B, Roberts S, Sparks D, Yu H (2019) Influence diagnostics for high-dimensional Lasso regression. J Comput Graph Stat 28(4):877–890. https://doi.org/10.1080/10618600.2019.1598869
Ransom CJ, Kitchen NR, Camberato JJ (2019) Statistical and machine learning methods evaluated for incorporating soil and weather into corn nitrogen recommendations. Comput Electron Agr 164:1–15. https://doi.org/10.1016/j.compag.2019.104872
Rojas-Downing MM, Nejadhashemi AP, Harrigan T, Woznicki SA (2017) Climate change and livestock: Impacts, adaptation, and mitigation. Clim Risk Manag 16:145–163. https://doi.org/10.1016/j.crm.2017.02.001
Rybarczyk Y, Zalakeviciute R (2018) Machine learning approaches for outdoor air quality modelling: a systematic review. Appl Sci 8(12):1–28. https://doi.org/10.3390/app8122570
Shahriar SA, Kayes I, Hasan K, Salam MA, Chowdhury S (2020) Applicability of machine learning in modeling of atmospheric particle pollution in Bangladesh. Air Qual Atmos Health 13:1247–1256. https://doi.org/10.1007/s11869-020-00878-8
Silva KAP, Almeida LMW (2018) The exponential function meaning in mathematical modeling activities: a semiotic approach. J Res Math Educ 7(2):195–215. https://doi.org/10.4471/redimat.2018.2762
Tran TTK, Lee T, Shin J, Kim J, Kamruzzaman M (2020) Deep learning-based maximum temperature forecasting assisted with meta-learning for hyperparameter optimization. Atmosphere 11(5):1–21. https://doi.org/10.3390/atmos11050487
Tuomisto HL, Scheelbeek PFD, Chalabi Z, Green R, Smith RD, Haines A, Dangour AD (2017) Effects of environmental change on population nutrition and health: a comprehensive framework with a focus on fruits and vegetables. Wellcome Open Res 2(21):1-33. https://doi.org/10.12688/wellcomeopenres.11190.2
Van Der Walt VDS, Colbert SC, Varoquaux G (2011) The NumPy array: a structure for efficient numerical computation. Comput Sci Eng 13(2):22–30. https://doi.org/10.1109/MCSE.2011.37
Van Mierlo K, Baert L, Bracquené E, De Tavernier J, Geeraerd A (2021) The influence of farm characteristics and feed compositions on the environmental impact of pig production in flanders: productivity, energy use and protein choices are key. Sustainability 13(21):1–28. https://doi.org/10.3390/su132111623
Wieringen WN (2020) Lecture notes on ridge regression. Version 0.31, July 17, 2020. Department of Epidemiology and Data Science, Amsterdam Public Health research institute, Amsterdam UMC, location VUmc. https://arxiv.org/pdf/1509.09169.pdf. Accessed 5 December 2020
Xiao-wen D, Sun Z, Müller D (2021) Driving factors of direct greenhouse gas emissions from China’s pig industry from 1976 to 2016. J Integr Agric 20(1):319–329. https://doi.org/10.1016/S2095-3119(20)63425-6
Zhou Y, Zhang J, Hu S (2021) Regression analysis and driving force model building of CO2 emissions in China. Sci Rep 11:6715. https://doi.org/10.1038/s41598-021-86183-5
Zou H, Hastie T (2005) Regularization and variable selection via the elastic net. J R Statist Soc B 67(2):301–320 (https://www.jstor.org/stable/3647580)
Funding
This research has been financially supported by the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) through Agriculture, Food and Rural Affairs Convergence Technologies Program for Educating Creative Global Leader, funded by the Ministry of Agriculture, Food and Rural Affairs (MAFRA) (717001-7) and in part by Brain Pool program through the National Research Foundation of Korea (2021H1D3A2A02038875).
Author information
Authors and Affiliations
Contributions
Jayanta Kumar Basak is responsible for all of the calculations, figures, writing, and large portions of the text. Na Eun Kim, Bhola Paudel and Byeong Eun Moon helped during the experimental setup and data collection period, and Shihab Ahmad Shahriar reviewed the article. Hyeon Tae Kim reviewed and edited the manuscript as well as guided the experiment.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The current research was conducted at the smart farm systems laboratory of Gyeongsang National University. The Institutional Animal Care and Use Committee (IACU) (GNU-150508-R0029) approved the experimental procedure and collection of data. All the authors followed the ethical guidelines and safety procedures carefully during the experimental period.
Consent for publication
All the authors have read and agreed to the published version of the manuscript in the journal of Air Quality, Atmosphere, and Health.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Basak, J.K., Kim, N.E., Shahriar, S.A. et al. Applicability of statistical and machine learning–based regression algorithms in modeling of carbon dioxide emission in experimental pig barns. Air Qual Atmos Health 15, 1899–1912 (2022). https://doi.org/10.1007/s11869-022-01225-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11869-022-01225-9